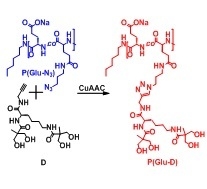
Synthesis of Dendronized Poly(l-Glutamate) via Azide-Alkyne Click Chemistry
Abstract
:1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Methods
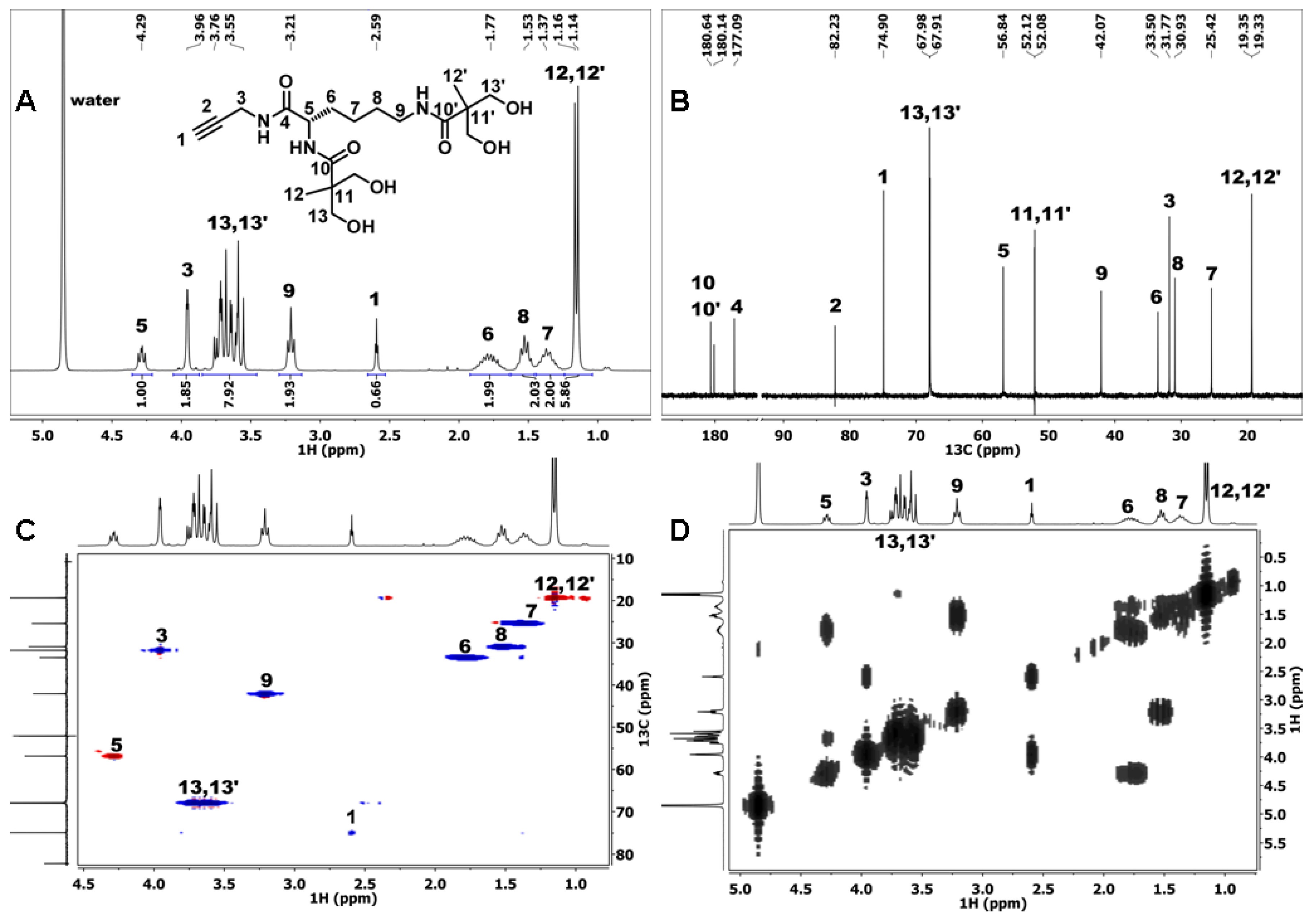
2.2.1. NMR
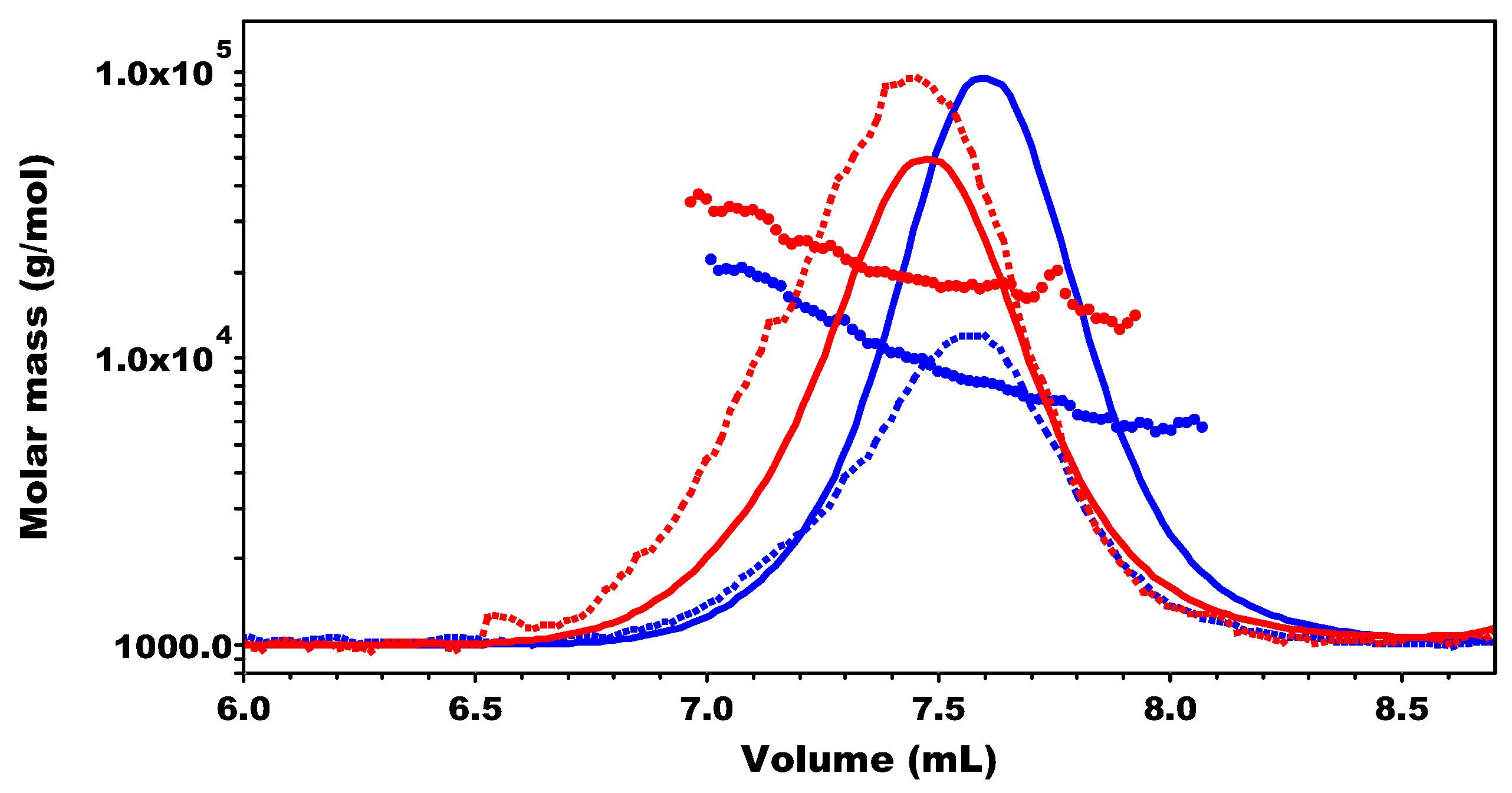
2.2.2. SEC-MALS
2.2.3. MALDI-TOF MS
2.2.4. FT-IR
3. Results and Discussion
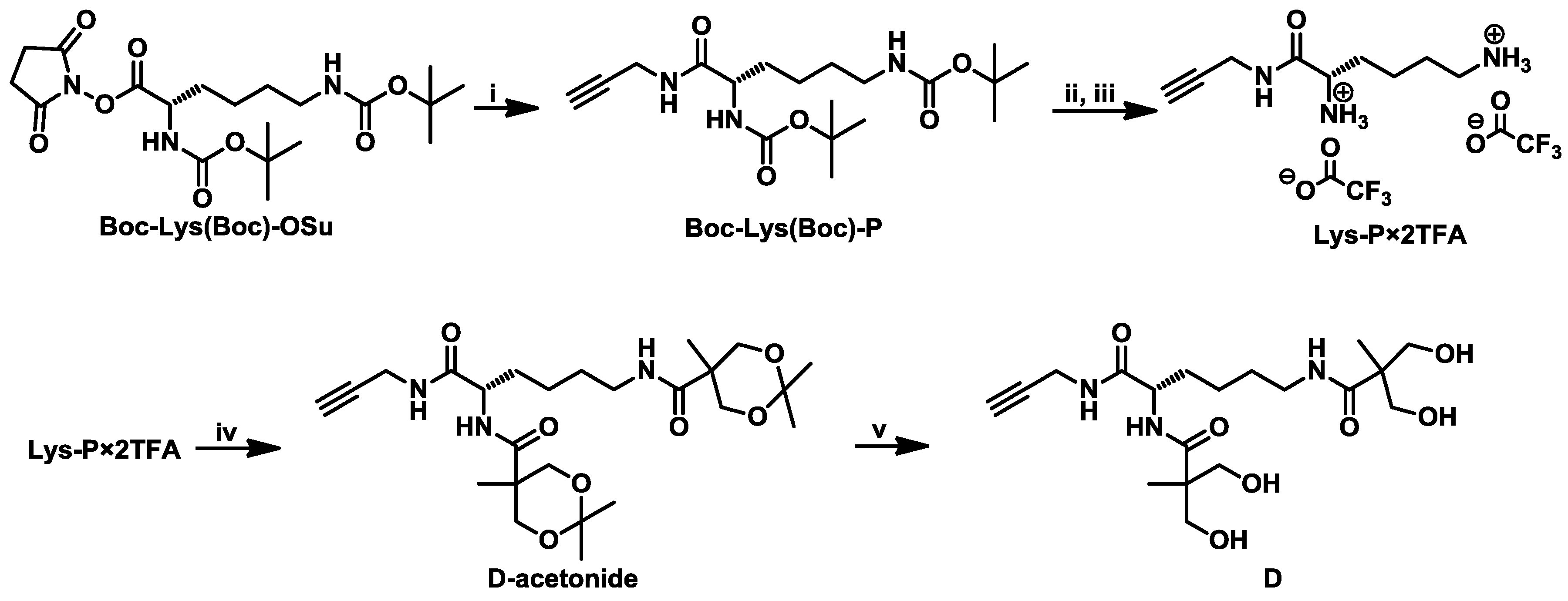
3.1. Synthesis of the Second-Generation Dendron (D) with Acetylene Focal Group
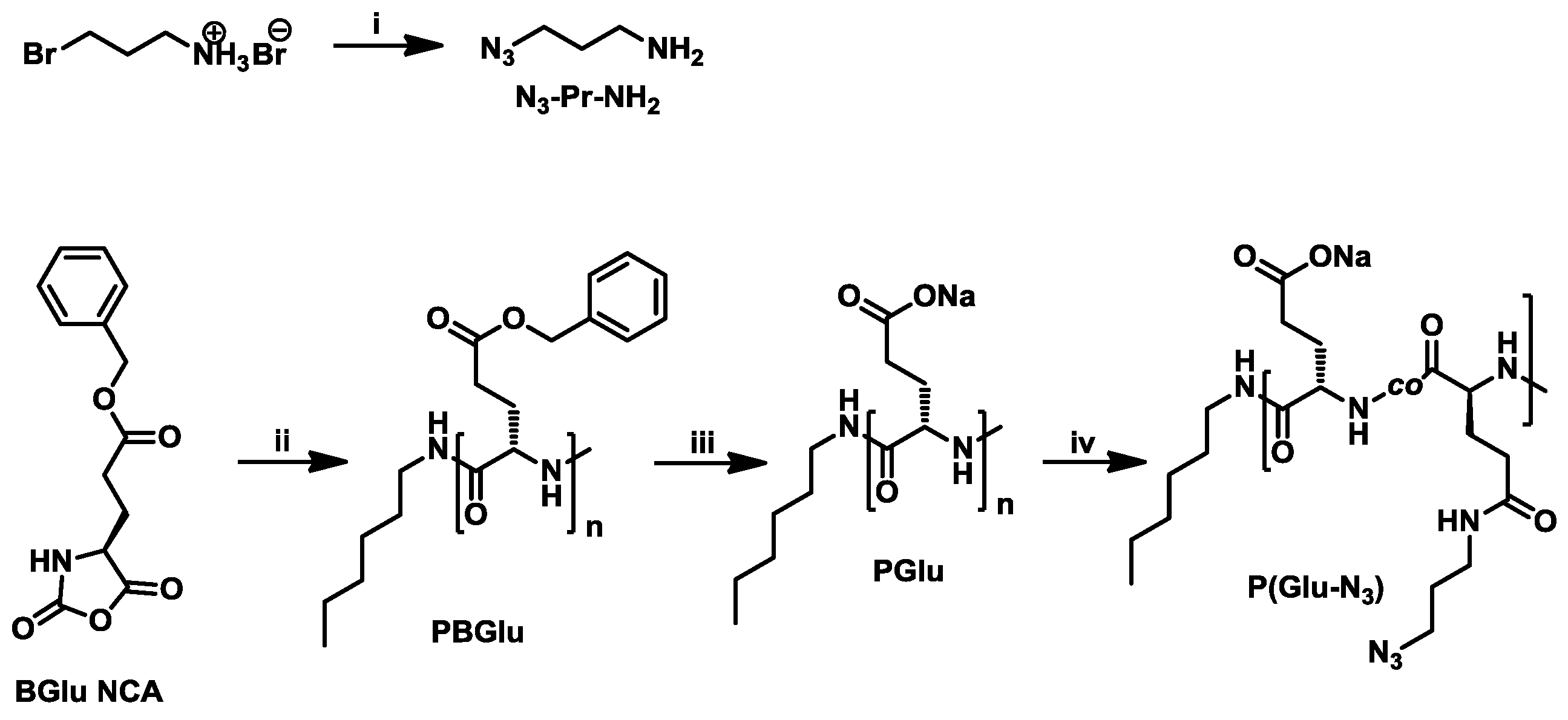
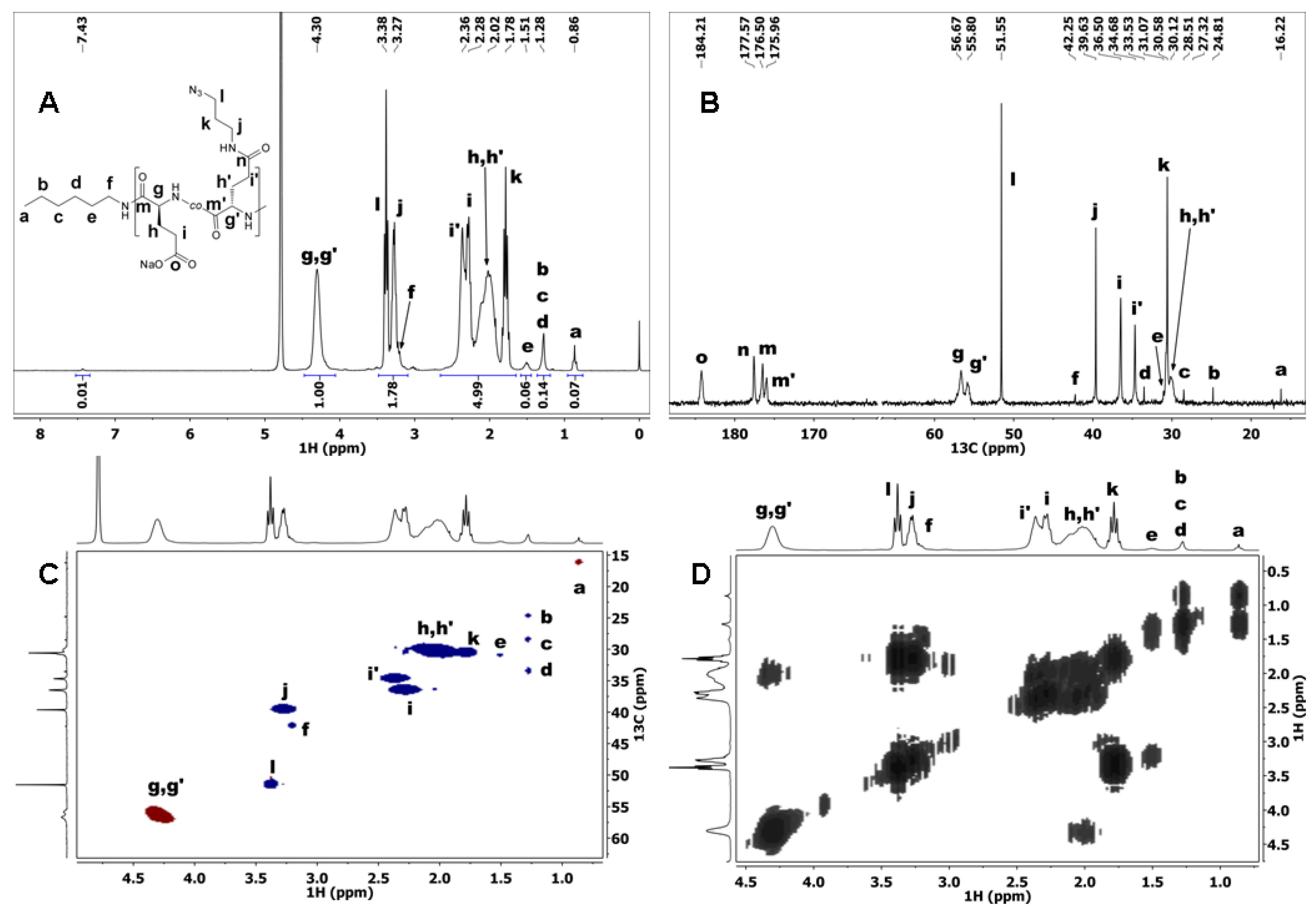
3.2. Synthesis of Poly[l-Glutamate-co-(N-3-Azidopropyl)-l-Glutamine] (P(Glu-N3))
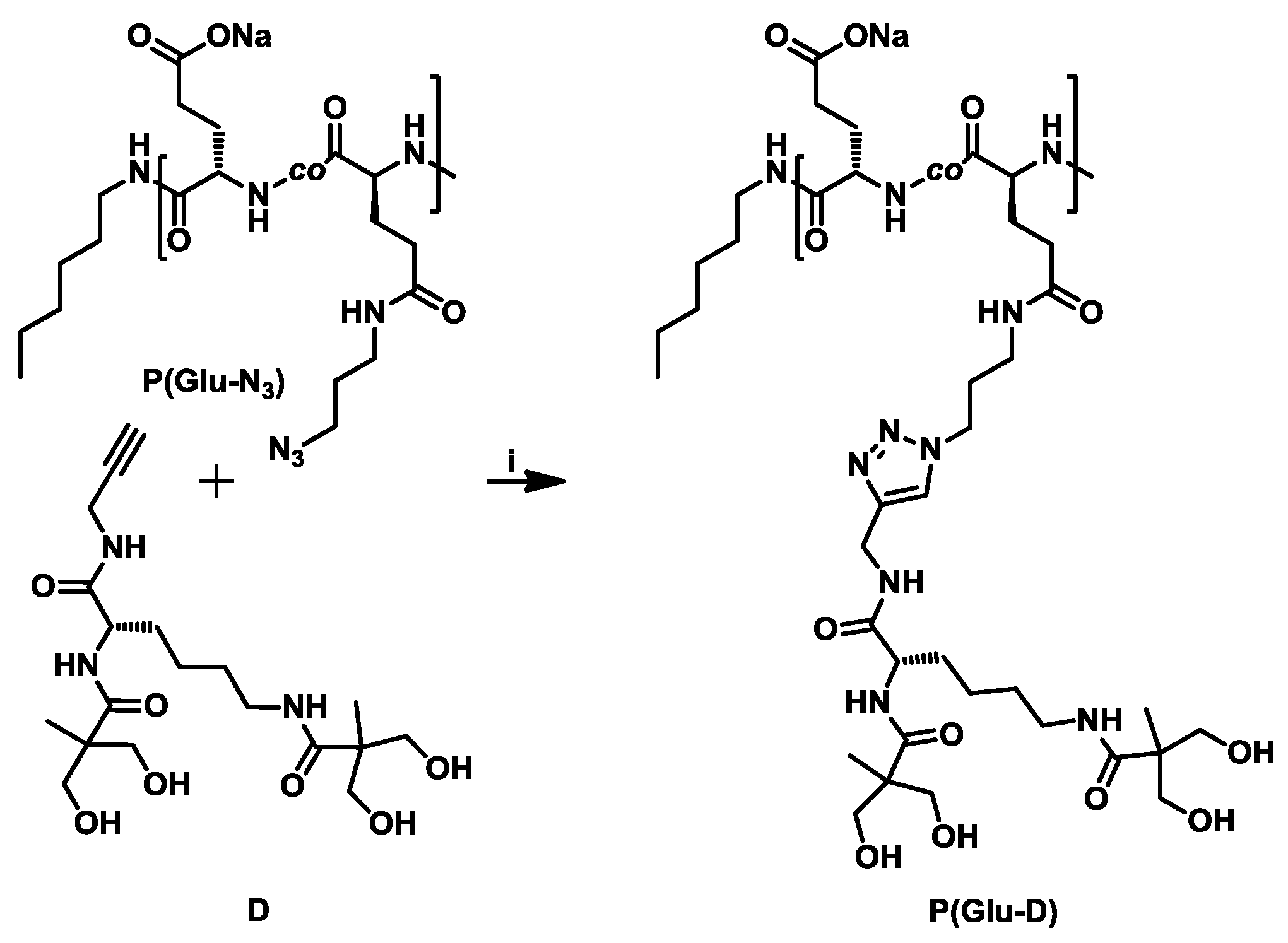
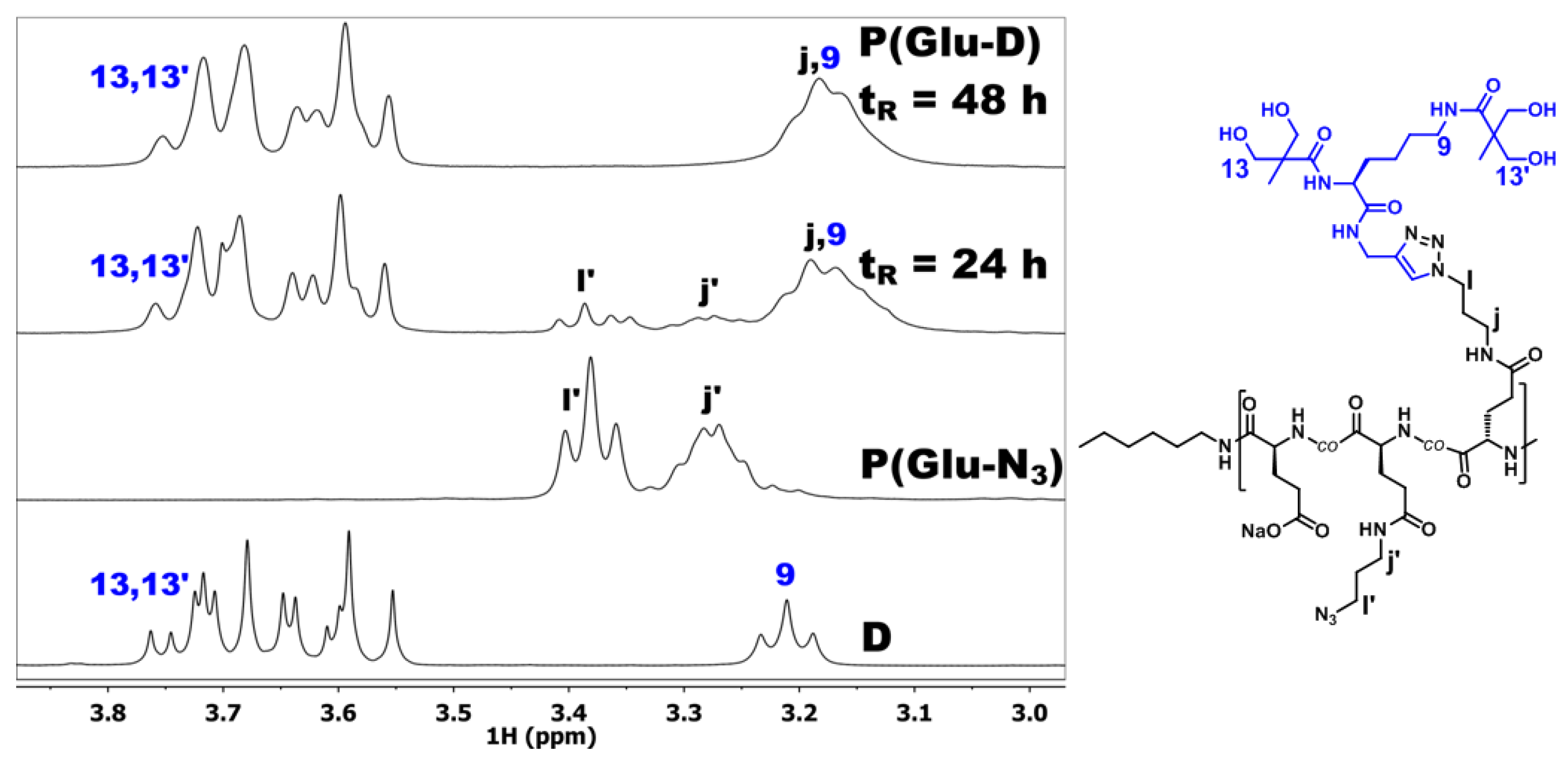
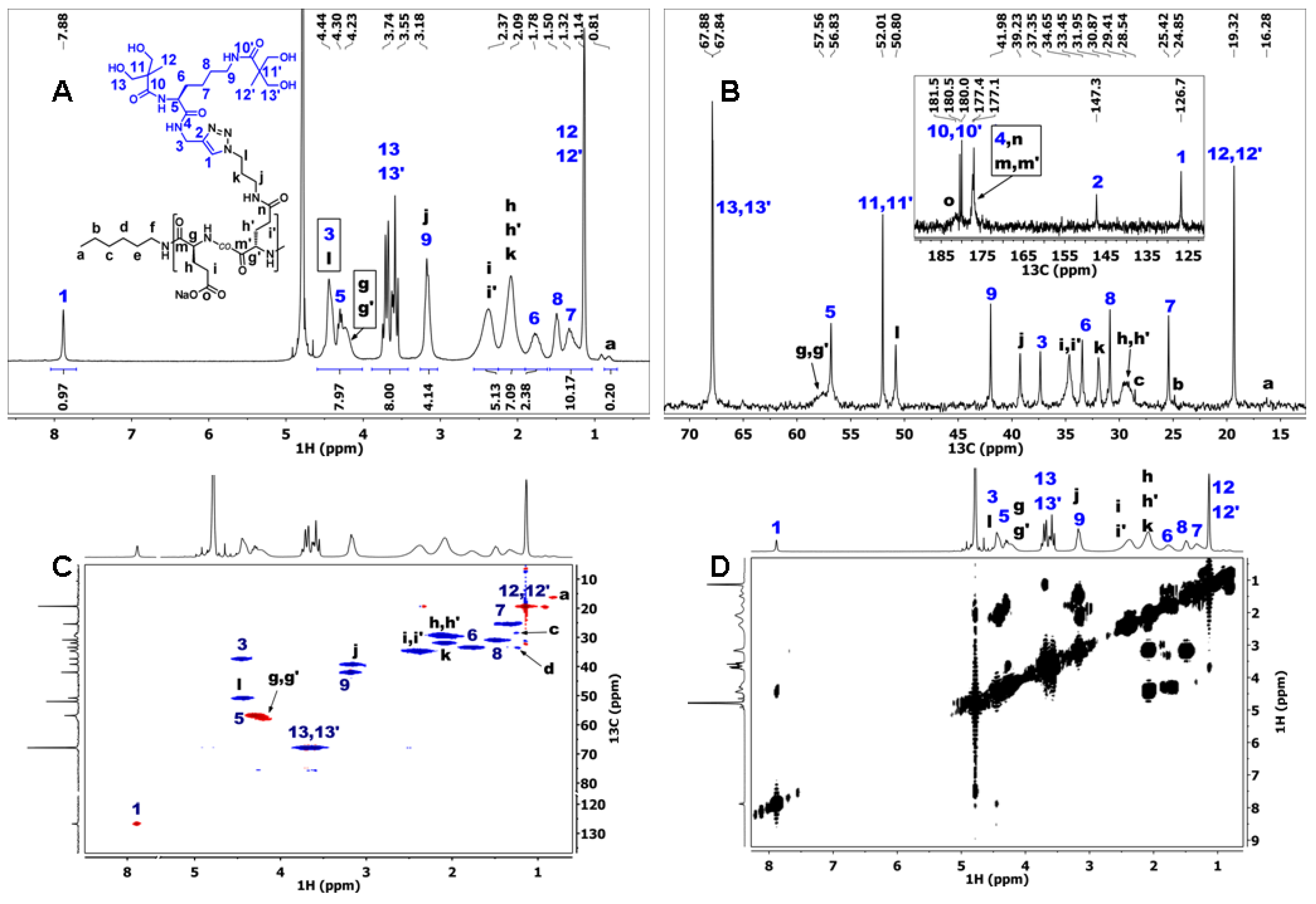
3.3. Synthesis of Dendronized Poly(l-Glutamate) (P(Glu-D))
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Lübbert, A.; Nguyen, T.Q.; Sun, F.; Sheiko, S.S.; Klok, H.A. l-lysine dendronized polystyrene. Macromolecules 2005, 38, 2064–2071. [Google Scholar] [CrossRef]
- Chen, Y.; Xiong, X. Tailoring dendronized polymers. Chem. Commun. 2010, 46, 5049–5060. [Google Scholar] [CrossRef] [PubMed]
- Frauenrath, H. Dendronized polymers—Building a new bridge from molecules to nanoscopic objects. Prog. Polym. Sci. 2005, 30, 325–384. [Google Scholar] [CrossRef]
- Schlüter, A.D. Dendrimers with polymeric core: Towards nanocylinders. In Dendrimers; Springer: Berlin/Heidelberg, Germany, 1998; pp. 165–191. [Google Scholar]
- Lee, C.C.; Fréchet, J.M.J. Synthesis and conformations of dendronized poly(l-lysine). Macromolecules 2006, 39, 476–481. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.; van Beek, J.D.; Zhang, B.; Colussi, M.; Walde, P.; Zhang, A.; Kröger, M.; Halperin, A.; Schlüter, A.D. Tuning polymer thickness: Synthesis and scaling theory of homologous series of dendronized polymers. J. Am. Chem. Soc. 2009, 131, 11841–11854. [Google Scholar] [CrossRef] [PubMed]
