Optimizing Conservation Strategies for a Threatened Tree Species: In Situ Conservation of White Ash (Fraxinus americana L.) Genetic Diversity through Insecticide Treatment
Abstract
1. Introduction
2. Materials and Methods
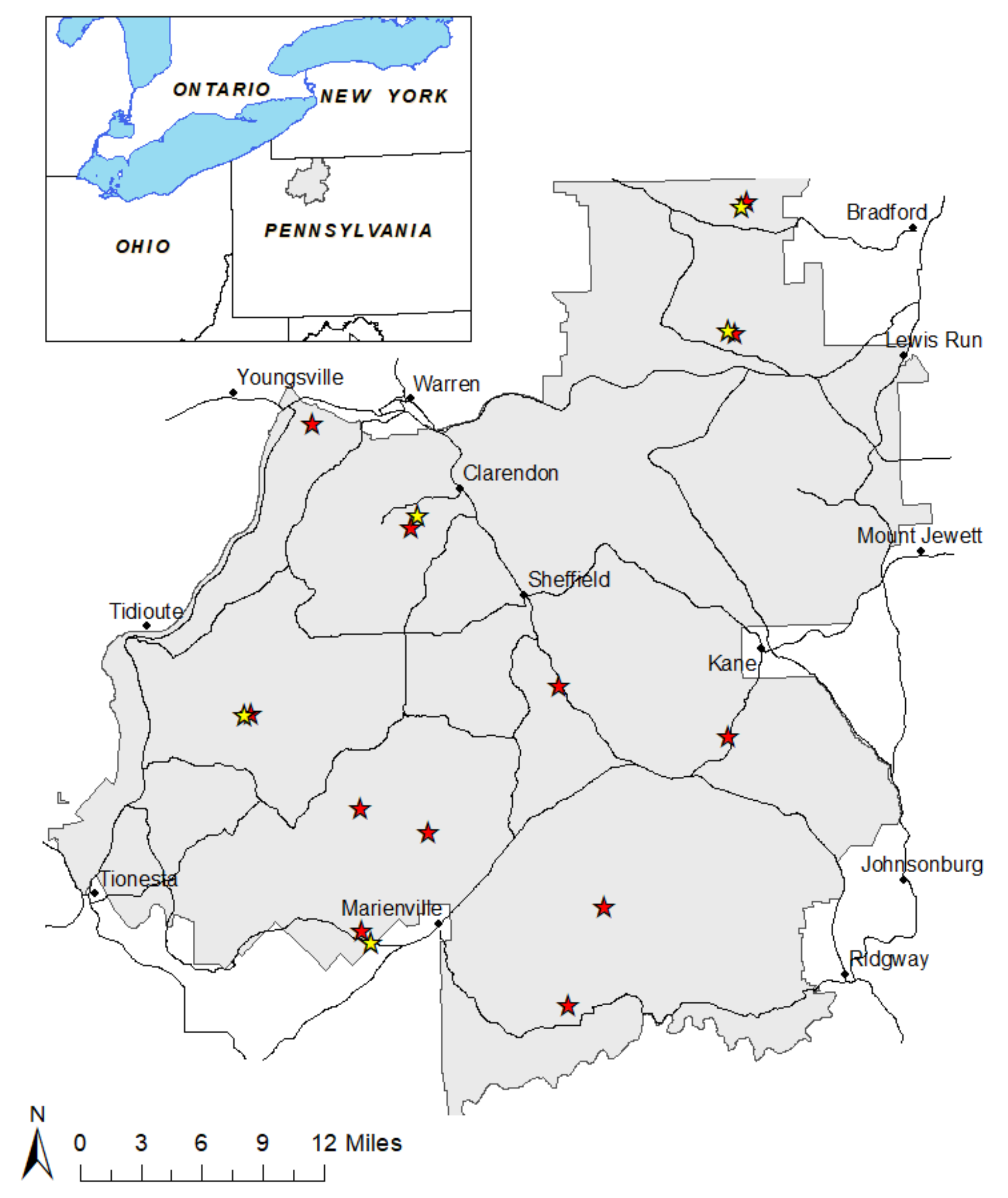
2.1. Site Selection
2.2. Sample Collection
2.3. Genetic Analysis
2.4. Data Analysis
2.4.1. Assessing the Success of In Situ Treatments
2.4.2. Optimizing In Situ Treatments
3. Results
3.1. White Ash Genetic Variability across the ANF
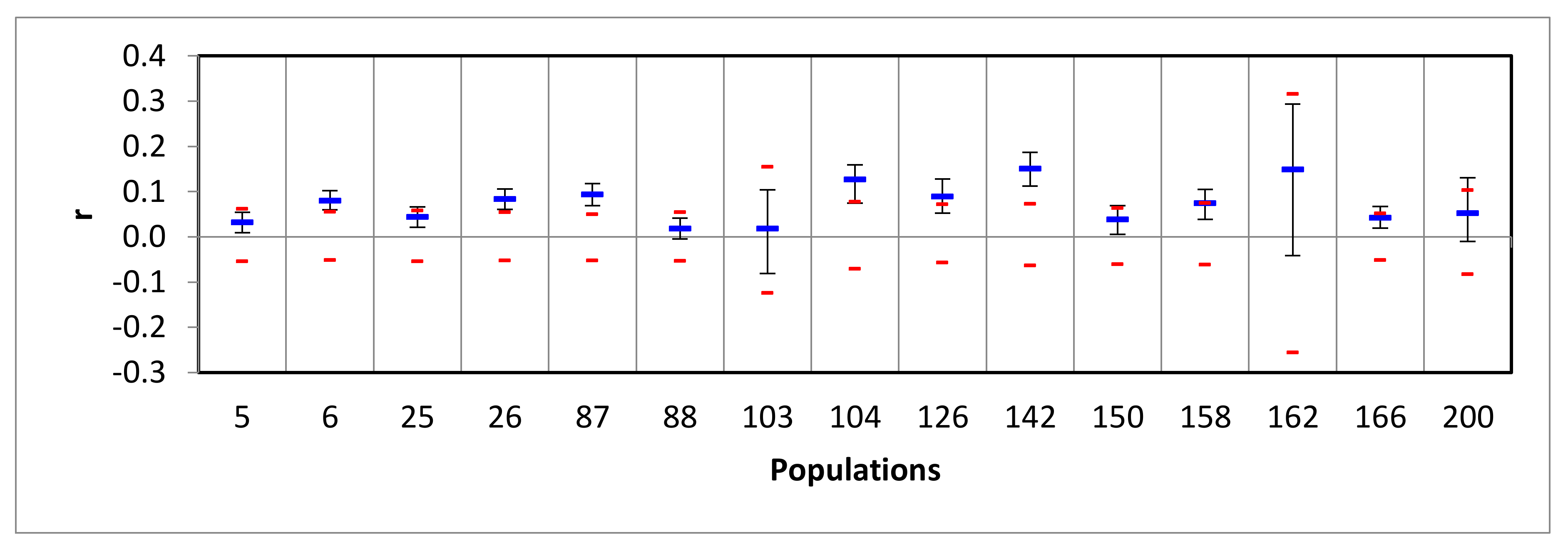
3.2. Assessing the Success of In Situ Efforts
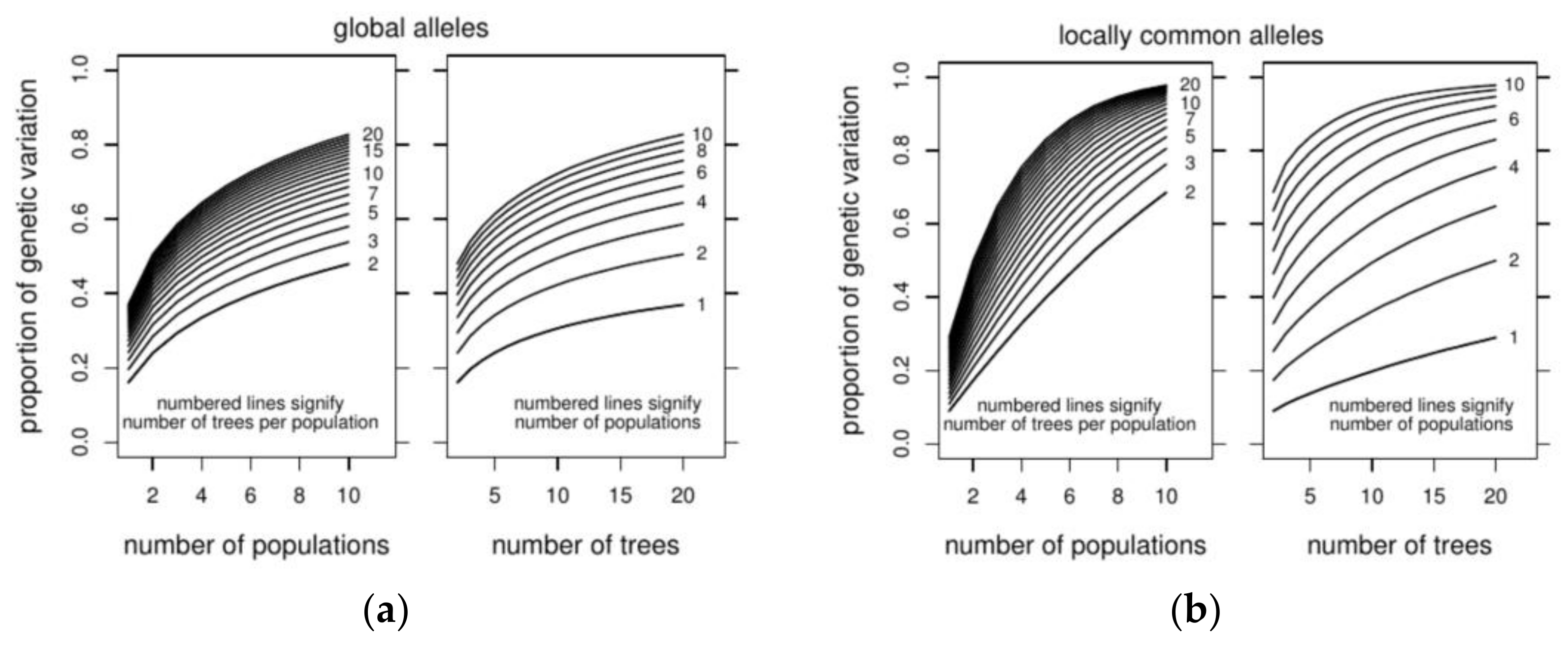
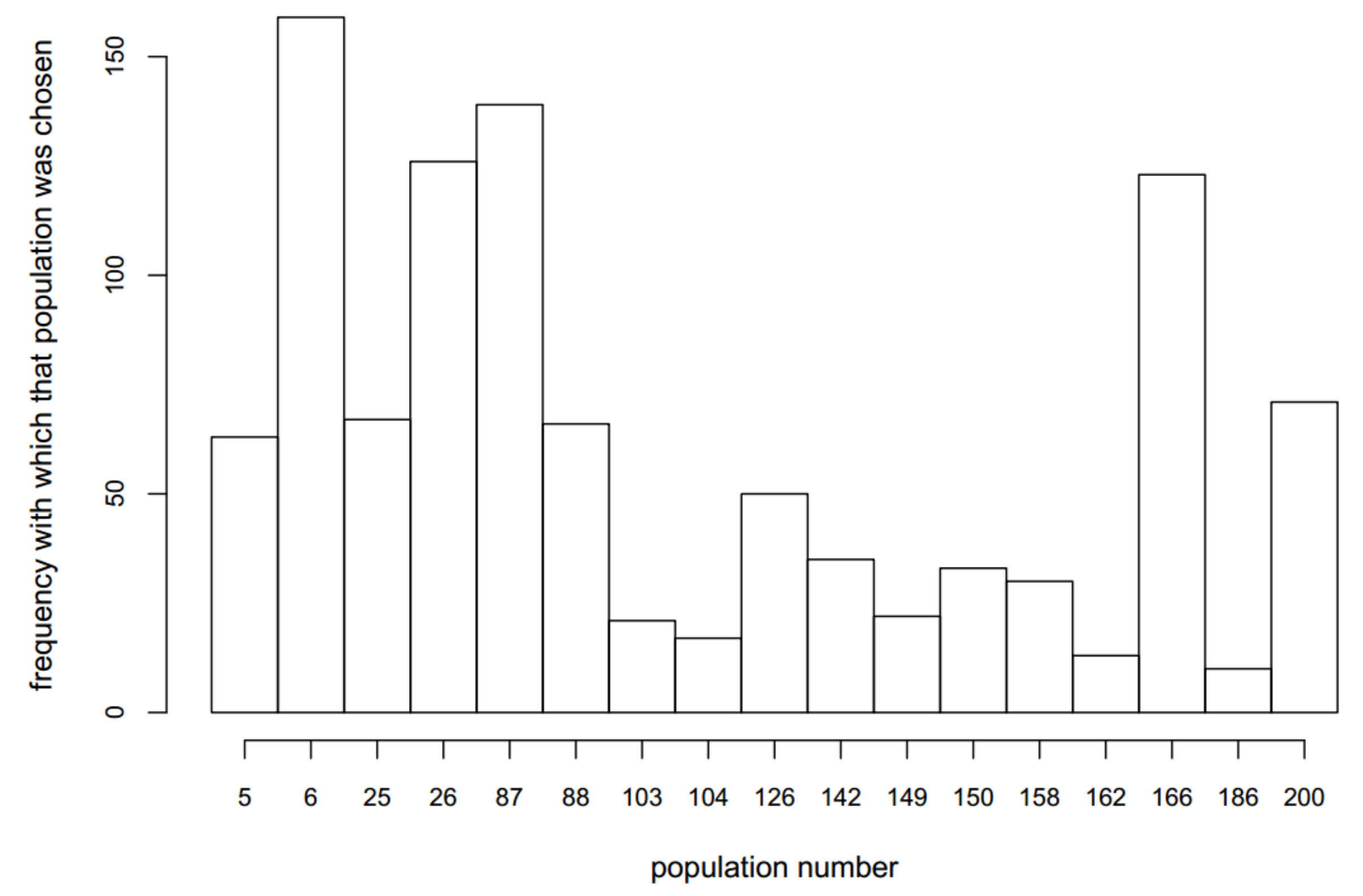
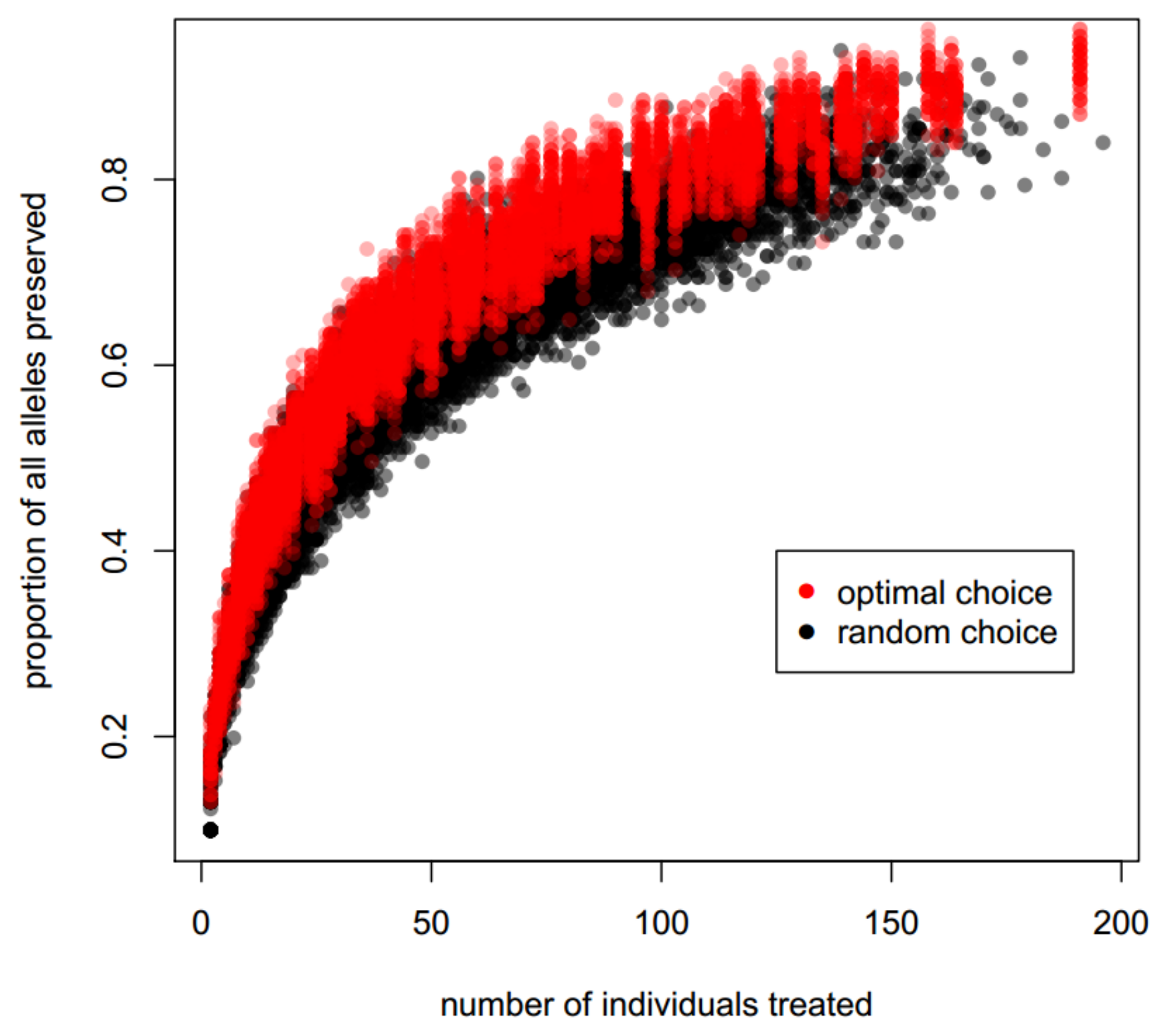
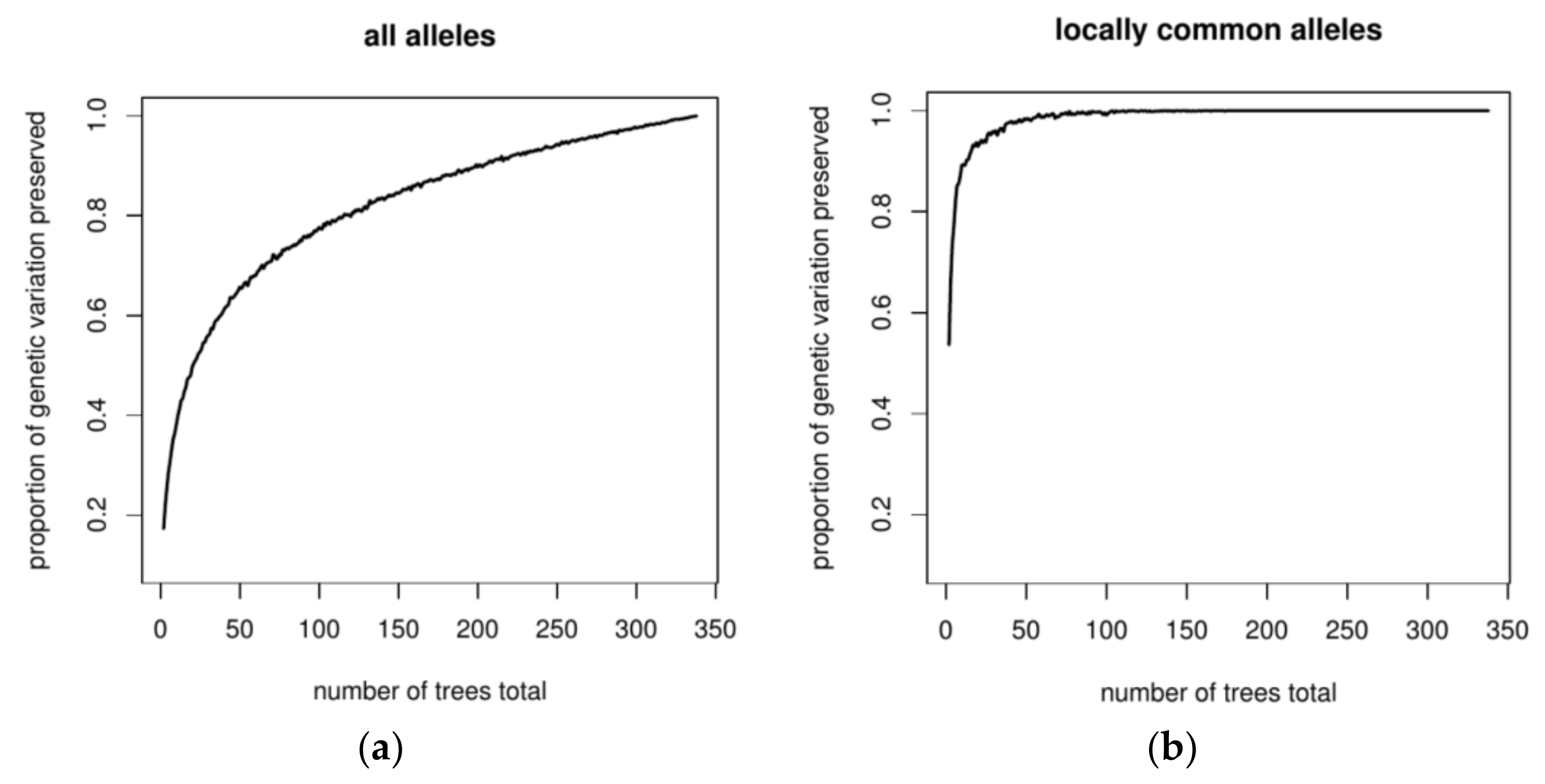
3.3. Optimizing Insecticide Treatment
4. Discussion
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- McKinney, M.L. Measuring floristic homogenization by non-native plants in North America. Glob. Ecol. Biogeogr. 2004, 13, 47–53. [Google Scholar] [CrossRef]
- Corvalan, C.; Hales, S.; McMichael, A. Ecosystems and human well-being: Biodiversity synthesis. In Millenium Ecosystem Assessment; World Resources Institute: Washington, DC, USA, 2005. [Google Scholar]
- Lovett, G.M.; Canham, C.D.; Arthur, M.A.; Weathers, K.C.; Fitzhugh, R.D. Forest ecosystem responses to exotic pests and pathogens in eastern North America. Bioscience 2006, 56, 395–405. [Google Scholar] [CrossRef]
