Seasonal Shifts in Soil Microbiome Structure Are Associated with the Cultivation of the Local Runner Bean Variety around the Lake Mikri Prespa
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
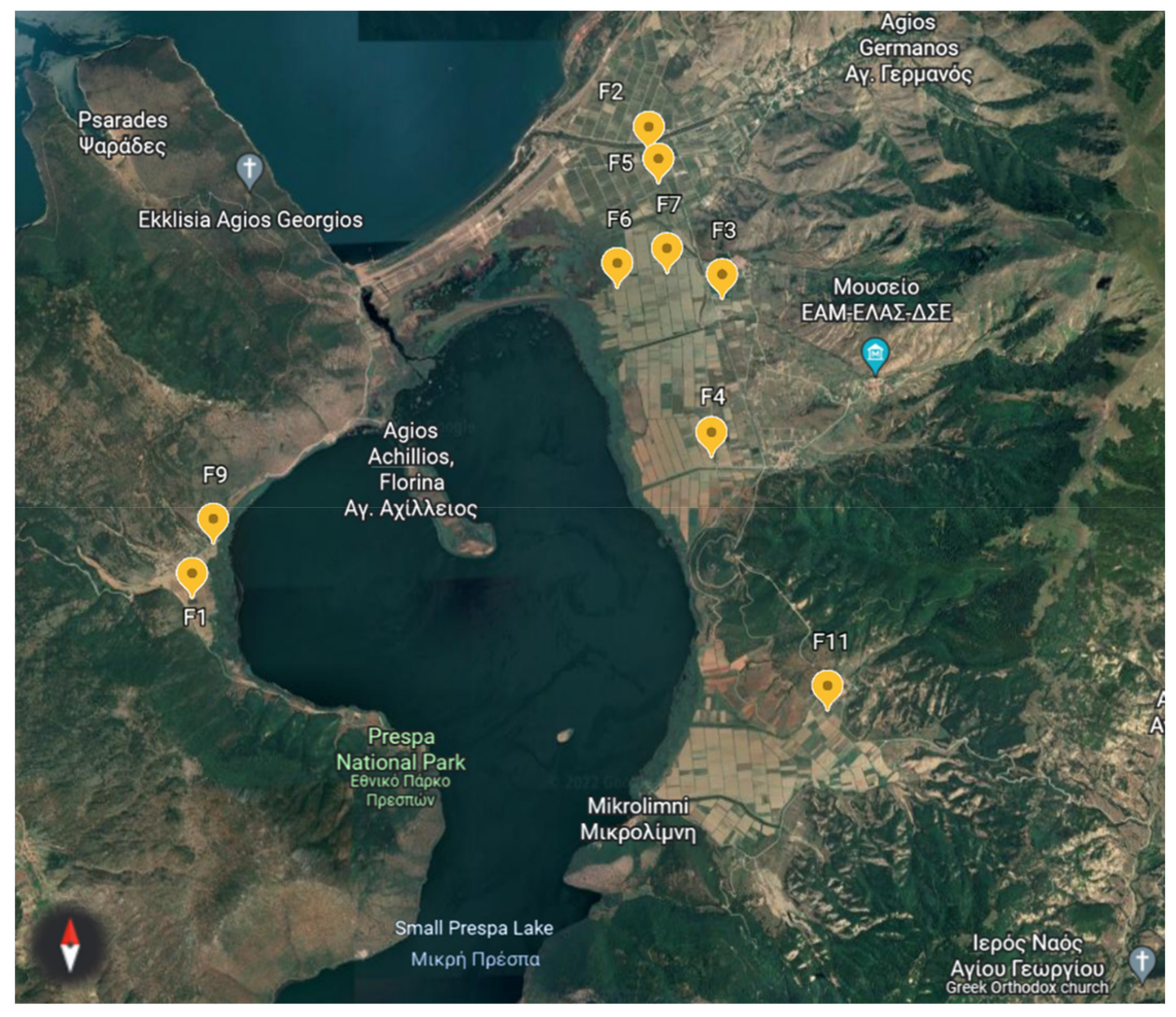
2.1. Experimental Site and Soil Samples
2.2. DNA Isolation, Library Preparation and Sequencing
2.3. Bioinformatic Analysis
2.4. Phylogeny-Based Functional Annotation
3. Results
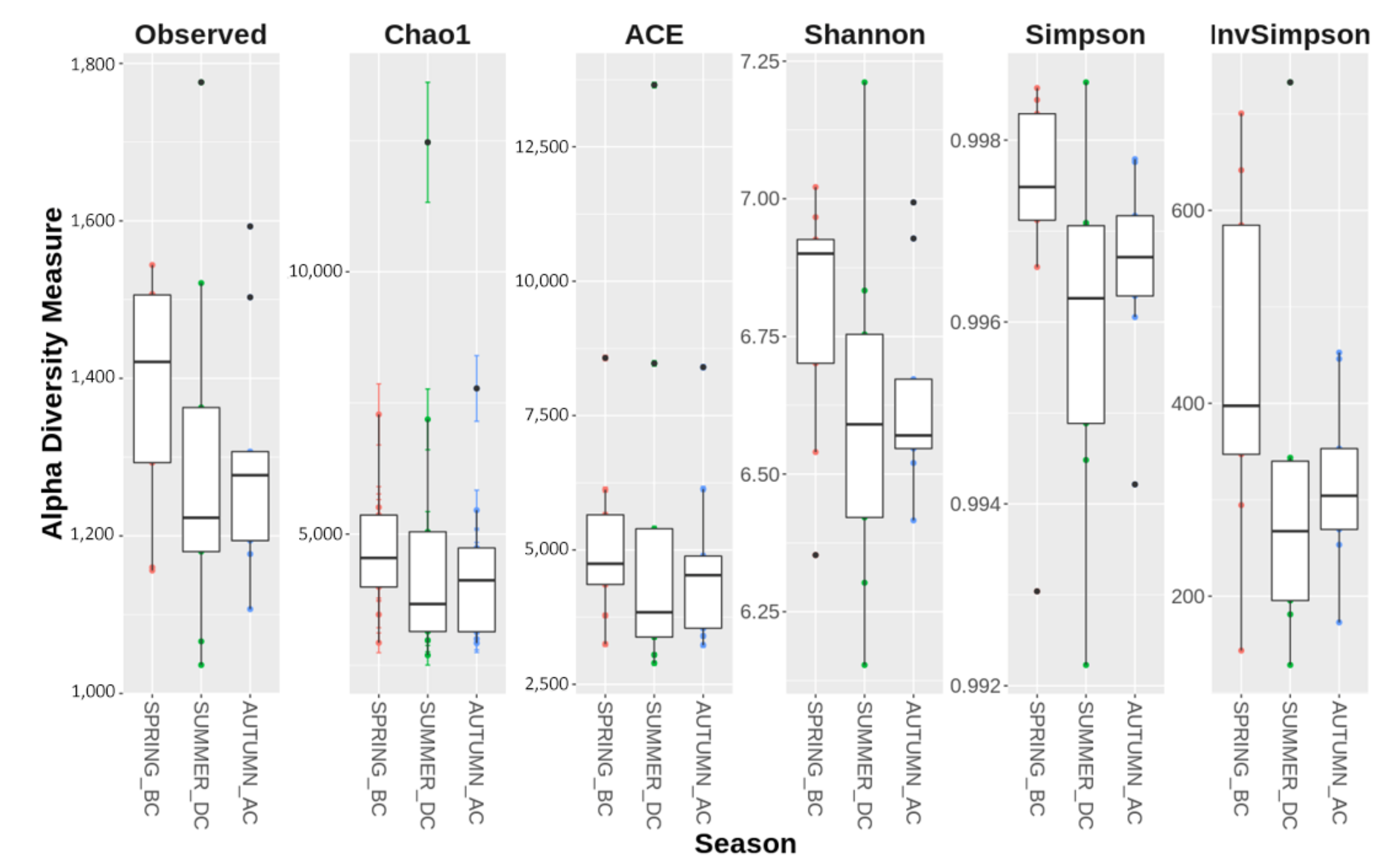
3.1. Seasonal Shifts in Microbial Communities’ Diversity
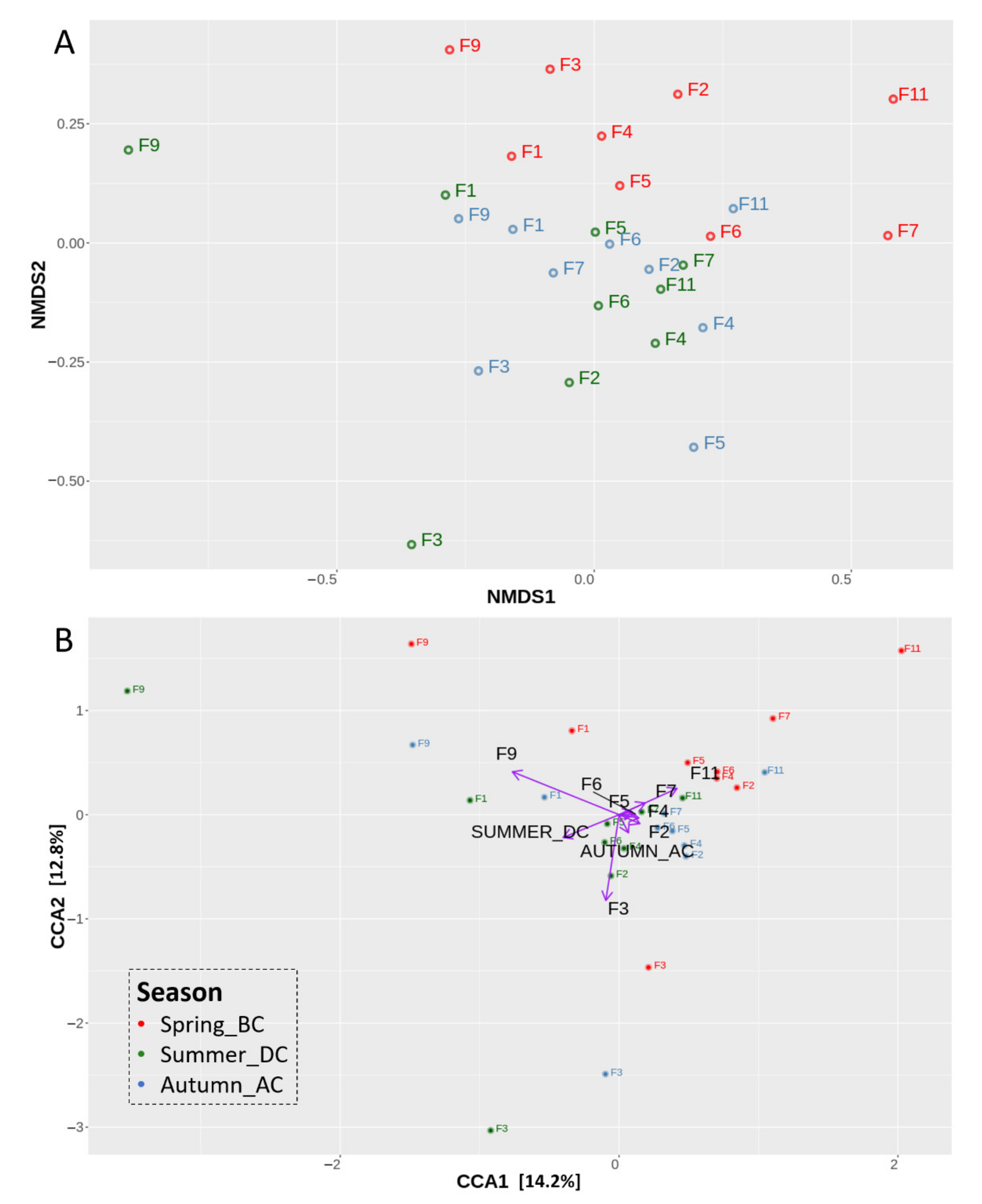
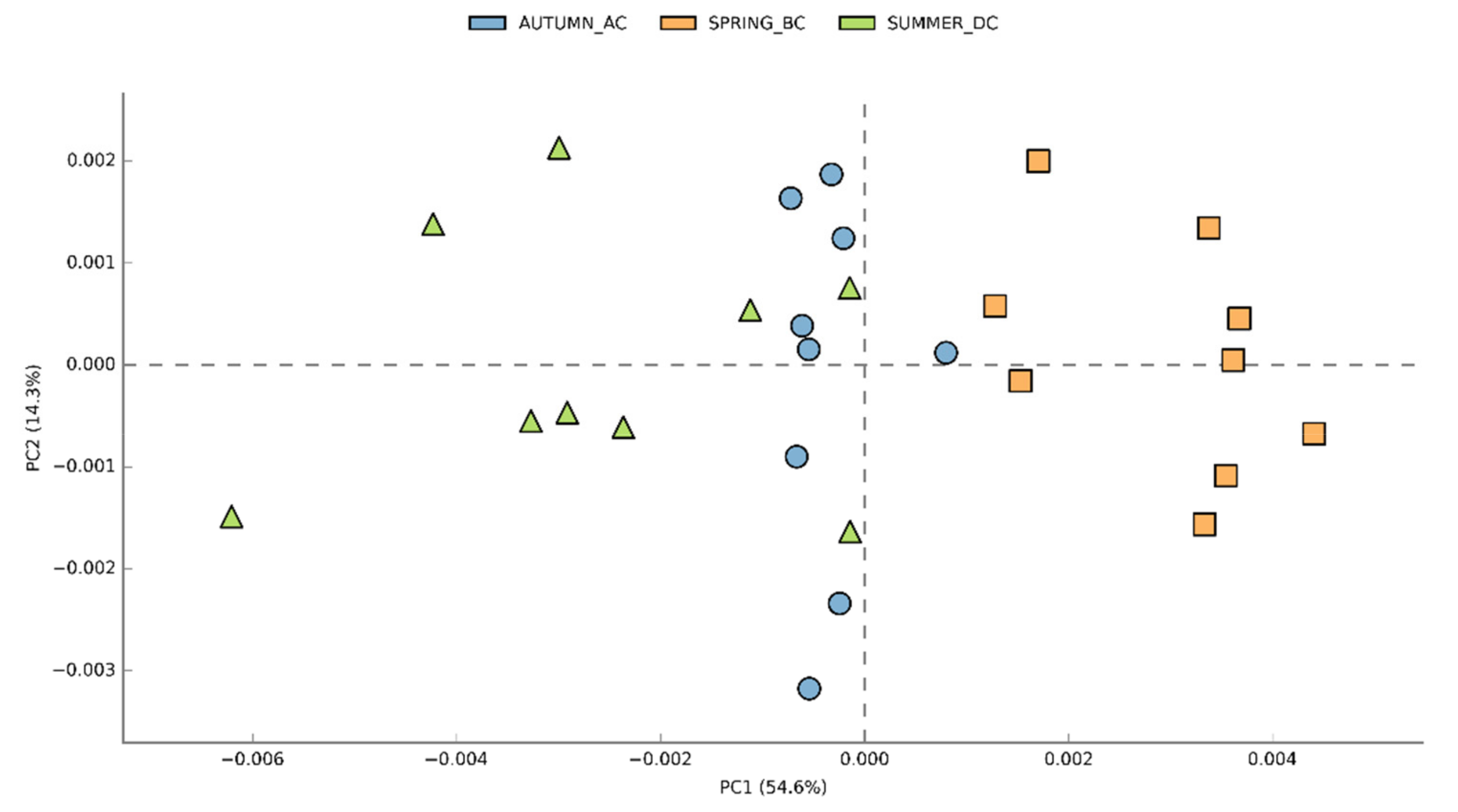
3.2. Effects of Cultivation Period on the Bacterial Community Structure
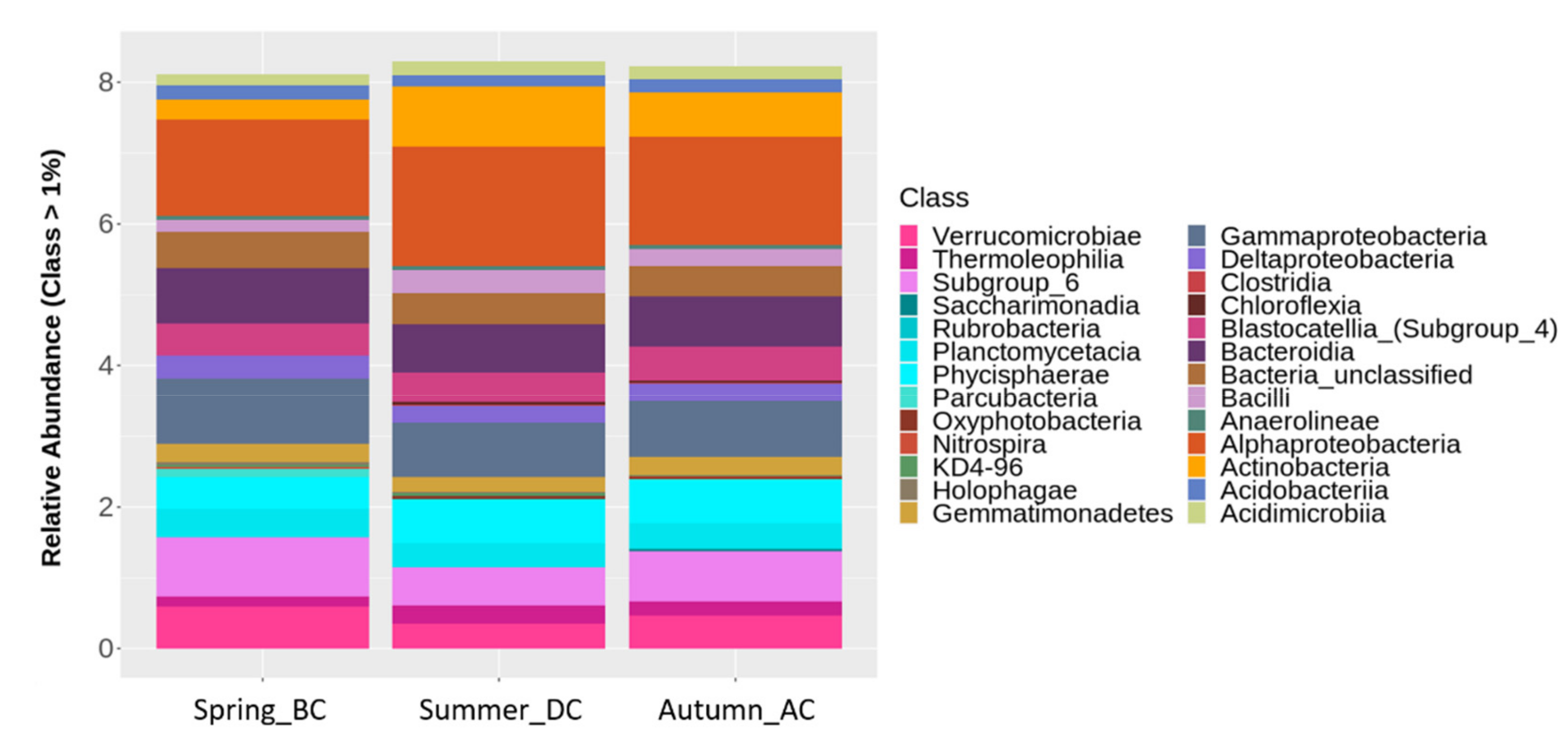
3.3. Effects of Cultivation Period on Different Taxonomical Levels
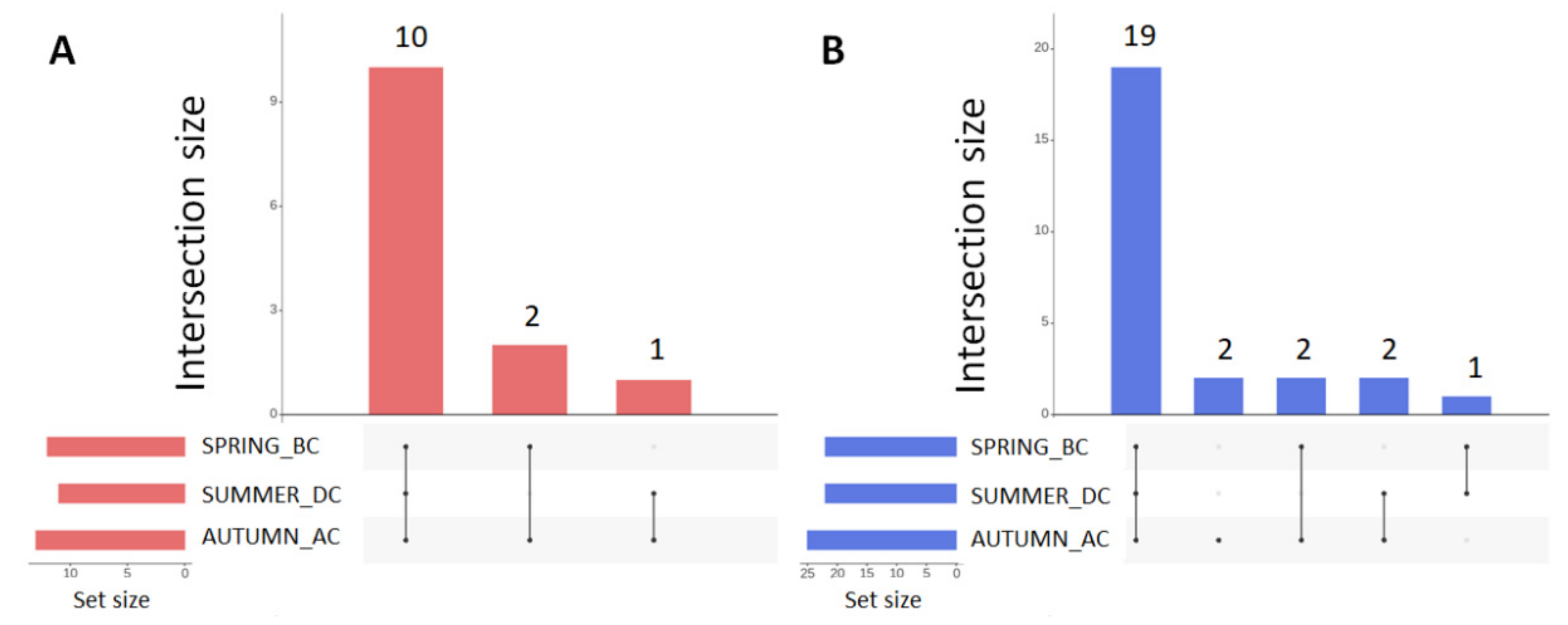
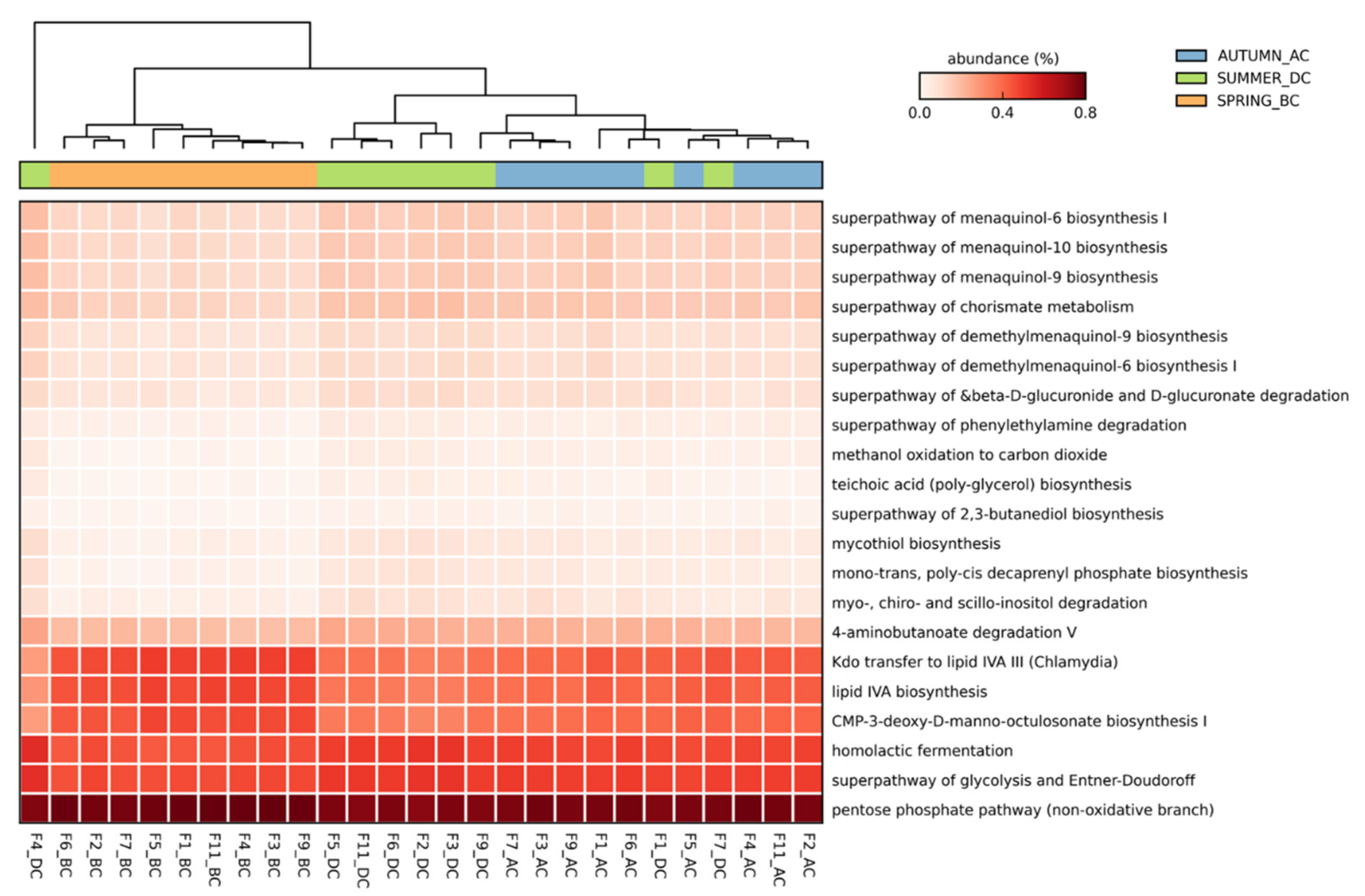
3.4. Predicted Functional Diversity of the Microbiome Present in the Different Field Sites
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Garbeva, P.; Van Veen, J.A.; Van Elsas, J.D. MICROBIAL DIVERSITY IN SOIL: Selection of Microbial Populations by Plant and Soil Type and Implications for Disease Suppressiveness. Ann. Rev. 2004, 42, 243–270. [Google Scholar] [CrossRef] [PubMed]
- Torsvik, V.; Øvreås, L. Microbial diversity and function in soil: From genes to ecosystems. Curr. Opin. Microbiol. 2002, 5, 240–245. [Google Scholar] [CrossRef]
- Lori, M.; Symnaczik, S.; Mäder, P.; De Deyn, G.; Gattinger, A. Organic farming enhances soil microbial abundance and activity—A meta-analysis and meta-Regression. PLoS ONE 2017, 12, 1–25. [Google Scholar] [CrossRef] [PubMed]
- Howard, M.M.; Muñoz, C.A.; Kao-Kniffin, J.; Kessler, A. Soil Microbiomes From Fallow Fields Have Species-Specific Effects on Crop Growth and Pest Resistance. Front. Plant Sci. 2020, 11, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Orgiazzi, A.; Ballabio, C.; Panagos, P.; Jones, A.; Fernández-Ugalde, O. LUCAS Soil, the largest expandable soil dataset for Europe: A review. Eur. J. Soil Sci. 2018, 69, 140–153. [Google Scholar] [CrossRef]
- Mendes, L.W.; Braga, L.P.P.; Navarrete, A.A.; de Souza, D.G.; Silva, G.G.Z.; Tsai, S.M. Using Metagenomics to Connect Microbial Community Biodiversity and Functions. Curr. Issues Mol. Biol. 2017, 24, 103–118. [Google Scholar] [CrossRef]
