Role of Biofilm Formation by Bacillus pumilus HR10 in Biocontrol against Pine Seedling Damping-Off Disease Caused by Rhizoctonia solani
Abstract
1. Introduction
2. Materials and Methods
2.1. Strains and Culture Conditions
2.2. Mutagenesis of B. pumilus HR10 and Screening of Biofilm-Deficient Mutants
2.3. Colony Morphology, Growth Curve and Spore Formation
2.4. Analysis of Biofilm Formation and Swarming Motility
2.5. Quantitative Estimation of Biofilm Polysaccharides
2.6. Quantitative Estimation of Biofilm Proteins
2.7. qRT-PCR Assay of Biofilm-Related Gene Expression
2.8. Antifungal Activity Assay
2.9. Cultivation of P. massoniana Seedlings
2.10. Root Colonization Assay
2.11. Assay of Plant Defense Gene Expression
2.12. Biocontrol Assay
total number of seedlings in each treatment) × 100.
disease incidence of untreated seedlings) × 100. Each treatment contained 16 P. massoniana seedlings.
2.13. Statistical Analyses
3. Results
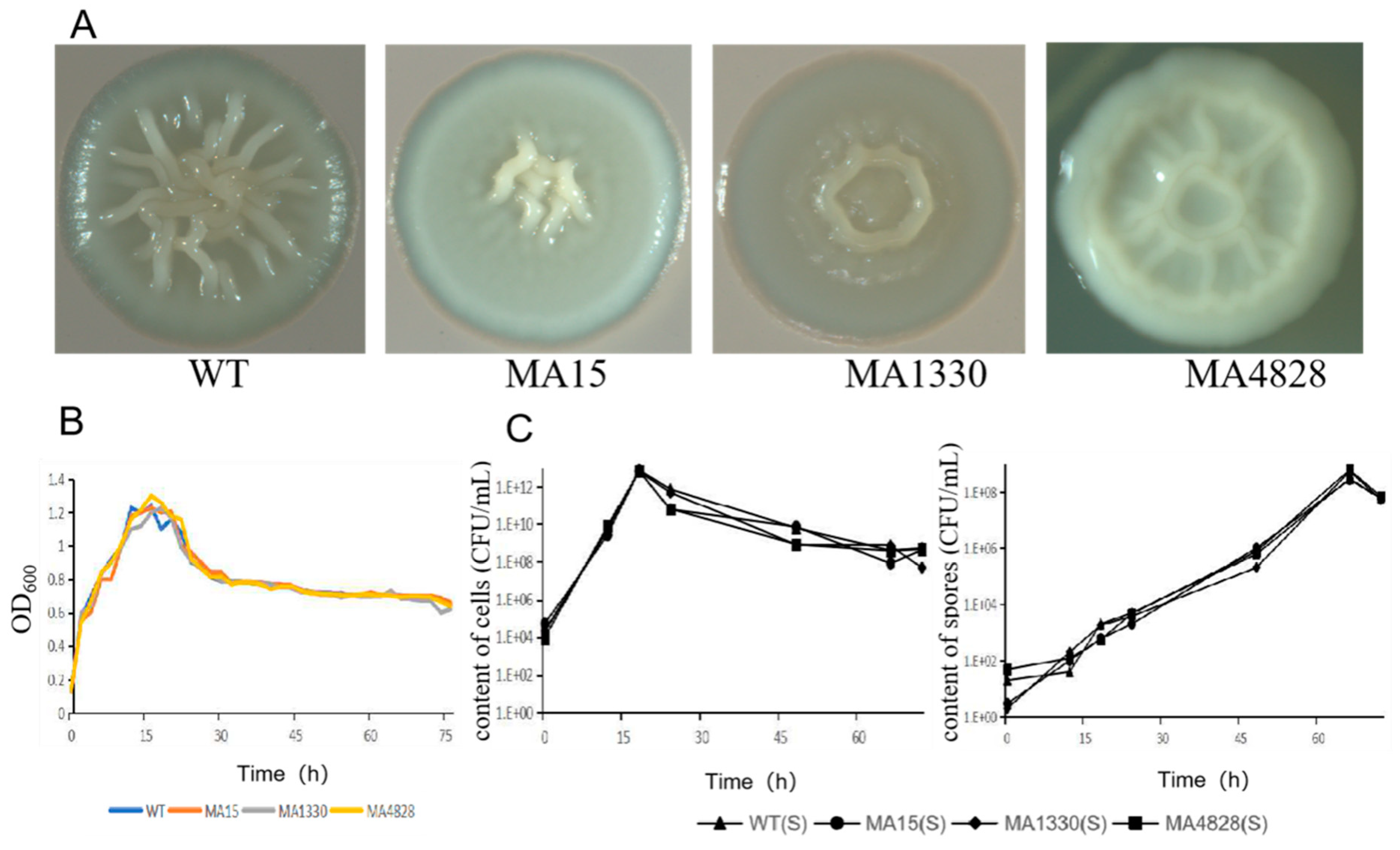
3.1. Colony Morphology and Growth Curve of Cells and Spores
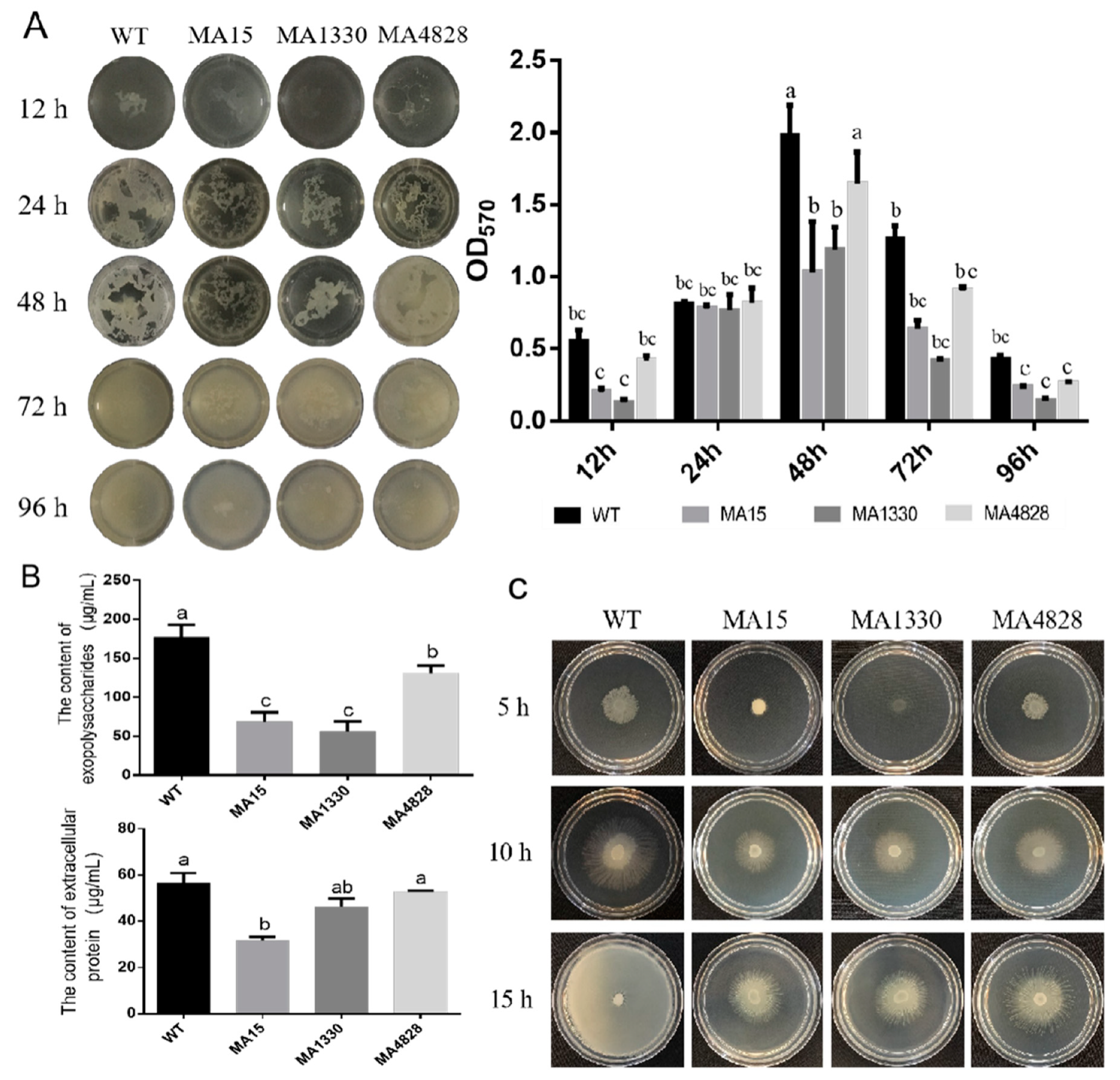
3.2. The Biofilm Substance Contents and Swarming Motility of the Biofilm-Deficient Mutants Declined
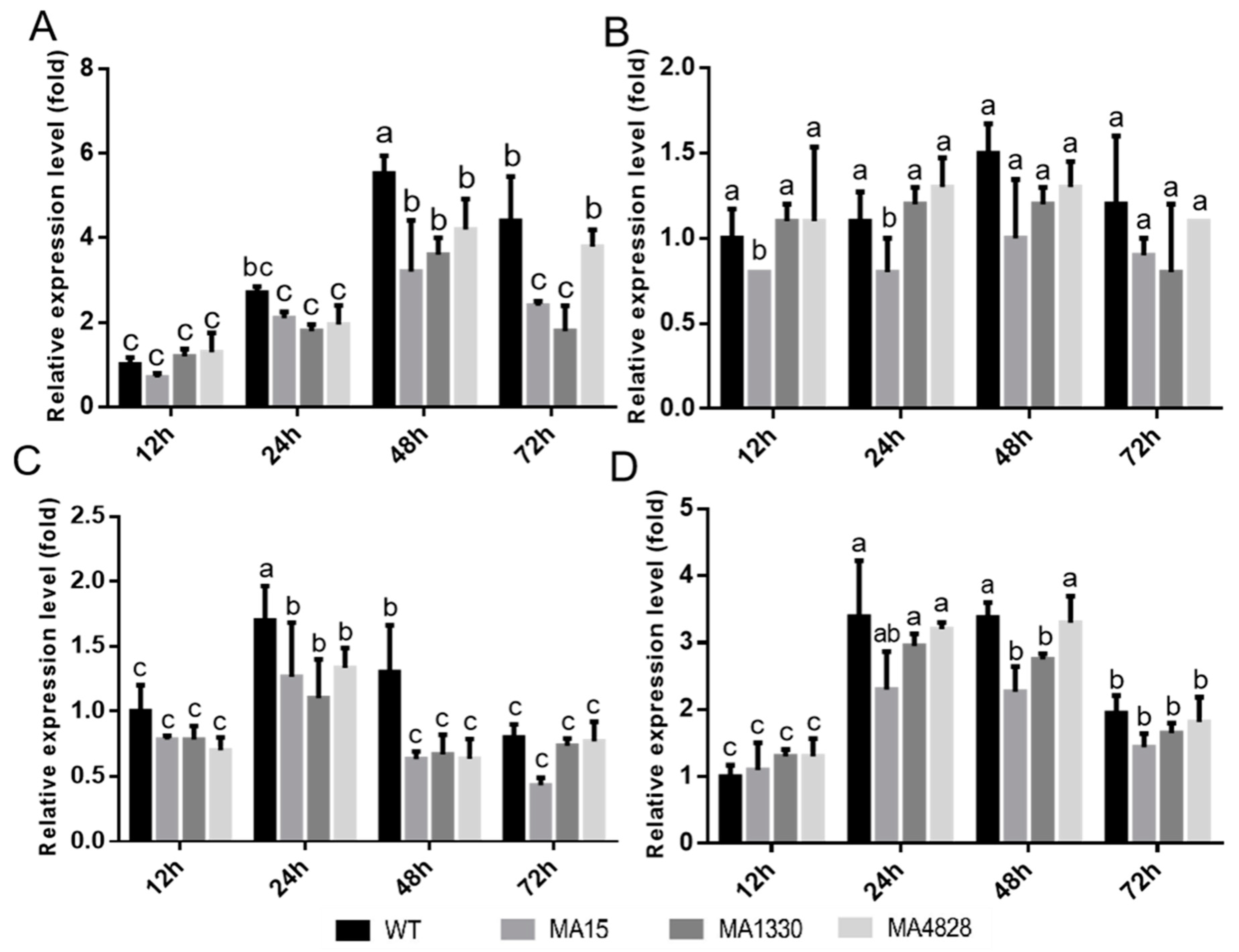
3.3. Biofilm-Related Genes of HR10 Wild-Type and Mutants Showed Different Expression Levels
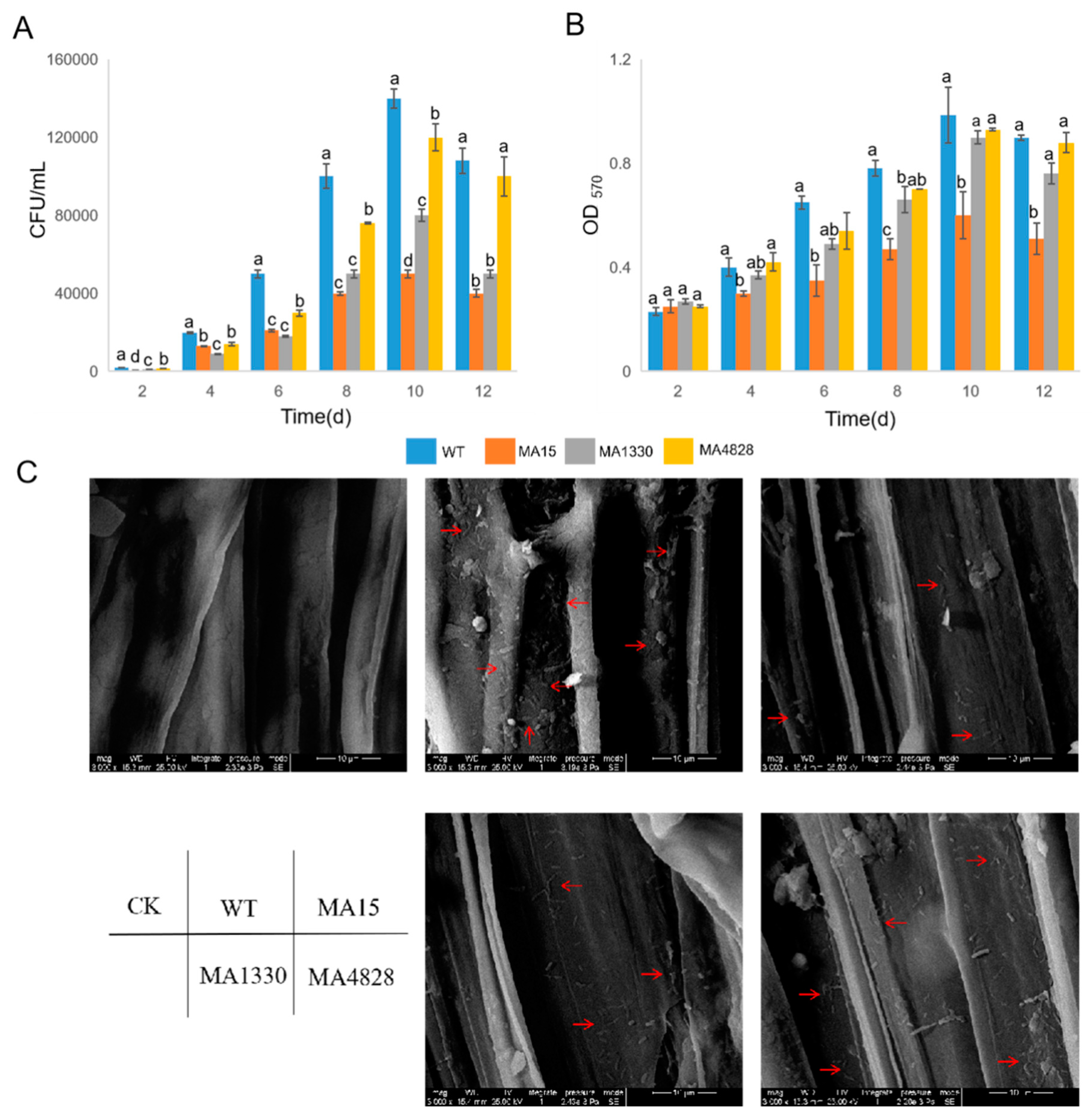
3.4. Deficiency in Biofilm Formation Reduced Root Colonization on P. massoniana Seedlings
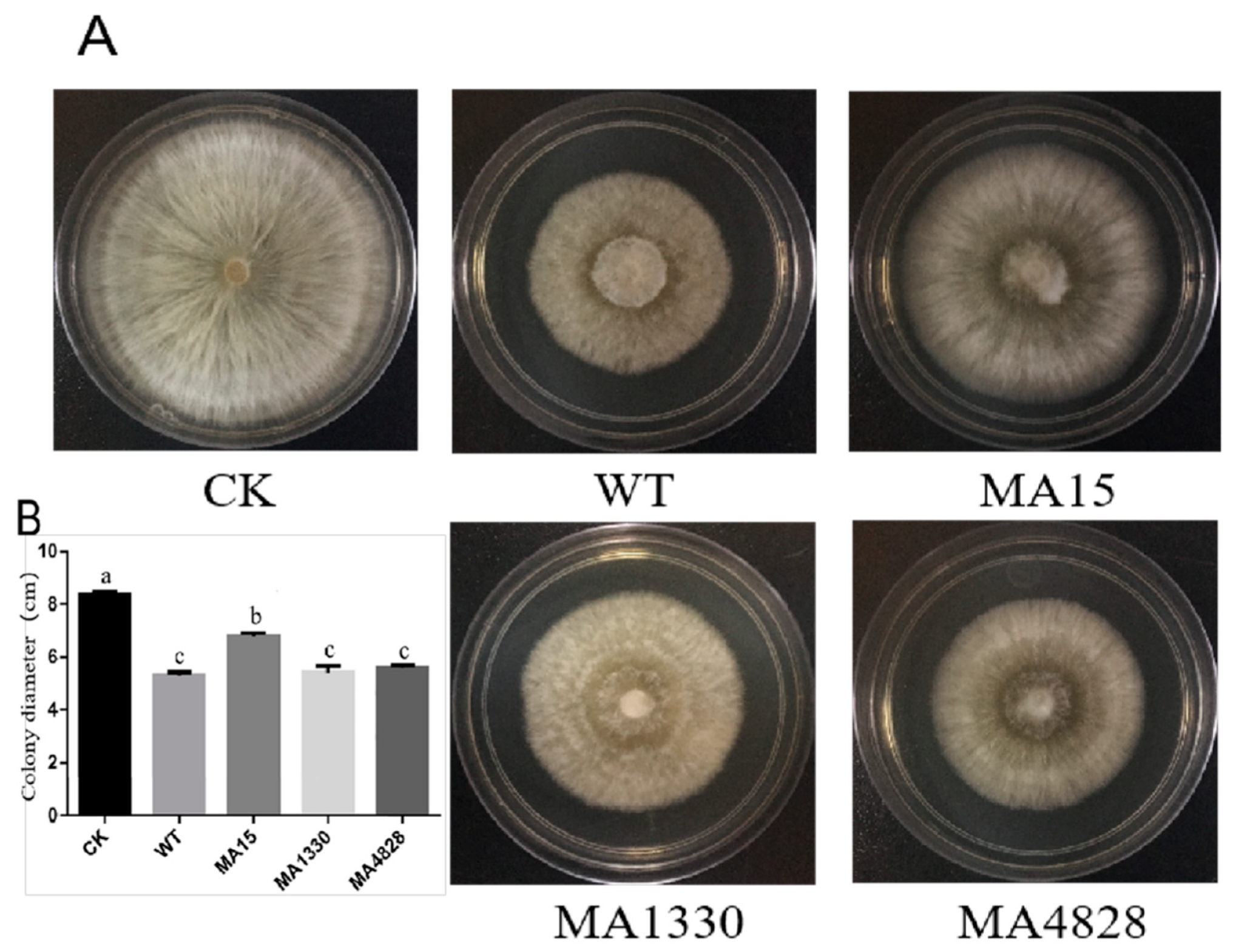
3.5. Mutants Exhibited Different Inhibitory Activities against R. Solani
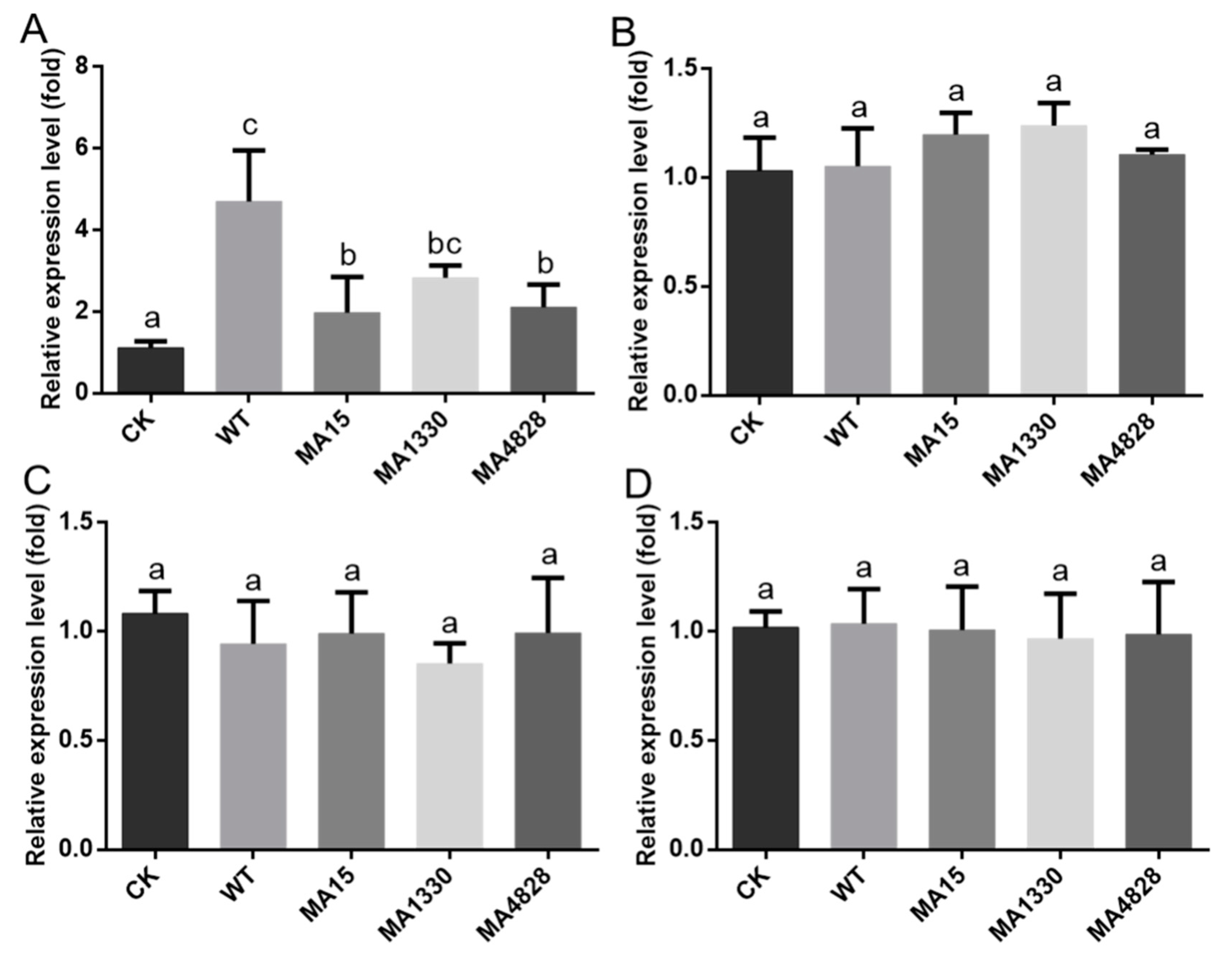
3.6. Biofilm Formation Contributes to Inducing the Systemic Resistance of Plants
3.7. Biofilm Formation and Inhibitory Activity Are Both Required for Biocontrol against Pine Seedling Damping-Off Disease
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Phazna Devi, T.A.; Sahoo, D.; Setti, A.; Sharma, C.; Kalita, M.C. Bacterial rhizosphere community profile at different growth stages of Umorok (Capsicum chinense) and its response to the root exudates. Int. Microbiol. 2019, 23, 241–251. [Google Scholar] [CrossRef]
