Detailed Characterization of Early HIV-1 Replication Dynamics in Primary Human Macrophages
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Culture
2.2. Plasmids
2.3. Antisera and Reagents
2.4. Virus Production and Characterization
2.5. Virus Infection and Transduction, Click-Labeling, and Immunostaining
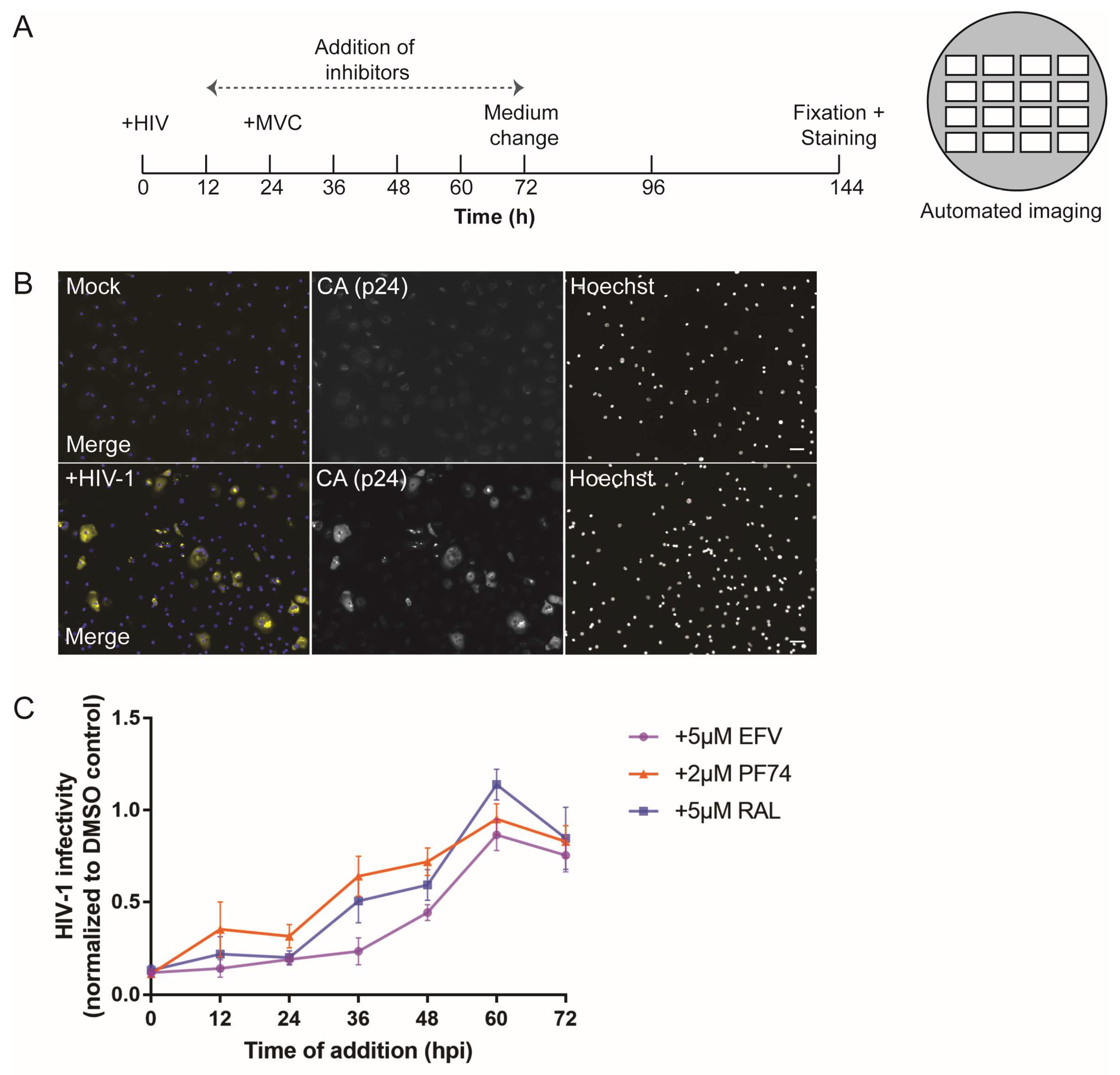
2.6. Infectivity Assays and Time-of-Addition Experiments
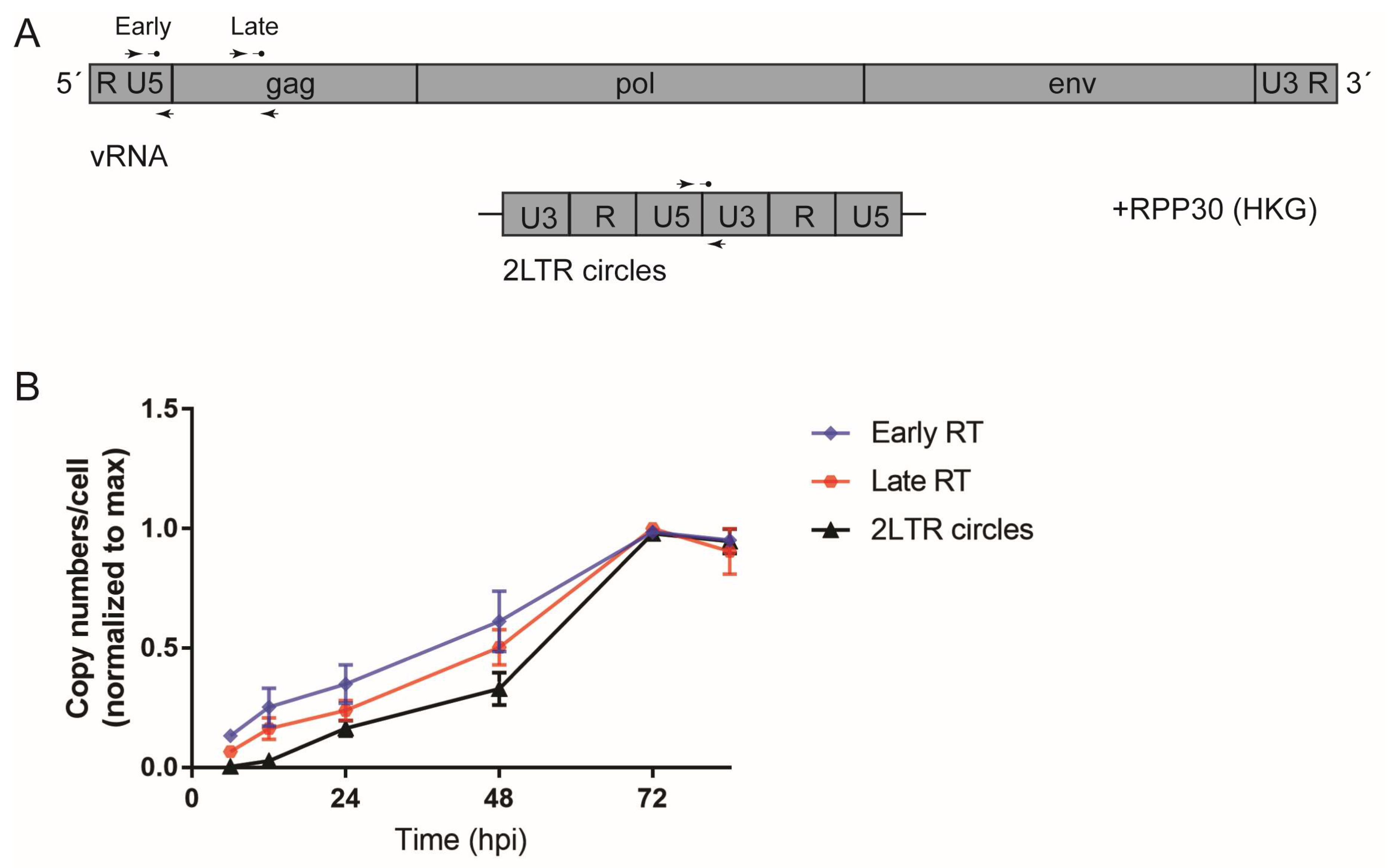
2.7. Detection of HIV-1 RT Products with ddPCR
2.8. Flow Cytometry
2.9. SDS-PAGE and Western Blot
2.10. Microscopy
2.11. Image Analysis
3. Results
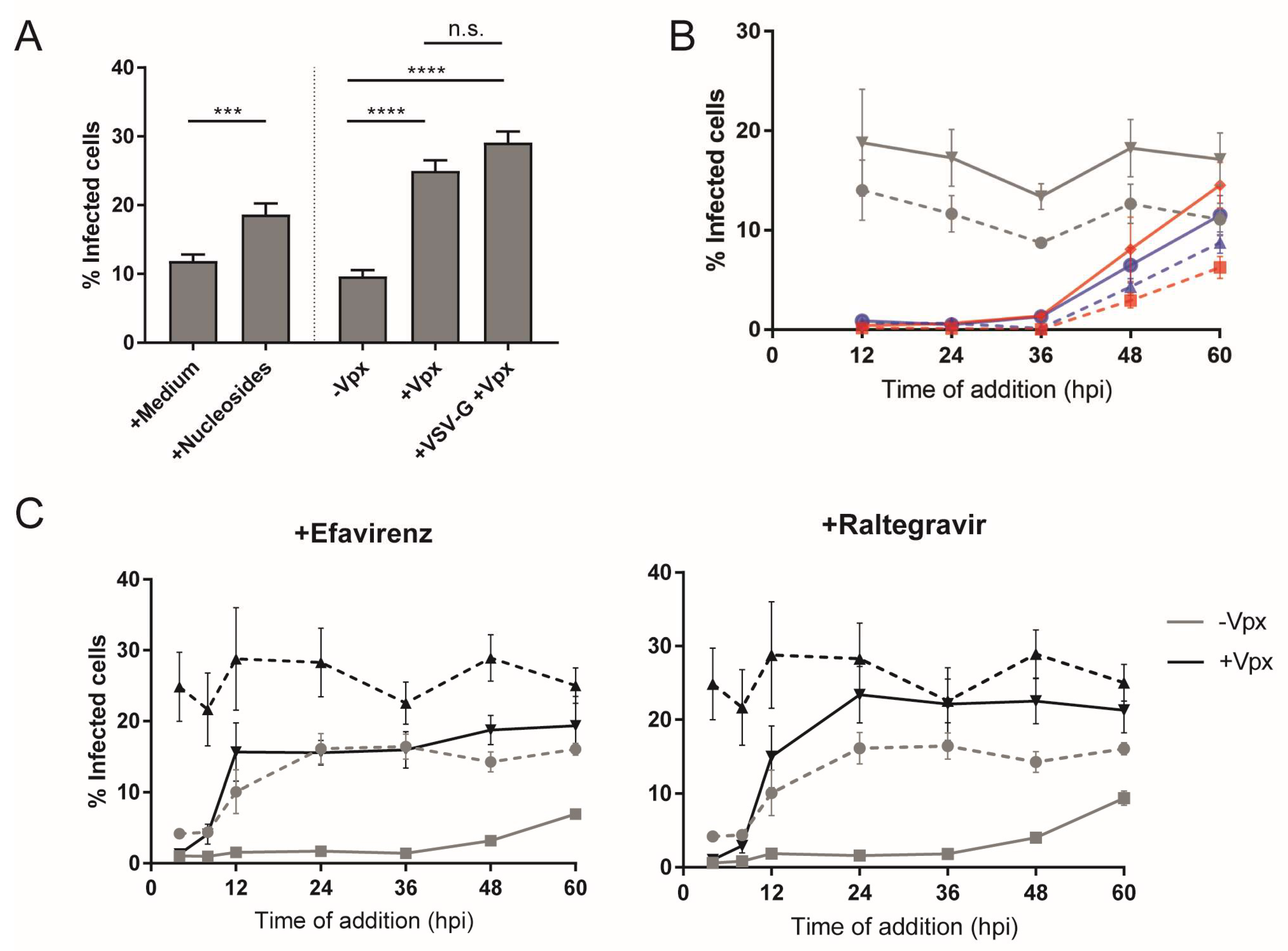
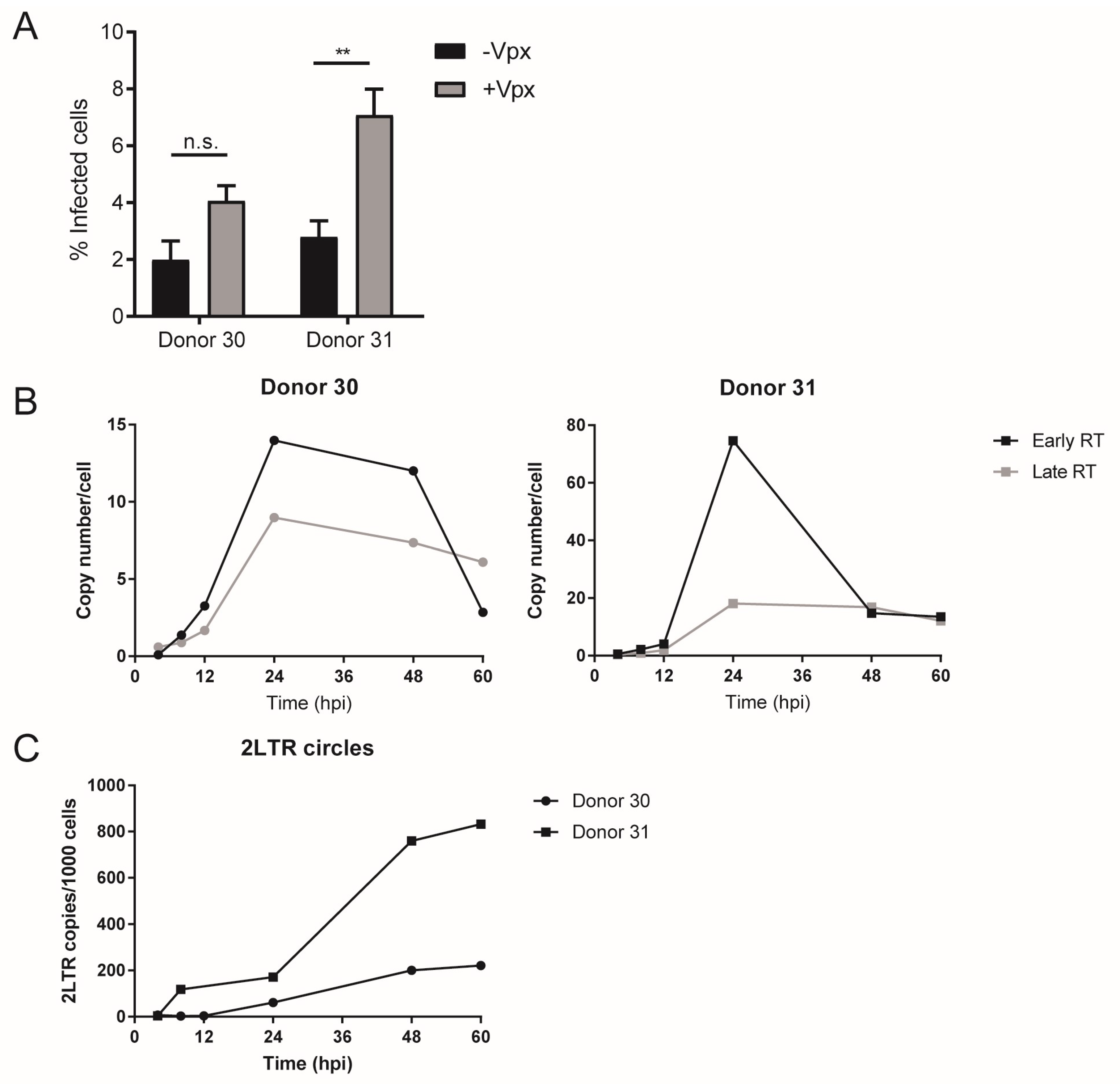
3.1. Early HIV-1 Replication Is Slow in Primary Macrophages
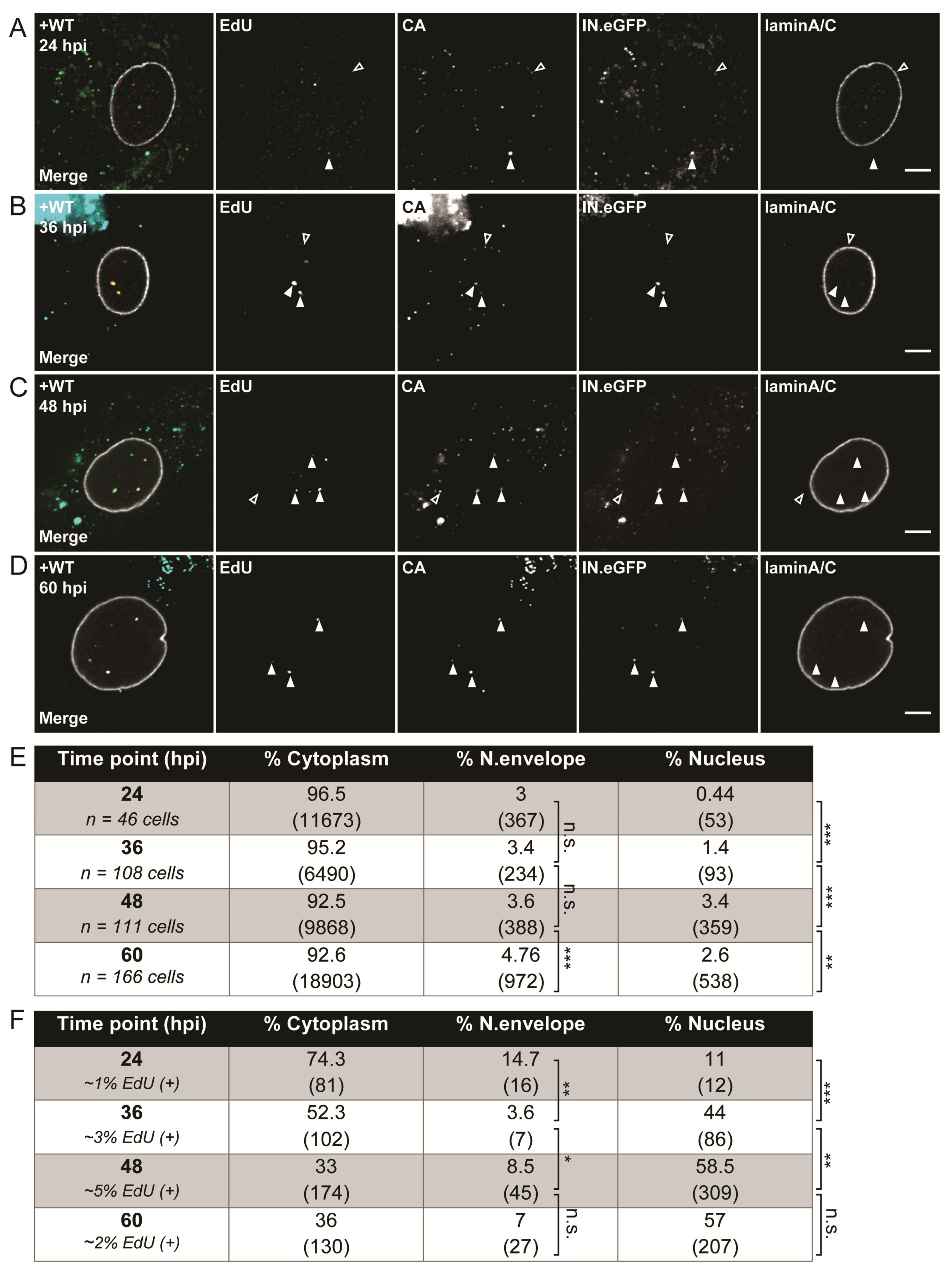
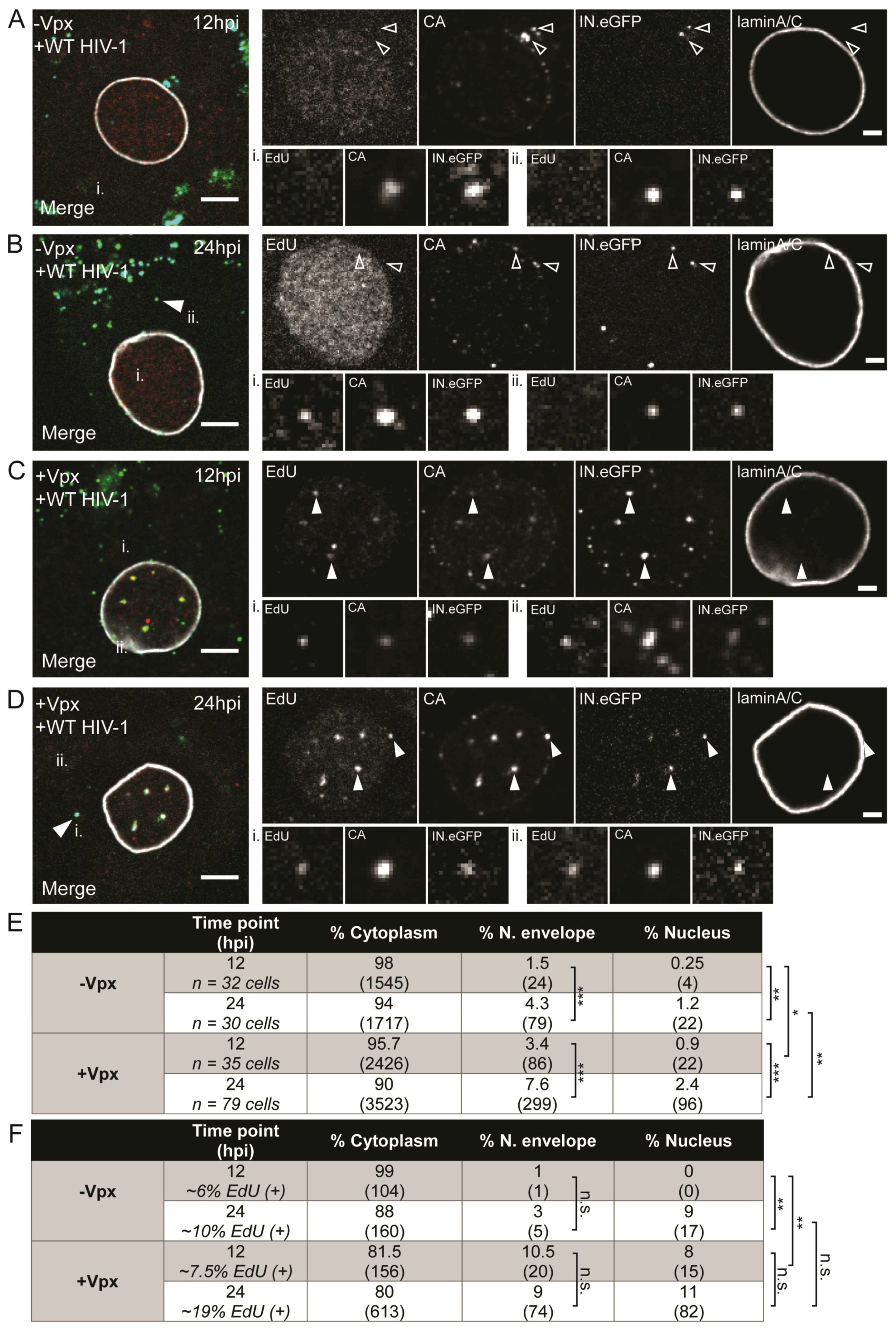
3.2. Visualization of HIV-1 RTC/PIC Revealed Timeframe of Nuclear Translocation
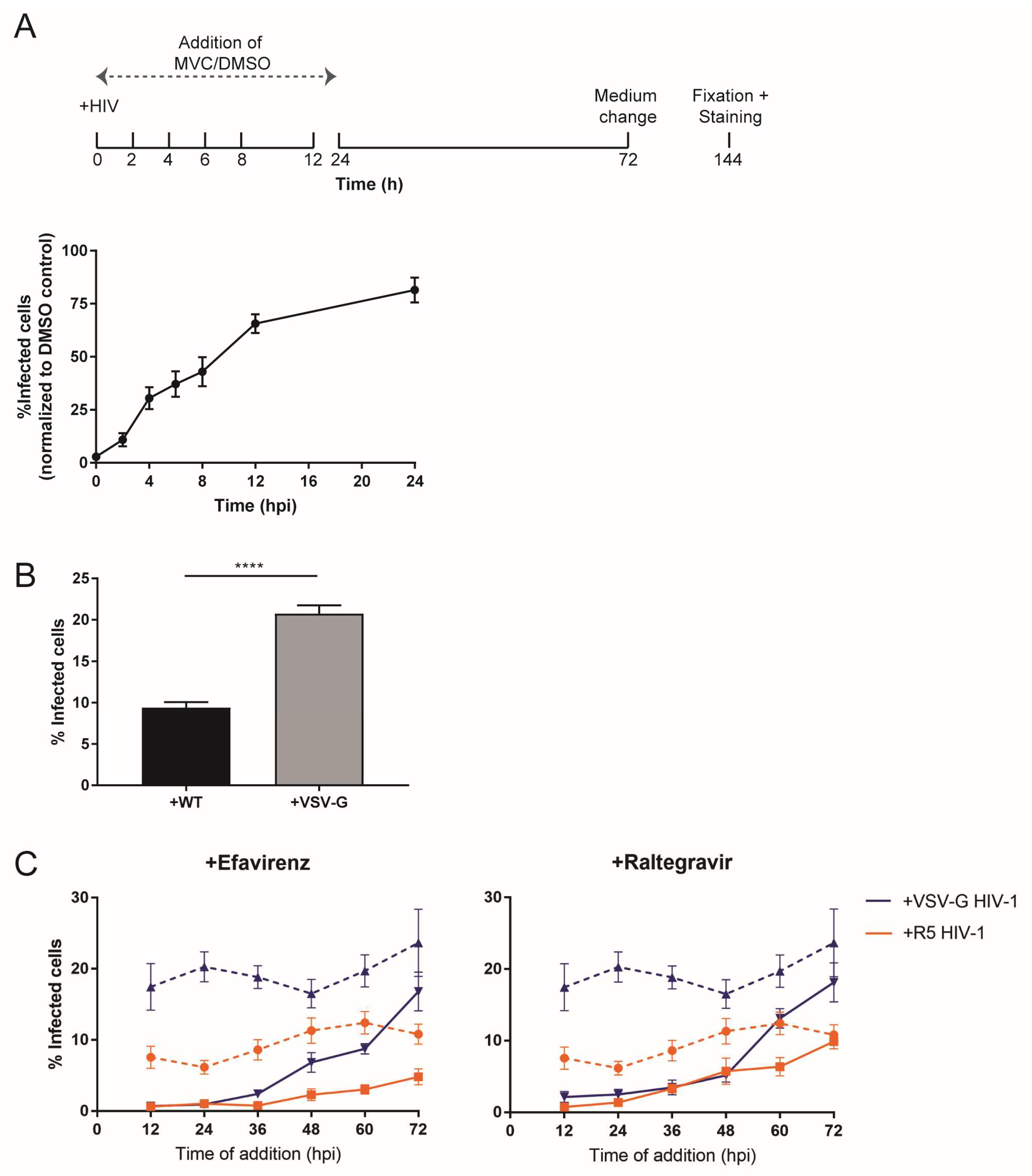
3.3. Type and Time of Cell Entry Does Not Affect Replication Kinetics in Macrophages
3.4. Low Availability of dNTPs Restricts HIV-1 Replication in MDMs
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Stevenson, M. HIV-1 pathogenesis. Nat. Med. 2003, 9, 853–860. [Google Scholar] [CrossRef] [PubMed]
- Gartner, S.; Markovits, P.; Markovitz, D.M.; Kaplan, M.H.; Gallo, R.C.; Popovic, M. The role of mononuclear phagocytes in HTLV-III/LAV infection. Science 1986, 233, 215–219. [Google Scholar] [CrossRef] [PubMed]
- Wiley, C.A.; Schrier, R.D.; Nelson, J.A.; Lampert, P.W.; Oldstone, M.B. Cellular localization of human immunodeficiency virus infection within the brains of acquired immune deficiency syndrome patients. Proc. Natl. Acad. Sci. USA 1986, 83, 7089–7093. [Google Scholar] [CrossRef] [PubMed]
- Weinberg, J.B.; Matthews, T.J.; Cullen, B.R.; Malim, M.H. Productive human immunodeficiency virus type 1 (HIV-1) infection of nonproliferating human monocytes. J. Exp. Med. 1991, 174, 1477–1482. [Google Scholar] [CrossRef] [PubMed]
- Igarashi, T.; Brown, C.R.; Endo, Y.; Buckler-White, A.; Plishka, R.; Bischofberger, N.; Hirsch, V.; Martin, M.A. Macrophage are the principal reservoir and sustain high virus loads in rhesus macaques after the depletion of CD4+ T cells by a highly pathogenic simian immunodeficiency virus/HIV type 1 chimera (SHIV): Implications for HIV-1 infections of humans. Proc. Natl. Acad. Sci. USA 2001, 98, 658–663. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, K.K.; Fernandez-Larsson, R.; Zinkus, D.M.; Robinson, H.L. Human immunodeficiency virus type 1 NL4-3 replication in four T-cell lines: Rate and efficiency of entry, a major determinant of permissiveness. J. Virol. 1991, 65, 3900–3902. [Google Scholar] [PubMed]
