Biocompatibility of ZrO2 vs. Y-TZP Alloys: Influence of Their Composition and Surface Topography
Abstract
:1. Introduction
2. Material and Method
2.1. Preparation of ZrO2 and Y-TZP Discs
2.1.1. Polishing Surfaces
2.1.2. Surface Characterization with Tactile Profilometer
- i.
- Measurement length: 1000 μm;
- ii.
- Long cut-off: 0.8 μm; short cut-off: 0.08 µm.
2.1.3. Scanning Electron Microscopy of Discs
2.2. Cell Culture
2.3. Cell Proliferation
2.4. Scanning Electron Microscopy of Saos-2 Cells Exposed to Discs
2.5. Total RNA Extraction and Purification
2.6. Microarray Hybridization
2.7. Statistical and Functional Analyses of Microarray Data
3. Results
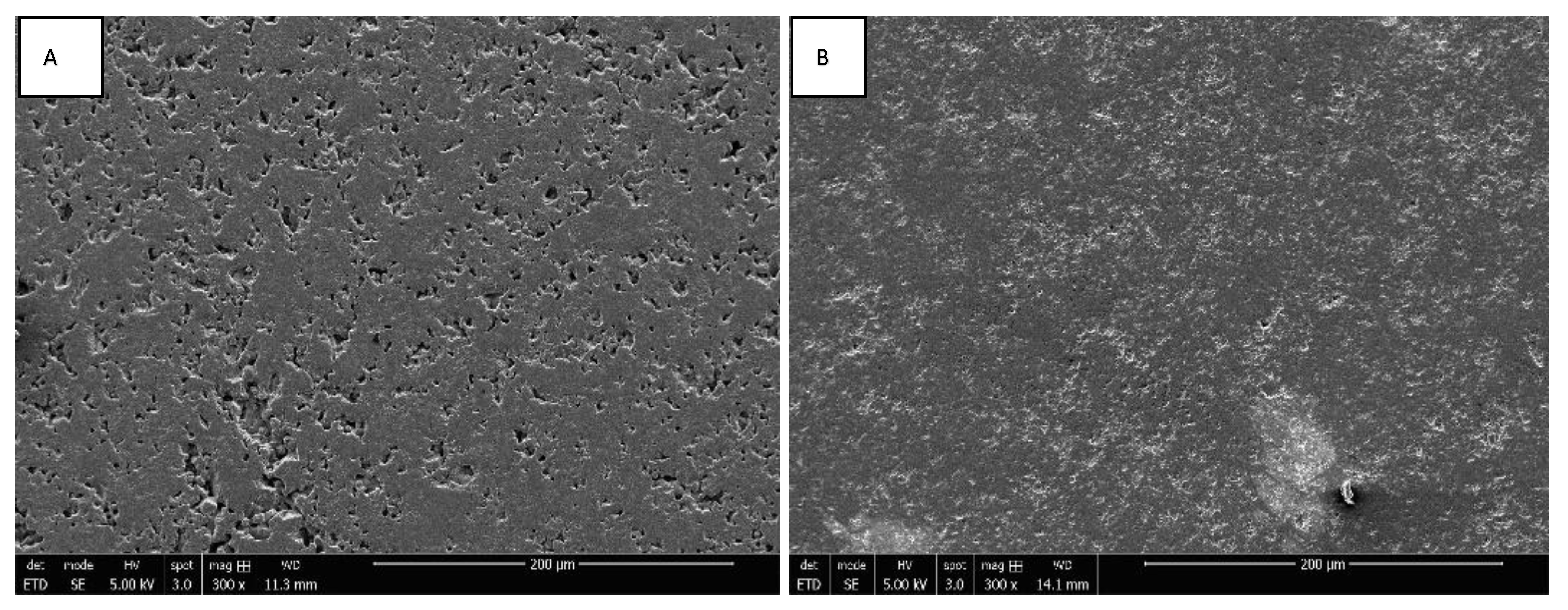
3.1. Morphology of ZrO2 and Y-TZP Discs
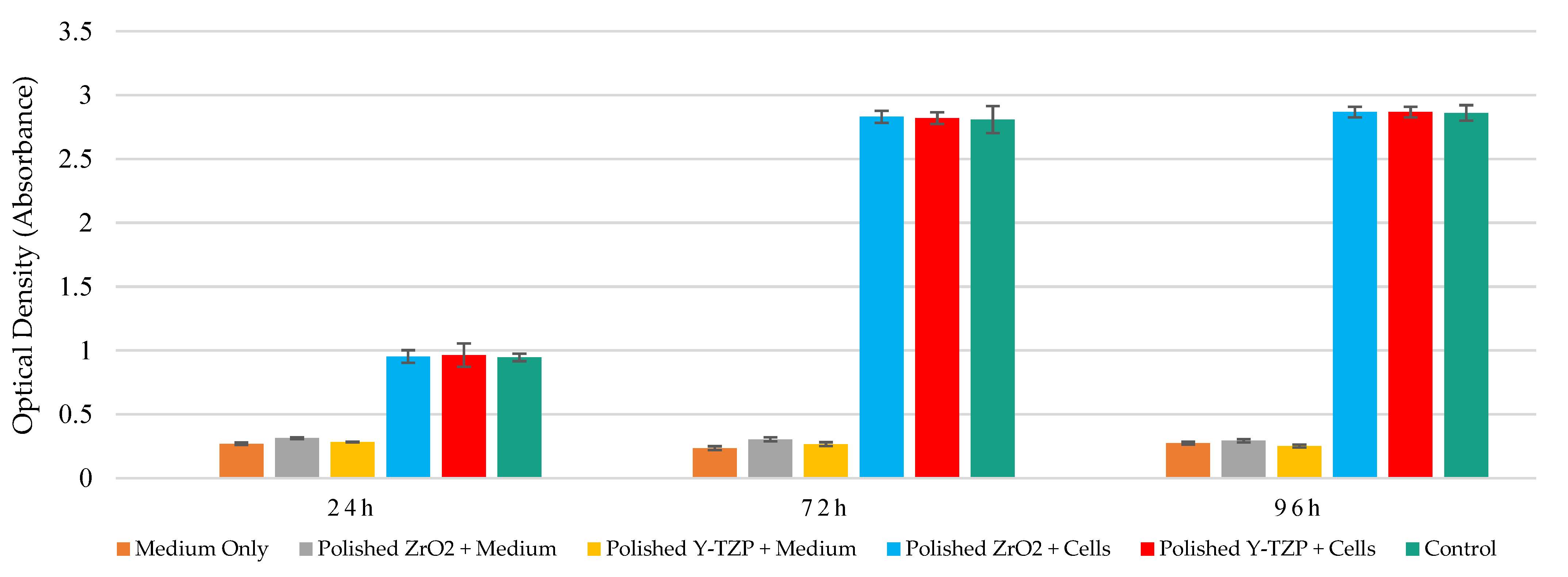
3.2. Measurement of Metabolic Activity of Saos-2 Cells in Contact with Smooth and Rough Surfaces of ZrO2 and Y-TZP Discs
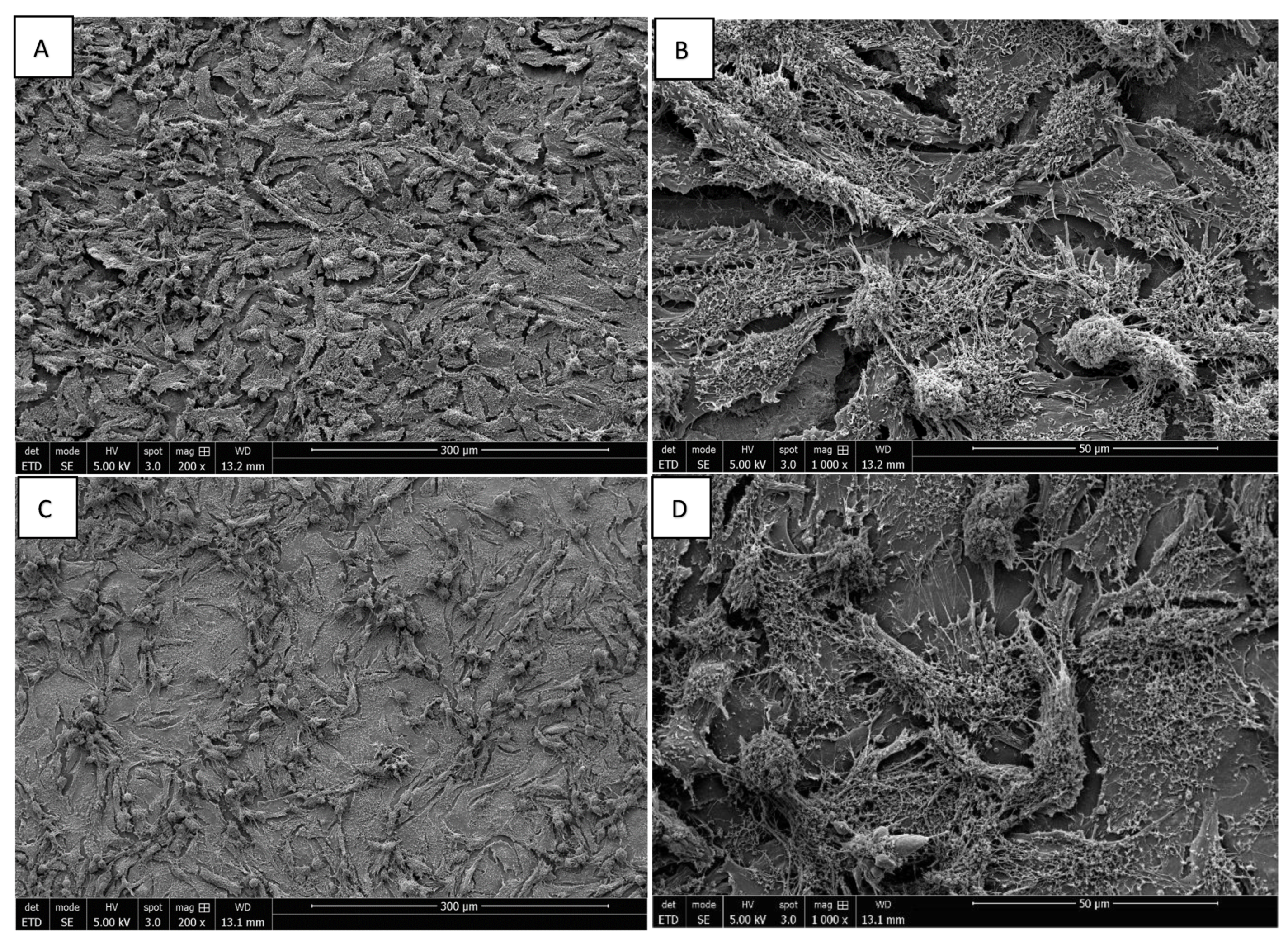
3.3. Morphology of Saos-2 Cells Exposed to ZrO2 and Y-TZP Discs
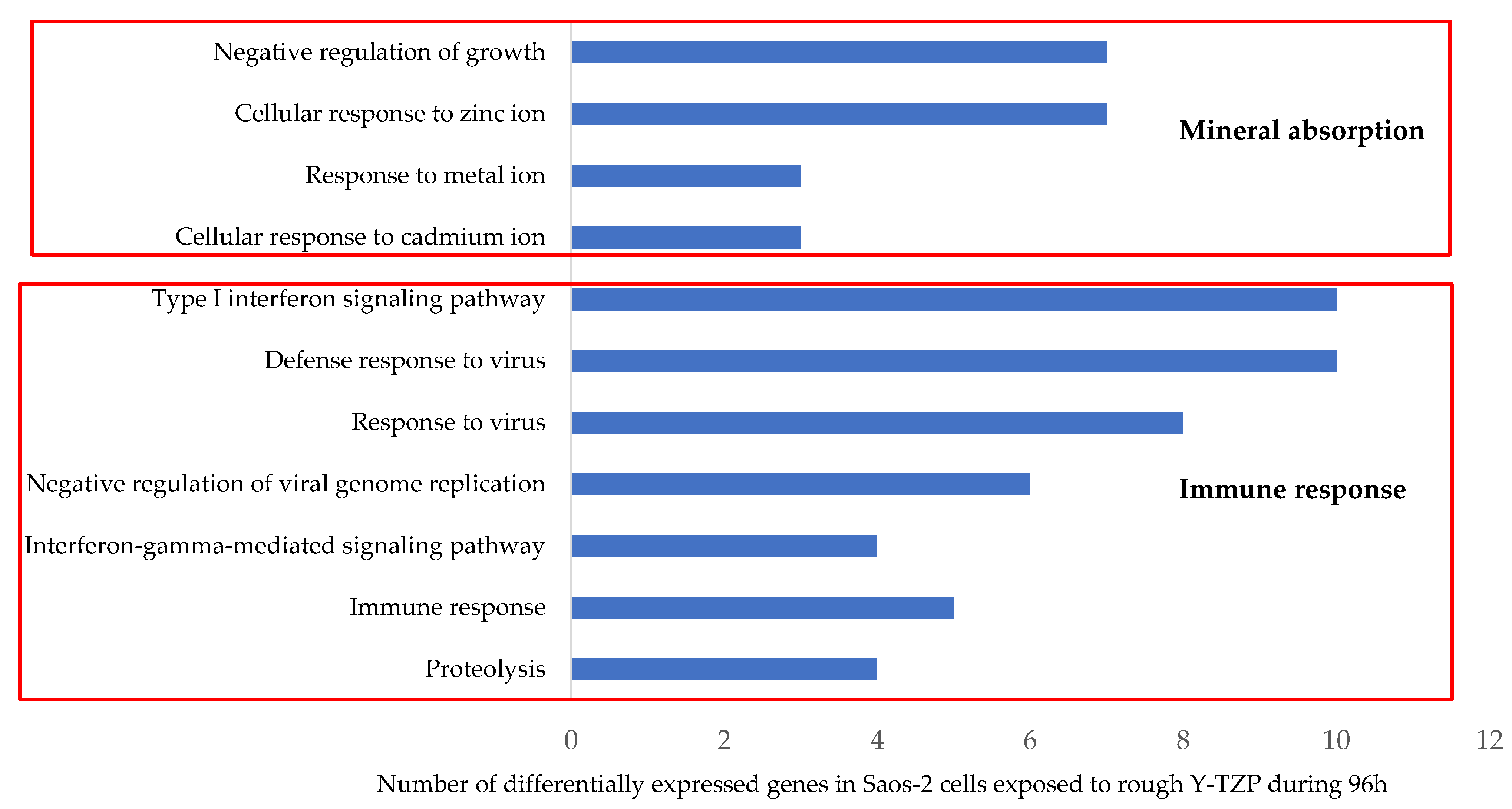
