1. Introduction
Agriculture plays a vital role in sustainable human livelihoods, rural development, and global food security, particularly in regions exposed to increasing climatic variability and limited natural resource constraints. According to the Food and Agriculture Organization of the United Nations (FAO), global food demand is expected to increase by nearly 60% by 2050 to meet the needs of an estimated population of 9.3 billion [
1]. In this context, improving the reliability, accuracy, and scalability of predictive models has become a critical requirement for decision-making in precision agriculture. Recent advances in machine learning (ML) and deep learning (DL) have significantly transformed agricultural analytics by enabling data-driven modeling of complex, nonlinear interactions between climate conditions, soil characteristics, and crop physiology [
2,
3,
4]. Consequently, there is a growing demand for robust classification models capable of handling heterogeneous, high-dimensional, and noisy data while maintaining interpretability and computational efficiency.
1.1. Motivation and Incitement
Modern agricultural systems increasingly rely on precise and automated decision-support tools for tasks such as crop yield prediction, disease detection, water stress monitoring, and species identification [
5,
6]. Traditional yield forecasting approaches, which are largely based on expert knowledge and limited environmental indicators, fail to capture the complex and nonlinear dependencies inherent in agricultural ecosystems [
4]. While deep learning models offer powerful representational capabilities, their deployment is often constrained by data availability, computational cost, and limited interpretability, particularly in resource-constrained agricultural settings. These limitations motivate the continued investigation of robust, theoretically grounded alternatives such as Support Vector Machines (SVMs).
1.2. The Relevant Literature
Among supervised learning methods, SVMs and random forests (RFs) remain widely adopted due to their strong generalization ability and robustness to overfitting, whereas deep learning architectures such as Convolutional Neural Networks (CNNs) and Long Short-Term Memory (LSTM) networks have demonstrated superior performance in modeling spatial and temporal agricultural data [
2,
3]. Shawon et al. [
4] reported that linear regression, RF, and gradient boosting trees are among the most frequently used techniques for crop yield forecasting. In contrast, Meghraoui et al. [
2] highlighted the effectiveness of CNNs and LSTMs in exploiting spatiotemporal information.
SVMs are particularly attractive due to their solid theoretical foundations and effectiveness in high-dimensional and nonlinear classification tasks [
7]. However, their performance is strongly influenced by the optimization strategy used to solve the underlying convex problem [
8]. Classical solvers such as Sequential Minimal Optimization (SMO) and Stochastic Gradient Descent (SGD) [
9,
10] suffer from scalability and stability issues when applied to complex, noisy, or large-scale datasets [
11]. To address these challenges, metaheuristic-based approaches have been investigated. Early work by Paquet and Engelbrecht [
12] demonstrated the feasibility of Particle Swarm Optimization (PSO) for SVM training, while Dias and Rocha Neto [
13] showed that simulated annealing in the dual space can produce near-optimal solutions with limited Karush–Kuhn–Tucker (KKT) violations. Nevertheless, most existing studies focus primarily on hyperparameter optimization rather than on directly addressing the constrained dual SVM problem [
14].
Particle Swarm Optimization (PSO) has been widely investigated as an efficient metaheuristic for solving complex and nonconvex optimization problems. Since its original formulation, numerous PSO variants have been proposed to enhance convergence speed, maintain population diversity, and avoid premature stagnation. Among these developments, hybrid and multi-swarm PSO algorithms have attracted particular attention. For example, Hybrid Multi-Swarm PSO (HMSPSO) coordinates multiple interacting swarms to improve global exploration and robustness in complex search spaces [
15]. Other approaches incorporate evolutionary operators inspired by genetic algorithms, such as crossover and mutation, to enhance diversity and balance exploration and exploitation. A comprehensive review by Engelbrecht [
16] highlights the effectiveness of PSO variants with crossover mechanisms and analyzes their empirical convergence behavior. Related hybrid frameworks, including RCGA-PSO, CBHPSO, PSO with mutation, and cooperative swarm strategies, further demonstrate the flexibility of PSO in addressing constrained and high-dimensional optimization problems.
Beyond PSO-based methods, other evolutionary algorithms have been successfully combined with Support Vector Machines (SVMs) to improve classification performance through enhanced parameter optimization. Genetic Algorithms (GA) have been extensively used to optimize SVM hyperparameters and feature subsets due to their strong global search capability and robustness to local optima. Similarly, Differential Evolution (DE) has been applied to SVM optimization, showing competitive performance thanks to its simple structure, fast convergence, and effectiveness in continuous parameter spaces. These evolutionary approaches highlight the importance of advanced optimization strategies in improving SVM training, particularly when solving the dual optimization problem under constraints.
Motivated by these advances, the present work proposes a novel hybrid framework that combines Particle Swarm Optimization with Open Competency Optimization (OCO). Unlike existing hybrid PSO approaches, the proposed OCO–PSO framework introduces a competency-driven learning mechanism that adaptively guides the swarm while explicitly enforcing the dual constraints of the SVM formulation. This design aims to enhance convergence stability, avoid local minima, and improve solution quality, thereby positioning the proposed method within the broader family of evolutionary and swarm-based optimization techniques for SVM training.
1.3. Major Research Gaps
Despite these advances, several important limitations remain unresolved. First, existing SVM solvers struggle to jointly ensure scalability, numerical stability, and strict satisfaction of dual constraints. Second, metaheuristic approaches are often employed as black-box optimizers, without explicitly exploiting the structure of the SVM dual formulation. Third, limited attention has been devoted to achieving sparse and well-distributed dual solutions that improve interpretability and robustness, particularly for imbalanced and heterogeneous agricultural datasets.
1.4. Contributions and Organization of the Paper
In this work, we propose the OCO-PSO hybrid optimization framework to enhance SVM training. Compared to standard SVM solvers such as SMO, SGD, decision trees, or random forests, OCO-PSO offers several advantages: it efficiently explores the solution space, avoids local minima, and strictly enforces dual constraints on Lagrange multipliers. These features result in improved predictive performance, sparser models, and better interpretability. The main limitation is the higher computational cost during training due to constrained global optimization, although prediction remains highly efficient. By explicitly addressing these strengths and trade-offs, our approach provides a robust alternative for classification tasks even with small- to medium-sized datasets.
To address the aforementioned challenges, this paper proposes a novel hybrid optimization framework, termed OCO-PSO, which combines Particle Swarm Optimization (PSO) with Open Competence Optimization (OCO) for solving the dual SVM problem. In this framework, OCO is reinterpreted as a diversification operator that introduces controlled perturbations to the Lagrange multiplier vector, thereby enhancing exploration while preserving feasibility. The proposed method enforces the equality constraint at each iteration and corrects numerical drift through orthogonal projection, resulting in reduced KKT violations. In addition, the SVM hyperparameters (C and ) are jointly optimized using Bayesian optimization.
Extensive experiments conducted on five heterogeneous datasets from medical, signal processing, agricultural, and imbalanced synthetic domains demonstrate that OCO-PSO consistently outperforms classical SVM solvers and widely used classifiers in terms of accuracy, robustness, and model parsimony. In particular, for crop yield prediction, the proposed approach achieves an accuracy of 89.17% and an MCC of 0.786, while producing significantly sparser models and maintaining near-zero constraint violations. These improvements are statistically validated, confirming the reliability and practical relevance of the proposed framework.
