High Per Parameter: A Large-Scale Study of Hyperparameter Tuning for Machine Learning Algorithms
Abstract
:1. Introduction
2. Previous Work
The Current Study
- Consider significantly more algorithms;
- Consider significantly more datasets;
- Consider Bayesian optimization, rather than weaker-performing random search or grid search.
3. Experimental Setup
- Datasets;
- Algorithms;
- Metrics;
- Hyperparameter tuning;
- Overall flow.
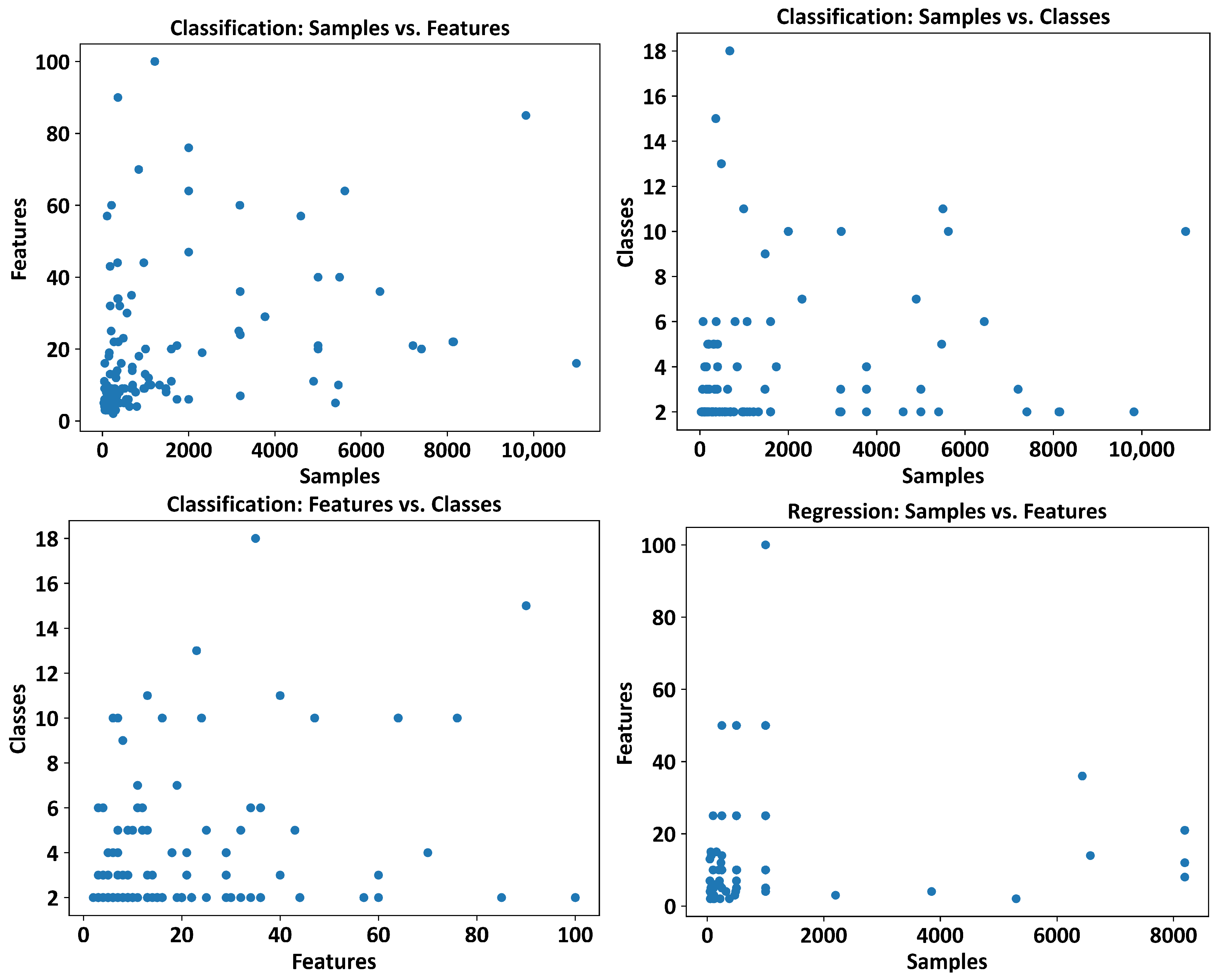
3.1. Datasets
3.2. Algorithms
3.3. Metrics
- Accuracy: a fraction of correct predictions (∈).
