Abstract
Background and Objectives: Polyploidisation and frequent hybridisation play an important role in speciation processes and evolutionary history and have a large impact on reproductive systems in the genus Crataegus. Reproductive modes in selected diploid and polyploid taxa in eastern Slovakia were investigated and analysed for the first time. Materials and Methods: Diploid, triploid, and tetraploid hawthorns were tested for self-pollination, self-compatibility, and self-fertilisation. Pollination experiments were performed within and between diploid and triploid species to determine the possibilities and directions of pollen transfer under natural conditions. Seeds from crossing experiments and open pollinations were analysed using the flow cytometric seed screen method. Results: These experiments demonstrated that sexual reproduction, cross-pollination, and self-incompatibility are typical of the diploid species Crataegus monogyna and C. kyrtostyla. Seeds produced by self-fertile tetraploid C. subsphaerica were derived from both meiotically reduced and unreduced megagametophytes. Conclusions: Experimental results concerning triploid C. subsphaerica and C. laevigata × C. subsphaerica are ambiguous but suggest that seeds are almost exclusively created through apomixis, although a few sexually generated seeds were observed. In the genus Crataegus, pseudogamy is a common feature of polyploid taxa, as in all cases pollination is essential for regular seed development. Research Highlights: We suggest that all studied Crataegus taxa produce reduced pollen irrespective of ploidy level. Moreover, we emphasise that triploids produce apparently aneuploid pollen grains as a result of irregular meiosis. They are also capable of utilising pollen from 2x, 3x, or 4x donors for pseudogamous formation of endosperm.
1. Introduction
Polyploidisation and hybridisation have played an important role in the evolution and speciation of angiosperms [1,2,3,4,5,6,7,8]. Both processes can often affect not only genomic, morphological, and physiological properties in plants [9] but are also associated with changes in the reproductive system [10,11,12,13,14]. In certain species, the shift from diploid to polyploid biotypes can be correlated with changes from sexuality to asexuality [12,15,16] or from self-incompatibility to self-compatibility [17,18,19,20,21,22]. Most asexually reproducing plants are polyploids and possibly hybrids [23,24]. Sexual reproduction has several benefits in the formation of genetic variability, which allows adaptations to ecological and climatic changes and improves survivability [25,26]. Despite these advantages, many plants, mainly in the families Asteraceae Bercht. et J. Presl, Poaceae Barnhart, and Rosaceae Juss., reproduce apomictically [12]. The success of apomictic plants is mainly because of their ability to occupy more geographical areas than their sexual progenitors [15,27,28,29], to fix successful genotypes [30], and to prevent reduced fitness because of loss of recombination and infertility in odd polyploids [14]. Apomixis (agamospermy) is an asexual reproduction via seeds, resulting in a progeny genetically identical to the maternal plant [31]. It is well known as a way to maintain hybrid genotypes and facilitate reproductive isolation from parental taxa [22,32]. Therefore, apomixis is essential for the formation of new species arising by polyploidy and hybridisation [12,15,33].
Extensive hybridisation, polyploidy, and apomixis have been observed in Crataegus L. (hawthorn), a monophyletic genus [34], which belongs to the family Rosaceae, subfamily Amygdaloideae Arn., tribe Maleae Small. [35] and subtribe Malinae Reveal [36,37,38]. The genus Crataegus is taxonomically complex and the number of species varies between 150 and 1200, depending on the interpretation of species boundaries by different authors [39,40,41,42,43]. The morphological differences between many species are relatively minor [44,45]. Hawthorns are deciduous, small trees and ruderal shrubs primarily distributed in the temperate regions of Europe, Asia, Africa, and North America [41,42]. Hawthorns grow abundantly in regions with high light intensity, alongside rivers or lakes or in anthropogenic areas. Flowers are insect-pollinated, and small, fleshy fruits containing 1–5 seeds [42,46] are an important source of food, not only for different kinds of animals but also for humans [47,48]. Seeds are produced either by allogamy, which is common in diploid hawthorns, when self-fertilisation is prevented by gametophytic self-incompatibility [18,49], or by autogamy and apomixis common in polyploids [50]. Gametophytic self-incompatibility found in the family Rosaceae [51,52,53] is a genetically conditioned biochemical reaction used by plants to prevent self-fertilisation and consequent inbreeding depression [54].
For the genus Crataegus, the occurrence of diploids, triploids, and tetraploids prevails, although pentaploids and one hexaploid have been recorded [55]. Seeds of diploid hawthorns are known to form sexually with triploid endosperm tissue; having the Polygonum type of embryo sac, the endosperm develops from fertilisation of the binucleate central cell [56]. In contrast, triploids and tetraploids primarily reproduce by gametophytic apomixis [10]. In unreduced megagametophytes with the Polygonum type morphology [57], an embryo develops from an unreduced egg parthenogenetically, either without (autonomously) or with pollination where fertilisation of polar nuclei by one or two sperm cell nuclei is essential (pseudogamy) for successful development of endosperm [18,31,58,59]. Many angiosperms require balanced dosages of maternal-to-paternal genome contributions (2m:1p) to the endosperm for successful seed development [14,60]. Overrepresentation of either parental genome could affect embryo and endosperm sizes and lead to endosperm failure and consequently, seed abortion [61,62]. This problem is bypassed in many apomictic Asteraceae, where the development of endosperm does not require fertilisation [31], whereas in some apomictic Poaceae, megagametophytes contain one rather than two central cell nuclei [63]. Fertilisation of the central cell by one or both sperm cells seems to be necessary for apomictic polyploid Crataegus [10]; thus, the endosperm balance requirement appears to be relaxed or absent in apomicts [11].
Extensive research on the genus Crataegus (e.g., sect. Coccineae Loudon and sect. Douglasia Loudon) has been conducted in North America [10,11,45,50,55,64,65,66,67,68], including the first pollination experiments on hawthorns described by Brown [69] and Love and Feigen [70]. Mating systems in European hawthorns have been studied with the emphasis mainly on hybridisation [71,72,73,74] and on embryology [75,76]. We present an analysis of reproductive systems in Crataegus sect. Crataegus occurring in Slovakia (central Europe), where we performed controlled and open pollinations between taxa of diploid and triploid ploidy levels and tested for self-pollinations of tetraploid species to determine the mating system of seed progeny, self-compatibility, and pollen transfer between different ploidy levels. Consequently, the ploidy levels of the embryo and endosperm were evaluated by the flow cytometric seed screen (FCSS), which can reveal whether the embryo sac originated meiotically or apomeiotically and whether fertilisation of the egg cell and central cell had occurred. To further understand the fertilisation process in megagametophytes, the ploidy level of the pollen was determined.
2. Materials and Methods
2.1. Plant Material
The mother trees used for pollination experiments in this paper, Crataegus monogyna Jacq., C. subsphaerica Gand. s.l., C. kyrtostyla Fingerh. (C. monogyna × C. subsphaerica), and hybrid C. laevigata × C. subsphaerica, originated from three localities in eastern Slovakia (Table 1): Košice, Botanical garden of Pavol Jozef Šafárik University (population code BOTZ), Prešov, south-eastern margin of village Hermanovce (GOCAL), and north-eastern margin of village Hermanovce (PS-VYH, VYH). Trees were selected for their accessibility and had inflorescences easily reachable from the ground. Voucher specimens of 13 trees (out of 14) used in our experiments are deposited in the herbarium of Botanical Garden of P. J. Šafárik University in Košice (KO). The ploidy levels of all trees and their pollen were investigated (see below). Fruits produced after pollinations were collected from all 14 individual trees. In total, 232 seeds were analysed from 248 mature fruits collected.

Table 1.
Number of seeds produced by controlled (Contr. Poll.) and open pollinations (Open Poll.) which were analysed using the flow cytometric seed screen (FCSS) and localities of the Crataegus individuals analysed in the study. Cytotype diversity at sites based on Kolarčik et al. (unpublished).