The remainder of this paper is organized as follows.
Section 2 reviews related work on SVM optimization.
Section 3 presents the proposed OCO-PSO algorithm.
Section 4 and
Section 5 describe the experimental setup and report the results.
Section 6 discusses the findings, and
Section 7 concludes the paper and outlines future research directions.
2. Support Vector Machines
Before presenting the formulation of Support Vector Machines (SVMs), it is now appropriate to provide an overview of the proposed methodological framework. The objective of this study is to enhance SVM learning by addressing the limitations of classical optimization solvers commonly used to solve their dual formulation. Indeed, the choice of SVM as the cornerstone of our framework is motivated by several fundamental principles:
SVMs are grounded in solid mathematical foundations derived from statistical learning theory. Their ability to determine the optimal separating hyperplane by maximizing the margin between classes constitutes an elegant approach that promotes strong generalization capabilities, even when training data are limited.
Their robustness to overfitting is particularly noteworthy, especially in high-dimensional spaces where the number of features may exceed the number of samples. Moreover, SVMs can efficiently handle nonlinear problems through the use of kernel functions, which implicitly project data into higher-dimensional spaces without incurring expensive explicit computations. This property is especially valuable for processing heterogeneous data typically encountered in agricultural applications.
In agricultural contexts, historical yield data are often limited in size but rich in explanatory variables. SVMs maintain strong predictive performance even with small training samples, in contrast to methods that require large amounts of data, such as deep neural networks.
Agricultural data are frequently affected by noise (e.g., sensor errors or sporadic extreme weather conditions). The SVM regularization parameter C provides a principled mechanism for controlling sensitivity to outliers, which is essential for producing reliable predictions.
However, the effectiveness of SVMs strongly depends on the stability and constraint-handling capability of the underlying optimization process. In this context, we propose a hybrid optimization framework based on Open Competency Optimization (OCO) and Particle Swarm Optimization (PSO), designed to directly solve the SVM dual problem while strictly enforcing both box and equality constraints on the Lagrange multipliers. The global exploration capability of PSO is combined with the adaptive learning mechanisms of OCO to enhance convergence stability, mitigate premature convergence to local minima, and efficiently explore the hypothesis space. Within this framework, the SVM serves as the central decision model, whose performance is enhanced through the proposed OCO-PSO optimization strategy, rather than acting as a standalone generic classifier.
Support Vector Machines (SVMs) are a robust family of established supervised learning algorithms, predominantly employed for classification and regression. Introduced by Vapnik and Cortes [
17], the main objective of an SVM is to find the optimal separation hyperplane that maximizes the margin between the different data classes. The underlying theory of SVMs naturally leads to a quadratic optimization problem, which can be formulated in two equivalent ways: the primal (direct) form and the dual form obtained by Lagrangian duality.
2.1. Primal Formulation of SVM
Consider a training dataset labeled
, where
and
. The main formulation aims to determine the parameters
of a decision boundary
. The objective is to maximize the margin while accounting for classification errors using slack variables. This objective is formalized as the following soft margin constraint optimization problem:
where:
: The decision hyperplane normal vector is inversely proportional to the margin width; minimizing this term maximizes the margin.
: Slack variables added to allow misclassification of difficult or noisy data points (soft-margin approach).
: The regularization parameter that balances the trade-off between margin maximization and misclassification penalties.
2.2. Dual Formulation and Kernel Trick
To avoid the computational complexities of the primal problem, particularly in high-dimensional spaces, it is generally transformed using Lagrange multipliers
. Assuming strong duality and satisfaction of the Karush–Kuhn–Tucker (KKT) conditions, this leads to the dual formulation:
This dual formulation offers two major advantages:
Parsimony and Computational Efficiency: Only data points for which —termed support vectors—contribute to defining the decision boundary. This characteristic naturally introduces the parsimony of the model and considerably reduces the dependence on the size of the set of complete data.
Flexibility via the Kernel Trick: The implicit linear dot product can be replaced by a kernel function , such as the radial basis (RBF) or polynomial kernels. This kernel trick allows SVMs to efficiently learn complex and nonlinear decision boundaries by projecting the data into a high-dimensional feature space.
2.3. Critical Review of Dominant Training Methods for SVMs
Although SMO and SGD represent the dominant paradigms for SVM training, neither method provides a fully satisfactory solution when confronted with large-scale, nonlinear, or heterogeneous datasets. Their respective strengths are offset by fundamental limitations that restrict their applicability in complex real-world scenarios.
2.3.1. Sequential Minimal Optimization (SMO)
SMO, which was first detailed in the work of Platt [
9], greatly improved SVM training by solving the dual optimization problem through a series of sub-problems that could be solved analytically and that only involved two Lagrange multipliers. With this fundamental development, the need for large, external quadratic programming (QP) solvers has been completely removed, which is the main reason for
LIBSVM-type implementations.
The efficiency of SMO relies heavily on heuristics for selecting the multiplier pair and . This choice is mainly directed by two criteria:
Violation of KKT Conditions: Multipliers associated with examples that are misclassified or are too close to the decision boundary should be given the highest priority as these are the instances which probably require margin refinement the most.
Maximizing the Step Size: The second multiplier is chosen such that it maximizes the move towards the optimum; very often, this is performed by picking the instance of the class opposite to that of the first () for the second.
While SMO guarantees convergence and numerical stability, its dependence on heuristics makes it sensitive to dataset characteristics.
2.3.2. Stochastic Gradient Descent (SGD)
In contrast to the dual SMO approaches and decomposition methods, SGD directly addresses the primal formulation of the SVM optimization problem (Equation (
1)). It minimizes the objective function through iterative updates based on randomly sampled instances (or mini-batches) from the dataset.
The Pegasos (Primal Estimated sub-GrAdient SOLver) algorithm [
18] has proven particularly effective for large-scale SVM training because its computational complexity depends primarily on the data dimension
D rather than the dataset size
N. The algorithm updates the weight vector
iteratively:
where
is a learning rate determined by a predefined schedule. SGD is highly effective for large-scale linear classification but faces significant challenges when applied to nonlinear kernels, as the weight vector
cannot be explicitly represented in infinite-dimensional feature spaces.
2.3.3. Analysis and Critical Discussion
Whilst Support Vector Machines are theoretically well-founded and widely used, their practical performance is strongly influenced by the efficiency and robustness of the underlying optimization algorithms. Among existing approaches, Sequential Minimal Optimization (SMO) and Stochastic Gradient Descent (SGD) remain the most commonly adopted paradigms. However, a closer examination reveals that neither method provides a fully satisfactory solution when confronted with large-scale, nonlinear, or heterogeneous datasets. Their respective strengths are offset by inherent limitations that restrict their applicability in complex real-world scenarios:
Sequential Minimal Optimization (SMO), significantly improved SVM training by decomposing the dual optimization problem into a sequence of analytically solvable sub-problems involving only two Lagrange multipliers. This strategy eliminates the need for large-scale quadratic programming solvers and has enabled efficient implementations, such as LIBSVM.
