A Computational Model for Drug Release from PLGA Implant
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials
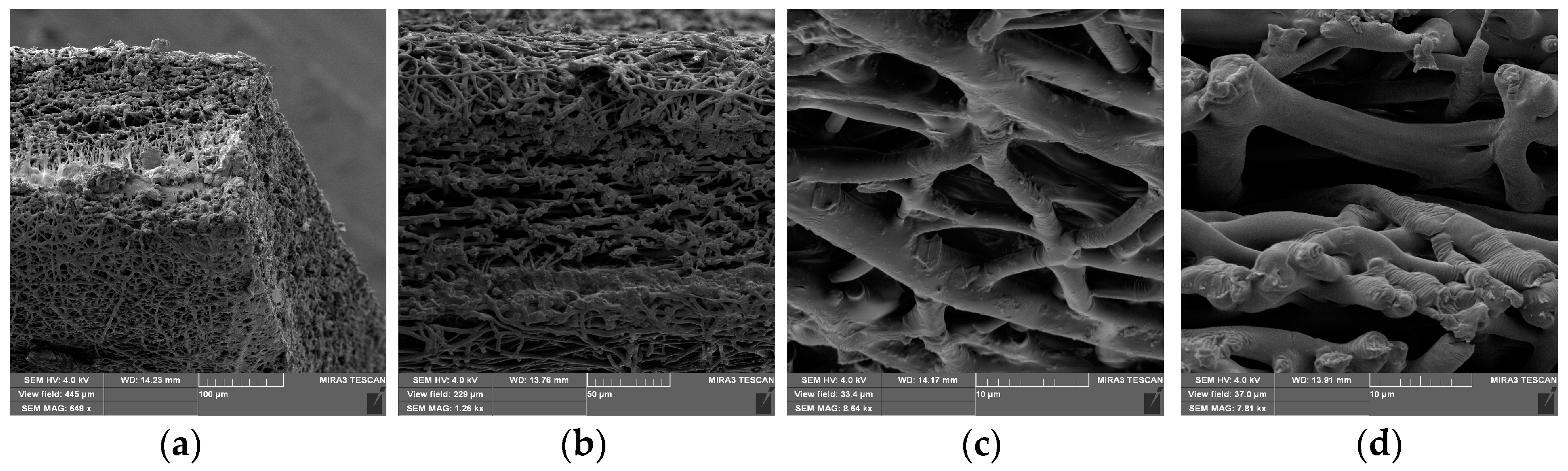
2.1.1. Preparation of PLGA Nanofibers Produced via Emulsion Electrospinning
2.1.2. Drug Loading Efficiency
2.1.3. In Vitro Drug Release Studies
3. Computational Models
3.1. Fundamental Equations