- Kielak, A.; Pijl, A.S.; Van Veen, J.A.; Kowalchuk, G.A. Differences in vegetation composition and plant species identity lead to only minor changes in soil-borne microbial communities in a former arable field. FEMS Microbiol. Ecol. 2008, 63, 372–382. [Google Scholar] [CrossRef]
- Fitzpatrick, C.R.; Mikhailitchenko, A.V.; Anstett, D.N.; Johnson, M.T.J. The influence of range-wide plant genetic variation on soil invertebrate communities. Ecography 2018, 41, 1135–1146. [Google Scholar] [CrossRef]
- Philippot, L.; Raaijmakers, J.M.; Lemanceau, P.; Van Der Putten, W.H. Going back to the roots: The microbial ecology of the rhizosphere. Nat. Rev. Microbiol. 2013, 11, 789–799. [Google Scholar] [CrossRef]
- Bardgett, R.D.; Van Der Putten, W.H. Belowground biodiversity and ecosystem functioning. Nature 2014, 515, 505–511. [Google Scholar] [CrossRef]
- Bender, S.F.; Wagg, C.; van der Heijden, M.G.A. An Underground Revolution: Biodiversity and Soil Ecological Engineering for Agricultural Sustainability. Trends Ecol. Evol. 2016, 31, 440–452. [Google Scholar] [CrossRef]
- Jiao, S.; Li, J.; Li, Y.; Xu, Z.; Kong, B.; Li, Y.; Shen, Y. Variation of soil organic carbon and physical properties in relation to land uses in the Yellow River Delta, China. Sci. Rep. 2020, 10, 1–12. [Google Scholar] [CrossRef]
- Fierer, N. Embracing the unknown: Disentangling the complexities of the soil microbiome. Nat. Rev. Microbiol. 2017, 15, 579–590. [Google Scholar] [CrossRef]
- Widder, S.; Allen, R.J.; Pfeiffer, T.; Curtis, T.P.; Wiuf, C.; Sloan, W.T.; Cordero, O.X.; Brown, S.P.; Momeni, B.; Shou, W.; et al. Challenges in microbial ecology: Building predictive understanding of community function and dynamics. ISME J. 2016, 10, 2557. [Google Scholar] [CrossRef]
- Valencia, E.; Gross, N.; Quero, J.L.; Carmona, C.P.; Ochoa, V.; Gozalo, B.; Delgado-Baquerizo, M.; Dumack, K.; Hamonts, K.; Singh, B.K.; et al. Cascading effects from plants to soil microorganisms explain how plant species richness and simulated climate change affect soil multifunctionality. Glob. Chang. Biol. 2018, 24, 5642–5654. [Google Scholar] [CrossRef]
- Li, X.; Jousset, A.; de Boer, W.; Carrión, V.J.; Zhang, T.; Wang, X.; Kuramae, E.E. Legacy of land use history determines reprogramming of plant physiology by soil microbiome. ISME J. 2019, 13, 738–751. [Google Scholar] [CrossRef]
- Nannipieri, P.; Ascher, J.; Ceccherini, M.T.; Landi, L.; Pietramellara, G.; Renella, G. Microbial diversity and soil functions. Eur. J. Soil Sci. 2003, 54, 655–670. [Google Scholar] [CrossRef]
