Abstract
As for most other temperate broadleaved tree species, large-scale genetic inventories of pedunculate oak (Quercus robur L.) have focused on the plastidial genome, which showed the impact of post-glacial recolonization and manmade seed transfer. However, how have pollen mediated gene flow and introgression impacted the large-scale genetic structure? To answer these questions, we did a genetic inventory on 1970 pedunculate oak trees from 197 locations in 13 European countries. All samples were screened with a targeted sequencing approach on a set of 381 polymorphic loci (356 nuclear SNPs, 3 nuclear InDels, 17 chloroplast SNPs, and 5 mitochondrial SNPs). In a former analysis with additional 1763 putative Quercus petraea trees screened for the same gene markers we obtained estimates on the species admixture of all pedunculate oak trees. We identified 13 plastidial haplotypes, which showed a strong spatial pattern with a highly significant autocorrelation up to a range of 1250 km. Significant spatial genetic structure up to 1250 km was also observed at the nuclear loci. However, the differentiation at the nuclear gene markers was much lower compared to the organelle gene markers. The matrix of genetic distances among locations was partially correlated between nuclear and organelle genomes. Bayesian clustering analysis revealed the best fit to the data for a sub-division into two gene pools. One gene pool is dominating the west and the other is the most abundant in the east. The western gene pool was significantly influenced by introgression from Quercus petraea in the past. In Germany, we identified a contact zone of pedunculate oaks with different introgression intensity, likely resulting from different historical levels of introgression in glacial refugia or during postglacial recolonization. The main directions of postglacial recolonization were south to north and south to northwest in West and Central Europe, and for the eastern haplotypes also east to west in Central Europe. By contrast, the pollen mediated gene flow and introgression from Q. petraea modified the large-scale structure at the nuclear gene markers with significant west–east direction.
Keywords:
gene flow; haplotype; hybridization; pollen; spatial genetic autocorrelation; Quercus petraea; SNPs 1. Introduction
The genetic differentiation of the maternally inherited chloroplast genome of European white oak species has been intensively studied for more than 20 years [1,2,3]. The screening of chloroplast haplotype variation of sessile (Quercus petraea) and pedunculate oaks (Quercus robur) at the continental level has revealed a clear large-scale structure, which most probably resulted from postglacial recolonization [4]. During the last glacial maximum, pedunculate oaks survived in refugia in southern and south-eastern parts of Europe. The oaks were fixed to different haplotypes in these refugia, and the directions of recolonization can be reconstructed using the large-scale genetic structure of the haplotypes [4]. More precisely, one lineage from the Iberian Peninsula has colonized many parts of western Europe, a lineage from Italy has migrated north to many parts of Central Europe and other haplotype lineages have migrated from the Balkan-region to the north in Central and East Europe [4]. The haplotype migration history of pedunculate oak in the eastern part of the natural distribution range in Russia and neighboring countries is less studied. Recently two studies have shown that the area in Belarus, Ukraine and western Russia have been colonized by haplotypes from the Balkan region, whereas the eastern and south-eastern part up to the Ural Mountains is showing other haplotypes with an origin in the southern Ural and southern Russia [2,5]. In summary, the pattern of oak haplotypes in Europe resulted from south to north, south to north-west and also south to north-east seed migration routes. For several other temperate tree species as such as Fagus sylvatica, Fraxinus excelsior, Picea abies, Abies alba, and Pinus sylvestris similar recolonization routes as for white oaks have been found in Europe [6].
Other factors that have influenced the present haplotype pattern are the hybridization among different white oak species [7] and to a smaller extent a manmade gene flow by seed transfer [8,9].
In contrast to the abundant literature relying on the chloroplast genome, there are no large-scale surveys on the genetic differentiation of pedunculate oak for the nuclear genome. Most studies are focused on comparison between a few regional stands using nuclear microsatellites, such as comparisons between different stands in Russia [10], Ireland [7], Finland [11], Germany [12,13], Bulgaria, and Greece [9]. The few works with SNP-arrays for pedunculate oak have used either only a few individuals or individuals from a specific region or from a single country, and are addressing the question of species hybridizations or adaptation following candidate gene approaches [13,14,15] For other temperate tree species there are also only few comparative studies dealing with the large-scale genetic structure at gene markers of the chloroplast and nuclear genome. For example, Bai et al. [16] observed a weaker but still clear genetic structure using nuclear markers in contrast to a prominent phylogeographic structure with chloroplast markers in a Juglans species and Thomson et al. [17] studied nuclear microsatellites and chloroplast markers for several birch species in North America characterizing postglacial recolonization and level of introgression.
The large-scale geographic pattern of the nuclear genome is probably influenced by several population genetic processes in different ways: genetic drift and migration barriers are expected to create local departures from a large-scale pattern. The effects of hybridization and introgression on the gene pool of the European white oaks are well known [14,18] but have never been studied for their impact on the large-scale genetic pattern. Finally, selection should only shape a large-scale genetic pattern in the presence of local adaptation (genotype environment association). However, the last should only involve a part of the genome. We therefore expect stronger effects by gene flow. There are several studies showing the impact of limited pollen and seed dispersal of pedunculate oak on small scale genetic structure [19,20], but the effectiveness of long-distance gene flow, particularly by pollen, has been poorly studied so far [21,22].
The objective of this paper was to analyze the genetic differentiation and spatial genetic structure of pedunculate oak for a large number of trees from many locations of the natural distribution range, using both an array of nuclear SNPs and plastid SNPs [23]. In a first manuscript, we focused on the large-scale structure of Quercus robur in Russia and neighboring countries [5]. In the present paper, we merged this data with those from 100 additional locations in Central and West Europe. The pedunculate oak trees in this paper were also part of a study on species admixture using genotype data of 4848 Q. robur and Q. petraea trees (unpublished data). By combining genotype data of the nuclear and organelle genomes with data on the level of individual admixture we aim to identify the impact of gene flow by seeds and pollen as well as the influence of introgression on the range-wide genetic structure of Q. robur.
We expected, in contrast to the known strong spatial differentiation at the plastidial genome, only limited genetic differentiation at the nuclear SNPs, due to extensive long-distance gene flow by pollen leading to gene exchange among individuals of different plastidial lineages. The impact of introgression on the large-scale genetic structure of the nuclear genome is linked to several questions:
Was introgression from other species limited to a particular geographic location and historical time period, such as a single glacial refugium? In this case, we would expect additional genetic differentiation and a visible contact zone of pedunculate oaks with different introgression history. Or did introgression occur several times and independently at different places? In that case, we would not expect any additional genetic differentiation of the pedunculate oak populations.
2. Materials and Methods
2.1. Sampling
We collected samples from 1970 pedunculate oak trees in 197 locations (10 individuals at each location) in 13 European countries (Figure 1, Supplementary S1). The majority of the samples (188 locations) were collected in forest stands, whereas the others (nine locations) were taken from provenance trials. To collect mature and unrelated individuals, we defined a minimum diameter at breast height of 20 cm and a minimum distance of 50 m between trees with exception of location 503 where only saplings could be found. The majority of the samples came from Germany and Russia due to the focus of the projects on those regions. The pedunculate oak samples of this study were part of a broader collection of 4848 pedunculate and sessile oaks in Europe. The species admixture of all 4848 samples was checked based on the genetic data with a principal component analysis and Bayesian clustering using the software STRUCTURE (Supplementary S2). All 1970 samples of this study had an admixture proportion of the Q. robur gene pool of at least 80%.
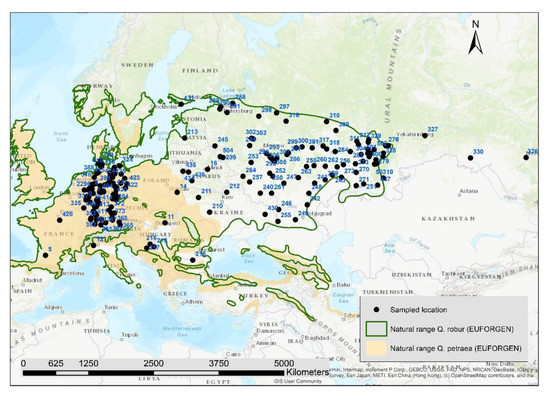
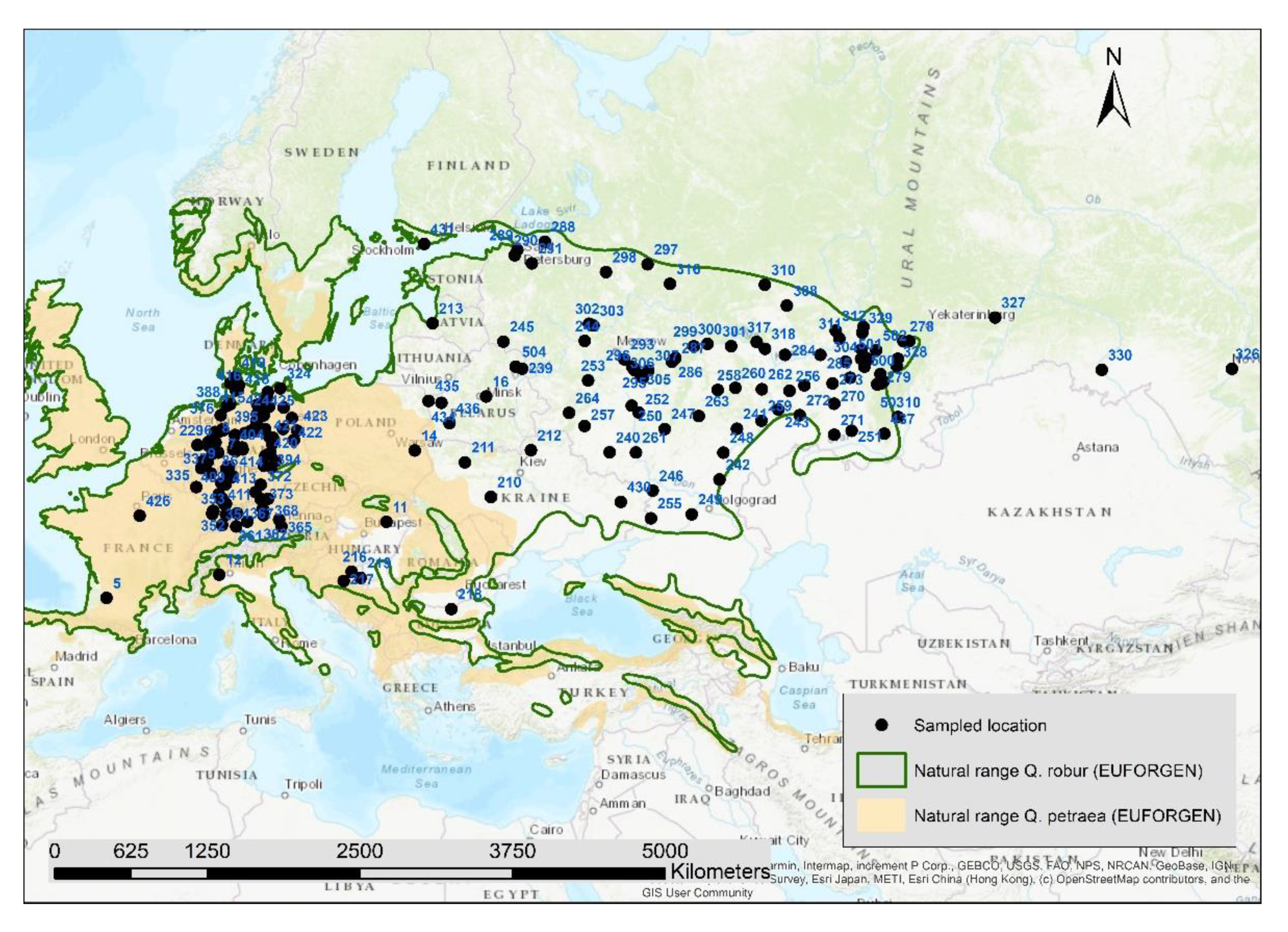
Figure 1.
