Figure 1.
Overall architecture of DCDRNet. The network adopts a parallel dual-stream design that explicitly decouples detail-sensitive and context-aware representations from the input stage. The Detail Branch preserves high-resolution geometric cues throughout encoding at H/2, H/4, and H/8 resolutions. The Context Branch performs progressive abstraction through LSKBlock stages to model long-range semantic dependencies. The structured interaction module fuses detail features (H/8) with context features (H/8 and H/16) through channel alignment, spatial attention, and channel attention mechanisms. The decoder progressively upsamples the fused representation.
Figure 1.
Overall architecture of DCDRNet. The network adopts a parallel dual-stream design that explicitly decouples detail-sensitive and context-aware representations from the input stage. The Detail Branch preserves high-resolution geometric cues throughout encoding at H/2, H/4, and H/8 resolutions. The Context Branch performs progressive abstraction through LSKBlock stages to model long-range semantic dependencies. The structured interaction module fuses detail features (H/8) with context features (H/8 and H/16) through channel alignment, spatial attention, and channel attention mechanisms. The decoder progressively upsamples the fused representation.
Figure 2.
Detail encoder with training–inference re-parameterization. The Detail Encoder is designed to preserve geometric fidelity by maintaining a high-resolution feature stream throughout encoding.
Figure 2.
Detail encoder with training–inference re-parameterization. The Detail Encoder is designed to preserve geometric fidelity by maintaining a high-resolution feature stream throughout encoding.
Figure 3.
Adaptive receptive field modeling in the Context Encoder. The Context Encoder captures global semantic dependencies by adaptively selecting contextual information at different spatial extents. Multiple contextual responses are generated in parallel and dynamically weighted according to the input content, forming an aggregated representation.
Figure 3.
Adaptive receptive field modeling in the Context Encoder. The Context Encoder captures global semantic dependencies by adaptively selecting contextual information at different spatial extents. Multiple contextual responses are generated in parallel and dynamically weighted according to the input content, forming an aggregated representation.
Figure 4.
Training convergence analysis of DCDRNet on the three benchmark datasets over 150 epochs. Each subplot shows training loss, validation loss, Dice score, and IoU. The model exhibits rapid convergence within the first 50 epochs and maintains stable performance thereafter, demonstrating the effectiveness of the proposed parallel dual-stream architecture.
Figure 4.
Training convergence analysis of DCDRNet on the three benchmark datasets over 150 epochs. Each subplot shows training loss, validation loss, Dice score, and IoU. The model exhibits rapid convergence within the first 50 epochs and maintains stable performance thereafter, demonstrating the effectiveness of the proposed parallel dual-stream architecture.
