Abstract
The major immediate-early (IE) gene of human cytomegalovirus (CMV) is believed to have a decisive role in acute infection and its activity is an important indicator of viral reactivation from latency. Although a variety of gene products are expressed from this region, the 72-kDa IE1 and the 86-kDa IE2 nuclear phosphoproteins are the most abundant and important. Both proteins have long been recognized as promiscuous transcriptional regulators. More recently, a critical role of the IE1 and IE2 proteins in counteracting non-adaptive host cell defense mechanisms has been revealed. In this review we will briefly summarize the available literature on IE1- and IE2-dependent mechanisms contributing to CMV evasion from intrinsic and innate immune responses.
1. Human Cytomegalovirus (CMV) Is a Significant Pathogen
CMV, the prototype β-herpesvirus, is the cause of a “silent pandemic” that continuously inflicts suffering upon people including immunocompromised patients as well as pregnant mothers and their unborn or prematurely newborn babies (reviewed in [1]). In the absence of an approved vaccine viral DNA polymerase inhibitors, including the nucleoside analogon ganciclovir, have provided major advances in CMV disease management. However, use of these drugs is limited by significant toxicity and relatively modest effectiveness due to poor bioavailability and problems with viral drug resistance (reviewed in [2]). Furthermore, no drug has been licensed to treat congenital CMV infection, and recent trials with newly developed antivirals targeting CMV have not lived up to their expectations. Thus, it is still imperative to develop new anti-CMV strategies directed at appropriate viral molecular targets.
2. The CMV Major Immediate-Early (IE) Proteins Are Multifunctional Key Regulators
2.1. Structure and Importance of the Major IE Gene
Within the >230,000 base pair CMV DNA genome, the major IE gene is believed to have a decisive role in acute infection and reactivation from viral latency. Through differential splicing, polyadenylation and promoter usage this viral genomic region produces multiple mRNAs (reviewed in [3]). Although a variety of protein products expressed from these mRNAs have been identified [4-7] (Figure 1), the 72-kDa IE1 and the 86-kDa IE2 nuclear phosphoproteins are the most abundant and important. They share 85 amino-terminalamino acids corresponding to major IE exons 2 and3 but have distinct carboxy-terminal parts encoded by exon 4(IE1) or exon 5 (IE2) (Figure 1). While IE2 is absolutely indispensable for productive viral replication [8,9], IE1 is only conditionally essential. In fact, IE1-null viruses replicate efficiently in fibroblasts at a high multiplicity of infection (MOI). However, the absence of IE1 results in severely attenuated viral replication under low MOI conditions [10,11].
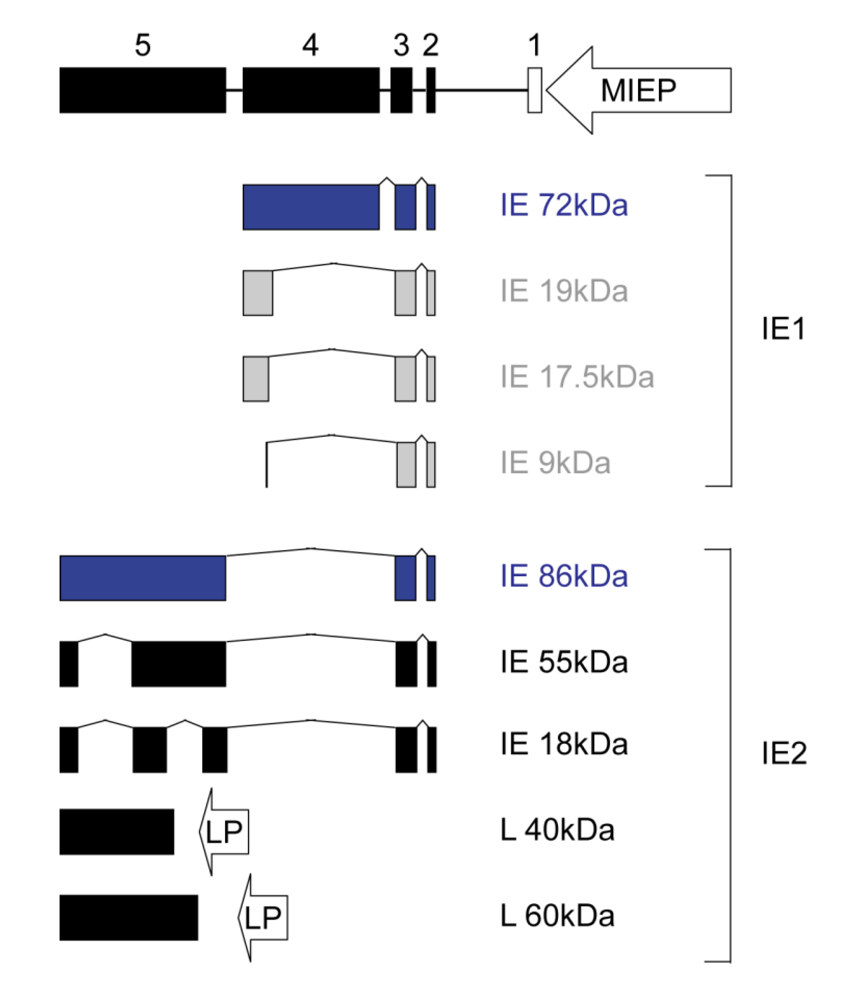
Figure 1.
Structural organization and protein products of the CMV major IE locus. At the top of the diagram, the lengths and relative positions of exons 1 to 5 (the four coding exons are presented as black boxes and non-coding exon 1 as open box) and the location of the major IE promoter-enhancer (MIEP) are shown. Proteins are subdivided into the IE1 (containing exon 4 sequences) and IE2 (containing exon 5 sequences) subfamilies. The predominant major IE protein species, which are the subject of this review, are highlighted in blue. It is uncertain whether the IE1 isoforms shown in gray are present in CMV-infected cells [4]. All IE proteins are expressed from differentially spliced mRNAs, but exon 5 also encodes at least two different late proteins whose mRNAs are transcribed from internal promoters (LP). None of the minor IE1 and IE2 protein isoforms has been characterized with respect to activities in intrinsic or innate immune evasion.
2.2. Functional Activities of the IE1 and IE2 Proteins
IE2 is the principal transcriptional activator of theCMV early genes [8,9]. In addition, IE2 negatively regulates viral gene expression including its own transcription (reviewed in [3,12]). IE1 appears to synergize with IE2 to promote transcriptional activation of the viral early genes, at least after low MOI infection [10,13]. Moreover, both IE1 and IE2 have been shown to activate certain host cell promoters (e.g., [6,14-20]) (reviewed in [3,12]). More recently, it has also been demonstrated that the two major IE proteins can individually block induction of distinct sets of potentially antiviral host genes [21-25]. In IE1, this activity depends at least partially on interactions with human signal transducer and activator of transcription (STAT) proteins. Beyond that, transcriptional regulation by IE1 and IE2 appears to involve multiple interactions with basal and accessory cellular transcription factors (reviewed in [1,3,12]) including histone modifying enzymes [24,26-30]. The latter are important for IE1- and IE2-dependent regulation of core histone tail acetylation and/or methylation [26-28,31]. In addition to their effects on the covalent modification and function of chromatin-associated proteins, IE1 and IE2 may also more directly act on DNA metabolism and structure. IE2 has been shown to block host cell DNA replication [32-34]. Furthermore, both major IE proteins seem to introduce mutations in cellular DNA [35], although the relevance of this observation remains to be determined. The chromatin-based activities of IE1 and IE2 are linked to their intranuclear localizations in ways that have not been fully elucidated. While IE2 binds sequence-specifically to DNA [36], IE1 does not seem to interact with DNA directly. However, IE1 associates with (mitotic) host cell chromatin [37,38], presumably via protein-protein interaction. In addition, both IE1 and IE2 target to interchromatinic matrix-associated nuclear domain 10 (ND10) compartments of the cell nucleus. IE1 disrupts these compartments most likely via interaction with the promyelocytic leukemia (PML) protein, the main structural organizer of ND10 [39-42].