Position of the 197 sampled locations, the numbers are the location-IDs in Table S1 (Supplementary), natural ranges of Quercus robur and Quercus petraea according to EUFORGEN.
2.2. DNA Extraction and Genotyping
The DNA was extracted following a slightly modified protocol of Dumolin et al. [24]. For all samples, 359 polymorphic nuclear loci (356 SNPs, 3 InDels), 17 polymorphic chloroplast SNPs (cp-SNPs), and five mitochondrial SNPs (mt-SNPs) were analyzed using targeted genotyping by sequencing. The gene markers were initially developed using 95 Q. robur and Q. petraea individuals sampled over the species’ ranges [23].
Sequencing was conducted by LGC Genomics GmbH with Illumina NextSeq, using single primer enrichment technology producing 75 bp single reads at 200x average coverage per sample. Reads were trimmed for adapter sequences and quality. Reads with average phred score of <30 over a window of 10 bases were trimmed. Reads shorter than 65 bp or reads containing Ns were discarded. Read alignments against self-assembled contigs were created with Bowtie2 v2.2.3 [25]. Variant calling was done with Freebayes v1.3.2 [26]. Nuclear variants were called with ploidy = 2 and organelle variants were called with ploidy = 1. Genotypes were filtered for a minimum coverage of 3 reads with vcftools [27]. All 1970 individuals had complete data for at least 90% of the loci. The proportion of missing data for each locus over all individuals was less than 5%.
2.3. Statistical Analysis
2.3.1. Haplotypes
We tested if there was any recombination between the 17 chloroplast SNPs and the five mitochondrial SNPs. For that, we pooled all 1970 individuals to one single population and computed the p-values for linkage equilibrium using a probability test implemented in the program GenPop [28]. This test checks for association between haplotypes at two loci, also called a test of the composite linkage disequilibrium. There was linkage (p < 0.000001) between all 22 organelle loci. Thus, we considered the combination of alleles at the 22 plastid loci as multilocus haplotypes, called just “haplotypes” in the following text.
We used the DNA-sequences of the 17 chloroplast SNPs and the five mitochondrial SNPs to construct a haplotype network with the minimum spanning method [29] as implemented in the software POPART [30]. The haplotype network was aimed to give us insights into the phylogenetic relationship of the different haplotypes.
2.3.2. Genetic Differentiation
Genetic distance (D0) was calculated using the allele frequencies at the 359 nuclear gene loci and the frequencies of haplotypes to quantify the genetic differentiation between pairs of locations [31]:
where i and j represent two sampled locations, n is the number of alleles or haplotypes, pik is the relative frequency of the k-th allele or haplotype.
Further, we calculated delta [32] and Wright’s FST [33], as measures of genetic differentiation and fixation among populations. The genetic differentiation delta was computed as
where j is the sampled location, pi is the frequency of allele i at locus k or frequency of haplotype i and is the frequency at allele i or haplotype i of all other locations (complement).
By shifting individual genotypes among the sampled locations, we estimated the statistical significance of delta, FST and D0 with 1000 permutations. The calculations were done with the computer program GDA_NT 2021 [34].
2.3.3. Bayesian Clustering Analysis
We used the Bayesian clustering method implemented in the software STRUCTURE v.2.3.4, Stanford, California, USA [35] to check the number of genetic groups present in the 197 sampled locations. The model assumption was that the involved loci are not linked and that they are in Hardy–Weinberg Equilibrium. We included all 359 nuclear gene markers in this analysis. We set the length of burn-in and Markov chain Monte Carlo simulations to 15,000 and 100,000, respectively, and tested K values from 1 to 7 each 20 times. We ran the admixture model with correlated allele frequencies. Then, we computed delta K values to find the optimal number of genetic clusters in the data [36]. For each tested K value, data from the 20 STRUCTURE runs were summarized and graphically presented to obtain the final Q values for each individual with the programs CLUMPP v.1.1.2 [37] and CLUMPAK [38].
2.3.4. Spatial Structure and Maps
We studied the spatial structure of allele frequencies at the nuclear gene markers and the frequencies of multilocus haplotypes using the software SGS [39,40]. The genetic distance (Do) was used for the genotypes and haplotypes to test spatial patterns in different geographic distance classes. By shifting the geographic co-ordinates of the sampled locations, the statistical significance was tested by 1000 permutations. The maps were drawn with ArcGIS 10.8 (Esri).
3. Results
3.1. Haplotypes
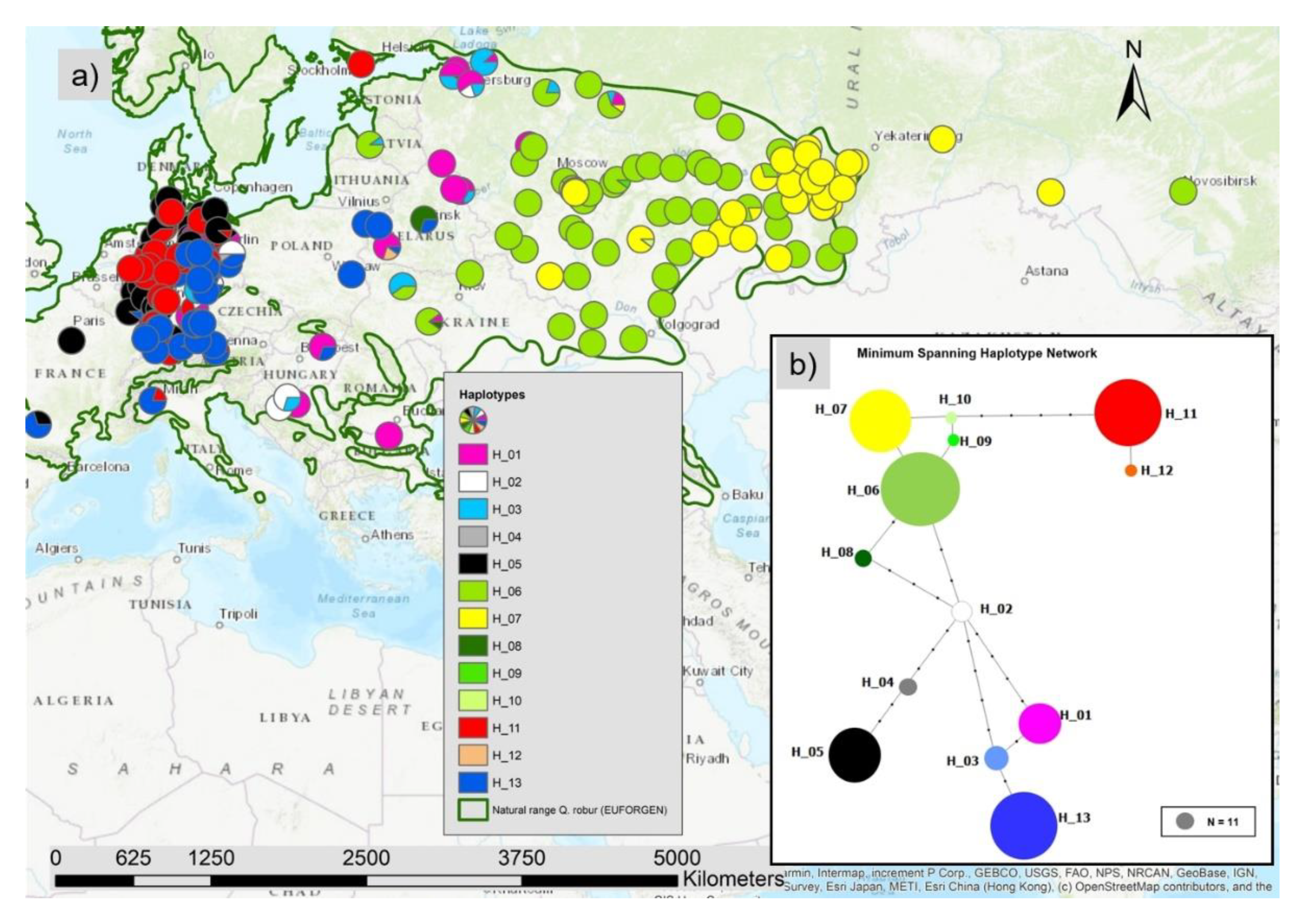
We observed 13 different haplotypes in all 1970 individuals based on the 17 cp-SNPs and the five mt-SNPs with a clear spatial pattern (Figure 2a). The haplotype network (Figure 2b) showed different lineages: an eastern lineage with the green and yellow marked haplotypes (H06, H07; H08, H09, H10), the lineage including the blue and pink colored haplotypes H01, H03 and H13, the lineage with the black (H05) and grey (H04) haplotype, and the lineage with the red haplotypes (H11, H12). The yellow haplotype H07 is dominating the east of Russia and the green haplotype H06 was most frequent in the central and western parts of the species distribution range in Russia including a part of Ukraine. In the middle belt including the Balkan and Baltic countries as well as the north-west of Russia the haplotypes H01 and H03 were more common. In eastern Poland and southern and south-east Germany, haplotype H13 was the most frequent one (Figure 7a). In West and North Germany and in part of France, haplotype H5 was frequent. In West and North Germany large areas were dominated by the genetically quite different haplotype H11 (Figure 2a and Figure 7a).
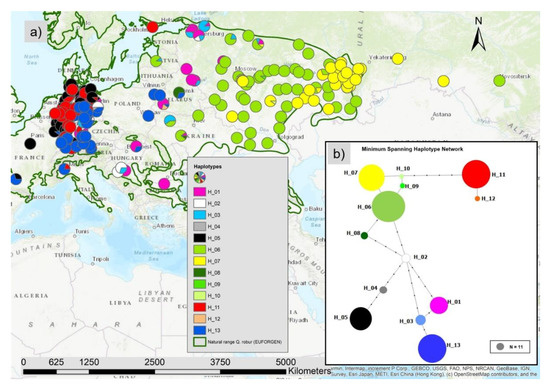
Figure 2.
(a) Map with the spatial distribution of the 13 different haplotypes and (b) haplotype network constructed with the minimum spanning method (small black dots represent the number of mutations, the diameter of the colored dots is proportional to the absolute number of sampled individuals with the haplotype).
