Genome-Wide Identification and Expression Profiling of AP2/ERF Transcription Factor Genes in Prunus armeniaca L.
Abstract
1. Introduction
2. Materials and Methods
2.1. Genome-Wide Identification of AP2/ERF Transcription Factors in Apricot
2.2. Phylogenetic and Evolutionary Analysis
2.3. Structural Characterization of AP2/ERF Genes
2.4. Chromosomal Localization and Synteny Analysis of AP2/ERF Gene Family
2.5. Analysis of Regulatory Elements and Functional Annotation
2.6. Transcriptional Profiling and Experimental Validation
2.7. Oil Content Quantification
2.8. Statistical Analysis
- (1)
- Differences in gene expression (RT-qPCR) and oil content across sample groups were assessed using one-way ANOVA.
- (2)
- Pearson correlation analyses were performed to evaluate the concordance between RNA-seq and RT-qPCR data, and the relationships between gene expression patterns and oil accumulation profiles.
2.9. Targets Prediction of PaWRI1
3. Results
3.1. Identification of AP2/ERF Family Members in Apricot
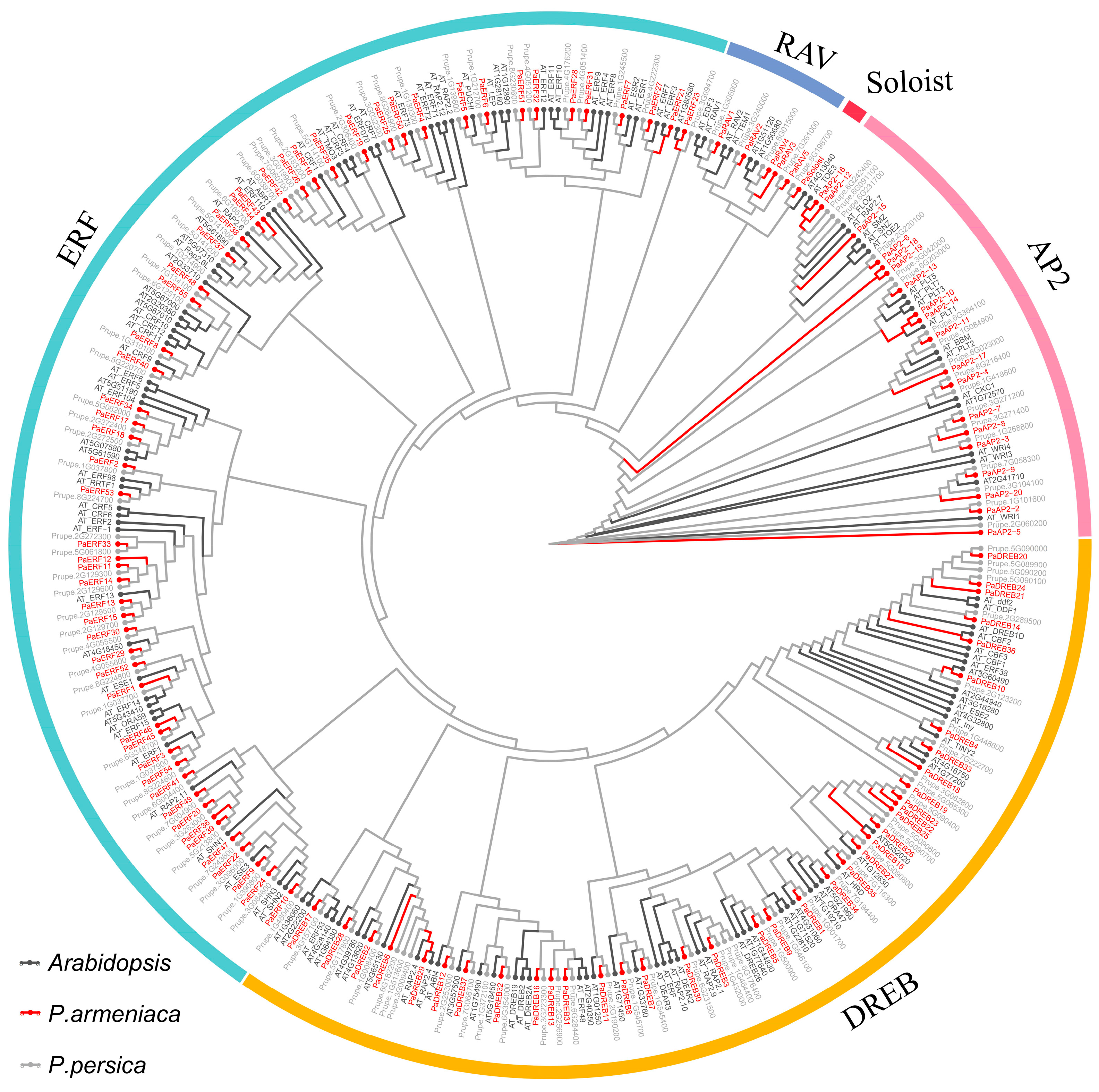
3.2. Phylogenetic Analysis of the PaAP2/ERF Family
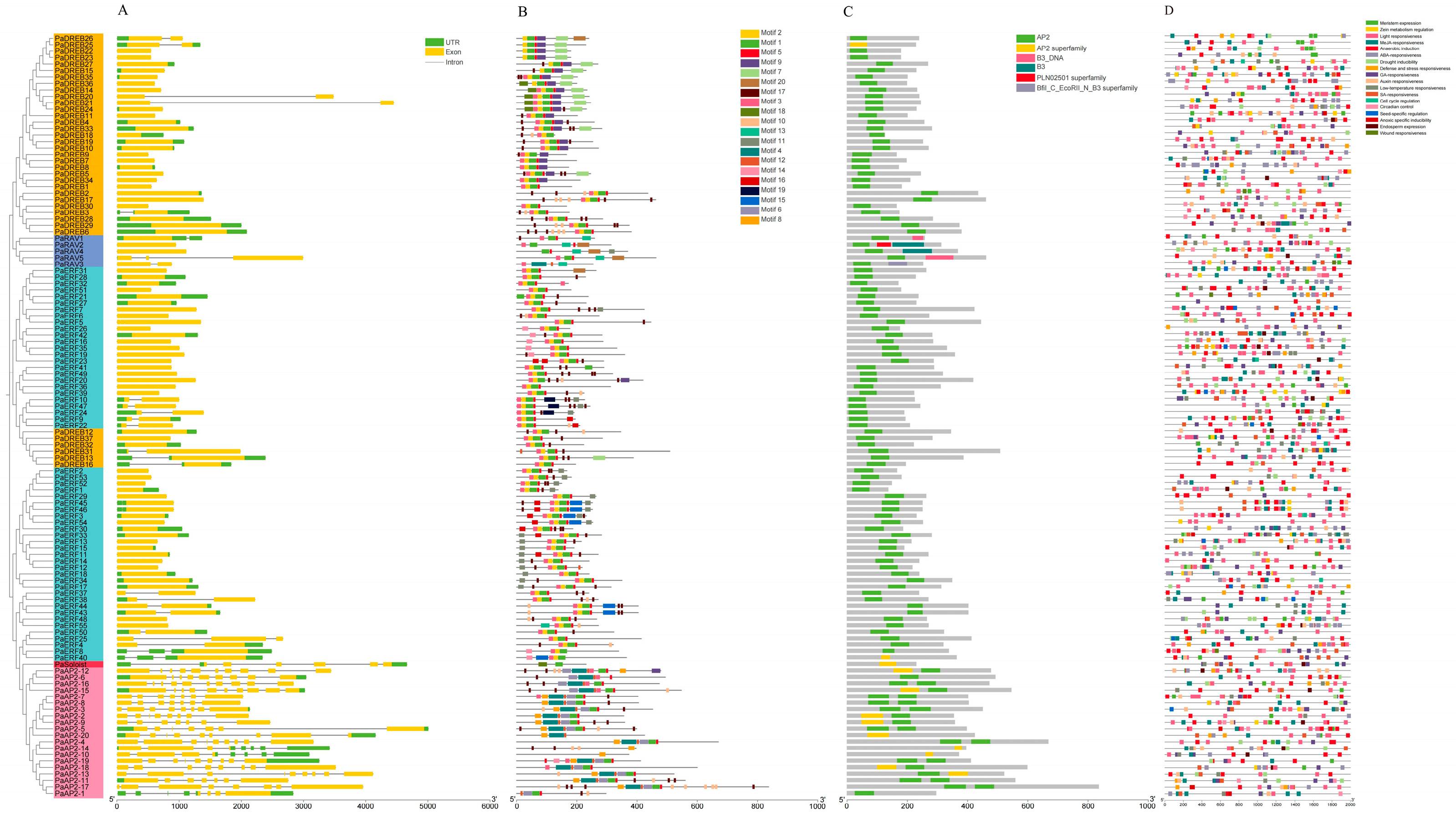
3.3. Gene Structure, Conserved Motif, and Domain Analysis of the AP2/ERF Family in Apricot
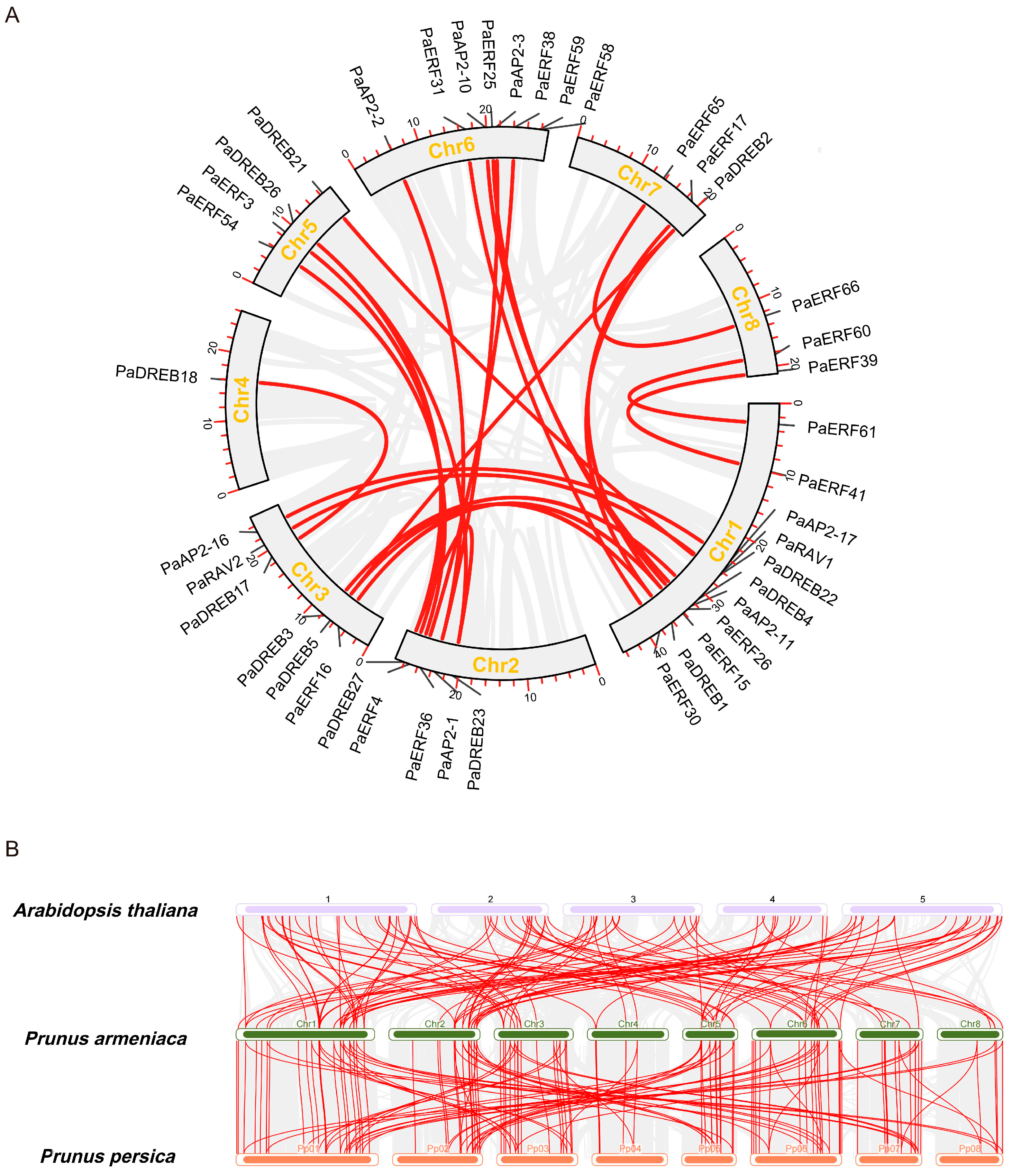