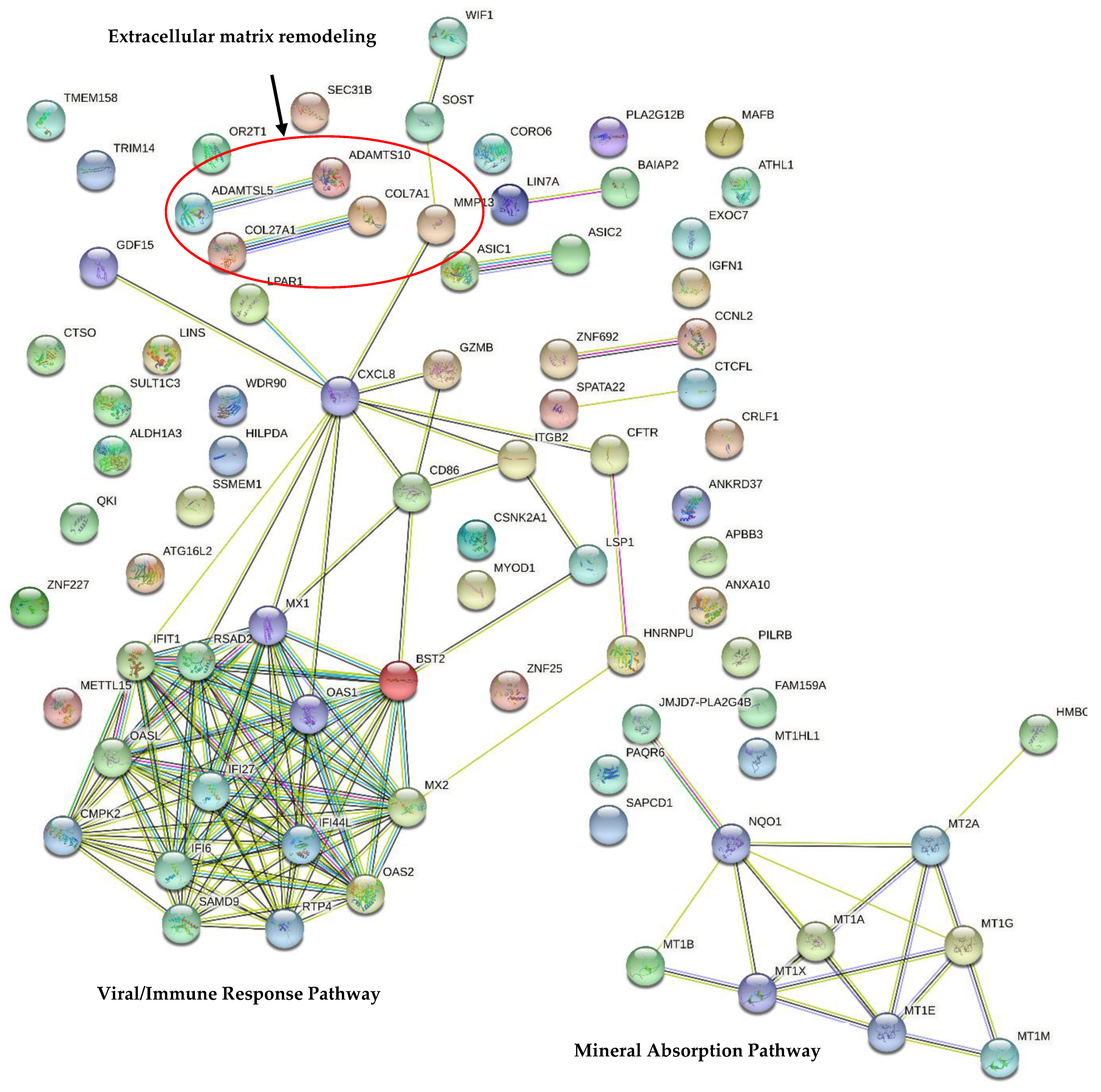
3.4. Transcriptomic Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Del Suárez-López Amo, F.; Garaicoa-Pazmiño, C.; Fretwurst, T.; Castilho, R.M.; Squarize, C.H. Dental Implants-Associated Release of Titanium Particles: A Systematic Review. Clin. Oral Implant. Researc. 2018, 29, 1085–1100. [Google Scholar] [CrossRef] [PubMed]
- Dias Corpa Tardelli, J.; da Lima Costa Valente, M.; de Theodoro Oliveira, T.; Cândido dos Reis, A. Influence of Chemical Composition on Cell Viability on Titanium Surfaces: A Systematic Review. J. Prosthet. Dent. 2021, 125, 421–425. [Google Scholar] [CrossRef] [PubMed]
- Jung, R.E.; Zembic, A.; Pjetursson, B.E.; Zwahlen, M.; Thoma, D.S. Systematic Review of the Survival Rate and the Incidence of Biological, Technical, and Aesthetic Complications of Single Crowns on Implants Reported in Longitudinal Studies with a Mean Follow-up of 5 Years. Clin. Oral Implant. Res. 2012, 23, 2–21. [Google Scholar] [CrossRef] [PubMed]
