Nearest Neighbours Graph Variational AutoEncoder
Abstract
1. Introduction
- A pooling operation for graph data that takes into account graph connectivity and, at the same time, is lightweight and scalable to a large graph;
- A graph generative model based on an encoder–decoder architecture;
- A decoding solution that is based on the message passing algorithm.
- Simple, symmetrical, and geometry-based pooling and unpooling operations on graphs, which allow for the creation of bottlenecks in neural network architectures and that are scalable to large graphs;
- A Variational AutoEncoder for regular graph data, where both the encoding and decoding modules use graph convolutional layers to fully exploit the graph structure during the learning process.
2. Materials and Methods
2.1. Nearest Neighbour Graph VAE
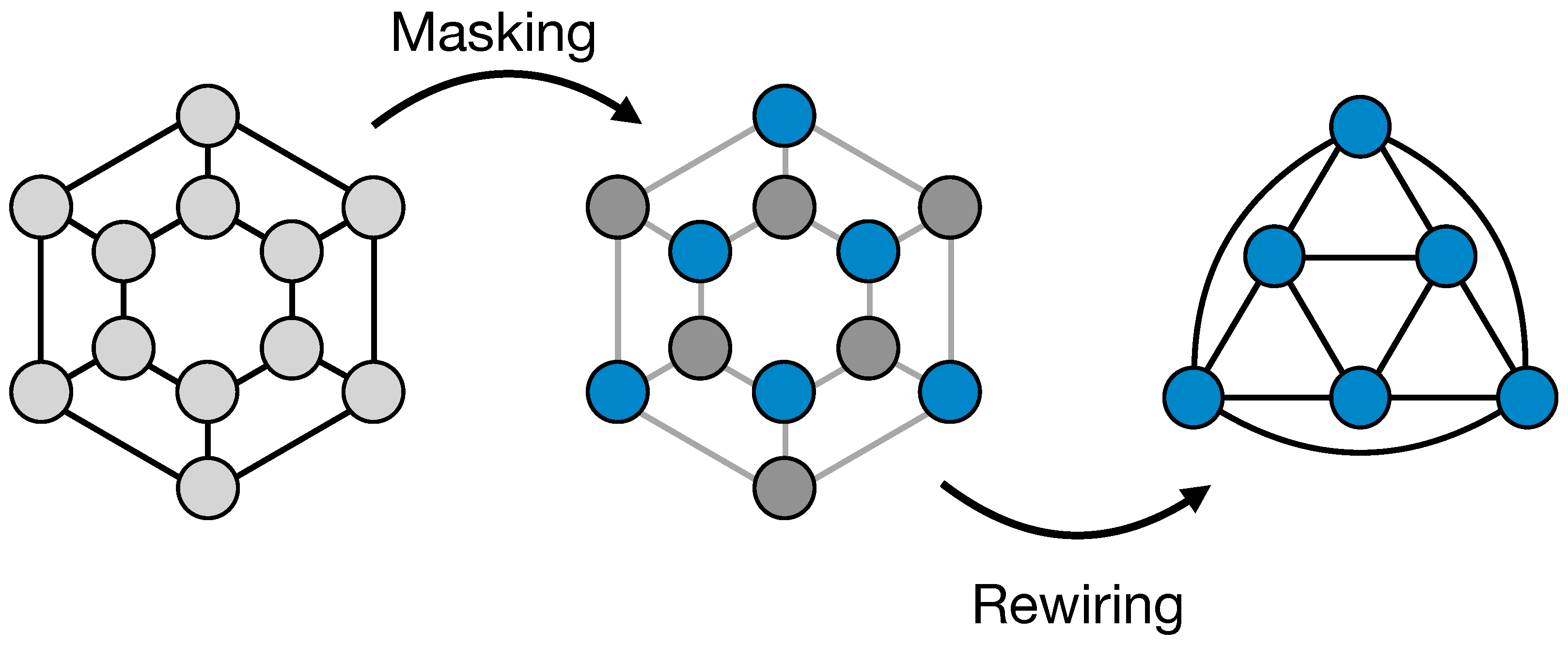
2.1.1. ReNN-Pool and Un-Pool
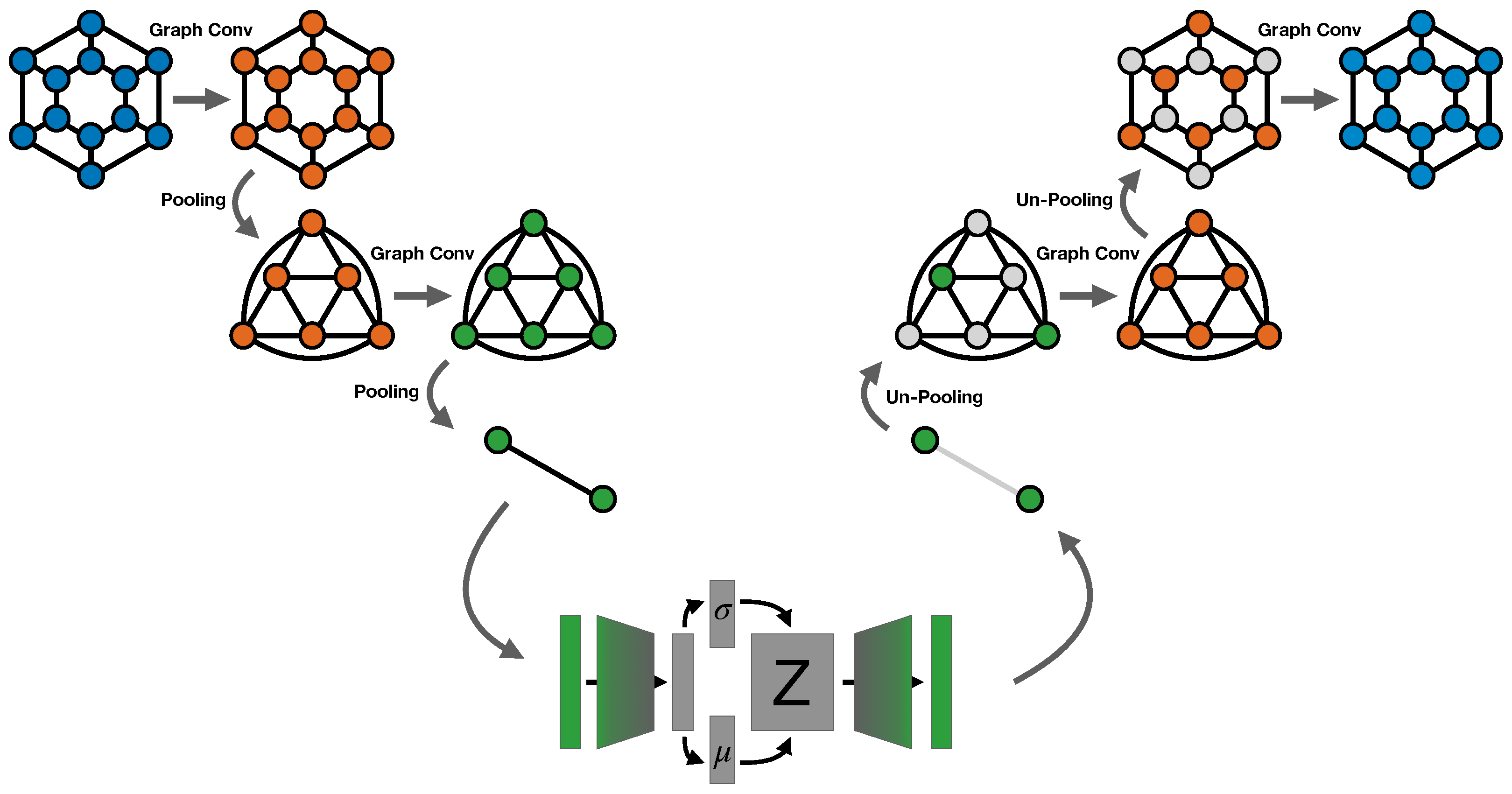
2.1.2. ReNN Graph VAE Architecture
2.2. Datasets
2.2.1. Energy Deposition Datasets
2.2.2. Sprite Dataset
3. Results
3.1. Results on Energy Deposition Datasets
- Total energy: computed by summing the features of all nodes.
- z profile: computed by integrating, i.e., by summing, the features of all nodes along the r and axes.
- r profile: computed by integrating, i.e., by summing, the features of all nodes along the z and axes.