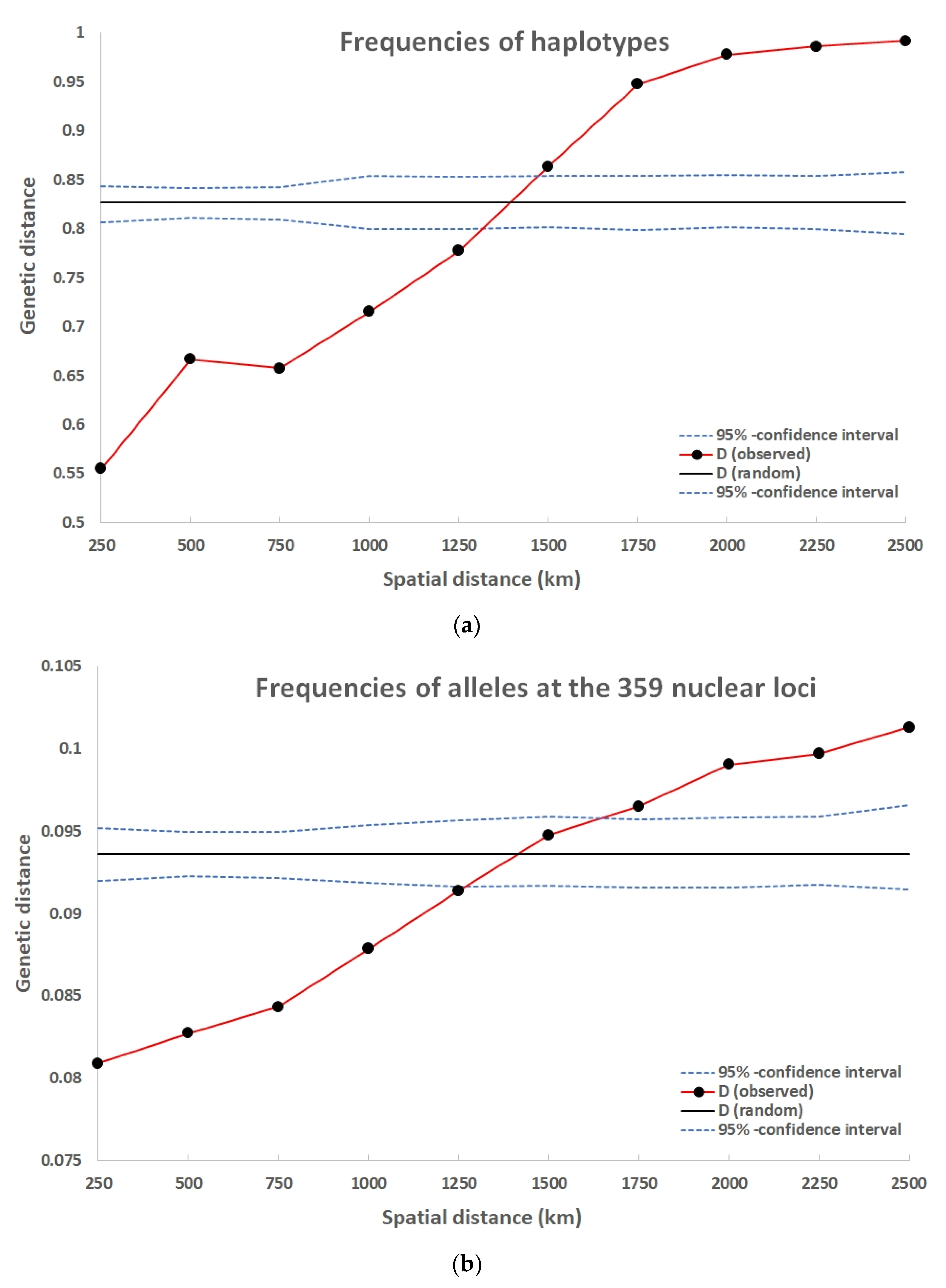
In a range up to 1250 km the genetic distance among the frequencies of haplotypes at the locations were significantly smaller than expected for a random spatial distribution (Figure 3a).
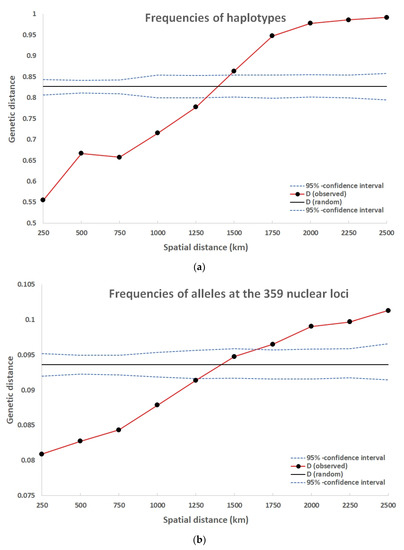
Figure 3.
Distograms of the spatial pattern for (a) frequencies of haplotypes and (b) frequencies of alleles at the 359 nuclear loci.
We calculated the frequencies of haplotypes at each location. Based on these haplotype frequencies, we computed the genetic distances among locations, delta and FST. The genetic distances D0 had an arithmetic mean of 0.83 and varied between 0.00 and 1.00. The population fixation FST for the haplotypes was 0.83 and delta 0.78. Both values were highly significant (p < 0.001).
3.2. Nuclear Markers
We computed the allele frequencies at the 359 nuclear gene loci at each of the 197 locations. Based on these allele frequencies we calculated the genetic distances D0 among locations, delta and FST. The permutation tests of the gene pool distances among all 19,306 pairs of locations were statistically significant (p < 0.05) in 87% of all cases. The gene pool distances had an arithmetic mean of 0.094 and varied between 0.062 and 0.159.