- Wang, B.; Shen, Z.; Zhang, F.; Raza, W.; Yuan, J.; Huang, R.; Ruan, Y.; Li, R.; Shen, Q. Bacillus amyloliquefaciens Strain W19 can Promote Growth and Yield and Suppress Fusarium Wilt in Banana Under Greenhouse and Field Conditions. Pedosphere 2016, 26, 733–744. [Google Scholar] [CrossRef]
- Liu, Y.; Teng, K.; Wang, T.; Dong, E.; Zhang, M.; Tao, Y.; Zhong, J. Antimicrobial Bacillus velezensis HC6: production of three kinds of lipopeptides and biocontrol potential in maize. J. Appl. Microbiol. 2019, 128, 242–254. [Google Scholar] [CrossRef] [PubMed]
- Fan, H.; Zhang, Z.; Li, Y.; Zhang, X.; Duan, Y.; Wang, Q. Biocontrol of Bacterial Fruit Blotch by Bacillus subtilis 9407 via Surfactin-Mediated Antibacterial Activity and Colonization. Front. Microbiol. 2017, 8, 8. [Google Scholar] [CrossRef]
- Chowdhury, S.P.; Hartmann, A.; Gao, X.; Borriss, R. Biocontrol mechanism by root-associated Bacillus amyloliquefaciens FZB42—A review. Front. Microbiol. 2015, 6, 780. [Google Scholar] [CrossRef]
- Xu, Z.; Shao, J.; Li, B.; Yan, X.; Shen, Q.; Zhang, R. Contribution of Bacillomycin D in Bacillus amyloliquefaciens SQR9 to Antifungal Activity and Biofilm Formation. Appl. Environ. Microbiol. 2012, 79, 808–815. [Google Scholar] [CrossRef]
- Li, L.; Tan, J.; Chen, F. Bacillus pumilus strain LYMC-3 shows nematicidal activity against Bursaphelenchus xylophilus via the production of a guanidine compound. Biocontrol Sci. Technol. 2018, 28, 1128–1139. [Google Scholar] [CrossRef]
- Fan, B.; Blom, J.; Klenk, H.-P.; Borriss, R. Bacillus amyloliquefaciens, Bacillus velezensis, and Bacillus siamensis Form an “Operational Group B. amyloliquefaciens” within the B. subtilis Species Complex. Front. Microbiol. 2017, 8, 1807. [Google Scholar] [CrossRef]