Other activities associated with IE1 and IE2 expression concern effects on cell cycle progression and cell survival (reviewed in [43]). Ectopic IE1 can trigger a p53-dependent G1 growth arrest response [44], while it induces quiescent cells to enter S phase in p53-negative cells [45]. IE1 might stimulate S phase entry via interaction with the pocket protein p107 [46-48] and/or phosphorylation of p107, p130, and E2Fs through a reported kinase activity [49]. Similarly, transient IE2 expression may either promote cell cycle progression to the G1/S interface or growth arrest at G1/S or G2/M [33,44,50-52] (reviewed in [53]). IE2 may even induce premature cellular senescence [54]. At least some of the IE2-dependent effects on the cell cycle likely involve interactions with p53 and/or the retinoblastoma (Rb) tumor suppressor protein [55-59]. Finally, both IE1 and IE2 have been shown to counteract apoptotic cell death [60-65].
A comparative summary of activities that have been ascribed to the CMV IE1 and IE2 proteins is presented as Table 1.

Table 1.
Common and distinct activities of the CMV IE1 and IE2 proteins.
| Activities | IE1 | IE2 | Selected References |
|---|---|---|---|
| CMV replication | |||
| Requirement for viral replication at low MOI | + | + | [8-11] |
| Requirement for viral replication at high MOI | – | + | [8-11] |
| Cell cycle and apoptosis | |||
| Inhibition of cell cycle progression | + | + | [33, 44,50-52] |
| Induction of cell cycle progression | + | + | [33, 45,51,66] |
| Inhibition of apoptosis | + | + | [60-65] |
| Induction of senescence | – | + | [54] |
| Nuclear structures | |||
| ND10 targeting | + | + | [39,40,42] |
| ND10 disruption | + | – | [39,40,42] |
| Mitotic chromatin association | + | – | [37,38] |
| DNA binding | – | + | [36] |
| Histone modification | |||
| Core histone tail acetylation | + | (+) | [26] |
| Core histone tail methylation | – | + | [27,31] |
| DNA metabolism | |||
| Inhibition of cellular DNA replication | – | + | [32-34] |
| Induction of mutations in cellular DNA | + | + | [35] |
| Transcription | |||
| Activation of viral genes | + | + | reviewed in [1,3,12] |
| Repression of viral genes | – | + | reviewed in [1,3,12] |
| Activation of cellular genes | + | + | [6,14-20] |
| Inhibition of cellular gene activation | + | + | [21-25] |
| Other activities | |||
| Kinase activity | + | – | [49] |
+, positive; (+), likely positive; –, negative or unknown. Shading indicates activities most relevant to this review.
2.3. IE1 and IE2 in Innate and Intrinsic Immunity
The innate (in the broader sense) immune system of many organisms includes both inducible (innate in the stricter sense) and constitutive (intrinsic) mechanisms. Inducible innate mechanisms largely depend on pathogen-initiated cytokine production and signaling. Intrinsic responses like apoptosis, autophagy, and viral genome-directed repression mechanisms have only recently been recognized as an essential component of immunity which gives all cells the capacity to respond to infection. In the following paragraphs we will expand on the activities of CMV IE1 and IE2 that have established the two viral proteins as antagonists of innate and intrinsic antiviral host responses.
3. The CMV IE1 and IE2 Proteins Exhibit Antiapoptotic Potential
3.2. Role of IE1 and IE2 in Apoptosis Inhibition
Although it is now clear that cytomegaloviruses employ multiple strategies to delay cell death in infected cells (reviewed in [73]), the IE1 and IE2 proteins were the first CMV gene products reported to block apoptosis [60]. Following transient or stable expression in HeLa cells, IE1 and IE2 individually inhibited induction of extrinsic apoptosis by short exposure to TNF-α and cycloheximide or by infection with a proapoptotic mutant (E1B-19kDa-deficient) adenovirus. However, the viral proteins did not protect HeLa cells from TNF-α- or Fas-mediated apoptosis under more stringent experimental conditions [74] or when cell death was triggered by irradiation with ultraviolet (UV) light [60]. Subsequently, the anti-apoptotic effects of ectopic IE1 and/or IE2 expression have been confirmed in other cancer cell lines and in primary cells [28,61-64,75]. However, the impact of the IE1- and IE2-associated pro-life activities on CMV infection remain unexplored.
There is no evidence that IE1 or IE2 interfere with mitochondria-related apoptotic processes, and the two proteins do not alter the expression of Bcl-2 or Bax [60]. Instead, IE1 and IE2, expressed in concert or individually, inhibit apoptosis by activating the phosphatidylinositide 3’-OH kinase (PI3K) pro-survival pathway [61,62], which is also induced by CMV infection [62,76,77]. This has been demonstrated in the ts13 cell line, which carries a temperature-sensitive allele of the gene encoding TAFII250 and therefore undergoes apoptosis at the non-permissive temperature. Furthermore, combined expression of the viral IE proteins increased the activity of the serine/threonine kinase Akt (also known as protein kinase B), a major PI3K downstream target (reviewed in [78,79]). Akt promotes cell survival in part by targeting IκB kinase, which phoshorylates IκB resulting in nuclear localization of NFκB and activation of NFκB-responsive promoters of antiapoptotic genes. In fact, several cellular [15-17], viral [80-82] and artificial [62] promoters have been shown to be transcriptionally activated by CMV IE1 in an NFκB-dependent fashion. However, IE2 may rather repress than stimulate transcription from NFκB-regulated promoters [21,22,25,83]. On the other hand, IE2 appears to activate expression of cellular FLIP in CMV-infected retinal pigment epithelial cells and human retina tissue [65]. FLIP blocks the apoptotic pathway by interacting with caspase-8 at the death-inducing signaling complex. Notably, IE2-specific up-regulation of FLIP in CMV-infected retinal cells depends on PI3K [65].
In addition, mechanisms involving the tumor suppressor protein p53 have been proposed to account for the observed inhibitory effects of CMV IE2 on cellular apoptosis. IE2 binds to p53 and interferes with the tumor suppressor protein’s transcriptional activator function [55,56,84]. It was further demonstrated that IE2 can repress the acetylase activity of p300/cAMP response element binding protein binding protein (CBP) towards p53, rendering the tumor suppressor protein unable to execute UV-dependent apoptosis of colon cancer cells [28]. Moreover, expression of IE2, but not IE1, protects smooth muscle cells from p53-mediated apoptosis [63]. Rather than p53, IE1 targets the tumor suppressor protein PML [41], but the functional impact of this interaction on cell survival has not been evaluated. This potential link warrants future investigation since PML is known to affect PI3K signaling, p53 activity, and apoptosis (reviewed in [85,86]).
In summary, it appears that each of the CMV major IE proteins can block extrinsic apoptosis pathways via activation of PI3K signaling, although no physical interaction partner (besides PML) of IE1 or IE2 has so far been identified in this pathway. Beyond that, additional mechanisms likely contribute to inhibition of apoptosis by the viral proteins that may involve IE2-p53 complex formation and other known or unidentified interactions. Despite the fact that the antiapoptotic potential of the two major IE proteins has clearly been established in several overexpression settings, its true relevance for viral infection and pathogenesis remains to be determined.