- Flower, C.E.; Gonzalez-Meler, M.A. Responses of temperate forest productivity to insect and pathogen disturbances. Annu. Rev. Plant Biol. 2015, 66, 547–569. [Google Scholar] [CrossRef] [PubMed]
- Liebhold, A.M.; Brockerhoff, E.G.; Kalisz, S.; Nuñez, M.A.; Wardle, D.A.; Wingfield, M.J. Biological invasions in forest ecosystems. Biol. Invasions 2017, 19, 3437–3458. [Google Scholar] [CrossRef]
- Schaberg, P.G.; DeHayes, D.H.; Hawley, G.J.; Nijensohn, S.E. Anthropogenic alterations of genetic diversity within tree populations: Implications for forest ecosystem resilience. For. Ecol. Manag. 2008, 256, 855–862. [Google Scholar] [CrossRef]
- Hughes, A.R.; Inouye, B.D.; Johnson, M.T.J.; Underwood, N.; Vellend, M. Ecological consequences of genetic diversity. Ecol. Lett. 2008, 11, 609–623. [Google Scholar] [CrossRef] [PubMed]
- Hughes, A.R.; Stachowicz, J.J. Genetic diversity enhances the resistance of a seagrass ecosystem to disturbance. Proc. Natl. Acad. Sci. USA 2004, 101, 8998–9002. [Google Scholar] [CrossRef] [PubMed]
- Clark, J.S. Individuals and the variation needed for high species diversity in forest trees. Science 2010, 327, 1129–1132. [Google Scholar] [CrossRef] [PubMed]