- Balanced accuracy: an accuracy score that takes into account class imbalances, essentially the accuracy score with class-balanced sample weights [14] (∈).
- F1 score: a harmonic mean of precision and recall; in the multi-class case, this is the average of the F1 score per class with weighting (∈)
- R2 score: an (coefficient of determination) regression score function (∈).
- Adjusted R2 score: a modified version of the R2 score that adjusts for the number of predictors in a regression model. It is defined as , with being the R2 score, n being the number of samples, and p being the number of features (∈).
- Complement RMSE: a complement of root mean squared error (RMSE), defined as (∈). This has the same range as the previous two metrics.
3.4. Hyperparameter Tuning
3.5. Overall Flow
- Optuna is run over the training set for 50 trials to tune the model’s hyperparameters, the best model is retained, and the best model’s test-set metric score is computed.
- Fifty models are evaluated over the training set with default parameters, the best model is retained, and the best model’s test-set metric score is computed. Strictly speaking, a few algorithms—decision tree, KNN, Bayesian—are essentially deterministic. For consistency, we still performed the 50 default hyperparameter trials. Further, our examination of the respective implementations revealed possible randomness, e.g., for decision tree, when max_features < n_features, the algorithm will select max_features at random; though the default is max_features = n_features we still took no chances of there being some hidden randomness deep within the code.
| Algorithm 1 Experimental setup (per algorithm and dataset) |
|
| # 'metric1', 'metric2', 'metric3' are, respectively: # · For classification: accuracy, balanced accuracy, F1 # · For regression: R2, adjusted R2, complement RMSE #eval_score is 5-fold cross-validation score |
|
4. Results
- The 13 algorithms and 9 measures of Table 2 are considered (separately for classifiers and regressors) as a dataset with 13 samples and the following 9 features: metric1_median, metric2_median, metric3_median, metric1_mean, metric2_mean, metric3_mean, metric1_std, metric2_std, metric3_std.
- Scikit-learn’s RobustScaler is applied, which scales features using statistics that are robust to outliers: “This Scaler removes the median and scales the data according to the quantile range (defaults to IQR: Interquartile Range). The IQR is the range between the 1st quartile (25th quantile) and the 3rd quartile (75th quantile). Centering and scaling happen independently on each feature…” [14].
- The hp_score of an algorithm is then simply the mean of its nine scaled features.
5. Discussion
- Decide how much to invest in hyperparameter tuning of a particular algorithm;
- Select algorithms that require less tuning to hopefully save time—as well as energy [15].
6. Concluding Remarks
- Algorithms may be added to the study.
- Datasets may be added to the study.
- Hyperparameters that have not been considered herein may be added.
- Specific components of the setup may be managed (e.g., the metrics and the scaler of Algorithm 1).
- Additional summary scores, like the hp_score, may be devised.
- For algorithms at the top of the lists in Table 3, we may inquire as to whether particular hyperparameters are the root cause of their hyperparameter sensitivity; further, we may seek out better defaults. For example, [16] recently focused on hyperparameter tuning for KernelRidge, which is at the top of the regressor list in Table 3. Ref [6] discussed the tunability of a specific hyperparameter, though they noted the problem of hyperparameter dependency.