2.2. Determination of Ploidy Level of Crataegus Mother Trees and Their Pollen
Flow cytometry (FCM) was used to establish the ploidy levels of 14 hawthorn trees and their pollen. Sample preparation for flow cytometry was done as per the modified protocol of Loureiro et al. [77]. Briefly, cell nuclei were simultaneously isolated from the leaves or petioles of target individual and the reference standard Solanum lycopersicum L. ’Stupnické polní tyčkové rané‘ (2C = 1.96 pg DNA, [78]) in 1 mL of general purpose buffer (GPB, [77]) in Petri dish by using the chopping technique [79]. The suspension was then filtered through a 42 µm nylon filter and supplemented with 2 µL β-mercaptoethanol, 30 µL RNAse and 30 µL propidium iodide. After approximately 30 min incubation at 4 °C, the fluorescence signal was measured by using a Partec CyFlow ML (Partec Gmbh, Münster, Germany) flow cytometer (Institute of Biological and Ecological Sciences, P. J. Šafárik University in Košice, Slovakia) equipped with a 532 nm (150 mW) green laser. Measurements were performed with FloMax ver. 2.70 (Partec Gmbh, Münster, Germany). The quality of histograms was ascertained according to the following criteria: 1300 particles recorded, CV below 5% and symmetric peaks of both sample and standard. DNA content (2C value) was determined according to the following formula: DNA amount in the sample = DNA amount in the internal standard × ((G0/G1 peak mean of sample)/(G0/G1 peak mean of internal standard)) (G0/G1 refers to the population of nuclei in G0 or G1 phases of the cell cycle). The ploidy level was then inferred on the basis of the obtained DNA amount interpreted with the help of the DNA amount ranges for the cytotypes provided by Talent and Dickinson [55]: 1.37–1.67 pg for diploids, 2.05–2.51 pg for triploids, 2.74–3.34 pg for tetraploids, and 3.42–4.18 pg for pentaploids.
Pollen was obtained from all studied mother trees, and nuclei were extracted using the general filter bursting method [80]. Whole flowers (3–5) with dehiscing anthers were placed in 0.5 ml GPB and then vortexed for a few seconds. Flowers were removed, and the pollen suspension was passed through a first filter into a clean tube to prevent debris from entering the sample. This filter (100 µm) was large enough to allow pollen to pass through. The suspension was then passed through a second filter (30 µm), which had pore size small enough that the pollen grains were collected on it but nuclei could pass through. The pollen grains were then gently rubbed against the filter for 20 s, using a glass rod with a rounded end. GPB (0.5 ml) was added for the second time to help nuclei pass through the filter. Next, the nuclei were stained and incubated as described above and the samples were analysed using flow cytometry. To determine the ploidy level of pollen grains, the signal of somatic nuclei from the same individual was then used as a reference for pollen grain nuclei signals.
2.3. Pollination Experiments
Pollination experiments were performed between diploid and triploid Crataegus individuals (Figure 1) to observe pollen transfer between different ploidy levels. Flowers of two tetraploid C. subsphaerica trees were bagged in pollinator exclusion bags to prevent cross-pollination and to determine whether tetraploids are self-compatible and able to self-fertilise.
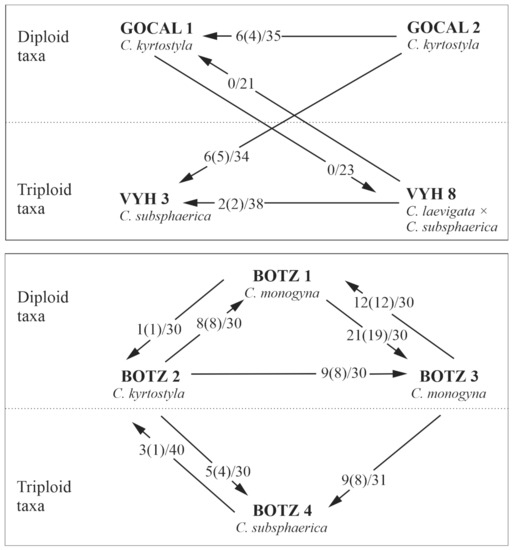
Figure 1.
Schemes of pollination experiments on selected diploid and triploid Crataegus individuals. Arrows represent the direction of pollen transfer to the mother tree, and fractions indicate the number of seeds produced in all the treated flowers (number of seeds successfully measured using FCSS is given in brackets).
The pollinations included (a) self-pollination to test for self-incompatibility (intact flowers bagged and left for self-pollination); (b) tree pollinated by pollen of different cytotype (cross-pollination); (c) tree pollinated by pollen of the same cytotype but different species (cross-pollination); (d) tree pollinated by its own pollen (controlled self-pollination), and (e) flowers left for pollination without the use of pollinator exclusion bags (open pollination). Firstly, flowers at the ‘balloon’ stage (Figure 2) were collected from all studied trees, knowing cross-pollination had not happened yet. Flowers were left to dry overnight to release pollen, which was used for pollination the following day. Consequently, manual pollination of flowers (at the ‘balloon’ stage) was performed by brushing dehiscing anthers against stigmas of mother trees until pollen could be seen on the surface of the stigma. Immediately after pollination, inflorescences were covered with pollinator exclusion bags to prevent cross-pollination. In these pollination experiments, emasculation was not done because it could cause parthenocarpy [79,81,82] and possibly affect reproduction [11]. Fruit formation was monitored, and fruits were collected when they turned red (ripe). Seeds were used to determine reproductive modes using the flow cytometric seed screen method [83]. In total, the number of pollinated flowers was 1354, whereas the number of flowers pollinated on one tree was approximately 180, and certain flowers were left for open pollination. Seeds from experimentally manipulated and open-pollinated flowers were counted and monitored for viability; only large, firm seeds filled with white embryo and endosperm were considered viable.
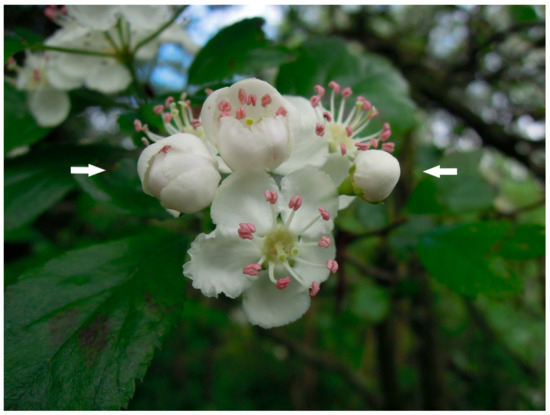

Figure 2.
Flowers of C. laevigata × C. subsphaerica at different stages of opening; flowers at the ‘balloon’ stage before anthesis used for pollination experiments are indicated by white arrows.
2.4. The Flow Cytometric Seed Screen
The flow cytometric seed screen has become an important method for identification of reproductive modes in flowering plants [83]. It provides information about embryo sac origin by estimating embryo and endosperm DNA content. This method thus facilitates distinction between meiotically and apomeiotically developed megagametophytic, parthenogenetic, and zygotic origins of an embryo and autonomous or pseudogamous development of endosperm and detection of meiotic irregularities [82,83,84].
Flow cytometry was used to analyse 232 seeds from collected fruits. The seeds were either analysed whole (sample containing both embryo and endosperm tissue) or the endosperm and embryo were prepared separately. The same FCM protocol as described above was applied. The nuclei of endosperm may sometimes be hidden among nuclei of the reference standard on FCM records in cases when the whole seed is used for sample preparation. Therefore, one of the reference standards (Solanum pseudocapsicum L., 2C = 2.59 pg DNA [85]; Solanum lycopersicum) was used for the determination of embryo ploidy level. In few cases, the ploidy level of plants (sample + reference standard) was analysed via preparation of ‘bulked‘ samples consisting of two, three, or even five individual seeds to accelerate the process. Ploidy levels determined in this way then served as a reference for the determination of endosperm ploidy level in separate endosperm + embryo measurements (without the reference standard). In cases where the whole embryo was used for the first sample, the second sample contained endosperm and reference standard tissue. Estimated DNA ploidies of endosperm and embryo were acquired and compared to determine the mode of reproduction for each analysed seed. The potential presence of endopolyploidy in seed tissues may generally compromise the results of FCSS measurements [84,86]; however, this was shown to be minimal in hawthorn seeds, allowing accurate identification of embryo and endosperm nuclei on FCSS records [10].
