Mating System Analysis and Genetic Diversity of Parkia multijuga Benth. One Native Tree Species of the Amazon
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Area and Sampling
2.2. Amplification of Microsatellite Markers
2.3. Data Analysis
3. Results
3.1. Crossing System
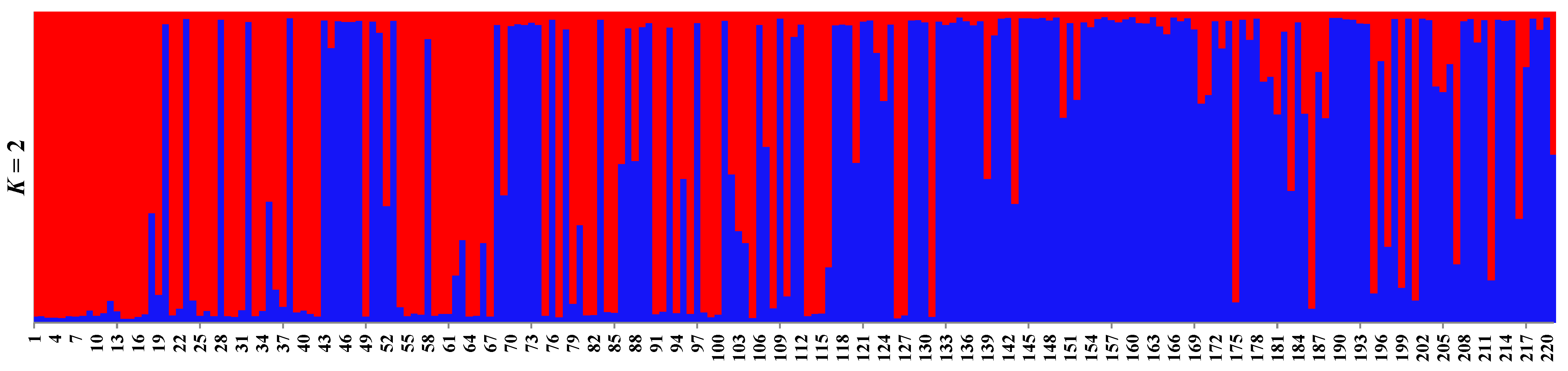
3.2. Diversity and Genetic Structure
4. Discussion
4.1. Crossing System
4.2. Diversity and Genetic Structure
4.3. Implications for Conservation
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Carvalho, P.E.R. Espécies Arbóreas Brasileiras; Embrapa Informação Tecnológica Brasília: Brasília, Brazil, 2008; Volume 3. [Google Scholar]
- Canuto, D.S.O.; Zaruma, D.U.G.; Moraes, M.A.; Silva, A.M.; Moraes, M.L.T.; Freitas, M.L.M. Genetic characterization of a progeny test of Dipteryx alata Vog., from a forest fragment of Estação Ecológica Paulo de Faria, São Paulo State, Brazil. Hoehnea 2015, 42, 641–648. [Google Scholar] [CrossRef]
- Projeto PRODES Monitoramento da Floresta Amazônica Brasileira por Satélite. Available online: http://terrabrasilis.dpi.inpe.br/downloads (accessed on 22 August 2023).