- Laikre, L.; Allendorf, F.W.; Aroner, L.C.; Baker, C.S.; Gregovich, D.P.; Hansen, M.M.; Jackson, J.A.; Kendall, K.C.; Mckelvey, K.; Neel, M.C.; et al. Neglect of genetic diversity in implementation of the convention on biological diversity. Conserv. Biol. 2009, 24, 86–88. [Google Scholar] [CrossRef] [PubMed]
- Schoen, D.J.; Brown, A.H. The Conservation of Wild Plant Species in Seed Banks: Attention to both taxonomic coverage and population biology will improve the role of seed banks as conservation tools. Bioscience 2001, 51, 960–966. [Google Scholar] [CrossRef]
- Fant, J.; Havens, K.; Kramer, A.; Walsh, S.; Callicrate, T.; Lacy, R.; Maunder, M.; Meyer, A.; Smith, P. What to do when we can’t bank on seeds: What botanic gardens can learn from the zoo community about conserving plants in living collections. Am. J. Bot. 2016, 103, 1541–1543. [Google Scholar] [CrossRef] [PubMed]
- Basey, A.C.; Fant, J.B.; Kramer, A.T. Producing native plant materials for restoration: 10 rules to collect and maintain genetic diversity. Nativ. Plants J. 2015, 16, 37–53. [Google Scholar] [CrossRef]
- Ricketts, M.P.; Flower, C.E.; Knight, K.S.; Gonzalez-Meler, M.A. Evidence of ash tree (Fraxinus spp.) association with soil bacterial community structure and function. Forests 2018, 9, 187. [Google Scholar] [CrossRef]
- Slesak, R.A.; Lenhart, C.F.; Brooks, K.N.; D’Amato, A.W.; Palik, B.J. Water table response to harvesting and simulated emerald ash borer mortality in black ash wetlands in Minnesota, USA. Can. J. For. Res. 2014, 44, 961–968. [Google Scholar] [CrossRef]