3. Results and Discussion
3.1. Cytogenetic Screening of Mother Trees and Their Progeny
Analysis of the DNA content and ploidy level of both leaves and seeds yielded good-quality histograms with high-resolution peaks (Figure 3). Results of ploidy level determinations for mother trees are given in Table 1. Apart from 15 seeds where the embryo or endosperm signals were not detected (and therefore omitted from further analyses), endosperm and embryo ploidy levels could be very well distinguished (Supplementary Materials Table S1 and Table S2).
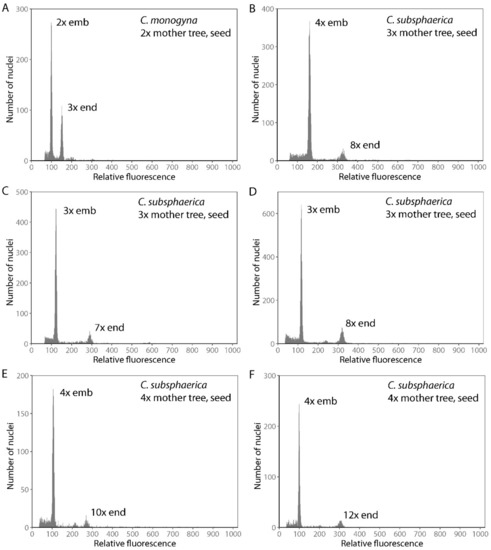
Figure 3.
Selected flow cytometric histograms of Crataegus seeds that originated sexually (A,B) and apomictically (C–F). The first peak represents the embryo nuclei and their ploidy level and the second peak represents the endosperm nuclei and their ploidy level.
3.2. The Flow Cytometric Screen of Crataegus Pollen
FCM screening of pollen always allows recognition of two peaks, which correspond to the nuclei of vegetative and generative cells. The quality of pollen FCM records was slightly lower than that of FCM records of somatic cells. Clearly, diploid mother trees produced reduced pollen, as evidenced by the presence of 1x vegetative and 2x generative nuclei on FCM histograms (Figure 4; mitosis of the generative cell has already occurred in mature binucleate pollen grain typical for Rosaceae [80,87,88]). Similarly, polyploids also produced reduced pollen; we recorded approximately 1.5x vegetative and 3x generative nuclei in triploids and further, 2x vegetative and 4x generative nuclei in tetraploids. We did not record any ’4x‘, ’6x‘, or ’8x‘ peaks on FCM histograms for diploid, triploid, and tetraploid hawthorns, respectively, which could potentially correspond to generative nuclei of unreduced pollen. This indicates that unreduced pollen is extremely rarely produced in investigated hawthorns. To our knowledge, this is the first ploidy level investigation of pollen grains in Crataegus except the successful test on C. macrosperma Ashe performed by Kron and Husband [80]. Comparable results were obtained in Sorbus L., a genus with similar breeding systems to Crataegus [89]. Their data suggest production of reduced pollen and absence of unreduced pollen in studied taxa.
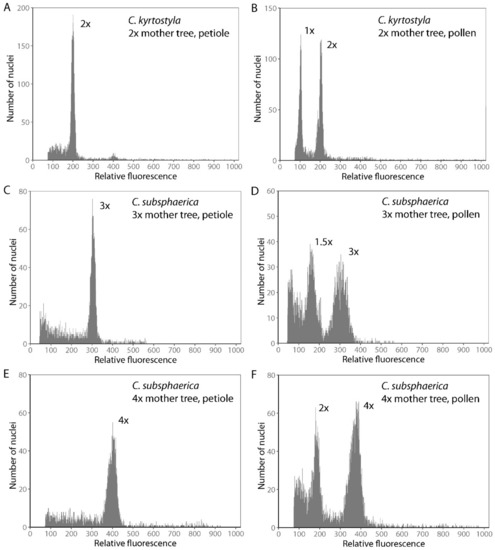
Figure 4.
Selected flow cytometric histograms of pollen grains of diploid (A,B), triploid (C,D), and tetraploid (E,F) mother trees. Histograms on the left represent the FCM of petiole nuclei and histograms on the right represent the FCM of pollen nuclei where the first peak corresponds to vegetative cell nuclei and the second peak to generative cell nuclei.
3.3. Pollination Experiments
In total, 13 pollinations were performed between diploid and triploid Crataegus trees, and flowers from two tetraploid trees were covered with pollinator exclusion bags and tested for self-pollination and self-compatibility. Two out of 13 cross-pollinations resulted in no seeds (Figure 1), including reciprocal crosses of diploid C. kyrtostyla (GOCAL 1) with the triploid hybrid C. laevigata × C. subsphaerica (VYH 8). The experiment on the tree GOCAL 2 was destroyed, and no seeds from this individual were obtained. The reproductive systems in diploid, triploid, and tetraploid trees were also observed with open pollinations.
3.4. Seed-Set Analysis and Reproductive Modes in Diploid Crataegus Mother Trees
Pollination experiments in diploid C. monogyna and C. kyrtostyla included self-pollinations, open pollinations, and pollinations with pollen from diploid and triploid species. Self-pollinations when inflorescences were enclosed in pollinator exclusion bags resulted in four seeds out of 606 pollinated flowers/648.4 carpels (pyrenes in fruits, approximately 0.7%/0.6%; number of pyrenes per fruit was comparable in diploids, 1.07 pyrenes per fruit (n = 383 analysed fruits)). Note, that only two seeds were measured successfully using FCSS. The result suggests self-incompatibility in diploid Crataegus taxa. Diploid C. monogyna was considered to be self-incompatible in other studies as well [81,90,91,92]. Self-incompatibility in the family Rosaceae is of the gametophytic type [52,53], defined by germination of the pollen grain on the stigma and consequent cessation of pollen tube growth in the style [93]. Both seeds were analysed by the FCSS method and yielded the diploid signal for the embryo (1.46, 1.48 pg DNA) and the triploid signal for the endosperm (2.22, 2.24 pg DNA), as expected in normal sexual reproduction. Similar results were reported in a North American experiment [10], where only 2% of self-pollinated flowers (from diploid C. punctata Jacq. and C. monogyna) produced fruits, and seeds originated sexually.
Pollination of a diploid mother tree with pollen from another diploid individual of the same species (C. monogyna × C. monogyna and C. kyrtostyla × C. kyrtostyla) resulted in 39 seeds out of 95 pollinated flowers/102 carpels (approximately 36.8%/34.3%, note: 35 seeds measured successfully using FCSS). Pollination of diploid C. kyrtostyla by diploid C. monogyna resulted in only one seed out of 30 pollinated flowers/32 carpels (approximately 3.3%/3.1%) while pollination of diploid C. monogyna by diploid C. kyrtostyla was more successful and resulted in 17 seeds out of 60 pollinated flowers/64 carpels (approximately 28.3%/26.6%; note: Sixteen seeds measured successfully using FCSS). After analysis, all produced seeds yielded the diploid embryo signal and the triploid endosperm signal, which indicates sexual reproduction. In these seeds, the DNA content ranged from 1.40 to 1.57 pg for diploid embryos and from 2.11 to 2.38 pg for triploid endosperms. Pollination of diploid C. kyrtostyla by triploid C. subsphaerica resulted in three seeds from 40 pollinated flowers/42.8 carpels (approximately 7.5%/7.0%). Only a single seed yielded a good-quality histogram and it originated sexually, with 2x embryo (1.56 pg) and 3x endosperm (2.33 pg). We speculate that this seed originated via self-fertilisation; note that we did not remove anthers from diploid mother trees. We analysed the ploidy level of the embryo and endosperm in 72 seeds to determine modes of reproduction in diploid Crataegus taxa without our interference (open pollination). After separate analysis of each seed, we observed diploid peaks for the embryo and triploid peaks for the endosperm on every histogram, which proves that seeds of diploid trees originate exclusively sexually, and double fertilisation by reduced 1x pollen occurred (Table 2). The ploidy-level screening of pollen from diploid trees performed in the present study supports this explanation. Sexual reproduction in diploid Crataegus taxa was also reported in other studies [10,11,75]. Sexual reproduction in diploids is rather a rule among angiosperms, and only rarely exceptions have been evidenced, e.g., in Paspalum L. [94] or Boechera Á.Löve et D.Löve [95,96,97]. In the Rosaceae family, where Crataegus belongs, diploids are mostly sexual, as observed by FCM in Amelanchier Medik. [98], Potentilla L. s.l. [82], Rubus L. [25,99], and Sorbus [100], but few exceptions were reported [e.g., 82,98].