- Bertran, O.; Zhang, B.; Schlüter, A.D.; Halperin, A.; Kröger, M.; Alemán, C. Computer simulation of dendronized polymers: Organization and characterization at the atomistic level. RSC Adv. 2012, 3, 126–140. [Google Scholar] [CrossRef]
- Bertran, O.; Zhang, B.; Schlüter, A.D.; Kröger, M.; Alemán, C. Computer simulation of fifth generation dendronized polymers: Impact of charge on internal organization. J. Phys. Chem. B 2013, 117, 6007–6017. [Google Scholar] [CrossRef] [PubMed]
- Gössl, I.; Shu, L.; Schlüter, A.D.; Rabe, J.P. Molecular structure of single DNA complexes with positively charged dendronized polymers. J. Am. Chem. Soc. 2002, 124, 6860–6865. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.C.; Yoshida, M.; Fréchet, J.M.J.; Dy, E.E.; Szoka, F.C. In vitro and in vivo evaluation of hydrophilic dendronized linear polymers. Bioconjug. Chem. 2005, 16, 535–541. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.; Schade, B.; Kumar, S.; Böttcher, C.; Sharma, S.K.; Haag, R. Non-ionic dendronized multiamphiphilic polymers as nanocarriers for biomedical applications. Small 2013, 9, 894–904. [Google Scholar] [CrossRef] [PubMed]
- Kumari, M.; Singh, A.K.; Kumar, S.; Achazi, K.; Gupta, S.; Haag, R.; Sharma, S.K. Synthesis of amphiphilic dendronized polymers to study their self-assembly and transport behavior. Polym. Adv. Technol. 2014, 25, 1208–1215. [Google Scholar] [CrossRef]
- Ossenbach, A.; Rüegger, H.; Zhang, A.; Fischer, K.; Schlüter, A.D.; Schmidt, M. Ion-induced stretching of low generation dendronized polymers with crown ether branching units. Macromolecules 2009, 42, 8781–8793. [Google Scholar] [CrossRef]
- Qin, C.; Wu, X.; Tong, H.; Wang, L. High solubility and photoluminescence quantum yield water-soluble polyfluorenes with dendronized amino acid side chains: Synthesis, photophysical, and metal ion sensing properties. J. Mater. Chem. 2010, 20, 7957–7964. [Google Scholar] [CrossRef]