- Fernandez-Larsson, R.; Srivastava, K.K.; Lu, S.; Robinson, H.L. Replication of patient isolates of human immunodeficiency virus type 1 in T cells: A spectrum of rates and efficiencies of entry. Proc. Natl. Acad. Sci. USA 1992, 89, 2223–2226. [Google Scholar] [CrossRef] [PubMed]
- Murray, J.M.; Kelleher, A.D.; Cooper, D.A. Timing of the components of the HIV life cycle in productively infected CD4+ T cells in a population of HIV-infected individuals. J. Virol. 2011, 85, 10798–10805. [Google Scholar] [CrossRef] [PubMed]
- Holmes, M.; Zhang, F.; Bieniasz, P.D. Single-Cell and Single-Cycle Analysis of HIV-1 Replication. PLoS Pathog. 2015, 11, e1004961. [Google Scholar] [CrossRef] [PubMed]
- Tomaras, G.D.; Lacey, S.F.; McDanal, C.B.; Ferrari, G.; Weinhold, K.J.; Greenberg, M.L. CD8+ T cell-mediated suppressive activity inhibits HIV-1 after virus entry with kinetics indicating effects on virus gene expression. Proc. Natl. Acad. Sci. USA 2000, 97, 3503–3508. [Google Scholar] [CrossRef] [PubMed]
- Collman, R.; Hassan, N.F.; Walker, R.; Godfrey, B.; Cutilli, J.; Hastings, J.C.; Friedman, H.; Douglas, S.D.; Nathanson, N. Infection of monocyte-derived macrophages with human immunodeficiency virus type 1 (HIV-1). Monocyte-tropic and lymphocyte-tropic strains of HIV-1 show distinctive patterns of replication in a panel of cell types. J. Exp. Med. 1989, 170, 1149–1163. [Google Scholar] [CrossRef] [PubMed]
- Rich, E.A.; Chen, I.S.; Zack, J.A.; Leonard, M.L.; O’Brien, W.A. Increased susceptibility of differentiated mononuclear phagocytes to productive infection with human immunodeficiency virus-1 (HIV-1). J. Clin. Investig. 1992, 89, 176–183. [Google Scholar] [CrossRef] [PubMed]
- Fouchier, R.A.; Brouwer, M.; Kootstra, N.A.; Huisman, H.G.; Schuitemaker, H. HIV-1 macrophage tropism is determined at multiple levels of the viral replication cycle. J. Clin. Investig. 1994, 94, 1806–1814. [Google Scholar] [CrossRef] [PubMed]
- Koyanagi, Y.; O’Brien, W.A.; Zhao, J.Q.; Golde, D.W.; Gasson, J.C.; Chen, I.S. Cytokines alter production of HIV-1 from primary mononuclear phagocytes. Science 1988, 241, 1673–1675. [Google Scholar] [CrossRef] [PubMed]
- Cassol, E.; Cassetta, L.; Rizzi, C.; Alfano, M.; Poli, G. M1 and M2a polarization of human monocyte-derived macrophages inhibits HIV-1 replication by distinct mechanisms. J. Immunol. 2009, 182, 6237–6246. [Google Scholar] [CrossRef] [PubMed]
- Munis, J.R.; Kornbluth, R.S.; Guatelli, J.C.; Richman, D.D. Ordered appearance of human immunodeficiency virus type 1 nucleic acids following high multiplicity infection of macrophages. J. Gen. Virol. 1992, 73 Pt 8, 1899–1906. [Google Scholar] [CrossRef]