Table 2.
Summary of results of pollination experiments in the genus Crataegus, including type of crosses, number of seeds and pollinations, hypothesized seed, embryo and endosperm origin, and embryo, endosperm, egg, central cell, and pollen ploidies. In case of alternative explanations, the most probable is given in bold.
3.5. Seed-Set Analysis and Reproductive Modes in Triploid Crataegus Mother Trees
Similarly, as in diploid Crataegus taxa, self-pollination in triploid C. subsphaerica and C. laevigata × C. subsphaerica resulted in no viable seeds out of 282 pollinated flowers. We suggest that this result indicates poor pollen quality resulting from irregular meiosis in triploid species (see Section 3.2, Supplementary Materials Figure S1), although gametophytic self-incompatibility or general lower fruit production in triploids (as a result of shift in flowering phenology and consequent period of unfavourable climatic conditions) are also possible explanations. Although cross-pollinations of the 3x × 3x type resulted in only two seeds out of 38 pollinated flowers, pollen viability tests would still be necessary to exclude male infertility. Male sterility (no pollen produced) and male infertility (i.e., poorly stainable pollen produced) were recorded in triploid Crataegus [11]. Two seeds (38 pollinated flowers/40.7 carpels, approximately 5.3%/4.9%; number of pyrenes per fruit was 1.07 (n = 297 analysed fruits)) were produced from pollination of a C. subsphaerica mother tree by C. laevigata × C. subsphaerica (3x × 3x). Flow cytometry revealed a 3x embryo signal (2.40, 2.46 pg) and a 9x endosperm signal (7.20, 7.29 pg). Seeds with 3x embryos and 9x endosperms were also recorded in other studies in the genera Crataegus [66] and Sorbus [89,100], where triploids use not only 1x or 2x but also 1.5x or 3x sperm cells for fertilisation. This indicates their apomictic origin when embryo developed parthenogenetically and endosperm possibly resulted from fertilisation of binucleate central cell (6x) either by one unreduced sperm cell (3x) or by two sperm cells with the aneuploid cytotype (approximately 1.5x). Our pollen ploidy-level screening indicates that the 3x trees produced regular pollen with approximately 1.5x. High CV values for both peaks (of vegetative nuclei and generative nuclei), compared with that for the peak of somatic nuclei (see Figure 4), may be a result of the presence of various aneuploid pollen grains ranging from 1x to 2x, with the highest probability of 1.5x. These data suggest that some irregular male meiosis occurs in triploids. Triploids are generally meiotically unstable and can often produce aneuploid gametes with incomplete or unbalanced chromosome sets [101,102,103]. In polyploid plants, aneuploidy is an extremely frequent phenomenon [104,105] and especially odd-numbered polyploids or hybrids may have pollen with high rates of aneuploidy [80]. Aneuploid pollen grains were recorded, e.g., in triploid Arabidopsis thaliana L. (Heynh.), and proven to be fertile after crossing experiments with diploids and tetraploids [106] and in allotriploid poplar (Populus alba × P. berolinensis ‘Yinzhong’), where pollen fertility is also suggested [107].
Pollinations of triploid individuals with pollen from diploids resulted in 20 well-developed seeds. All originated only on C. subsphaerica, out of all 95 pollinated flowers/100.7 carpels (approximately 21.1%/19.7%, note: 17 seeds measured successfully using FCSS; C. subsphaerica (1.06 pyrenes per fruit, n = 297 analysed fruits), C. laevigata × C. subsphaerica (2.11 pyrenes per fruit, n = 299 analysed fruits)). Seeds were found with different ploidy levels of embryos and endosperms. In this case, we can observe that triploids of the genus Crataegus are almost exclusively apomictic because only three seeds originated sexually, whereas 14 seeds were produced via apomixis. Two seeds with aneuploid approximately 3.7x embryos and approximately 5x endosperms possibly originated sexually when the egg cell and central cell were fertilised by unreduced sperm cells. A single seed, which yielded a 4x embryo signal and a second 8x signal, probably originated sexually (BIII hybrid) when the egg cell was fertilised with a 1x sperm cell, and the second signal might be either the G2 phase of the embryo nuclei or, less probably, the endoreduplicated nuclei (Figure 4B). Fertilisation of an unreduced triploid egg cell, although rare because of the probable low viability of the seeds, was documented in the genera Crataegus [10,11] and Sorbus [89,100]. In pseudogamous apomictic plants, if the unreduced egg cell is fertilised by a sperm cell, ploidy levels can increase [10,31,108,109]. The other 14 seeds were produced via apomixis with 3x embryos and 6x, 7x, and 8x endosperms. One seed with a 3x embryo and 6x endosperm possibly originated autonomously without fertilisation by sperm cells in unreduced megagametophyte, or the 6x peak could represent the G2 phase of embryonic nuclei or endoreduplicated nuclei. The origin of another single seed was probably pseudogamous apomixis when a 3x embryo developed parthenogenetically and 7x endosperm was produced from fertilisation of the binucleate central cell by a single 1x sperm cell. The majority of seeds were derived from meiotically unreduced megagametophytes: A 3x embryo developed parthenogenetically and 8x endosperm originated from the fertilisation of the binucleate central cell (6x) by two reduced sperm cells (1x), originating from diploid pollen donors (or by one unreduced sperm cell (2x), but note extremely low production of unreduced pollen grains in diploids).
Similarly, as in diploids, we analysed ploidy levels of embryos and endosperms in triploid individuals produced from open pollinations. The analysis of open-pollinated seeds yielded 3x embryos and endosperms with five different ploidy levels (5x, 7x, 8x, 9x, and 10x), and the 8x endosperm was the most common. All seeds are probably of apomictic origin. A single seed with a 3x embryo and 5x endosperm probably originated via apomixis where the embryo developed parthenogenetically, while the endosperm resulted from fertilisation of a single, 3x central cell with two reduced sperm cells (1x) from a diploid. This seed could have also originated sexually, where the egg cell and the central cell were fertilised by reduced sperm cells (Table 2). The inferred origin of seeds with 3x embryos and 7x, 8x, and 9x endosperms corresponds to the same cases in 3x × 3x crosses mentioned above. The origin of a single seed with a triploid embryo and decaploid endosperm is consistent with a binucleate central cell and two reduced 2x sperm cells contributing to the endosperm (originating most probably from 4x plants). Seeds with 3x embryos and 10x endosperms are rare but have been recorded in the genera Crataegus [10,11,66] and Sorbus [89,100]. The majority of seeds in pollination experiments in triploid Crataegus are suggested to develop unreduced megagametophytes, and the observed variation in ploidy levels of embryo and endosperm stems from the natural cross-compatibility of triploids; they can utilise pollen from 2x, 3x, and possibly even from 4x donors (Table 2) for pseudogamous development of functional endosperm.
3.6. Seed-Set Analysis and Reproductive Modes in Tetraploid Crataegus Mother Trees
Self-pollination experiments were performed in tetraploid Crataegus individuals where intact flowers of tetraploid C. subsphaerica that were enclosed in pollinator exclusion bags produced 24 seeds out of 101 flowers/102 carpels (approximately 23.8%/23.5%; number of pyrenes per fruit was comparable in tetraploids, 1.01 pyrenes per fruit (n = 164 analysed fruits)). Note, that histograms of three seeds did not yield embryo or endosperm signals. After FCSS analysis, histograms of 21 seeds yielded tetraploid embryo signals (3.22–3.32 pg DNA) and 10x or 12x endosperm signals (8.02–10.30 pg DNA), which indicates the formation of unreduced megagametophytes. The majority of seeds developed endosperms with the 12x ploidy level. Embryos developed parthenogenetically wherein the binucleate central cell was fertilised (pseudogamy) by one meiotically reduced sperm cell (10x) or by one unreduced or two reduced sperm cells (12x). These experiments indicate that tetraploid Crataegus taxa are pseudogamous apomicts able to self-pollinate and that pollination is essential for seed formation, as reported in other studies [10,11,110].