3.2. Diffusion within Fibers
3.2.1. Axial Diffusion
3.2.2. Radial Diffusion
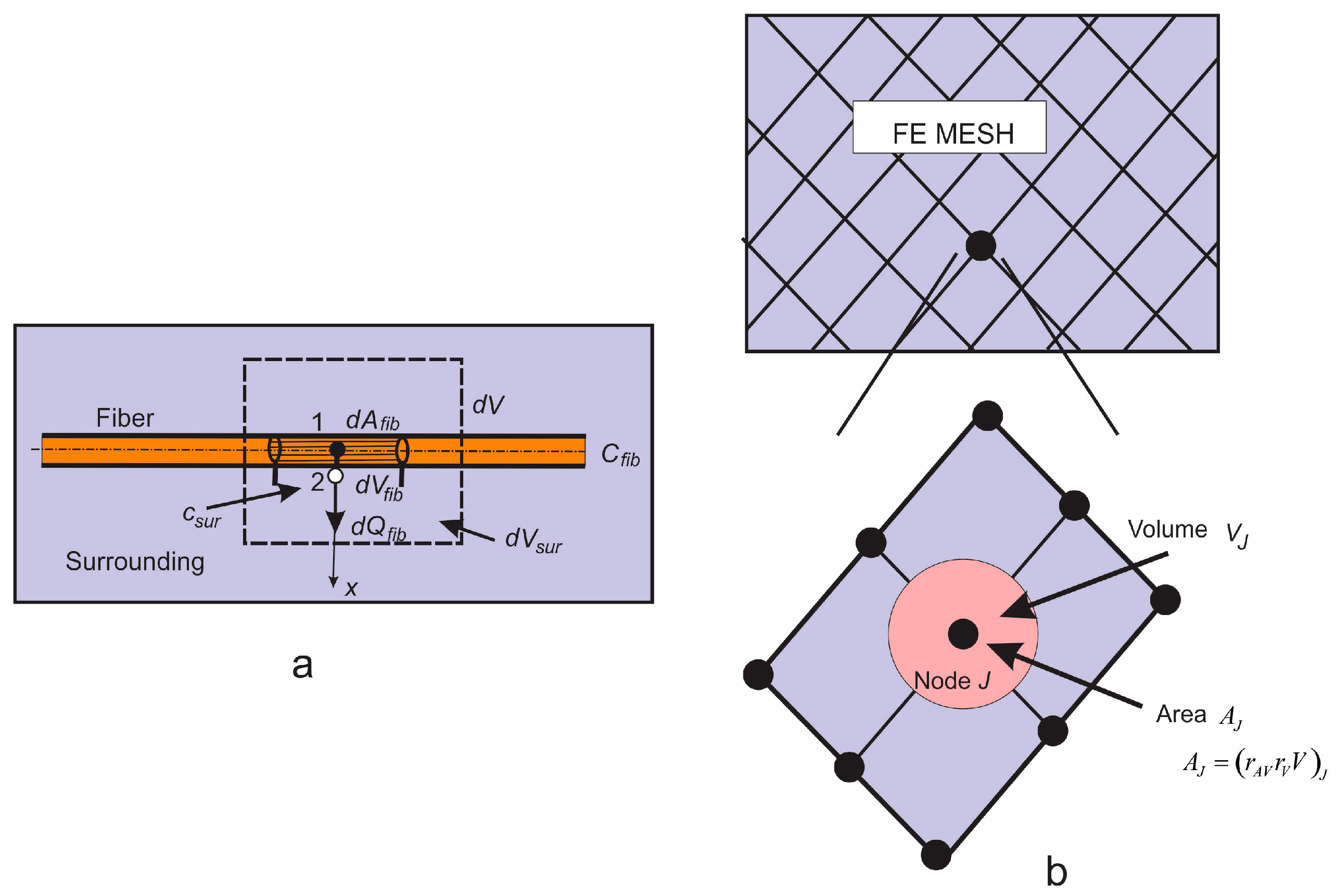
3.3. Fundamental Equations for CSFE
4. Numerical and Experimental Results of Drug Release
- Hydrophobicity (partitioning) of drug transport within PLGA1 is lower than for PLGA2;
- Degradation of PLGA1 is much slower than degradation of PLGA2.