3.2. Results on Sprite Dataset
3.3. Ablation Study on Pooling
- ReNN-Pool: the one proposed in this work with only the simple node masking;
- ReNN Mean Pool: mean pooling on clusters defined by the masking operation of ReNN-Pool;
- ReNN Max Pool: max pooling on clusters defined by the masking operation of ReNN-Pool;
- Random Pool: dropping random nodes in the graph;
- Top-k Pool: defined in [22], dropping nodes on the base of features’ alignment with a learnable vector.
4. Discussion
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations and Symbols
| DL | Deep Learning |
| CNN | Convolutional Neural Network |
| GAN | Generative Adversarial Networks |
| VAE | Variational AutoEncoder |
| RT | Radiotherapy |
| GNN | Graph Neural Networks |
| GCN | Graph Convolutional layers |
| GAE | Graph AutoEncoders |
| VGAE | Variational Graph AutoEncoders |
| ReNN-Pool | Recursive Nearest Neighbour Pooling |
| ELBO | Evidence Lower Bound |
| Cylindrical coordinate system | |
| Node feature vector | |
| Adjacency matrix | |
| M | ReNN-Pool masking vector |
| N | Number of nodes in the graph |
| Number of neighbours of node i | |
| Weight of the edge between node i and j | |
| Generic weight of the Neural network | |
| Z | Latent space variable |
| -index |
Appendix A. Full Model Description
| Layers | Parameters | N Nodes | N Edges | |
|---|---|---|---|---|
| Graph Encoder | GraphConv (1, 16, ‘mean’) | 48 | 21,952 | 128,576 |
| ReNN-Pool | - | 10,976 | 188,216 | |
| GraphConv (16, 32, ‘mean’) | 1056 | 10,976 | 188,216 | |
| ReNN-Pool | - | 1470 | 21,952 | |
| GraphConv (32, 64, ‘mean’) | 4160 | 1470 | 21,952 | |
| ReNN-Pool | - | 236 | 6206 | |
| Linear (64 × 236, 64) | 966,720 | - | - | |
| Linear (64, 2*) | 130 | - | - | |
| Linear (64, 2*) | 130 | - | - | |
| Graph Decoder | Linear (2*, 64) | 192 | - | - |
| Linear (64, 64 × 236) | 981,760 | - | - | |
| ReNN-Unpool | - | 1470 | 21,952 | |
| GraphConv (64, 32, ‘mean’) | 4128 | 1470 | 21,952 | |
| ReNN-Unpool | - | 10,976 | 188,216 | |
| GraphConv (32, 16, ‘mean’) | 1040 | 10,976 | 188,216 | |
| ReNN-Unpool | - | 21,952 | 128,576 | |
| GraphConv (16, 1, ‘mean’) | 33 | 21,952 | 128,576 |
| Layers | Parameters | N Nodes | N Edges | |
|---|---|---|---|---|
| Graph Encoder | GraphConv (3, 16, ‘mean’) | 112 | 4096 | 16,128 |
| ReNN-Pool | - | 2048 | 15,874 | |
| Linear (1, 15,874) | 31,748 | - | - | |
| GraphConv (16, 32, ‘mean’, ) | 1056 | 2048 | 15,874 | |
| ReNN-Pool | - | 528 | 3906 | |
| Linear (1, 3906) | 7812 | - | - | |
| GraphConv (32, 64, ‘mean’, ) | 4160 | 528 | 3906 | |
| ReNN-Pool | - | 136 | 930 | |
| Linear (64 × 136, 64) | 557,120 | - | - | |
| Linear (64, 5) | 325 | - | - | |
| Linear (64, 5) | 325 | - | - | |
| Graph Decoder | Linear (5, 64) | 384 | - | - |
| Linear (64, 64 × 136) | 565,760 | - | - | |
| ReNN-Unpool | - | 528 | 3906 | |
| Linear (1, 3906) | 7812 | - | - | |
| GraphConv (64, 32, ‘mean’, ) | 4128 | 528 | 3906 | |
| ReNN-Unpool | - | 2048 | 15,874 | |
| Linear (1, 15,874) | 31,748 | - | - | |
| GraphConv (32, 16, ‘mean’, ) | 1040 | 2048 | 15,874 | |
| ReNN-Unpool | - | 4096 | 16,128 | |
| GraphConv (16, 3, ‘mean’) | 99 | 4096 | 16,128 |
Appendix B. Variational AutoEncoder