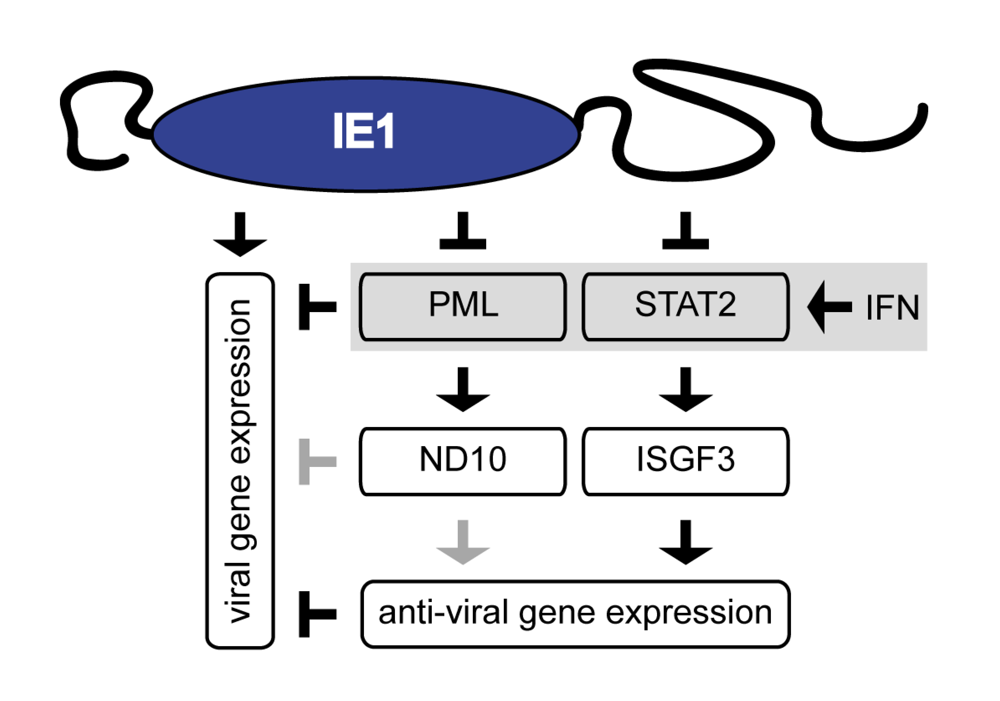
6. Conclusions
The innate host cell response to viral infection is in large parts characterized by the induction of cytokines and inflammatory chemokines. In addition, constitutively active intrinsic antiviral defense mechanisms (e.g., apoptosis and chromatin-based repression) ubiquitously exist. The IE1 and IE2 proteins are among the first de novo synthesized proteins following primary CMV infection or viral reactivation. It emerges that a principal task of IE1 and IE2 is to counteract intrinsic and innate host responses that would otherwise terminate the viral life cycle in its very beginnings. In particular, IE1 antagonizes apoptosis, ND10-related transcription silencing, and type I IFN signaling. Likewise, IE2 inhibits apoptosis and inflammatory cytokine/chemokine induction. In comparison to the long recognized direct effects of IE1 and IE2 on viral transcription, their rather recently discovered immune evasion activities may turn out to be equally important to assure viral replicative success. Consequently, the interactions of IE1/IE2 with host cell innate and intrinsic defense pathways may provide new opportunities for antiviral intervention.
Acknowledgments
We apologize to our many colleagues whose publications were not cited. This work was partly supported by the European Union TargetHerpes grant (LSHG-CT-2006-037517). We thank Michael Schnitzbauer and Carla Winterling for helpful comments on the manuscript and Hans Wolf for continuous support.
References and Notes
- Mocarski, E.S.; Shenk, T.; Pass, R.F. Cytomegaloviruses. In Fields Virology, 5th; Knipe, D.M., Howley, P.M., Griffin, D.E., Lamb, R.A., Martin, M.A., Eds.; 2007; Lippincott Williams & Wilkins: Philadelphia, PA, USA; Volume 2, pp. 2701–2773. [Google Scholar]
- Mercorelli, B.; Sinigalia, E.; Loregian, A.; Palu, G. Human cytomegalovirus DNA replication: antiviral targets and drugs. Rev. Med. Virol. 2008, 18, 177–210. [Google Scholar] [CrossRef] [PubMed]
- Stinski, M.F.; Meier, J.L. Immediate-early viral gene regulation and function. In Human Herpesviruses - Biology, Therapy, and Immunoprophylaxis, 1st; Arvin, A., Campadelli-Fiume, G., Mocarski, E., Moore, P.S., Roizman, B., Whitley, R., Yamanishi, K., Eds.; 2007; Cambridge University Press: Cambridge, UK; pp. 241–263. [Google Scholar]
- Awasthi, S.; Isler, J.A.; Alwine, J.C. Analysis of splice variants of the immediate-early 1 region of human cytomegalovirus. J. Virol. 2004, 78, 8191–8200. [Google Scholar] [CrossRef] [PubMed]
- Pizzorno, M.C.; Mullen, M.A.; Chang, Y.N.; Hayward, G.S. The functionally active IE2 immediate-early regulatory protein of human cytomegalovirus is an 80-kilodalton polypeptide that contains two distinct activator domains and a duplicated nuclear localization signal. J. Virol. 1991, 65, 3839–3852. [Google Scholar] [PubMed]
- Shirakata, M.; Terauchi, M.; Ablikim, M.; Imadome, K.; Hirai, K.; Aso, T.; Yamanashi, Y. Novel immediate-early protein IE19 of human cytomegalovirus activates the origin recognition complex I promoter in a cooperative manner with IE72. J. Virol. 2002, 76, 3158–3167. [Google Scholar] [CrossRef] [PubMed]
- Kerry, J.A.; Sehgal, A.; Barlow, S.W.; Cavanaugh, V.J.; Fish, K.; Nelson, J.A.; Stenberg, R.M. Isolation and characterization of a low-abundance splice variant from the human cytomegalovirus major immediate-early gene region. J. Virol. 1995, 69, 3868–3872. [Google Scholar] [PubMed]
- Heider, J.A.; Bresnahan, W.A.; Shenk, T.E. Construction of a rationally designed human cytomegalovirus variant encoding a temperature-sensitive immediate-early 2 protein. Proc. Natl. Acad. Sci. USA 2002, 99, 3141–3146. [Google Scholar] [CrossRef]
- Marchini, A.; Liu, H.; Zhu, H. Human cytomegalovirus with IE-2 (UL122) deleted fails to express early lytic genes. J. Virol. 2001, 75, 1870–1878. [Google Scholar] [CrossRef] [PubMed]
- Greaves, R.F.; Mocarski, E.S. Defective growth correlates with reduced accumulation of a viral DNA replication protein after low-multiplicity infection by a human cytomegalovirus ie1 mutant. J. Virol. 1998, 72, 366–379. [Google Scholar] [PubMed]
- Mocarski, E.S.; Kemble, G.W.; Lyle, J.M.; Greaves, R.F. A deletion mutant in the human cytomegalovirus gene encoding IE1(491aa) is replication defective due to a failure in autoregulation. Proc. Natl. Acad. Sci. USA 1996, 93, 11321–11326. [Google Scholar] [CrossRef]
- White, E.A.; Spector, D.H. Early viral gene expression and function. In >Human Herpesviruses - Biology, Therapy, and Immunoprophylaxis, 1st; Arvin, A., Campadelli-Fiume, G., Mocarski, E., Moore, P.S., Roizman, B., Whitley, R., Yamanishi, K., Eds.; 2007; pp. 264–294. Cambridge University Press: Cambridge, UK. [Google Scholar]
- Gawn, J.M.; Greaves, R.F. Absence of IE1 p72 protein function during low-multiplicity infection by human cytomegalovirus results in a broad block to viral delayed-early gene expression. J. Virol. 2002, 76, 4441–4455. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.J.; Stinski, M.F. Effect of the human cytomegalovirus IE86 protein on expression of E2F-responsive genes: a DNA microarray analysis. Proc. Natl. Acad. Sci. USA 2002, 99, 2836–2841. [Google Scholar] [PubMed]
- Murayama, T.; Mukaida, N.; Sadanari, H.; Yamaguchi, N.; Khabar, K.S.; Tanaka, J.; Matsushima, K.; Mori, S.; Eizuru, Y. The immediate early gene 1 product of human cytomegalovirus is sufficient for up-regulation of interleukin-8 gene expression. Biochem. Biophys. Res. Commun. 2000, 279, 298–304. [Google Scholar] [CrossRef] [PubMed]
- Geist, L.J.; Dai, L.Y. Cytomegalovirus modulates interleukin-6 gene expression. Transplantation 1996, 62, 653–658. [Google Scholar] [CrossRef] [PubMed]