- Chaparro, J.M.; Sheflin, A.M.; Manter, D.K.; Vivanco, J.M. Manipulating the soil microbiome to increase soil health and plant fertility. Biol. Fertil. Soils 2012, 48, 489–499. [Google Scholar] [CrossRef]
- Louca, S.; Polz, M.F.; Mazel, F.; Albright, M.B.N.; Huber, J.A.; O’Connor, M.I.; Ackermann, M.; Hahn, A.S.; Srivastava, D.S.; Crowe, S.A.; et al. Function and functional redundancy in microbial systems. Nat. Ecol. Evol. 2018, 2, 936–943. [Google Scholar] [CrossRef]
- Tsiknia, M.; Tsikou, D.; Papadopoulou, K.K.; Ehaliotis, C. Multi-species relationships in legume roots: From pairwise legume-symbiont interactions to the plant – microbiome – soil continuum. FEMS Microbiol. Ecol. 2021, 97, fiaa222. [Google Scholar] [CrossRef]
- Banerjee, S.; Schlaeppi, K.; van der Heijden, M.G.A. Keystone taxa as drivers of microbiome structure and functioning. Nat. Rev. Microbiol. 2018, 16, 567–576. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, S.; Walder, F.; Büchi, L.; Meyer, M.; Held, A.Y.; Gattinger, A.; Keller, T.; Charles, R.; van der Heijden, M.G.A. Agricultural intensification reduces microbial network complexity and the abundance of keystone taxa in roots. ISME J. 2019, 13, 1722–1736. [Google Scholar] [CrossRef] [PubMed]
- Herren, C.M.; McMahon, K.D. Keystone taxa predict compositional change in microbial communities. Environ. Microbiol. 2018, 20, 2207–2217. [Google Scholar] [CrossRef]
- Schlaeppi, K.; Bulgarelli, D. The plant microbiome at work. Mol. Plant. Microbe. Interact. 2015, 28, 212–217. [Google Scholar] [CrossRef] [PubMed]
- Hardoim, P.R.; van Overbeek, L.S.; Elsas, J.D. van Properties of bacterial endophytes and their proposed role in plant growth. Trends Microbiol. 2008, 16, 463–471. [Google Scholar] [CrossRef] [PubMed]
- Vandenkoornhuyse, P.; Baldauf, S.L.; Leyval, C.; Straczek, J.; Young, J.P.W. Extensive fungal diversity in plant roots. Science 2002, 295, 2051. [Google Scholar] [CrossRef]
- Shi, S.; Nuccio, E.E.; Shi, Z.J.; He, Z.; Zhou, J.; Firestone, M.K. The interconnected rhizosphere: High network complexity dominates rhizosphere assemblages. Ecol. Lett. 2016, 19, 926–936. [Google Scholar] [CrossRef]
- Bulgarelli, D.; Schlaeppi, K.; Spaepen, S.; Van Themaat, E.V.L.; Schulze-Lefert, P. Structure and functions of the bacterial microbiota of plants. Annu. Rev. Plant Biol. 2013, 64, 807–838. [Google Scholar] [CrossRef]