3.4. Chromosomal Localization and Syntenic Relationships of the PaAP2/ERF Family
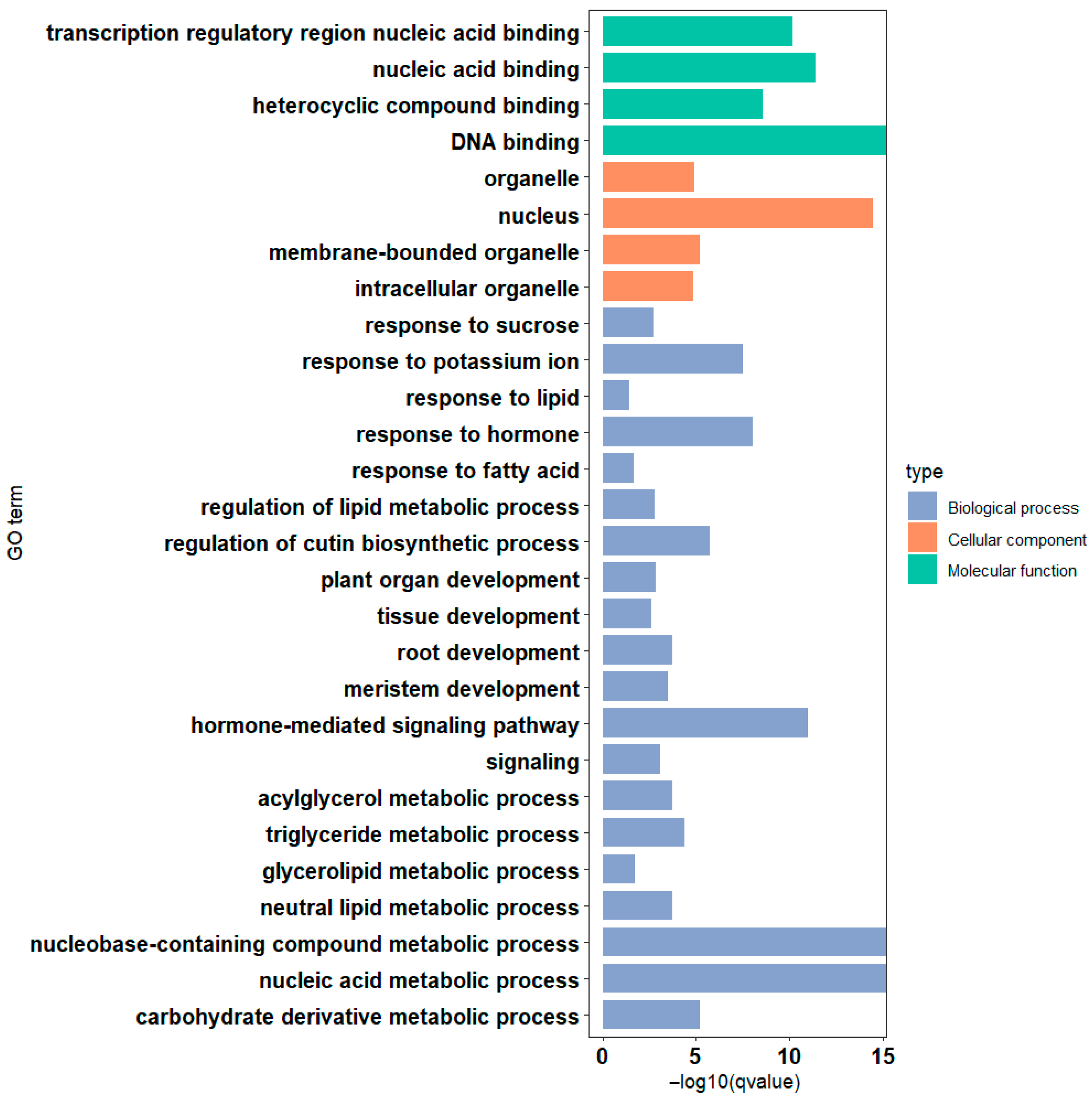
3.5. Analysis of Cis-Regulatory Elements and Functional Annotation of the PaAP2/ERF Family
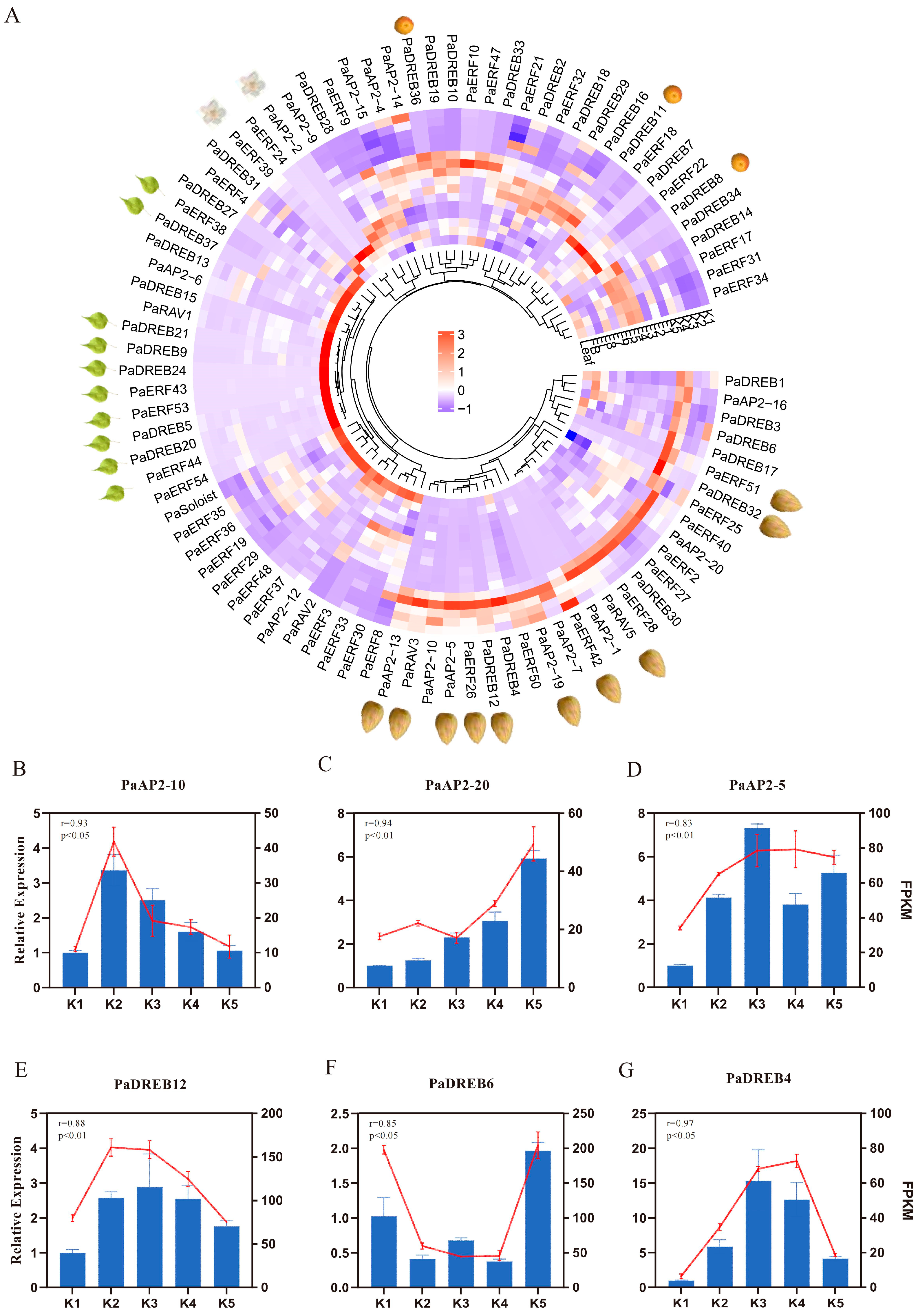
3.6. Expression Patterns of AP2/ERF Genes Across Tissues and Their Roles in Apricot Oil Accumulation
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Erdogan-Orhan, I.; Kartal, M. Insights into research on phytochemistry and biological activities of Prunus armeniaca L. (apricot). Food Res. Int. 2011, 44, 1238–1243. [Google Scholar] [CrossRef]
- Pawar, K.R.; Nema, P.K. Apricot kernel characterization, oil extraction, and its utilization: A review. Food Sci. Biotechnol. 2023, 32, 249–263. [Google Scholar] [CrossRef]
- Lee, H.; Ahn, J.; Kwon, A.; Lee, E.S.; Kwak, J.; Min, Y. Chemical composition and antimicrobial activity of the essential oil of apricot seed. Phytotherapy Res. 2014, 28, 1867–1872. [Google Scholar] [CrossRef]
- Kiralan, M.; Özkan, G.; Kucukoner, E.; Ozcelik, M.M. Apricot (Prunus armeniaca L.) Oil. In Fruit Oils: Chemistry and Functionality; Springer: Berlin/Heidelberg, Germany, 2019; pp. 505–519. Available online: https://link.springer.com/chapter/10.1007/978-3-030-12473-1_25 (accessed on 17 August 2025).