- Fady, B.; Aravanopoulos, F.A.; Alizoti, P.; Mátyás, C.; Von Wühlisch, G.; Westergren, M.; Belletti, P.; Cvjetkovic, B.; Ducci, F.; Huber, G.; et al. Evolution-based approach needed for the conservation and silviculture of peripheral forest tree populations. For. Ecol. Manag. 2016, 375, 66–75. [Google Scholar] [CrossRef]
- Herms, D.A.; McCullough, D.G. Emerald ash borer invasion of North America: History, biology, ecology, impacts, and management. Annu. Rev. Entomol. 2014, 59, 13–30. [Google Scholar] [CrossRef] [PubMed]
- MacFarlane, D.W.; Meyer, S.P. Characteristics and distribution of potential ash tree hosts for emerald ash borer. For. Ecol. Manag. 2005, 213, 15–24. [Google Scholar] [CrossRef]
- Flower, C.E.; Knight, K.S.; Gonzalez-Meler, M.A. Impacts of the emerald ash borer (Agrilus planipennis Fairmaire) induced ash (Fraxinus spp.) mortality on forest carbon cycling and successional dynamics in the eastern United States. Biol. Invasions 2013, 15, 931–944. [Google Scholar] [CrossRef]
- Flower, C.E.; Knight, K.S.; Rebbeck, J.; Gonzalez-Meler, M.A. The relationship between the emerald ash borer (Agrilus planipennis) and ash (Fraxinus spp.) tree decline: Using visual canopy condition assessments and leaf isotope measurements to assess pest damage. For. Ecol. Manag. 2013, 303, 143–147. [Google Scholar] [CrossRef]