- Fan, B.; Wang, C.; Song, X.; Ding, X.; Wu, L.; Wu, H.; Gao, X.; Borriss, R. Bacillus velezensis FZB42 in 2018: The Gram-Positive Model Strain for Plant Growth Promotion and Biocontrol. Front. Microbiol. 2018, 9, 2491. [Google Scholar] [CrossRef]
- Beneduzi, A.; Ambrosini, A.; Passaglia, L.M.P. Plant growth-promoting rhizobacteria (PGPR): their potential as antagonists and biocontrol agents. Genet. Mol. Boil. 2012, 35, 1044–1051. [Google Scholar] [CrossRef]
- Saleem, M.; Arshad, M.; Hussain, S.; Bhatti, A.S. Perspective of plant growth promoting rhizobacteria (PGPR) containing ACC deaminase in stress agriculture. J. Ind. Microbiol. Biotechnol. 2007, 34, 635–648. [Google Scholar] [CrossRef] [PubMed]
- Jin, Y.; Zhu, H.; Luo, S.; Yang, W.; Zhang, L.; Li, S.; Jin, Q.; Cao, Q.; Sun, S.; Xiao, M. Role of Maize Root Exudates in Promotion of Colonization of Bacillus velezensis Strain S3-1 in Rhizosphere Soil and Root Tissue. Curr. Microbiol. 2019, 76, 855–862. [Google Scholar] [CrossRef] [PubMed]
- Chowdhury, S.P.; Dietel, K.; Rändler, M.; Schmid, M.; Junge, H.; Borriss, R.; Hartmann, A.; Grosch, R. Effects of Bacillus amyloliquefaciens FZB42 on Lettuce Growth and Health under Pathogen Pressure and Its Impact on the Rhizosphere Bacterial Community. PLoS ONE 2013, 8, e68818. [Google Scholar] [CrossRef] [PubMed]
- Weng, J.; Wang, Y.; Li, J.; Shen, Q.; Zhang, R. Enhanced root colonization and biocontrol activity of Bacillus amyloliquefaciens SQR9 by abrB gene disruption. Appl. Microbiol. Biotechnol. 2012, 97, 8823–8830. [Google Scholar] [CrossRef] [PubMed]
- Flemming, H.-C.; Wingender, J.; Szewzyk, U.; Steinberg, P.; Rice, S.A.; Kjelleberg, S. Biofilms: an emergent form of bacterial life. Nat. Rev. Genet. 2016, 14, 563–575. [Google Scholar] [CrossRef] [PubMed]
- Flemming, H.-C.; Wingender, J. The biofilm matrix. Nat. Rev. Genet. 2010, 8, 623–633. [Google Scholar] [CrossRef]
- Guttenplan, S.B.; Blair, K.M.; Kearns, D.B. The EpsE Flagellar Clutch Is Bifunctional and Synergizes with EPS Biosynthesis to Promote Bacillus subtilis Biofilm Formation. PLoS Genet. 2010, 6, e1001243. [Google Scholar] [CrossRef]