- Roeser, J.; Heinrich, B.; Bourgogne, C.; Rawiso, M.; Michel, S.; Hubscher-Bruder, V.; Arnaud-Neu, F.; Méry, S. Dendronized polymers with silver and mercury cations recognition: Complexation studies and polyelectrolyte behavior. Macromolecules 2013, 46, 7075–7085. [Google Scholar] [CrossRef]
- Grotzky, A.; Nauser, T.; Erdogan, H.; Schlüter, A.D.; Walde, P. A fluorescently labeled dendronized polymer-enzyme conjugate carrying multiple copies of two different types of active enzymes. J. Am. Chem. Soc. 2012, 134, 11392–11395. [Google Scholar] [CrossRef] [PubMed]
- Laurino, P.; Kikkeri, R.; Azzouz, N.; Seeberger, P.H. Detection of bacteria using glyco-dendronized polylysine prepared by continuous flow photofunctionalization. Nano Lett. 2011, 11, 73–78. [Google Scholar] [CrossRef] [PubMed]
- Yan, J.; Li, W.; Zhang, A. Dendronized supramolecular polymers. Chem. Commun. 2014, 50, 12221–12233. [Google Scholar] [CrossRef] [PubMed]
- Schlüter, A.D.; Rabe, J.P. Dendronized polymers: Synthesis, characterization, assembly at interfaces, and manipulation. Angew. Chem. Int. Ed. 2000, 39, 864–883. [Google Scholar] [CrossRef]
- Golas, P.L.; Matyjaszewski, K. Marrying click chemistry with polymerization: Expanding the scope of polymeric materials. Chem. Soc. Rev. 2010, 39, 1338–1354. [Google Scholar] [CrossRef] [PubMed]
- Tasdelen, M.A. Diels-Alder “click” reactions: Recent applications in polymer and material science. Polym. Chem. 2011, 2, 2133–2145. [Google Scholar] [CrossRef]
- Lowe, A.B. Thiol-ene “click” reactions and recent applications in polymer and materials synthesis: A first update. Polym. Chem. 2014, 5, 4820–4870. [Google Scholar] [CrossRef]
- Helms, B.; Mynar, J.L.; Hawker, C.J.; Fréchet, J.M.J. Dendronized linear polymers via “click chemistry”. J. Am. Chem. Soc. 2004, 126, 15020–15021. [Google Scholar] [CrossRef] [PubMed]
- Tonga, M.; Yesilbag Tonga, G.; Seber, G.; Gok, O.; Sanyal, A. Dendronized polystyrene via orthogonal double-click reactions. J. Polym. Sci. A Polym. Chem. 2013, 51, 5029–5037. [Google Scholar] [CrossRef]
- Zhang, X.; Li, W.; Zhao, X.; Zhang, A. Thermoresponsive dendronized polyprolines via the “grafting to” route. Macromol. Rapid Commun. 2013, 34, 1701–1707. [Google Scholar] [CrossRef] [PubMed]
