Biogeographic Changes in Forest Soil Microbial Communities of Offshore Islands—A Case Study of Remote Islands in Taiwan
Abstract
1. Introduction
2. Materials and Methods
3. Results
3.1. Soil Properties and Microbial Biomass
3.2. Soil Enzyme Activities
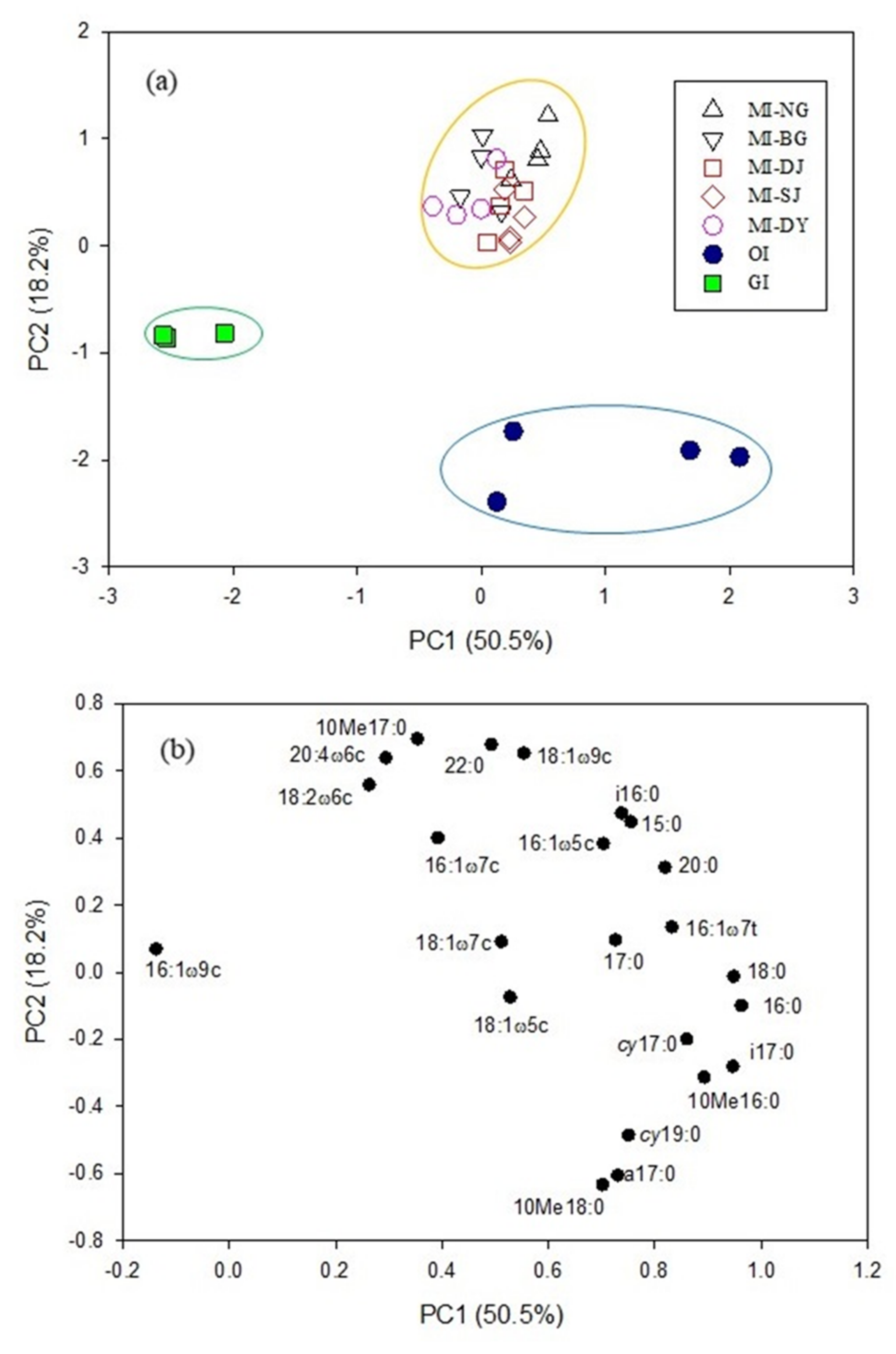
3.3. PLFA Biomarkers
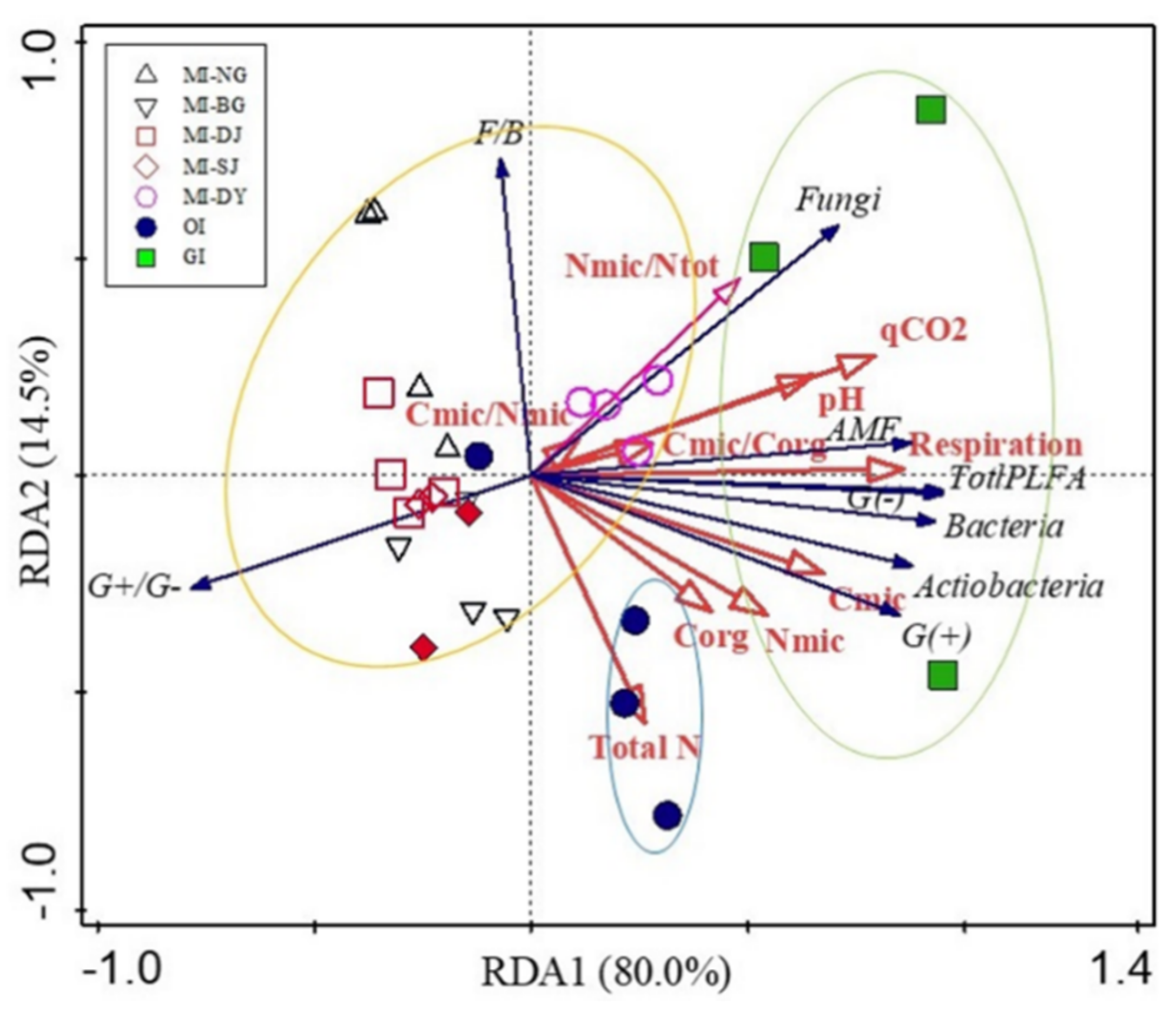
3.4. Soil Microbial Community Structure
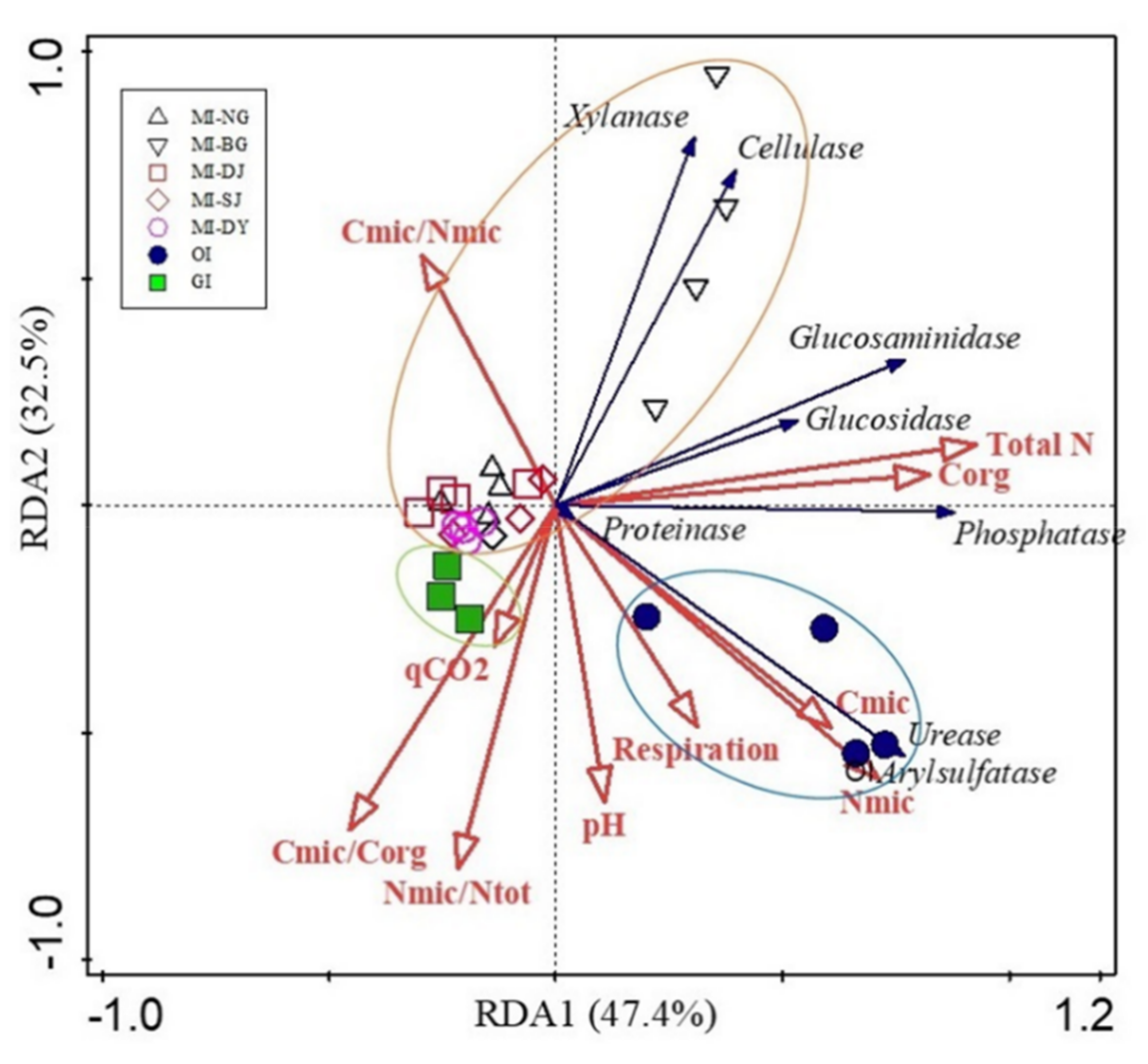
3.5. Correlation among Soil Properties and Microbial Communities
4. Discussion
4.1. Soil Chemistry and Biological Properties of the Different Islands
4.2. The Differences in Soil Enzyme Activity among the Islands
4.3. Soil Microbial Community Structure of Different Islands
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ochoa-Hueso, R.; Arca, V.; Delgado-Baquerizo, M.; Hamonts, K.; Piñeiro, J.; Serrano-Grijalva, L.; Shawyer, J.; Power, S.A. Links between soil microbial communities, functioning, and plant nutrition under altered rainfall in Australian grassland. Ecol. Monogr. 2020, 90, e01424. [Google Scholar] [CrossRef]