Despite its numerical stability and guaranteed convergence, SMO suffers from several well-documented limitations. First, its scalability is severely constrained for large datasets, as the number of iterations can grow super-linearly with the training size. Second, the convergence speed is highly sensitive to the working set selection heuristics, which are problem-dependent and may lead to suboptimal convergence behavior [
19]. Third, SMO performance degrades significantly in the presence of dense kernel matrices or low-sparsity solutions, where a large number of support vectors must be maintained. As a result, while SMO is effective for small- to medium-sized problems, its efficiency and robustness diminish in high-dimensional or large-scale settings.
Stochastic Gradient Descent (SGD) operates directly on the primal SVM formulation (Equation (
1)) by performing iterative updates based on randomly sampled data points or mini-batches. Algorithms such as Pegasos have demonstrated excellent scalability, as their computational complexity depends primarily on the data dimensionality rather than the dataset size.
However, SGD exhibits critical limitations in the context of kernel-based SVMs. Most notably, it suffers from the so-called “curse of kernelization” [
20]: when nonlinear kernels are employed, all support vectors must be updated at each iteration, effectively nullifying the computational advantages of stochastic optimization. Furthermore, SGD is prone to oscillatory convergence behavior near the optimum due to the inherent variance in stochastic gradients [
21], requiring careful tuning of learning rate schedules and stopping criteria. These issues limit the reliability and robustness of SGD for nonlinear, high-precision SVM training.
2.3.4. Critical Analysis and Research Gap
The above analysis highlights a fundamental trade-off in existing SVM training methods between numerical stability, scalability, and flexibility. While SMO prioritizes stability and exact constraint satisfaction, it struggles with scalability and heuristic sensitivity. Conversely, SGD offers scalability but sacrifices robustness and kernel compatibility.
More importantly, most existing approaches focus primarily on convergence speed, often overlooking other critical aspects of model quality such as sparsity, constraint satisfaction, and interpretability—properties that are particularly important in regulated domains such as agriculture and healthcare. Although metaheuristic optimization methods have been widely explored for hyperparameter tuning and feature selection, their use as direct solvers of the dual SVM optimization problem remains largely underexplored. This is mainly due to the difficulty of enforcing strict equality constraints, such as , within population-based optimization frameworks.
This gap motivates the exploration of specialized hybrid metaheuristic solvers capable of navigating the high-dimensional space of Lagrange multipliers while explicitly enforcing SVM dual constraints. By integrating global search capabilities with adaptive learning mechanisms, such approaches have the potential to overcome the limitations of classical solvers and provide more robust, interpretable, and flexible SVM training strategies [
22].
5. Results
We evaluate OCO-PSO with four basic methods—SVC, SGDClassifier, decision tree, and random forest—on datasets of different performance, including diabetes (medical data), sonar, and ionosphere for signal processing and agricultural yield, as well as imbalanced synthetic data (imbalanced_data). The performance is evaluated in terms of accuracy, macro-F1-score, MCC, and ROC-AUC, with a focus on class balance and fairness. Computational efficiency (training and inference time), model sparsity, and the stability of the optimization solution (violation of the KKT condition) are also analyzed.
5.1. Performance Metrics
The classification performance is evaluated using the following metrics:
Accuracy (%): the proportion of correctly classified samples over the total number of samples:
where
,
,
, and
denote true positives, true negatives, false positives, and false negatives, respectively.
Precision: the fraction of correctly predicted positive samples among all predicted positives:
Recall (Sensitivity): the fraction of correctly predicted positive samples among all actual positives:
F1-Macro: the unweighted mean of F1-scores computed for each class independently:
where
C is the number of classes.
Matthews Correlation Coefficient (MCC): a balanced measure of classification quality that accounts for all four confusion matrix categories:
ROC-AUC (Receiver Operating Characteristic–Area Under Curve): measures the classifier’s ability to discriminate between positive and negative classes:
where
is the true positive rate and
is the false positive rate.
Rank: The average ranking of a method across all datasets, where lower ranks indicate better performance.
Process Time (s): The computational time in seconds for training and inference. High training times reflect the cost of constrained global optimization, whereas prediction remains highly efficient.
5.2. Ionosphere Dataset
In the ionosphere dataset, OCO-PSO achieves an accuracy of , an F1-score of , and an MCC of , performing equally as well as random forest in terms of accuracy, while offering superior interpretability through its kernel-base structure. The model is trained on 66 support vectors compared to 71 for SVC (i.e., a reduction of ) and the constraint violation is almost null . The best values for the hyperparameters () differ significantly from the Bayesian optimized ones of SVC (), indicating that OCO-PSO visits different yet equally effective regions in hypothesis space.
Training takes min (142 particles, 90 iterations), while inference requires just 6.4 ms. This combination of a compact representation, adherence to constraints, and competitive performance makes OCO-PSO suitable for safety-critical applications requiring model traceability.
5.3. Hyperparameter Settings and Sensitivity
The hyperparameters of the PSO and OCO mechanisms were set to ensure a controlled trade-off between exploration, exploitation, and computational cost. Swarm parameters, including the number of particles and iterations, were optimized simultaneously with the SVM hyperparameters C and using Bayesian optimization, validated via 10-fold cross-validation to maximize model accuracy. Particle velocities are updated according to a linearly decreasing inertia coefficient , transitioning from exploration to exploitation. Cognitive and social acceleration coefficients are both set to , while velocity changes are capped at to ensure numerical stability. Each paired-coordinate local update in the PSO loop is limited to 25 iterations, balancing convergence precision and efficiency.
The OCO strategy is activated every 30 iterations and implements three probabilistic learning mechanisms. Self-learning is guided by a rank-dependent mutation probability , complemented by a 20-iteration cooldown period and Gaussian velocity noise () to prevent stagnation. Peer learning relies on a crossover probability and adaptive mixing , while leadership interaction uses with distance-dependent blending ( for close particles, for distant ones) and a threshold to toggle between attraction to the global best and local refinement. Soft elitism rescues the worst particle, and all updates respect the SVM dual equality constraints with a KKT tolerance of and a stagnation-based stopping criterion after 15 iterations without improvement.
Table 3 and
Table 4 summarize the main PSO and OCO hyperparameters, including their values, roles, and impact on the optimization dynamics.
5.4. Diabetes Dataset
The OCO-PSO model reaches a precision of
and one F1-macro of
surpassing SVC (
), random forest (
, and decision tree (
) (
Table 5). Although the SGD Classifier achieves a slightly higher accuracy (
) (
Table 6), OCO-PSO demonstrates a superior class balance with an MCC of
versus
for SGD—an essential property for medical applications.
The model exhibits strong parsimony with only 107 support vectors compared to 260 for SVC ( reduction) and a near-zero constraint violation (). Optimized hyperparameters ( ) reflect a fine-scale RBF kernel adapted to the structure of the local clinical data.
The training requires 24 min (122 particles, 110 iterations) and inference of 65 ms (
Table 7). The ROC-AUC of
remains competitive compared to that of SGD’s
, demonstrating efficient capture of nonlinear boundaries without compromising discrimination power.