- Xiong, X.; Chen, Y. Clickable dendronized copolymers for introducing structural heterogeneity. Eur. Polym. J. 2012, 48, 569–579. [Google Scholar] [CrossRef]
- Tonga, M.; Cengiz, N.; Kose, M.M.; Dede, T.; Sanyal, A. Dendronized polymers via Diels-Alder “click” reaction. J. Polym. Sci. A Polym. Chem. 2010, 48, 410–416. [Google Scholar] [CrossRef]
- Walter, M.V.; Lundberg, P.; Hult, A.; Malkoch, M. Novel macrothiols for the synthesis of a structurally comprehensive dendritic library using thiol-ene click chemistry. J. Polym. Sci. A Polym. Chem. 2011, 49, 2990–2995. [Google Scholar] [CrossRef]
- Polaske, N.W.; McGrath, D.V.; McElhanon, J.R. Thermally reversible dendronized linear AB step-polymers via “click” chemistry. Macromolecules 2011, 44, 3203–3210. [Google Scholar] [CrossRef]
- Polaske, N.W.; McGrath, D.V.; McElhanon, J.R. Thermally reversible dendronized step-polymers based on sequential huisgen 1,3-dipolar cycloaddition and Diels-Alder “click” reactions. Macromolecules 2010, 43, 1270–1276. [Google Scholar] [CrossRef]
- Shu, L.; Schäfer, A.; Schlüter, A.D. Dendronized polymers: Increasing of dendron generation by the attach-to approach. Macromolecules 2000, 33, 4321–4328. [Google Scholar] [CrossRef]
- Lee, C.C.; Grayson, S.M.; Fréchet, J.M.J. Synthesis of narrow-polydispersity degradable dendronized aliphatic polyesters. J. Polym. Sci. A Polym. Chem. 2004, 42, 3563–3578. [Google Scholar] [CrossRef]
- Malkoch, M.; Carlmark, A.; Woldegiorgis, A.; Hult, A.; Malmström, E.E. Dendronized aliphatic polymers by a combination of ATRP and divergent growth. Macromolecules 2004, 37, 322–329. [Google Scholar] [CrossRef]
- Yu, H.; Schlüter, A.D.; Zhang, B. Synthesis of dendronized polymers by a “n + 2” approach. Macromolecules 2012, 45, 8555–8560. [Google Scholar] [CrossRef]
- Yu, H.; Schlüter, A.D.; Zhang, B. Synthesis of high generation dendronized polymers and quantification of their structure perfection. Macromolecules 2014, 47, 4127–4135. [Google Scholar] [CrossRef]