- Collin, M.; Gordon, S. The kinetics of human immunodeficiency virus reverse transcription are slower in primary human macrophages than in a lymphoid cell line. Virology 1994, 200, 114–120. [Google Scholar] [CrossRef] [PubMed]
- Stultz, R.D.; Cenker, J.J.; McDonald, D. Imaging HIV-1 Genomic DNA from Entry through Productive Infection. J. Virol. 2017, 91, e00034-17. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, W.A.; Namazi, A.; Kalhor, H.; Mao, S.H.; Zack, J.A.; Chen, I.S. Kinetics of human immunodeficiency virus type 1 reverse transcription in blood mononuclear phagocytes are slowed by limitations of nucleotide precursors. J. Virol. 1994, 68, 1258–1263. [Google Scholar] [PubMed]
- Meyerhans, A.; Vartanian, J.P.; Hultgren, C.; Plikat, U.; Karlsson, A.; Wang, L.; Eriksson, S.; Wain-Hobson, S. Restriction and enhancement of human immunodeficiency virus type 1 replication by modulation of intracellular deoxynucleoside triphosphate pools. J. Virol. 1994, 68, 535–540. [Google Scholar] [PubMed]
- Diamond, T.L.; Roshal, M.; Jamburuthugoda, V.K.; Reynolds, H.M.; Merriam, A.R.; Lee, K.Y.; Balakrishnan, M.; Bambara, R.A.; Planelles, V.; Dewhurst, S.; et al. Macrophage tropism of HIV-1 depends on efficient cellular dNTP utilization by reverse transcriptase. J. Biol. Chem. 2004, 279, 51545–51553. [Google Scholar] [CrossRef] [PubMed]
- Goldstone, D.C.; Ennis-Adeniran, V.; Hedden, J.J.; Groom, H.C.; Rice, G.I.; Christodoulou, E.; Walker, P.A.; Kelly, G.; Haire, L.F.; Yap, M.W.; et al. HIV-1 restriction factor SAMHD1 is a deoxynucleoside triphosphate triphosphohydrolase. Nature 2011, 480, 379–382. [Google Scholar] [CrossRef] [PubMed]
- Laguette, N.; Sobhian, B.; Casartelli, N.; Ringeard, M.; Chable-Bessia, C.; Segeral, E.; Yatim, A.; Emiliani, S.; Schwartz, O.; Benkirane, M. SAMHD1 is the dendritic- and myeloid-cell-specific HIV-1 restriction factor counteracted by Vpx. Nature 2011, 474, 654–657. [Google Scholar] [CrossRef] [PubMed]
- Hollenbaugh, J.A.; Tao, S.; Lenzi, G.M.; Ryu, S.; Kim, D.H.; Diaz-Griffero, F.; Schinazi, R.F.; Kim, B. dNTP pool modulation dynamics by SAMHD1 protein in monocyte-derived macrophages. Retrovirology 2014, 11, 63. [Google Scholar] [CrossRef] [PubMed]
- Lahouassa, H.; Daddacha, W.; Hofmann, H.; Ayinde, D.; Logue, E.C.; Dragin, L.; Bloch, N.; Maudet, C.; Bertrand, M.; Gramberg, T.; et al. SAMHD1 restricts the replication of human immunodeficiency virus type 1 by depleting the intracellular pool of deoxynucleoside triphosphates. Nat. Immunol. 2012, 13, 223–228. [Google Scholar] [CrossRef] [PubMed]