- Pedersoli, G.D.; Pádua Teixeira, S. Floral development of Parkia multijuga and Stryphnodendron adstringens, two andromonoecious mimosoid trees (Leguminosae). Int. J. Plant Sci. 2016, 177, 60–75. [Google Scholar] [CrossRef]
- Hopkins, H.C. Floral biology and pollination ecology of the neotropical species of Parkia. J. Ecol. 1984, 72, 1–23. [Google Scholar] [CrossRef]
- Borpuzazri, P.P.; Singh, M.K. A review on pollination study of Parkia- An economically and ecologically important genus of food tree species. Indian J. Agrofor. 2022, 24, 93–99. [Google Scholar]
- Kottek, M.; Grieser, J.; Beck, C.; Rudolf, B.; Rubel, F. World Map of the Köppen-Geiger climate classification updated. Meteorol. Z. 2006, 15, 259–263. [Google Scholar] [CrossRef] [PubMed]
- Beck, H.; Zimmermann, N.; McVicar, T.; Vergopolan, N.; Berg, A.; Wood, E.F. Present and future Köppen-Geiger climate classification maps at 1-km resolution. Sci. Data 2018, 5, 180214. [Google Scholar] [CrossRef] [PubMed]
- Serrote, C.M.L.; Reiniger, L.R.S.; Stefanel, C.M.; da Silva, K.B.; Golle, D.P. Microsatellites are important for forest genetic resources conservation in Brazilian biomes. Acta Bot. Bras. 2023, 37, e20220176. [Google Scholar] [CrossRef]
- Silva, D.Y.B.D.O.; Farias, S.G.G.D.; Araújo, P.C.D.; Sousa, M.B.D.; Silva, R.B.E.; Oliveira, C.V.D.A. Genetic variability of Parkia platycephala populations: Support for defining seed collection areas. Rev. Caatinga 2022, 35, 905–914. [Google Scholar] [CrossRef]
- Guimarães, Z.T.M.; Lopes, K.F.L.; Barbosa, M.S.; Santos, V.A.H.F.; Silva, T.V.M.; Oliveira, R.G.; Lima Júnior, M.J.V.; Martins, N.O.A.; Ferreira, M.J. Silvicultural performance of Parkia multijuga Benth. progenies in the Amazonas three years after planting. Cienc. Florest. 2022, 32, 43–70. [Google Scholar] [CrossRef]
- Chaves, S.R.; Santos, R.R.; Silva, A.L.G. Reproductive biology of Parkia platycephala Benth (Leguminosae, Caesal-pinioideae, clado mimosoide). Braz. J. Dev. 2020, 6, 79442–79458. [Google Scholar] [CrossRef]
- Barros, H.S.D.; Cruz, E.D.; Pereira, A.G.; Silva, E.A.A. Fruit and seed morphometry, seed germination and seedling vigor of Parkia gigantocarpa. Rev. Florest. 2019, 50, 877–886. [Google Scholar] [CrossRef]
- Ouborg, N.J.; Pertoldi, C.; Loeschcke, V.; Bijlsma, R.; Hedrick, P.W. Conservation genetics in transition to conservation genomics. Trends Genet. 2010, 26, 177–187. [Google Scholar] [CrossRef] [PubMed]
- Caixeta, E.T.; Oliveira, A.C.B.; Brito, G.G.; Sakiyama, N.S. Tipos de Marcadores Moleculares. In Marcadores Moleculares; Borem, A., Caixeta, E.T., Eds.; Editora UFV: Vicosa, Brazil, 2009; pp. 11–93. [Google Scholar]
- Coelho, N.H.P.; Tambarussi, E.V.; Aguiar, B.I.; Roque, R.H.; Portela, R.M.; Braga, R.C.; Sanson, D.; Silva, R.A.R.; Ferraz, E.M.; Moreno, M.A.; et al. Understanding genetic diversity, spatial genetic structure, and mating system through microsatellite markers for the conservation and sustainable use of Acrocomia aculeata (Jacq.) Lodd. Ex Mart. Conserv. Genet. 2018, 19, 879–891. [Google Scholar] [CrossRef]
- Ramos, S.L.F.; Lopes, M.T.G.; Lopes, R.; Dequigiovanni, G.; Macêdo, J.L.V.D.; Sebbenn, A.M.; Silva, E.B.D.; Garcia, J.N. Mating system analysis of Açaí-do-Amazonas (Euterpe precatoria Mart.) using molecular markers. Crop Breed. Appl. Biotechnol. 2019, 19, 126–130. [Google Scholar] [CrossRef]
- Alves, R.M.; Artero, A.S.; Sebbenn, A.M.; Figueira, A. Mating system in a natural population of Theobroma grandiflorum (Willd. ex Spreng.) Schum., by microsatellite markers. Genet. Mol. Biol. 2003, 26, 373–379. [Google Scholar] [CrossRef]
- Campos, T.; Cunha, M.O.; De Sousa, A.C.B.; Teixeira, R.B.; Raposo, A.; Sebbenn, A.M.; Wadt, L.H.O. Mating system parameters in a high density population of andirobas in the Amazon forest. Pesqui. Agropecu. Bras. 2013, 48, 504–509. [Google Scholar] [CrossRef][Green Version]
- Peel, M.C.; Finlayson, B.L.; McMahon, T.A. Updated world map of the Köppen-Geiger climate classification. Hydrol. Earth Syst. Sci. 2007, 11, 1633–1644. [Google Scholar] [CrossRef]
- Doyle, J.J.; Doyle, J.L. Isolation of plant DNA from fresh tissue. Focus 1990, 12, 13–15. [Google Scholar]
- Luettmann, K.; Michalczyk, I.M.; Mengel, C.; Ziegenhagen, B.; Heymann, E.W.; Saboya, P.P.P.; Bialozyt, R. Characterization of nuclear microsatellite loci in the neotropical tree Parkia panurensis (Fabaceae). Am. J. Bot. 2010, 4, e34–e36. [Google Scholar] [CrossRef]
- Schuelke, M. An economic method for the fluorescent labeling of PCR fragments. Nat. Biotechnol. 2000, 18, 233–234. [Google Scholar] [CrossRef]
- Andrade, R.L.; Pico-Mendoza, J.; Morillo, E.; Buitrón, J.; Meneses, S.; Navarrete, B.; Pinoargote, M.; Carrasco, B. Molecular characterization of mahogany tree (Swietenia macrophylla King, Meliaceae) in the remnant natural forest of Ecuador. Neotrop. Biodivers. 2022, 8, 222–228. [Google Scholar] [CrossRef]
- Ritland, K. Multilocus Mating System Program—MLTR Version 3.0. Vancouver. 2004. Available online: http://genetics.forestry.ubc.ca/ritland/programs.htlm (accessed on 10 August 2023).