- Flower, C.E.; Lynch, D.J.; Knight, K.S.; Gonzalez-Meler, M.A. Biotic and abiotic drivers of sap flux in mature green ash trees (Fraxinus pennsylvanica) experiencing varying levels of emerald ash borer (Agrilus planipennis) infestation. Forests 2018, in press. [Google Scholar]
- Knight, K.S.; Brown, J.P.; Long, R.P. Factors affecting the survival of ash trees (Fraxinus spp.) infested by emerald ash borer (Agrilus planipennis). Biol. Invasions 2013, 15, 371–383. [Google Scholar] [CrossRef]
- Klooster, W.S.; Herms, D.A.; Knight, K.S.; Herms, C.P.; McCullough, D.G.; Smith, A.; Gandhi, K.J.K.; Cardina, J. Ash (Fraxinus spp.) mortality, regeneration, and seed bank dynamics in mixed hardwood forests following invasion by emerald ash borer (Agrilus planipennis). Biol. Invasions 2013, 16, 859–873. [Google Scholar] [CrossRef]
- McCullough, D.G.; Tanis, S.R.; Robinett, M.; Limback, C.; Poland, T.M. White ash—Is EAB always a death sentence? In Proceedings of the 2014 Emerald Ash Borer National Research and Technology Development Meeting, Wooster, OH, USA, 15–16 October 2014; Buck, J., Parra, G., Lance, D., Reardon, R., Binion, D., Eds.; U.S. Department of Agriculture, Forest Service: Washington, DC, USA, 2015; pp. 10–11. [Google Scholar]
- The IUCN Red List of Threatened Species. Available online: http://www.iucnredlist.org/ (accessed on 11 April 2018).