- Morikawa, M.; Kagihiro, S.; Haruki, M.; Takano, K.; Branda, S.; Kolter, R.; Kanaya, S. Biofilm formation by a Bacillus subtilis strain that produces gamma-polyglutamate. Microbiol SGM 2006, 152, 2801–2807. [Google Scholar] [CrossRef]
- Romero, D.; Aguilar, C.; Losick, R.; Kolter, R. Amyloid fibers provide structural integrity to Bacillus subtilis biofilms. Proc. Natl. Acad. Sci. USA 2010, 107, 2230–2234. [Google Scholar] [CrossRef]
- Luo, C.; Zhou, H.; Zou, J.; Wang, X.; Zhang, R.; Xiang, Y.; Chen, Z. Bacillomycin L and surfactin contribute synergistically to the phenotypic features of Bacillus subtilis 916 and the biocontrol of rice sheath blight induced by Rhizoctonia solani. Appl. Microbiol. Biotechnol. 2014, 99, 1897–1910. [Google Scholar] [CrossRef]
- Zeriouh, H.; Romero, D.; García-Gutiérrez, L.; Cazorla, F.M.; De Vicente, A.; Pérez-García, A. The Iturin-like Lipopeptides Are Essential Components in the Biological Control Arsenal of Bacillus subtilis Against Bacterial Diseases of Cucurbits. Mol. Plant Microbe Interact. 2011, 24, 1540–1552. [Google Scholar] [CrossRef] [PubMed]
- Hobley, L.; Harkins, C.; Macphee, C.E.; Stanley-Wall, N.R. Giving structure to the biofilm matrix: an overview of individual strategies and emerging common themes. FEMS Microbiol Rev. 2015, 39, 649–669. [Google Scholar] [CrossRef] [PubMed]
- Sheng, J.M.; Wu, X.Q.; Hou, L.L. Screening and identification of mycorrhiza helper bacteria from rhizosphere of Pinus thunbergii-Rhizopogen luteous. J. Northeast For. Univ. 2014, 42, 110–114. (In Chinese) [Google Scholar]
- Hou, L.L.; Wu, X.Q.; Sheng, J.M. Inhibition of 4 mycorrhizal-helper bacteria strains against seedling damping-off disease. J. Nanjing For. Univ. 2011, 35, 43–46. (In Chinese) [Google Scholar]
- Huang, Y.M.; Yang, X.; Zhang, D.J.; Zhang, J. The effects of gap size and litter species on colonization of soil fauna during litter decomposition in Pinus massoniana plantations. Appl. Soil Ecol. 2020, 155, 103611. [Google Scholar] [CrossRef]