- Ritland, K. Extensions of models for the estimation of mating systems using in independent loci. Heredity 2002, 88, 221–228. [Google Scholar] [CrossRef]
- Ritland, K. Correlated Matings in the Partial Selfer Mimulus guttatus. Evolution 1989, 43, 848–859. [Google Scholar] [CrossRef]
- Feres, J.M.; Sebbenn, A.M.; Guidugli, M.C.; Mestriner, M.A.; Moraes, M.L.T.; Alzate-Marin, A.L. Mating system parameters at hierarchical levels of fruits, individuals and populations in the Brazilian insect-pollinated tropical tree, Tabebuia roseaalba (Bignoniaceae). Conserv. Genet. 2012, 13, 393–405. [Google Scholar] [CrossRef]
- Cockerham, C.C. Variance of gene frequency. Evolution 1969, 23, 72–84. [Google Scholar] [CrossRef]
- Nei, M. F-statistics and analysis of gene diversity in subdivided populations. Ann. Hum. Genet. 1977, 41, 225–233. [Google Scholar] [CrossRef]
- Wadt, L.H.O.; Baldoni, A.B.; Silva, V.S.; Campos, T.; Martins, K.; Azevedo, V.C.R.; Mata, L.R.; Botin, A.A.; Hoogerheide, E.S.S.; Tonini, T.; et al. Mating system variation among populations, individuals and within and among fruits in Bertholletia excelsa. Silvae Genet. 2015, 64, 5–6. [Google Scholar] [CrossRef]
- Nunney, L.; Campbell, K.A. Assessing minimum viable population size: Demography meets population genetics. Tree 1993, 8, 234–239. [Google Scholar] [CrossRef]
- Peakall, R.; Smouse, P.E. GENALEX 6: Genetic analysis in Excel. Population genetic software for teaching and research. Mol. Ecol. Notes 2006, 6, 288–295. [Google Scholar] [CrossRef]
- Peakall, R.; Smouse, P.E. GenAlEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research—An update. Bioinformatics 2012, 28, 2537–2539. [Google Scholar] [CrossRef]
- Wright, S. The Genetical Structure of Populations. Ann. Eugen. 1951, 15, 323–354. [Google Scholar] [CrossRef]
- Weir, B.S.; Cockerham, C.C. Estimating F-statistic for the analysis of population structure. Evolution 1984, 38, 1358–1370. [Google Scholar]
- Pritchard, J.K.; Stephens, M.; Donnelly, P.J. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar] [CrossRef]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software structure: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef]
- Jakobsson, M.; Rosenberg, N.A. CLUMPP: A cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 2007, 23, 1801–1806. [Google Scholar] [CrossRef]
- Rosenberg, N.A. Distruct: A program for the graphical display of population structure. Mol. Ecol. Notes 2004, 4, 137–138. [Google Scholar] [CrossRef]
- Mori, N.T.; Mori, E.S.; Tambarussi, E.V.; Moraes, M.L.T.; Sebbenn, A.M. Mating system in Handroanthus heptaphyllus (Vell.) Mattos populations and its implications for seed collection to genetic conservation and breeding. Sci. For. 2015, 43, 675–681. [Google Scholar]
- Guidice-Neto, J.D.; Sebbenn, A.M.; Kageyama, P.Y. Mating system in Caesalpinia echinata Lam. implanted in experimental arboretum. Rev. Bras. Bot. 2005, 28, 409–418. [Google Scholar] [CrossRef]
