Multi-Objective Optimized Differential Privacy with Interpretable Machine Learning for Brain Stroke and Heart Disease Diagnosis
Abstract
1. Introduction
- Comprehensive analysis of BS and HD prediction using four ensemble ML models.
- Implementation of DP techniques to safeguard individual data.
- Optimization of the trade-offs between privacy and model performance through PFMOO.
- Utilization of XAI techniques to provide transparency and interpretability of the decision-making process of the best-performing models.
2. Methodology
2.1. Dataset Collection
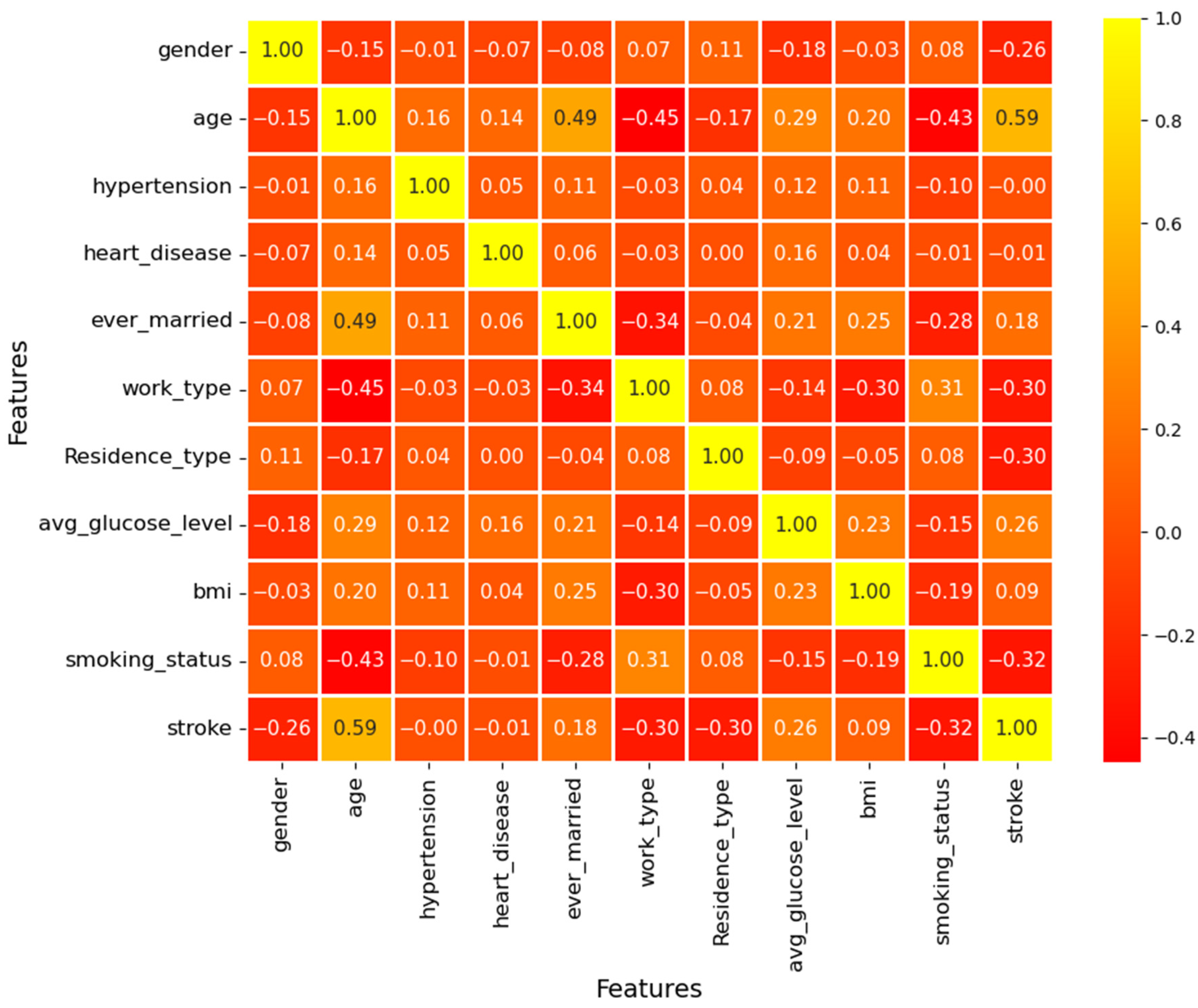
2.2. Data Visualization
2.3. Data Preprocessing
- 10-fold cross-validation with minimal accuracy variance across folds (e.g., ±0.58% for XGB on BSP), confirming consistent performance;
- Ensemble models with built-in regularization (e.g., XGB’s Ω term and RF’s bagging) that control complexity and reduce sensitivity to synthetic samples;
- Comparative evaluation against baseline models trained on imbalanced data to verify that performance improvements were not overfitting artifacts.