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Bergstra, J.; Yamins, D.; Cox, D.D. Hyperopt: A python library for optimizing the hyperparameters of machine learning algorithms. In Proceedings of the 12th Python in Science Conference, Austin, TX, USA, 11–17 July 2013; Volume 13, p. 20. [Google Scholar]
- Akiba, T.; Sano, S.; Yanase, T.; Ohta, T.; Koyama, M. Optuna: A Next-Generation Hyperparameter Optimization Framework. In Proceedings of the 25th ACM SIGKDD International Conference on Knowledge Discovery & Data Mining, Anchorage, AK, USA, 4–8 August 2019; Association for Computing Machinery: New York, NY, USA, 2019; pp. 2623–2631. [Google Scholar]
- Sipper, M.; Moore, J.H. AddGBoost: A gradient boosting-style algorithm based on strong learners. Mach. Learn. Appl. 2022, 7, 100243. [Google Scholar] [CrossRef]
- Sipper, M. Neural networks with à la carte selection of activation functions. SN Comput. Sci. 2021, 2, 1–9. [Google Scholar] [CrossRef]
- Bischl, B.; Binder, M.; Lang, M.; Pielok, T.; Richter, J.; Coors, S.; Thomas, J.; Ullmann, T.; Becker, M.; Boulesteix, A.L.; et al. Hyperparameter Optimization: Foundations, Algorithms, Best Practices and Open Challenges. arXiv 2021, arXiv:2107.05847. [Google Scholar]
- Probst, P.; Boulesteix, A.L.; Bischl, B. Tunability: Importance of Hyperparameters of Machine Learning Algorithms. J. Mach. Learn. Res. 2019, 20, 1–32. [Google Scholar]
- Weerts, H.J.P.; Mueller, A.C.; Vanschoren, J. Importance of Tuning Hyperparameters of Machine Learning Algorithms. arXiv 2020, arXiv:2007.07588. [Google Scholar]
- Turner, R.; Eriksson, D.; McCourt, M.; Kiili, J.; Laaksonen, E.; Xu, Z.; Guyon, I. Bayesian Optimization is Superior to Random Search for Machine Learning Hyperparameter Tuning: Analysis of the Black-Box Optimization Challenge 2020. In Proceedings of the NeurIPS 2020 Competition and Demonstration Track, Virtual Event/Vancouver, BC, Canada, 6–12 December 2020; Volume 133, pp. 3–26. [Google Scholar]
- Bergstra, J.; Bengio, Y. Random Search for Hyper-Parameter Optimization. J. Mach. Learn. Res. 2012, 13, 281–305. [Google Scholar]
- Romano, J.D.; Le, T.T.; La Cava, W.; Gregg, J.T.; Goldberg, D.J.; Chakraborty, P.; Ray, N.L.; Himmelstein, D.; Fu, W.; Moore, J.H. PMLB v1.0: An open source dataset collection for benchmarking machine learning methods. arXiv 2021, arXiv:2012.00058v2. [Google Scholar]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; et al. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Chen, T.; Guestrin, C. XGBoost: A Scalable Tree Boosting System. In Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco, CA, USA, 13–17 August 2016; ACM: New York, NY, USA, 2016; pp. 785–794. [Google Scholar] [CrossRef]
- Ke, G.; Meng, Q.; Finley, T.; Wang, T.; Chen, W.; Ma, W.; Ye, Q.; Liu, T.Y. LightGBM: A highly efficient gradient boosting decision tree. Adv. Neural Inf. Process. Syst. 2017, 30, 3146–3154. [Google Scholar]
- Scikit-Learn: Machine Learning in Python. 2022. Available online: https://scikit-learn.org/ (accessed on 22 June 2022).