- Hossain, Z.; Sugiyama, S. Geographical structure of soil microbial communities in northern Japan: Effects of distance, land use type and soil properties. Eur. J. Soil Biol. 2011, 47, 88–94. [Google Scholar] [CrossRef]
- Bossio, D.A.; Scow, K.M.; Gunapala, N.; Graham, K.J. Determinants of soil microbial communities: Effects of agricultural management, season and soil type on phospholipid fatty acid profiles. Microbiol. Ecol. 1998, 36, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Byers, A.K.; Condron, L.; Donavan, T.; O’Callaghan, M.; Patuawa, T.; Waipara, N.; Black, A. Soil microbial diversity in adjacent forest systems—contrasting native, old growth kauri (Agathis australis) forest with exotic pine (Pinus radiata) plantation forest. FEMS Microbial. Ecol. 2020, 96, fiaa047. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.T.; Lin, Y.F.; Tsai, I.J.; Chang, E.H.; Jien, S.H.; Lin, Y.J.; Chiu, C.Y. Structure and Diversity of Soil Bacterial Communities in Offshore Islands. Sci. Rep. 2019, 9, 4689. [Google Scholar] [CrossRef] [PubMed]
- Cao, H.; Chen, R.; Wang, L.; Jiang, L.; Yang, F.; Zheng, S.; Wang, G.; Lin, X. Soil pH, total phosphorus, climate and distance are the major factors influencing microbial activity at a regional spatial scale. Sci. Rep. 2016, 6, 25815. [Google Scholar] [CrossRef]
- Jia, X.; He, Z.; Weiser, M.D.; Yin, T.; Akbar, S.; Kong, X.; Tian, K.; Jia, Y.; Lin, H.; Yu, M.; et al. Indoor evidence for the contribution of soil microbes and corresponding environments to the decomposition of Pinus massoniana and Castanopsis sclerophylla litter from Thousand Island Lake. Eur. J. Soil Biol. 2016, 77, 44–52. [Google Scholar] [CrossRef]
- Li, S.P.; Wang, P.; Chen, Y.; Wilson, M.C.; Yang, X.; Ma, C.; Lu, J.; Chen, X.Y.; Wu, J.; Shu, W.S.; et al. Island biogeography of soil bacteria and fungi: Similar patterns, but different mechanisms. ISME J. 2020, 14, 886–1896. [Google Scholar] [CrossRef]