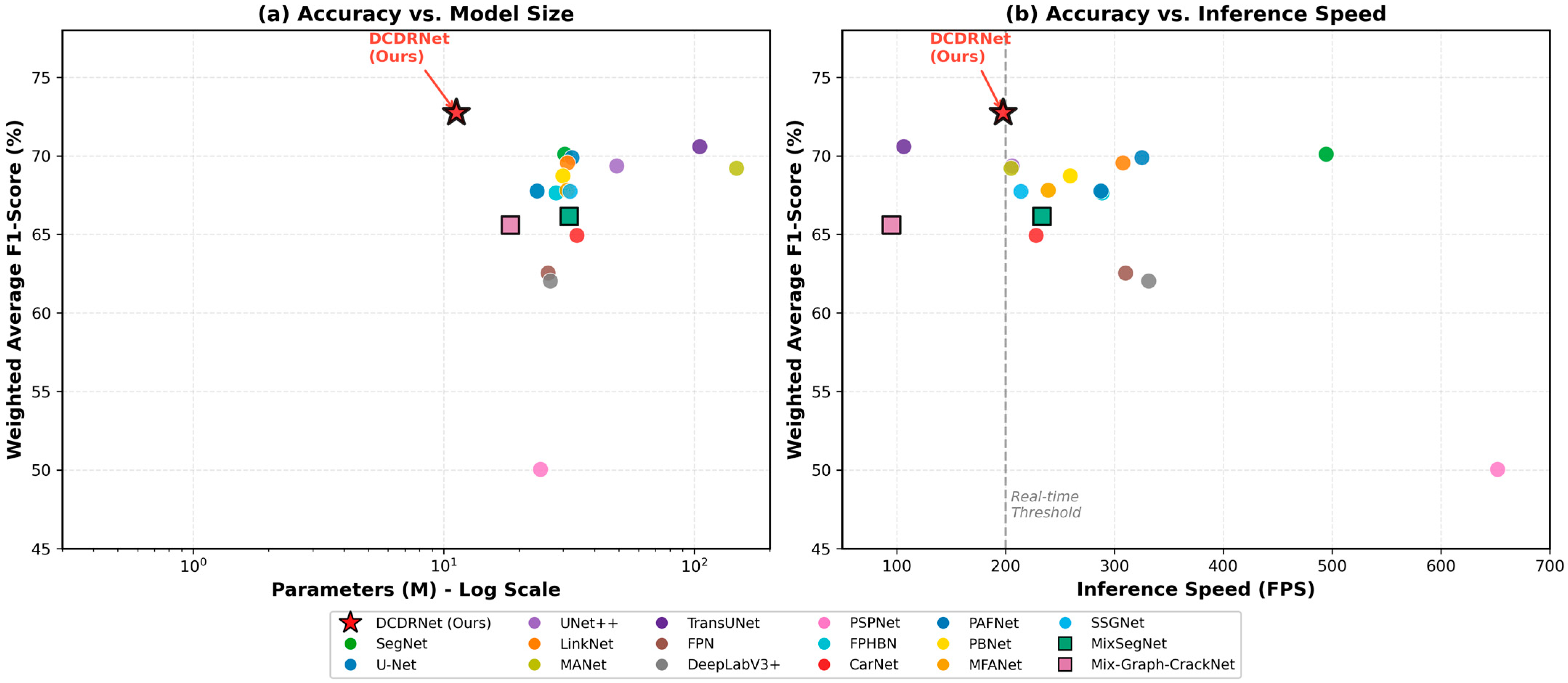
Figure 5.
Efficiency analysis of crack segmentation methods. (a) Average F1-score vs. model parameters (log scale). DCDRNet achieves the highest accuracy among lightweight models. (b) Average F1-score vs. inference speed. The dashed line indicates the real-time threshold (200 FPS). DCDRNet achieves superior accuracy while maintaining real-time performance.
Figure 5.
Efficiency analysis of crack segmentation methods. (a) Average F1-score vs. model parameters (log scale). DCDRNet achieves the highest accuracy among lightweight models. (b) Average F1-score vs. inference speed. The dashed line indicates the real-time threshold (200 FPS). DCDRNet achieves superior accuracy while maintaining real-time performance.
Figure 6.
Multi-dimensional performance comparison using radar chart. Five metrics are evaluated: F1-scores on DeepCrack, CrackForest, and CrackTree260 datasets, model efficiency (1/Parameters), and inference speed (FPS). The area value of each model’s coverage polygon is displayed in parentheses in the legend, enabling quantitative comparison. DCDRNet (Ours) achieves an area of 1.47, demonstrating balanced excellence across all evaluation dimensions, with particularly strong performance on the challenging CrackTree260 benchmark.
Figure 6.
Multi-dimensional performance comparison using radar chart. Five metrics are evaluated: F1-scores on DeepCrack, CrackForest, and CrackTree260 datasets, model efficiency (1/Parameters), and inference speed (FPS). The area value of each model’s coverage polygon is displayed in parentheses in the legend, enabling quantitative comparison. DCDRNet (Ours) achieves an area of 1.47, demonstrating balanced excellence across all evaluation dimensions, with particularly strong performance on the challenging CrackTree260 benchmark.
Figure 7.
Visual comparison of crack segmentation results on DeepCrack test images. The first row shows the original images, the second row shows ground truth labels, and subsequent rows display predictions from different methods. Red boxes highlight regions where baseline methods exhibit notable errors (missed detections, discontinuities, or false positives), while yellow boxes indicate areas where our model (DCDRNet) underperforms. DCDRNet maintains accurate segmentation with better crack continuity and boundary precision in the remaining regions.
Figure 7.
Visual comparison of crack segmentation results on DeepCrack test images. The first row shows the original images, the second row shows ground truth labels, and subsequent rows display predictions from different methods. Red boxes highlight regions where baseline methods exhibit notable errors (missed detections, discontinuities, or false positives), while yellow boxes indicate areas where our model (DCDRNet) underperforms. DCDRNet maintains accurate segmentation with better crack continuity and boundary precision in the remaining regions.