- Quentin, G. Influence des Etats de Surface sur le Developpement des Peri Implantites; Université Claude Bernard-Lyon 1 U.F.R. d’odontologie: Lyon, France, 2014; Volume 74, Available online: http://bibnum.univ-lyon1.fr/nuxeo/nxfile/default/9a2e93b1-b59e-4efe-ae4a-578610e64b21/file:content/THod_2014_GHIBAUDO_Quentin.pdf (accessed on 27 April 2022).
- Kissa, J.; El Kholti, W.; Chemlali, S.; Kawtari, H.; Laalou, Y.; Albandar, J.M. Prevalence and Risk Indicators of Peri-implant Diseases in a Group of Moroccan Patients. J. Periodontol. 2021, 92, 1096–1106. [Google Scholar] [CrossRef] [PubMed]
- Tchinda, A.; Pierson, G.; Chezeau, L.; Kouitat-Njiwa, R.; Rihn, B.H.; Bravetti, P. Desulfovibrio Fairfieldensis Adhesion on Implantable Titanium Used in Odontology: A Preliminary Study. Cell. Mol. Biol. 2021, 67, 56–65. [Google Scholar] [CrossRef]
- Comino-Garayoa, R.; Cortés-Bretón Brinkmann, J.; Peláez, J.; López-Suárez, C.; Martínez-González, J.M.; Suárez, M.J. Allergies to Titanium Dental Implants: What Do We Really Know about Them? A Scoping Review. Biology 2020, 9, 404. [Google Scholar] [CrossRef]