- Seoane, C.E.S.; Sebbenn, A.M.; Kageyama, P.Y. Mating system in two Euterpe edulis M. natural populations under different forest fragmentation conditions. Sci. For. 2005, 69, 13–24. [Google Scholar]
- Dequigiovanni, G.; Ramos, S.L.F.; Alves-Pereira, A.; Fabri, E.G.; Picanço-Rodrigues, D.; Clement, C.R.; Gepts, P.; Veasey, E.A. Highly structured genetic diversity of Bixa orellana var. urucurana, the wild ancestor of annatto, in Brazilian Amazonia. PLoS ONE 2018, 13, e0198593. [Google Scholar] [CrossRef]
- Siqueira, M.V.B.M.; Costa, F.M.; Novaes, C.R.D.B.; Bustamante, P.G.; de Moraes Silvestre, M.A. In Situ, Ex Situ and On Farm Conservation of Plant Genetic Resources in Neotropics. In Conservation Genetics in the Neotropics, 1st ed.; Galetti, P.M., Jr., Ed.; Springer: Cham, Switzerland, 2023; Volume 1, pp. 201–225. [Google Scholar]
- Nazareno, A.G.; dos Reis, M.S. At Risk of Population Decline? An Ecological and Genetic Approach to the Threatened Palm Species Butia eriospatha (Arecaceae) of Southern Brazil. J. Hered. 2014, 105, 120–129. [Google Scholar] [CrossRef] [PubMed]
- Hartl, D.L.; Clark, A.G.; Clark, A.G. Principles of Population Genetics; Sinauer associates Sunderland: Sunderland, MA, USA, 1997; Volume 116. [Google Scholar]
- Pannell, J.R.; Fields, P.D. Evolution in subdivided plant populations: Concepts, recent advances and future directions. New Phytol. 2014, 201, 417–432. [Google Scholar] [CrossRef] [PubMed]
- Kalinowski, S.T. Counting alleles with rarefaction: Private alleles and hierarchical sampling designs. Conserv. Genet. 2004, 5, 539–543. [Google Scholar] [CrossRef]
- Leducq, J.B.; Llaurens, V.; Castric, V.; Saumitou-Laprade, P.; Hardy, O.J.; Vekemans, X. Effect of balancing selection on spatial genetic structure within populations: Theoretical investigations on the self-incompatibility locus and empirical studies in Arabidopsis halleri. Heredity 2011, 106, 319–329. [Google Scholar] [CrossRef] [PubMed]
- Gonçalves, A.L.; García, M.V.; Barrandeguy, M.E.; González-Martínez, S.C.; Heuertz, M. Spatial genetic structure and mating system in forest tree populations from seasonally dry tropical forests: A review. Tree Genet. Genomes 2022, 18, 18. [Google Scholar] [CrossRef]
- Browne, L.; Ottewell, K.M.; Sork, V.L.; Karubian, J. The relative contributions of seed and pollen dispersal to gene flow and genetic diversity in seedlings of a tropical palm. Mol. Ecol. 2018, 27, 3159–3173. [Google Scholar] [CrossRef]
- Li, F.; Chen, H.; Liu, S.; Zhang, H.; Zhou, Z. Mating systems of single families and population genetic diversity of endangered Ormosia hosiei in south China. Genes 2022, 13, 2117. [Google Scholar] [CrossRef]
| Locus | Repeat Motif | Primer Sequences (5′-3′) | AT °C | (bp) | AN GenBank |
|---|---|---|---|---|---|
| Parpan 3 | (GT)15 | F: CACGTTAATTCAATCAAAATGGTG | 56.5 | 155–209 | GU735073 |
| R: TTTTGCCTTTTTCGGACTTG | |||||