- Hacquard, S.; Garrido-Oter, R.; González, A.; Spaepen, S.; Ackermann, G.; Lebeis, S.; McHardy, A.C.; Dangl, J.L.; Knight, R.; Ley, R.; et al. Microbiota and host nutrition across plant and animal kingdoms. Cell Host Microbe 2015, 17, 603–616. [Google Scholar] [CrossRef]
- Tian, T.; Reverdy, A.; She, Q.; Sun, B.; Chai, Y. The role of rhizodeposits in shaping rhizomicrobiome. Environ. Microbiol. Rep. 2020, 12, 160–172. [Google Scholar] [CrossRef]
- Santos, L.F.; Olivares, F.L. Plant microbiome structure and benefits for sustainable agriculture. Curr. Plant Biol. 2021, 26, 100198. [Google Scholar] [CrossRef]
- Parniske, M. Arbuscular mycorrhiza: The mother of plant root endosymbioses. Nat. Rev. Microbiol. 2008, 6, 763–775. [Google Scholar] [CrossRef]
- Ikeda, S.; Okubo, T.; Anda, M.; Nakashita, H.; Yasuda, M.; Sato, S.; Kaneko, T.; Tabata, S.; Eda, S.; Momiyama, A.; et al. Community- and genome-based views of plant-associated bacteria: Plant-bacterial interactions in soybean and rice. Plant Cell Physiol. 2010, 51, 1398–1410. [Google Scholar] [CrossRef]
- Vandana, U.K.; Chopra, A.; Bhattacharjee, S.; Mazumder, P.B. Microbial Biofertilizer: A Potential Tool for Sustainable Agriculture; Panpatte, D.G., Jhala, Y.K., Vyas, R.V., Shelat, H.N., Eds.; Springer: Singapore, 2017; Volume 1, ISBN 9789811062407. [Google Scholar]
- Muresu, R.; Porceddu, A.; Concheri, G.; Stevanato, P.; Squartini, A. Legumes of the Sardinia Island: Knowledge on Symbiotic and Endophytic Bacteria and Interactive Software Tool for Plant Species Determination. Plants 2022, 11, 1521. [Google Scholar] [CrossRef]
- Soares, R.; Trejo, J.; Lorite, M.J.; Figueira, E.; Sanjuán, J.; E Castro, I.V. Diversity, Phylogeny and Plant Growth Promotion Traits of Nodule Associated Bacteria Isolated from Lotus parviflorus. Microorganisms 2020, 8, 499. [Google Scholar] [CrossRef]
- Pang, J.; Palmer, M.; Sun, H.J.; Seymour, C.O.; Zhang, L.; Hedlund, B.P.; Zeng, F. Diversity of Root Nodule-Associated Bacteria of Diverse Legumes Along an Elevation Gradient in the Kunlun Mountains, China. Front. Microbiol. 2021, 12, 168. [Google Scholar] [CrossRef]
- Stagnari, F.; Maggio, A.; Galieni, A.; Pisante, M. Multiple benefits of legumes for agriculture sustainability: An overview. Chem. Biol. Technol. Agric. 2017, 4, 1–13. [Google Scholar] [CrossRef]
- Foyer, C.H.; Lam, H.M.; Nguyen, H.T.; Siddique, K.H.M.; Varshney, R.K.; Colmer, T.D.; Cowling, W.; Bramley, H.; Mori, T.A.; Hodgson, J.M.; et al. Neglecting legumes has compromised human health and sustainable food production. Nat. Plants 2016, 2, 1–10. [Google Scholar] [CrossRef]