Up to a range of 1250 km, the gene pool distances among locations were smaller than expected for a random pattern (Figure 3b). This illustrates the range of a positive spatial autocorrelation. The population fixation over all loci FST was 0.077 and delta 0.069. Both values were highly significant (p < 0.001).
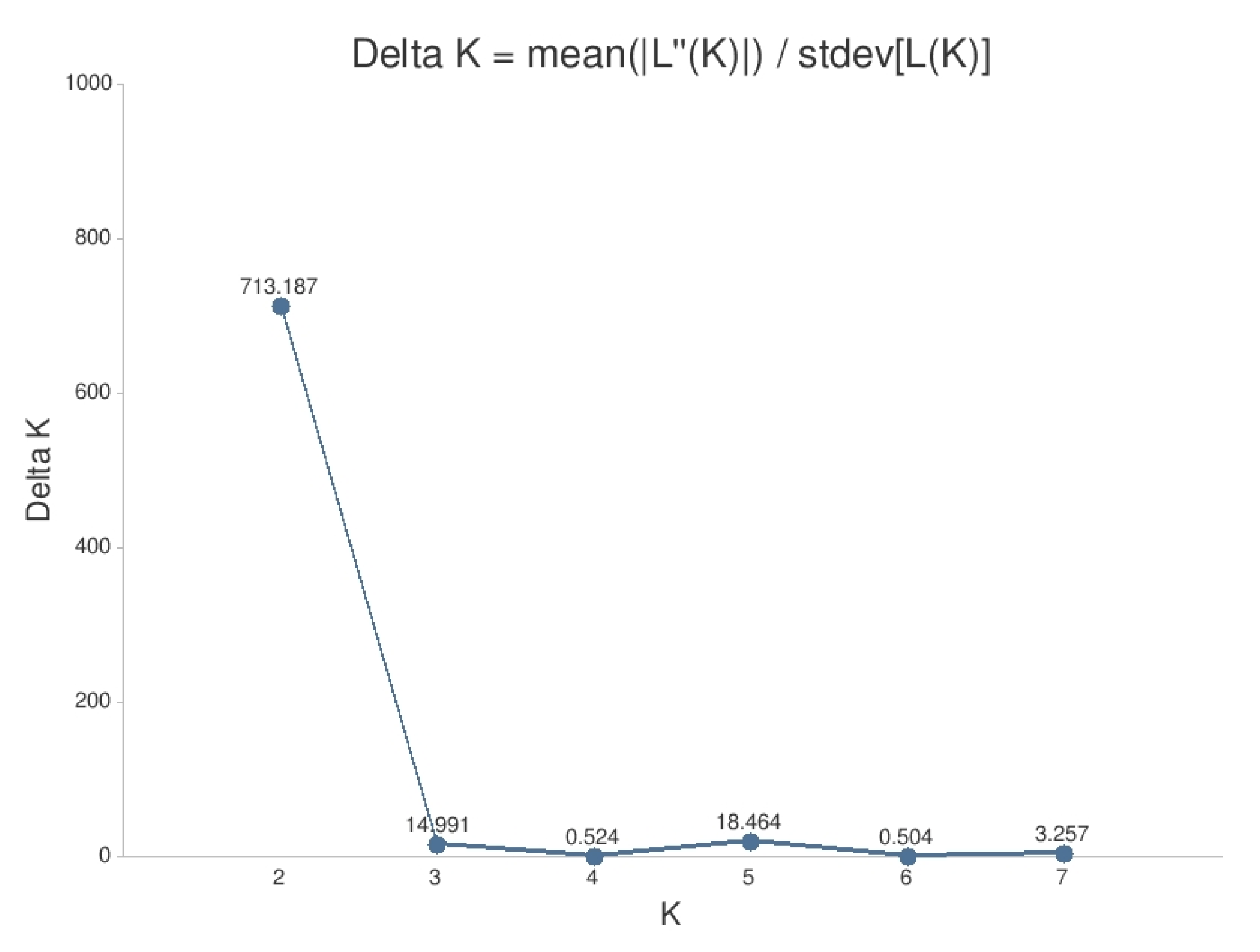
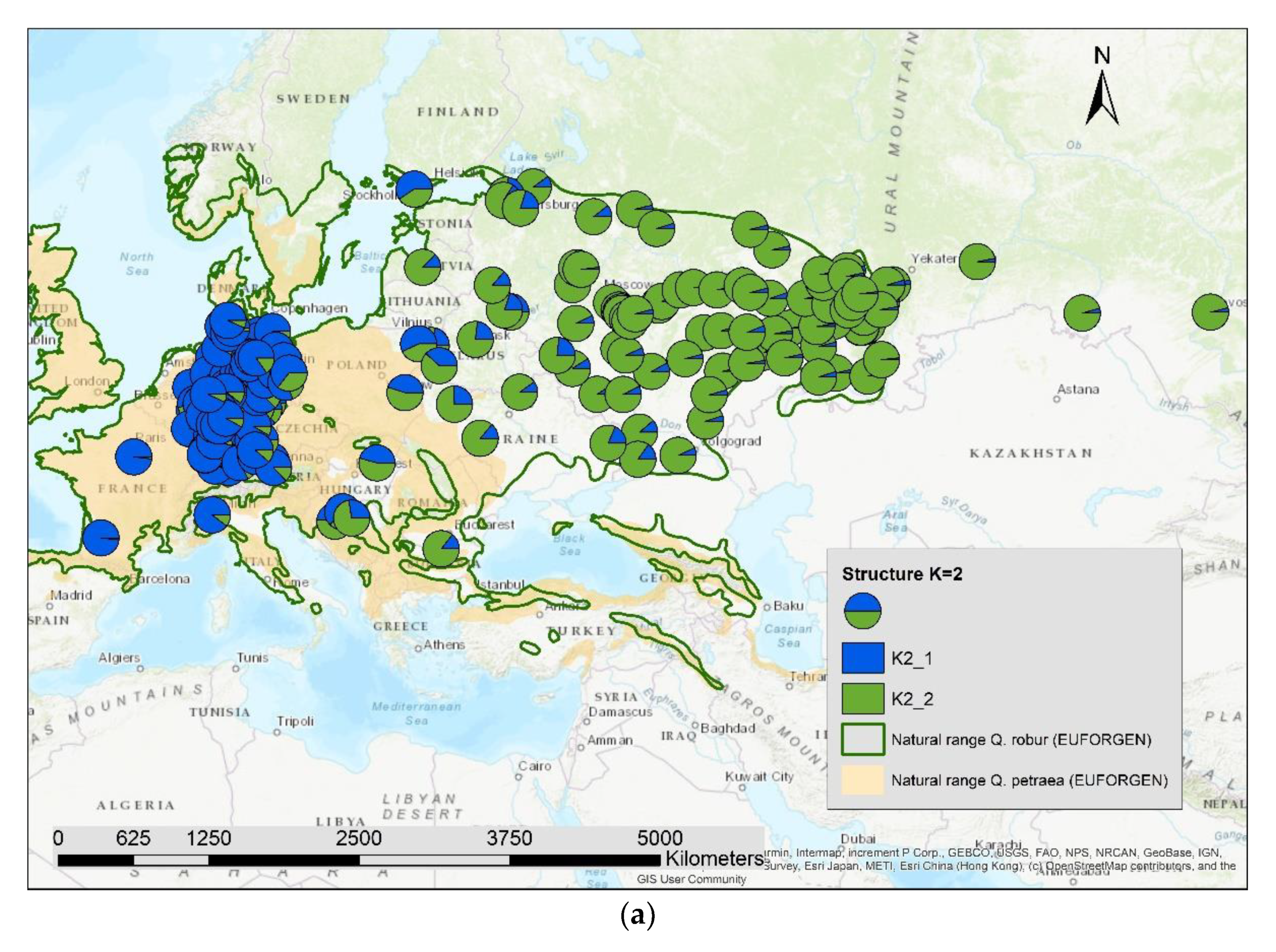
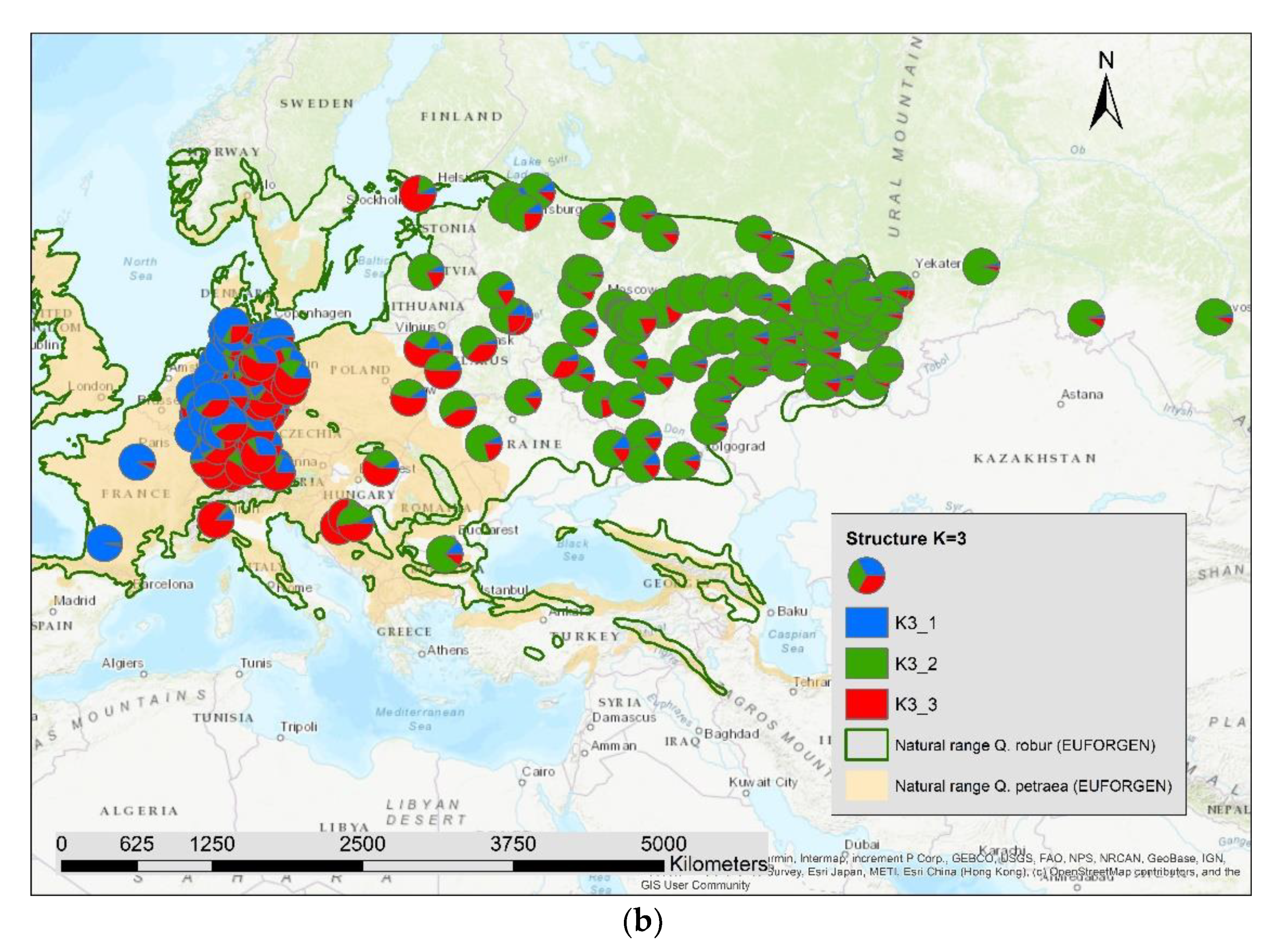
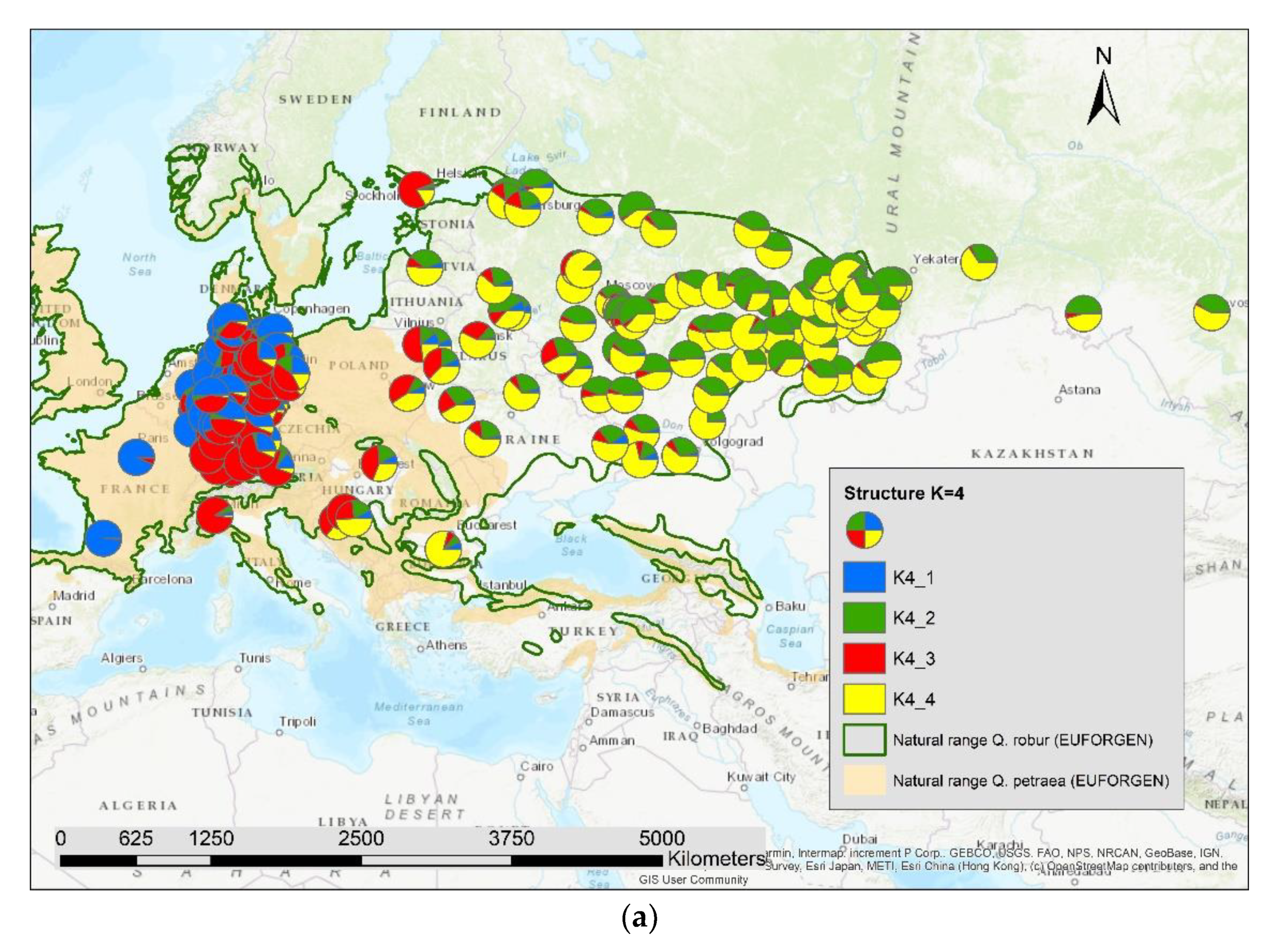
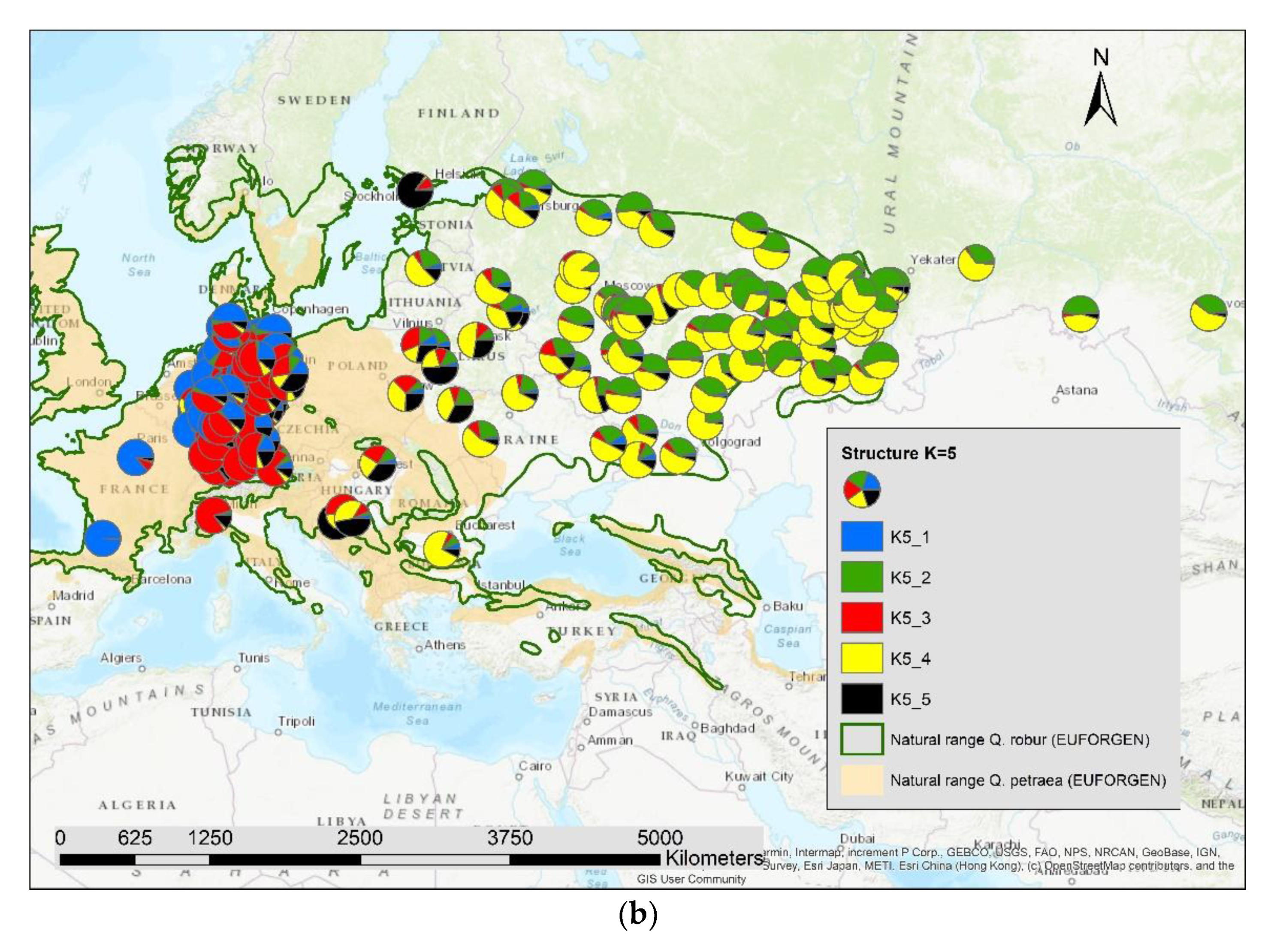
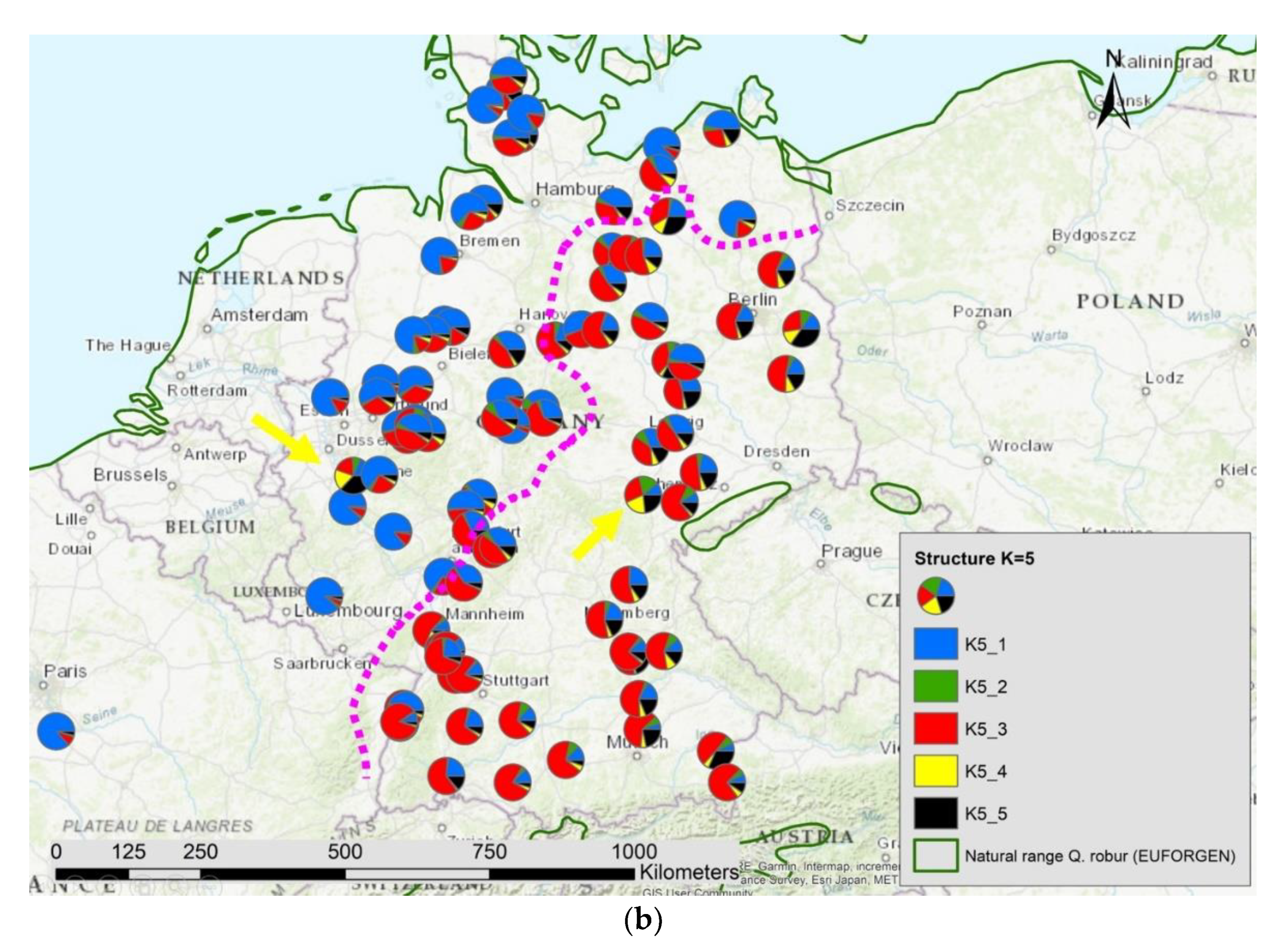
The results of the STRUCTURE analysis showed that based on delta K [36], the best representation of the data is achieved if two groups are considered and there was a second much smaller peak for K = 5 (Figure 4). With K = 2, we observed a clear geographic pattern with one gene pool dominating the west (blue color in Figure 5a) and another gene pool dominating the east (green color). There was an area including parts of Germany, Poland, the Balkan and Baltic states, Belarus, and Ukraine with locations that had a mixture of both gene