- Alves, M.S.; Dadalto, S.P.; Gonçalves, A.B.; De Souza, G.B.; Barros, V.A.; Fietto, L.G. Transcription factor functional protein-protein interactions in plant defense responses. Proteomes 2014, 2, 85–106. [Google Scholar] [CrossRef]
- Nakano, T.; Suzuki, K.; Fujimura, T.; Shinshi, H. Genome-Wide Analysis of the ERF Gene Family in Arabidopsis and Rice. Plant Physiol. 2006, 140, 411–432. [Google Scholar] [CrossRef] [PubMed]
- Licausi, F.; Ohme-Takagi, M.; Perata, P. APETALA2/Ethylene Responsive Factor (AP2/ERF) transcription factors: Mediators of stress responses and developmental programs. New Phytol. 2013, 199, 639–649. [Google Scholar] [CrossRef]
- Sakuma, Y.; Liu, Q.; Dubouzet, J.G.; Abe, H.; Shinozaki, K.; Yamaguchi-Shinozaki, K. DNA-binding specificity of the ERF/AP2 domain of Arabidopsis DREBs, transcription factors involved in dehydration-and cold-inducible gene expression. Biochem. Biophys. Res. Commun. 2002, 290, 998–1009. [Google Scholar] [CrossRef] [PubMed]
- Ohme-Takagi, M.; Shinshi, H. Ethylene-inducible DNA binding proteins that interact with an ethylene-responsive element. Plant Cell 1995, 7, 173. [Google Scholar] [CrossRef]
- Rashid, M.; Guangyuan, H.; Guangxiao, Y.; Hussain, J.; Xu, Y. AP2/ERF transcription factor in rice: Genome-wide canvas and syntenic relationships between monocots and eudicots. Evol. Bioinform. 2012, 8, 321–355. [Google Scholar] [CrossRef]
- Allen, M.D.; Yamasaki, K.; Ohme-Takagi, M.; Tateno, M.; Suzuki, M. A novel mode of DNA recognition by a β-sheet revealed by the solution structure of the GCC-box binding domain in complex with DNA. EMBO J. 1998, 17, 5484–5496. [Google Scholar] [CrossRef]
- Giri, M.K.; Swain, S.; Gautam, J.K.; Singh, S.; Singh, N.; Bhattacharjee, L.; Nandi, A.K. The Arabidopsis thaliana At4g13040 gene, a unique member of the AP2/EREBP family, is a positive regulator for salicylic acid accumulation and basal defense against bacterial pathogens. J. Plant Physiol. 2014, 171, 860–867. [Google Scholar] [CrossRef]
- Magnani, E.; SjölAnder, K.; Hake, S. From endonucleases to transcription factors: Evolution of the AP2 DNA binding domain in plants. Plant Cell 2004, 16, 2265–2277. [Google Scholar] [CrossRef] [PubMed]
- Sharoni, A.M.; Nuruzzaman, M.; Satoh, K.; Shimizu, T.; Kondoh, H.; Sasaya, T.; Choi, I.-R.; Omura, T.; Kikuchi, S. Gene structures, classification and expression models of the AP2/EREBP transcription factor family in rice. Plant Cell Physiol. 2010, 52, 344–360. [Google Scholar] [CrossRef]
- Liu, Q.; Wang, Z.; Xu, X.; Zhang, H.; Li, C.; Prasad, M. Genome-wide analysis of C2H2 zinc-finger family transcription factors and their responses to abiotic stresses in poplar (Populus trichocarpa). PLoS ONE 2015, 10, e0134753. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, J.; Cai, B.; Peng, R.-H.; Zhu, B.; Jin, X.-F.; Xue, Y.; Gao, F.; Fu, X.-Y.; Tian, Y.-S.; Zhao, W.; et al. Genome-wide analysis of the AP2/ERF gene family in Populus trichocarpa. Biochem. Biophys. Res. Commun. 2008, 371, 468–474. [Google Scholar] [CrossRef] [PubMed]
- Verde, I.; Jenkins, J.; Dondini, L.; Micali, S.; Pagliarani, G.; Vendramin, E.; Paris, R.; Aramini, V.; Gazza, L.; Rossini, L.; et al. The Peach v2.0 release: High-resolution linkage mapping and deep resequencing improve chromosome-scale assembly and contiguity. BMC Genom. 2017, 18, 1–18. [Google Scholar] [CrossRef]
- Gao, Z.; Li, Q.; Li, J.; Chen, Y.; Luo, M.; Li, H.; Wang, J.; Wu, Y.; Duan, S.; Wang, L.; et al. Characterization of the ABA receptor VlPYL1 that regulates anthocyanin accumulation in grape berry skin. Front. Plant Sci. 2018, 9, 592. [Google Scholar] [CrossRef]
- Tatsuki, M.; Nakajima, N.; Fujii, H.; Shimada, T.; Nakano, M.; Hayashi, K.-I.; Hayama, H.; Yoshioka, H.; Nakamura, Y. Increased levels of IAA are required for system 2 ethylene synthesis causing fruit softening in peach (Prunus persica L. Batsch). J. Exp. Bot. 2013, 64, 1049–1059. [Google Scholar] [CrossRef]
- Chuck, G.; Meeley, R.B.; Hake, S. The control of maize spikelet meristem fate by the APETALA2-like gene indeterminate spikelet1. Genes Dev. 1998, 12, 1145–1154. [Google Scholar] [CrossRef]
- El-Sharkawy, I.; Kim, W.S.; El-Kereamy, A.; Jayasankar, S.; Svircev, A.M.; Brown, D.C.W. Isolation and characterization of four ethylene signal transduction elements in plums (Prunus salicina L.). J. Exp. Bot. 2007, 58, 3631–3643. [Google Scholar] [CrossRef]