- Peoples, M.B.; Brockwell, J.; Herridge, D.F.; Rochester, I.J.; Alves, B.J.R.; Urquiaga, S.; Boddey, R.M.; Dakora, F.D.; Bhattarai, S.; Maskey, S.L.; et al. The contributions of nitrogen-fixing crop legumes to the productivity of agricultural systems. Symbiosis 2009, 48, 1–17. [Google Scholar] [CrossRef]
- Graham, P.H.; Vance, C.P. Legumes: Importance and constraints to greater use. Plant Physiol. 2003, 131, 872–877. [Google Scholar] [CrossRef]
- Głodowska, M.; Wozniak, M. Changes in Soil Microbial Activity and Community Composition as a Result of Selected Agricultural Practices. Agric. Sci. 2019, 10, 330–351. [Google Scholar] [CrossRef][Green Version]
- Cassman, K.G. Nitrogen fixation in tropical cropping systems. F. Crop. Res. 1993, 34, 230–232. [Google Scholar] [CrossRef]
- Giller, K.E.; Cadisch, G. Future benefits from biological nitrogen fixation: An ecological approach to agriculture. In Management of Biological Nitrogen Fixation for the Development of More Productive and Sustainable Agricultural Systems; Springer: Dordrecht, Switzerland, 1995; pp. 255–277. [Google Scholar]
- Weese, D.J.; Heath, K.D.; Dentinger, B.T.M.; Lau, J.A. Long-term nitrogen addition causes the evolution of less-cooperative mutualists. Evolution 2015, 69, 631–642. [Google Scholar] [CrossRef] [PubMed]
- Epihov, D.Z.; Saltonstall, K.; Batterman, S.A.; Hedin, L.O.; Hall, J.S.; van Breugel, M.; Leake, J.R.; Beerling, D.J. Legume-microbiome interactions unlock mineral nutrients in regrowing tropical forests. Proc. Natl. Acad. Sci. USA 2021, 118, e2022241118. [Google Scholar] [CrossRef]
- Miranda-Sánchez, F.; Rivera, J.; Vinuesa, P. Diversity patterns of Rhizobiaceae communities inhabiting soils, root surfaces and nodules reveal a strong selection of rhizobial partners by legumes. Environ. Microbiol. 2016, 18, 2375–2391. [Google Scholar] [CrossRef]
- Sinkovič, L.; Pipan, B.; Vasić, M.; Antić, M.; Todorović, V.; Ivanovska, S.; Brezeanu, C.; Šuštar-Vozlič, J.; Meglič, V. Morpho-Agronomic characterisation of Runner Bean (Phaseolus coccineus L.) from South-Eastern Europe. Sustainability 2019, 11, 6165. [Google Scholar] [CrossRef]
- Schwember, A.R.; Carrasco, B.; Gepts, P. Unraveling agronomic and genetic aspects of runner bean (Phaseolus coccineus L.). Field Crop. Res. 2017, 206, 86–94. [Google Scholar] [CrossRef]
- Crivelli, A.J.; Catsadorakis, G. Lake Prespa, Northwestern Greece, 1st ed.; Springer: Dordrecht, Switzerland, 1997; ISBN 978-94-011-5180-1. [Google Scholar]