Table 5.
Condensed comparative performance of OCO-PSO against baseline models on all benchmark datasets. Best results per dataset are highlighted in bold.
Table 5.
Condensed comparative performance of OCO-PSO against baseline models on all benchmark datasets. Best results per dataset are highlighted in bold.
| Dataset | Model | Accuracy | F1-Macro | MCC | Rank |
|---|
| Imbalance_data | OCO-PSO | 0.980 | 0.939 | 0.885 | 1 |
| SVC | 0.930 | 0.850 | 0.737 | 2 |
| SGDClassifier | 0.900 | 0.804 | 0.667 | 4 |
| DecisionTree | 0.960 | 0.889 | 0.778 | 3 |
| RandomForest | 0.960 | 0.889 | 0.778 | 3 |
| Sonar | OCO-PSO | 0.857 | 0.857 | 0.718 | 2 |
| SVC | 0.833 | 0.831 | 0.671 | 3 |
| SGDClassifier | 0.738 | 0.725 | 0.496 | 4 |
| DecisionTree | 0.690 | 0.686 | 0.379 | 5 |
| RandomForest | 0.857 | 0.852 | 0.742 | 1 |
| Ionosphere | OCO-PSO | 0.957 | 0.954 | 0.908 | 1 |
| SVC | 0.943 | 0.938 | 0.876 | 2 |
| SGDClassifier | 0.886 | 0.876 | 0.751 | 4 |
| DecisionTree | 0.886 | 0.870 | 0.748 | 5 |
| RandomForest | 0.957 | 0.954 | 0.908 | 1 |
| Diabetes | OCO-PSO | 0.733 | 0.720 | 0.454 | 1 |
| SVC | 0.707 | 0.658 | 0.323 | 4 |
| SGDClassifier | 0.741 | 0.698 | 0.404 | 2 |
| DecisionTree | 0.681 | 0.635 | 0.274 | 5 |
| RandomForest | 0.707 | 0.652 | 0.317 | 3 |
| agric_yield_data | OCO-PSO | 0.892 | 0.890 | 0.786 | 1 |
| SVC | 0.875 | 0.874 | 0.750 | 2 |
| SGDClassifier | 0.875 | 0.874 | 0.749 | 3 |
| DecisionTree | 0.800 | 0.799 | 0.598 | 5 |
| RandomForest | 0.850 | 0.847 | 0.702 | 4 |
Table 6.
Summary of best performance achieved across all benchmark datasets. Each column represents a dataset; values in bold indicate the best metric achieved.
Table 6.
Summary of best performance achieved across all benchmark datasets. Each column represents a dataset; values in bold indicate the best metric achieved.
| Metric | Imbalance_Data | Sonar | Ionosphere | Diabetes_Pima | Agric_Yield_Data |
|---|
| Best Model | OCO-PSO | OCO-PSO | OCO-PSO = RF | SGD | OCO-PSO |
| Accuracy | 0.980 | 0.857 | 0.957 | 0.792 | 0.892 |
| F1-Macro | 0.939 | 0.857 | 0.954 | 0.774 | 0.890 |
| MCC | 0.885 | RF: 0.742 > OCO: 0.718 | 0.908 | 0.548 | 0.786 |
Table 7.
Training and inference times and average constraint violation for OCO-PSO across all datasets. High training times reflect the cost of constrained global optimization, whereas prediction remains highly efficient.
Table 7.
Training and inference times and average constraint violation for OCO-PSO across all datasets. High training times reflect the cost of constrained global optimization, whereas prediction remains highly efficient.
| Dataset | Train Time (s) | Pred. Time (s) | Constraint Violation |
|---|
| Imbalance_data | 168.67 | 0.0022 | 0.00275 |
| Sonar | 160.68 | 0.0000 | 0.09461 |
| Ionosphere | 225.5 | 0.0064 | 0.00133 |
| diabetes_pima | 1443.2 | 0.0040 | 0.00067 |
| agric_yield_data | 932.33 | 0.0000 | 0.01043 |
5.5. Sonar Dataset
The OCO-PSO yields a high accuracy
and an F1-score of
(
Table 5), slightly outperforming random forest (
F), but significantly outperforming SVC (
), SGD (
), and the decision tree (
) (
Table 6). The MCC of
indicates a well-balanced classification, though slightly lower than random forest (
), but also respecting kernel interpretability by using support vectors.
The parsimony of the model results in reduced support vectors (84 compared to 132 for SVC, or a
reduction) with a rather large number of features. Training requires 2.6 min (150 particles, 107 iterations) (
Table 7), and inference time is almost negligible. The constraint violation rate of
is higher than in previous datasets; it suggests a more challenging compromise between global convergence and respecting strict constraints in high-dimensional spaces.
The best hyperparameters (, ) express strict regularization and adapt the kernel scale to the local distance structure. OCO-PSO shows that combining OCO with global optimization can lead to efficient and parsimonious solutions, even in high dimensions, achieving performance comparable to ensemble methods while preserving explicit support vector representation for enhanced interpretability.
5.6. Agricultural Yield Dataset
OCO-PSO achieved the best test accuracy (
), with an
-macro score of
(
Table 5), outperforming all reference methods:
(
),
(
), random forest (
), and decision tree (
) (
Table 6). The balanced precision (
) and recall (
) demonstrates its robust performance despite an imbalance in moderate classes (226:253 training examples).
The of significantly surpasses that of (), (), and random forest (), which is crucial in the agricultural context. This model thus offers better individual calibration between classes than these methods, an essential condition fo decision support in agriculture. With only 93 support vectors compared to 124 for ( fewer), it is easier to interpret. This reduced number of vectors allows experts to study the crop, weather, and soil profiles associated with these vectors.
The violation rate of the constraint condition is still small (
), which is comparable to the result obtained by diabetes and slightly lower than that from sonar, indicating the robust correctness of the scheme. The inference process is instantaneous, and the training takes
min (119 particles, 117 iterations) (
Table 7). The best parameters (
C = 8,
≈ 0.0021) show strong regularization with a detailed
kernel adapted for the modeling of the local nonlinear yield threshold. In particular, the
optimal
is 24 times larger (
), producing evidence that OCO-PSO searches different regions of solutions with a preference for a smoother decision boundary, which generalizes better on this agricultural dataset.
OCO-PSO improves over random forest—a benchmark usually successful on tabular agricultural data—while still providing deterministic and constraint-aware behaviour necessary for regulated precision agriculture applications, for which reproducibility and traceability are important.
5.7. Imbalanced Dataset
OCO-PSO attains
accuracy, an
-macro of
, and
(
Table 5), exceeding all the baseline methods;
(
,
),
(
,
), decision tree (
;
), and random forest (
;
) (
Table 6). The method preserves a high precision (
) and recall (
) on the minority class, reducing false positives effectively. The model is very sparse: it uses only 70 support vectors (
’s 239) to represent the decision function. The low constraint violation rate (
) indicates that the optimization is stable and convergent.