- Hafezeqoran, A.; Koodaryan, R. Effect of Zirconia Dental Implant Surfaces on Bone Integration: A Systematic Review and Meta-Analysis. BioMed Res. Int. 2017, 2017, 1–12. [Google Scholar] [CrossRef]
- Zhang, Y.; Lawn, B.R. Novel Zirconia Materials in Dentistry. J. Dent. Res. 2018, 97, 140–147. [Google Scholar] [CrossRef]
- Ban, S. Classification and Properties of Dental Zirconia as Implant Fixtures and Superstructures. Materials 2021, 14, 4879. [Google Scholar] [CrossRef]
- Yoshinari, M. Future Prospects of Zirconia for Oral Implants—A Review. Dent. Mater. J. 2020, 39, 37–45. [Google Scholar] [CrossRef] [Green Version]
- Piconi, C.; Maccauro, G. Zirconia as a Ceramic Biomaterial. Biomaterials 1999, 20, 1–25. [Google Scholar] [CrossRef]
- Duchatelard, P. La Zircone Est-Elle Une Alternative Au Titane En Implantologie Orale ? L’Information Dentaire. Available online: https://www.information-dentaire.fr/formations/la-zircone-est-elle-une-alternative-au-titane-en-implantologie-orale%e2%80%89/ (accessed on 5 June 2021).