- García-Martín, E.; Rodrigues, C.F.; Riley, G.; Grahn, H. Estimation of energy consumption in machine learning. J. Parallel Distrib. Comput. 2019, 134, 75–88. [Google Scholar] [CrossRef]
- Stuke, A.; Rinke, P.; Todorović, M. Efficient hyperparameter tuning for kernel ridge regression with Bayesian optimization. Mach. Learn. Sci. Technol. 2021, 2, 035022. [Google Scholar] [CrossRef]
| Classification | ||
|---|---|---|
| Algorithm | Hyperparameter | Values |
| AdaBoostClassifier | n_estimators | [10, 1000] (log) |
| learning_rate | [0.1, 10] (log) | |
| DecisionTreeClassifier | max_depth | [2, 10] |
| min_impurity_decrease | [0.0, 0.5] | |
| criterion | {gini, entropy} | |
| GradientBoostingClassifier | n_estimators | [10, 1000] (log) |
| learning_rate | [0.01, 0.3] | |
| subsample | [0.1, 1] | |
| KNeighborsClassifier | weights | {uniform, distance} |
| algorithm | {auto, ball_tree, kd_tree, brute} | |
| n_neighbors | [2, 20] | |
| LGBMClassifier | n_estimators | [10, 1000] (log) |
| learning_rate | [0.01, 0.2] | |
| bagging_fraction | [0.5, 0.95] | |
| LinearSVC | max_iter | [10, 10,000] (log) |
| tol | [1 × 10, 0.1] (log) | |
| C | [0.01, 10] (log) | |
| LogisticRegression | penalty | {l1, l2} |
| solver | {liblinear, saga} | |
| MultinomialNB | alpha | [0.01, 10] (log) |
| fit_prior | {True, False} | |
| PassiveAggressiveClassifier | C | [0.01, 10] (log) |
| fit_intercept | {True, False} | |
| max_iter | [10, 1000] (log) | |
| RandomForestClassifier | n_estimators | [10, 1000] (log) |
| min_weight_fraction_leaf | [0.0, 0.5] | |
| max_features | {auto, sqrt, log2} | |
| RidgeClassifier | solver | {auto, svd, cholesky, lsqr, sparse_cg, sag, saga} |
| alpha | [0.001, 10] (log) | |
| SGDClassifier | penalty | {l2, l1, elasticnet} |
| alpha | [1 × 10, 1] (log) | |
| XGBClassifier | n_estimators | [10, 1000] (log) |
| learning_rate | [0.01, 0.2] | |
| gamma | [0.0, 0.4] | |
| Regression | ||
| Algorithm | Hyperparameter | Values |
| AdaBoostRegressor | n_estimators | [10, 1000] (log) |
| learning_rate | [0.1, 10] (log) | |
| BayesianRidge | n_liter | [10, 1000] (log) |
| alpha_1 | [1 × 10, 1 × 10] (log) | |
| lambda_1 | [1 × 10, 1 × 10] (log) | |
| tol | [1 × 10, 0.1] (log) | |
| DecisionTreeRegressor | max_depth | [2, 10] |
| min_impurity_decrease | [0.0, 0.5] | |
| criterion | {squared_error, friedman_mse, absolute_error} | |
| GradientBoostingRegressor | n_estimators | [10, 1000] (log) |
| learning_rate | [0.01, 0.3] | |
| subsample | [0.1, 1] | |
| KNeighborsRegressor | weights | {uniform, distance} |
| algorithm | {auto, ball_tree, kd_tree, brute} | |
| n_neighbors | [2, 20] | |