2.4. Preprocessed Dataset
2.5. Privacy-Preserving Mechanism
2.6. Non-Secure Server
2.7. ML Model Formation
2.7.1. XGB
2.7.2. RF
2.7.3. LGBM
2.7.4. CAT
| Algorithm 1: Privacy-preserving model training with Gaussian DP | |
| Input: Dataset D, privacy budget ε, probability of failure δ, sensitivity S Output: Trained model M satisfying (ε,δ)-DP | |
| 1: | Split D into training set D_train (80%) and test set D_test (20%) |
| 2: | Normalize D_train to [0, 1] range using min-max scaling |
| 3: | Compute noise scale σ using Equation (4): σ = (S/ε) · 1/√(2·ln(1.25/δ)) |
| 4: | For each feature vector x_i in D_train: Generate Gaussian noise n_i ~ N(0, σ2) Add noise: x_i′ = x_i + n_i |
| 5: | Create noisy training set D_noisy = {(x_i′, y_i)} |
| 6: | Train ensemble model M (XGB, RF, LGBM, or CAT) on D_noisy |
| 7: | Evaluate M on clean test set D_test using metrics from Table 3 |
| 8: | Return M |
2.8. ML Model Evaluation
2.9. Optimal Privacy Parameter Selection
2.10. Best Model Selection
2.11. Outcome Analysis with XAI
3. Results and Discussion
- Original imbalanced data without DP (No SMOTE, No DP);
- SMOTE-balanced data without DP (SMOTE only);
- Original imbalanced data with DP (DP only);
- SMOTE-balanced data with DP (SMOTE + DP, our proposed approach).
4. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| Abbreviation | Full Term |
| ANNAR | Artificial Neural Network Autoregressive |
| AUC | Area Under the Curve |
| BMI | Body Mass Index |
| BS | Brain Stroke |
| BSP | Brain Stroke Prediction |
| CAT | Categorical Boosting |
| CM | Confusion Matrix |
| CV | Cross-Validation |
| DP | Differential Privacy |
| DT | Decision Tree |
| FN | False Negative |
| FP | False Positive |
| HD | Heart Disease |
| HDP | Heart Disease Prediction |
| LGBM | Light Gradient Boosting Machine |
| LIME | Local Interpretable Model-Agnostic Explanations |
| ML | Machine Learning |
| PB | Privacy Budget |
| PC | Privacy Concerns |
| PFMOO | Pareto Frontier Multi-Objective Optimization |
| POF | Probability of Failure |
| RF | Random Forest |
| ROC | Receiver Operating Characteristic |
| SHAP | Shapley Additive Explanations |
| SMOTE | Synthetic Minority Over-sampling Technique |
| SNR | Signal-to-Noise Ratio |
| TN | True Negative |
| TP | True Positive |
| XAI | Explainable Artificial Intelligence |
| XGB | Extreme Gradient Boosting |
References
- Donkor, E.S. Stroke in the 21st century: A snapshot of the burden, epidemiology, and quality of life. Stroke Res. Treat. 2018, 2018, 3238165. [Google Scholar] [CrossRef]
- Moraes, M.D.A.; Jesus, P.A.P.D.; Muniz, L.S.; Costa, G.A.; Pereira, L.V.; Nascimento, L.M.; de Souza Teles, C.A.; Baccin, C.A.; Mussi, F.C. Ischemic stroke mortality and time for hospital arrival: Analysis of the first 90 days. Rev. Esc. Enferm. USP 2023, 57, e20220309. [Google Scholar] [CrossRef] [PubMed]
- Centers for Disease Control and Prevention. Heart Disease Facts and Statistics. Centers for Disease Control and Prevention. 24 October 2024. Available online: https://www.cdc.gov/heart-disease/data-research/facts-stats/index.html (accessed on 5 November 2024).