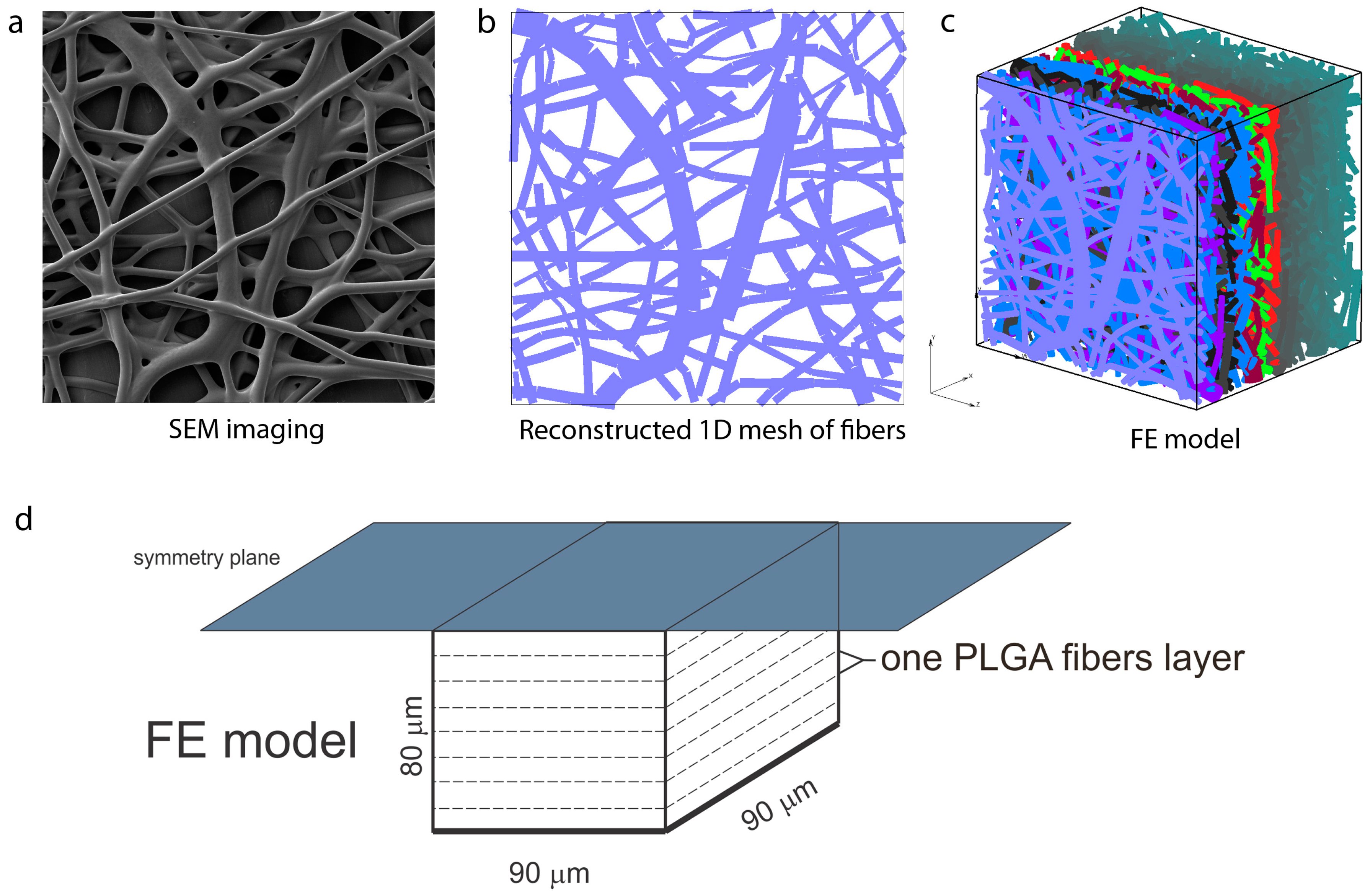
4.1. Preparation and Numerical Simulation of Detailed FE Models
- Dimensions: 80 µm × 90 µm × 90 µm;
- FE mesh: 40 × 48 × 48 divisions;
- Diffusion coefficient within fibers: Dfiber = 0.04 µm2/s;
- Diffusion coefficient in between fibers: Dliquid = 0.04 µm2/s;
- Mean diameter: D = 2.5 µm.