Open pollination resulted in 30 seeds with different ploidy levels of embryos: Most were tetraploid, one was diploid, and another one was hexaploid. The endosperm ploidy levels greatly varied; 6x, 8x, 10x, 11x, 12x, 14x, and 16x were recorded. Open-pollinated apomictic tetraploids as well as triploids seem to always require fertilisation of the central cell involving either one or two sperm cells [10,11]. The majority of seeds produced by open pollination are suggested to be apomictic: 4x embryos (and a single 6x embryo with 10x endosperm) with 10x, 11x, 12x, 14x, and 16x endosperms were recorded. The origin of seeds with 10x and 12x endosperms is proposed above. A single seed had 11x endosperm, suggesting fertilisation with pollen from triploids (either one unreduced 3x sperm cell or two reduced, probably approximately 1.5x sperm cells). Endosperm ploidy levels higher than 12x can be explained by the presence of a trinucleate central cell, which was observed in Crataegus [111]. Endosperms of these four seeds are then suggested to develop through fertilisation of a trinucleate central cell by either one reduced 2x sperm cell (14x) or by one unreduced 4x or two reduced 2x sperm cells (16x). A single seed with a 6x embryo and 10x endosperm was probably derived from fertilisation of both an unreduced egg cell and a binucleate central cell by a reduced 2x sperm cell. Seeds with 6x embryos representing BIII hybrids seem to be rare [10], and the prevailing ploidy level of the embryo in tetraploids is 4x. In addition to seeds of apomictic origin, reduced megagametophytes were observed in tetraploid seeds. A single seed with a 4x embryo and 6x endosperm was likely to have resulted from a reduced megagametophyte where both the egg cell and the central cell were fertilised by reduced 2x sperm cells. Another seed with a reduced embryo sac was recorded where the embryo ploidy level was 2x and could therefore have developed parthenogenetically, whereas the endosperm with ploidy level 8x was derived from fertilisation of the central cell by either one unreduced 4x sperm cell or two reduced 2x sperm cells. These results indicate that tetraploid Crataegus subsphaerica frequently produce unreduced megagametophytes, but sometimes sexual megagametophytes occur (Table 2).
4. Conclusions
In this first analysis of reproductive systems of Crataegus taxa in Europe and Crataegus sect. Crataegus overall (except C. monogyna introduced to North America), we performed controlled and open pollinations to establish that the diploid species C. monogyna and C. kyrtostyla are self-incompatible; however, after being pollinated by pollen from other diploid or triploid trees, they produced viable seed progeny. The flow cytometric seed screen method was used to identify ploidy levels of trees and their pollen grains and modes of reproduction, providing fast, accurate analysis with relatively low expanses. Flow cytometry analyses demonstrated that seeds produced by both controlled and open pollinations originated sexually in these diploids. Our data suggest that polyploid taxa are almost exclusively pseudogamous apomicts. The results of self-pollination in triploid C. subsphaerica and C. laevigata × C. subsphaerica reflect either self-incompatibility or more likely, poor pollen quality. Conversely, tetraploid C. subsphaerica can use their own pollen for fertilisation, i.e., they are self-compatible. To determine reproductive systems, particularly in the polyploid Crataegus, measurements of the ploidy levels of pollen used for fertilisation are essential. We observed that mother trees more frequently produce reduced pollen, as evidenced on FCM histograms. The study indicates that reproductive systems of central European diploid and polyploid Crataegus species of sect. Crataegus are similar to those of hawthorns in North America of sect. Douglasia, sect. Coccineae, sect. Crus-galli Loudon or sect. Pruinosae (Sarg.) Eggl. Apomixis and polyploidisation documented in the study, as well as the hybridisation inferred from morphological comparisons performed in other studies, are probably responsible for the taxonomic complexity of European hawthorns. This genus can provide valuable insights into reproductive interactions between cytotypes. The next step should be to determine the interactions among several apomictic tetraploid species with partial sexuality and to experimentally test for interspecific cross-compatibility and hybridisation.
Supplementary Materials
The following are available online at https://www.mdpi.com/1999-4907/10/12/1059/s1. Table S1: FCM results of cross-pollination experiments in studied Crataegus taxa. Table S2: FCM results of open pollinations in studied Crataegus taxa. Figure S1: Light microscopy photographs of Crataegus pollen grains of diploid (C. kyrtostyla), triploid (C. laevigata × C. subsphaerica) and tetraploid (C. subsphaerica) accessions.
Author Contributions
Conceptualization, D.V. and V.K.; Methodology, D.V. and V.K.; Validation, D.V.; Formal analysis, V.K.; Investigation, D.V.; Resources, D.V. and V.K.; Data curation, D.V.; Writing—original draft preparation, D.V.; Writing—review and editing, D.V. and V.K.; Visualization, D.V. and V.K.; Supervision, V.K.; Project administration, V.K.; Funding acquisition, V.K.
Funding
This research was funded by the Grant Agency for Science, Bratislava (VEGA, No. 1/0741/19).
Acknowledgments
We would like to thank anonymous reviewers for their helpful suggestions on the manuscript; Albert Rákai for field assistance and Enago (www.enago.com) for English language editing.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Rieseberg, L.H.; Willis, J.H. Plant speciation. Science 2007, 317, 910–914. [Google Scholar] [CrossRef] [PubMed]
- Doyle, J.J.; Flagel, L.E.; Paterson, A.H.; Rapp, R.A.; Soltis, D.E.; Soltis, P.S.; Wendel, J.F. Evolutionary genetics of genome merger and doubling in plants. Annu. Rev. Genet. 2008, 42, 443–461. [Google Scholar] [CrossRef] [PubMed]
- Soltis, P.S.; Soltis, D.E. The role of hybridization in plant speciation. Annu. Rev. Plant. Biol. 2009, 60, 561–588. [Google Scholar] [CrossRef] [PubMed]
- Jiao, Y.; Wickett, N.J.; Ayyampalayam, S.; Chanderbali, A.S.; Landherr, L.; Ralph, P.E.; Tomsho, L.P.; Hu, Y.; Liang, H.; Soltis, P.S.; et al. Ancestral polyploidy in seed plants and angiosperms. Nature 2011, 473, 97–100. [Google Scholar] [CrossRef] [PubMed]
- Husband, B.C.; Baldwin, S.J.; Suda, J. The incidence of polyploidy in natural plant populations: Major patterns and evolutionary processes. In Plant Genome Diversity: Physical Structure, Behaviour and Evolution of Plant Genomes; Leitch, I.J., Greilhuber, J., Doležel, J., Wendel, J.F., Eds.; Springer: Wien, Austria, 2013; Volume 2, pp. 255–276. [Google Scholar]