- Ghannam, A.; Alwidian, J. A Predictive Model of Stroke Diseases using Machine Learning Techniques. Int. J. Recent Technol. Eng. 2022, 11, 53–59. [Google Scholar] [CrossRef]
- Rodríguez, J.A.T. Stroke Prediction Through Data Science and Machine Learning Algorithms. Available online: https://www.researchgate.net/profile/Jose_A_Tavares/publication/352261064_Stroke_prediction_through_Data_Science_and_Machine_Learning_Algorithms/links/60c1071f92851ca6f8d6100b/Stroke-prediction-through-Data-Science-and-Machine-Learning-Algorithms.pdf (accessed on 5 November 2024).
- Ikpea, O.W.; Han, D. Performance of Machine Learning Algorithms for Heart Disease Prediction: Logistic Regressions Regularized by Elastic Net, SVM, Random Forests, and Neural Networks. 2022. Available online: https://www.semanticscholar.org/paper/Performance-of-Machine-Learning-Algorithms-for-by-Dr-Han/65fb6f5c92a7c987ab048e8d4b461cfe7503c9e9 (accessed on 5 November 2024).
- Abdulsalam, G.; Meshoul, S.; Shaiba, H. Explainable heart disease prediction using ensemble-quantum machine learning approach. Intell. Autom. Soft Comput 2023, 36, 761–779. [Google Scholar] [CrossRef]
- Ju, C.; Zhao, R.; Sun, J.; Wei, X.; Zhao, B.; Liu, Y.; Li, H.; Chen, T.; Zhang, X.; Gao, D.; et al. Privacy-preserving technology to help millions of people: Federated prediction model for stroke prevention. arXiv 2020, arXiv:2006.10517. [Google Scholar] [CrossRef]
- Hussain, I.; Qureshi, M.; Ismail, M.; Iftikhar, H.; Zywiołek, J.; López-Gonzales, J.L. Optimal features selection in the high dimensional data based on robust technique: Application to different health database. Heliyon 2024, 10, e37241. [Google Scholar] [CrossRef]
- Qureshi, M.; Ishaq, K.; Daniyal, M.; Iftikhar, H.; Rehman, M.Z.; Salar, S.A. Forecasting cardiovascular disease mortality using artificial neural networks in Sindh, Pakistan. BMC Public Health 2025, 25, 34. [Google Scholar] [CrossRef]
- Soriano, F. Stroke Prediction Dataset [Data Set]. Kaggle. 2019. Available online: https://www.kaggle.com/datasets/fedesoriano/stroke-prediction-dataset (accessed on 31 October 2024).
- Rousseeuw, P.J.; Ruts, I.; Tukey, J.W. The bagplot: A bivariate boxplot. Am. Stat. 1999, 53, 382–387. [Google Scholar] [CrossRef]
- Gu, Z. Complex heatmap visualization. Imeta 2022, 1, e43. [Google Scholar] [CrossRef]
- Zhang, Z. Missing data imputation: Focusing on single imputation. Ann. Transl. Med. 2016, 4, 9. [Google Scholar] [CrossRef]
- Zlatić, L. An alternative for one-hot encoding in neural network models. arXiv 2023, arXiv:2311.05911. [Google Scholar] [CrossRef]
- Blagus, R.; Lusa, L. SMOTE for high-dimensional class-imbalanced data. BMC Bioinform. 2013, 14, 106. [Google Scholar] [CrossRef] [PubMed]
- Honkela, A.; Melkas, L. Gaussian processes with differential privacy. arXiv 2021, arXiv:2106.00474. [Google Scholar] [CrossRef]
- Chen, T.; He, T.; Benesty, M.; Khotilovich, V.; Tang, Y.; Cho, H.; Chen, K.; Mitchell, R.; Cano, I.; Zhou, T. Xgboost: Extreme Gradient Boosting. R Package Version 0.4-2. 2015. Available online: https://cir.nii.ac.jp/crid/1370869856033678496 (accessed on 5 November 2024).