- Geist, L.J.; Hopkins, H.A.; Dai, L.Y.; He, B.; Monick, M.M.; Hunninghake, G.W. Cytomegalovirus modulates transcription factors necessary for the activation of the tumor necrosis factor-alpha promoter. Am. J. Respir. Cell Mol. Biol. 1997, 16, 31–37. [Google Scholar] [PubMed]
- Margolis, M.J.; Pajovic, S.; Wong, E.L.; Wade, M.; Jupp, R.; Nelson, J.A.; Azizkhan, J.C. Interaction of the 72-kilodalton human cytomegalovirus IE1 gene product with E2F1 coincides with E2F-dependent activation of dihydrofolate reductase transcription. J. Virol. 1995, 69, 7759–7767. [Google Scholar] [PubMed]
- Straat, K.; Liu, C.; Rahbar, A.; Zhu, Q.; Liu, L.; Wolmer-Solberg, N.; Lou, F.; Liu, Z.; Shen, J.; Jia, J.; Kyo, S.; Bjorkholm, M.; Sjoberg, J.; Soderberg-Naucler, C.; Xu, D. Activation of telomerase by human cytomegalovirus. J. Natl. Cancer Inst. 2009, 101, 488–497. [Google Scholar] [CrossRef] [PubMed]
- Hayhurst, G.P.; Bryant, L.A.; Caswell, R.C.; Walker, S.M.; Sinclair, J.H. CCAAT box-dependent activation of the TATA-less human DNA polymerase alpha promoter by the human cytomegalovirus 72-kilodalton major immediate-early protein. J. Virol. 1995, 69, 182–188. [Google Scholar] [PubMed]
- Taylor, R.T.; Bresnahan, W.A. Human cytomegalovirus IE86 attenuates virus- and tumor necrosis factor alpha-induced NFkappaB-dependent gene expression. J. Virol. 2006, 80, 10763–10771. [Google Scholar] [CrossRef] [PubMed]
- Taylor, R.T.; Bresnahan, W.A. Human cytomegalovirus immediate-early 2 gene expression blocks virus-induced beta interferon production. J. Virol. 2005, 79, 3873–3877. [Google Scholar] [CrossRef] [PubMed]
- Paulus, C.; Krauss, S.; Nevels, M. A human cytomegalovirus antagonist of type I IFN-dependent signal transducer and activator of transcription signaling. Proc. Natl. Acad. Sci. USA 2006, 103, 3840–3845. [Google Scholar] [CrossRef]
- Huh, Y.H.; Kim, Y.E.; Kim, E.T.; Park, J.J.; Song, M.J.; Zhu, H.; Hayward, G.S.; Ahn, J.H. Binding STAT2 by the acidic domain of human cytomegalovirus IE1 promotes viral growth and is negatively regulated by SUMO. J. Virol. 2008, 82, 10444–10454. [Google Scholar] [CrossRef] [PubMed]
- Taylor, R.T.; Bresnahan, W.A. Human cytomegalovirus immediate-early 2 protein IE86 blocks virus-induced chemokine expression. J. Virol. 2006, 80, 920–928. [Google Scholar] [CrossRef] [PubMed]
- Nevels, M.; Paulus, C.; Shenk, T. Human cytomegalovirus immediate-early 1 protein facilitates viral replication by antagonizing histone deacetylation. Proc. Natl. Acad. Sci. USA 2004, 101, 17234–17239. [Google Scholar] [CrossRef]
- Reeves, M.; Murphy, J.; Greaves, R.; Fairley, J.; Brehm, A.; Sinclair, J. Autorepression of the human cytomegalovirus major immediate-early promoter/enhancer at late times of infection is mediated by the recruitment of chromatin remodeling enzymes by IE86. J. Virol. 2006, 80, 9998–10009. [Google Scholar] [CrossRef] [PubMed]
- Hsu, C.H.; Chang, M.D.; Tai, K.Y.; Yang, Y.T.; Wang, P.S.; Chen, C.J.; Wang, Y.H.; Lee, S.C.; Wu, C.W.; Juan, L.J. HCMV IE2-mediated inhibition of HAT activity downregulates p53 function. EMBO J. 2004, 23, 2269–2280. [Google Scholar] [CrossRef] [PubMed]
- Bryant, L.A.; Mixon, P.; Davidson, M.; Bannister, A.J.; Kouzarides, T.; Sinclair, J.H. The human cytomegalovirus 86-kilodalton major immediate-early protein interacts physically and functionally with histone acetyltransferase P/CAF. J. Virol. 2000, 74, 7230–7237. [Google Scholar] [CrossRef] [PubMed]
- Park, J.J.; Kim, Y.E.; Pham, H.T.; Kim, E.T.; Chung, Y.H.; Ahn, J.H. Functional interaction of the human cytomegalovirus IE2 protein with histone deacetylase 2 in infected human fibroblasts. J. Gen. Virol. 2007, 88, 3214–3223. [Google Scholar] [CrossRef] [PubMed]
- Cuevas-Bennett, C.; Shenk, T. Dynamic histone H3 acetylation and methylation at human cytomegalovirus promoters during replication in fibroblasts. J. Virol. 2008, 82, 9525–9536. [Google Scholar] [CrossRef] [PubMed]
- Wiebusch, L.; Hagemeier, C. The human cytomegalovirus immediate early 2 protein dissociates cellular DNA synthesis from cyclin-dependent kinase activation. EMBO J. 2001, 20, 1086–1098. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.J.; Stinski, M.F. Inhibition of cell division by the human cytomegalovirus IE86 protein: role of the p53 pathway or cyclin-dependent kinase 1/cyclin B1. J. Virol. 2005, 79, 2597–2603. [Google Scholar] [CrossRef] [PubMed]
- Petrik, D.T.; Schmitt, K.P.; Stinski, M.F. Inhibition of cellular DNA synthesis by the human cytomegalovirus IE86 protein is necessary for efficient virus replication. J. Virol. 2006, 80, 3872–3883. [Google Scholar] [CrossRef] [PubMed]
- Shen, Y.; Zhu, H.; Shenk, T. Human cytomagalovirus IE1 and IE2 proteins are mutagenic and mediate "hit-and-run" oncogenic transformation in cooperation with the adenovirus E1A proteins. Proc. Natl. Acad. Sci. USA 1997, 94, 3341–3345. [Google Scholar] [CrossRef]
- Macias, M.P.; Stinski, M.F. An in vitro system for human cytomegalovirus immediate early 2 protein (IE2)-mediated site-dependent repression of transcription and direct binding of IE2 to the major immediate early promoter. Proc. Natl. Acad. Sci. USA 1993, 90, 707–711. [Google Scholar] [CrossRef]
- Lafemina, R.L.; Pizzorno, M.C.; Mosca, J.D.; Hayward, G.S. Expression of the acidic nuclear immediate-early protein (IE1) of human cytomegalovirus in stable cell lines and its preferential association with metaphase chromosomes. Virology 1989, 172, 584–600. [Google Scholar] [CrossRef] [PubMed]
- Reinhardt, J.; Smith, G.B.; Himmelheber, C.T.; Azizkhan-Clifford, J.; Mocarski, E.S. The carboxyl-terminal region of human cytomegalovirus IE1491aa contains an acidic domain that plays a regulatory role and a chromatin-tethering domain that is dispensable during viral replication. J. Virol. 2005, 79, 225–233. [Google Scholar] [CrossRef] [PubMed]