- Akagawa, Y.; Hosokawa, R.; Sato, Y.; Kamayama, K. Comparison between Freestanding and Tooth-Connected Partially Stabilized Zirconia Implants after Two Years’ Function in Monkeys: A Clinical and Histologic Study. J. Prosthet. Dent. 1998, 80, 551–558. [Google Scholar] [CrossRef]
- Magne, P.; Belser, U. Restaurations Adhésives En Céramique Sur Dents Antérieures: Approche Biomimétique; Quintessence International: Paris, France, 2003. [Google Scholar]
- Kohal, R.J.; Weng, D.; Bächle, M.; Strub, J.R. Loaded Custom-Made Zirconia and Titanium Implants Show Similar Osseointegration: An Animal Experiment. J. Periodontol. 2004, 75, 1262–1268. [Google Scholar] [CrossRef]
- Scarano, A.; Piattelli, M.; Caputi, S.; Favero, G.A.; Piattelli, A. Bacterial Adhesion on Commercially Pure Titanium and Zirconium Oxide Disks: An In Vivo Human Study. J. Periodontol. 2004, 75, 292–296. [Google Scholar] [CrossRef]
- Siddiqi, A.; Khan, A.S.; Zafar, S. Thirty Years of Translational Research in Zirconia Dental Implants: A Systematic Review of the Literature. J. Oral Implantol. 2017, 43, 314–325. [Google Scholar] [CrossRef] [PubMed]
- Sivaraman, K.; Chopra, A.; Narayan, A.I.; Balakrishnan, D. Is Zirconia a Viable Alternative to Titanium for Oral Implant? A Critical Review. J. Prosthodont. Res. 2018, 62, 121–133. [Google Scholar] [CrossRef] [PubMed]
- Rimondini, L.; Cerroni, L.; Carrassi, A.; Torricelli, P. Bacterial Colonization of Zirconia Ceramic Surfaces: An in Vitro and in Vivo Study. Int. J. Oral Maxillofac. Implants. 2002, 17, 793–798. [Google Scholar]
- Roehling, S.; Schlegel, K.A.; Woelfler, H.; Gahlert, M. Zirconia Compared to Titanium Dental Implants in Preclinical Studies-A Systematic Review and Meta-Analysis. Clin. Oral Implants. Res. 2019, 30, 365–395. [Google Scholar] [CrossRef]
- Andreiotelli, M.; Wenz, H.J.; Kohal, R.-J. Are Ceramic Implants a Viable Alternative to Titanium Implants? A Systematic Literature Review. Clin. Oral Implant. Res. 2009, 20, 32–47. [Google Scholar] [CrossRef]
- Martins, R.; Cestari, T.M.; Arantes, R.V.N.; Santos, P.S.; Taga, R.; Carbonari, M.J.; Oliveira, R.C. Osseointegration of Zirconia and Titanium Implants in a Rabbit Tibiae Model Evaluated by Microtomography, Histomorphometry and Fluorochrome Labeling Analyses. J. Periodont. Res. 2018, 53, 210–221. [Google Scholar] [CrossRef]
- Sadowsky, S.J. Has Zirconia Made a Material Difference in Implant Prosthodontics? A Review. Dent. Mater. 2020, 36, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Kligman, S.; Ren, Z.; Chung, C.-H.; Perillo, M.A.; Chang, Y.-C.; Koo, H.; Zheng, Z.; Li, C. The Impact of Dental Implant Surface Modifications on Osseointegration and Biofilm Formation. JCM 2021, 10, 1641. [Google Scholar] [CrossRef] [PubMed]
- Kniha, K.; Heussen, N.; Modabber, A.; Hölzle, F.; Möhlhenrich, S.C. The Effect of Zirconia and Titanium Surfaces on Biofilm Formation and on Host-Derived Immunological Parameters. Int. J. Oral Maxillofac. Surg. 2021, 50, 1361–1374. [Google Scholar] [CrossRef] [PubMed]
- Bhola, M. Effets de la macro et de la nano-géométrie sur la stabilité primaire des implants. In 59ème Congrès de la SFMBCB; EDP Sciences: Ulys, France, 2012; p. 01003. [Google Scholar] [CrossRef] [Green Version]
- Le Guéhennec, L.; Soueidan, A.; Layrolle, P.; Amouriq, Y. Surface Treatments of Titanium Dental Implants for Rapid Osseointegration. Dent. Mater. 2007, 23, 844–854. [Google Scholar] [CrossRef]
- Matos, G.R.M. Surface Roughness of Dental Implant and Osseointegration. J. Maxillofac. Oral Surg. 2021, 20, 1–4. [Google Scholar] [CrossRef]
- Bosshardt, D.D.; Chappuis, V.; Buser, D. Osseointegration of Titanium, Titanium Alloy and Zirconia Dental Implants: Current Knowledge and Open Questions. Periodontology 2000 2017, 73, 22–40. [Google Scholar] [CrossRef]
- Noronha Oliveira, M.; Schunemann, W.V.H.; Mathew, M.T.; Henriques, B.; Magini, R.S.; Teughels, W.; Souza, J.C.M. Can Degradation Products Released from Dental Implants Affect Peri-implant Tissues? J. Periodont. Res. 2018, 53, 1–11. [Google Scholar] [CrossRef]