- Carlmark, A.; Hawker, C.; Hult, A.; Malkoch, M. New methodologies in the construction of dendritic materials. Chem. Soc. Rev. 2009, 38, 352–362. [Google Scholar] [CrossRef] [PubMed]
- Carlmark, A.; Malmström, E.; Malkoch, M. Dendritic architectures based on bis-MPA: Functional polymeric scaffolds for application-driven research. Chem. Soc. Rev. 2013, 42, 5858–5879. [Google Scholar] [CrossRef] [PubMed]
- Ohsaki, M.; Okuda, T.; Wada, A.; Hirayama, T.; Niidome, T.; Aoyagi, H. In vitro gene transfection using dendritic poly(l-lysine). Bioconjug. Chem. 2002, 13, 510–517. [Google Scholar] [CrossRef] [PubMed]
- Okuda, T.; Kidoaki, S.; Ohsaki, M.; Koyama, Y.; Yoshikawa, K.; Niidome, T.; Aoyagi, H. Time-dependent complex formation of dendritic poly(l-lysine) with plasmid DNA and correlation with in vitro transfection efficiencies. Org. Biomol. Chem. 2003, 1, 1270–1273. [Google Scholar] [CrossRef] [PubMed]
- Kadlecova, Z.; Baldi, L.; Hacker, D.; Wurm, F.M.; Klok, H.A. Comparative study on the in vitro cytotoxicity of linear, dendritic, and hyperbranched polylysine analogues. Biomacromolecules 2012, 13, 3127–3137. [Google Scholar] [CrossRef] [PubMed]
- Chapman, T.M.; Hillyer, G.L.; Mahan, E.J.; Shaffer, K.A. Hydraamphiphiles: Novel linear dendritic block copolymer surfactants. J. Am. Chem. Soc. 1994, 116, 11195–11196. [Google Scholar] [CrossRef]
- Choi, J.S.; Lee, E.J.; Choi, Y.H.; Jeong, Y.J.; Park, J.S. Poly(ethylene glycol)-block-poly(l-lysine) dendrimer: novel linear polymer/dendrimer block copolymer forming a spherical water-soluble polyionic complex with DNA. Bioconjug. Chem. 1999, 10, 62–65. [Google Scholar] [CrossRef] [PubMed]
- Choi, J.S.; Joo, D.K.; Kim, C.H.; Kim, K.; Park, J.S. Synthesis of a barbell-like triblock copolymer, poly(l-lysine) dendrimer-block-poly(ethylene glycol)-block-poly(l-lysine) dendrimer, and its self-assembly with plasmid DNA. J. Am. Chem. Soc. 2000, 122, 474–480. [Google Scholar] [CrossRef]