- Cappaert, D.; Mccullough, D.G.; Poland, T.M.; Siegert, N.W. Emerald ash borer in North America: A research and regulatory challenge. Am. Entomol. 2005, 51, 152–165. [Google Scholar] [CrossRef]
- O’Brien, E.M. Conserving Ash (Fraxinus) Populations and Genetic Variation in Forests Invaded by Emerald Ash Borer Using Large-Scale Insecticide Applications; Ohio State University: Columbus, OH, USA, 2017. [Google Scholar]
- Flower, C.E.; Aubihl, E.; Fant, J.; Forry, S.; Hille, A.; Knight, K.S.; Oldland, W.K.; Royo, A.A.; Richard, M. In-situ Genetic Conservation of White Ash (Fraxinus americana) at the Allegheny National Forest. In Proceedings of the Workshop on Gene Conservation of Tree Species-Banking on the Future, Chicago, IL, USA, 16–19 May 2016; Sniezko, R.A., Man, G., Hipkins, V., Woeste, K., Gwaze, D., Kliejunas, J.T., McTeague, B.A., Eds.; U.S. Department of Agriculture, Forest Service: Washington, DC, USA; Pacific Northwest Research Station: Portland, OR, USA, 2016; pp. 165–169. [Google Scholar]
- Herms, D.A.; Mccullough, D.G.; Smitley, D.R.; Sadof, C.S.; Cranshaw, W. Insecticide Options for Protecting Ash Trees from Emerald Ash Borer, 2nd ed.; North Central IPM Center Bulletin: Champaign, IL, USA, 2014; 16p. [Google Scholar]
- Flower, C.E.; Dalton, J.E.; Knight, K.S.; Brikha, M.; Gonzalez-Meler, M.A. To treat or not to treat: Diminishing effectiveness of emamectin benzoate tree injections in ash trees heavily infested by emerald ash borer. Urban For. Urban Green. 2015, 14. [Google Scholar] [CrossRef]
- Gregorious, H.-R. The probability of losing an allele when diploid genotypes are sampled. Bioinformatics 1980, 36, 643–652. [Google Scholar] [CrossRef]
- Bashalkhanov, S.; Pandley, M.; Rahjora, O.P. A simple method for estimating genetic diversity in large populations from finite sample sizes. BMC Genet. 2009, 10, 84. [Google Scholar] [CrossRef] [PubMed]
- Brown, A.; Marshall, D. A basic sampling strategy: Theory and practice. In Collecting Plant Genetic Diversity: Technical Guidelines; Guarino, L., Rao, V.R., Reid, R., Eds.; CAB International: Wallingford, UK, 1995; pp. 75–91. [Google Scholar]
- Richards, C.M.; Antolin, M.F.; Reilley, A.; Poole, J.; Walters, C. Capturing genetic diversity of wild populations for ex situ conservation: Texas wild rice (Zizania texana) as a model. Genet. Resour. Crop Evol. 2007, 54, 837–848. [Google Scholar] [CrossRef]
- Hoban, S.; Strand, A. Ex situ seed collections will benefit from considering spatial sampling design and species’ reproductive biology. Biol. Conserv. 2015, 187, 182–191. [Google Scholar] [CrossRef]
- Griffith, M.P.; Calonje, M.; Meerow, A.W.; Francisco-Ortega, J.; Knowles, L.; Aguilar, R.; Tut, F.; Sánchez, V.; Meyer, A.; Noblick, L.R.; et al. Will the same ex situ protocols give similar results for closely related species? Biodivers. Conserv. 2017, 26, 2951–2966. [Google Scholar] [CrossRef]
- McGlaughlin, M.E.; Riley, L.; Brandsrud, M.; Arcibal, E.; Helenurm, M.K.; Helenurm, K. How much is enough? Minimum sampling intensity required to capture extant genetic diversity in ex situ seed collections: Examples from the endangered plant Sibara filifolia (Brassicaceae). Conserv. Genet. 2015, 16, 253–266. [Google Scholar] [CrossRef]
- Kashimshetty, Y.; Pelikan, S.; Rogstad, S.H. Effective seed harvesting strategies for the ex situ genetic diversity conservation of rare tropical tree populations. Biodivers. Conserv. 2017, 26, 1311–1331. [Google Scholar] [CrossRef]
- Schoen, D.J.; Brown, A.H.D. Intraspecific variation in population gene diversity and effective population size correlates with the mating system in plants. Proc. Natl. Acad. Sci. USA 1991, 88, 4494–4497. [Google Scholar] [CrossRef] [PubMed]
- Doyle, J.J.; Doyle, J.L. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem. Bull. 1987, 19, 11–15. [Google Scholar]
- Lefort, F.; Brachet, S.; Frascaria-Lacoste, N.; Edwards, K.J.; Douglas, G. Identification and characterization of microsatellite loci in ash (Fraxinus excelsior L.) and their conservation in the olive family (Oleaceae). Mol. Ecol. 1999, 8, 1088–1089. [Google Scholar] [CrossRef]
- Harbourne, M.E.; Douglas, C.G.; Waldren, S.; Hodkinson, T.R. Characterization and primer development for amplification of chloroplast microsatellite regions of Fraxinus excelsior. J. Plant Res. 2005, 118, 339–341. [Google Scholar] [CrossRef] [PubMed]
- Sutherland, B.G.; Belaj, A.; Nier, S.; Cottrell, J.E.; P Vaughan, S.; Hubert, J.; Russell, K. Molecular biodiversity and population structure in common ash (Fraxinus excelsior L.) in Britain: Implications for conservation. Mol. Ecol. 2010, 19, 2196–2211. [Google Scholar] [CrossRef] [PubMed]
- Beatty, G.E.; Brown, J.A.; Cassidy, E.M.; Finlay, C.M.V.; McKendrick, L.; Montgomery, W.I.; Reid, N.; Tosh, D.G.; Provan, J. Lack of genetic structure and evidence for long-distance dispersal in ash (Fraxinus excelsior) populations under threat from an emergent fungal pathogen: Implications for restorative planting. Tree Genet. Genomes 2015, 11, 1–13. [Google Scholar] [CrossRef]