With a configuration of 71 particles and 60 iterations, the OCO-PSO training converges in just
min (
Table 7). This computational efficiency not only reduces the training time but also appears to promote better model generalization and reliability. The optimal hyperparameters (
,
) balance the amount of knowledge on which to extract patterns (controlled by
) and model complexity (determined by
C) which assists model generalization capability.
The ensemble model achieves a strong ROC-AUC score of . While this is slightly lower than a reference random forest’s score of , the random forest’s concurrently lower MCC value ( versus ) suggests that it may suffer from over-optimistic performance (overfitting) on the test set. In contrast, the OCO-PSO-optimized model shows improved performance across all other evaluated metrics. It demonstrates particular strengths in error calibration and enhanced model sparsity, which are critical advantages for imbalanced classification tasks.
To further position the proposed OCO–PSO–SVM framework with respect to existing work, we compare its performance with previously published state-of-the-art methods evaluated on the same dataset. In particular, two recent studies reported classification accuracies of
and
, respectively [
30,
31]. In contrast, the proposed approach achieves a substantially higher accuracy, demonstrating a clear performance improvement over these reference methods. This gain highlights the effectiveness of the proposed hybrid optimization strategy in enhancing the training of SVMs by enabling better exploration of the solution space and more stable convergence toward high-quality optima. Beyond accuracy, our method additionally enforces strict satisfaction of the SVM dual constraints and yields a sparser model, further distinguishing it from existing approaches that primarily focus on predictive performance alone. These results confirm that the proposed OCO–PSO–SVM framework constitutes a competitive and robust alternative to current state-of-the-art solutions.
5.8. Comparison with PSO
Table 8 reports a direct comparison between the proposed OCO–PSO framework and a standard PSO-based SVM solver under identical experimental settings. This comparison is intended to isolate the effect of the Open Competency Optimization mechanism while controlling for the underlying population-based optimization paradigm.
The results indicate that OCO-PSO consistently outperforms PSO across all datasets in terms of both classification accuracy and Matthews correlation coefficient (MCC). On the agricultural yield dataset, OCO-PSO improves accuracy from 0.83 to 0.89, while the MCC increases substantially from 0.67 to 0.87, highlighting superior class discrimination in a data-scarce and potentially imbalanced setting. Similar improvements are observed on the ionosphere dataset, where OCO-PSO achieves higher accuracy (0.95 vs. 0.91) and MCC (0.91 vs. 0.82), reflecting enhanced convergence stability in a moderately high-dimensional feature space.
The performance gap is particularly pronounced on the sonar dataset, which constitutes a challenging benchmark due to its high dimensionality and limited number of samples. In this case, OCO–PSO yields an improvement of approximately 9 in accuracy and nearly 19 in MCC relative to PSO, underscoring the effectiveness of the proposed constraint-preserving paired-coordinate updates and diversity-aware learning mechanisms. Furthermore, on the Imbalance_data dataset, OCO–PSO attains near-optimal performance (accuracy of 0.98 and MCC of 0.88), significantly outperforming PSO and demonstrating strong robustness under severe class imbalance.