- Wilkinson, G.W.; Kelly, C.; Sinclair, J.H.; Rickards, C. Disruption of PML-associated nuclear bodies mediated by the human cytomegalovirus major immediate early gene product. J. Gen. Virol. 1998, 79, 1233–1245. [Google Scholar] [PubMed]
- Korioth, F.; Maul, G.G.; Plachter, B.; Stamminger, T.; Frey, J. The nuclear domain 10 (ND10) is disrupted by the human cytomegalovirus gene product IE1. Exp. Cell Res. 1996, 229, 155–158. [Google Scholar] [CrossRef] [PubMed]
- hn, J.H.; Brignole, E.J.; Hayward, G.S. A Disruption of PML subnuclear domains by the acidic IE1 protein of human cytomegalovirus is mediated through interaction with PML and may modulate a RING finger-dependent cryptic transactivator function of PML . Mol. Cell. Biol. 1998, 18 , 4899–4913. [Google Scholar] [PubMed]
- Ahn, J.H.; Hayward, G.S. The major immediate-early proteins IE1 and IE2 of human cytomegalovirus colocalize with and disrupt PML-associated nuclear bodies at very early times in infected permissive cells. J. Virol. 1997, 71, 4599–4613. [Google Scholar] [PubMed]
- Sanchez, V.; Spector, D.H. Subversion of cell cycle regulatory pathways. Curr. Top. Microbiol. Immunol. 2008, 325, 243–262. [Google Scholar] [PubMed]
- Castillo, J.P.; Frame, F.M.; Rogoff, H.A.; Pickering, M.T.; Yurochko, A.D.; Kowalik, T.F. Human cytomegalovirus IE1-72 activates ataxia telangiectasia mutated kinase and a p53/p21-mediated growth arrest response. J. Virol. 2005, 79, 11467–11475. [Google Scholar] [CrossRef] [PubMed]
- Castillo, J.P.; Yurochko, A.D.; Kowalik, T.F. Role of human cytomegalovirus immediate-early proteins in cell growth control. J. Virol. 2000, 74, 8028–8037. [Google Scholar] [CrossRef] [PubMed]
- Poma, E.E.; Kowalik, T.F.; Zhu, L.; Sinclair, J.H.; Huang, E.S. The human cytomegalovirus IE1-72 protein interacts with the cellular p107 protein and relieves p107-mediated transcriptional repression of an E2F-responsive promoter. J. Virol. 1996, 70, 7867–7877. [Google Scholar] [PubMed]
- Zhang, Z.; Huong, S.M.; Wang, X.; Huang, D.Y.; Huang, E.S. Interactions between human cytomegalovirus IE1-72 and cellular p107: functional domains and mechanisms of up-regulation of cyclin E/cdk2 kinase activity. J. Virol. 2003, 77, 12660–12670. [Google Scholar] [CrossRef] [PubMed]
- Johnson, R.A.; Yurochko, A.D.; Poma, E.E.; Zhu, L.; Huang, E.S. Domain mapping of the human cytomegalovirus IE1-72 and cellular p107 protein-protein interaction and the possible functional consequences. J. Gen. Virol. 1999, 80, 1293–1303. [Google Scholar] [PubMed]
- Pajovic, S.; Wong, E.L.; Black, A.R.; Azizkhan, J.C. Identification of a viral kinase that phosphorylates specific E2Fs and pocket proteins. Mol. Cell. Biol. 1997, 17, 6459–6464. [Google Scholar] [PubMed]
- Wiebusch, L.; Hagemeier, C. Human cytomegalovirus 86-kilodalton IE2 protein blocks cell cycle progression in G(1). J. Virol. 1999, 73, 9274–9283. [Google Scholar] [PubMed]
- Wiebusch, L.; Asmar, J.; Uecker, R.; Hagemeier, C. Human cytomegalovirus immediate-early protein 2 (IE2)-mediated activation of cyclin E is cell-cycle-independent and forces S-phase entry in IE2-arrested cells. J. Gen. Virol. 2003, 84, 51–60. [Google Scholar] [CrossRef] [PubMed]
- Murphy, E.A.; Streblow, D.N.; Nelson, J.A.; Stinski, M.F. The human cytomegalovirus IE86 protein can block cell cycle progression after inducing transition into the S phase of permissive cells. J. Virol. 2000, 74, 7108–7118. [Google Scholar] [CrossRef] [PubMed]
- Stinski, M.F.; Petrik, D.T. Functional roles of the human cytomegalovirus essential IE86 protein. Curr. Top. Microbiol. Immunol. 2008, 325, 133–152. [Google Scholar] [PubMed]
- Noris, E.; Zannetti, C.; Demurtas, A.; Sinclair, J.; De Andrea, M.; Gariglio, M.; Landolfo, S. Cell cycle arrest by human cytomegalovirus 86-kDa IE2 protein resembles premature senescence. J. Virol. 2002, 76, 12135–12148. [Google Scholar] [CrossRef] [PubMed]
- Speir, E.; Modali, R.; Huang, E.S.; Leon, M.B.; Shawl, F.; Finkel, T.; Epstein, S.E. Potential role of human cytomegalovirus and p53 interaction in coronary restenosis. Science 1994, 265, 391–394. [Google Scholar] [PubMed]
- Bonin, L.R.; McDougall, J.K. Human cytomegalovirus IE2 86-kilodalton protein binds p53 but does not abrogate G1 checkpoint function. J. Virol. 1997, 71, 5861–5870. [Google Scholar] [PubMed]
- Fortunato, E.A.; Sommer, M.H.; Yoder, K.; Spector, D.H. Identification of domains within the human cytomegalovirus major immediate-early 86-kilodalton protein and the retinoblastoma protein required for physical and functional interaction with each other. J. Virol. 1997, 71, 8176–8185. [Google Scholar] [PubMed]
- Hagemeier, C.; Caswell, R.; Hayhurst, G.; Sinclair, J.; Kouzarides, T. Functional interaction between the HCMV IE2 transactivator and the retinoblastoma protein. EMBO J. 1994, 13, 2897–2903. [Google Scholar] [PubMed]
- Sommer, M.H.; Scully, A.L.; Spector, D.H. Transactivation by the human cytomegalovirus IE2 86-kilodalton protein requires a domain that binds to both the TATA box-binding protein and the retinoblastoma protein. J. Virol. 1994, 68, 6223–6231. [Google Scholar] [PubMed]
- Zhu, H.; Shen, Y.; Shenk, T. Human cytomegalovirus IE1 and IE2 proteins block apoptosis. J. Virol. 1995, 69, 7960–7970. [Google Scholar] [PubMed]
- Lukac, D.M.; Alwine, J.C. Effects of human cytomegalovirus major immediate-early proteins in controlling the cell cycle and inhibiting apoptosis: studies with ts13 cells. J. Virol. 1999, 73, 2825–2831. [Google Scholar] [PubMed]