- Kumar, P.S. Systemic Risk Factors for the Development of Periimplant Diseases. Implant. Dent. 2019, 28, 115–119. [Google Scholar] [CrossRef]
- Chézeau, L.; Sébillaud, S.; Safar, R.; Seidel, C.; Dembélé, D.; Lorcin, M.; Langlais, C.; Grossmann, S.; Nunge, H.; Michaux, S.; et al. Short- and Long-Term Gene Expression Profiles Induced by Inhaled TiO2 Nanostructured Aerosol in Rat Lung. Toxicol. Appl. Pharmacol. 2018, 356, 54–64. [Google Scholar] [CrossRef]
- Brazma, A.; Hingamp, P.; Quackenbush, J.; Sherlock, G.; Spellman, P.; Stoeckert, C.; Aach, J.; Ansorge, W.; Ball, C.A.; Causton, H.C.; et al. Minimum Information about a Microarray Experiment (Miame)—Toward Standards for Microarray Data. Nat. Genet. 2001, 29, 365–371. [Google Scholar] [CrossRef]
- Barrett, T.; Edgar, R. [19] Gene Expression Omnibus: Microarray Data Storage, Submission, Retrieval, and Analysis. Methods Enzymol. 2006, 411, 352–369. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gene Expression Omnibus. Available online: http://www.ncbi.nlm.nih.gov/geo/ (accessed on 23 May 2022).
- Institut de Génétique et de Biologie Moléculaire et Cellulaire, Strasbourg. Available online: http://www-microarrays.u-strasbg.fr/Solo/index.html (accessed on 8 May 2022).
- Dembélé, D.; Kastner, P. Fold Change Rank Ordering Statistics: A New Method for Detecting Differentially Expressed Genes. BMC Bioinform. 2014, 15, 14. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- The Human Gene Database. Available online: https://www.genecards.org (accessed on 15 May 2022).
- Bioinformatics Resources. Available online: https://david.ncifcrf.gov/. (accessed on 19 May 2022).
- Huang, D.W.; Sherman, B.T.; Zheng, X.; Yang, J.; Imamichi, T.; Stephens, R.; Lempicki, R.A. Extracting Biological Meaning from Large Gene Lists with DAVID. Curr. Protoc. Bioinform. 2009, 27, 13.11.1–13.11.13. [Google Scholar] [CrossRef]
- Protein-Protein Interaction Networks Functional Enrichment Analysis. Available online: https://string-db.org (accessed on 21 May 2022).
- Szklarczyk, D.; Franceschini, A.; Wyder, S.; Forslund, K.; Heller, D.; Huerta-Cepas, J.; Simonovic, M.; Roth, A.; Santos, A.; Tsafou, K.P.; et al. STRING V10: Protein–Protein Interaction Networks, Integrated over the Tree of Life. Nucleic Acids Res. 2015, 43, D447–D452. [Google Scholar] [CrossRef] [PubMed]
- Chézeau, L.; Kohlstaedt, L.A.; Le Faou, A.; Cosnier, F.; Rihn, B.; Gaté, L. Proteomic Analysis of Bronchoalveolar Lavage Fluid in Rat Exposed to TiO2 Nanostructured Aerosol by Inhalation. J. Proteom. 2019, 207, 103451. [Google Scholar] [CrossRef] [PubMed]
- Baus-Domínguez, M.; Gómez-Díaz, R.; Corcuera-Flores, J.-R.; Torres-Lagares, D.; Ruiz-Villandiego, J.-C.; Machuca-Portillo, G.; Gutiérrez-Pérez, J.-L.; Serrera-Figallo, M.-A. Metallothioneins in Failure of Dental Implants and Periodontitis Down Syndrome Patients. Genes 2019, 10, 711. [Google Scholar] [CrossRef] [Green Version]
- Krause, C.; Korchynskyi, O.; de Rooij, K.; Weidauer, S.E.; de Gorter, D.J.J.; van Bezooijen, R.L.; Hatsell, S.; Economides, A.N.; Mueller, T.D.; Löwik, C.W.G.M.; et al. Distinct Modes of Inhibition by Sclerostin on Bone Morphogenetic Protein and Wnt Signaling Pathways. J. Biol. Chem. 2010, 285, 41614–41626. [Google Scholar] [CrossRef] [Green Version]
- Eger, M.; Hiram-Bab, S.; Liron, T.; Sterer, N.; Carmi, Y.; Kohavi, D.; Gabet, Y. Mechanism and Prevention of Titanium Particle-Induced Inflammation and Osteolysis. Front. Immunol. 2018, 9, 2963. [Google Scholar] [CrossRef] [Green Version]
- Wang, X.; Li, Y.; Feng, Y.; Cheng, H.; Li, D. Macrophage Polarization in Aseptic Bone Resorption around Dental Implants Induced by Ti Particles in a Murine Model. J. Periodont. Res. 2019, 54, 329–338. [Google Scholar] [CrossRef]
- Messous, R.; Henriques, B.; Bousbaa, H.; Silva, F.S.; Teughels, W.; Souza, J.C.M. Cytotoxic Effects of Submicron- and Nano-Scale Titanium Debris Released from Dental Implants: An Integrative Review. Clin. Oral Invest. 2021, 25, 1627–1640. [Google Scholar] [CrossRef]