- Jofuku, K.D.; Omidyar, P.K.; Gee, Z.; Okamuro, J.K. Control of seed mass and seed yield by the floral homeotic gene APETALA2. Proc. Natl. Acad. Sci. USA 2005, 102, 3117–3122. [Google Scholar] [CrossRef]
- Hirota, A.; Kato, T.; Fukaki, H.; Aida, M.; Tasaka, M. The auxin-regulated AP2/EREBP gene PUCHI is required for morphogenesis in the early lateral root primordium of Arabidopsis. Plant Cell 2007, 19, 2156–2168. [Google Scholar] [CrossRef]
- van Raemdonck, D.; Pesquet, E.; Cloquet, S.; Beeckman, H.; Boerjan, W.; Goffner, D.; El Jaziri, M.; Baucher, M. Molecular changes associated with the setting up of secondary growth in aspen. J. Exp. Bot. 2005, 56, 2211–2227. [Google Scholar] [CrossRef]
- Shukla, R.K.; Raha, S.; Tripathi, V.; Chattopadhyay, D. Expression of CAP2, an APETALA2-family transcription factor from chickpea, enhances growth and tolerance to dehydration and salt stress in transgenic tobacco. Plant Physiol. 2006, 142, 113–123. [Google Scholar] [CrossRef]
- Dietz, K.-J.; Vogel, M.O.; Viehhauser, A. AP2/EREBP transcription factors are part of gene regulatory networks and integrate metabolic, hormonal and environmental signals in stress acclimation and retrograde signalling. Protoplasma 2010, 245, 3–14. [Google Scholar] [CrossRef]
- Yamaguchi-Shinozaki, K.; Shinozaki, K. A novel cis-acting element in an Arabidopsis gene is involved in responsiveness to drought, low-temperature, or high-salt stress. Plant Cell 1994, 6, 251–264. [Google Scholar] [PubMed]
- Finkelstein, R.R.; Wang, M.L.; Lynch, T.J.; Rao, S.; Goodman, H.M. The Arabidopsis abscisic acid response locus ABI4 encodes an APETALA2 domain protein. Plant Cell 1998, 10, 1043. [Google Scholar] [CrossRef] [PubMed]
- Kitomi, Y.; Ito, H.; Hobo, T.; Aya, K.; Kitano, H.; Inukai, Y. The auxin responsive AP2/ERF transcription factor CROWN ROOTLESS5 is involved in crown root initiation in rice through the induction of OsRR1, a type-A response regulator of cytokinin signaling. Plant J. 2011, 67, 472–484. [Google Scholar] [CrossRef]
- Wang, G.; Wang, H.; Zhu, J.; Zhang, J.; Zhang, X.; Wang, F.; Tang, Y.; Mei, B.; Xu, Z.; Song, R. An expression analysis of 57 transcription factors derived from ESTs of developing seeds in Maize (Zea mays). Plant Cell Rep. 2010, 29, 545–559. [Google Scholar] [CrossRef]
- Boutilier, K.; Offringa, R.; Sharma, V.K.; Kieft, H.; Ouellet, T.; Zhang, L.; Hattori, J.; Liu, C.-M.; van Lammeren, A.A.M.; Miki, B.L.A.; et al. Ectopic Expression of BABY BOOM Triggers a Conversion from Vegetative to Embryonic Growth. Plant Cell 2002, 14, 1737–1749. [Google Scholar] [CrossRef]
- Edgar, R.C. Muscle5: High-accuracy alignment ensembles enable unbiased assessments of sequence homology and phylogeny. Nat. Commun. 2022, 13, 6968. [Google Scholar] [CrossRef]
- Vilella, A.J.; Severin, J.; Ureta-Vidal, A.; Heng, L.; Durbin, R.; Birney, E. EnsemblCompara GeneTrees: Complete, duplication-aware phylogenetic trees in vertebrates. Genome Res. 2009, 19, 327–335. [Google Scholar] [CrossRef]
- Darriba, D.; Posada, D.; Kozlov, A.M.; Stamatakis, A.; Morel, B.; Flouri, T.; Crandall, K. ModelTest-NG: A New and Scalable Tool for the Selection of DNA and Protein Evolutionary Models. Mol. Biol. Evol. 2020, 37, 291–294. [Google Scholar] [CrossRef]
- Minh, B.Q.; Schmidt, H.A.; Chernomor, O.; Schrempf, D.; Woodhams, M.D.; von Haeseler, A.; Lanfear, R. IQ-TREE 2: New Models and Efficient Methods for Phylogenetic Inference in the Genomic Era. Mol. Biol. Evol. 2020, 37, 1530–1534. [Google Scholar] [CrossRef]
- Letunic, I.; Life, P.B. Interactive Tree of Life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 2021, 49, W293–W296. Available online: https://pubmed.ncbi.nlm.nih.gov/33885785/ (accessed on 17 August 2025). [CrossRef]
- Chen, C.J.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.H.; Xia, R. TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Marchler-Bauer, A.; Bo, Y.; Han, L.; He, J.; Lanczycki, C.J.; Lu, S.; Chitsaz, F.; Derbyshire, M.K.; Geer, R.C.; Gonzales, N.R.; et al. CDD/SPARCLE: Functional classification of proteins via subfamily domain architectures. Nucleic Acids Res. 2017, 45, D200–D203. [Google Scholar] [CrossRef]
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouzé, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef] [PubMed]