- Rigatti, S.J. Random forest. J. Insur. Med. 2017, 47, 31–39. [Google Scholar] [CrossRef]
- Minastireanu, E.A.; Mesnita, G. Light gbm machine learning algorithm to online click fraud detection. J. Inform. Assur. Cybersecur. 2019, 2019, 263928. [Google Scholar] [CrossRef]
- Hancock, J.T.; Khoshgoftaar, T.M. CatBoost for big data: An interdisciplinary review. J. Big Data 2020, 7, 94. [Google Scholar] [CrossRef]
- Hossain, M.M.; Swarna, R.A.; Mostafiz, R.; Shaha, P.; Pinky, L.Y.; Rahman, M.M.; Rahman, W.; Hossain, M.S.; Hossain, M.E.; Iqbal, M.S. Analysis of the performance of feature optimization techniques for the diagnosis of machine learning-based chronic kidney disease. Mach. Learn. Appl. 2022, 9, 100330. [Google Scholar] [CrossRef]
- Giagkiozis, I.; Fleming, P.J. Methods for multi-objective optimization: An analysis. Inf. Sci. 2015, 293, 338–350. [Google Scholar] [CrossRef]
- Gunning, D.; Aha, D. DARPA’s explainable artificial intelligence (XAI) program. AI Mag. 2019, 40, 44–58. [Google Scholar] [CrossRef]
- Goenka, N.; Tiwari, S. Deep learning for Alzheimer prediction using brain biomarkers. Artif. Intell. Rev. 2021, 54, 4827–4871. [Google Scholar] [CrossRef]
- Kaur, I.; Sachdeva, R. Prediction models for early detection of Alzheimer: Recent trends and future prospects. Arch. Comput. Methods Eng. 2025, 32, 3565–3592. [Google Scholar] [CrossRef]
- Volkmann, H.; Höglinger, G.U.; Grön, G.; Bârlescu, L.A.; Müller, H.P.; Kassubek, J.; DESCRIBE-PSP Study Group. MRI classification of progressive supranuclear palsy, Parkinson disease and controls using deep learning and machine learning algorithms for the identification of regions and tracts of interest as potential biomarkers. Comput. Biol. Med. 2025, 185, 109518. [Google Scholar] [CrossRef]
- Tasnim, S.; Mamun, M.; Chowdhury, S.H.; Hussain, M.I.; Hossain, M.M. Advancing Interpretable AI for Cardiovascular Risk Assessment: A Stacking Regression Approach in Clinical Data from Bangladesh. Medinformatics 2025. [Google Scholar] [CrossRef]
- Hussain, M.I.; Chowdhury, S.H.; Hossain, M.M.; Mamun, M. NeuroBlend-3: Hybrid Deep and Machine Learning Framework with Explainable AI for Multi-class Brain Tumor Detection Using MRI Scans. Medinformatics 2026, 3, 56–66. [Google Scholar] [CrossRef]
- Hussain, M.I.; Chowdhury, S.H.; Shovon, M.; Morzina, M.S.; Hossain, M.M.; Mamun, M. SENet-Augmented Explainable Deep Feature Framework with Machine Learning for Breast Tumor Detection in Ultrasound Imaging. In 2025 International Conference on Quantum Photonics, Artificial Intelligence, and Networking (QPAIN); IEEE: New York, NY, USA, 2025; pp. 1–6. [Google Scholar] [CrossRef]











| Authors | Technique | Affiliate | Limitations |
|---|---|---|---|
| Ghannam and Alwidian [4] | DT | BSP | Class imbalance and lack of CV. |