Appendix C. ReNN Graph VAE vs. CNN VAE

| Layers | Parameters | |
|---|---|---|
| Graph Encoder | Conv2d (3, 16, kernel_size = (3, 3, 3)) | 448 |
| AvgPool2d (kernel_size = 2, stride = 2) | - | |
| Conv2d (16, 32, kernel_size = (3, 3, 3)) | 4640 | |
| AvgPool2d (kernel_size = 2, stride = 1) | - | |
| Conv2d (32, 64, kernel_size = (3, 3, 3)) | 18,496 | |
| AvgPool2d (kernel_size = 2, stride = 2) | - | |
| Linear (64 × 169, 64) | 692,288 | |
| Linear (64, 5) | 325 | |
| Linear (64, 5) | 325 | |
| Graph Decoder | Linear (5, 64) | 384 |
| Linear (64, 64 × 169) | 703,040 | |
| Upsample (size = (26, 26), mode = ‘bilinear’) | - | |
| ConvTranspose2d (64, 32, kernel_size = (3, 3, 3)) | 18,464 | |
| Upsample (size = (29, 29), mode = ‘bilinear’) | - | |
| ConvTranspose2d (32, 16, kernel_size = (3, 3, 3)) | 4624 | |
| Upsample (size = (62, 62), mode = ‘bilinear’) | - | |
| ConvTranspose2d (16, 3, kernel_size = (3, 3, 3)) | 435 |
References
- Goodfellow, I.J.; Pouget-Abadie, J.; Mirza, M.; Xu, B.; Warde-Farley, D.; Ozair, S.; Courville, A.; Bengio, Y. Generative Adversarial Networks. arXiv 2014, arXiv:1406.2661. [Google Scholar] [CrossRef]
- Kingma, D.P.; Welling, M. Auto-Encoding Variational Bayes. arXiv 2014, arXiv:1312.6114. [Google Scholar]
- Rezende, D.; Mohamed, S. Variational Inference with Normalizing Flows. In Proceedings of the Machine Learning Research (PMLR), Proceedings of the 32nd International Conference on Machine Learning, Lille, France, 6–11 July 2015; Volume 37, pp. 1530–1538. [Google Scholar]
- Bond-Taylor, S.; Leach, A.; Long, Y.; Willcocks, C.G. Deep Generative Modelling: A Comparative Review of VAEs, GANs, Normalizing Flows, Energy-Based and Autoregressive Models. IEEE Trans. Pattern Anal. Mach. Intell. 2022, 44, 7327–7347. [Google Scholar] [CrossRef]
- Mentzel, F.; Kröninger, K.; Lerch, M.; Nackenhorst, O.; Paino, J.; Rosenfeld, A.; Saraswati, A.; Tsoi, A.C.; Weingarten, J.; Hagenbuchner, M.; et al. Fast and accurate dose predictions for novel radiotherapy treatments in heterogeneous phantoms using conditional 3D-UNet generative adversarial networks. Med. Phys. 2022, 49, 3389–3404. [Google Scholar] [CrossRef]
- Zhang, X.; Hu, Z.; Zhang, G.; Zhuang, Y.; Wang, Y.; Peng, H. Dose calculation in proton therapy using a discovery cross-domain generative adversarial network (DiscoGAN). Med. Phys. 2021, 48, 2646–2660. [Google Scholar] [CrossRef] [PubMed]
- Mendonça, R.V.; Silva, J.C.; Rosa, R.L.; Saadi, M.; Rodriguez, D.Z.; Farouk, A. A lightweight intelligent intrusion detection system for industrial internet of things using deep learning algorithms. Expert Syst. 2022, 39, e12917. [Google Scholar] [CrossRef]
- Beniczky, S.; Karoly, P.; Nurse, E.; Ryvlin, P.; Cook, M. Machine learning and wearable devices of the future. Epilepsia 2021, 62, S116–S124. [Google Scholar] [CrossRef]
- Khan, N.; Ullah, A.; Haq, I.U.; Menon, V.G.; Baik, S.W. SD-Net: Understanding overcrowded scenes in real-time via an efficient dilated convolutional neural network. J. Real-Time Image Process. 2021, 18, 1729–1743. [Google Scholar] [CrossRef]
- Francescato, S.; Giagu, S.; Riti, F.; Russo, G.; Sabetta, L.; Tortonesi, F. Model compression and simplification pipelines for fast deep neural network inference in FPGAs in HEP. Eur. Phys. J. C 2021, 81, 969. [Google Scholar] [CrossRef]