- Brachet, S.; Jubier, M.; Richard, M.; Jung-Muller, B.; Frascaria-Lacoste, N. Rapid identification of microsatellite loci using 5’ anchored PCR in the common ash Fraxinus excelsior. Microb. Ecol. 1999, 8, 160–163. [Google Scholar]
- Noakes, A.G.; Best, T.; Staton, M.E.; Koch, J.; Romero-Severson, J. Cross amplification of 15 EST-SSR markers in the genus Fraxinus. Conserv. Genet. Resour. 2014, 6, 969–970. [Google Scholar] [CrossRef]
- Schuelke, M. An economic method for the fluorescent labeling of PCR fragments. Nat. Biotechnol. 2000, 18, 233–234. [Google Scholar] [CrossRef] [PubMed]
- Peakall, R.; Smouse, P. GenAlEx 6: Genetic analysis in Excel. Population genetic software for teaching and research. Mol. Ecol. Notes 2006, 6, 288–295. [Google Scholar] [CrossRef]
- Weir, B.; Cockerham, C. Estimating F-statistics for the analysis of population structure. Evolution 1984, 38, 1358–1370. [Google Scholar] [PubMed]
- Queller, D.C.; Goodnight, K.F. Estimating relatedness using genetic markers. Evolution 1989, 43, 258. [Google Scholar] [CrossRef] [PubMed]
- Bates, D.; Maechler, M.; Bolker, B.; Walker, S. Fitting linear mixed-effects models using lme4. J. Stat. Softw. 2015, 67, 1–48. [Google Scholar] [CrossRef]
- Pritchard, J.; Stephens, M.; Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar] [PubMed]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef] [PubMed]
- Earl, D.A.; von Holdt, B.M. STRUCTURE HARVESTER: A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 2012, 4, 359–361. [Google Scholar] [CrossRef]
- Jombart, T. Adegenet: A R package for the multivariate analysis of genetic markers. Bioinformatics 2008, 24, 1403–1405. [Google Scholar] [CrossRef] [PubMed]
- Heuertz, M.; Hausman, J.F.; Tsvetkov, I.; Frascaria-Lacoste, N.; Vekemans, X. Assessment of genetic structure within and among Bulgarian populations of the common ash (Fraxinus excelsior L.). Mol. Ecol. 2001, 10, 1615–1623. [Google Scholar] [CrossRef] [PubMed]
- Heuertz, M.; Hausman, J.F.; Hardy, O.; Vendramin, G.G.; Frascaria-Lacoste, N.; Vekemans, X. Nuclear microsatellites reveal contrasting patterns of genetic structure between western and southeastern European populations of the common ash (Fraxinus excelsior L.). Evolution 2004, 58, 976–988. [Google Scholar] [PubMed]
- Craft, K.J.; Ashley, M.V. Landscape genetic structure of bur oak (Quercus macrocarpa) savannas in Illinois. For. Ecol. Manag. 2007, 239, 13–20. [Google Scholar] [CrossRef]
- Streiff, R.; Ducousso, A.; Lexer, C.; Steinkellner, H.; Gloessl, J.; Kremer, A. Pollen dispersal inferred from paternity analysis in a mixed oak stand of Quercus robur L. and Q. petraea (Matt.) Liebl. Mol. Ecol. 1999, 8, 831–841. [Google Scholar] [CrossRef]
- Nadeau, S.; Meirmans, P.G.; Aitken, S.N.; Ritland, K.; Isabel, N. The challenge of separating signatures of local adaptation from those of isolation by distance and colonization history: The case of two white pines. Ecol. Evol. 2016, 6, 8649–8664. [Google Scholar] [CrossRef] [PubMed]
- Bower, A.D.; Aitken, S.N. Ecological genetics and seed transfer guidelines for Pinus albicaulis (Pinaceae). Am. J. Bot. 2008, 95, 66–76. [Google Scholar] [CrossRef] [PubMed]
- Jump, A.S.; Penuelas, J. Genetic effects of chronic habitat fragmentation in a wind-pollinated tree. Proc. Natl. Acad. Sci. USA 2006, 103, 8096–8100. [Google Scholar] [CrossRef] [PubMed]
- Brown, A.H.D. Isozymes, plant population genetic structure and genetic conservation. Theor. Appl. Genet. 1978, 52, 145–157. [Google Scholar] [CrossRef] [PubMed]
- Hoban, S.; Schlarbaum, S. Optimal sampling of seeds from plant populations for ex-situ conservation of genetic biodiversity, considering realistic population structure. Biol. Conserv. 2014, 177, 90–99. [Google Scholar] [CrossRef]
- Guerrant, E.O.; Fiedler, P.L.; Havens, K.; Maunder, M. Revised genetic sampling guidelines for conservation collections of rare and endangered plants. In Ex Situ Plant Conservation: Supporting Species Survival in the Wild; Island Press: Washington, DC, USA, 2004; pp. 419–448. [Google Scholar]
- Vinceti, B.; Loo, J.; Gaisberger, H.; van Zonneveld, M.J.; Schueler, S.; Konrad, H.; Kadu, C.A.C.; Geburek, T. Conservation priorities for prunus Africana defined with the aid of spatial analysis of genetic data and climatic variables. PLoS ONE 2013, 8, e59987. [Google Scholar] [CrossRef] [PubMed]
- Schlottfeldt, S.; Walter, M.E.M.T.; de Carvalho, A.C.P.L.F.; Soares, T.N.; Telles, M.P.C.; Loyola, R.D.; Diniz-Filho, J.A.F. Multi-objective optimization for plant germplasm collection conservation of genetic resources based on molecular variability. Tree Genet. Genomes 2015, 11, 16. [Google Scholar] [CrossRef][Green Version]
- Jump, A.S.; Hunt, J.M.; Martínez-Izquierdo, J.; Peñuelas, J. Natural selection and climate change: Temperature-Linked spatial and temporal trends in gene frequency in Fagus sylvatica. Mol. Ecol. 2006, 15, 3469–3480. [Google Scholar] [CrossRef] [PubMed]