- Javakhishvili, I.; Binder, W.H.; Tanner, S.; Hvilsted, S. Facile synthesis of linear-dendritic cholesteryl-poly(ε-caprolactone)-b-(l-lysine)G2 by thiol-ene and azide-alkyne “click” reactions. Polym. Chem. 2010, 1, 506–513. [Google Scholar] [CrossRef]
- Aharoni, S.M.; Crosby, C.R.; Walsh, E.K. Size and solution properties of globular tert-butyloxycarbonyl-poly(α,iε-l-lysine). Macromolecules 1982, 15, 1093–1098. [Google Scholar] [CrossRef]
- Sakthivel, T.; Toth, I.; Florence, A.T. Synthesis and physicochemical properties of lipophilic polyamide dendrimers. Pharm. Res. 1998, 15, 776–782. [Google Scholar] [CrossRef] [PubMed]
- Shah, D.S.; Sakthivel, T.; Toth, I.; Florence, A.T.; Wilderspin, A.F. DNA transfection and transfected cell viability using amphipathic asymmetric dendrimers. Int. J. Pharm. 2000, 208, 41–48. [Google Scholar] [CrossRef]
- Kaneshiro, T.L.; Wang, X.; Lu, Z.R. Synthesis, characterization, and gene delivery of poly-l-lysine octa(3-aminopropyl)silsesquioxane dendrimers: Nanoglobular drug carriers with precisely defined molecular architectures. Mol. Pharm. 2007, 4, 759–768. [Google Scholar] [CrossRef] [PubMed]
- Luo, K.; Li, C.; Wang, G.; Nie, Y.; He, B.; Wu, Y.; Gu, Z. Peptide dendrimers as efficient and biocompatible gene delivery vectors: Synthesis and in vitro characterization. J. Control. Release 2011, 155, 77–87. [Google Scholar] [CrossRef] [PubMed]
- Liu, K.; Zhang, X.; Tao, X.; Yan, J.; Kuang, G.; Li, W.; Zhang, A. Lysine-based dendronized polymers with preferred chirality. Polym. Chem. 2012, 3, 2708–2711. [Google Scholar] [CrossRef]
- Zhang, A. High-molar-mass, first and second generation l-lysine dendronized polymethacrylates. Macromol. Rapid Commun. 2008, 29, 839–845. [Google Scholar] [CrossRef]
- Zhuravel, M.A.; Davis, N.E.; Nguyen, S.T.; Koltover, I. Dendronized protein polymers: Synthesis and self-assembly of monodisperse cylindrical macromolecules. J. Am. Chem. Soc. 2004, 126, 9882–9883. [Google Scholar] [CrossRef] [PubMed]