- Yu, Y.; Alwine, J.C. Human cytomegalovirus major immediate-early proteins and simian virus 40 large T antigen can inhibit apoptosis through activation of the phosphatidylinositide 3'-OH kinase pathway and the cellular kinase Akt. J. Virol. 2002, 76, 3731–3738. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, K.; Zou, J.P.; Takeda, K.; Ferrans, V.J.; Sandford, G.R.; Johnson, T.M.; Finkel, T.; Epstein, S.E. Effects of human cytomegalovirus immediate-early proteins on p53-mediated apoptosis in coronary artery smooth muscle cells. Circulation 1999, 99, 1656–1659. [Google Scholar] [PubMed]
- Kim, J.; Kwon, Y.J.; Park, E.S.; Sung, B.; Kim, J.H.; Park, C.G.; Hwang, E.S.; Cha, C.Y. Human cytomegalovirus (HCMV) IE1 plays role in resistance to apoptosis with etoposide in cancer cell line by Cdk2 accumulation. Microbiol. Immunol. 2003, 47, 959–967. [Google Scholar] [PubMed]
- Chiou, S.H.; Yang, Y.P.; Lin, J.C.; Hsu, C.H.; Jhang, H.C.; Yang, Y.T.; Lee, C.H.; Ho, L.L.; Hsu, W.M.; Ku, H.H.; Chen, S.J.; Chen, S.S.; Chang, M.D.; Wu, C.W.; Juan, L.J. The immediate early 2 protein of human cytomegalovirus (HCMV) mediates the apoptotic control in HCMV retinitis through up-regulation of the cellular FLICE-inhibitory protein expression. J. Immunol. 2006, 177, 6199–6206. [Google Scholar] [PubMed]
- Cobbs, C.S.; Soroceanu, L.; Denham, S.; Zhang, W.; Kraus, M.H. Modulation of oncogenic phenotype in human glioma cells by cytomegalovirus IE1-mediated mitogenicity. Cancer Res. 2008, 68, 724–730. [Google Scholar] [CrossRef] [PubMed]
- Kroemer, G.; Galluzzi, L.; Brenner, C. Mitochondrial membrane permeabilization in cell death. Physiol. Rev. 2007, 87, 99–163. [Google Scholar] [CrossRef] [PubMed]
- Elmore, S. Apoptosis: a review of programmed cell death. Toxicol. Pathol. 2007, 35, 495–516. [Google Scholar] [CrossRef] [PubMed]
- Youle, R.J.; Strasser, A. The BCL-2 protein family: opposing activities that mediate cell death. Nat. Rev. Mol. Cell Biol. 2008, 9, 47–59. [Google Scholar] [CrossRef]
- Laptenko, O.; Prives, C. Transcriptional regulation by p53: one protein, many possibilities. Cell Death Differ. 2006, 13, 951–961. [Google Scholar] [CrossRef] [PubMed]
- Wilson, N.S.; Dixit, V.; Ashkenazi, A. Death receptor signal transducers: nodes of coordination in immune signaling networks. Nat. Immunol. 2009, 10, 348–355. [Google Scholar] [CrossRef] [PubMed]
- Galluzzi, L.; Brenner, C.; Morselli, E.; Touat, Z.; Kroemer, G. Viral control of mitochondrial apoptosis . PLoS Pathog. 2008, 4 , e1000018. [Google Scholar] [CrossRef] [PubMed]
- McCormick, A.L. Control of apoptosis by human cytomegalovirus. Curr. Top. Microbiol. Immunol. 2008, 325, 281–295. [Google Scholar] [PubMed]
- Goldmacher, V.S.; Bartle, L.M.; Skaletskaya, A.; Dionne, C.A.; Kedersha, N.L.; Vater, C.A.; Han, J.W.; Lutz, R.J.; Watanabe, S.; Cahir McFarland, E.D.; Kieff, E.D.; Mocarski, E.S.; Chittenden, T. A cytomegalovirus-encoded mitochondria-localized inhibitor of apoptosis structurally unrelated to Bcl-2. Proc. Natl. Acad. Sci. USA 1999, 96, 12536–12541. [Google Scholar] [CrossRef]
- Bai, Z.; Li, L.; Wang, B.; Liu, Z.; Wang, H.; Yan, Z.; Qian, D.; Ding, S.; Song, X. Effect of inducible expressed human cytomegalovirus immediate early 86 protein on cell apoptosis. Biosci. Biotechnol. Biochem. 2009, 73, 1268–1273. [Google Scholar] [CrossRef] [PubMed]
- Kudchodkar, S.B.; Yu, Y.; Maguire, T.G.; Alwine, J.C. Human cytomegalovirus infection alters the substrate specificities and rapamycin sensitivities of raptor- and rictor-containing complexes. Proc. Natl. Acad. Sci. USA 2006, 103, 14182–14187. [Google Scholar] [CrossRef]
- Johnson, R.A.; Wang, X.; Ma, X.L.; Huong, S.M.; Huang, E.S. Human cytomegalovirus up-regulates the phosphatidylinositol 3-kinase (PI3-K) pathway: inhibition of PI3-K activity inhibits viral replication and virus-induced signaling. J. Virol. 2001, 75, 6022–6032. [Google Scholar] [CrossRef] [PubMed]
- Alwine, J.C. Modulation of host cell stress responses by human cytomegalovirus. Curr. Top. Microbiol. Immunol. 2008, 325, 263–279. [Google Scholar] [PubMed]
- Buchkovich, N.J.; Yu, Y.; Zampieri, C.A.; Alwine, J.C. The TORrid affairs of viruses: effects of mammalian DNA viruses on the PI3K-Akt-mTOR signalling pathway. Nat. Rev. Microbiol. 2008, 6, 266–275. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Yu, S.S.; Kim, V.N. Essential role of NF-kappa B in transactivation of the human immunodeficiency virus long terminal repeat by the human cytomegalovirus 1E1 protein. J. Gen. Virol. 1996, 77, 83–91. [Google Scholar] [CrossRef] [PubMed]
- Sambucetti, L.C.; Cherrington, J.M.; Wilkinson, G.W.; Mocarski, E.S. NF-kappa B activation of the cytomegalovirus enhancer is mediated by a viral transactivator and by T cell stimulation. EMBO J. 1989, 8, 4251–4258. [Google Scholar] [PubMed]
- Cherrington, J.M.; Mocarski, E.S. Human cytomegalovirus ie1 transactivates the alpha promoter-enhancer via an 18-base-pair repeat element. J. Virol. 1989, 63, 1435–1440. [Google Scholar] [PubMed]
- Gealy, C.; Humphreys, C.; Dickinson, V.; Stinski, M.; Caswell, R. An activation-defective mutant of the human cytomegalovirus IE2p86 protein inhibits NF-kappaB-mediated stimulation of the human interleukin-6 promoter. J. Gen. Virol. 2007, 88, 2435–2440. [Google Scholar] [CrossRef] [PubMed]
- Tsai, H.L.; Kou, G.H.; Chen, S.C.; Wu, C.W.; Lin, Y.S. Human cytomegalovirus immediate-early protein IE2 tethers a transcriptional repression domain to p53. J. Biol. Chem. 1996, 271, 3534–3540. [Google Scholar] [CrossRef] [PubMed]
- Borden, K.L.; Culjkovic, B. Perspectives in PML: a unifying framework for PML function. Front. Biosci. 2009, 14, 497–509. [Google Scholar] [CrossRef] [PubMed]
- Borden, K.L. Pondering the puzzle of PML (promyelocytic leukemia) nuclear bodies: can we fit the pieces together using an RNA regulon? Biochim. Biophys. Acta 2008, 1783, 2145–2154. [Google Scholar] [CrossRef] [PubMed]