- Xu, Z.; Yu, G.; Zhang, X.; He, N.; Wang, Q.; Wang, S.; Xu, X.; Wang, R.; Zhao, N. Biogeographical patterns of soil microbial community as influenced by soil characteristics and climate across Chinese forest biomes. Appl. Soil Ecol. 2018, 124, 298–305. [Google Scholar] [CrossRef]
- Xu, Z.; Yu, G.; Zhang, X.; He, N.; Wang, Q.; Wang, S.; Xu, X.; Wang, R.; Zhao, N. Divergence of dominant factors in soil microbial communities and functions in forest ecosystems along a climatic gradient. Biogeosciences 2018, 15, 1217–1228. [Google Scholar] [CrossRef]
- Zhou, Y.; Staver, A.C. Enhanced activity of soil nutrient-releasing enzymes after plant invasion: A meta-analysis. Ecology 2019, 100, e02830. [Google Scholar] [CrossRef] [PubMed]
- Fanin, N.; Kardol, P.; Farrell, M.; Nilsson, M.C.; Gundale, M.J.; Wardle, D.A. The ratio of Gram-positive to Gram-negative bacterial PLFA markers as an indicator of carbon availability in organic soils. Soil Biol. Biochem. 2019, 128, 111–114. [Google Scholar] [CrossRef]
- Soil Survey Staff. Keys to Soil Taxonomy, 12th ed.; USDA-Natural Resources Conservation Service: Washington, DC, USA, 2014.