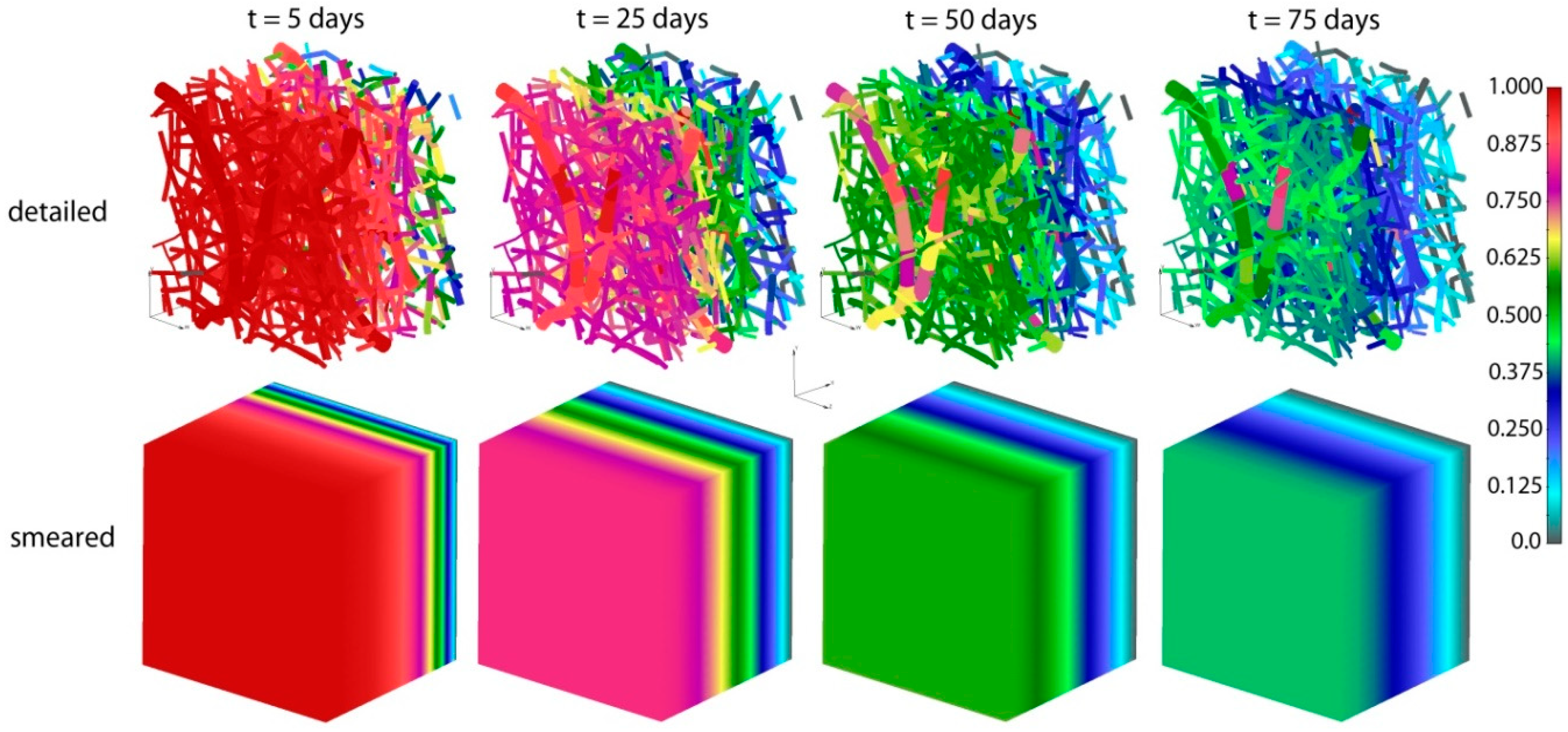
4.2. Application of Smeared Modeling for Drug Transport in PLGA Implant
- Fiber domain—equivalent domain of fibers;
- Surrounding domain—equivalent “pore” space surrounding fibers.
- Volume fraction of fibers in PLGA layers;
- Diffusion coefficient within PLGA fiber, for either 24 wt.% 50:50 and 65:35 emulsion;
- Diffusion coefficient of drug within the surrounding domain. Coefficient of hydrophobicity (partitioning);
- Mean diameter of PLGA fibers.