pools. For K = 3 (Figure 5b) there was still a gene pool frequent in the west (blue color) and in the east (green color) but there appeared a third gene pool (red color) most frequent in the area between those two groups. With four gene pools (K = 4, Figure 6a), the eastern gene pool was further subdivided into two gene pools (green + yellow color). Finally, with five gene pools (K = 5, Figure 6b and Figure 7b) an additional gene pool (black color) was visible in the central area (East Germany, Balkan, Poland, Baltic States, Belarus, Ukraine). Within Germany we observed two zones: the north-west with higher frequency of K1 and the south and north-east with K3 as the most abundant gene pool (Figure 7b).
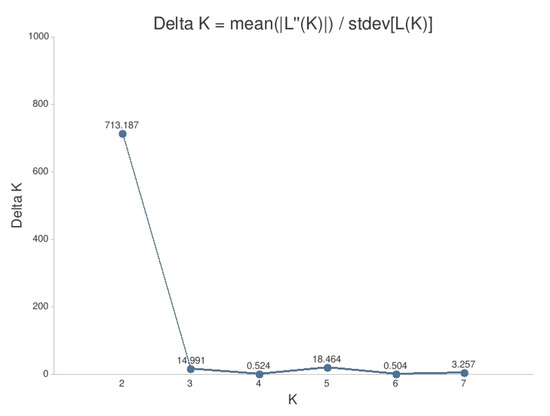
Figure 4.
Delta K values for the results of the analysis with STRUCTURE.
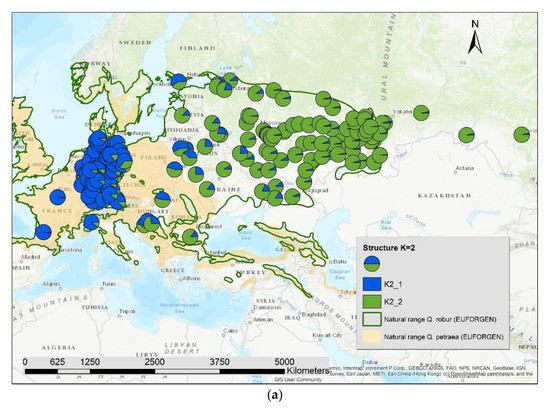
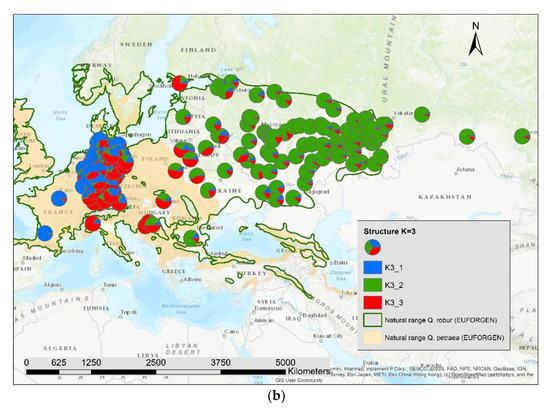
Figure 5.
Spatial distribution of the results of STRUCTURE for (a) K = 2 and (b) K = 3.
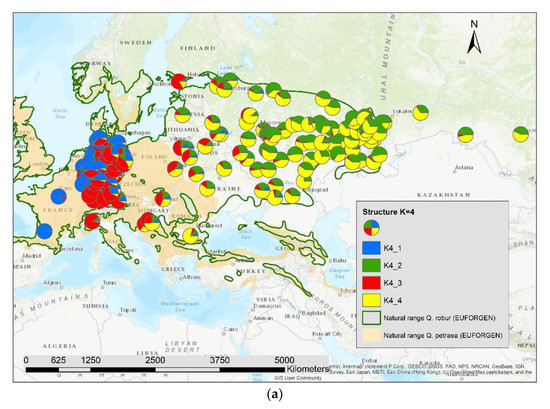
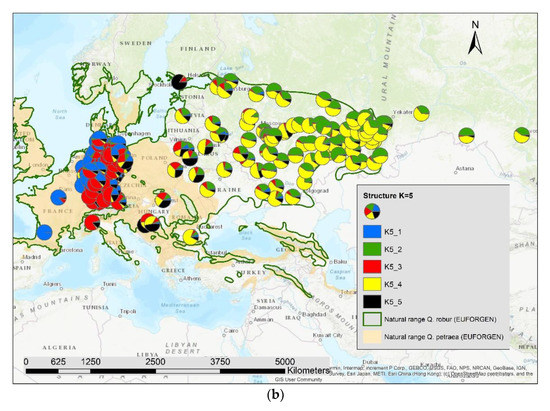
Figure 6.
Spatial distribution of the results of STRUCTURE for (a) K = 4 and (b) K = 5.