- McLean, E.O. Soil pH and lime requirement. In Methods of Soil Analysis, Part 2 Chemical and Microbiological Properties; Page, A.L., Ed.; American society of Agronomy: Madison, WI, USA, 1982; pp. 199–224. [Google Scholar]
- Huang, C.Y.; Jien, S.H.; Chen, T.H.; Tian, G.; Chiu, C.Y. Soluble organic C and N and their relationships with soil organic C and N and microbial characteristics in moso bamboo (Phyllostachys edulis) plantations along an elevation gradient in central Taiwan. J. Soils Sediments 2014, 14, 1061–1070. [Google Scholar] [CrossRef]
- Tabatabai, M.A.; Bremner, J.M. Use of p-nitrophenyl phosphate for assay of soil phosphatase activity. Soil Biol. Biochem. 1969, 1, 301–307. [Google Scholar] [CrossRef]
- Schinner, F.; von Mersi, W. Xylanase-, CM-cellulase- and invertase activity in soil: An improved method. Soil Biol. Biochem. 1990, 22, 511–515. [Google Scholar] [CrossRef]
- Tabatabai, M.A.; Bremner, J.M. Arylsulphatase activity of soils. Soil Sci. Soc. Am. Proc. 1970, 34, 427–429. [Google Scholar] [CrossRef]
- Kandeler, E.; Gerber, H. Short-term assay of soil urease activity using colorimetric determination of ammonium. Biol Fertil. Soils 1988, 8, 199–202. [Google Scholar] [CrossRef]
- Ladd, J.N.; Butler, J.H.A. Short-term assays of soil proteolytic enzyme activities using proteins and dipeptide derivatives as substrates. Soil Biol. Biochem. 1972, 4, 19–30. [Google Scholar] [CrossRef]
- Parham, J.A.; Deng, S.P. Detection, quantification and characterization of β-glucosaminidase activity in soil. Soil Biol. Biochem. 2000, 32, 1183–1190. [Google Scholar] [CrossRef]
- Frostegård, A.; Baath, E.; Tunlid, A. Shifts in the structure of soil microbial communities in limed forests as revealed by phospholipid fatty acid analysis. Soil Biol. Biochem. 1993, 25, 723–730. [Google Scholar] [CrossRef]
- Chang, E.H.; Chen, C.T.; Chen, T.H.; Chiu, C.Y. Soil microbial communities and activities in sand dunes of subtropical coastal forests. Appl. Soil Ecol. 2011, 49, 256–262. [Google Scholar] [CrossRef]
- Zogg, G.P.; Zak, D.R.; Ringleberg, D.B.; MacDonald, N.W.; Pregitzer, K.S.; White, D.C. Compositional and functional shifts in microbial communities due to soil warming. Soil Sci. Soc. Am. J. 1997, 61, 475–481. [Google Scholar] [CrossRef]
- Zelles, L. Fatty acid patterns of phospholipids and lipopolysaccharides in the characterization of microbial communities in soil: A review. Biol. Fertil. Soils 1999, 29, 111–129. [Google Scholar] [CrossRef]
- Tonon, G.; Boldreghini, P.; Gioacchini, P. Seasonal changes in microbial nitrogen in an old broadleaf forest and in a neighbouring young plantation. Biol. Fertil. Soils 2005, 41, 101–108. [Google Scholar] [CrossRef]
- Hofmann, K.; Lamprecht, A.; Pauli, H.; Illmer, P. Distribution of prokaryotic abundance and microbial nutrient cycling across a high-alpine altitudinal gradient in the Austrian central Alps is affected by vegetation, temperature, and soil nutrients. Microb. Ecol. 2016, 72, 704–716. [Google Scholar] [CrossRef] [PubMed]