- Tavalai, N.; Stamminger, T. New insights into the role of the subnuclear structure ND10 for viral infection. Biochim. Biophys. Acta 2008, 1783, 2207–2221. [Google Scholar] [CrossRef] [PubMed]
- Ishov, A.M.; Stenberg, R.M.; Maul, G.G. Human cytomegalovirus immediate early interaction with host nuclear structures: definition of an immediate transcript environment. J. Cell Biol. 1997, 138, 5–16. [Google Scholar] [CrossRef] [PubMed]
- Sourvinos, G.; Tavalai, N.; Berndt, A.; Spandidos, D.A.; Stamminger, T. Recruitment of human cytomegalovirus immediate-early 2 protein onto parental viral genomes in association with ND10 in live-infected cells. J. Virol. 2007, 81, 10123–10136. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.R.; Kim, D.J.; Lee, J.M.; Choi, C.Y.; Ahn, B.Y.; Hayward, G.S.; Ahn, J.H. Ability of the human cytomegalovirus IE1 protein to modulate sumoylation of PML correlates with its functional activities in transcriptional regulation and infectivity in cultured fibroblast cells. J. Virol. 2004, 78, 6527–6542. [Google Scholar] [CrossRef] [PubMed]
- Muller, S.; Dejean, A. Viral immediate-early proteins abrogate the modification by SUMO-1 of PML and Sp100 proteins, correlating with nuclear body disruption. J. Virol. 1999, 73, 5137–5143. [Google Scholar] [PubMed]
- Ahn, J.H.; Hayward, G.S. Disruption of PML-associated nuclear bodies by IE1 correlates with efficient early stages of viral gene expression and DNA replication in human cytomegalovirus infection. Virology 2000, 274, 39–55. [Google Scholar] [CrossRef] [PubMed]
- Tavalai, N.; Papior, P.; Rechter, S.; Leis, M.; Stamminger, T. Evidence for a role of the cellular ND10 protein PML in mediating intrinsic immunity against human cytomegalovirus infections. J. Virol. 2006, 80, 8006–8018. [Google Scholar] [CrossRef] [PubMed]
- Tavalai, N.; Papior, P.; Rechter, S.; Stamminger, T. Nuclear domain 10 components promyelocytic leukemia protein and hDaxx independently contribute to an intrinsic antiviral defense against human cytomegalovirus infection. J. Virol. 2008, 82, 126–137. [Google Scholar] [CrossRef] [PubMed]
- Saffert, R.T.; Kalejta, R.F. Inactivating a cellular intrinsic immune defense mediated by Daxx is the mechanism through which the human cytomegalovirus pp71 protein stimulates viral immediate-early gene expression. J. Virol. 2006, 80, 3863–3871. [Google Scholar] [CrossRef] [PubMed]
- Saffert, R.T.; Kalejta, R.F. Human cytomegalovirus gene expression is silenced by Daxx-mediated intrinsic immune defense in model latent infections established in vitro. J. Virol. 2007, 81, 9109–9120. [Google Scholar] [CrossRef] [PubMed]
- Lukashchuk, V.; McFarlane, S.; Everett, R.D.; Preston, C.M. Human cytomegalovirus protein pp71 displaces the chromatin-associated factor ATRX from nuclear domain 10 at early stages of infection. J. Virol. 2008, 82, 12543–12554. [Google Scholar] [CrossRef] [PubMed]
- Torok, D.; Ching, R.W.; Bazett-Jones, D.P. PML nuclear bodies as sites of epigenetic regulation. Front. Biosci. 2009, 14, 1325–1336. [Google Scholar] [CrossRef] [PubMed]
- Hollenbach, A.D.; McPherson, C.J.; Mientjes, E.J.; Iyengar, R.; Grosveld, G. Daxx and histone deacetylase II associate with chromatin through an interaction with core histones and the chromatin-associated protein Dek. J. Cell Sci. 2002, 115, 3319–3330. [Google Scholar] [PubMed]
- Xue, Y.; Gibbons, R.; Yan, Z.; Yang, D.; McDowell, T.L.; Sechi, S.; Qin, J.; Zhou, S.; Higgs, D.; Wang, W. The ATRX syndrome protein forms a chromatin-remodeling complex with Daxx and localizes in promyelocytic leukemia nuclear bodies. Proc. Natl. Acad. Sci. USA 2003, 100, 10635–10640. [Google Scholar] [CrossRef]
- Groves, I.; Reeves, M.; Sinclair, J. Lytic infection of permissive cells with human cytomegalovirus is regulated by an intrinsic "pre-immediate early" repression of viral gene expression mediated by histone post-translational modification. J. Gen. Virol. 2009, 90, 2364–2374. [Google Scholar] [CrossRef] [PubMed]
- Tang, Q.; Maul, G.G. Mouse cytomegalovirus immediate-early protein 1 binds with host cell repressors to relieve suppressive effects on viral transcription and replication during lytic infection. J. Virol. 2003, 77, 1357–1367. [Google Scholar] [CrossRef] [PubMed]
- Randall, R.E.; Goodbourn, S. Interferons and viruses: an interplay between induction, signalling, antiviral responses and virus countermeasures. J. Gen. Virol. 2008, 89, 1–47. [Google Scholar] [CrossRef]
- Fensterl, V.; Sen, G.C. Interferons and viral infections. Biofactors 2009, 35, 14–20. [Google Scholar] [CrossRef] [PubMed]
- Bartee, E.; Mohamed, M.R.; McFadden, G. Tumor necrosis factor and interferon: cytokines in harmony. Curr. Opin. Microbiol. 2008, 11, 378–383. [Google Scholar] [CrossRef] [PubMed]
- Powers, C.; DeFilippis, V.; Malouli, D.; Fruh, K. Cytomegalovirus immune evasion. Curr. Top. Microbiol. Immunol. 2008, 325, 333–359. [Google Scholar] [PubMed]
- DeFilippis, V.R. Induction and evasion of the type I interferon response by cytomegaloviruses. Adv. Exp. Med. Biol. 2007, 598, 309–324. [Google Scholar] [PubMed]
- Schindler, C.; Plumlee, C. Inteferons pen the JAK-STAT pathway. Semin. Cell Dev. Biol. 2008, 19, 311–318. [Google Scholar] [CrossRef] [PubMed]
- Sehgal, P.B. Paradigm shifts in the cell biology of STAT signaling. Semin. Cell Dev. Biol. 2008, 19, 329–340. [Google Scholar] [CrossRef] [PubMed]
- Boyman, O.; Purton, J.F.; Surh, C.D.; Sprent, J. Cytokines and T-cell homeostasis. Curr. Opin. Immunol. 2007, 19, 320–326. [Google Scholar] [CrossRef] [PubMed]
- Browne, E.P.; Wing, B.; Coleman, D.; Shenk, T. Altered cellular mRNA levels in human cytomegalovirus-infected fibroblasts: viral block to the accumulation of antiviral mRNAs. J. Virol. 2001, 75, 12319–12330. [Google Scholar] [CrossRef] [PubMed]