| Rodríguez [5] | RF | BSP | Fails to handle outliers and lacks CV. |
| Ikpea and Han [6] | RF | HDP | Low recall value. |
| Abdulsalam et al. [7] | BQSVC | HDP | Class imbalance. |
| Ju et al. [8] | CT | BSP | Class imbalance. |
| Hussain et al. [9] | Hybrid Feature Selection (SNR + Mood’s Median Test) | Gene Selection in High-Dim Data | Focused on genomic data, not clinical risk prediction for specific diseases like BS/HD. |
| Qureshi et al. [10] | Time Series (Naïve, SES, Holt, ANNAR) | HD Mortality Forecasting | Small sample size, single-region focus. |
| Features | Details | Data Type | Metric |
|---|---|---|---|
| ID | Patient ID number | Numerical | - |
| Gender | Patient gender | Nominal | - |
| Age | Patient age | Numerical | Years |
| Heart Disease | HD (yes/no) | Nominal | - |
| Hypertension | Hypertension (yes/no) | Nominal | - |
| Ever Married | Marriage status (married/single) | Nominal | - |
| Work Type | Employment type (child, government, unemployed, private, self-employed) | Nominal | - |
| Residence | Location type (rural/urban) | Nominal | - |
| Average Glucose | Blood sugar | Numerical | mg/dL |
| BMI | Patient’s body mass index | Numerical | kg/m2 |
| Smoking Status | Patient smoking background (former, never, current, unknown) | Nominal | - |
| Stroke | Stroke outcome (yes/no) | Nominal | - |
| Metrics | Expression | Explanation |
|---|---|---|
| ACC | The proportion of correct predictions | |
| Precision | Predicted positives that are actually correct | |
| Recall | Actual positives that are correctly identified | |
| Specificity | Actual negatives that are correctly identified | |
| F1 Score | The harmonic means of precision and recall |
| Metric | BS Prediction (XGB, ε = 2, δ = 10−6) | HD Prediction (RF, ε = 0.5, δ = 10−5) |
|---|---|---|
| Accuracy (%) | Mean ± SD: 92.30 ± 0.58 95% CI: [91.72, 92.88] CoV: 0.63% | Mean ± SD: 95.61 ± 0.47 95% CI: [95.13, 96.09] CoV: 0.49% |
| Precision (%) | Mean ± SD: 92.45 ± 0.62 95% CI: [91.83, 93.07] CoV: 0.67% | Mean ± SD: 97.80 ± 0.35 95% CI: [97.45, 98.15] CoV: 0.36% |
| Recall (%) | Mean ± SD: 92.28 ± 0.71 95% CI: [91.57, 92.99] CoV: 0.77% | Mean ± SD: 52.14 ± 1.98 95% CI: [50.16, 54.12] CoV: 3.80% |
| F1 Score (%) | Mean ± SD: 92.29 ± 0.64 95% CI: [91.65, 92.93] CoV: 0.69% | Mean ± SD: 52.95 ± 1.85 95% CI: [51.10, 54.80] CoV: 3.49% |
| Specificity (%) | Mean ± SD: 89.54 ± 0.93 95% CI: [88.61, 90.47] CoV: 1.04% | Mean ± SD: 100.0 ± 0.00 95% CI: [100.0, 100.0] CoV: 0.00% |
| Comparison | Mean Difference | t-Statistic | p-Value (t-Test) | p-Value (Wilcoxon) | Cohen’s d | Effect Size |
|---|---|---|---|---|---|---|
| BS: XGB vs. LGBM | 0.60% | 4.32 | 0.0019 | 0.0021 | 0.89 | Large |
| HD: RF vs. LGBM | 0.41% | 3.87 | 0.0037 | 0.0039 | 0.76 | Large |
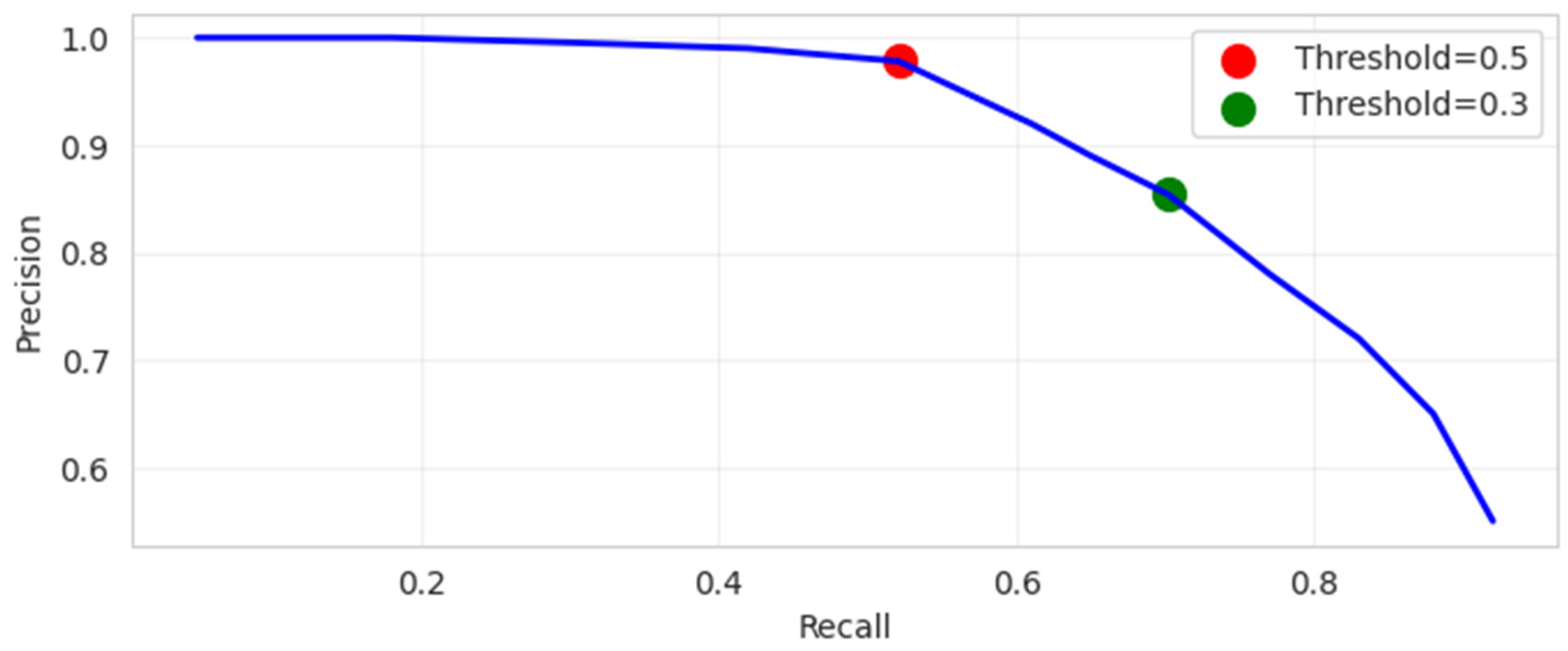