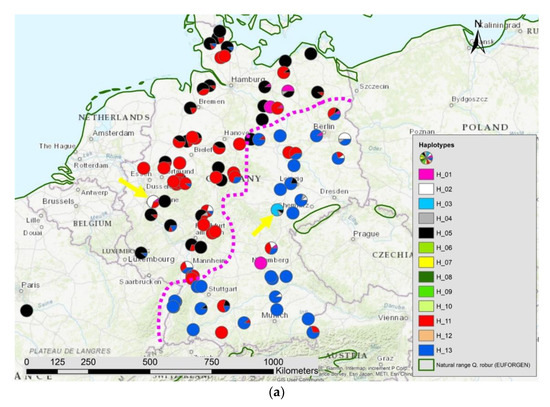
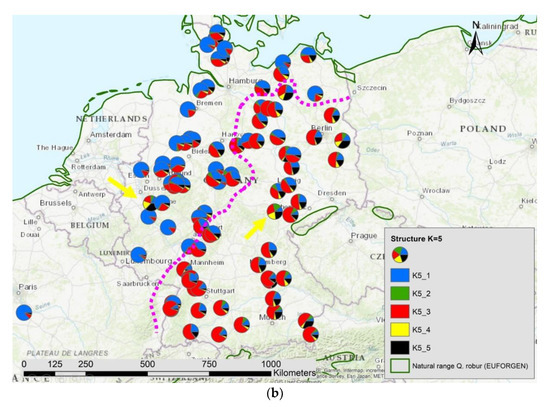
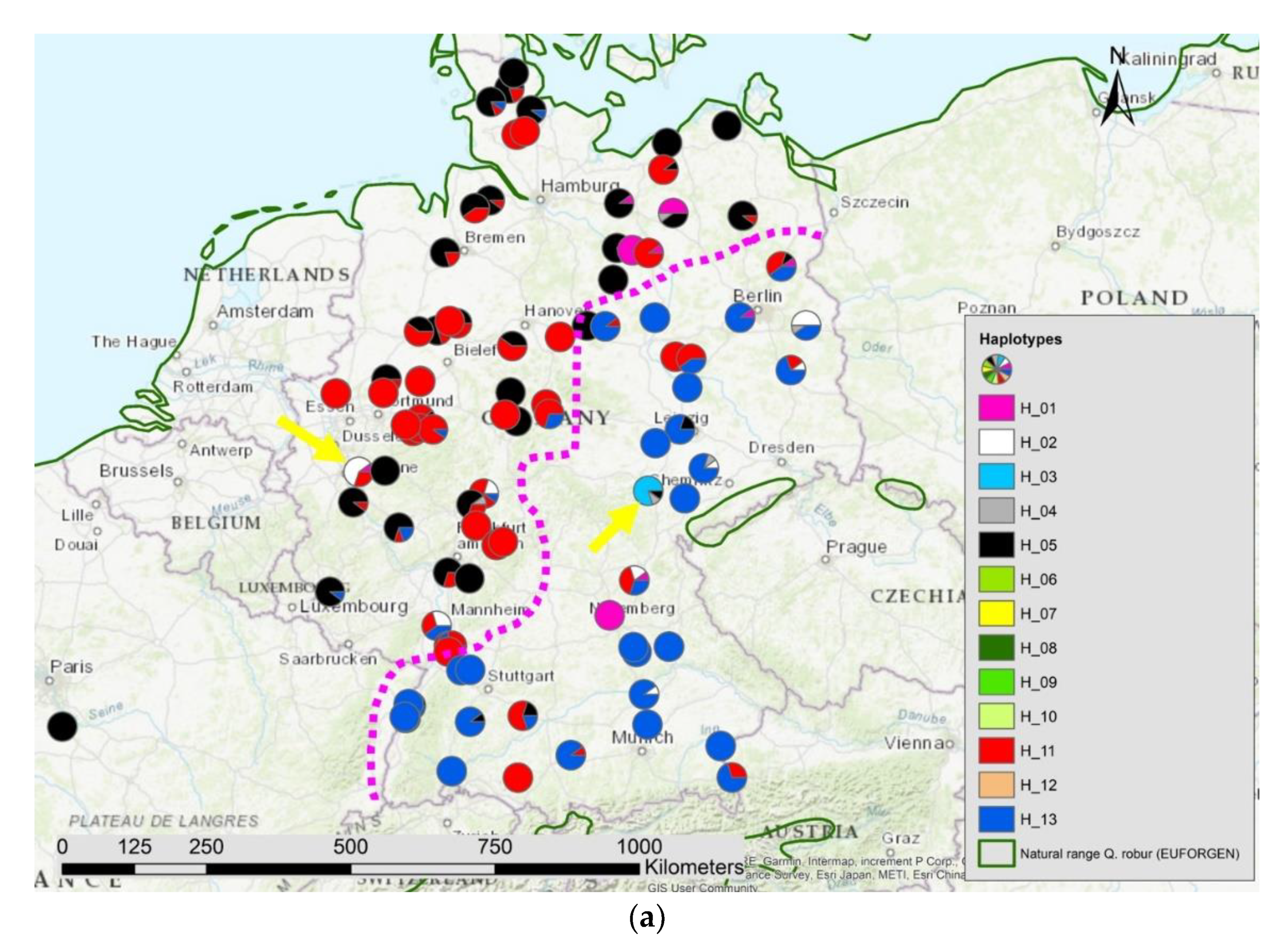
Figure 7.
Spatial distribution of (a) haplotypes and (b) the results of STRUCTURE K = 5 in Germany; the pink dotted line indicates the contact zone of different haplotypes and nuclear gene pools; the yellow arrows point to two locations with manmade gene flow.
We computed the Pearson correlation coefficient between the different genetic groups for K2 to K5 and the admixture proportion of Quercus petraea (Supplementary S2) for all 197 locations and observed a significant positive correlation (p < 0.001) with the first genetic group indicated as blue color in Figure 5a,b, Figure 6a,b and Figure 7b: r = 0.51 for K2_1, r = 0.72 for K3_1, r = 0.77 for K4_1, and r = 0.81 for K5_1.
3.3. Genetic Correlation among Nuclear and Organelle Genome
We observed a significant positive Pearson’s correlation coefficient of r = 0.37 (p < 0.0001) among the pairs of gene pool distances at the 359 nuclear gene markers and the genetic distances of the haplotypes. In Germany, we found a similar pattern of genetic differentiation for both types of genomes with a nearly identical contact zone (Figure 7a,b). However, due to the high level of haplotype diversity, the correlation coefficient among the matrix of genetic distances of nuclear gene pools and haplotypes including only locations in Germany had a correlation of only r = 0.15 (p < 0.001).
4. Discussion
4.1. Impact of Seed and Pollen Mediated Gene Flow
For both the nuclear and organelle SNPs a positive spatial autocorrelation up to 1250 km was observed. The strength of genetic differentiation was much larger for the haplotypes. The distribution pattern of haplotypes is best explained by recolonization after the last glacial period from refugia in the Iberian Peninsula, Italy, the Balkan region, southern Russia, and the southern Ural region [1,2,4]. The locations of refugia and the postglacial recolonization history of white oaks in Europe has also been confirmed by fossil pollen profiles [41,42,43]. In Central Europe, different haplotype lineages came together and increased the mixture and diversity [8]. This was also the case for several other tree species [44].
Thus, main directions of this gene flow by seeds were south to north-east in West and Central Europe and south-north and east-west in East Europe. Besides this large-scale pattern, also the influence of manmade seed transfer is visible in Central Europe on a local scale [8,9]. Therefore, it is known that pedunculate oaks have been introduced to western Germany on purpose from Slavonia and this seed transfer is still visible by different haplotypes compared to the oak stands in the neighborhood [45]. However, so far, the impact of this long-distance manmade seed transfer on the regional and large scale spatial genetic structure of pedunculate oak is limited (see two examples marked with arrows in the maps of Figure 7a,b). The limited impact might be explained by the difficulties for humans to store oak acorns for longer time and the weight of acorns. Both were factors influencing transport distances in the past, especially in the time before railway carrying was an option. In contrast, for conifers with small seeds that can be stored for several years the impact of manmade seed transfer all over Europe is much more pronounced [46].
For oaks with their heavy acorns, natural seed dispersal by wind and to a smaller extent by birds, especially the Eurasian jay, is limited. Wind dispersal of seeds is below 50 m in most cases [47] and the jay can transport seeds up to a few hundred meters and in rare cases a few kilometers [48]. Apparently, the capacities of seed dispersal were not strong enough during the time since recolonization to create a stronger mixture of haplotypes. We observed a mixture of haplotypes where different recolonization pathways came together but no further mixture occurred in the way that eastern haplotypes were present far in the west and vice versa.
In contrast to the gene flow by seeds, we expected a gene flow by pollen over much larger distances due to wind-dispersal. Most experimental studies on pollen-mediated gene flow were done on rather local scales. The effective pollen dispersal was often less than 100 m and in rare cases up to a few hundred meters [49,50]. There are, however, contrasting findings on the importance of long-distance gene flow by pollen of pedunculate oaks. Buschbom, Yanbaev and Degen [21] found, in a small isolated population of Q. robur in the southern Urals, that about 30% of the effective pollen came from a distance of at least 80 km. Additionally, Gerber, et al. [51] demonstrated relevant proportions of successful pollen in oaks coming from outside the studied stands. Moracho et al. [52], on the other hand, found little evidence for gene flow from outside the local stand in fragmented pedunculate oak populations in Spain. Several factors influence the impact of external long-distance pollen [22,53]: the synchronization of flowering, abundance of local pollen compared to external pollen, viability of pollen in relation to UV radiation, wind direction and strength. Some of these factors have been considered in a modeling approach on dispersal of oak pollen by Schueler and Schlunzen [54]. They found that physical pollen dispersal up to 100 km is possible, but to which extent this pollen could be part of fertilization was still unclear.
For the nuclear genome of the oaks which is dispersed by pollen and seeds we would have expected either no or only a relatively weak spatial autocorrelation on larger scales [55,56]. Nevertheless, we found spatial autocorrelation on the same scale up to 1250 km as for the maternal haplotypes. The genetic differentiation at the nuclear gene loci was much weaker compared to the organelle gene loci which can be interpreted as an effect of the stronger gene flow by pollen. The weak positive correlation of 0.37 among the matrix of gene pool distances at nuclear and organelle gene markers underlines that at least partly different forces or different geographic directions of gene flow are responsible for the observed genetic differentiation of the two parts of the genome.
The STRUCTURE analysis with an aggregation of the data at the nuclear gene loci to different gene pools did show a higher mixture of gene pools compared to the distribution pattern of haplotypes. In contrast to the main direction of recolonization (south–north) there was evidence of an intensive pollen-mediated gene flow from east to west and west to east. Nevertheless, there was still a clear isolation by distance effect [57] that produced the significant spatial autocorrelation at the nuclear gene loci and that enables the detection of different gene pools with STRUCTURE. Additionally, the relatively sharp contact zone of different nuclear gene pools in Germany (Figure 7b) raises questions on the effectiveness of long-distance pollen-mediated gene flow.
4.2. Hybridization and Introgression
Hybridization among Q. robur and Q. petraea occurs with low frequency in mixed oak stands [18,58]. Hybridization leads to introgression if the interspecific gene flow confers an adaptive advantage or is selectively neutral at least [14,59]. For the maternal haplotypes, it is well established that the same haplotypes can be found in the same regions in different white oak species, especially Q. robur and Q. petraea, and this is explained by a certain level of hybridization among related oak species [60], which could even be asymmetric [61]. The impacts of hybridization and introgression on shared haplotypes have also been reported for other deciduous tree species such as Betula in North America [62].
The natural distribution range of pedunculate and sessile oak is different: Quercus robur has a distribution a few thousand km more to the east up to the Ural Mountains (Figure 1). Hence, if hybridization among these two oak species is shaping the gene pool, it should not be the same in all regions but mainly plays a role in West and Central Europe.