- Madlung, A. Polyploidy and its effect on evolutionary success: Old questions revisited with new tools. Heredity 2013, 110, 99–104. [Google Scholar] [CrossRef] [PubMed]
- Alix, K.; Gérard, P.R.; Schwarzacher, T.; Heslop-Harrison, J.S.P. Polyploidy and interspecific hybridization: Partners for adaptation, speciation and evolution in plants. Ann. Bot. 2017, 120, 183–194. [Google Scholar] [CrossRef]
- Šarhanová, P.; Sharbel, T.F.; Sochor, M.; Vašut, R.J.; Dancák, M.; Trávnícek, B. Hybridization drives evolution in Rubus subgenus Rubus: Evidence from microsatellite markers. Ann. Bot. 2017, 120, 317–328. [Google Scholar] [CrossRef]
- Weiss-Schneeweiss, H.; Emadzade, K.; Jang, T.S.; Schneeweiss, G.M. Evolutionary consequences, constraints and potential of polyploidy in plants. Cytogenet. Genome Res. 2013, 140, 137–150. [Google Scholar] [CrossRef]
- Talent, N.; Dickinson, T.A. Endosperm formation in aposporous Crataegus (Rosaceae, Spiraeoideae, tribe Pyreae): Parallels to Ranunculaceae and Poaceae. New Phytol. 2007, 173, 231–249. [Google Scholar] [CrossRef]
- Talent, N.; Dickinson, T.A. The potential for ploidy level increases and decreases in Crataegus (Rosaceae, Spiraeoideae, tribe Pyreae). Can. J. Bot. 2007, 85, 570–584. [Google Scholar] [CrossRef]
- Whitton, J.; Sears, C.J.; Baack, E.J.; Otto, S.P. The dynamic nature of apomixis in the angiosperms. Int. J. Plant Sci. 2008, 169, 169–182. [Google Scholar] [CrossRef]
- Hörandl, E. A combinational theory for maintenance of sex. Heredity 2009, 103, 445–457. [Google Scholar] [CrossRef] [PubMed]
- Hojsgaard, D.; Greilhuber, J.; Pellino, M.; Paun, O.; Sharbel, T.F.; Hörandl, E. Emergence of apospory and bypass of meiosis via apomixis after sexual hybridisation and polyploidisation. New Phytol. 2014, 204, 1000–1012. [Google Scholar] [CrossRef] [PubMed]
- Hörandl, E. The complex causality of geographical parthenogenesis. New Phytol. 2006, 171, 525–538. [Google Scholar] [CrossRef] [PubMed]
- Cosendai, A.C.; Hörandl, E. Cytotype stability, facultative apomixis and geographical parthenogenesis in Ranunculus kuepferi (Ranunculaceae). Ann. Bot. 2010, 105, 457–470. [Google Scholar] [CrossRef][Green Version]
- Marhold, K.; Lihová, J. Polyploidy, hybridization and reticulate evolution: Lessons from the Brassicaceae. Plant Syst. Evol. 2006, 259, 143–174. [Google Scholar] [CrossRef]
- Dickinson, T.A.; Lo, E.; Talent, N. Polyploidy, reproductive biology, and Rosaceae: Understanding evolution and making classifications. Plant Syst. Evol. 2007, 266, 59–78. [Google Scholar] [CrossRef]
- Alix, K.; Joets, J.; Ryder, C.D.; Moore, J.; Barker, G.C.; Bailey, J.P.; King, G.J.; Pat Heslop-Harrison, J.S. The CACTA transposon Bot1 played a major role in Brassica genome divergence and gene proliferation. Plant J. 2008, 56, 1030–1044. [Google Scholar] [CrossRef]
- Husband, B.C.; Ozimec, B.; Martin, S.L.; Pollock, L. Mating consequences of polyploid evolution in flowering plants: Current trends and insights from synthetic polyploids. Int. J. Plant Sci. 2008, 169, 195–206. [Google Scholar] [CrossRef]
- Robertson, K.; Goldberg, E.E.; Igić, B. Comparative evidence for the correlated evolution of polyploidy and self-compatibility in Solanaceae. Evolution 2011, 65, 139–155. [Google Scholar] [CrossRef]
- Ludwig, S.; Robertson, A.; Rich, T.C.G.; Djordević, M.; Cerović, R.; Houston, L.; Harris, S.A.; Hiscock, S.J. Breeding systems, hybridization and continuing evolution in Avon Gorge Sorbus. Ann. Bot. 2013, 111, 563–575. [Google Scholar] [CrossRef] [PubMed]
- Asker, S.E.; Jerling, L. Apomixis in Plants, 1st ed.; CRC Press: Boca Raton, FL, USA, 1992; p. 320. [Google Scholar]
- Simon, J.C.; Delmotte, F.; Rispe, C.; Crease, T. Phylogenetic relationships between parthenogens and their sexual relatives: The possible routes to parthenogenesis in animals. Biol. J. Linn. Soc. 2003, 79, 151–163. [Google Scholar] [CrossRef]
- Šarhanová, P.; Vašut, R.J.; Dančák, M.; Bureš, P.; Trávníček, B. New insights into the variability of reproduction modes in European populations of Rubus subgen. Rubus: How sexual are the polyploid brambles? Sex. Plant Reprod. 2012, 25, 319–335. [Google Scholar] [CrossRef] [PubMed]
- Lovell, J.T.; Aliyu, O.M.; Mau, M.; Schranz, M.E.; Koch, M.; Kiefer, C.; Song, B.-H.; Mitchell-Olds, T.; Sharbel, T.F. On the origin and evolution of apomixis in Boechera. Plant Reprod. 2013, 26, 309–315. [Google Scholar] [CrossRef] [PubMed]
- Stebbins, G.L. Variation and Evolution in Plants; Columbia University Press: New York, NY, USA, 1950. [Google Scholar]
- Hörandl, E. Geographical parthenogenesis: Opportunities for asexuality. In Lost Sex; Schön, I., Martens, K., Van Dijk, P.J., Eds.; Springer: Heidelberg, Germany, 2009; pp. 161–186. [Google Scholar]
- Hörandl, E. The classification of asexual organisms: Old myths, new facts, and a novel pluralistic approach. Taxon 2018, 67, 1066–1081. [Google Scholar] [CrossRef]
- Smith, J.M. The Evolution of Sex; Cambridge University Press: Cambridge, NY, USA, 1978. [Google Scholar]
- Nogler, G.A. Gametophytic apomixis. In Embryology of Angiosperms; Johri, B.M., Ed.; Springer: Berlin/Heidelberg, Germany, 1984; pp. 475–518. [Google Scholar]
- Coyne, J.A.; Orr, H.A. Speciation; Sinauer Associates, Inc.: Sunderland, MA, USA, 2004. [Google Scholar]
- Grant, V. Plant Speciation, 2nd ed.; Columbia University Press: New York, NY, USA, 1981. [Google Scholar]
- Evans, R.; Campbell, C.S. The origin of the apple subfamily (Rosaceae: Maloideae) is clarified by DNA sequence data from duplicated GBSSI genes. Am. J. Bot. 2002, 89, 1478–1484. [Google Scholar] [CrossRef]
- Kalkman, C. Rosaceae. In The Families and Genera of Vascular Plants; Kubitzki, K., Ed.; Springer: Berlin/Heidelberg, Germany, 2004; Volume 6, pp. 343–386. [Google Scholar]
- Campbell, C.S.; Evans, R.C.; Morgan, D.R.; Dickinson, T.A.; Arsenault, M.P. Phylogeny of subtribe Pyrinae (formerly the Maloideae, Rosaceae): Limited resolution of a complex evolutionary history. Plant Syst. Evol. 2007, 266, 119–145. [Google Scholar] [CrossRef]
- Potter, D.; Eriksson, T.; Evans, R.C.; Oh, S.; Smedmark, J.E.E.; Morgan, D.R.; Kerr, M.; Robertson, K.R.; Arsenault, M.; Dickinson, T.A.; et al. Phylogeny and classification of Rosaceae. Plant Syst. Evol. 2007, 266, 5–43. [Google Scholar] [CrossRef]
- Reveal, J.L. Newly required infrafamilial names mandated by changes in the Code of Nomenclature for Algae, Fungi, and Plants. Phytoneuron 2012, 33, 1–32. [Google Scholar]
- Bailey, L.H. The Standard Cyclopedia of Horticulture, 2nd ed.; The Macmillan Company: New York, NY, USA, 1963. [Google Scholar]
- Phipps, J.B.; Robertson, K.R.; Smith, P.G.; Roher, J.R. A checklist of the subfamily Maloideae (Rosaceae). Can. J. Bot. 1990, 68, 2209–2269. [Google Scholar] [CrossRef]
- Christensen, K.I. Revision of Crataegus sect. Crataegus and nothosect. Crataeguineae (Rosaceae-Maloideae) in the Old World. Syst. Bot. Monogr. 1992, 35, 1–199. [Google Scholar] [CrossRef]