- Volume fraction of fibers: = 0.4223;
- Mean diameter of fibers: D = 2.5 µm;
- Diffusion coefficient within fibers: Dwall = 0.04 µm2/s;
- Equivalent diffusion coefficient in surrounding domain: Dliquid = 0.004 µm2/s;
- Partitioning: P = 1.
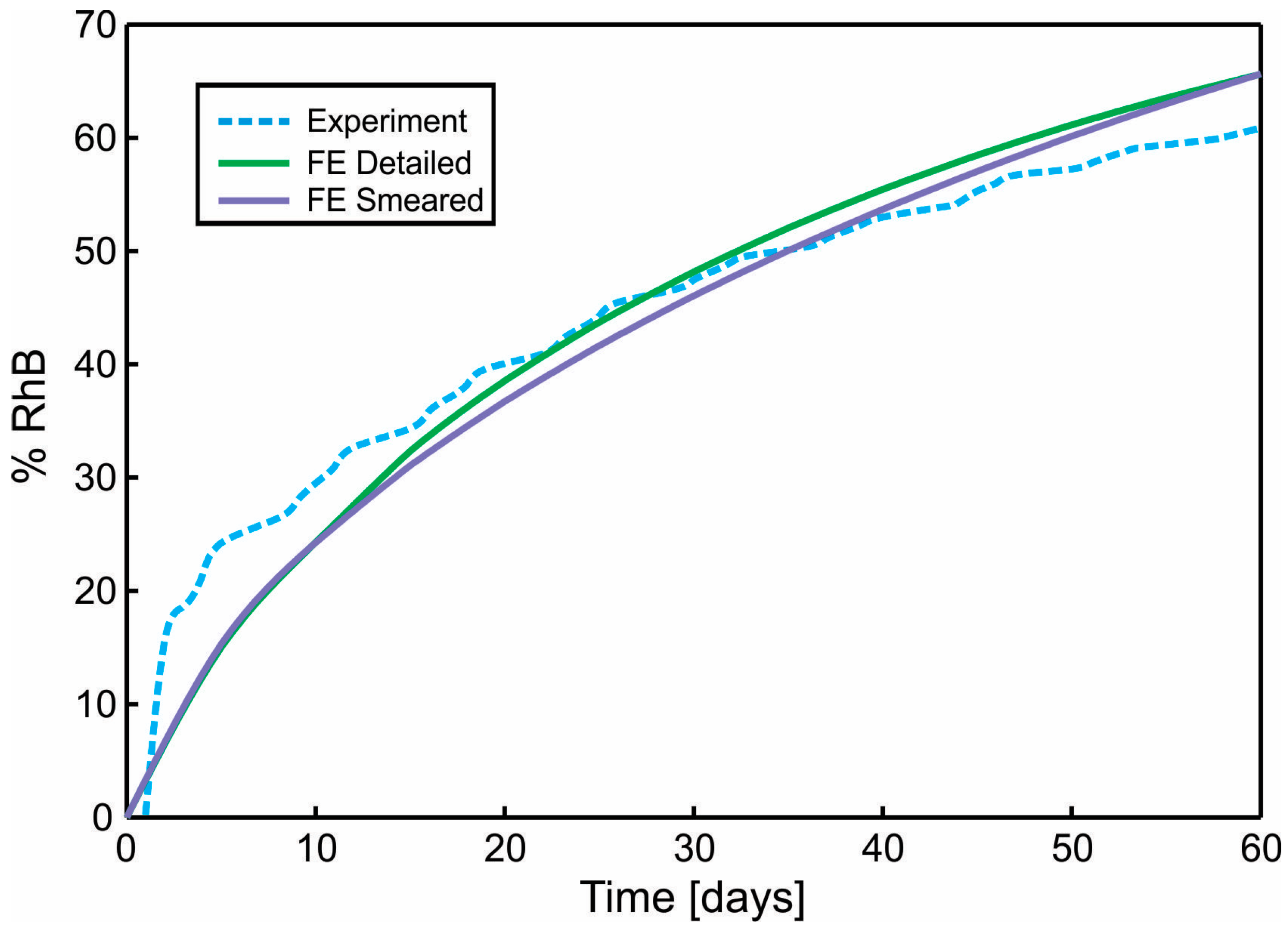
4.3. Comparation of Numerical and Experimental Results
5. Discussion
6. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A. Initial Concentrations for Radial Subelements
References
- Langer, R. Biomaterials in drug delivery and tissue engineering: One laboratory’s experience. Acc. Chem. Res. 2000, 33, 94–101. [Google Scholar] [CrossRef] [PubMed]
- Park, T.G.; Alonso, M.J.; Langer, R. Controlled release of proteins from poly (l-lactic acid) coated polyisobutylcyanoacrylate microcapsules. J. Appl. Polym. Sci. 1994, 52, 1797–1807. [Google Scholar] [CrossRef]
- Borden, M.; Attawia, M.; Laurencin, C.T. The sintered microsphere matrix for bone tissue engineering: In vitro osteoconductivity studies. J. Biomed. Mater. Res. 2002, 61, 421–429. [Google Scholar] [CrossRef] [PubMed]
- Kenawy, E.R.; Bowlin, G.L.; Mansfield, K.; Layman, J.; Simpson, D.G.; Sanders, E.H.; Wnek, G.E. Release of tetracycline hydrochloride from electrospun poly (ethylene-co-vinylacetate), poly (lactic acid), and a blend. J. Control. Release 2002, 81, 57–64. [Google Scholar] [CrossRef]
- Luo, X.; Xie, C.; Wang, H.; Liu, C.; Yan, S.; Li, X. Antitumor activities of emulsion electrospun fibers with core loading of hydroxycamptothecin via intratumoral implantation. Int. J. Pharm. 2012, 425, 19–28. [Google Scholar] [CrossRef] [PubMed]
- Katti, D.S.; Robinson, K.W.; Ko, F.K.; Laurencin, C.T. Bioresorbable nanofiber-based systems for wound healing and drug delivery: Optimization of fabrication parameters. J. Biomed. Mater. Res. Part B Appl. Biomater. 2004, 70, 286–296. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Xu, W.; Li, D.; Liu, T.; Zhang, Y.S.; Ding, J.; Chen, X. A locally deployable nanofiber patch for sequential drug delivery in treatment of primary and advanced orthotopic hepatomas. ACS Nano 2018, 12, 6685–6699. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Xu, W.; Chen, J.; Li, D.; Zhang, K.; Liu, T.; Ding, J.; Chen, X. Highly bioadhesive polymer membrane continuously releases cytostatic and anti-inflammatory drugs for peritoneal adhesion prevention. ACS Biomater. Sci. Eng. 2017, 4, 2026–2036. [Google Scholar] [CrossRef]