Using genetic data of 4848 sessile and pedunculate oaks from the whole species distribution ranges, we estimated the admixture level of all trees involved in this study (Supplementary S2). We only included in the present paper those Q. robur trees with an admixture of Q. petraea of less than 20%. The analysis with STRUCTURE showed that parts of the western gene pool of the pedunculate oaks were correlated with the admixture level of Q. petraea (Supplementary S2). Especially in Germany, we found a sharp border between pedunculate oaks in the north-west influenced by this introgression and pedunculate oaks in the south-west and east composed by other gene pools with no sign of introgression. This sharp border was also found in respect to the composition of haplotypes. We conclude from the similarity of these contact zone (Figure 7a,b) that the present day spatial genetic structure of nuclear genetic information still has footprints of postglacial recolonization. The introgression happened either in the refugia or during the recolonization phase [63]. West and north-west Germany have been colonized from the Iberian-lineage that has still signs of introgression, whereas the south-east of Germany has been colonized by oaks from another refugia lineage (refugia from Italy and the Balkan region) that did not experience the same level of introgression from Q. petraea. Gene flow by pollen was not strong enough to equalize these genetic differences. Thus, we conclude that the introgression occurred in the past in the west and migrated in direction north-east.
For other broadleaved tree species, e.g., Betula alleghaniensis and Betula papyrifera, in North America also large-scale patterns of different levels of introgression have been observed [17]. This study on the Betula species showed how ongoing hybridization could impact the distribution of haplotypes and obliterate the large-scale genetic structure resulting from recolonization from glacial refugia. In contrast, our results showed that historical introgression could even strengthen the large-scale genetic pattern caused by recolonization from glacial refugia.
One factor influencing the level of hybridization is simply the level of mixture and abundance of both oak species [58]. In this aspect, we would had expected higher ratios of hybridization in the south-west of Germany. However, this region is part of the area with a low level of introgression further emphasizing the importance of historic rather than current hybridization events shaping large-scale genetic patterns.
4.3. Genetic Selection
Can genetic selection explain the large-scale pattern of the nuclear gene loci? First, it should be noted that we did not follow a candidate gene approach for the gene marker development as in other studies [15,55]. Moreover, our selection of candidate SNPs in the nuclear genome was determined by the occurrence of restriction sites in the ddRAD approach [23]. In a blast search of all probe sequences including our SNPs against the reference genome of Q. robur, we estimated a uniform genome-wide genetic marker distribution [64]. In principle, many of the identified markers might also be adaptive as we found about half of all SNPs to be located within gene annotations. It is unlikely, however, that many of the studied SNPs are under selection for traits that are linked to large-scale environmental gradients. Future studies with whole genome inventories may clarify the impact of adaptation on the genetic structure.
5. Conclusions
Pedunculate oak exhibits marked genetic patterns on the European continental scale for both the nuclear and the organelle genomes. The pattern of the organelle DNA is still reflecting the impact of post-glacial recolonization from different refugia in the south. Central Europe is a region with a high level of haplotype diversity because of the merging of different recolonization lineages. Recent gene flow by seeds did not erase this historical recolonization pattern of haplotypes and, so far, also manmade seed transfer had only local influence on the spatial genetic pattern. The intensity of historical introgression of Q. petraea into the gene pool of Q. robur was not the same for all recolonization lineages and thus a part of the large-scale genetic structure for the nuclear genome can be explained by differences of historical introgression. This added an intraspecific gene flow from west to east. In addition, the pollen-mediated gene flow modified the large-scale structure at the nuclear gene markers with distinct east–west and west–east gradients leading to increased admixture of the gene pools.
Supplementary Materials
The following are available online at https://www.mdpi.com/article/10.3390/f12101425/s1, Table S1: Information on sampled locations; S2: Description of genetic identification of Quercus robur, Quercus petraea and hybrids.
Author Contributions
Conceptualization and sample design, B.D. and Y.Y.; sampling, Y.Y., R.I., S.B., C.B.-J. and H.S.; gene marker development and bioinformatics, M.M., C.B.-J. and H.S.; lab work, R.I. and S.B., population genetic data analysis, B.D.; writing—original draft preparation, B.D.; writing—review and editing, Y.Y., M.M., R.I., S.B., H.S. and C.B.-J.; project administration and funding acquisition, B.D. and Y.Y. All authors have read and agreed to the published version of the manuscript.
Funding
The study was funded through the grant No 19-16-00084 from the Russian Science Foundation (project members: B. Degen, Y. Yanbaev, R. Ianbaev and S. Bakhtina) and by the grant WKF-WF04-22WC4111 01 from the German Federal Ministry of Food and Agriculture and German Federal German Ministry for the Environment, Nature Conservation and Nuclear Safety.
Data Availability Statement
The genotype data including geographic co-ordinates have been uploaded at DYRAD: https://doi:10.5061/dryad.8931zcrrd (accessed on 18 October 2021).
Acknowledgments
We thank Alwin Janßen, Jörg Kleinschmit, Ralf Kätzel, Dagmar Schneck, Wilfried Steiner, Bernd Rose, and Hans-Peter Ehrhart for supporting the sampling and selection of oak stands in Germany and we thank Antoine Kremer, Jaroslaw Burczyk and Giovanni Giuseppe Vendramin for providing samples from France, Poland and Italy collected in frame of the EU FORGER project. We are also thankful to former Evoltree members providing us with additional oak material from France, Hungary, Switzerland, and Finland and to our colleague Muhidin Seho for material from Bulgaria, Bosnia and Herzegovina, and Croatia. We are grateful to Stefan Jencsik, Petra Hoffmann and Susanne Hoppe for technical assistance with laboratory work and Niels Müller for helpful comments and suggestions on a former version of the manuscript.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Petit, R.J.; Csaikl, U.M.; Bordács, S.; Burg, K.; Coart, E.; Cottrell, J.; van Dam, B.; Deans, J.D.; Lapegue, S.; Fineschi, S.; et al. Chloroplast DNA variation in European white oaks-Phylogeography and patterns of diversity based on data from over 2600 populations. For. Ecol. Manag. 2002, 156, 5–26. [Google Scholar] [CrossRef]
- Semerikova, S.A.; Isakov, I.Y.; Semerikov, V.L. Chloroplast DNA variation and phylogeography of pedunculate oak Quercus robur L. in the eastern part of the range. Russ. J. Genet. 2021, 57, 47–60. [Google Scholar] [CrossRef]
- Deguilloux, M.-F.; Pemonge, M.-H.; Petit, R.J. Use of chloroplast microsatellites to differentiate oak populations. Ann. For. Sci. 2004, 61, 825–830. [Google Scholar] [CrossRef] [Green Version]
- Petit, R.J.; Brewer, S.; Sándorbordácsde, S.; Burge, K.; Cheddadibc, R.; Coartf, E.; Cottrellg, J.; Csaikl, U.M.; van Dam, B.; Deans, J.D.; et al. Identification of refugia and post-glacial colonisation routes of European white oaks based on chloroplast DNA and fossil pollen evidence. For. Ecol. Manag. 2002, 156, 49–74. [Google Scholar] [CrossRef]
- Degen, B.; Thuenen-Institute of Forest Genetics, Grosshansdorf, Germany; Yanbaev, Y.; Bashkir State Agrarian University, Ufa, Russia; Ianbaev, R.; Bashkir State Agrarian University, Ufa, Russia; Blanc-Jolivet, C.; Thuenen-Institute of Forest Genetics, Grosshansdorf, Germany; Mader, M.; Thuenen-Institute of Forest Genetics, Grosshansdorf, Germany; Bakhtina, S.; Bashkir State Agrarian University, Ufa, Russia. Targeted sequencing reveals large scale genetic structure of Quercus robur in Russia and neighbouring countries. Personal communication, 2021. [Google Scholar]
- Gomory, D.; Zhelev, P.; Brus, R. The Balkans: A genetic hotspot but not a universal colonization source for trees. Plant. Syst. Evol. 2020, 306, 9. [Google Scholar] [CrossRef]
- Beatty, G.E.; Montgomery, W.I.; Spaans, F.; Tosh, D.G.; Provan, J. Pure species in a continuum of genetic and morphological variation: Sympatric oaks at the edge of their range. Ann. Bot. 2016, 117, 541–549. [Google Scholar] [CrossRef] [Green Version]
- Konig, A.O.; Ziegenhagen, B.; van Dam, B.C.; Csaikl, U.M.; Coart, E.; Degen, B.; Burg, K.; de Vries, S.M.G.; Petit, R.J. Chloroplast DNA variation of oaks in western Central Europe and genetic consequences of human influences. For. Ecol. Manag. 2002, 156, 147–166. [Google Scholar] [CrossRef]
- Neophytou, C.; Aravanopoulos, F.A.; Fink, S.; Dounavi, A. Detecting interspecific and geographic differentiation patterns in two interfertile oak species (Quercus petraea (Matt.) Liebl. and Q. robur L.) using small sets of microsatellite markers. For. Ecol. Manag. 2010, 259, 2026–2035. [Google Scholar] [CrossRef] [Green Version]
- Degen, B.; Yanbaev, R.; Yanbaev, Y. Genetic differentiation of Quercus robur in the South-Ural. Silvae Genet. 2019, 68, 111–115. [Google Scholar] [CrossRef] [Green Version]
- Pohjanmies, T.; Elshibli, S.; Pulkkinen, P.; Rusanen, M.; Vakkari, P.; Korpelainen, H.; Roslin, T. Fragmentation-related patterns of genetic differentiation in pedunculate oak (Quercus robur) at two hierarchical scales. Silva. Fenn. 2016, 50, 15. [Google Scholar] [CrossRef] [Green Version]
- Degen, B.; Streiff, R.; Ziegenhagen, B. Comparative study of genetic variation and differentiation of two pedunculate oak (Quercus robur) stands using microsatellite and allozyme loci. Heredity 1999, 83, 597–603. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gregorius, H.R.; Degen, B.; Konig, A. Problems in the analysis of genetic differentiation among populations—A case study in Quercus robur. Silvae Genet. 2007, 56, 190–199. [Google Scholar] [CrossRef] [Green Version]
- Leroy, T.; Louvet, J.M.; Lalanne, C.; Le Provost, G.; Labadie, K.; Aury, J.M.; Delzon, S.; Plomion, C.; Kremer, A. Adaptive introgression as a driver of local adaptation to climate in European white oaks. New Phytol. 2020, 226, 1171–1182. [Google Scholar] [CrossRef]