- Luo, X.; Yu, C. First report of damping-off disease caused by Fusarium oxysporum in Pinus massoniana in China. J. Plant Dis. Prot. 2020, 127, 401–409. [Google Scholar] [CrossRef]
- Xu, W.X.; Liu, H.P.; Zhuang, Z.H. Mutagenesis of Bacillus pumilus AS 1.271 by 1-methyl-3-nitro-1-nitrosoguanidine. Acta Microbiol. Sin. 1977, 1, 23–30. (In Chinese) [Google Scholar]
- Meng, L.; Xu, D.; Chen, W.Z.; Zhang, X.C. Screening and character analysis of a new strain 07-2 of Asparagus schoberioides. Period. Ocean Univ. China 2009, 39, 94–98. (In Chinese) [Google Scholar]
- Wang, N.; Wang, L.; Zhu, K.; Hou, S.; Chen, L.; Mi, D.; Gui, Y.; Qi, Y.; Jiang, C.; Guo, J. Plant Root Exudates Are Involved in Bacillus cereus AR156 Mediated Biocontrol Against Ralstonia solanacearum. Front. Microbiol. 2019, 10, 98. [Google Scholar] [CrossRef]
- Gu, D.; Meng, H.; Li, Y.; Ge, H.; Jiao, X. A GntR Family Transcription Factor (VPA1701) for Swarming Motility and Colonization of Vibrio parahaemolyticus. Pathogens 2019, 8, 235. [Google Scholar] [CrossRef]
- Liu, L.; Hu, Q.; Le, Y.; Chen, G.; Tong, Z.; Xu, Q.; Wang, G. Chlorination-mediated EPS excretion shapes early-stage biofilm formation in drinking water systems. Process. Biochem. 2017, 55, 41–48. [Google Scholar] [CrossRef]
- Velmourougane, K.; Prasanna, R.; Singh, S.; Chawla, G.; Kumar, A.; Saxena, A.K. Modulating rhizosphere colonisation, plant growth, soil nutrient availability and plant defense enzyme activity through Trichoderma viride-Azotobacter chroococcum biofilm inoculation in chickpea. Plant Soil 2017, 421, 157–174. [Google Scholar] [CrossRef]
- Hirao, T.; Fukatsu, E.; Watanabe, A. Characterization of resistance to pine wood nematode infection in Pinus thunbergii using suppression subtractive hybridization. BMC Plant Boil. 2012, 12, 13. [Google Scholar] [CrossRef] [PubMed]