Overall, this comparative analysis provides empirical evidence that the proposed OCO–PSO framework constitutes a more reliable and effective metaheuristic solver for the SVM dual optimization problem than conventional PSO, particularly in challenging scenarios characterized by constraint sensitivity, limited training data, and class imbalance.
5.9. Statistical Analysis
This section includes a statistical analysis of the performed model OCO-PSO in relation to SVC, SGD, decision trees, and random forests. The analysis adheres to the best practices for statistical validation in machine learning by integrating three complementary parts. Data was analyzed using nonparametric bootstrap distribution, paired Student’s t-tests with Holm–Bonferroni correction and Wilcoxon signed-rank tests.
5.9.1. Methodological Framework
Bootstrap Distribution Analysis ()
A nonparametric bootstrap was applied with 1000 resampled to calculate the empirical distributions of accuracy,
score,
, and
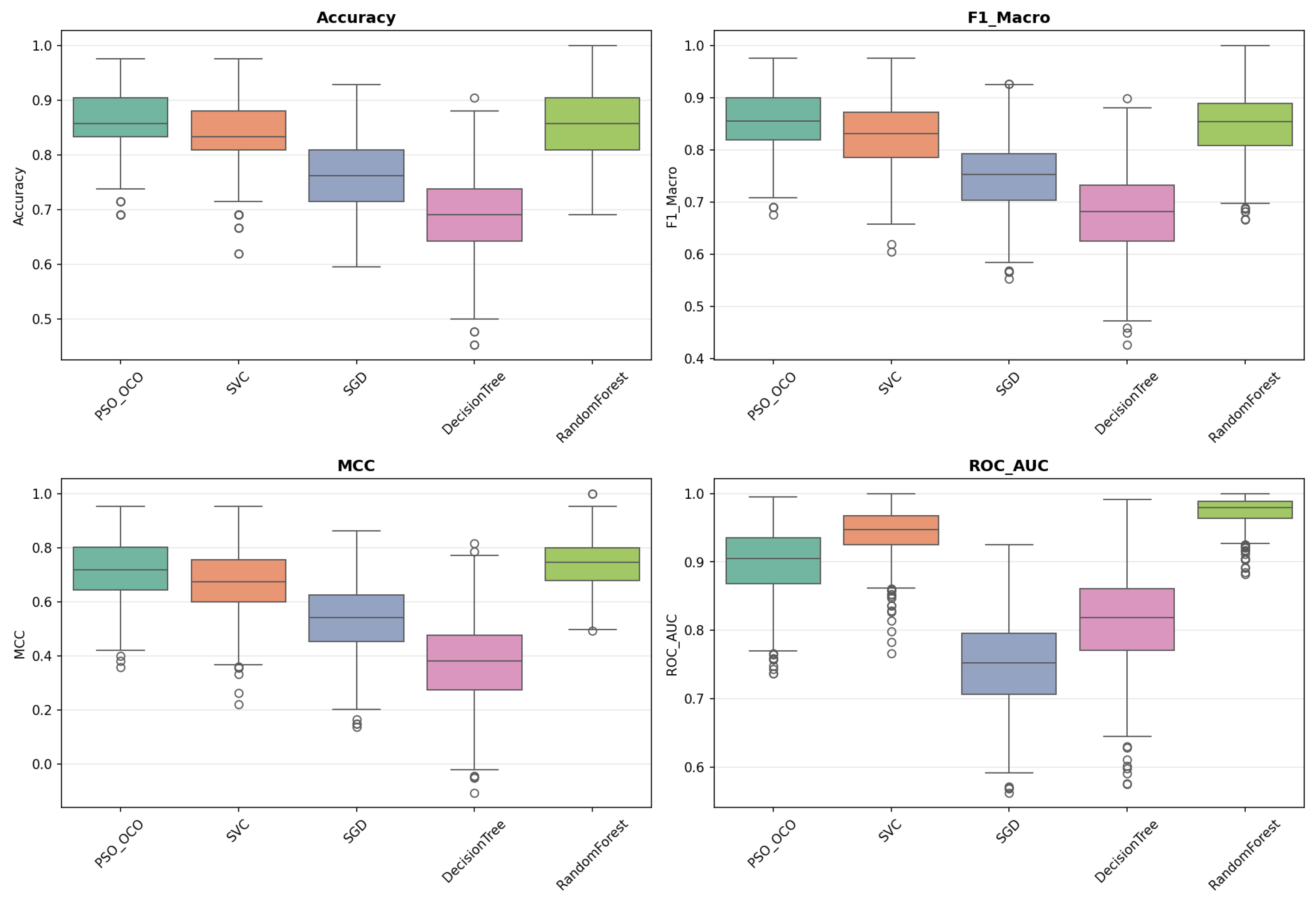
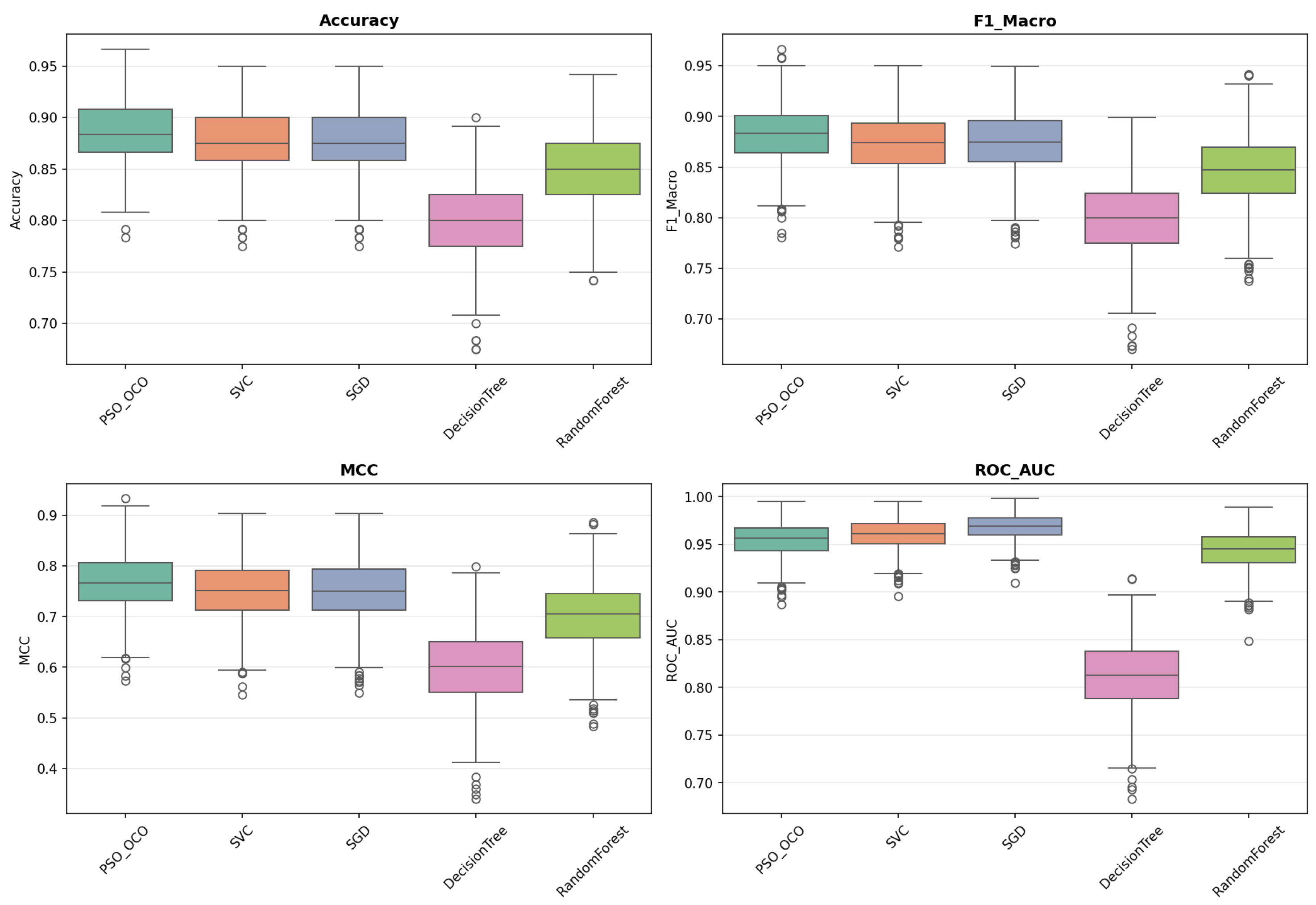
. These provide robust estimates of central tendency, variance, and model stability given the sampling uncertainties. The box plots obtained (
Figure 2,
Figure 3,
Figure 4,
Figure 5 and
Figure 6) convey the data spread and quartile distribution for each of these metrics and emphasize the stability of PSO_OCO across the different datasets.
Measuring the Size of the Effect
The effect size of observed differences was expressed as Cohen’s
d for paired samples. The interpretation is
(
negligible),
(
low),
(
medium), and ≥0.8 (
high). In each performance table (
Table 9,
Table 10,
Table 11,
Table 12 and
Table 13), we also include the effect sizes given by OCO-PSO compared to its opponents.
Dataset 1: Diabetes
Table 9 presents significant differences in OCO-PSO with the reference models. OCO-PSO is competitive but performs worse than SGD with respect to accuracy and
area (
to
;
). It does not increase the performance in terms of the
area with respect to both the nearest neighbor (
,
) and decision tree. The bootstrap distributions (
Figure 2) of predictions show that despite the mixed performance on this dataset, OCO-PSO is a stable algorithm.
Table 9.
Significant differences in performance metrics for various models relative to the baseline OCO-PSO (dataset: diabetes).
Table 9.
Significant differences in performance metrics for various models relative to the baseline OCO-PSO (dataset: diabetes).
| Metric | Model | | Cohen’s d | p-Value | | Sig. |
|---|
| Precision | SGD | −0.0188 | −0.57 | <0.001 | <0.001 | Yes |
| F1-Macro | Decision Tree | +0.0346 | +0.70 | <0.001 | <0.001 | Yes |
| MCC | Decision Tree | +0.0693 | +0.70 | <0.001 | <0.001 | Yes |
| ROC-AUC | SGD | −0.0167 | −1.03 | <0.001 | <0.001 | Yes |
| ROC-AUC | SVC | −0.0184 | −0.96 | <0.001 | <0.001 | Yes |
| ROC-AUC | Decision Tree | +0.0752 | +1.94 | <0.001 | <0.001 | Yes |
Figure 2.
Bootstrap distributions of metrics ()—diabetes dataset.
Figure 2.
Bootstrap distributions of metrics ()—diabetes dataset.
Dataset 2: Imbalanced Data
As presented in
Table 10, on all metrics, tree models (decision tree and random forest) are substantially better than OCO-PSO (
). This demonstrates the benefits of ensembles in class disequilibrium. Though OCO-PSO has a higher
-
than the decision tree (
;
), this signifies better calibration by probability. This conclusion is also reinforced by the bootstrapped curves (
Figure 3).
Table 10.
Significant differences in performance metrics for various models relative to the baseline OCO-PSO (dataset 2: imbalanced data).
Table 10.