- Morris, C.; Ritzert, M.; Fey, M.; Hamilton, W.L.; Lenssen, J.E.; Rattan, G.; Grohe, M. Weisfeiler and Leman Go Neural: Higher-order Graph Neural Networks. arXiv 2021, arXiv:1810.02244. [Google Scholar] [CrossRef]
- Kipf, T.N.; Welling, M. Semi-Supervised Classification with Graph Convolutional Networks. arXiv 2017, arXiv:1609.02907. [Google Scholar]
- Defferrard, M.; Bresson, X.; Vandergheynst, P. Convolutional Neural Networks on Graphs with Fast Localized Spectral Filtering. arXiv 2017, arXiv:1606.09375. [Google Scholar]
- Veličković, P.; Cucurull, G.; Casanova, A.; Romero, A.; Liò, P.; Bengio, Y. Graph Attention Networks. arXiv 2018, arXiv:1710.10903. [Google Scholar]
- Zhu, Y.; Du, Y.; Wang, Y.; Xu, Y.; Zhang, J.; Liu, Q.; Wu, S. A Survey on Deep Graph Generation: Methods and Applications. arXiv 2022, arXiv:2203.06714. [Google Scholar]
- Kipf, T.N.; Welling, M. Variational Graph Auto-Encoders. arXiv 2016, arXiv:1611.07308. [Google Scholar]
- Dhillon, I.S.; Guan, Y.; Kulis, B. Weighted Graph Cuts without Eigenvectors A Multilevel Approach. IEEE Trans. Pattern Anal. Mach. Intell. 2007, 29, 1944–1957. [Google Scholar] [CrossRef]
- Zhang, M.; Cui, Z.; Neumann, M.; Chen, Y. An End-to-End Deep Learning Architecture for Graph Classification. In Proceedings of the AAAI Conference on Artificial Intelligence, New Orleans, LA, USA, 2–7 February 2018; Volume 32. [Google Scholar] [CrossRef]
- Bianchi, F.M.; Grattarola, D.; Livi, L.; Alippi, C. Hierarchical Representation Learning in Graph Neural Networks with Node Decimation Pooling. IEEE Trans. Neural Netw. Learn. Syst. 2022, 33, 2195–2207. [Google Scholar] [CrossRef]
- Bravo-Hermsdorff, G.; Gunderson, L.M. A Unifying Framework for Spectrum-Preserving Graph Sparsification and Coarsening. arXiv 2020, arXiv:1902.09702. [Google Scholar]
- Ying, R.; You, J.; Morris, C.; Ren, X.; Hamilton, W.L.; Leskovec, J. Hierarchical Graph Representation Learning with Differentiable Pooling. arXiv 2019, arXiv:1806.08804. [Google Scholar]
- Gao, H.; Ji, S. Graph U-Nets. arXiv 2019, arXiv:1905.05178. [Google Scholar] [CrossRef]
- Ranjan, E.; Sanyal, S.; Talukdar, P. Asap: Adaptive structure aware pooling for learning hierarchical graph representations. In Proceedings of the AAAI Conference on Artificial Intelligence, New York, NY, USA, 7–12 February 2020; Volume 34, pp. 5470–5477. [Google Scholar]
- Guo, Y.; Zou, D.; Lerman, G. An Unpooling Layer for Graph Generation. arXiv 2022, arXiv:2206.01874. [Google Scholar]
- Liu, Q.; Allamanis, M.; Brockschmidt, M.; Gaunt, A.L. Constrained Graph Variational Autoencoders for Molecule Design. arXiv 2019, arXiv:1805.09076. [Google Scholar]
- Bresson, X.; Laurent, T. A Two-Step Graph Convolutional Decoder for Molecule Generation. arXiv 2019, arXiv:1906.03412. [Google Scholar]
- Guo, X.; Zhao, L.; Qin, Z.; Wu, L.; Shehu, A.; Ye, Y. Interpretable Deep Graph Generation with Node-Edge Co-Disentanglement. In Proceedings of the 26th ACM SIGKDD International Conference on Knowledge Discovery & Data Mining, Virtual, 6–10 July 2020; pp. 1697–1707. [Google Scholar] [CrossRef]