- Cao, F.X.; Wang, M.; Long, J.X. Molecular cloning and sequence analysis of phenylalanine ammonia-lyase cDNA from Pinus massoniana. J. Cent. South Univ. For. Technol. 2010, 30, 79–83. (In Chinese) [Google Scholar]
- Du, M.; Ding, G.; Cai, Q. The Transcriptomic Responses of Pinus massoniana to Drought Stress. Forests 2018, 9, 326. [Google Scholar] [CrossRef]
- Wei, Y.; Liu, Q.; Dong, H.; Zhou, Z.; Hao, Y.; Chen, X.; Xu, L. Selection of Reference Genes for Real-Time Quantitative PCR in Pinus massoniana Post Nematode Inoculation. PLoS ONE 2016, 11, e0147224. [Google Scholar] [CrossRef] [PubMed]
- Porter, M.K.; Steinberg, A.P.; Ismagilov, R.F. Interplay of motility and polymer-driven depletion forces in the initial stages of bacterial aggregation. Soft Matter 2019, 15, 7071–7079. [Google Scholar] [CrossRef]
- Yánez-Mendizábal, V.; Falconi, C. Efficacy of Bacillus spp. to biocontrol of anthracnose and enhance plant growth on Andean lupin seeds by lipopeptide production. Boil. Control. 2018, 122, 67–75. [Google Scholar] [CrossRef]
- Jayakumar, A.; Krishna, A.; Mohan, M.; Nair, I.C.; Radhakrishnan, E.K. Plant Growth Enhancement, Disease Resistance, and Elemental Modulatory Effects of Plant Probiotic Endophytic Bacillus sp. Fcl1. Probiotics Antimicrob Proteins 2018, 11, 526–534. [Google Scholar] [CrossRef]
- Lou, Y.; Guo, Q.; Peng, C.; Shi, M.D.; Li, H.Y.; Li, X.; Xue, Q.H.; Lai, H.X. Effects of three Bacillus strains on growth promoting and rhizosphere soil microflora of tomato. J. Appl. Ecol. 2018, 29, 260–268. [Google Scholar]
- Liu, Y.; Chen, L.; Wu, G.; Feng, H.; Zhang, G.; Shen, Q.; Zhang, R. Identification of Root-Secreted Compounds Involved in the Communication Between Cucumber, the Beneficial Bacillus amyloliquefaciens, and the Soil-Borne Pathogen Fusarium oxysporum. Mol. Plant Microbe Interact. 2017, 30, 53–62. [Google Scholar] [CrossRef] [PubMed]