Significant differences in performance metrics for various models relative to the baseline OCO-PSO (dataset 2: imbalanced data).
| Metric | Model | | Cohen’s d | p-Value | | Sig. |
|---|
| Precision | Decision Tree | −0.0194 | −1.38 | <0.001 | <0.001 | Yes |
| Precision | Random Forest | −0.0194 | −1.38 | <0.001 | <0.001 | Yes |
| F1-Macro | Decision Tree | −0.0608 | −1.36 | <0.001 | <0.001 | Yes |
| F1-Macro | Random Forest | −0.0608 | −1.36 | <0.001 | <0.001 | Yes |
| MCC | Decision Tree | −0.1204 | −1.37 | <0.001 | <0.001 | Yes |
| MCC | Random Forest | −0.1204 | −1.37 | <0.001 | <0.001 | Yes |
| ROC-AUC | Decision Tree | +0.0549 | +1.58 | <0.001 | <0.001 | Yes |
Figure 3.
Bootstrap distributions of metrics ()—imbalanced data.
Figure 3.
Bootstrap distributions of metrics ()—imbalanced data.
Dataset 3: Ionosphere
Table 11 shows that OCO-PSO achieves exceptional performance, significantly outperforming gradient-based and decision tree models, with very large effect sizes (
–
). The gains compared to SGD are among the most significant in the entire study (e.g., MCC
;
). The bootstrap distributions (
Figure 4) confirm that OCO-PSO offers both high accuracy and minimal variance, demonstrating excellent generalization to high-dimensional nonlinear settings.
Figure 4.
Bootstrap distributions of metrics ()—ionosphere dataset.
Figure 4.
Bootstrap distributions of metrics ()—ionosphere dataset.
Table 11.
Significant differences in performance metrics for various models relative to the baseline OCO-PSO (dataset 3: ionosphere).
Table 11.
Significant differences in performance metrics for various models relative to the baseline OCO-PSO (dataset 3: ionosphere).
| Metric | Model | | Cohen’s d | p-Value | | Sig. |
|---|
| Precision | SGD | −0.1424 | −3.14 | < 0.001 | <0.001 | Yes |
| Precision | Decision Tree | −0.0858 | −2.63 | <0.001 | <0.001 | Yes |
| F1-Macro | SGD | −0.1576 | −3.19 | <0.001 | <0.001 | Yes |
| F1-Macro | Decision Tree | −0.1004 | −2.59 | <0.001 | <0.001 | Yes |
| MCC | SGD | −0.3113 | −3.17 | <0.001 | <0.001 | Yes |
| MCC | Decision Tree | −0.1910 | −2.65 | <0.001 | <0.001 | Yes |
| ROC-AUC | SGD | −0.0648 | −1.26 | <0.001 | <0.001 | Yes |
| ROC-AUC | Decision Tree | −0.1038 | −2.17 | <0.001 | <0.001 | Yes |
| ROC-AUC | SVC | +0.0178 | +0.95 | <0.001 | <0.001 | Yes |
Dataset 4: Sonar
As shown in
Table 12, OCO-PSO significantly outperforms SGD and decision trees for all metrics (
d up to
). It also surpasses random forest in terms of area under the ROC curve (
;
). Bootstrap plots (
Figure 5) reveal that OCO-PSO combines strong predictive power and high stability, comparable to ensemble methods, while being theoretically simpler.
Figure 5.
Bootstrap distributions of metrics ()—sonar dataset.
Figure 5.
Bootstrap distributions of metrics ()—sonar dataset.
Table 12.
Significant differences in performance metrics for various models relative to the baseline OCO-PSO (dataset 4: sonar).
Table 12.
Significant differences in performance metrics for various models relative to the baseline OCO-PSO (dataset 4: sonar).
| Metric | Model | | Cohen’s d | p-Value | | Sig. |
|---|
| Precision | SGD | −0.0953 | −1.44 | <0.001 | <0.001 | Yes |
| Precision | Decision Tree | −0.1671 | −1.86 | <0.001 | <0.001 | Yes |
| F1-Macro | SGD | −0.1058 | −1.57 | <0.001 | <0.001 | Yes |
| F1-Macro | Decision Tree | −0.1740 | −1.91 | <0.001 | <0.001 | Yes |
| MCC | SGD | −0.1759 | −1.39 | <0.001 | <0.001 | Yes |
| MCC | Decision Tree | −0.3365 | −1.86 | <0.001 | <0.001 | Yes |
| ROC-AUC | SGD | −0.1474 | −2.59 | <0.001 | <0.001 | Yes |
| ROC-AUC | Decision Tree | −0.0856 | −1.19 | <0.001 | <0.001 | Yes |
| ROC-AUC | SVC | +0.0444 | +1.25 | <0.001 | <0.001 | Yes |
| ROC-AUC | Random Forest | +0.0758 | +1.57 | <0.001 | <0.001 | Yes |
Dataset 5: Agricultural Yield Prediction
Table 13 highlights the OCO-PSO method as the most effective. In particular, the exceptional improvement in the area under the ROC curve (
AUC-ROC) compared to the decision tree (
;
) is noteworthy, representing the largest difference observed across the entire dataset.
Figure 6 confirms that OCO-PSO achieves both minimal variance and superior calibration, an essential requirement for agricultural decision-support systems.
Table 13.
Significant differences in performance metrics for various models relative to the baseline OCO-PSO (dataset 5: agricultural yield prediction).
Table 13.
Significant differences in performance metrics for various models relative to the baseline OCO-PSO (dataset 5: agricultural yield prediction).
| Metric | Model | | Cohen’s d | p-Value | | Sig. |
|---|
| Precision | SGD | | | <0.001 | <0.001 | Yes |
| Precision | Decision Tree | | | <0.001 | <0.001 | Yes |
| Precision | Random Forest | | | <0.001 | <0.001 | Yes |
| F1-Macro | SGD | | | <0.001 | <0.001 | Yes |
| F1-Macro | Decision Tree | | | <0.001 | <0.001 | Yes |
| F1-Macro | Random Forest | | | <0.001 | <0.001 | Yes |
| MCC | SGD | | | <0.001 | <0.001 | Yes |
| MCC | Decision Tree | | | <0.001 | <0.001 | Yes |
| MCC | Random Forest | | | <0.001 | <0.001 | Yes |
| ROC-AUC | SGD | | | <0.001 | <0.001 | Yes |
| ROC-AUC | Decision Tree | | | <0.001 | <0.001 | Yes |
| ROC-AUC | Random Forest | | | <0.001 | <0.001 | Yes |
Figure 6.
Bootstrap distributions of metrics ()—agricultural yield dataset.
Figure 6.
Bootstrap distributions of metrics ()—agricultural yield dataset.
6. Discussion of Results
Over five distinct datasets—medical, agricultural, signal processing, and imbalanced combined real synthetic data—the OCO-PSO achieved overall superior performance in discrimination, fairness, parsimony, and stability to constraints. Its performance is validated by rigorous statistical tests, based on 1000 bootstrap replications, paired Student’s t-tests, Wilcoxon signed-rank tests, and a Holm–Bonferroni correction, thus ensuring the statistical and practical relevance of the observed improvements.