- Cordeiro, J.M.; Barão, V.A.R. Is There Scientific Evidence Favoring the Substitution of Commercially Pure Titanium with Titanium Alloys for the Manufacture of Dental Implants? Mater. Sci. Eng. C 2017, 71, 1201–1215. [Google Scholar] [CrossRef] [PubMed]
- Bozoglan, A.; Dundar, S. ComparisoSAn of Osseointegration of Ti–Al6V4 and Ti–Al6Nb7 Implants: An Experimental Study. J. Oral Biol. Craniofac. Res. 2021, 11, 624–627. [Google Scholar] [CrossRef] [PubMed]
- Pieralli, S.; Kohal, R.J.; Jung, R.E.; Vach, K.; Spies, B.C. Clinical Outcomes of Zirconia Dental Implants: A Systematic Review. J. Dent. Res. 2017, 96, 38–46. [Google Scholar] [CrossRef] [PubMed]
- Kazi, G.A.S.; Yamagiwa, R. Cytotoxicity and Biocompatibility of High Mol% Yttria Containing Zirconia. Restor. Dent. Endod. 2020, 45, e52. [Google Scholar] [CrossRef] [PubMed]
- Albrektsson, T.; Wennerberg, A. Oral Implant Surfaces: Part 1—Review Focusing on Topographic and Chemical Properties of Different Surfaces and in Vivo Responses to Them. Int. J. Prosthodont. 2004, 17, 536–543. [Google Scholar]
- Thirumoorthy, N. Metallothionein: An Overview. WJG 2007, 13, 993. [Google Scholar] [CrossRef] [Green Version]
- Rahman, M.T.; Hossain, A.; Pin, C.H.; Yahya, N.A. Zinc and Metallothionein in the Development and Progression of Dental Caries. Biol. Trace Elem. Res. 2019, 187, 51–58. [Google Scholar] [CrossRef]
- Liu, A.; Zhang, Z.; Zhu, B.; Liao, Z.; Liu, Z. Metallothionein Protects Bone Marrow Stromal Cells against Hydrogen Peroxide-Induced Inhibition of Osteoblastic Differentiation. Cell Biol. Int. 2004, 28, 905–911. [Google Scholar] [CrossRef]
- Liu, W.; Konermann, A.; Guo, T.; Jäger, A.; Zhang, L.; Jin, Y. Canonical Wnt Signaling Differently Modulates Osteogenic Differentiation of Mesenchymal Stem Cells Derived from Bone Marrow and from Periodontal Ligament under Inflammatory Conditions. Biochim. Biophys. Acta (BBA)-Gen. Subj. 2014, 1840, 1125–1134. [Google Scholar] [CrossRef]
- Carinci, F.; Pezzetti, F.; Volinia, S.; Francioso, F.; Arcelli, D.; Farina, E.; Piattelli, A. Zirconium Oxide: Analysis of MG63 Osteoblast-like Cell Response by Means of a Microarray Technology. Biomaterials 2004, 25, 215–228. [Google Scholar] [CrossRef]
- Alford, A.I.; Kozloff, K.M.; Hankenson, K.D. Extracellular Matrix Networks in Bone Remodeling. Int. J. Biochem. Cell Biol. 2015, 65, 20–31. [Google Scholar] [CrossRef] [PubMed]
- Stadlinger, B.; Pilling, E.; Huhle, M.; Mai, R.; Bierbaum, S.; Bernhardt, R.; Scharnweber, D.; Kuhlisch, E.; Hempel, U.; Eckelt, U. Influence of Extracellular Matrix Coatings on Implant Stability and Osseointegration: An Animal Study. J. Biomed. Mater. Res. 2007, 83B, 222–231. [Google Scholar] [CrossRef] [PubMed]
- Fei, L.; Wang, C.; Xue, Y.; Lin, K.; Chang, J.; Sun, J. Osteogenic Differentiation of Osteoblasts Induced by Calcium Silicate and Calcium Silicate/β-Tricalcium Phosphate Composite Bioceramics. J. Biomed. Mater. Res. Part. B Appl. Biomater. 2012, 100B, 1237–1244. [Google Scholar] [CrossRef]
- Bou Assaf, R.; Fayyad-Kazan, M.; Al-Nemer, F.; Makki, R.; Fayyad-Kazan, H.; Badran, B.; Berbéri, A. Evaluation of the Osteogenic Potential of Different Scaffolds Embedded with Human Stem Cells Originated from Schneiderian Membrane: An In Vitro Study. BioMed Res. Int. 2019, 2019, 1–10. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kannan, S.; Ghosh, J.; Dhara, S.K. Osteogenic Differentiation Potential of Porcine Bone Marrow Mesenchymal Stem Cell Subpopulations Selected in Different Basal Media. Biol. Open 2020, 9, bio053280. [Google Scholar] [CrossRef]
- Ferreira, J.A.; Sartori, E.M.; Piola Rizzante, F.A.; Siqueira, R.; Silveira Mendonça, D.B.; de Magalhães, D.; Mendonça, G. Analysis In Vivo of Bone Modulation in Hydrophilic and Hydrophobic Titanium Implant Surfaces. Int. J. Oral Maxillofac. Implant. 2021, 36, 650–659. [Google Scholar] [CrossRef] [PubMed]
- Ashifa, N.; Viswanathan, K.; Sundaram, R.; Srinivasan, S. Sclerostin and Its Role as a Bone Modifying Agent in Periodontal Disease. J. Oral Biosci. 2021, 63, 104–110. [Google Scholar] [CrossRef]