- Sapre, S.; Gontia-Mishra, I.; Tiwari, S. Klebsiella sp. confers enhanced tolerance to salinity and plant growth promotion in oat seedlings (Avena sativa). Microbiol. Res. 2018, 206, 25–32. [Google Scholar] [CrossRef] [PubMed]
- Houry, A.; Briandet, R.; Aymerich, S.; Gohar, M. Involvement of motility and flagella in Bacillus cereus biofilm formation. Microbiology 2010, 156, 1009–1018. [Google Scholar] [CrossRef] [PubMed]
- Zeriouh, H.; De Vicente, A.; Pérez-García, A.; Romero, D.; Romero, D. Surfactin triggers biofilm formation ofBacillus subtilisin melon phylloplane and contributes to the biocontrol activity. Environ. Microbiol. 2013, 16, 2196–2211. [Google Scholar] [CrossRef]
- Cao, Y.; Pi, H.; Chandrangsu, P.; Li, Y.; Wang, Y.; Zhou, H.; Xiong, H.; Helmann, J.D.; Cai, Y. Antagonism of Two Plant-Growth Promoting Bacillus velezensis Isolates Against Ralstonia solanacearum and Fusarium oxysporum. Sci. Rep. 2018, 8, 4360. [Google Scholar] [CrossRef]
- Nie, P.; Li, X.; Wang, S.; Guo, J.; Zhao, H.; Niu, D. Induced Systemic Resistance against Botrytis cinerea by Bacillus cereus AR156 through a JA/ET-and NPR1-Dependent Signaling Pathway and Activates PAMP-Triggered Immunity in Arabidopsis. Front. Plant Sci. 2017, 8, 759. [Google Scholar] [CrossRef]
- Liu, Y.; Feng, H.; Fu, R.; Zhang, N.; Du, W.; Shen, Q.; Zhang, R. Induced root-secreted D-galactose functions as a chemoattractant and enhances the biofilm formation of Bacillus velezensis SQR9 in an McpA-dependent manner. Appl. Microbiol. Biotechnol. 2019, 104, 785–797. [Google Scholar] [CrossRef]
- Zhang, N.; Wang, D.; Liu, Y.; Li, S.; Shen, Q.; Zhang, R. Effects of different plant root exudates and their organic acid components on chemotaxis, biofilm formation and colonization by beneficial rhizosphere-associated bacterial strains. Plant Soil 2013, 374, 689–700. [Google Scholar] [CrossRef]
| Primer a | Sequence (5′-3′) |
|---|---|
| epsA-qF | GCCACAAGTACAGAAAATGAGTC |
| epsA-qR | GATGTGAACGAATTTACGTGAGT |
| tasA-qF | GAATTCCTCAGCCAATTTCAG |
| tasA-qR | GGTGGCAACATTAATTTTACCA |
| fliG-qF | CAGCATGAGCATCCACAAAC |