- Franchini, G.; Gurgo, C.; Guo, H.G.; Gallo, R.C.; Collalti, E.; Fargnoli, K.A.; Hall, L.F.; Wong-Staal, F.; Reitz, M.S., Jr. Sequence of simian immunodeficiency virus and its relationship to the human immunodeficiency viruses. Nature 1987, 328, 539–543. [Google Scholar] [CrossRef] [PubMed]
- Guyader, M.; Emerman, M.; Sonigo, P.; Clavel, F.; Montagnier, L.; Alizon, M. Genome organization and transactivation of the human immunodeficiency virus type 2. Nature 1987, 326, 662–669. [Google Scholar] [CrossRef] [PubMed]
- Goujon, C.; Arfi, V.; Pertel, T.; Luban, J.; Lienard, J.; Rigal, D.; Darlix, J.L.; Cimarelli, A. Characterization of Simian Immunodeficiency Virus SIVSM/Human Immunodeficiency Virus Type 2 Vpx Function in Human Myeloid Cells. J. Virol. 2008, 82, 12335–12345. [Google Scholar] [CrossRef] [PubMed]
- Hrecka, K.; Hao, C.; Gierszewska, M.; Swanson, S.K.; Kesik-Brodacka, M.; Srivastava, S.; Florens, L.; Washburn, M.P.; Skowronski, J. Vpx relieves inhibition of HIV-1 infection of macrophages mediated by the SAMHD1 protein. Nature 2011, 474, 658–661. [Google Scholar] [CrossRef] [PubMed]
- Hofmann, H.; Logue, E.C.; Bloch, N.; Daddacha, W.; Polsky, S.B.; Schultz, M.L.; Kim, B.; Landau, N.R. The Vpx Lentiviral Accessory Protein Targets SAMHD1 for Degradation in the Nucleus. J. Virol. 2012, 86, 12552–12560. [Google Scholar] [CrossRef] [PubMed]
- Triques, K.; Stevenson, M. Characterization of restrictions to human immunodeficiency virus type 1 infection of monocytes. J. Virol. 2004, 78, 5523–5527. [Google Scholar] [CrossRef] [PubMed]
- Kim, B.; Nguyen, L.A.; Daddacha, W.; Hollenbaugh, J.A. Tight interplay among SAMHD1 protein level, cellular dNTP levels, and HIV-1 proviral DNA synthesis kinetics in human primary monocyte-derived macrophages. J. Biol. Chem. 2012, 287, 21570–21574. [Google Scholar] [CrossRef] [PubMed]
- Pear, W.S.; Nolan, G.P.; Scott, M.L.; Baltimore, D. Production of high-titer helper-free retroviruses by transient transfection. Proc. Natl. Acad. Sci. USA 1993, 90, 8392–8396. [Google Scholar] [CrossRef] [PubMed]
- Wei, X.; Decker, J.M.; Liu, H.; Zhang, Z.; Arani, R.B.; Kilby, J.M.; Saag, M.S.; Wu, X.; Shaw, G.M.; Kappes, J.C. Emergence of resistant human immunodeficiency virus type 1 in patients receiving fusion inhibitor (T-20) monotherapy. Antimicrob. Agents Chemother. 2002, 46, 1896–1905. [Google Scholar] [CrossRef] [PubMed]
- Adachi, A.; Gendelman, H.E.; Koenig, S.; Folks, T.; Willey, R.; Rabson, A.; Martin, M.A. Production of acquired immunodeficiency syndrome-associated retrovirus in human and nonhuman cells transfected with an infectious molecular clone. J. Virol. 1986, 59, 284–291. [Google Scholar] [PubMed]