- Fineschi, S.; Salvini, D.; Turchini, D.; Pastorelli, R.; Vendramin, G.G. Crataegus monogyna Jacq. and C. laevigata (Poir.) DC. (Rosaceae, Maloideae) display low level of genetic diversity assessed by chloroplast markers. Plant Syst. Evol. 2005, 250, 187–196. [Google Scholar] [CrossRef]
- Depypere, L.; Vander Mijnsbrugge, K.; De Cock, K.; Verschelde, P.; Quataert, P.; Van Slycken, J.; Goetghebeur, P. Indigenous species of Crataegus (Rosaceae-Maloideae) in Flanders (Belgium). An explorative morphometric study. Belg. J. Bot. 2006, 139, 139–152. [Google Scholar] [CrossRef]
- Camp, W.H. The Crataegus problem. Castanea 1942, 7, 51–55. [Google Scholar]
- Zarrei, M.; Stefanović, S.; Dickinson, T.A. Reticulate evolution in North American black-fruited hawthorns (Crataegus section Douglasia; Rosaceae): Evidence from nuclear ITS2 and plastid sequences. Ann. Bot. 2014, 114, 253–269. [Google Scholar] [CrossRef] [PubMed]
- Baranec, T. Crataegus L. In Flóra Slovenska IV/3; Bertová, L., Ed.; Veda: Bratislava, Slovakia, 1992; pp. 465–491. [Google Scholar]
- Li, L.-Z.; Gao, P.-Y.; Song, S.-J.; Yuan, Y.-Q.; Liu, C.-T.; Huang, X.-X.; Liu, Q.-B. Monoterpenes and flavones from the leaves of Crataegus pinnatifida with anticoagulant activities. J. Funct. Foods 2015, 12, 237–245. [Google Scholar] [CrossRef]
- Qiao, A.; Wang, Y.; Xiang, L.; Zhang, Z.; He, X. Novel triterpenoids isolated from hawthorn berries functioned as antioxidant and antiproliferative activities. J. Funct. Foods 2015, 13, 308–313. [Google Scholar] [CrossRef]
- Clapham, A.R.; Tutin, T.G.; Moore, D.M. Flora of the British Isles; Cambridge University Press: Cambridge, UK, 1990. [Google Scholar]
- Lo, E.Y.Y.; Stefanović, S.; Ritland, K.; Dickinson, T.A. Fine-scale comparisons of genetic variability in seed families of asexually and sexually reproducing Crataegus (Hawthorn; Rosaceae). Am. J. Bot. 2010, 97, 1014–1024. [Google Scholar] [CrossRef]
- Lewis, D. Competition and dominance of incompatibility alleles in diploid pollen. Heredity 1947, 1, 85–108. [Google Scholar] [CrossRef]
- De Nettancourt, D. Incompatibility in angiosperms. Sex. Plant Reprod. 1977, 10, 185–199. [Google Scholar] [CrossRef]
- Raspé, O.; Kohn, J.R. S-allele diversity in Sorbus aucuparia and Crataegus monogyna (Rosaceae: Maloideae). Heredity 2002, 88, 458–465. [Google Scholar] [CrossRef] [PubMed]
- Erdelská, O.; Švubová, R.; Mártonfiová, L.; Lux, A. Embryológia krytosemenných rastlín; Veda: Bratislava, Slovakia, 2017; p. 100. [Google Scholar]
- Talent, N.; Dickinson, T.A. Polyploidy in Crataegus and Mespilus (Rosaceae, Maloideae): Evolutionary inferences from flow cytometry of nuclear DNA amounts. Can. J. Bot. 2005, 83, 1268–1304. [Google Scholar] [CrossRef]
- Maheshwari, P. An Introduction to the Embryology of Angiosperms; McGraw-Hill Book Company, Inc.: New York, NY, USA, 1950; pp. 87–95. [Google Scholar]
- Czapik, R. Problems of apomictic reproduction in the families Compositae and Rosaceae. Folia Geobot. Phytotax. 1996, 31, 381–387. [Google Scholar] [CrossRef]
- Muniyamma, M.; Phipps, J.B. Cytological proof of apomixis in Crataegus (Rosaceae). Am. J. Bot. 1979, 66, 149–155. [Google Scholar] [CrossRef]
- Smith, P.G.; Phipps, J.B. Studies in Crataegus (Rosaceae, Maloideae). XIX. Breeding behaviour in Ontario Crataegus series Rotundifoliae. Can. J. Bot. 1988, 66, 1914–1923. [Google Scholar] [CrossRef]
- Scott, R.J. Polyspermy in apomictic Crataegus: Yes and no. New Phytol. 2007, 173, 227–229. [Google Scholar] [CrossRef]
- Scott, R.J.; Spielman, M.; Bailey, J.; Dickinson, H.G. Parent-of-origin effects on seed development in Arabidopsis thaliana. Development 1998, 125, 3329–3341. [Google Scholar]
- Kradolfer, D.; Hennig, L.; Köhler, C. Increased maternal genome dosage bypasses the requirement of the FIS Polycomb Repressive Complex 2 in Arabidopsis seed development. PLoS Genet. 2013, 9, e1003163. [Google Scholar] [CrossRef]
- Savidan, Y.H. Apomixis: Genetics and breeding. Plant Breed. Rev. 2000, 18, 13–86. [Google Scholar] [CrossRef]
- Lo, E.Y.Y.; Stefanović, S.; Dickinson, T.A. Population genetic structure of diploid sexual and polyploid apomicts of Crataegus (hawthorns; Rosaceae) in the Pacific Northwest. Mol. Ecol. 2009, 18, 1145–1160. [Google Scholar] [CrossRef]
- Lo, E.Y.Y.; Stefanović, S.; Dickinson, T.A. Reconstructing reticulation history in a phylogenetic framework and the potential of allopatric speciation driven by polyploidy in an agamic complex in Crataegus (Rosaceae). Evolution 2010, 64, 3593–3608. [Google Scholar] [CrossRef] [PubMed]
- Lo, E.Y.Y.; Stefanović, S.; Dickinson, T.A. Geographical parthenogenesis in Pacific Northwest hawthorns (Crataegus; Rosaceae). Botany 2013, 91, 107–116. [Google Scholar] [CrossRef]
- McGoey, B.V.; Chau, K.; Dickinson, T.A. Stomata size in relation to ploidy level in North American hawthorns (Crataegus, Rosaceae). Madroño 2014, 61, 177–193. [Google Scholar] [CrossRef]
- Coughlan, J.M.; Han, S.; Stefanović, S.; Dickinson, T.A. Widespread generalist clones are associated with range and niche expansion in allopolyploids of Pacific Northwest hawthorns (Crataegus L.). Mol. Ecol. 2017, 26, 5484–5499. [Google Scholar] [CrossRef] [PubMed]
- Brown, H.B. The genus Crataegus with some theories of the origin of its species. Bull. Torrey Bot. Club 1910, 37, 251–260. [Google Scholar] [CrossRef]
- Love, R.; Feigen, M. Interspecific hybridization between native and naturalized Crataegus (Rosaceae) in western Oregon. Madroño 1978, 25, 211–217. [Google Scholar]
- Byatt, J.I. Hybridization between Crataegus monogyna Jacq. and C. laevigata (Poiret) DC. in southeastern England. Watsonia 1975, 10, 253–264. [Google Scholar]
- Christensen, K.I. A biometric study of some hybridizing Crataegus populations in Denmark. Nord. J. Bot. 1983, 2, 537–548. [Google Scholar] [CrossRef]
- Christensen, K.I. A reanalysis of the status of Crataegus eremitagensis, C. raavadensis and C. schumacheri (Rosaceae). Symb. Bot. Ups. 1996, 31, 211–220. [Google Scholar]
- Christensen, K.I.; Zarrei, M.; Kuzmina, M.; Talent, N.; Lin, C.; Dickinson, T.A. Crataegus ×ninae-celottiae and C. ×cogswellii (Rosaceae, Maleae), two spontaneously formed intersectional nothospecies. PhytoKeys 2014, 36, 1–26. [Google Scholar] [CrossRef]
- Ptak, K. Cyto-embryological investigations on the Polish representatives of the genus Crataegus L. I. Chromosome numbers; embryology of diploid and tetraploid species. Acta Biol. Cracov. Ser. Bot. 1986, 28, 107–122. [Google Scholar]
- Ptak, K. Cyto-embryological investigations on the Polish representatives of the genus Crataegus L. II. Embryology of triploid species. Acta Biol. Cracov. Ser. Bot. 1989, 31, 97–112. [Google Scholar]
- Loureiro, J.; Rodriguez, E.; Doležel, J.; Santos, C. Two new nuclear isolation buffers for plant DNA flow cytometry: A test with 37 species. Ann. Bot. 2007, 100, 875–888. [Google Scholar] [CrossRef] [PubMed]