| KernelRidge | kernel | {linear, poly, rbf, sigmoid} |
| alpha | [0.1, 10] (log) | |
| gamma | [0.1, 10] (log) | |
| LGBMRegressor | lambda_l1 | [1 × 10, 10.0] (log) |
| lambda_l2 | [1 × 10, 10.0] (log) | |
| num_leaves | [2, 256] | |
| LinearRegression | fit_intercept | {True, False} |
| normalize | {True, False} | |
| LinearSVR | loss | {epsilon_insensitive, squared_epsilon_insensitive} |
| tol | [1 × 10, 0.1] (log) | |
| C | [0.01, 10] (log) | |
| PassiveAggressiveRegressor | C | [0.01, 10] (log) |
| fit_intercept | {True, False} | |
| max_iter | [10, 1000] (log) | |
| RandomForestRegressor | n_estimators | [10, 1000] (log) |
| min_weight_fraction_leaf | [0.0, 0.5] | |
| max_features | {auto, sqrt, log2} | |
| SGDRegressor | alpha | [1 × 10, 1] (log) |
| penalty | {l2, l1, elasticnet} | |
| XGBRegressor | n_estimators | [10, 1000] (log) |
| learning_rate | [0.01, 0.2] | |
| gamma | [0.0, 0.4] | |
| Classification | |||||||
|---|---|---|---|---|---|---|---|
| Algorithm | Acc | Bal | F1 | Reps | |||
| Median | Mean(std) | Median | Mean(std) | Median | Mean(std) | ||
| AdaBoostClassifier | 1.9 | 20.9 (65.3) | 2.2 | 21.5 (57.4) | 1.9 | 39.3 (150.1) | 4320 |
| DecisionTreeClassifier | 0.0 | 115.6 (2.3 × 103) | 0.0 | 96.0 (2.2 × 103) | 0.0 | 55.7 (1.3 × 103) | 4220 |
| GradientBoostingClassifier | 0.5 | 45.1 (1.4 × 103) | 0.6 | 48.9 (1.4 × 103) | 0.6 | 42.0 (1.1 × 103) | 4096 |
| KNeighborsClassifier | 0.8 | 3.8 (13.8) | 1.8 | 5.9 (16.7) | 1.5 | 5.1 (17.7) | 4254 |
| LGBMClassifier | 0.0 | 1.2 (11.5) | 0.0 | 1.0 (11.9) | 0.0 | 0.9 (12.1) | 4287 |
| LinearSVC | 0.0 | 1.0 (8.3) | 0.0 | 1.9 (8.5) | 0.0 | 1.7 (8.2) | 4299 |
| LogisticRegression | 0.0 | 1.5 (8.2) | 0.0 | 3.4 (12.5) | 0.0 | 3.4 (11.8) | 4307 |
| MultinomialNB | 0.0 | 9.8 (58.9) | 8.5 | 27.5 (48.9) | 10.5 | 40.5 (128.6) | 4149 |
| PassiveAggressiveClassifier | 1.9 | 7.8 (24.0) | 1.8 | 5.9 (18.6) | 3.0 | 10.8 (28.8) | 4301 |
| RandomForestClassifier | 0.0 | 153.6 (2.3 × 103) | 0.0 | 218.8 (3.0 × 103) | 0.0 | 134.1 (2.0 × 103) | 4320 |
| RidgeClassifier | 0.0 | 1.0 (6.8) | 0.0 | 1.4 (7.3) | 0.0 | 1.9 (7.9) | 4273 |
| SGDClassifier | 1.2 | 5.0 (20.6) | 1.6 | 5.2 (16.7) | 2.0 | 8.6 (26.7) | 4212 |
| XGBClassifier | 0.0 | 13.5 (643.1) | 0.0 | 11.4 (431.2) | 0.0 | 10.1 (467.6) | 4111 |
| Regression | |||||||
| Algorithm | R2 | Adj | C-RMSE | Reps | |||
| median | mean(std) | median | mean(std) | median | mean(std) | ||
| AdaBoostRegressor | 2.0 | 3.6 (33.5) | 2.1 | −741.3 (9.5 × 103) | 3.8 | 5.1 (20.9) | 3179 |
| BayesianRidge | 0.0 | 6.8 × 103 (3.7 × 105) | −0.0 | −3.3 (55.3) | 0.0 | 1.0 (9.8) | 3117 |