- Catsadorakis, G.; Malakou, M. Conservation and management issues of Prespa National Park. Hydrobiol. 1997, 351, 175–196. [Google Scholar]
- Lima, P.L.T.; Silva, M.L.N.; Curi, N.; Quinton, J. Soil loss by water erosion in areas under maize and jack beans intercropped and monocultures. Ciência e Agrotecnologia 2014, 38, 129–139. [Google Scholar] [CrossRef]
- Klindworth, A.; Pruesse, E.; Schweer, T.; Peplies, J.; Quast, C.; Horn, M.; Glöckner, F.O. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 2013, 41, e1. [Google Scholar] [CrossRef]
- Schloss, P.D.; Westcott, S.L.; Ryabin, T.; Hall, J.R.; Hartmann, M.; Hollister, E.B.; Lesniewski, R.A.; Oakley, B.B.; Parks, D.H.; Robinson, C.J.; et al. Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 2009, 75, 7537–7541. [Google Scholar] [CrossRef]
- Douglas, G.M.; Maffei, V.J.; Zaneveld, J.R.; Yurgel, S.N.; Brown, J.R.; Taylor, C.M.; Huttenhower, C.; Langille, M.G.I. PICRUSt2 for prediction of metagenome functions. Nat. Biotechnol. 2020, 38, 685–688. [Google Scholar] [CrossRef]
- Parks, D.H.; Tyson, G.W.; Hugenholtz, P.; Beiko, R.G. STAMP: Statistical analysis of taxonomic and functional profiles. Bioinformatics 2014, 30, 3123–3124. [Google Scholar] [CrossRef]
- Ondov, B.D.; Bergman, N.H.; Phillippy, A.M. Interactive metagenomic visualization in a Web browser. BMC Bioinform. 2011, 12, 1–10. [Google Scholar] [CrossRef]
- Aguilar-Paredes, A.; Valdés, G.; Nuti, M. Ecosystem functions of microbial consortia in sustainable agriculture. Agronomy 2020, 10, 1902. [Google Scholar] [CrossRef]
- Inbaraj, M.P. Plant-Microbe Interactions in Alleviating Abiotic Stress—A Mini Review. Front. Agron. 2021, 3, 1–11. [Google Scholar] [CrossRef]
- Bulgarelli, D.; Garrido-Oter, R.; Münch, P.C.; Weiman, A.; Dröge, J.; Pan, Y.; McHardy, A.C.; Schulze-Lefert, P. Structure and function of the bacterial root microbiota in wild and domesticated barley. Cell Host Microbe 2015, 17, 392–403. [Google Scholar] [CrossRef]
- Lauber, C.L.; Ramirez, K.S.; Aanderud, Z.; Lennon, J.; Fierer, N. Temporal variability in soil microbial communities across land-use types. ISME J. 2013, 7, 1641–1650. [Google Scholar] [CrossRef]
- Cardinale, M.; Grube, M.; Erlacher, A.; Quehenberger, J.; Berg, G. Bacterial networks and co-occurrence relationships in the lettuce root microbiota. Environ. Microbiol. 2015, 17, 239–252. [Google Scholar] [CrossRef]