- Ramakrishna, S.; Fujihara, K.; Teo, W.; Lim, T.; Ma, Z. An Introduction to Electrospinning and Nanofibers; World Scientific: Singapore, 2005. [Google Scholar]
- Fong, H. Electrospun polymer, ceramic, carbon/graphite nanofibers and their applications. In Polymeric Nanostructures and Their Applications; American Scientific Publishers: Valencia, CA, USA, 2007; pp. 451–474. [Google Scholar]
- Greiner, A.; Wendorff, J.H. Electrospinning: A fascinating method for the preparation of ultrathin fibers. Angew. Chem. Int. Ed. 2007, 46, 5670–5703. [Google Scholar] [CrossRef] [PubMed]
- Liao, Y.; Zhang, L.; Gao, Y.; Zhu, Z.T.; Fong, H. Preparation, characterization, and encapsulation/release studies of a composite nanofiber mat electrospun from an emulsion containing poly (lactic-co-glycolic acid). Polymer 2008, 49, 5294–5299. [Google Scholar] [CrossRef] [PubMed]
- Mano, J.F.; Sousa, R.A.; Boesel, L.F.; Neves, N.M.; Reis, R.L. Bloinert, biodegradable and injectable polymeric matrix composites for hard tissue replacement: State of the art and recent developments. Compos. Sci. Technol. 2004, 64, 789–817. [Google Scholar] [CrossRef]
- Lin, H.R.; Kuo, C.J.; Yang, C.Y.; Shaw, S.Y.; Wu, Y.J. Preparation of macroporous biodegradable PLGA scaffolds for cell attachment with the use of mixed salts as porogen additives. J. Biomed. Mater. Res. A 2002, 63, 271–279. [Google Scholar] [CrossRef] [PubMed]
- Lu, L.; Peter, S.J.; Lyman, M.D.; Lai, H.; Leite, S.M.; Tamada, J.A.; Uyama, S.; Vacanti, J.P.; Langer, R.; Mikos, A.G. In vitro and in vivo degradation of porous poly(-lactic-co-glycolic acid) foams. Biomaterials 2000, 21, 1837–1845. [Google Scholar] [CrossRef]
- Lu, L.; Garcia, C.A.; Mikos, A.G. In vitro degradation of thin poly (d,l-lactic-co-glycolic acid) films. J. Biomed. Mater. Res. 1999, 46, 236–244. [Google Scholar] [CrossRef]
- Park, T.G. Degradation of poly(lactic- co-glycolic acid) microspheres: Effect of copolymer composition. Biomaterials 1995, 16, 1123–1130. [Google Scholar] [CrossRef]
- Makadia, H.K.; Siegel, S.J. Poly Lactic-co-glycolic acid (PLGA) as biodegradable controlled drug delivery carrier. Polymers 2011, 3, 1377–1397. [Google Scholar] [CrossRef] [PubMed]