- Doležel, J.; Sgorbati, S.; Lucretti, S. Comparison of three DNA fluorochromes for flow cytometric estimation of nuclear DNA content in plants. Physiol. Plant. 1992, 85, 625–631. [Google Scholar] [CrossRef]
- Galbraith, D.W.; Harkins, K.R.; Maddox, J.M.; Ayres, N.M.; Sharma, D.P.; Firoozabady, E. Rapid flow cytometric analysis of the cell cycle in intact plant tissues. Science 1983, 220, 1049–1051. [Google Scholar] [CrossRef]
- Kron, P.; Husband, B.C. Using flow cytometry to estimate pollen DNA content: Improved methodology and applications. Ann. Bot. 2012, 110, 1067–1078. [Google Scholar] [CrossRef]
- Dickinson, T.A.; Phipps, J.B. Studies in Crataegus (Rosaceae: Maloideae). XIV. The breeding system of Crataegus crus-galli sensu lato in Ontario. Am. J. Bot. 1986, 73, 116–130. [Google Scholar] [CrossRef]
- Dobeš, C.; Lückl, A.; Hülber, K.; Paule, J. Prospects and limits of the flow cytometric seed screen-insights from Potentilla sensu lato (Potentilleae, Rosaceae). New Phytol. 2013, 198, 605–616. [Google Scholar] [CrossRef]
- Matzk, F.; Meister, A.; Schubert, I. An efficient screen for reproductive pathways using mature seeds of monocots and dicots. Plant J. 2000, 21, 97–108. [Google Scholar] [CrossRef]
- Kolarčik, V.; Kocová, V.; Vašková, D. Flow cytometric seed screen data are consistent with models of chromosome inheritance in asymmetrically compensating allopolyploids. Cytometry A 2018, 93, 737–748. [Google Scholar] [CrossRef]
- Temsch, E.M.; Greilhuber, J.; Krisai, R. Genome size in liverworts. Preslia 2010, 82, 63–80. [Google Scholar]
- Krahulcová, A.; Rotreklová, O. Use of flow cytometry in research on apomictic plants. Preslia 2010, 82, 23–39. [Google Scholar]
- Brewbaker, J.L. The distribution and phylogenetic significance of binucleate and trinucleate pollen grains in the Angiosperms. Am. J. Bot. 1967, 54, 1069–1083. [Google Scholar] [CrossRef]
- Bino, R.J.; Van Tuyl, J.M.; De Vries, J.N. Flow cytometric determination of relative nuclear DNA contents in bicellulate and tricellulate pollen. Ann. Bot. 1990, 65, 3–8. [Google Scholar] [CrossRef]
- Lepší, M.; Koutecký, P.; Nosková, J.; Lepší, P.; Urfus, T.; Rich, T.C.G. Versatility of reproductive modes and ploidy level interactions in Sorbus s.l. (Malinae, Rosaceae). Bot. J. Linn. Soc. 2019, 191, 502–522. [Google Scholar] [CrossRef]
- Dickinson, T.A. Crataegu Crus-Galli L. Sensu Lato in Southern Ontario: Phenotypic Variation and Variability in Relation to Reproductive Behaviour. Ph.D. Thesis, University of Western Ontario, London, ON, Canada, 1983. [Google Scholar]
- Celotti, N. The Pollen Tube Pathway and Obturator in Hawthorn Sexual Reproduction. Ph.D. Thesis, Queen’s University, Kingston, ON, Canada, 1995. [Google Scholar]
- Purich, M.A. Characterizing Hybridization between Native and Non-Native Crataegus Species. Master’s Thesis, University of Toronto, Toronto, ON, Canada, 2005. [Google Scholar]
- Luu, D.T.; Qin, X.K.; Morse, D.; Cappadocia, M. S-RNase uptake by compatible pollen tubes in gametophytic self-incompatibility. Nature 2000, 407, 649–651. [Google Scholar] [CrossRef]
- Delgado, L.; Galdeano, F.; Sartor, M.E.; Quarin, C.L.; Espinoza, F.; Ortiz, J.P. Analysis of variation for apomictic reproduction in diploid Paspalum rufum. Ann. Bot. 2014, 113, 1211–1218. [Google Scholar] [CrossRef]
- Böcher, T.W. Cytological and embryological studies in the amphi-apomictic Arabis holboellii complex. Kongel. Danske Vidensk.-Selskab. Biol. Skr. 1951, 6, 1–59. [Google Scholar]
- Böcher, T.W. Experimental taxonomical studies in the Arabis holboellii complex. Sven. Bot. Tidskr. 1954, 48, 31–44. [Google Scholar]
- Sharbel, T.F.; Mitchell-Olds, T. Recurrent polyploid origins and chloroplast phylogeography in the Arabis holboellii complex (Brassicaceae). Heredity 2001, 87, 59–68. [Google Scholar] [CrossRef]
- Burgess, M.B.; Cushman, K.R.; Doucette, E.T.; Talent, N.; Frye, C.T.; Campbell, C.S. Effects of apomixis and polyploidy on diversification and geographic distribution in Amelanchier (Rosaceae). Am. J. Bot. 2014, 101, 1375–1387. [Google Scholar] [CrossRef] [PubMed]
- Sochor, M.; Trávníček, B. Melting pot of diversity: First insights into evolutionary patterns of the Colchic bramble flora (Rubus subgenus Rubus, Rosaceae). Bot. J. Linn. Soc. 2016, 181, 610–620. [Google Scholar] [CrossRef]
- Hajrudinović, A.; Siljak-Yakovlev, S.; Brown, S.C.; Pustahija, F.; Bourge, M.; Ballian, D.; Bogunić, F. When sexual meets apomict: Genome size, ploidy level and reproductive mode variation of Sorbus aria s.l. and S. austriaca (Rosaceae) in Bosnia and Herzegovina. Ann. Bot. 2015, 116, 301–312. [Google Scholar] [CrossRef] [PubMed]
- Ramsey, J.; Schemske, D.W. Pathways, mechanisms, and rates of polyploid formation in flowering plants. Annu. Rev. Ecol. Syst. 1998, 29, 467–501. [Google Scholar] [CrossRef]
- Singh, R. Plant Cytogenetics; CRC Press: Boca Raton, FL, USA, 2003. [Google Scholar]
- Henry, I.M.; Dilkes, B.P.; Young, K.; Watson, B.; Wu, H.; Comai, L. Aneuploidy and genetic variation in the Arabidopsis thaliana triploid response. Genetics 2005, 170, 1979–1988. [Google Scholar] [CrossRef] [PubMed]
- Ramsey, J.; Schemske, D.W. Neopolyploidy in flowering plants. Annu. Rev. Ecol. Syst. 2002, 33, 589–639. [Google Scholar] [CrossRef]
- Henry, I.M.; Dilkes, B.P.; Comai, L. Molecular karyotyping and aneuploidy detection in Arabidopsis thaliana using quantitative fluorescent polymerase chain reaction. Plant J. 2006, 48, 307–319. [Google Scholar] [CrossRef]
- Henry, I.M.; Dilkes, B.P.; Tyagi, A.P.; Lin, H.-Y.; Comai, L. Dosage and parent-of-origin effects shaping aneuploid swarms in A. thaliana. Heredity 2009, 103, 458–468. [Google Scholar] [CrossRef]
- Wang, J.; Huo, B.; Liu, W.; Li, D.; Liao, L. Abnormal meiosis in an intersectional allotriploid of Populus L. and segregation of ploidy levels in 2x × 3x progeny. PLoS ONE 2017, 12. [Google Scholar] [CrossRef]
- De Storme, N.; Mason, A. Plant speciation through chromosome instability and ploidy change: Cellular mechanisms, molecular factors and evolutionary relevance. Curr. Plant Biol. 2014, 1, 10–33. [Google Scholar] [CrossRef]
- Dickinson, T.A. Sex and Rosaceae apomicts. Taxon 2018, 67, 1093–1107. [Google Scholar] [CrossRef]
- Dickinson, T.A.; Belaoussoff, S.; Love, R.M.; Muniyamma, M. North American black-fruited hawthorns: I. Variation in floral construction, breeding system correlates, and their possible evolutionary significance in Crataegus sect. Douglasii Loudon. Folia Geobot. Phytotax. 1996, 31, 355–371. [Google Scholar] [CrossRef]
- Muniyamma, M.; Phipps, J.B. Studies in Crataegus. XI. Further cytological evidence for the occurrence of apomixis in North American hawthorns. Can. J. Bot. 1984, 62, 2316–2324. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).