| Threshold | TP | FN | FP | TN | Accuracy | Precision | Recall | Specificity | Balanced Accuracy | MCC | AUC |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 0.5 | 48 | 44 | 1 | 929 | 95.61% | 97.8% | 52.14% | 99.89% | 76.02% | 0.715 | 0.87 |
| 0.38 | 59 | 33 | 5 | 925 | 96.28% | 92.1% | 64.50% | 99.46% | 81.98% | 0.751 | 0.87 |
| 0.3 | 65 | 27 | 11 | 919 | 96.28% | 85.4% | 70.20% | 98.82% | 84.51% | 0.758 | 0.87 |
| Prediction Task | Optimal Model | Rank | Feature | Mean SHAP Value | Mean LIME Contribution | Rank Variance | Clinical Interpretation of Top Features |
|---|---|---|---|---|---|---|---|
| BS | XGB (ε = 2, δ = 10−6) | 1 | Age | 0.324 | +0.215 | Very High (σ2 = 0.11) | Advanced age is the single most critical non-modifiable risk factor, reflecting cumulative vascular damage and increased susceptibility. |
| 2 | Avg. Glucose Level | 0.187 | +0.142 | High (σ2 = 0.23) | Elevated blood sugar, often indicating diabetes or prediabetes, is a major contributor to vascular endothelial dysfunction and atherosclerosis. | ||
| 3 | Hypertension | 0.156 | +0.108 | High (σ2 = 0.19) | Chronic high blood pressure is a primary driver of cerebrovascular damage, including small vessel disease and increased risk of vessel rupture or blockage. | ||
| 4 | BMI | 0.098 | +0.065 | Moderate (σ2 = 0.35) | Higher BMI, associated with obesity, exacerbates hypertension, diabetes, and systemic inflammation, increasing stroke risk. | ||
| 5 | Heart Disease | 0.076 | +0.051 | Moderate (σ2 = 0.41) | A history of heart disease (e.g., atrial fibrillation) is a known embolic source for ischemic strokes, confirming the link between cardiac and cerebrovascular health. | ||
| HD | RF (ε = 0.5, δ = 10−5) | 1 | Age | 0.291 | +0.188 | Very High (σ2 = 0.09) | Age is the bedrock of cardiovascular risk, with arterial stiffness and plaque burden increasing progressively over a lifetime. |
| 2 | Avg. Glucose Level | 0.203 | +0.151 | High (σ2 = 0.21) | Hyperglycemia’s high rank underscores its direct role in promoting coronary artery disease through oxidative stress and glycation of vessel walls. | ||
| 3 | Hypertension | 0.142 | +0.095 | High (σ2 = 0.18) | High blood pressure is a primary mechanism for left ventricular hypertrophy and the progression of coronary atherosclerosis. | ||
| 4 | BMI | 0.089 | +0.058 | Moderate (σ2 = 0.33) | Elevated BMI’s contribution reflects its strong correlation with metabolic syndrome, a cluster of conditions (including hypertension and dyslipidemia) that directly increase heart disease risk. | ||
| 5 | Smoking Status | 0.067 | +0.044 | Moderate (σ2 = 0.44) | Smoking’s presence in the top five confirms its potent role in promoting endothelial injury, inflammation, and thrombus formation, which are central to heart disease pathogenesis. |
| Model | XGB | RF | LGBM | CAT | |