- Sackett, C.K.; Narasimhan, B. Mathematical modeling of polymer erosion: Consequences for drug delivery. Int. J. Pharm. 2011, 418, 104–114. [Google Scholar] [CrossRef] [PubMed]
- Siepmann, J.; Geopferich, A. Mathematical modeling of bioerodible, polymeric drug delivery systems. Adv. Drug Deliv. Rev. 2001, 48, 229–247. [Google Scholar] [CrossRef]
- Zhu, X.; Braatz, R.D. A mechanistic model for drug release in PLGA biodegradable stent coatings coupled with polymer degradation and erosion. J. Biomed. Mater. Res. Part A 2015, 103, 2269–2279. [Google Scholar] [CrossRef] [PubMed]
- Kojic, M.; Milosevic, M.; Simic, V.; Stojanovic, D.; Uskokovic, P. A radial 1D finite element for drug release from drug loaded nanofibers. J. Serb. Soc. Comput. Mech. 2017, 11, 82–93. [Google Scholar] [CrossRef]
- Kojic, M.; Milosevic, M.; Simic, V.; Koay, E.J.; Fleming, J.B.; Nizzero, S.; Kojic, N.; Ziemys, A.; Ferrari, M. A composite smeared finite element for mass transport in capillary systems and biological tissue. Comput. Methods Appl. Mech. Eng. 2017, 324, 413–437. [Google Scholar] [CrossRef] [PubMed]
- Kojic, M.; Milosevic, M.; Simic, V.; Koay, E.J.; Kojic, N.; Ziemys, A.; Ferrari, M. Multiscale smeared finite element model for mass transport in biological tissue: From blood vessels to cells and cellular organelles. Comput. Biol. Med. 2018, 99, 7–23. [Google Scholar] [CrossRef] [PubMed]
- Milosevic, M.; Simic, V.; Milicevic, B.; Koay, E.J.; Ferrari, M.; Ziemys, A.; Kojic, M. Correction function for accuracy improvement of the Composite Smeared Finite Element for diffusive transport in biological tissue systems. Comput. Meth. Appl. Mech. Eng. 2018, 338, 97–116. [Google Scholar] [CrossRef]
- Fouad, H.; Elsarnagawy, T.; Almajhdi, F.N.; Khalil, K.A. Preparation and in vitro thermo-mechanical characterization of electrospun PLGA nanofibers for soft and hard tissue replacement. Int. J. Electrochem. Sci. 2013, 8, 2293–2304. [Google Scholar]
- Sun, S.B.; Liu, P.; Shao, F.M.; Miao, Q.L. Formulation and evaluation of PLGA nanoparticles loaded capecitabine for prostate cancer. Int. J. Clin. Exp. Med. 2015, 8, 19670. [Google Scholar] [PubMed]