Figure 8.
Precision–Recall curves for crack segmentation methods on the DeepCrack, CrackForest, CrackTree260 dataset. Each curve represents the precision–recall trade-off at different classification thresholds. The Area Under the Curve (AUC) values are shown in the legend. DCDRNet (Ours) achieves the highest AUC of 0.881, 0.867, 0.844, demonstrating superior overall performance across all threshold settings. The red star represents our model.
Figure 8.
Precision–Recall curves for crack segmentation methods on the DeepCrack, CrackForest, CrackTree260 dataset. Each curve represents the precision–recall trade-off at different classification thresholds. The Area Under the Curve (AUC) values are shown in the legend. DCDRNet (Ours) achieves the highest AUC of 0.881, 0.867, 0.844, demonstrating superior overall performance across all threshold settings. The red star represents our model.
Table 1.
Performance comparison on DeepCrack dataset.
Table 1.
Performance comparison on DeepCrack dataset.
| Method | Prec (%) | Recall (%) | F1 (%) | IoU (%) |
|---|
| DCDRNet (Ours) | 88.13 | 78.47 | 81.12 | 70.10 |
| TransUNet [33] | 86.45 | 78.42 | 80.82 | 69.01 |
| LinkNet [34] | 86.82 | 77.66 | 80.38 | 68.79 |
| FPHBN [35] | 85.27 | 77.58 | 80.04 | 67.67 |
| U-Net [36] | 86.15 | 77.89 | 80.00 | 68.41 |
| SegNet [37] | 86.74 | 76.55 | 79.60 | 67.60 |
| UNet++ [38] | 86.84 | 76.27 | 79.58 | 67.60 |
| MANet [39] | 86.34 | 76.85 | 79.41 | 67.60 |
| SSGNet [40] | 84.38 | 75.33 | 78.65 | 65.05 |
| PBNet [41] | 83.94 | 76.85 | 78.78 | 65.96 |
| MFANet [42] | 84.94 | 74.48 | 78.07 | 64.74 |
| PAFNet [43] | 85.16 | 74.17 | 77.75 | 64.53 |
| CarNet [44] | 83.81 | 74.98 | 77.71 | 64.72 |
| FPN [45] | 79.44 | 74.59 | 75.22 | 62.17 |
| DeepLabV3+ [46] | 78.53 | 74.89 | 74.37 | 61.46 |
| PSPNet [47] | 71.28 | 59.31 | 61.81 | 47.36 |
| MixSegNet [15] | 79.90 | 78.07 | 77.23 | 64.44 |
| Mix-Graph-CrackNet [48] | 75.23 | 76.89 | 76.05 | 61.35 |
Table 2.
Performance comparison on CrackForest dataset.
Table 2.
Performance comparison on CrackForest dataset.
| Method | Prec (%) | Recall (%) | F1 (%) | IoU (%) |
|---|
| DCDRNet (Ours) | 66.15 | 65.74 | 63.11 | 47.98 |
| TransUNet | 63.02 | 66.00 | 63.12 | 46.86 |
| LinkNet | 63.49 | 61.79 | 62.01 | 46.10 |
| FPHBN | 55.99 | 71.89 | 62.02 | 45.74 |
| U-Net | 63.18 | 67.47 | 64.38 | 48.50 |
| SegNet | 67.21 | 69.45 | 67.56 | 51.82 |
| UNet++ | 65.20 | 66.65 | 65.01 | 49.37 |
| MANet | 67.07 | 61.74 | 62.23 | 46.92 |
| SSGNet | 59.93 | 64.03 | 60.27 | 44.01 |
| PBNet | 60.56 | 65.90 | 61.00 | 45.22 |
| MFANet | 62.46 | 64.55 | 61.74 | 45.41 |
| PAFNet | 62.11 | 63.78 | 61.60 | 45.22 |
| CarNet | 53.79 | 63.75 | 55.82 | 38.57 |
| FPN | 54.81 | 62.52 | 56.99 | 40.67 |
| DeepLabV3+ | 56.96 | 63.84 | 58.53 | 42.43 |
| PSPNet | 41.66 | 38.05 | 36.62 | 23.57 |
| MixSegNet | 56.12 | 71.32 | 61.66 | 45.28 |
| Mix-Graph-CrackNet | 56.12 | 60.45 | 58.27 | 41.12 |
Table 3.