- Wu, T.; Hu, E.; Xu, S.; Chen, M.; Guo, P.; Dai, Z.; Feng, T.; Zhou, L.; Tang, W.; Zhan, L.; et al. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation 2021, 2, 100141. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Tang, H.; DeBarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.-H.; Jin, H.; Marler, B.; Guo, H.; et al. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357–360. [Google Scholar] [CrossRef] [PubMed]
- Luo, J.; Nvsvrot, T.; Wang, N. Comparative transcriptomic analysis uncovers conserved pathways involved in adventitious root formation in poplar. Physiol. Mol. Biol. Plants 2021, 27, 1903–1918. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Lu, Y.; Wang, J.; Hu, J.; Wuyun, T. Genome-wide screening of long non-coding RNAs involved in rubber biosynthesis in Eucommia ulmoides. J. Integr. Plant Biol. 2018, 60, 1070–1082. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Gayas, B.; Kaur, G.; Singh, A. Ultrasound assisted extraction of apricot kernel oil: Effect on physicochemical, morphological characteristics, and fatty acid composition. Acta Aliment. 2020, 49, 23–31. [Google Scholar] [CrossRef]
- Maeo, K.; Tokuda, T.; Ayame, A.; Mitsui, N.; Kawai, T.; Tsukagoshi, H.; Ishiguro, S.; Nakamura, K. An AP2-type transcription factor, WRINKLED1, of Arabidopsis thaliana binds to the AW-box sequence conserved among proximal upstream regions of genes involved in fatty acid synthesis. Plant J. 2009, 60, 476–487. [Google Scholar] [CrossRef]
- Feng, K.; Hou, X.; Xing, G.; Liu, J.; Duan, A.; Xu, Z.; Li, M.; Zhuang, J.; Xiong, A. Advances in AP2/ERF super-family transcription factors in plant. Crit. Rev. Biotechnol. 2020, 40, 750–776. [Google Scholar] [CrossRef]
- Okamuro, J.K.; Caster, B.; Villarroel, R.; Van Montagu, M.; Jofuku, K.D. The AP2 domain of APETALA2 defines a large new family of DNA binding proteins in Arabidopsis. Proc. Natl. Acad. Sci. USA 1997, 94, 7076–7081. [Google Scholar] [CrossRef]
- Shigyo, M.; Hasebe, M.; Ito, M. Molecular evolution of the AP2 subfamily. Gene 2006, 366, 256–265. [Google Scholar] [CrossRef]
- Kim, S.; Soltis, P.S.; Wall, K.; Soltis, D.E. Phylogeny and Domain Evolution in the APETALA2-like Gene Family. Mol. Biol. Evol. 2006, 23, 107–120. [Google Scholar] [CrossRef]
- Han, Y.; Cai, M.; Zhang, S.; Chai, J.; Sun, M.; Wang, Y.; Xie, Q.; Chen, Y.; Wang, H.; Chen, T. Genome-wide identification of AP2/ERF transcription factor family and functional analysis of DcAP2/ERF#96 associated with abiotic stress in Dendrobium catenatum. Int. J. Mol. Sci. 2022, 23, 13603. [Google Scholar] [CrossRef]
- Roy, S.W.; Gilbert, W. The evolution of spliceosomal introns: Patterns, puzzles and progress. Nat. Rev. Genet. 2006, 7, 211–221. [Google Scholar] [CrossRef] [PubMed]
- Kordiš, D. Extensive intron gain in the ancestor of placental mammals. Biol. Direct 2011, 6, 59. [Google Scholar] [CrossRef] [PubMed]
- Binford, G.J.; Bodner, M.R.; Cordes, M.H.; Baldwin, K.L.; Rynerson, M.R.; Burns, S.N.; Zobel-Thropp, P.A. Molecular evolution, functional variation, and proposed nomenclature of the gene family that includes sphingomyelinase D in sicariid spider venoms. Mol. Biol. Evol. 2009, 26, 547–566. [Google Scholar] [CrossRef]
- Huang, Y.-L.; Zhang, L.-K.; Zhang, K.; Chen, S.-M.; Hu, J.-B.; Cheng, F. The impact of tandem duplication on gene evolution in Solanaceae species. J. Integr. Agric. 2022, 21, 1004–1014. [Google Scholar] [CrossRef]
- Rensing, S.A. Gene duplication as a driver of plant morphogenetic evolution. Curr. Opin. Plant Biol. 2014, 17, 43–48. [Google Scholar] [CrossRef]
- Qiao, X.; Li, Q.; Yin, H.; Qi, K.; Li, L.; Wang, R.; Zhang, S.; Paterson, A.H. Gene duplication and evolution in recurring polyploidization–diploidization cycles in plants. Genome Biol. 2019, 20, 38. [Google Scholar] [CrossRef]
- Van de Peer, Y.; Mizrachi, E.; Marchal, K. The evolutionary significance of polyploidy. Nat. Rev. Genet. 2017, 18, 411–424. [Google Scholar] [CrossRef]
- Riaz, M.W.; Lu, J.; Shah, L.; Yang, L.; Chen, C.; Mei, X.D.; Xue, L.; Manzoor, M.A.; Abdullah, M.; Rehman, S.; et al. Expansion and Molecular Characterization of AP2/ERF Gene Family in Wheat (Triticum aestivum L.). Front. Genet. 2021, 12, 632155. [Google Scholar] [CrossRef]
- Lawton-Rauh, A. Evolutionary dynamics of duplicated genes in plants. Mol. Phylogenetics Evol. 2003, 29, 396–409. [Google Scholar] [CrossRef]