- Rellstab, C.; Zoller, S.; Walthert, L.; Lesur, I.; Pluess, A.R.; Graf, R.; Bodenes, C.; Sperisen, C.; Kremer, A.; Gugerli, F. Signatures of local adaptation in candidate genes of oaks (Quercus spp.) with respect to present and future climatic conditions. Mol. Ecol. 2016, 25, 5907–5924. [Google Scholar] [CrossRef] [PubMed]
- Bai, W.N.; Wang, W.T.; Zhang, D.Y. Contrasts between the phylogeographic patterns of chloroplast and nuclear DNA highlight a role for pollen-mediated gene flow in preventing population divergence in an East Asian temperate tree. Mol. Phylogenet. Evol. 2014, 81, 37–48. [Google Scholar] [CrossRef] [PubMed]
- Thomson, A.M.; Dick, C.W.; Pascoini, A.L.; Dayanandan, S. Despite introgressive hybridization, North American birches (Betula spp.) maintain strong differentiation at nuclear microsatellite loci. Tree Genet. Genomes 2015, 11, 1–12. [Google Scholar] [CrossRef] [Green Version]
- Curtu, A.L.; Gailing, O.; Finkeldey, R. Evidence for hybridization and introgression within a species-rich oak (Quercus spp.) community. BMC Evol. Biol. 2007, 7, 1–15. [Google Scholar] [CrossRef] [Green Version]
- Chybicki, I.J.; Burczyk, J. Realized gene flow within mixed stands of Quercus robur L. and Q-petraea (Matt.) L. revealed at the stage of naturally established seedling. Mol. Ecol. 2010, 19, 2137–2151. [Google Scholar] [CrossRef]
- Olrik, D.C.; Hauser, T.P.; Kjaer, E.D. Natural colonisation of an open area by Quercus robur L-From where did the vectors disperse the seed? Scand. J. For. Res. 2012, 27, 350–360. [Google Scholar] [CrossRef]
- Buschbom, J.; Yanbaev, Y.; Degen, B. Efficient long-distance gene flow into an isolated relict oak stand. J. Hered. 2011, 102, 464–472. [Google Scholar] [CrossRef] [Green Version]
- Schueler, S.; Schlunzen, K.; Scholz, F. Viability and sunlight sensitivity of oak pollen and its implications for pollen-mediated gene flow. Trees 2005, 19, 154–161. [Google Scholar] [CrossRef]
- Degen, B.; Blanc-Jolivet, C.; Bakhtina, S.; Ianbaev, R.; Yanbaev, Y.; Mader, M.; Nürnberg, S.; Schröder, H. Applying targeted genotyping by sequencing with a new set of nuclear and plastid SNP and indel loci for Quercus robur and Quercus petraea. Conserv. Genet. Resour. 2021, 13, 345–347. [Google Scholar] [CrossRef]
- Dumolin, S.; Demesure, B.; Petit, R.J. Inheritance of chloroplast and mitochondrial genomes in pedunculate oak investigated with an efficient PCR method. Theor. Appl. Genet. 1995, 91, 1253–1256. [Google Scholar] [CrossRef] [PubMed]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Garrison, E.; Marth, G. Haplotype-based variant detection from short-read sequencing. arXiv 2012, arXiv:1207.3907. [Google Scholar]
- Danecek, P.; Auton, A.; Abecasis, G.; Albers, C.A.; Banks, E.; DePristo, M.A.; Handsaker, R.E.; Lunter, G.; Marth, G.T.; Sherry, S.T.; et al. The variant call format and VCFtools. Bioinformatics 2011, 27, 2156–2158. [Google Scholar] [CrossRef] [PubMed]
- Rousset, F. GENEPOP’ 007: A complete re-implementation of the GENEPOP software for Windows and Linux. Mol. Ecol. Resour. 2008, 8, 103–106. [Google Scholar] [CrossRef]
- Kruskal, J.B. On the shortest spanning subtree of a graph and the traveling salesman problem. Proc. Am. Math. Soc. 1956, 7, 48–50. [Google Scholar] [CrossRef]
- Leigh, J.W.; Bryant, D. POPART: Full-feature software for haplotype network construction. Methods Ecol. Evol. 2015, 6, 1110–1116. [Google Scholar] [CrossRef]
- Gregorius, H.R. A unique genetic distance. Biom. J. 1984, 26, 13–18. [Google Scholar] [CrossRef]
- Gregorius, H.R. The relationship between the concepts of genetic diversity and differentiation. Theor. Appl. Genet. 1987, 74, 397–401. [Google Scholar] [CrossRef] [PubMed]
- Wright, S. Evolution and the Genetics of Populations: A Treatise in Four Volumes: Variability within and Among Natural Populations; University of Chicago Press: Chicargo, IL, USA, 1978; Volume 4. [Google Scholar]
- Degen, B. GDA-NT 2021: Genetic Data Analysis and Numercical Tests. Available online: https://www.thuenen.de/en/fg/software/gda-nt-2021/ (accessed on 1 August 2021).
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar] [CrossRef] [PubMed]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jakobsson, M.; Rosenberg, N.A. CLUMPP: A cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 2007, 23, 1801–1806. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kopelman, N.M.; Mayzel, J.; Jakobsson, M.; Rosenberg, N.A.; Mayrose, I. Clumpak: A program for identifying clustering modes and packaging population structure inferences across K. Mol. Ecol. Resour. 2015, 15, 1179–1191. [Google Scholar] [CrossRef] [Green Version]
- Degen, B. Spatial Genetic Software Version 2–2021. Available online: https://www.thuenen.de/en/fg/software/sgs/ (accessed on 1 August 2021).
- Degen, B.; Petit, R.J.; Kremer, A. SGS-Spatial genetic software: A computer program for analysis of spatial genetic and phenotypic structures of individuals and populations. J. Hered. 2001, 92, 447–449. [Google Scholar] [CrossRef] [Green Version]
- Brewer, S.; Cheddadi, R.; de Beaulieu, J.L.; Reille, M. The spread of deciduous Quercus throughout Europe since the last glacial period. For. Ecol. Manag. 2002, 156, 27–48. [Google Scholar] [CrossRef]
- Kozharinov, A.V.; Borisov, P.V. Distribution of oak forests in Eastern Europe over the last 13000 years. Contemp. Probl. Ecol. 2013, 6, 755–760. [Google Scholar] [CrossRef]
- Magyari, E.O.K.; Kunes, P.; Jakab, G.; Sumegi, P.; Pelankova, B.; Schabitz, F.; Braun, M.; Chytry, M. Late Pleniglacial vegetation in eastern-central Europe: Are there modern analogues in Siberia? Quat. Sci. Rev. 2014, 95, 60–79. [Google Scholar] [CrossRef]
- Petit, R.J.; Aguinagalde, I.; de Beaulieu, J.L.; Bittkau, C.; Brewer, S.; Cheddadi, R.; Ennos, R.; Fineschi, S.; Grivet, D.; Lascoux, M.; et al. Glacial refugia: Hotspots but not melting pots of genetic diversity. Science 2003, 300, 1563–1565. [Google Scholar] [CrossRef] [Green Version]
- Gailing, O.; Wachter, H.; Heyder, J.; Schmittz, H.P.; Finkeldey, R. Chloroplast DNA analysis in oak stands (Quercus robur L.) in North Rhine-Westphalia with presumably Slavonian origin: Is there an association between geographic origin and bud phenology? J. Appl. Bot. Food Qual. 2007, 81, 165–171. [Google Scholar]
- Jansen, S.; Konrad, H.; Geburek, T. The extent of historic translocation of Norway spruce forest reproductive material in Europe. Ann. For. Sci. 2017, 74, 1–17. [Google Scholar] [CrossRef]
- Eusemann, P.; Liesebach, H. Small-scale genetic structure and mating patterns in an extensive sessile oak forest (Quercus petraea (Matt.) Liebl.). Ecol. Evol. 2021, 11, 7796–7809. [Google Scholar] [CrossRef] [PubMed]
- Olszewski, A.; Brzeziecki, B. Role of jay (Garrulus glandarius) in initializing successional changes in forest communities with the participation of oak (Quercus sp.). Sylwan 2019, 163, 479–488. [Google Scholar] [CrossRef]
- Chybicki, I.J.; Burczyk, J. Seeing the forest through the trees: Comprehensive inference on individual mating patterns in a mixed stand of Quercus robur and Q-petraea. Ann. Bot. 2013, 112, 561–574. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Streiff, R.; Ducousso, A.; Lexer, C.; Steinkellner, H.; Gloessl, J.; Kremer, A. Pollen dispersal inferred from paternity analysis in a mixed oak stand of Quercus robur L-and Q-petraea (Matt.) Liebl. Mol. Ecol. 1999, 8, 831–841. [Google Scholar] [CrossRef]
- Gerber, S.; Chadoeuf, J.; Gugerli, F.; Lascoux, M.; Buiteveld, J.; Cottrell, J.; Dounavi, A.; Fineschi, S.; Forrest, L.L.; Fogelqvist, J.; et al. High rates of gene flow by pollen and seed in oak populations across Europe. PLoS ONE 2014, 9, e85130. [Google Scholar] [CrossRef]
- Moracho, E.; Moreno, G.; Jordano, P.; Hampe, A. Unusually limited pollen dispersal and connectivity of Pedunculate oak (Quercus robur) refugial populations at the species’ southern range margin. Mol. Ecol. 2016, 25, 3319–3331. [Google Scholar] [CrossRef] [PubMed]
- Buschbom, J.; Gimmerthal, S.; Kirschner, P.; Michalczyk, I.M.; Sebbenn, A.; Schueler, S.; Schlunzen, K.H.; Degen, B. Spatial composition of pollen-mediated gene flow in sessile oak. Forstarchiv 2012, 83, 12–18. [Google Scholar]
- Schueler, S.; Schlunzen, K.H. Modeling of oak pollen dispersal on the landscape level with a mesoscale atmospheric model. Environ. Modeling Assess. 2006, 11, 179–194. [Google Scholar] [CrossRef]
- Homolka, A.; Schueler, S.; Burg, K.; Fluch, S.; Kremer, A. Insights into drought adaptation of two European oak species revealed by nucleotide diversity of candidate genes. Tree Genet. Genomes 2013, 9, 1179–1192. [Google Scholar] [CrossRef]
- Savolainen, O.; Pyhajarvi, T.; Knurr, T. Gene flow and local adaptation in trees. Annu. Rev. Ecol. Evol. Syst. 2007, 38, 595–619. [Google Scholar] [CrossRef]
- Wright, S. Isolation by distance. Genetics 1943, 28, 114. [Google Scholar] [CrossRef] [PubMed]
- Lepais, O.; Petit, R.J.; Guichoux, E.; Lavabre, J.E.; Alberto, F.; Kremer, A.; Gerber, S. Species relative abundance and direction of introgression in oaks. Mol. Ecol. 2009, 18, 2228–2242. [Google Scholar] [CrossRef] [PubMed]
- Guichoux, E.; Garnier-Gere, P.; Lagache, L.; Lang, T.; Boury, C.; Petit, R.J. Outlier loci highlight the direction of introgression in oaks. Mol. Ecol. 2013, 22, 450–462. [Google Scholar] [CrossRef]
- Petit, R.J.; Bodenes, C.; Ducousso, A.; Roussel, G.; Kremer, A. Hybridization as a mechanism of invasion in oaks. New Phytol. 2004, 161, 151–164. [Google Scholar] [CrossRef] [Green Version]
- Jensen, J.; Larsen, A.; Nielsen, L.R.; Cottrell, J. Hybridization between Quercus robur and Q. petraea in a mixed oak stand in Denmark. Ann. For. Sci. 2009, 66, 1–12. [Google Scholar] [CrossRef] [Green Version]
- Thomson, A.M.; Dick, C.W.; Dayanandan, S. A similar phylogeographical structure among sympatric North American birches (Betula) is better explained by introgression than by shared biogeographical history. J. Biogeogr. 2015, 42, 339–350. [Google Scholar] [CrossRef] [Green Version]
- Leroy, T.; Roux, C.; Villate, L.; Bodenes, C.; Romiguier, J.; Paiva, J.A.P.; Dossat, C.; Aury, J.M.; Plomion, C.; Kremer, A. Extensive recent secondary contacts between four European white oak species. New Phytol. 2017, 214, 865–878. [Google Scholar] [CrossRef] [Green Version]
- Plomion, C.; Aury, J.M.; Amselem, J.; Leroy, T.; Murat, F.; Duplessis, S.; Faye, S.; Francillonne, N.; Labadie, K.; Le Provost, G.; et al. Oak genome reveals facets of long lifespan. Nat. Plants 2018, 4, 440–452. [Google Scholar] [CrossRef] [Green Version]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).