- Hoban, S.; Kelley, J.L.; Lotterhos, K.E.; Antolin, M.F.; Bradburd, G.; Lowry, D.B.; Poss, M.L.; Reed, L.K.; Storfer, A.; Whitlock, M.C. Finding the genomic basis of local adaptation: pitfalls, practical solutions, and future directions. Am. Nat. 2016, 188, 379–397. [Google Scholar] [CrossRef] [PubMed]
- Way, M.J. Collecting seed from non-domesticated plants for long-term conservation. In Seed Conservation: Turning Science into Practice; Smith, R., Dickie, J., Linington, S., Pritchard, H., Probert, R., Eds.; Royal Botanic Gardens Kew: London, UK, 2003; pp. 165–201. [Google Scholar]
- Vanderwel, M.C.; Coomes, D.A.; Purves, D.W. Quantifying variation in forest disturbance, and its effects on aboveground biomass dynamics, across the eastern United States. Glob. Chang. Biol. 2013, 19, 1504–1517. [Google Scholar] [CrossRef] [PubMed]
- Flanagan, S.P.; Forester, B.R.; Latch, E.K.; Aitken, S.N.; Hoban, S. Guidelines for planning genomic assessment and monitoring of locally adaptive variation to inform species conservation. Evol. Appl. 2017, 1–18. [Google Scholar] [CrossRef]
- Margulies, E.; Bauer, L.; Ibáñez, I. Buying time: Preliminary assessment of biocontrol in the recovery of native forest vegetation in the aftermath of the invasive emerald ash borer. Forests 2017, 8, 369. [Google Scholar] [CrossRef]
- Schoettle, A.W.; Klutsch, J.G.; Sniezko, R.A. Integrating regeneration, genetic resistance, and timing of intervention for the long-term sustainability of ecosystems challenged by non-native Pests—A novel proactive approach. In Proceedings of the 4th International Workshop on the Genetics of Host-Parasite Interactions in Forestry: Disease and Insect Resistance in Forest Trees, Eugene, OR, USA, 31 July–5 August 2011; Sniezko, R.A., Yanchuk, A.D., Kliejunas, J.T., Palmieri, K.M., Alexander, J.M., Frankel, S.J., Eds.; Pacific Southwest Research Station: Redding, CA, USA; Forest Service, U.S. Department of Agriculture: Washington, DC, USA, 2012; pp. 112–123. [Google Scholar]
| PopID | Insecticide Treated | Topography | n | Na | Ne | He | P | Fis |
|---|---|---|---|---|---|---|---|---|
| 104 | N | Lowland | 16 | 4.88 | 2.89 | 0.60 | 0 | 0.00 |
| 150 | N | Lowland | 23 | 5.88 | 3.13 | 0.64 | 0 | 0.13 |
| 162 | N | Lowland | 3 | 2.63 | 2.11 | 0.48 | 0 | −0.09 |
| 186 | N | Lowland | 2 | 1.63 | 1.58 | 0.23 | 0 | −0.64 |
| 200 | N | Lowland | 23 | 5.00 | 2.88 | 0.61 | 2 | 0.04 |
| 103 | N | Upland | 7 | 4.13 | 2.83 | 0.60 | 0 | 0.16 |
| 149 | N | Upland | 1 | 1.38 | 1.38 | 0.25 | 0 | −1.00 |
| 6 | Y | Lowland | 30 | 8.25 | 3.40 | 0.62 | 7 | 0.08 |
| 26 | Y | Lowland | 29 | 7.50 | 3.14 | 0.61 | 7 | 0.07 |
| 88 | Y | Lowland | 30 | 7.50 | 3.32 | 0.65 | 0 | 0.07 |
| 126 | Y | Lowland | 24 | 6.13 | 2.89 | 0.61 | 1 | 0.11 |
| 142 | Y | Lowland | 22 | 5.38 | 2.76 | 0.59 | 1 | 0.09 |
| 158 | Y | Lowland | 23 | 5.75 | 2.99 | 0.63 | 0 | 0.02 |
| 166 | Y | Lowland | 30 | 7.50 | 3.45 | 0.64 | 2 | 0.08 |
| 5 | Y | Upland | 29 | 7.13 | 3.26 | 0.65 | 1 | 0.08 |
| 25 | Y | Upland | 30 | 6.63 | 3.10 | 0.63 | 2 | 0.08 |
| 87 | Y | Upland | 30 | 7.25 | 3.58 | 0.64 | 3 | 0.17 |
| Management Option | No. of Populations | All | Very Common | Common | Low Freq | Rare | Locally Common |
|---|---|---|---|---|---|---|---|
| Option 1 | 4 | 64.4% | ~100% | 99.4% | 84.8% | 31.4% | 75.5% |
| Option 2 | 10 | 66.6% | ~100% | 99.9% | 91.1% | 31.4% | 88.4% |
| Difference | 2.2% | <0.1% | 0.5% | 6.3% | <0.1% | 12.9% |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Flower, C.E.; Fant, J.B.; Hoban, S.; Knight, K.S.; Steger, L.; Aubihl, E.; Gonzalez-Meler, M.A.; Forry, S.; Hille, A.; Royo, A.A. Optimizing Conservation Strategies for a Threatened Tree Species: In Situ Conservation of White Ash (Fraxinus americana L.) Genetic Diversity through Insecticide Treatment. Forests 2018, 9, 202. https://doi.org/10.3390/f9040202
Flower CE, Fant JB, Hoban S, Knight KS, Steger L, Aubihl E, Gonzalez-Meler MA, Forry S, Hille A, Royo AA. Optimizing Conservation Strategies for a Threatened Tree Species: In Situ Conservation of White Ash (Fraxinus americana L.) Genetic Diversity through Insecticide Treatment. Forests. 2018; 9(4):202. https://doi.org/10.3390/f9040202
Chicago/Turabian StyleFlower, Charles E., Jeremie B. Fant, Sean Hoban, Kathleen S. Knight, Laura Steger, Elijah Aubihl, Miquel A. Gonzalez-Meler, Stephen Forry, Andrea Hille, and Alejandro A. Royo. 2018. "Optimizing Conservation Strategies for a Threatened Tree Species: In Situ Conservation of White Ash (Fraxinus americana L.) Genetic Diversity through Insecticide Treatment" Forests 9, no. 4: 202. https://doi.org/10.3390/f9040202
APA StyleFlower, C. E., Fant, J. B., Hoban, S., Knight, K. S., Steger, L., Aubihl, E., Gonzalez-Meler, M. A., Forry, S., Hille, A., & Royo, A. A. (2018). Optimizing Conservation Strategies for a Threatened Tree Species: In Situ Conservation of White Ash (Fraxinus americana L.) Genetic Diversity through Insecticide Treatment. Forests, 9(4), 202. https://doi.org/10.3390/f9040202