|---|---|---|---|---|---|
| BSP | Accuracy (%) | 94.44% ± 0.51 [94.06, 94.82] | 93.87% ± 0.62 [93.39, 94.35] | 94.19% ± 0.55 [93.76, 94.62] | 93.68% ± 0.68 [93.14, 94.22] |
| Precision (%) | 94.49% ± 0.53 [94.09, 94.89] | 93.92% ± 0.59 [93.46, 94.38] | 94.22% ± 0.57 [93.78, 94.66] | 93.72% ± 0.71 [93.16, 94.28] | |
| Recall (%) | 94.43% ± 0.58 [94.00, 94.86] | 93.86% ± 0.64 [93.36, 94.36] | 94.18% ± 0.61 [93.72, 94.64] | 93.68% ± 0.73 [93.10, 94.26] | |
| Specificity (%) | 92.95% ± 0.79 [92.36, 93.54] | 92.22% ± 0.88 [91.55, 92.89] | 93.15% ± 0.74 [92.59, 93.71] | 92.36% ± 0.92 [91.66, 93.06] | |
| F1 Score (%) | 94.44% ± 0.52 [94.05, 94.83] | 93.86% ± 0.63 [93.37, 94.35] | 94.19% ± 0.56 [93.76, 94.62] | 93.68% ± 0.70 [93.13, 94.23] | |
| HDP | Accuracy (%) | 96.09% ± 0.44 [95.77, 96.41] | 96.47% ± 0.38 [96.19, 96.75] | 96.29% ± 0.41 [95.96, 96.62] | 96.14% ± 0.48 [95.78, 96.50] |
| Precision (%) | 80.87% ± 1.92 [79.43, 82.31] | 90.25% ± 1.45 [89.13, 91.37] | 84.89% ± 1.78 [83.52, 86.26] | 85.24% ± 1.83 [83.86, 86.62] | |
| Recall (%) | 66.05% ± 2.34 [64.29, 67.81] | 64.26% ± 2.18 [62.62, 65.90] | 64.93% ± 2.27 [63.25, 66.61] | 62.24% ± 2.51 [60.35, 64.13] | |
| Specificity (%) | 99.12% ± 0.31 [98.89, 99.35] | 99.73% ± 0.19 [99.59, 99.87] | 99.46% ± 0.26 [99.26, 99.66] | 99.57% ± 0.23 [99.39, 99.75] | |
| F1 Score (%) | 70.77% ± 2.11 [69.17, 72.37] | 70.45% ± 2.05 [68.90, 72.00] | 70.41% ± 2.08 [68.84, 71.98] | 67.52% ± 2.42 [65.68, 69.36] |
| Task | Configuration | Model | Accuracy (%) | Accuracy σ (%) | Recall (%) | Recall σ (%) |
|---|---|---|---|---|---|---|
| BS | No SMOTE, No DP | XGB | 85.3 | 1.21 | 28.7 | 3.45 |
| SMOTE only | XGB | 94.4 | 0.58 | 94.4 | 0.61 | |
| DP only (ε = 2, δ = 10−6) | XGB | 83.1 | 1.43 | 26.3 | 4.02 | |
| SMOTE + DP | XGB | 92.3 | 0.71 | 92.3 | 0.74 | |
| HD | No SMOTE, No DP | RF | 89.7 | 0.95 | 31.2 | 2.88 |
| SMOTE only | RF | 96.5 | 0.49 | 64.3 | 1.55 | |
| DP only (ε = 0.5, δ = 10−5) | RF | 88.4 | 1.12 | 28.9 | 3.21 | |
| SMOTE + DP | RF | 95.6 | 0.63 | 52.1 | 1.98 |
| Authors | Dataset | Technique | Affiliate | Accuracy | PC | XAI | |
|---|---|---|---|---|---|---|---|
| Sample | Features | ||||||
| Ghannam and Alwidian [4] | 5110 | 11 | DT | BSP | 94.2% | NO | NO |
| Rodríguez [5] | 5110 | 11 | RF | BSP | 92.32% | NO | NO |
| Ikpea and Han [6] | 5110 | 11 | RF | HDP | 75% | NO | NO |
| Abdulsalam et al. [7] | 303 | 14 | BQSVC | HDP | 90.16% | NO | YES |
| Ju et al. [8] | - | 119 | CT | BSP | 81.4% | YES | NO |
| Hussain et al. [9] | 10937, 5470, 12534 | 413, 76, 148 | RF | Identifies optimal, significant genes | - | NO | NO |
| Qureshi et al. [10] | 23 (years 1999–2021) | 1 (death cases) + time | ANNAR | HD Mortality | - | NO | NO |
| This Study | 5110 | 11 | XGB with DP | BSP | 92.3% | YES | YES |
| RF with DP | HDP | 95.61% | |||||