- DeFilippis, V.R.; Robinson, B.; Keck, T.M.; Hansen, S.G.; Nelson, J.A.; Fruh, K.J. Interferon regulatory factor 3 is necessary for induction of antiviral genes during human cytomegalovirus infection. J. Virol. 2006, 80, 1032–1037. [Google Scholar] [CrossRef] [PubMed]
- Simmen, K.A.; Singh, J.; Luukkonen, B.G.; Lopper, M.; Bittner, A.; Miller, N.E.; Jackson, M.R.; Compton, T.; Fruh, K. Global modulation of cellular transcription by human cytomegalovirus is initiated by viral glycoprotein B. Proc. Natl. Acad. Sci. USA 2001, 98, 7140–7145. [Google Scholar] [CrossRef]
- Abate, D.A.; Watanabe, S.; Mocarski, E.S. Major human cytomegalovirus structural protein pp65 (ppUL83) prevents interferon response factor 3 activation in the interferon response. J. Virol. 2004, 78, 10995–11006. [Google Scholar] [CrossRef] [PubMed]
- Browne, E.P.; Shenk, T. Human cytomegalovirus UL83-coded pp65 virion protein inhibits antiviral gene expression in infected cells. Proc. Natl. Acad. Sci. USA 2003, 100, 11439–11444. [Google Scholar] [CrossRef]
- Miller, D.M.; Rahill, B.M.; Boss, J.M.; Lairmore, M.D.; Durbin, J.E.; Waldman, J.W.; Sedmak, D.D. Human cytomegalovirus inhibits major histocompatibility complex class II expression by disruption of the Jak/Stat pathway. J. Exp. Med. 1998, 187, 675–683. [Google Scholar] [CrossRef] [PubMed]
- Miller, D.M.; Zhang, Y.; Rahill, B.M.; Waldman, W.J.; Sedmak, D.D. Human cytomegalovirus inhibits IFN-alpha-stimulated antiviral and immunoregulatory responses by blocking multiple levels of IFN-alpha signal transduction. J. Immunol. 1999, 162, 6107–6113. [Google Scholar] [PubMed]
- Child, S.J.; Hakki, M.; De Niro, K.L.; Geballe, A.P. Evasion of cellular antiviral responses by human cytomegalovirus TRS1 and IRS1. J. Virol. 2004, 78, 197–205. [Google Scholar] [CrossRef] [PubMed]
- Child, S.J.; Jarrahian, S.; Harper, V.M.; Geballe, A.P. Complementation of vaccinia virus lacking the double-stranded RNA-binding protein gene E3L by human cytomegalovirus. J. Virol. 2002, 76, 4912–4918. [Google Scholar] [CrossRef] [PubMed]
- Chin, K.C.; Cresswell, P. Viperin (cig5), an IFN-inducible antiviral protein directly induced by human cytomegalovirus. Proc. Natl. Acad. Sci. USA 2001, 98, 15125–15130. [Google Scholar] [CrossRef]
- Jarvis, M.A.; Borton, J.A.; Keech, A.M.; Wong, J.; Britt, W.J.; Magun, B.E.; Nelson, J.A. Human cytomegalovirus attenuates interleukin-1beta and tumor necrosis factor alpha proinflammatory signaling by inhibition of NF-kappaB activation. J. Virol. 2006, 80, 5588–5598. [Google Scholar] [CrossRef] [PubMed]
- Montag, C.; Wagner, J.; Gruska, I.; Hagemeier, C. Human cytomegalovirus blocks tumor necrosis factor alpha- and interleukin-1beta-mediated NF-kappaB signaling. J. Virol. 2006, 80, 11686–11698. [Google Scholar] [CrossRef] [PubMed]
- Viola, A.; Luster, A.D. Chemokines and their receptors: drug targets in immunity and inflammation. Annu. Rev. Pharmacol. Toxicol. 2008, 48, 171–197. [Google Scholar] [CrossRef] [PubMed]
- Billstrom Schroeder, M.; Worthen, G.S. Viral regulation of RANTES expression during human cytomegalovirus infection of endothelial cells. J. Virol. 2001, 75, 3383–3390. [Google Scholar] [CrossRef] [PubMed]
- Randolph-Habecker, J.R.; Rahill, B.; Torok-Storb, B.; Vieira, J.; Kolattukudy, P.E.; Rovin, B.H.; Sedmak, D.D. The expression of the cytomegalovirus chemokine receptor homolog US28 sequesters biologically active CC chemokines and alters IL-8 production. Cytokine 2002, 19, 37–46. [Google Scholar] [CrossRef] [PubMed]
- Michelson, S.; Dal Monte, P.; Zipeto, D.; Bodaghi, B.; Laurent, L.; Oberlin, E.; Arenzana-Seisdedos, F.; Virelizier, J.L.; Landini, M.P. Modulation of RANTES production by human cytomegalovirus infection of fibroblasts. J. Virol. 1997, 71, 6495–6500. [Google Scholar] [PubMed]
- Chan, G.; Bivins-Smith, E.R.; Smith, M.S.; Smith, P.M.; Yurochko, A.D. Transcriptome analysis reveals human cytomegalovirus reprograms monocyte differentiation toward an M1 macrophage. J. Immunol. 2008, 181, 698–711. [Google Scholar] [PubMed]
- Wang, D.; Bresnahan, W.; Shenk, T. Human cytomegalovirus encodes a highly specific RANTES decoy receptor. Proc. Natl. Acad. Sci. USA 2004, 101, 16642–16647. [Google Scholar] [CrossRef]
- Bodaghi, B.; Jones, T.R.; Zipeto, D.; Vita, C.; Sun, L.; Laurent, L.; Arenzana-Seisdedos, F.; Virelizier, J.L.; Michelson, S. Chemokine sequestration by viral chemoreceptors as a novel viral escape strategy: withdrawal of chemokines from the environment of cytomegalovirus-infected cells. J. Exp. Med. 1998, 188, 855–866. [Google Scholar] [CrossRef] [PubMed]
- Hirsch, A.J.; Shenk, T. Human cytomegalovirus inhibits transcription of the CC chemokine MCP-1 gene. J. Virol. 1999, 73, 404–410. [Google Scholar] [PubMed]
- Krauss, S.; Kaps, J.; Czech, N.; Paulus, C.; Nevels, M. Physical requirements and functional consequences of complex formation between the cytomegalovirus IE1 protein and human STAT2 . J. Virol. 2009, 83 . [Google Scholar] [CrossRef]
- Hofmann, H.; Sindre, H.; Stamminger, T. Functional interaction between the pp71 protein of human cytomegalovirus and the PML-interacting protein human Daxx. J. Virol. 2002, 76, 5769–5783. [Google Scholar] [CrossRef] [PubMed]
- Hwang, J.; Kalejta, R. Proteasome-dependent, ubiquitin-independent degradation of Daxx by the viral pp71 protein of human cytomegalovirus-infected cells. Virology 2007, 367, 334–338. [Google Scholar] [CrossRef] [PubMed]
- Ishov, A.M.; Vladimirova, O.V.; Maul, G.G. Daxx-mediated accumulation of human cytomegalovirus tegument protein pp71 at ND10 facilitates initiation of viral infection at these nuclear domains. J. Virol. 2002, 76, 7705–7712. [Google Scholar] [CrossRef] [PubMed]
© 2009 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.