| DecisionTreeRegressor | 3.8 | 61.5 (788.0) | 4.0 | 49.1 (841.5) | 7.0 | 63.4 (1.3 × 103) | 3150 |
| GradientBoostingRegressor | 1.6 | 17.3 (430.9) | 1.7 | −6.6 × 105 (2.4 × 107) | 4.1 | 2.3 (126.4) | 3180 |
| KNeighborsRegressor | 3.5 | 77.8 (627.4) | 3.5 | 18.8 (471.6) | 4.5 | 203.5 (5.3 × 103) | 3160 |
| KernelRidge | 69.5 | −9.3 × 105 (5.0 × 107) | 65.9 | 3.6 × 103 (1.7 × 105) | 49.5 | 1.7 × 103 (8.1 × 104) | 3053 |
| LGBMRegressor | 0.0 | 0.0 (25.6) | 0.0 | −1.2 (34.4) | 0.0 | 0.4 (2.1) | 3179 |
| LinearRegression | 0.0 | 2.3 (70.4) | 0.0 | −35.1 (469.4) | 0.0 | −1.7 (62.8) | 3170 |
| LinearSVR | 25.1 | 86.4 (2.7 × 103) | 24.3 | 173.5 (2.8 × 103) | 23.9 | 159.7 (2.2 × 103) | 3161 |
| PassiveAggressiveRegressor | 71.6 | 180.7 (1.7 × 103) | 58.5 | −304.3 (4.1 × 103) | 62.0 | 331.9 (5.5 × 103) | 3167 |
| RandomForestRegressor | −0.1 | 1.5 (44.2) | −0.2 | −1.2 × 103 (4.6 × 104) | −0.5 | −1.5 (13.2) | 3180 |
| SGDRegressor | 0.0 | 2.6 (68.6) | 0.0 | −41.4 (2.0 × 103) | 0.0 | 2.2 (39.8) | 3167 |
| XGBRegressor | 0.9 | 20.0 (717.1) | 0.8 | −675.6 (7.4 × 103) | 2.3 | 6.8 (164.6) | 3180 |
| Classification | Regression | ||
|---|---|---|---|
| RandomForestClassifier | 3.89 | KernelRidge | 2110.75 |
| DecisionTreeClassifier | 2.43 | GradientBoostingRegressor | 183.97 |
| GradientBoostingClassifier | 1.52 | BayesianRidge | 35.19 |
| MultinomialNB | 1.36 | PassiveAggressiveRegressor | 5.34 |
| AdaBoostClassifier | 0.79 | LinearSVR | 2.13 |
| PassiveAggressiveClassifier | 0.56 | KNeighborsRegressor | 0.59 |
| XGBClassifier | 0.38 | DecisionTreeRegressor | 0.35 |
| SGDClassifier | 0.35 | RandomForestRegressor | 0.09 |
| KNeighborsClassifier | 0.27 | AdaBoostRegressor | −0.07 |
| LogisticRegression | −0.08 | XGBRegressor | −0.10 |
| LinearSVC | −0.09 | SGDRegressor | −0.23 |
| RidgeClassifier | −0.10 | LinearRegression | −0.25 |
| LGBMClassifier | −0.10 | LGBMRegressor | −0.26 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sipper, M. High Per Parameter: A Large-Scale Study of Hyperparameter Tuning for Machine Learning Algorithms. Algorithms 2022, 15, 315. https://doi.org/10.3390/a15090315
Sipper M. High Per Parameter: A Large-Scale Study of Hyperparameter Tuning for Machine Learning Algorithms. Algorithms. 2022; 15(9):315. https://doi.org/10.3390/a15090315
Chicago/Turabian StyleSipper, Moshe. 2022. "High Per Parameter: A Large-Scale Study of Hyperparameter Tuning for Machine Learning Algorithms" Algorithms 15, no. 9: 315. https://doi.org/10.3390/a15090315
APA StyleSipper, M. (2022). High Per Parameter: A Large-Scale Study of Hyperparameter Tuning for Machine Learning Algorithms. Algorithms, 15(9), 315. https://doi.org/10.3390/a15090315