Performance comparison on CrackTree260 dataset.
Table 3.
Performance comparison on CrackTree260 dataset.
| Method | Prec (%) | Recall (%) | F1 (%) | IoU (%) |
|---|
| DCDRNet (Ours) | 31.05 | 54.40 | 39.02 | 24.52 |
| TransUNet | 23.69 | 33.50 | 27.48 | 15.99 |
| LinkNet | 21.42 | 28.33 | 23.69 | 13.52 |
| FPHBN | 10.04 | 25.33 | 13.73 | 7.43 |
| U-Net | 22.54 | 33.18 | 26.34 | 15.23 |
| SegNet | 23.95 | 34.64 | 28.06 | 16.39 |
| UNet++ | 21.70 | 30.64 | 24.79 | 14.22 |
| MANet | 22.49 | 32.53 | 25.96 | 14.99 |
| SSGNet | 16.23 | 33.16 | 21.48 | 11.93 |
| PBNet | 20.22 | 40.58 | 26.51 | 15.36 |
| MFANet | 18.19 | 34.59 | 23.85 | 13.36 |
| PAFNet | 19.40 | 36.18 | 25.14 | 14.36 |
| CarNet | 8.56 | 17.87 | 10.92 | 5.80 |
| FPN | 10.72 | 6.65 | 7.38 | 3.88 |
| DeepLabV3+ | 10.53 | 6.80 | 7.46 | 3.91 |
| PSPNet | 4.98 | 2.07 | 2.58 | 1.32 |
| MixSegNet | 13.56 | 27.26 | 17.72 | 9.77 |
| Mix-Graph-CrackNet | 24.34 | 26.56 | 25.43 | 14.57 |
Table 4.
Summary of all methods sorted by weighted average F1-score. The weighted average is calculated as: , where Ni denotes the number of test samples for each dataset (DeepCrack: 237, CrackForest: 24, CrackTree260: 52).
Table 4.
Summary of all methods sorted by weighted average F1-score. The weighted average is calculated as: , where Ni denotes the number of test samples for each dataset (DeepCrack: 237, CrackForest: 24, CrackTree260: 52).
| Method | DeepCrack F1 (%) | CrackForest F1 (%) | CrackTree260 F1 (%) | Weighted Avg F1 (%) |
|---|
| DCDRNet (Ours) | 81.12 | 63.11 | 39.02 | 72.74 |
| TransUNet | 80.82 | 63.12 | 27.48 | 70.60 |
| LinkNet | 80.38 | 62.01 | 23.69 | 69.55 |
| FPHBN | 80.04 | 62.02 | 13.73 | 67.64 |
| U-Net | 80.00 | 64.38 | 26.34 | 69.89 |
| SegNet | 79.60 | 67.56 | 28.06 | 70.11 |
| UNet++ | 79.58 | 65.01 | 24.79 | 69.36 |
| MANet | 79.41 | 62.23 | 25.96 | 69.21 |
| SSGNet | 78.65 | 60.27 | 21.48 | 67.74 |
| PBNet | 78.78 | 61.00 | 26.51 | 68.73 |
| MFANet | 78.07 | 61.74 | 23.85 | 67.81 |
| PAFNet | 77.75 | 61.60 | 25.14 | 67.77 |
| CarNet | 77.71 | 55.82 | 10.92 | 64.94 |
| FPN | 75.22 | 56.99 | 7.38 | 62.55 |
| DeepLabV3+ | 74.37 | 58.53 | 7.46 | 62.04 |
| PSPNet | 61.81 | 36.62 | 2.58 | 50.04 |
| MixSegNet | 77.23 | 61.66 | 17.72 | 66.15 |
| Mix-Graph-CrackNet | 76.05 | 58.27 | 25.43 | 65.58 |
Table 5.
Ablation study on DeepCrack dataset.
Table 5.
Ablation study on DeepCrack dataset.
| Configuration | P | C | D | Prec (%) | Recall (%) | F1 (%) | Params (M) |
|---|
| Base + P | ✓ | | | 68.42 | 65.18 | 66.76 | 21.81 |
| Base + C | | ✓ | | 72.35 | 68.92 | 70.59 | 4.20 |
| Base + D | | | ✓ | 70.18 | 67.44 | 68.78 | 3.87 |
| Base + P + C | ✓ | ✓ | | 75.84 | 71.26 | 73.48 | 10.00 |
| Base + P + D | ✓ | | ✓ | 74.29 | 70.85 | 72.53 | 21.21 |
| Base + C + D | | ✓ | ✓ | 78.92 | 74.38 | 76.58 | 4.56 |
| Full (P + C + D) | ✓ | ✓ | ✓ | 88.13 | 78.47 | 81.12 | 11.23 |
Table 6.