- Assouel, R.; Ahmed, M.; Segler, M.H.; Saffari, A.; Bengio, Y. DEFactor: Differentiable Edge Factorization-based Probabilistic Graph Generation. arXiv 2018, arXiv:1811.09766. [Google Scholar]
- Du, Y.; Guo, X.; Cao, H.; Ye, Y.; Zhao, L. Disentangled Spatiotemporal Graph Generative Models. In Proceedings of the AAAI Conference on Artificial Intelligence, Virtual, 22 February–1 March 2022; Volume 36, pp. 6541–6549. [Google Scholar] [CrossRef]
- Shelhamer, E.; Long, J.; Darrell, T. Fully Convolutional Networks for Semantic Segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 2017, 39, 640–651. [Google Scholar] [CrossRef]
- Lin, B.; Gao, F.; Yang, Y.; Wu, D.; Zhang, Y.; Feng, G.; Dai, T.; Du, X. FLASH Radiotherapy: History and Future. Front. Oncol. 2021, 11. [Google Scholar] [CrossRef]
- Agostinelli, S.; Allison, J.; Amako, K.; Apostolakis, J.; Araujo, H.; Arce, P.; Asai, M.; Axen, D.; Banerjee, S.; Barrand, G.; et al. Geant4—A simulation toolkit. Nucl. Instruments Methods Phys. Res. Sect. A Accel. Spectrometers Detect. Assoc. Equip. 2003, 506, 250–303. [Google Scholar] [CrossRef]
- Li, Y.; Mandt, S. Disentangled Sequential Autoencoder. arXiv 2018, arXiv:1803.02991. [Google Scholar]
- Low, D.A.; Harms, W.B.; Mutic, S.; Purdy, J.A. A technique for the quantitative evaluation of dose distributions. Med. Phys. 1998, 25, 656–661. [Google Scholar] [CrossRef]
- Higgins, I.; Matthey, L.; Pal, A.; Burgess, C.; Glorot, X.; Botvinick, M.; Mohamed, S.; Lerchner, A. beta-VAE: Learning Basic Visual Concepts with a Constrained Variational Framework. In Proceedings of the International Conference on Learning Representations, Toulon, France, 24–26 April 2017. [Google Scholar]






| Dataset | z Profile Error | r Profile Error | Total Energy Error | < 3% |
|---|---|---|---|---|
| Water | 5.8 ± 3.4% | 2.6 ± 1.6% | 2.2 ± 1.6% | 99.3 ± 0.1% |
| Water + Slab | 6.9 ± 3.4% | 3.0 ± 1.2% | 2.2 ± 1.6% | 98.6 ± 0.3% |
| Pooling | z Profile Error | r Profile Error | Total Energy Error | |
|---|---|---|---|---|
| ReNN-Pool | 6.9 ± 3.4% | 3.0 ± 1.2% | 2.2 ± 1.6% | 98.6 ± 0.3% |
| ReNN Mean Pool | 6.4 ± 3.0% | 2.8 ± 1.1% | 2.0 ± 1.4% | 98.6 ± 0.3% |
| ReNN Max Pool | 22.6 ± 9.9% | 5.0 ± 1.8% | 3.2 ± 2.3% | 97.5 ± 0.7% |
| Random Pool | 172.6 ± 21.7% | 52.2 ± 3.7% | 2.0 ± 1.5% | 92.4 ± 0.4% |
| Top-k Pool | 51.7 ± 3.4% | 75.1 ± 9.1% | 4.0 ± 2.6% | 79.9 ± 1.3% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Arsini, L.; Caccia, B.; Ciardiello, A.; Giagu, S.; Mancini Terracciano, C. Nearest Neighbours Graph Variational AutoEncoder. Algorithms 2023, 16, 143. https://doi.org/10.3390/a16030143
Arsini L, Caccia B, Ciardiello A, Giagu S, Mancini Terracciano C. Nearest Neighbours Graph Variational AutoEncoder. Algorithms. 2023; 16(3):143. https://doi.org/10.3390/a16030143
Chicago/Turabian StyleArsini, Lorenzo, Barbara Caccia, Andrea Ciardiello, Stefano Giagu, and Carlo Mancini Terracciano. 2023. "Nearest Neighbours Graph Variational AutoEncoder" Algorithms 16, no. 3: 143. https://doi.org/10.3390/a16030143
APA StyleArsini, L., Caccia, B., Ciardiello, A., Giagu, S., & Mancini Terracciano, C. (2023). Nearest Neighbours Graph Variational AutoEncoder. Algorithms, 16(3), 143. https://doi.org/10.3390/a16030143