| Parpan 4 | (GT)13(GA)15 | F: TTGATGGGAGTGGGAAAAAG | 54.0 | 148–210 | GU735074 |
| R: CAGGAGGTGGTCTCTTCAGG | |||||
| Parpan 5 | (CA)17 | F: CTCAATAAGATACCCTTTACATTGC | 60.0 | 166–200 | GU735075 |
| R: TTGAATCGAGGAATGAGATTATTG | |||||
| Parpan 9 | (AC)12 | F: GGGGCTTGTGTCTCTCACTG | 58.0 | 204–262 | GU735076 |
| R: ACTTTGAAGGCACGAGATGG | |||||
| Parpan 11 | (TG)5CA(TG)16 | F: ACGTAGGGAATAGGGCCATC | 58.5 | 94–214 | GU735077 |
| R: CTACGTACGAGCCGACACTC | |||||
| Parpan 13 | (GT)17TT(GT)8 | F: CCTCCCTCGCTTCACAATC | 58.5 | 86–194 | GU735078 |
| R: CACATGCAAATGAAAATGGTG | |||||
| Parpan 14 | (GT)20 | F: ACATCAAAATGGTCGCTCAAC | 60.0 | 76–116 | GU735079 |
| R: CAAATGTTCTTGTATGGAGCAAG | |||||
| Parpan 15 | (AC)24 | F: TGGCCTCACTGCATACTGAC | 55.0 | 104–152 | GU735080 |
| R: TGGGATGAACAAAACTGTGC | |||||
| Parpan 21 | (GT)15 | F: TGCTTTGTGCGACTTGAATC | 58.0 | 153–185 | GU735081 |
| R: CATTGTTCAGCATATAGGCATACAG |
| Parameters | Mean (95% CI) ¹ |
|---|---|
| Maternal fixation index: | 0 (0–0) |
| Multilocus outcrossing rate: | 1.0 (1.0–1.0) |
| Single-locus outcrossing rate: | 0.998 (0.997–1.000) |
| Mating among relatives: | 0.002 (0.003–0.000) |
| Selfing correlation: | 0.11 (0.11–0.11) |
| Paternity correlation: | 0.307 (0.193–0.338) |
| Number of pollen donors: | 3.3 (3.0–5.2) |
| Frequency of self-sibs: | 0 (0–0) |
| Frequency of self-half-sibs: | 0 (0–0) |
| Frequency of half-sibs: | 0.693 (0.807–0.662) |
| Frequency of full-sibs: | 0.307 (0.193–0.338) |
| Coancestry within family: | 0.163 (0.149–0.167) |
| Variance effective size: | 2.730 (2.676–2.945) |
| Number of seed trees: | 55 (51–56) |
| Mothers | (SD) | (SD) | (SD) | |||
|---|---|---|---|---|---|---|
| Family 1 | 0.998 (0.005) | 0.247 (0.033) | 0.656 (0.106) | 1.52 | 0.207 | 2.271 |
| Family 2 | 0.998 (0.005) | 0.014 (0.001) | 0.224 (0.079) | 4.46 | 0.153 | 3.025 |
| Family 3 | 0.998 (0.005) | 0.020 (0.002) | 0.153 (0.065) | 6.54 | 0.145 | 3.170 |
| Family 4 | 0.998 (0.005) | 0.029 (0.004) | 0.383 (0.136) | 2.61 | 0.173 | 2.725 |
| Family 5 | 0.998 (0.005) | 0.021 (0.002) | 0.127 (0.077) | 7.87 | 0.141 | 3.302 |
| Family 6 | 0.998 (0.005) | 0.034 (0.006) | 0.267 (0.099) | 3.75 | 0.159 | 2.963 |
| Family 7 | 0.998 (0.005) | 0.053 (0.011) | 0.251 (0.114) | 3.98 | 0.157 | 2.941 |
| Family 8 | 0.998 (0.005) | 0.059 (0.013) | 0.281 (0.090) | 3.56 | 0.161 | 2.850 |
| Family 9 | 0.998 (0.005) | 0.034 (0.007) | 0.091 (0.044) | 10.99 | 0.137 | 3.355 |
| Family 10 | 0.998 (0.005) | 0.062 (0.013) | 0.233 (0.097) | 4.29 | 0.155 | 3.022 |
| Family 11 | 0.998 (0.005) | 0.096 (0.029) | 0.198 (0.062) | 5.05 | 0.150 | 3.060 |
| Family 12 | 0.998 (0.005) | 0.007 (0.000) | 0.192 (0.069) | 5.21 | 0.149 | 3.070 |
| Family 13 | 0.998 (0.005) | 0.045 (0.006) | 0.291 (0.092) | 3.44 | 0.162 | 2.879 |
| Family | N (SE) | A (SE) | NP (SE) | HO (SE) | HE (SE) | f (SE) |
|---|---|---|---|---|---|---|
| Fam1 | 15.111 (0.716) | 3.000 (0.373) | 0.000 (0.000) | 0.655 (0.085) | 0.543 (0.038) | –0.201 (0.126) |