Ablation study on CrackForest dataset.
Table 6.
Ablation study on CrackForest dataset.
| Configuration | P | C | D | Prec (%) | Recall (%) | F1 (%) | Params (M) |
|---|
| Base + P | ✓ | | | 52.48 | 58.21 | 55.20 | 21.81 |
| Base + C | | ✓ | | 55.62 | 60.45 | 57.93 | 4.20 |
| Base + D | | | ✓ | 53.19 | 59.02 | 55.95 | 3.87 |
| Base + P + C | ✓ | ✓ | | 58.74 | 62.38 | 60.50 | 10.00 |
| Base + P + D | ✓ | | ✓ | 57.28 | 61.55 | 59.34 | 21.21 |
| Base + C + D | | ✓ | ✓ | 61.45 | 63.82 | 62.61 | 4.56 |
| Full (P + C + D) | ✓ | ✓ | ✓ | 66.15 | 65.74 | 63.11 | 11.23 |
Table 7.
Ablation study on CrackTree260 dataset.
Table 7.
Ablation study on CrackTree260 dataset.
| Configuration | P | C | D | Prec (%) | Recall (%) | F1 (%) | Params (M) |
|---|
| Base + P | ✓ | | | 18.52 | 24.66 | 21.15 | 21.81 |
| Base + C | | ✓ | | 22.14 | 28.92 | 25.08 | 4.2 |
| Base + D | | | ✓ | 19.88 | 26.54 | 22.73 | 3.87 |
| Base + P + C | ✓ | ✓ | | 25.63 | 32.18 | 28.53 | 10.0 |
| Base + P + D | ✓ | | ✓ | 24.15 | 30.76 | 27.06 | 21.21 |
| Base + C + D | | ✓ | ✓ | 28.42 | 45.21 | 34.9 | 4.56 |
| Full (P + C + D) | ✓ | ✓ | ✓ | 31.05 | 54.40 | 39.02 | 11.23 |
Table 8.
Model complexity and efficiency comparison.
Table 8.
Model complexity and efficiency comparison.
| Method | Params (M) | Size (MB) | FPS | Weighted Avg F1 (%) | F1/Params |
|---|
| DCDRNet (Ours) | 11.23 | 42.84 | 197.63 | 72.74 | 6.48 |
| SegNet | 30.41 | 116.02 | 494.54 | 70.11 | 2.31 |
| TransUNet | 105.28 | 401.60 | 106.13 | 70.60 | 0.67 |
| U-Net | 32.51 | 124.03 | 325.03 | 69.89 | 2.15 |
| LinkNet | 31.17 | 118.91 | 307.69 | 69.55 | 2.23 |
| UNet++ | 48.98 | 186.84 | 205.93 | 69.36 | 1.42 |
| MANet | 147.43 | 562.42 | 204.85 | 69.21 | 0.47 |
| PBNet | 29.86 | 113.92 | 259.33 | 68.73 | 2.30 |
| MFANet | 31.08 | 118.58 | 239.14 | 67.81 | 2.18 |
| PAFNet | 23.60 | 90.04 | 287.32 | 67.77 | 2.87 |
| SSGNet | 31.87 | 121.58 | 213.89 | 67.74 | 2.13 |
| FPHBN | 28.03 | 106.94 | 288.64 | 67.64 | 2.41 |
| CarNet | 33.99 | 129.66 | 228.01 | 64.94 | 1.91 |
| FPN | 26.11 | 99.60 | 310.06 | 62.55 | 2.40 |
| DeepLabV3+ | 26.67 | 101.74 | 331.5 | 62.04 | 2.33 |
| PSPNet | 24.30 | 92.69 | 651.69 | 50.04 | 2.06 |
| MixSegNet | 31.65 | 120.76 | 233.30 | 66.15 | 2.09 |
| Mix-Graph-CrackNet | 18.44 | 70.35 | 95.00 | 65.58 | 3.56 |