- Schnell, G.; Joseph, S.; Spudich, S.; Price, R.W.; Swanstrom, R. HIV-1 replication in the central nervous system occurs in two distinct cell types. PLoS Pathog. 2011, 7, e1002286. [Google Scholar] [CrossRef] [PubMed]
- Albanese, A.; Arosio, D.; Terreni, M.; Cereseto, A. HIV-1 pre-integration complexes selectively target decondensed chromatin in the nuclear periphery. PLoS ONE 2008, 3, e2413. [Google Scholar] [CrossRef] [PubMed]
- Pertel, T.; Reinhard, C.; Luban, J. Vpx rescues HIV-1 transduction of dendritic cells from the antiviral state established by type 1 interferon. Retrovirology 2011, 8, 49. [Google Scholar] [CrossRef] [PubMed]
- Sunseri, N.; O’Brien, M.; Bhardwaj, N.; Landau, N.R. Human Immunodeficiency Virus Type 1 Modified To Package Simian Immunodeficiency Virus Vpx Efficiently Infects Macrophages and Dendritic Cells. J. Virol. 2011, 85, 6263–6274. [Google Scholar] [CrossRef] [PubMed]
- Pizzato, M.; Erlwein, O.; Bonsall, D.; Kaye, S.; Muir, D.; McClure, M.O. A one-step SYBR Green I-based product-enhanced reverse transcriptase assay for the quantitation of retroviruses in cell culture supernatants. J. Virol. Methods 2009, 156, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Moron-Lopez, S.; Puertas, M.C.; Galvez, C.; Navarro, J.; Carrasco, A.; Esteve, M.; Manye, J.; Crespo, M.; Salgado, M.; Martinez-Picado, J. Sensitive quantification of the HIV-1 reservoir in gut-associated lymphoid tissue. PLoS ONE 2017, 12, e0175899. [Google Scholar] [CrossRef] [PubMed]
- Hindson, B.J.; Ness, K.D.; Masquelier, D.A.; Belgrader, P.; Heredia, N.J.; Makarewicz, A.J.; Bright, I.J.; Lucero, M.Y.; Hiddessen, A.L.; Legler, T.C.; et al. High-throughput droplet digital PCR system for absolute quantitation of DNA copy number. Anal. Chem. 2011, 83, 8604–8610. [Google Scholar] [CrossRef] [PubMed]
- Liszewski, M.K.; Yu, J.J.; O’Doherty, U. Detecting HIV-1 integration by repetitive-sampling Alu-gag PCR. Methods 2009, 47, 254–260. [Google Scholar] [CrossRef] [PubMed]
- Palmer, S.; Wiegand, A.P.; Maldarelli, F.; Bazmi, H.; Mican, J.M.; Polis, M.; Dewar, R.L.; Planta, A.; Liu, S.; Metcalf, J.A.; et al. New real-time reverse transcriptase-initiated PCR assay with single-copy sensitivity for human immunodeficiency virus type 1 RNA in plasma. J. Clin. Microbiol. 2003, 41, 4531–4536. [Google Scholar] [CrossRef] [PubMed]
- Puertas, M.C.; Gomez-Mora, E.; Santos, J.R.; Molto, J.; Urrea, V.; Moron-Lopez, S.; Hernandez-Rodriguez, A.; Marfil, S.; Martinez-Bonet, M.; Matas, L.; et al. Impact of intensification with raltegravir on HIV-1-infected individuals receiving monotherapy with boosted PIs. J. Antimicrob. Chemother. 2018. [Google Scholar] [CrossRef] [PubMed]