- Zeng, H.; Little, H.C.; Tiambeng, T.N.; Williams, G.A.; Guan, Z. Multifunctional dendronized peptide polymer platform for safe and effective siRNA delivery. J. Am. Chem. Soc. 2013, 135, 4962–4965. [Google Scholar] [CrossRef] [PubMed]
- Liu, R.; Sanda, F.; Masuda, T. Synthesis and properties of ornithine- and lysine-based poly(N-propargylamides). Responsiveness of the helical structure to acids. Polymer 2007, 48, 6510–6518. [Google Scholar] [CrossRef]
- Sanda, F.; Terada, K.; Masuda, T. Synthesis, chiroptical properties, and pH responsibility of aspartic acid- and glutamic acid-based helical polyacetylenes. Macromolecules 2005, 38, 8149–8154. [Google Scholar] [CrossRef]
- Gao, G.; Sanda, F.; Masuda, T. Synthesis and properties of amino acid-based polyacetylenes. Macromolecules 2003, 36, 3932–3927. [Google Scholar] [CrossRef]
- Sanda, F.; Araki, H.; Masuda, T. Synthesis and properties of serine- and threonine-based helical polyacetylenes. Macromolecules 2004, 37, 8510–8516. [Google Scholar] [CrossRef]
- Brosch, O.; Weyhermüller, T.; Metzler-Nolte, N. Synthesis of alkynyl amino acids and peptides and their palladium-catalyzed coupling to ferrocene. Inorg. Chem. 1999, 38, 5308–5313. [Google Scholar] [CrossRef]
- Ihre, H.; Hult, A.; Fréchet, J.M.J.; Gitsov, I. Double-stage convergent approach for the synthesis of functionalized dendritic aliphatic polyesters based on 2,2-bis(hydroxymethyl)propionic acid. Macromolecules 1998, 31, 4061–4068. [Google Scholar] [CrossRef]
- Claridge, T.D.W. High-Resolution NMR Techniques in Organic Chemistry; Newnes: Philadelphia, PA, USA, 2009. [Google Scholar]
- Carboni, B.; Benalil, A.; Vaultier, M. Aliphatic amino azides as key building blocks for efficient polyamine syntheses. J. Org. Chem. 1993, 58, 3736–3741. [Google Scholar] [CrossRef]
- Agut, W.; Agnaou, R.; Lecommandoux, S.; Taton, D. Synthesis of block copolypeptides by click chemistry. Macromol. Rapid Commun. 2008, 29, 1147–1155. [Google Scholar] [CrossRef]