- Sarawaneeyaruk, S.; Lorliam, W.; Krajangsang, S.; Pringsulaka, O. Enhancing plant growth under municipal wastewater irrigation by plant growth promoting rhizospheric Bacillus spp. J. King Saud Univ.—Sci. 2019, 31, 384–389. [Google Scholar] [CrossRef]
- Ueki, A.; Kaku, N.; Ueki, K. Role of anaerobic bacteria in biological soil disinfestation for elimination of soil-borne plant pathogens in agriculture. Appl. Microbiol. Biotechnol. 2018, 102, 6309–6318. [Google Scholar] [CrossRef] [PubMed]
- Figueiredo, G.G.O.; Lopes, V.R.; Romano, T.; Camara, M.C. Clostridium. Benef. Microbes Agro-Ecology; Academic Press: Amsterdam, The Netherlands, 2020; pp. 477–491. [Google Scholar]
- Attard, E.; Poly, F.; Commeaux, C.; Laurent, F.; Terada, A.; Smets, B.F.; Recous, S.; Roux, X. Le Shifts between Nitrospira- and Nitrobacter-like nitrite oxidizers underlie the response of soil potential nitrite oxidation to changes in tillage practices. Environ. Microbiol. 2010, 12, 315–326. [Google Scholar] [CrossRef] [PubMed]
- Carbonetto, B.; Rascovan, N.; Álvarez, R.; Mentaberry, A.; Vázquez, M.P. Structure, composition and metagenomic profile of soil microbiomes associated to agricultural land use and tillage systems in Argentine Pampas. PLoS ONE 2014, 9, e99949. [Google Scholar] [CrossRef] [PubMed]
- Vartoukian, S.R.; Palmer, R.M.; Wade, W.G. Strategies for culture of ‘unculturable’ bacteria. FEMS Microbiol. Lett. 2010, 309, 1–7. [Google Scholar] [CrossRef]
- Chaudhary, D.K.; Khulan, A.; Kim, J. Development of a novel cultivation technique for uncultured soil bacteria. Sci. Reports 2019, 9, 1–11. [Google Scholar] [CrossRef]
- Kalam, S.; Basu, A.; Ahmad, I.; Sayyed, R.Z.; El-Enshasy, H.A.; Dailin, D.J.; Suriani, N.L. Recent Understanding of Soil Acidobacteria and Their Ecological Significance: A Critical Review. Front. Microbiol. 2020, 11, 580024. [Google Scholar] [CrossRef]
- Zhou, Y.; Zhu, H.; Fu, S.; Yao, Q. Variation in soil microbial community structure associated with different legume species is greater than that associated with different grass species. Front. Microbiol. 2017, 8, 1–13. [Google Scholar] [CrossRef]
- Lipson, D.A. Relationships between temperature responses and bacterial community structure along seasonal and altitudinal gradients. FEMS Microbiol. Ecol. 2007, 59, 418–427. [Google Scholar] [CrossRef]
- Weir, B.S. The current taxonomy of rhizobia. Available online: https://www.rhizobia.co.nz/taxonomy/rhizobia (accessed on 2 September 2021).