| XGB without DP | BSP | 94.44% | NO | NO | |||
| RF without DP | HDP | 96.47% | |||||
| Study | Best Model | Accuracy | Precision | Recall | F1 Score | Specificity | Error Rate | MAPE |
|---|---|---|---|---|---|---|---|---|
| [4] | DT | 94.2% | 83.2% | 86.8% | 84.9% | 95.9% | – | – |
| [5] | RF | 91.89% | – | – | 92.0% | – | – | – |
| [6] | RF | 75% | 12% | 59% | ~0.20 | 76% | – | – |
| [7] | BQSVC | 90.16% | 90% | 90% | 90% | – | – | – |
| [8] | CT | – | – | – | – | – | – | – |
| [9] | RF | – | – | – | – | – | 0.004 | – |
| [10] | ANNAR | – | – | – | – | – | – | 13.08 |
| This Study | RF (HDP, DP) | 95.61% | 97.8% | 70.2% (After Threshold Adjustment) | 52.95% | 100% | – | – |
| XGB (BSP, DP) | 92.3% | 92.45% | 92.28% | 92.29% | 89.54% | – | – |
| Authors | Models Used | Performance Metrics | DP | XAI | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| RF | Neural Networks | XGB | Support Vector Machine | Logistic Regression | Decision Tree | k-Nearest Neighbor | Naïve Bayes | Quantum ML Models | LGBM | CAT | Federated Learning Models | Accuracy | Precision | Recall | F1 Score | Specificity | Error | PB | POF | SHAP | LIME | |
| [4] | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ||||||||||||||
| [5] | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ||||||||||||
| [6] | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | |||||||||||||
| [7] | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ||||||||||||||
| [8] | ✓ | ✓ | ||||||||||||||||||||
| [9] | ✓ | ✓ | ✓ | |||||||||||||||||||
| [10] | ✓ | ✓ | ✓ | |||||||||||||||||||
| This Study | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | |||||||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Hussain, M.I.; Munir, A.; Chowdhury, S.H.; Mamun, M.; Hossain, M.M. Multi-Objective Optimized Differential Privacy with Interpretable Machine Learning for Brain Stroke and Heart Disease Diagnosis. Algorithms 2026, 19, 260. https://doi.org/10.3390/a19040260
Hussain MI, Munir A, Chowdhury SH, Mamun M, Hossain MM. Multi-Objective Optimized Differential Privacy with Interpretable Machine Learning for Brain Stroke and Heart Disease Diagnosis. Algorithms. 2026; 19(4):260. https://doi.org/10.3390/a19040260
Chicago/Turabian StyleHussain, Mohammed Ibrahim, Arslan Munir, Safiul Haque Chowdhury, Mohammad Mamun, and Muhammad Minoar Hossain. 2026. "Multi-Objective Optimized Differential Privacy with Interpretable Machine Learning for Brain Stroke and Heart Disease Diagnosis" Algorithms 19, no. 4: 260. https://doi.org/10.3390/a19040260
APA StyleHussain, M. I., Munir, A., Chowdhury, S. H., Mamun, M., & Hossain, M. M. (2026). Multi-Objective Optimized Differential Privacy with Interpretable Machine Learning for Brain Stroke and Heart Disease Diagnosis. Algorithms, 19(4), 260. https://doi.org/10.3390/a19040260