- Kato, Y.; Umemoto, N.; Kayama, Y.; Fukushima, H.; Takeda, Y.; Hara, T.; Tsukada, Y. A novel method of conjugation of daunomycin with antibody with a poly(l-glutamic acid) derivative as intermediate drug carrier. An anti-alpha-fetoprotein antibody-daunomycin conjugate. J. Med. Chem. 1984, 27, 1602–1607. [Google Scholar] [CrossRef] [PubMed]
- Krannig, K.S.; Sun, J.; Schlaad, H. Stimuli-responsivity of secondary structures of glycopolypeptides derived from poly(l-glutamate-co-allylglycine). Biomacromolecules 2014, 15, 978–984. [Google Scholar] [CrossRef] [PubMed]
- Jackson, C.A.; Simonsick, W.J., Jr. Application of mass spectrometry to the characterization of polymers. Curr. Opin. Solid State Mater. Sci. 1997, 2, 661–667. [Google Scholar] [CrossRef]
- Montaudo, G.; Samperi, F.; Montaudo, M.S. Characterization of synthetic polymers by MALDI-MS. Prog. Polym. Sci. 2006, 31, 277–357. [Google Scholar] [CrossRef]
- Weidner, S.M.; Trimpin, S. Mass spectrometry of synthetic polymers. Anal. Chem. 2008, 80, 4349–4361. [Google Scholar] [CrossRef] [PubMed]

| Sample | Calculated | SEC-MALS | ||
|---|---|---|---|---|
| Mn(theor.) a (104 Da) | Mn (104 Da) | Mw (104 Da) | ĐM | |
| P(Glu-N3) | - | 0.823 | 0.905 | 1.1 |
| P(Glu-D) | 1.65 | 1.67 | 2.00 | 1.2 |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Perdih, P.; Kržan, A.; Žagar, E. Synthesis of Dendronized Poly(l-Glutamate) via Azide-Alkyne Click Chemistry. Materials 2016, 9, 242. https://doi.org/10.3390/ma9040242
Perdih P, Kržan A, Žagar E. Synthesis of Dendronized Poly(l-Glutamate) via Azide-Alkyne Click Chemistry. Materials. 2016; 9(4):242. https://doi.org/10.3390/ma9040242
Chicago/Turabian StylePerdih, Peter, Andrej Kržan, and Ema Žagar. 2016. "Synthesis of Dendronized Poly(l-Glutamate) via Azide-Alkyne Click Chemistry" Materials 9, no. 4: 242. https://doi.org/10.3390/ma9040242
APA StylePerdih, P., Kržan, A., & Žagar, E. (2016). Synthesis of Dendronized Poly(l-Glutamate) via Azide-Alkyne Click Chemistry. Materials, 9(4), 242. https://doi.org/10.3390/ma9040242