| fliG-qR | GGTATAATCTTGCGTAAAGGTTG |
| sfp-qF | GAGAATATCACCGGAATTGAAAA |
| sfp-qR | GCTTTCCTTCCAGCCATAGC |
| gyrB-qF | GAGGGAGTCGGTAATGGTTCTT |
| gyrB-qR | CGAAGCTGGCTTTAAAACCG |
| Primer b. | Sequence (5′-3′) |
|---|---|
| PR2-qF | CGACAACATTCGCCCCTTCT |
| PR2-qR | CTGCAGCGCGGTTTGAATAT |
| PAL-qF | CCCCTGCTCGGTTTTGCTCTG |
| PAL-qR | TGGCCTGCGTCATTTCTGCTG |
| COI-qF | GCACCTGACATGGCATTTCC |
| COI-qR | GATAGACCGTCCACCCAGTGC |
| GPX-qF | TTCACAGTCAAGGACATCAGGGGTAA |
| GPX-qR | CACGGGAAAGCCAGTATTTCTAAC |
| U2af-qF | TCGGGAGGTTGGGTCTACAT |
| U2af-qR | ACCAGTCCTTCAGTCCCCTT |
| Treatment | Disease Incidence (%) | Biocontrol Efficacy (%) |
|---|---|---|
| Blank | 0 | -- |
| CK + Rsl | 81.3 ± 1.02 a | -- |
| WT + Rsl | 18.8 ± 1.52 d | 76.88 ± 1.42 a |
| MA15 + Rsl | 43.75 ± 1.37 b | 46.19 ± 1.28 c |
| MA1330 + Rsl | 31.25 ± 1.46 c | 61.56 ± 1.25 b |
| MA4828 + Rsl | 31.25 ± 1.33 c | 61.56 ± 1.21 b |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhu, M.-L.; Wu, X.-Q.; Wang, Y.-H.; Dai, Y. Role of Biofilm Formation by Bacillus pumilus HR10 in Biocontrol against Pine Seedling Damping-Off Disease Caused by Rhizoctonia solani. Forests 2020, 11, 652. https://doi.org/10.3390/f11060652
Zhu M-L, Wu X-Q, Wang Y-H, Dai Y. Role of Biofilm Formation by Bacillus pumilus HR10 in Biocontrol against Pine Seedling Damping-Off Disease Caused by Rhizoctonia solani. Forests. 2020; 11(6):652. https://doi.org/10.3390/f11060652
Chicago/Turabian StyleZhu, Mei-Ling, Xiao-Qin Wu, Ya-Hui Wang, and Yun Dai. 2020. "Role of Biofilm Formation by Bacillus pumilus HR10 in Biocontrol against Pine Seedling Damping-Off Disease Caused by Rhizoctonia solani" Forests 11, no. 6: 652. https://doi.org/10.3390/f11060652
APA StyleZhu, M.-L., Wu, X.-Q., Wang, Y.-H., & Dai, Y. (2020). Role of Biofilm Formation by Bacillus pumilus HR10 in Biocontrol against Pine Seedling Damping-Off Disease Caused by Rhizoctonia solani. Forests, 11(6), 652. https://doi.org/10.3390/f11060652