- Tufekcioglu, A.; Raich, J.W.; Isenhart, T.M.; Schultz, R.C. Soil respiration within riparian buffers and adjacent crop fields. Plant. Soil 2001, 229, 117–124. [Google Scholar] [CrossRef]
- Romanowicz, K.J.; Freedman, Z.B.; Upchurch, R.A.; Argiroff, W.A.; Zak, D.R. Active microorganisms in forest soils differ from the total community yet are shaped by the same environmental factors: The influence of pH and soil moisture. FEMS Microb. Ecol. 2016, 92, fiw149. [Google Scholar] [CrossRef]
- Verburg, P.S.J.; Van Dam, D.; Hefting, M.M.; Tietema, A. Microbial transformations of C and N in a boreal forest floor as affected by temperature. Plant. Soil 1999, 208, 187–197. [Google Scholar] [CrossRef]
- Davidson, E.A.; Janssens, I.A. Temperature sensitivity of soil carbon decomposition and feedbacks to climate change. Nature 2006, 440, 165–173. [Google Scholar] [CrossRef]
- Deng, Q.; Hui, D.; Zhang, D.; Zhou, G.; Liu, J.; Liu, S.; Chu, G.; Li, J. Effects of Precipitation Increase on Soil Respiration: A Three-Year Field Experiment in Subtropical Forests in China. PLoS ONE 2012, 7, e41493. [Google Scholar] [CrossRef]
- Kassim, G.; Martin, J.R.; Haider, K. Incorporation of a wide variety of organic substrate carbons into soil biomass as estimated by the fumigation procedure. Soil Sci. Soc. Am. J. 1982, 45, 1106–1112. [Google Scholar] [CrossRef]
- Wardle, D.A.; Ghani, A. A critique of the microbial metabolic quotient (qCO2) as a bioindicator of disturbance and ecosystem development. Soil Biol. Biochem. 1995, 27, 1601–1610. [Google Scholar] [CrossRef]
- Mahía, J.; Pérez-Ventura, L.; Cabaneiro, A.; Díaz-Raviña, M. Soil microbial biomass under pine forests in the north-western Spain: Influence of stand age, site index and parent material. For. Syst. 2006, 15, 152–159. [Google Scholar] [CrossRef]
- Hagerty, S.B.; van Groenigen, K.J.; Allison, S.D.; Hungate, B.A.; Schwartz, E.; Koch, G.W.; Kolka, R.K.; Dijkstra, P. Accelerated microbial turnover but constant growth efficiency with warming in soil. Nat. Clim. Chang. 2014, 4, 903–906. [Google Scholar] [CrossRef]
- Xu, X.; Schimel, J.P.; Janssens, I.A.; Song, X.; Song, C.; Yu, G.; Sinsabaugh, R.L.; Tang, D.; Zhang, X.; Thornton, P.E. Global pattern and controls of soil microbial metabolic quotient. Ecol. Monogr. 2017, 87, 429–441. [Google Scholar] [CrossRef]
- Lamarche, J.; Bradley, R.L.; Paré, D.; Légaré, S.; Bergeron, Y. Soil parent material may control forest floor properties more than stand type or stand age in mixedwood boreal forests. Écoscience 2004, 11, 228–237. [Google Scholar] [CrossRef]
- Alvarez, R.; Santanatoglia, O.J.; Garcîa, R. Effect of temperature on soil microbial biomass and its metabolic quotient in situ under different tillage systems. Biol. Fertil. Soils 1995, 19, 227–230. [Google Scholar] [CrossRef]
- Pommerening-Roser, A.; Koops, H.P. Environmental pH as an important factor for the distribution of urease positive ammonia-oxidizing bacteria. Microbiol. Res. 2005, 160, 27–35. [Google Scholar] [CrossRef]