- Jung, S.; Jiwan, D.; Cho, I.; Lee, T.; Abbott, A.; Sosinski, B.; Main, D. Synteny of Prunus and other model plant species. BMC Genom. 2009, 10, 76. [Google Scholar] [CrossRef]
- Nakashima, K.; Ito, Y.; Yamaguchi-Shinozaki, K. Transcriptional regulatory networks in response to abiotic stresses in arabidopsis and grasses. Plant Physiol. 2009, 149, 88–95. [Google Scholar] [CrossRef]
- Soares, V.L.F.; Rodrigues, S.M.; de Oliveira, T.M.; de Queiroz, T.O.; Lima, L.S.; Hora-Júnior, B.T.; Gramacho, K.P.; Micheli, F.; Cascardo, J.C.M.; Otoni, W.C.; et al. Unraveling new genes associated with seed development and metabolism in Bixa orellana L. by expressed sequence tag (EST) analysis. Mol. Biol. Rep. 2011, 38, 1329–1340. [Google Scholar] [CrossRef] [PubMed]
- Kanai, M.; Mano, S.; Kondo, M.; Hayashi, M.; Nishimura, M. Extension of oil biosynthesis during the mid-phase of seed development enhances oil content in Arabidopsis seeds. Plant Biotechnol. J. 2016, 14, 1241–1250. [Google Scholar] [CrossRef] [PubMed]
- Xie, Z.; Nolan, T.M.; Jiang, H.; Yin, Y. AP2/ERF Transcription Factor Regulatory Networks in Hormone and Abiotic Stress Responses in Arabidopsis. Front. Plant Sci. 2019, 10, 228. [Google Scholar] [CrossRef] [PubMed]
- Pegoraro, C.; Tadiello, A.; Girardi, C.L.; Chaves, F.C.; Quecini, V.; de Oliveira, A.C.; Trainotti, L.; Rombaldi, C.V. Transcriptional regulatory networks controlling woolliness in peach in response to preharvest gibberellin application and cold storage. BMC Plant Biol. 2015, 15, 279. [Google Scholar] [CrossRef]
- Liu, N. Effects of IAA and ABA on the Immature Peach Fruit Development Process. Hortic. Plant J. 2019, 5, 145–154. [Google Scholar] [CrossRef]
- Soto, A.; Ruiz, K.B.; Ravaglia, D.; Costa, G.; Torrigiani, P. ABA may promote or delay peach fruit ripening through modulation of ripening- and hormone-related gene expression depending on the developmental stage. Plant Physiol. Biochem. 2013, 64, 11–24. [Google Scholar] [CrossRef]
- Hu, Y.X.; Wang, Y.H.; Liu, X.F.; Li, J.Y. Arabidopsis RAV1 is down-regulated by brassinosteroid and may act as a negative regulator during plant development. Cell Res. 2004, 14, 8–15. [Google Scholar] [CrossRef]
- Woo, H.R.; Kim, J.H.; Kim, J.; Kim, J.; Lee, U.; Song, I.J.; Kim, J.H.; Lee, H.Y.; Nam, H.G.; Lim, P.O. The RAV1 transcription factor positively regulates leaf senescence in Arabidopsis. J. Exp. Bot. 2010, 61, 3947–3957. [Google Scholar] [CrossRef]
- Cernac, A.; Benning, C. WRINKLED1 encodes an AP2/EREB domain protein involved in the control of storage compound biosynthesis in Arabidopsis. Plant J. 2004, 40, 575–585. [Google Scholar] [CrossRef]
- Baud, S.; Mendoza, M.S.; To, A.; Harscoët, E.; Lepiniec, L.; Dubreucq, B. WRINKLED1 specifies the regulatory action of LEAFY COTYLEDON2 towards fatty acid metabolism during seed maturation in Arabidopsis. Plant J. 2007, 50, 825–838. [Google Scholar] [CrossRef]
- Deng, S.; Mai, Y.; Shui, L.; Niu, J. WRINKLED1 transcription factor orchestrates the regulation of carbon partitioning for C18:1 (oleic acid) accumulation in Siberian apricot kernel. Sci. Rep. 2019, 9, 2693. [Google Scholar] [CrossRef]
- Savadi, S.; Lambani, N.; Kashyap, P.L.; Bisht, D.S. Genetic engineering approaches to enhance oil content in oilseed crops. Plant Growth Regul. 2017, 83, 207–222. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
He, Y.; Wang, L.; Jiang, N.; Zhang, D.; Shi, X.; Wuyun, T.; Liu, H. Genome-Wide Identification and Expression Profiling of AP2/ERF Transcription Factor Genes in Prunus armeniaca L. Forests 2025, 16, 1353. https://doi.org/10.3390/f16081353
He Y, Wang L, Jiang N, Zhang D, Shi X, Wuyun T, Liu H. Genome-Wide Identification and Expression Profiling of AP2/ERF Transcription Factor Genes in Prunus armeniaca L. Forests. 2025; 16(8):1353. https://doi.org/10.3390/f16081353
Chicago/Turabian StyleHe, Yanguang, Lin Wang, Nan Jiang, Donglin Zhang, Xiaodan Shi, Tana Wuyun, and Huimin Liu. 2025. "Genome-Wide Identification and Expression Profiling of AP2/ERF Transcription Factor Genes in Prunus armeniaca L." Forests 16, no. 8: 1353. https://doi.org/10.3390/f16081353
APA StyleHe, Y., Wang, L., Jiang, N., Zhang, D., Shi, X., Wuyun, T., & Liu, H. (2025). Genome-Wide Identification and Expression Profiling of AP2/ERF Transcription Factor Genes in Prunus armeniaca L. Forests, 16(8), 1353. https://doi.org/10.3390/f16081353