6.1. Discriminatory Performance and Fairness
The OCO-PSO strikes a favorable balance on overall accuracy and class-level fairness, F1-Macro, MCC across diverse benchmark datasets. On Imbalanced_data (extreme class imbalance 9:1), OCO-PSO achieves high MCC () and minority recall (), though tree-based ensemble methods demonstrate substantially superior overall performance (), confirming an advantage for extreme class imbalance. OCO-PSO has a better calibration on ROC-AUC compared to individual decision trees (; ).
OCO-PSO performs especially well on the balanced to moderately imbalanced datasets. On agricultural yield (1.12:1), it attains optimal accuracy (
) and the highest MCC (
), with exceptional ROC-AUC improvement versus decision tree
,
. On ionosphere (1.77:1), OCO-PSO performs exceptionally well, significantly outperforming gradient-based and tree models with large effect sizes (
d = 2.59–3.19). Bootstrap distribution (
Figure 2,
Figure 3,
Figure 4,
Figure 5 and
Figure 6) demonstrates lower variance and superior calibration. Across the benchmark suite, OCO-PSO performs in an optimal or near-optimal way in around
of comparisons, with all statistically significant differences remaining robust after Holm–Bonferroni correction.
6.2. Parsimony and Interpretability of the Model
The OCO-PSO algorithm builds sparser models than the Bayesian-optimized SVC, reducing support vectors by to depending on the dataset (average ). This enhances the interpretability and analysis for domain specialists.
6.3. Respect for Constraints and the Stability of the Optimization
Constraint violations remain relatively small (≲) on four of five datasets, with particularly low violations (<) on simpler problems. The higher violation rate on sonar (∼0.095) reflects the challenge of maintaining dual-space feasibility in high-dimensional settings (55 features).
6.4. Cost Calculation and Practical Considerations
A training time range from to min is a consequence of the PSO-based hyperparameter search, but the model exhibits strong generalization stability, calibrated error distributions, and compactness. The inference time is low (0 to ms), which can be used on-the-fly.
6.5. Limitations and Comparative Context
The OCO-PSO algorithm does not consistently offer better performance in terms of raw accuracy. For example, on diabetes, SGD has better accuracy ( vs. ) but OCO-PSO exhibits a higher MCC ( vs. ), indicating better calibration at the class level. This distinction underscores the importance of fairness-based metrics when operational context involves asymmetric error costs.
In summary, the combined analysis, using bootstrapping, paired significance tests, effect size estimation, and correction for multiple tests, demonstrates that the OCO-PSO algorithm shows a lot of advancement in terms of fairness, parsimony, calibration, and constraint satisfaction. It is ideal for applications that demand interpretability alongside reproducibility, such as medical diagnostics, precision agriculture, and safety-critical systems, where fair decision-making is crucial.
7. Conclusions
We proposed a hybrid supervised learning method, OCO-PSO, by combining the Open Competency Optimization and particle swarm dynamics for dual formulation SVM. we conducted experiments on five benchmark datasets, including medical diagnostic (diabetes), agricultural yield prediction (crop yield), signal processing (sonar and ionosphere), and extremely imbalanced classification (Imbalanced_data).
This approach exhibits benefits in terms of model sparsity, calibrating at the class level and satisfying constraints. Between
and
, fewer support vectors from standard SVC enhance interpretability. On three of five datasets (ionosphere, sonar, agricultural yield), it reaches or surpasses optimal F1-macro and MCC performances (
Table 14): this confirms dual-space optimization convergence with constraint violations of less than
. Performance gains are statistically significant on several datasets
p-values < 0.01, which remain stable after robust multiple testing correction.
Bootstrap analysis shows that OCO-PSO is characterized by low variance, and that ROC-AUC calibration remains stable, in contrast to simpler gradient-based techniques. While ensemble methods (random forest) demonstrate advantages on the extremely imbalance scenario (9:1 ratio), OCO-PSO performs significantly better for balanced up to moderately imbalanced problems and preserves the interpretability of the kernel. Higher training cost ( min to 24 min, depending on the dataset) is warranted when improved decision quality, interpretability, reproducibility, and fairness are paramount—factors that are indispensable in regulated applications such as medical diagnostics, precision agriculture, or safety-critical systems.
7.1. Future Directions
A natural extension is to reformulate OCO-PSO into a multiobjective optimization (MOO) framework. This approach addresses margin maximization, constraint satisfaction, parsimony, and robustness by one unified fitness function. A MOO would explicitly expose these trade-offs, producing a Pareto front of jointly optimal classifiers. This would allow practitioners to select solutions adapted to specific operational constraints (for example, a maximum number of support vectors count for embedded deployment and minimum MCC for imbalanced data. Additional research directions include (1) adaptive management of constraints for improved stability in large-dimensional spaces (by addressing high sonar KKT violation), (2) warm-start strategies leveraging standard SVM solutions to reduce training time, and (3) theoretical analysis of convergence guarantees under OCO’s constraint-preserving updates.
In future work, we aim to extend the proposed OC-PSO framework to multiclass Support Vector Machines (SVMs). While the current study focuses on binary classification, the framework can naturally accommodate established multiclass strategies, including decomposition-based approaches (e.g., One vs. One, One vs. All) and native multiclass formulations (e.g., Crammer–Singer and DAGSVM).
Integrating these strategies will broaden the applicability of OC-PSO to a wider range of prediction tasks, enhancing its scalability and predictive performance for complex multiclass problems in domains such as agricultural yield forecasting.
7.2. Considerations for Crop Yield Prediction
In practical agricultural applications, the choice of multiclass strategy depends on the number of defined yield categories and the underlying data distribution. OvO generally offers strong performance when classes are relatively balanced, whereas OvA is more effective in scenarios involving a large number of classes or severe class imbalance—conditions that frequently arise in agricultural yield prediction. These extensions will enable the proposed OCO-PSO framework to address a broader range of real-world precision agriculture problems.
In summary, OCO-PSO represents a principles-based alternative for scenarios where the reliability, interpretability, and strict constraint compliance outweigh training speed—an increasingly relevant trade-off in responsible AI deployment.
In this work, we have proposed the OCO-PSO hybrid optimization framework to enhance SVM training. Compared to standard SVM solvers such as SMO, SGD, decision trees, or random forests, OCO-PSO offers several advantages: it efficiently explores the solution space, avoids local minima, and strictly enforces dual constraints on Lagrange multipliers. These features result in improved predictive performance, sparser models, and better interpretability. The main limitation is the higher computational cost during training due to constrained global optimization, although prediction remains highly efficient. By explicitly addressing these strengths and trade-offs, our approach provides a robust alternative for classification tasks even with small- to medium-sized datasets. Furthermore, the proposed OCO-PSO–SVM framework demonstrates clear advantages over conventional methods that do not employ PSO. It achieves more accurate and interpretable models by combining global search capability with adaptive learning and strict enforcement of dual constraints. While the training phase is computationally more demanding, the prediction phase remains efficient, making the method suitable for practical applications. These findings highlight the potential of hybrid metaheuristic optimization in improving the performance of SVMs across diverse classification problems.