- Shahbaz, M.; Kätterer, T.; Thornton, B.; Börjesson, G. Dynamics of fungal and bacterial groups and their carbon sources during the growing season of maize in a long-term experiment. Biol. Fertil. Soils 2020, 56, 759–770. [Google Scholar] [CrossRef]
- Fierer, N.; Jackson, R.B. The diversity and biogeography of soil bacterial communities. Proc. Natl. Acad. Sci. USA 2006, 103, 626–631. [Google Scholar] [CrossRef]
- Lauber, C.L.; Hamady, M.; Knight, R.; Fierer, N. Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale. Appl. Environ. Microbiol. 2009, 75, 5111–5120. [Google Scholar] [CrossRef] [PubMed]
- Rousk, J.; Brookes, P.C.; Bååth, E. Investigating the mechanisms for the opposing pH relationships of fungal and bacterial growth in soil. Soil Biol. Biochem. 2010, 42, 926–934. [Google Scholar] [CrossRef]
- Blagodatskaya, E.V.; Anderson, T.H. Interactive effects of pH and substrate quality on the fungal-to-bacterial ratio and qCO2 of microbial communities in forest soils. Soil Biol. Biochem. 1998, 30, 1269–1274. [Google Scholar] [CrossRef]
- Bååth, E.; Anderson, T.H. Comparison of soil fungal/bacterial ratios in a pH gradient using physiological and PLFA-based techniques. Soil Biol. Biochem. 2003, 35, 955–963. [Google Scholar] [CrossRef]
- Djukic, I.; Zehetner, F.; Mentler, A.; Gerzabek, M.H. Microbial community composition and activity in different Alpine vegetation zones. Soil Biol. Biochem. 2010, 42, 155–161. [Google Scholar] [CrossRef]
- Shen, C.; Ge, Y.; Yang, T.; Chu, H. Verrucomicrobial elevational distribution was strongly influenced by soil pH and carbon/nitrogen ratio. J. Soils Sediments 2017, 17, 2449–2456. [Google Scholar] [CrossRef]
- Wagai, R.; Kitayama, K.; Satomura, T.; Fujinuma, R.; Balser, T. Interactive influences of climate and parent material on soil microbial community structure in Bornean tropical forest ecosystems. Ecol. Res. 2011, 26, 627–636. [Google Scholar] [CrossRef]
- Iovieno, P.; Alfani, A.; Bååth, E. Soil microbial community structure and biomass as affected by Pinus pinea plantation in two Mediterranean areas. Appl. Soil Ecol. 2010, 45, 56–63. [Google Scholar] [CrossRef]
- Xiong, J.; Liu, Y.; Lin, X.; Zhang, H.; Zeng, J.; Hou, J.; Yang, Y.; Yao, T.; Knight, R.; Chu, H. Geographic distance and pH drive bacterial distribution in alkaline lake sediments across Tibetan Plateau. Environ. Microbiol. 2012, 14, 2457–2466. [Google Scholar] [CrossRef]
| Island | Abbreviation | pH § | Org C § (g kg−1) | Cmic § (µg g−1) | Cmic/Corg (%) | Respiration (µg g−1 h−1) | Metabolic Quotient (qCO2) |
|---|---|---|---|---|---|---|---|
| Matsu (Nangan) | MI-NG | 4.86 b | 24.0 c | 448.5 de | 1.90 a | 2.89 c | 5.75 c |
| Matsu (Beigan) | MI-BG | 4.24 b | 57.1 a | 641.7 c | 1.17 b | 3.66 bc | 5.67 c |
| Matsu (Donju) | MI-DJ | 4.81 b | 21.0 c | 412.7 e | 1.99 a | 2.32 c | 5.44 c |
| Matsu (Shiju) | MI-SJ | 4.47 b | 22.9 c | 503.9 d | 2.25 a | 2.73 c | 5.41 c |
| Matsu (Dongyin) | MI-DY | 4.85 b | 29.7 c | 565.8 c | 1.91 a | 5.4 b | 9.64 a |
| Orchid | OI | 6.10 a | 64.0 a | 1241.6 b | 1.95 a | 8.84 a | 7.17 b |