- Borner, K.; Hermle, J.; Sommer, C.; Brown, N.P.; Knapp, B.; Glass, B.; Kunkel, J.; Torralba, G.; Reymann, J.; Beil, N.; et al. From experimental setup to bioinformatics: An RNAi screening platform to identify host factors involved in HIV-1 replication. Biotechnol. J. 2010, 5, 39–49. [Google Scholar] [CrossRef] [PubMed]
- Martinez, F.O.; Sica, A.; Mantovani, A.; Locati, M. Macrophage activation and polarization. Front. Biosci. 2008, 13, 453–461. [Google Scholar] [CrossRef] [PubMed]
- Barros, M.H.; Hauck, F.; Dreyer, J.H.; Kempkes, B.; Niedobitek, G. Macrophage polarisation: An immunohistochemical approach for identifying M1 and M2 macrophages. PLoS ONE 2013, 8, e80908. [Google Scholar] [CrossRef] [PubMed]
- Gobeil, L.A.; Lodge, R.; Tremblay, M.J. Differential HIV-1 endocytosis and susceptibility to virus infection in human macrophages correlate with cell activation status. J. Virol. 2012, 86, 10399–10407. [Google Scholar] [CrossRef] [PubMed]
- Peng, K.; Muranyi, W.; Glass, B.; Laketa, V.; Yant, S.R.; Tsai, L.; Cihlar, T.; Muller, B.; Krausslich, H.G. Quantitative microscopy of functional HIV post-entry complexes reveals association of replication with the viral capsid. eLife 2014, 3, e04114. [Google Scholar] [CrossRef] [PubMed]
- Saito, A.; Ferhadian, D.; Sowd, G.A.; Serrao, E.; Shi, J.; Halambage, U.D.; Teng, S.; Soto, J.; Siddiqui, M.A.; Engelman, A.N.; et al. Roles of Capsid-Interacting Host Factors in Multimodal Inhibition of HIV-1 by PF74. J. Virol. 2016, 90, 5808–5823. [Google Scholar] [CrossRef] [PubMed]
- Shi, J.; Zhou, J.; Shah, V.B.; Aiken, C.; Whitby, K. Small-molecule inhibition of human immunodeficiency virus type 1 infection by virus capsid destabilization. J. Virol. 2011, 85, 542–549. [Google Scholar] [CrossRef] [PubMed]
- Sharkey, M.E.; Teo, I.; Greenough, T.; Sharova, N.; Luzuriaga, K.; Sullivan, J.L.; Bucy, R.P.; Kostrikis, L.G.; Haase, A.; Veryard, C.; et al. Persistence of episomal HIV-1 infection intermediates in patients on highly active anti-retroviral therapy. Nat. Med. 2000, 6, 76–81. [Google Scholar] [CrossRef] [PubMed]
- Burdick, R.C.; Delviks-Frankenberry, K.A.; Chen, J.; Janaka, S.K.; Sastri, J.; Hu, W.S.; Pathak, V.K. Dynamics and regulation of nuclear import and nuclear movements of HIV-1 complexes. PLoS Pathog. 2017, 13, e1006570. [Google Scholar] [CrossRef] [PubMed]
- Francis, A.C.; Melikyan, G.B. Single HIV-1 Imaging Reveals Progression of Infection through CA-Dependent Steps of Docking at the Nuclear Pore, Uncoating, and Nuclear Transport. Cell Host Microbe 2018, 23, 536–548. [Google Scholar] [CrossRef] [PubMed]