- Jonderian, A.; Maalouf, R. Formulation and in vitro interaction of Rhodamine-B loaded PLGA nanoparticles with cardiac myocytes. Front. Pharmacol. 2016, 7, 458. [Google Scholar] [CrossRef] [PubMed]
- Kojic, M.; Filipovic, N.; Stojanovic, B.; Kojic, N. Computer Modelling in Bioengineering—Theory, Examples and Software; John Wiley and Sons: Hoboken, NJ, USA, 2018. [Google Scholar]
- Kojic, M.; Slavkovic, R.; Zivkovic, M.; Grujovic, N.; Filipovic, N.; Milosevic, M. PAK-FE program for structural analysis, fluid mechanics, coupled problems and biomechanics. Bioengineering R&D Center for Bioengineering, Faculty of Technical Science: Kragujevac, Serbia. Available online: http://www.bioirc.ac.rs/index.php/software (accessed on 21 November 2018).
- Ruiz-Esparza, G.U.; Wu, S.; Segura-Ibarra, V.; Cara, F.E.; Evans, K.W.; Milosevic, M.; Ziemys, A.; Kojic, M.; Meric-Bernstam, F.; Ferrari, M.; et al. Polymer nanoparticles enhanced in a cyclodextrin complex shell for potential site- and sequence-specific drug release. Adv. Funct. Mater. 2014, 24, 4753–4761. [Google Scholar] [CrossRef]
- Ziemys, A.; Kojic, M.; Milosevic, M.; Kojic, N.; Hussain, F.; Ferrari, M.; Grattoni, A. Hierarchical modeling of diffusive transport through nanochannels by coupling molecular dynamics with finite element method. J. Comput. Phys. 2001, 230, 5722–5731. [Google Scholar] [CrossRef]







© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Milosevic, M.; Stojanovic, D.; Simic, V.; Milicevic, B.; Radisavljevic, A.; Uskokovic, P.; Kojic, M. A Computational Model for Drug Release from PLGA Implant. Materials 2018, 11, 2416. https://doi.org/10.3390/ma11122416
Milosevic M, Stojanovic D, Simic V, Milicevic B, Radisavljevic A, Uskokovic P, Kojic M. A Computational Model for Drug Release from PLGA Implant. Materials. 2018; 11(12):2416. https://doi.org/10.3390/ma11122416
Chicago/Turabian StyleMilosevic, Miljan, Dusica Stojanovic, Vladimir Simic, Bogdan Milicevic, Andjela Radisavljevic, Petar Uskokovic, and Milos Kojic. 2018. "A Computational Model for Drug Release from PLGA Implant" Materials 11, no. 12: 2416. https://doi.org/10.3390/ma11122416
APA StyleMilosevic, M., Stojanovic, D., Simic, V., Milicevic, B., Radisavljevic, A., Uskokovic, P., & Kojic, M. (2018). A Computational Model for Drug Release from PLGA Implant. Materials, 11(12), 2416. https://doi.org/10.3390/ma11122416