| Green | GI | 6.43 a | 44.9 b | 1963.5 a | 2.34 a | 10.3 a | 9.86 a |
| Island | Abbreviation | Cellulase (µg glucose g−1d−1) | Xylanase (µg glucose g−1d−1) | Urease (mmole NH4+-N g−1h−1) | Phosphatase (µg nitrophenol g−1h−1) | β–Glucosaminidase (µg nitrophenol g−1h−1) | Glucosidase (µg nitrophenol g−1h−1) | Arylsulfatase (μg nitrophenol g−1h−1) | Proteinase µg tyrosine g−12h−1) |
|---|---|---|---|---|---|---|---|---|---|
| Matsu (Nangan) | MI-NG | 762 b | 2955 b | 1.08 b | 1574 b | 260 bc | 101 a | 97.0 cd | 92.5 b |
| Matsu (Beigan) | MI-BG | 3070 a | 6608 a | 1.79 b | 1987 b | 403 a | 135 a | 97.3 cd | 199 ab |
| Matsu (Donju) | MI-DJ | 1149 b | 3190 b | 1.34 b | 870 c | 211 c | 93.0 a | 42.7 d | 153 b |
| Matsu (Shiju) | MI-SJ | 704 b | 2331 b | 0.85 b | 755 c | 253 bc | 120 a | 128 c | 141 b |
| Matsu (Dongyin) | MI-DY | 677 b | 2722 b | 2.18 b | 565 c | 87.0 d | 106 a | 119 cd | 162 b |
| Orchid | OI | 960 b | 2530 b | 10.5 a | 2762 a | 340 ab | 127 a | 788 a | 180 ab |
| Green | GI | 973 b | 3347 b | 2.37 b | 842 c | 95.3 d | 91 a | 308 b | 340 a |
| Island | Abbreviation | Total PLFAs | Bacteria | Fungi | AMF Fungi | Actinobacteria | G+ | G− | G+/G− | Fungi/ Bacteria |
|---|---|---|---|---|---|---|---|---|---|---|
| Matsu (Nangan) | MI-NG | 23.0 e | 9.43 d | 0.973 cd | 0.724 c | 0.374 c | 5.26 d | 3.84 c | 1.37 c | 0.10 a |
| Matsu (Beigan) | MI-BG | 37.2 cd | 15.9 cd | 0.948 cd | 1.14 bc | 0.572 c | 9.95 bc | 5.32 c | 1.91 a | 0.057 bc |
| Matsu (Donju) | MI-DJ | 22.7 e | 9.92 d | 0.448 c | 0.617 c | 0.449 c | 6.18 d | 3.37 c | 1.84 ab | 0.047 bc |
| Matsu (Shiju) | MI-SJ | 30.2 de | 13.7 d | 0.681 c | 1.01 bc | 0.504 c | 8.23 cd | 5.0 c | 1.64 b | 0.050 bc |
| Matsu (Dongyin) | MI-DY | 48.7 bc | 21.6 bc | 1.66 b | 1.44 b | 0.772 c | 11.4 bc | 9.56 b | 1.19 c | 0.080 ab |
| Orchid | OI | 50.6 b | 23.7 b | 0.753 c | 1.09 bc | 1.77 b | 12.8 b | 10.4 b | 1.27 c | 0.035 c |
| Green | GI | 95.2 a | 43.6 a | 2.66 a | 3.71 a | 2.73 a | 21.3 a | 20.7 a | 1.02 d | 0.067 abc |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chang, E.-H.; Tsai, I.J.; Jien, S.-H.; Tian, G.; Chiu, C.-Y. Biogeographic Changes in Forest Soil Microbial Communities of Offshore Islands—A Case Study of Remote Islands in Taiwan. Forests 2021, 12, 4. https://doi.org/10.3390/f12010004
Chang E-H, Tsai IJ, Jien S-H, Tian G, Chiu C-Y. Biogeographic Changes in Forest Soil Microbial Communities of Offshore Islands—A Case Study of Remote Islands in Taiwan. Forests. 2021; 12(1):4. https://doi.org/10.3390/f12010004
Chicago/Turabian StyleChang, Ed-Haun, Isheng Jason Tsai, Shih-Hao Jien, Guanglong Tian, and Chih-Yu Chiu. 2021. "Biogeographic Changes in Forest Soil Microbial Communities of Offshore Islands—A Case Study of Remote Islands in Taiwan" Forests 12, no. 1: 4. https://doi.org/10.3390/f12010004
APA StyleChang, E.-H., Tsai, I. J., Jien, S.-H., Tian, G., & Chiu, C.-Y. (2021). Biogeographic Changes in Forest Soil Microbial Communities of Offshore Islands—A Case Study of Remote Islands in Taiwan. Forests, 12(1), 4. https://doi.org/10.3390/f12010004