- Schroers, R.; Sinha, I.; Segall, H.; Schmidt-Wolf, I.G.; Rooney, C.M.; Brenner, M.K.; Sutton, R.E.; Chen, S.Y. Transduction of human PBMC-derived dendritic cells and macrophages by an HIV-1-based lentiviral vector system. Mol. Ther. 2000, 1, 171–179. [Google Scholar] [CrossRef] [PubMed]
- Neil, S.; Martin, F.; Ikeda, Y.; Collins, M. Postentry restriction to human immunodeficiency virus-based vector transduction in human monocytes. J. Virol. 2001, 75, 5448–5456. [Google Scholar] [CrossRef] [PubMed]
- Matlin, K.S.; Reggio, H.; Helenius, A.; Simons, K. Pathway of vesicular stomatitis virus entry leading to infection. J. Mol. Biol. 1982, 156, 609–631. [Google Scholar] [CrossRef]
- Aiken, C. Pseudotyping human immunodeficiency virus type 1 (HIV-1) by the glycoprotein of vesicular stomatitis virus targets HIV-1 entry to an endocytic pathway and suppresses both the requirement for Nef and the sensitivity to cyclosporin A. J. Virol. 1997, 71, 5871–5877. [Google Scholar] [PubMed]
- Amie, S.M.; Noble, E.; Kim, B. Intracellular Nucleotide Levels and the Control of Retroviral Infections. Virology 2013, 436, 247–254. [Google Scholar] [CrossRef] [PubMed]
- Ahn, J.; Hao, C.; Yan, J.; DeLucia, M.; Mehrens, J.; Wang, C.; Gronenborn, A.M.; Skowronski, J. HIV/simian immunodeficiency virus (SIV) accessory virulence factor Vpx loads the host cell restriction factor SAMHD1 onto the E3 ubiquitin ligase complex CRL4DCAF1. J. Biol. Chem. 2012, 287, 12550–12558. [Google Scholar] [CrossRef] [PubMed]
- Chin, C.R.; Perreira, J.M.; Savidis, G.; Portmann, J.M.; Aker, A.M.; Feeley, E.M.; Smith, M.C.; Brass, A.L. Direct Visualization of HIV-1 Replication Intermediates Shows that Capsid and CPSF6 Modulate HIV-1 Intra-nuclear Invasion and Integration. Cell Rep. 2015, 13, 1717–1731. [Google Scholar] [CrossRef] [PubMed]
- Herold, N.; Anders-Osswein, M.; Glass, B.; Eckhardt, M.; Muller, B.; Krausslich, H.G. HIV-1 entry in SupT1-R5, CEM-ss, and primary CD4+ T cells occurs at the plasma membrane and does not require endocytosis. J. Virol. 2014, 88, 13956–13970. [Google Scholar] [CrossRef] [PubMed]
- Miyauchi, K.; Kim, Y.; Latinovic, O.; Morozov, V.; Melikyan, G.B. HIV Enters Cells via Endocytosis and Dynamin-Dependent Fusion with Endosomes. Cell 2009, 137, 433–444. [Google Scholar] [CrossRef] [PubMed]
- Marechal, V.; Prevost, M.C.; Petit, C.; Perret, E.; Heard, J.M.; Schwartz, O. Human immunodeficiency virus type 1 entry into macrophages mediated by macropinocytosis. J. Virol. 2001, 75, 11166–11177. [Google Scholar] [CrossRef] [PubMed]
- Bol, S.M.; van Remmerden, Y.; Sietzema, J.G.; Kootstra, N.A.; Schuitemaker, H.; van’t Wout, A.B. Donor variation in in vitro HIV-1 susceptibility of monocyte-derived macrophages. Virology 2009, 390, 205–211. [Google Scholar] [CrossRef] [PubMed]