- Zhang, W.; Liu, Y.; Hu, H.; Huang, H.; Bao, Z.; Yang, P.; Wang, Y.; You, G.; Yan, W.; Jiang, T.; et al. ALDH1A3: A Marker of Mesenchymal Phenotype in Gliomas Associated with Cell Invasion. PLoS ONE 2015, 10, e0142856. [Google Scholar] [CrossRef]
- Palmieri, A.; Pezzetti, F.; Brunelli, G.; Zollino, I.; Lo Muzio, L.; Martinelli, M.; Scapoli, L.; Arlotti, M.; Masiero, E.; Carinci, F. Zirconium Oxide Regulates RNA Interfering of Osteoblast-like Cells. J. Mater. Sci Mater. Med. 2008, 19, 2471–2476. [Google Scholar] [CrossRef]
- Komorowski, P.; Sokołowska, P.; Siatkowska, M.; Elgalal, M.; Rosowski, M.; Makowski, K.; Lipińska, L.; Leszczewicz, M.; Styczyński, A.; Fogel, K.; et al. Designing Laser-modified Surface Structures on Titanium Alloy Custom Medical Implants Using a Hybrid Manufacturing Technology. J. Biomed. Mater. Res. 2020, 108, 1790–1800. [Google Scholar] [CrossRef]
- Terheyden, H.; Lang, N.P.; Bierbaum, S.; Stadlinger, B. Osseointegration—Communication of Cells. Clin. Oral Impl. Res. 2012, 23, 1127–1135. [Google Scholar] [CrossRef] [PubMed]





| Gene Name | FC2 | f-Value | Gene Ontology Biological Process (GO BP) |
|---|---|---|---|
| ENST00000450667 | 2.04 | 0.997 | / |
| IFI27 | 2.47 | 0.997 | Immune system process (cellular antiviral response) |
| IFI44L | 2.36 | 0.997 | Involved in immune response (defense response to the virus) |
| MT1E | 2.24 | 0.997 | Cellular response to metal ion |
| MX1 | 2.74 | 0.998 | Immune system process (cellular antiviral response) |
| MYOD1 | 2.36 | 0.998 | Involved in muscle organ development |
| OAS1 | 2.45 | 0.997 | Immune system process (cellular antiviral response) |
| OAS2 | 2.01 | 0.997 | Immune system process (cellular antiviral response) |
| RSAD2 | 2.18 | 0.997 | Immune system process (cellular antiviral response) |
| SOST | 2.33 | 0.997 | Involved in ossification |
| TRIM14 | 2.03 | 0.996 | Immune system process (protein ubiquitination) |
| XLOC_008559 | 2.11 | 0.997 | / |
| Gene Name | FC2 | f-Value | Gene Ontology Biological Process (GO BP) |
|---|---|---|---|
| ATHL1 (PGGHG) | 0.57 | 0.003 | Metabolic process |
| EXOC7 | 0.59 | 0.004 | Involved in regulation of entry of bacterium into the host cell (exocytosis) |
| GZMB | 0.51 | 0.003 | Proteolysis (apoptotic process) |
| H19 | 0.58 | 0.003 | Involved in the cellular response to virus |
| IGFN1 | 0.59 | 0.003 | Involved in homophilic cell adhesion via plasma membrane adhesion molecules |
| LSP1 | 0.57 | 0.003 | Defense response |
| MIR143HG | 0.59 | 0.003 | Involved in actin cytoskeleton organization |
| NEAT1 | 0.57 | 0.003 | Involved in positive regulation of inflammatory response |
| SAPCD1 | 0.6 | 0.004 | / |
| WDR90 | 0.59 | 0.004 | Cell projection organization (cilium assembly) |
| KEGG | Number of Genes | Genes | p-Value |
|---|---|---|---|
| Mineral absorption | 6 | MT2A, MT1A, MT1M, MT1X, MT1B, MT1E | 2.386 × 10−6 |
| Influenza A | 5 | CXCL8, RSAD2, OAS1, OAS2, MX1 | 0.011 |
| Measles | 4 | CSNK2A1, OAS1, OAS2, MX1 | 0.029 |
| Hepatitis C | 4 | CXCL8, OAS1, OAS2, IFIT1 | 0.029 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tchinda, A.; Chézeau, L.; Pierson, G.; Kouitat-Njiwa, R.; Rihn, B.H.; Bravetti, P. Biocompatibility of ZrO2 vs. Y-TZP Alloys: Influence of Their Composition and Surface Topography. Materials 2022, 15, 4655. https://doi.org/10.3390/ma15134655
Tchinda A, Chézeau L, Pierson G, Kouitat-Njiwa R, Rihn BH, Bravetti P. Biocompatibility of ZrO2 vs. Y-TZP Alloys: Influence of Their Composition and Surface Topography. Materials. 2022; 15(13):4655. https://doi.org/10.3390/ma15134655
Chicago/Turabian StyleTchinda, Alex, Laëtitia Chézeau, Gaël Pierson, Richard Kouitat-Njiwa, B H Rihn, and Pierre Bravetti. 2022. "Biocompatibility of ZrO2 vs. Y-TZP Alloys: Influence of Their Composition and Surface Topography" Materials 15, no. 13: 4655. https://doi.org/10.3390/ma15134655
APA StyleTchinda, A., Chézeau, L., Pierson, G., Kouitat-Njiwa, R., Rihn, B. H., & Bravetti, P. (2022). Biocompatibility of ZrO2 vs. Y-TZP Alloys: Influence of Their Composition and Surface Topography. Materials, 15(13), 4655. https://doi.org/10.3390/ma15134655