| Fam2 | 17.000 (0.000) | 4.667 (0.441) | 0.000 (0.000) | 0.837 (0.057) | 0.647 (0.034) | –0.291 (0.056) |
| Fam3 | 17.000 (0.000) | 4.556 (0.294) | 0.111 (0.111) | 0.837 (0.065) | 0.659 (0.038) | –0.265 (0.062) |
| Fam4 | 16.222 (0.434) | 4.000 (0.408) | 0.000 (0.000) | 0.820 (0.080) | 0.613 (0.047) | –0.323 (0.052) |
| Fam5 | 16.778 (0.147) | 5.111 (0.633) | 0.000 (0.000) | 0.906 (0.038) | 0.661 (0.031) | –0.379 (0.054) |
| Fam6 | 16.778 (0.222) | 4.333 (0.333) | 0.000 (0.000) | 0.876 (0.047) | 0.649 (0.031) | –0.350 (0.047) |
| Fam7 | 16.556 (0.242) | 4.889 (0.611) | 0.000 (0.000) | 0.802 (0.080) | 0.633 (0.043) | –0.249 (0.075) |
| Fam8 | 16.556 (0.338) | 4.111 (0.389) | 0.111 (0.111) | 0.725 (0.084) | 0.601 (0.043) | –0.187 (0.087) |
| Fam9 | 16.778 (0.147) | 5.222 (0.596) | 0.222 (0.147) | 0.850 (0.089) | 0.624 (0.051) | –0.317 (0.081) |
| Fam10 | 16.556 (0.176) | 4.556 (0.242) | 0.222 (0.147) | 0.814 (0.056) | 0.613 (0.034) | –0.331 (0.082) |
| Fam11 | 16.333 (0.441) | 5.333 (0.500) | 0.111 (0.111) | 0.766 (0.081) | 0.599 (0.057) | –0.268 (0.061) |
| Fam12 | 16.111 (0.423) | 5.000 (0.333) | 0.111 (0.111) | 0.836 (0.081) | 0.647 (0.049) | –0.270 (0.062) |
| Fam13 | 15.556 (0.930) | 4.444 (0.530) | 0.000 (0.000) | 0.788 (0.072) | 0.600 (0.051) | –0.307 (0.040) |
| FIS | FIT | FST | |
|---|---|---|---|
| Below all loci | −0.190 | −0.073 | 0.098 |
| p value | 1.000 | 1.000 | 0.001 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oliveira, A.M.; Ramos, S.L.F.; Ferreira, M.J.; Lopes, R.; Meneses, C.H.S.G.; Valente, M.S.F.; da Silva, R.F.; Batista, J.d.S.; Muniz, A.W.; Lopes, M.T.G. Mating System Analysis and Genetic Diversity of Parkia multijuga Benth. One Native Tree Species of the Amazon. Forests 2024, 15, 172. https://doi.org/10.3390/f15010172
Oliveira AM, Ramos SLF, Ferreira MJ, Lopes R, Meneses CHSG, Valente MSF, da Silva RF, Batista JdS, Muniz AW, Lopes MTG. Mating System Analysis and Genetic Diversity of Parkia multijuga Benth. One Native Tree Species of the Amazon. Forests. 2024; 15(1):172. https://doi.org/10.3390/f15010172
Chicago/Turabian StyleOliveira, Ariane Mendes, Santiago Linorio Ferreyra Ramos, Marciel José Ferreira, Ricardo Lopes, Carlos Henrique Salvino Gadelha Meneses, Magno Savio Ferreira Valente, Rogério Freire da Silva, Jacqueline da Silva Batista, Aleksander Westphal Muniz, and Maria Teresa Gomes Lopes. 2024. "Mating System Analysis and Genetic Diversity of Parkia multijuga Benth. One Native Tree Species of the Amazon" Forests 15, no. 1: 172. https://doi.org/10.3390/f15010172
APA StyleOliveira, A. M., Ramos, S. L. F., Ferreira, M. J., Lopes, R., Meneses, C. H. S. G., Valente, M. S. F., da Silva, R. F., Batista, J. d. S., Muniz, A. W., & Lopes, M. T. G. (2024). Mating System Analysis and Genetic Diversity of Parkia multijuga Benth. One Native Tree Species of the Amazon. Forests, 15(1), 172. https://doi.org/10.3390/f15010172