- Zennou, V.; Petit, C.; Guetard, D.; Nerhbass, U.; Montagnier, L.; Charneau, P. HIV-1 genome nuclear import is mediated by a central DNA flap. Cell 2000, 101, 173–185. [Google Scholar] [CrossRef]
- Arhel, N.; Munier, S.; Souque, P.; Mollier, K.; Charneau, P. Nuclear import defect of human immunodeficiency virus type 1 DNA flap mutants is not dependent on the viral strain or target cell type. J. Virol. 2006, 80, 10262–10269. [Google Scholar] [CrossRef] [PubMed]
- Goujon, C.; Jarrosson-Wuilleme, L.; Bernaud, J.; Rigal, D.; Darlix, J.L.; Cimarelli, A. With a little help from a friend: Increasing HIV transduction of monocyte-derived dendritic cells with virion-like particles of SIV(MAC). Gene Ther. 2006, 13, 991–994. [Google Scholar] [CrossRef] [PubMed]
- Butler, S.L.; Hansen, M.S.; Bushman, F.D. A quantitative assay for HIV DNA integration in vivo. Nat. Med. 2001, 7, 631–634. [Google Scholar] [CrossRef] [PubMed]
| Target | Sequence (5′–3′) | Reference |
|---|---|---|
| 5′-LTR Fw | TTAAGCCTCAATAAAGCTTGCC | [43] |
| 5′-LTR Rv | GTTCGGGCGCCACTGCTAG | |
| 5′-LTR Probe | FAM–CCAGAGTCACACAACAGACGGGCA–BHQ1 | |
| Gag Fw | CATGTTTTCAGCATTATCAGAAGGA | [44] |
| Gag Rv | TGCTTGATGTCCCCCCACT | |
| Gag Probe | HEX—CCACCCCACAAGATTTAAACACCATGCTAA–BHQ1 | |
| 2-LTR Fw | CTAACTAGGGAACCCACTGCT | [45] |
| 2-LTR Rv | GTAGTTCTGCCAATCAGGGAA | |
| 2-LTR Probe | FAM—AGCCTCAATAAAGCTTGCCTTGAGTGC–BHQ1 | |
| RPP30 Fw | GATTTGGACCTGCGAGCG | [42] |
| RPP30 Rv | GCGGCTGTCTCCACAAGT | |
| RPP30 Probe | FAM–CTGACCTGAAGGCTCT–BHQ1 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bejarano, D.A.; Puertas, M.C.; Börner, K.; Martinez-Picado, J.; Müller, B.; Kräusslich, H.-G. Detailed Characterization of Early HIV-1 Replication Dynamics in Primary Human Macrophages. Viruses 2018, 10, 620. https://doi.org/10.3390/v10110620
Bejarano DA, Puertas MC, Börner K, Martinez-Picado J, Müller B, Kräusslich H-G. Detailed Characterization of Early HIV-1 Replication Dynamics in Primary Human Macrophages. Viruses. 2018; 10(11):620. https://doi.org/10.3390/v10110620
Chicago/Turabian StyleBejarano, David Alejandro, Maria C. Puertas, Kathleen Börner, Javier Martinez-Picado, Barbara Müller, and Hans-Georg Kräusslich. 2018. "Detailed Characterization of Early HIV-1 Replication Dynamics in Primary Human Macrophages" Viruses 10, no. 11: 620. https://doi.org/10.3390/v10110620
APA StyleBejarano, D. A., Puertas, M. C., Börner, K., Martinez-Picado, J., Müller, B., & Kräusslich, H.-G. (2018). Detailed Characterization of Early HIV-1 Replication Dynamics in Primary Human Macrophages. Viruses, 10(11), 620. https://doi.org/10.3390/v10110620