- Sprent, J.I.; Ardley, J.; James, E.K. Biogeography of nodulated legumes and their nitrogen-fixing symbionts. New Phytol. 2017, 215, 40–56. [Google Scholar] [CrossRef]
- Ofek, M.; Hadar, Y.; Minz, D. Ecology of root colonizing Massilia (Oxalobacteraceae). PLoS ONE 2012, 7, e40117. [Google Scholar] [CrossRef]
- Rubiales, D.; Flores, F.; Emeran, A.A.; Kharrat, M.; Amri, M.; Rojas-Molina, M.M.; Sillero, J.C. Identification and multi-environment validation of resistance against broomrapes (Orobanche crenata and Orobanche foetida) in faba bean (Vicia faba). F. Crop. Res. 2014, 166, 58–65. [Google Scholar] [CrossRef]
- Hanada, S.; Pierson, B.K. The family chloroflexaceae. The prokaryotes 2006, 7, 815–842. [Google Scholar]
- Ward, N.L.; Challacombe, J.F.; Janssen, P.H.; Henrissat, B.; Coutinho, P.M.; Wu, M.; Xie, G.; Haft, D.H.; Sait, M.; Badger, J.; et al. Three genomes from the phylum Acidobacteria provide insight into the lifestyles of these microorganisms in soils. Appl. Environ. Microbiol. 2009, 75, 2046–2056. [Google Scholar] [CrossRef]
- Hausmann, B.; Pelikan, C.; Herbold, C.W.; Köstlbacher, S.; Albertsen, M.; Eichorst, S.A.; Glavina Del Rio, T.; Huemer, M.; Nielsen, P.H.; Rattei, T.; et al. Peatland Acidobacteria with a dissimilatory sulfur metabolism. ISME J. 2018, 12, 1729–1742. [Google Scholar] [CrossRef]
- Jiang, Y.; Li, S.; Li, R.; Zhang, J.; Liu, Y.; Lv, L.; Zhu, H.; Wu, W.; Li, W. Plant cultivars imprint the rhizosphere bacterial community composition and association networks. Soil Biol. Biochem. 2017, 109, 145–155. [Google Scholar] [CrossRef]
- Li, F.; Chen, L.; Zhang, J.; Yin, J.; Huang, S. Bacterial community structure after long-term organic and inorganic fertilization reveals important associations between soil nutrients and specific taxa involved in nutrient transformations. Front. Microbiol. 2017, 8, 187. [Google Scholar] [CrossRef]
- Da Rocha, U.N.; Plugge, C.M.; George, I.; Van Elsas, J.D.; Van Overbeek, L.S. The rhizosphere selects for particular groups of Acidobacteria and Verrucomicrobia. PLoS ONE 2013, 8, 16–20. [Google Scholar]
- Kielak, A.M.; Barreto, C.C.; Kowalchuk, G.A.; van Veen, J.A.; Kuramae, E.E. The ecology of Acidobacteria: Moving beyond genes and genomes. Front. Microbiol. 2016, 7, 1–16. [Google Scholar] [CrossRef]
- Pérez-Jaramillo, J.E.; Carrión, V.J.; Bosse, M.; Ferrão, L.F.V.; De Hollander, M.; Garcia, A.A.F.; Ramírez, C.A.; Mendes, R.; Raaijmakers, J.M. Linking rhizosphere microbiome composition of wild and domesticated Phaseolus vulgaris to genotypic and root phenotypic traits. ISME J. 2017, 11, 2244–2257. [Google Scholar] [CrossRef]
- Zhao, Z.B.; He, J.Z.; Quan, Z.; Wu, C.F.; Sheng, R.; Zhang, L.M.; Geisen, S. Fertilization changes soil microbiome functioning, especially phagotrophic protists. Soil Biol. Biochem. 2020, 148, 107863. [Google Scholar] [CrossRef]
- Stefan, L.; Hartmann, M.; Engbersen, N.; Six, J.; Schöb, C. Positive Effects of Crop Diversity on Productivity Driven by Changes in Soil Microbial Composition. Front. Microbiol. 2021, 12, 660749. [Google Scholar] [CrossRef]
- Yadav, S.K.; Soni, R.; Rajput, A.S. Role of Microbes in Organic Farming for Sustainable Agro-Ecosystem. In Microorganisms for Green Revolution; Springer: Singapore, 2018; pp. 241–252. ISBN 9789811071461. [Google Scholar]
- Raetz, C.R.H.; Guan, Z.; Ingram, B.O.; Six, D.A.; Song, F.; Wang, X.; Zhao, J. Discovery of new biosynthetic pathways: The lipid A story. J. Lipid Res. 2009, 50, 103–108. [Google Scholar] [CrossRef] [PubMed]
- Boersch, M.; Rudrawar, S.; Grant, G.; Zunk, M. Menaquinone biosynthesis inhibition: A review of advancements toward a new antibiotic mechanism. RSC Adv. 2018, 8, 5099–5105. [Google Scholar] [CrossRef] [PubMed]
- Johnston, J.M.; Bulloch, E.M. Advances in menaquinone biosynthesis: Sublocalisation and allosteric regulation. Curr. Opin. Struct. Biol. 2020, 65, 33–41. [Google Scholar] [CrossRef]
- Pelchovich, G.; Omer-Bendori, S.; Gophna, U. Menaquinone and Iron Are Essential for Complex Colony Development in Bacillus subtilis. PLoS ONE 2013, 8, e79488. [Google Scholar] [CrossRef]
- Farrand, S.K.; Taber, H.W. Changes in menaquinone concentration during growth and early sporulation in Bacillus subtilis. J. Bacteriol. 1974, 117, 324–326. [Google Scholar] [CrossRef]
- Wicaksono, W.A.; Cernava, T.; Berg, C.; Berg, G. Bog ecosystems as a playground for plant–microbe coevolution: Bryophytes and vascular plants harbour functionally adapted bacteria. Microbiome 2021, 9, 1–16. [Google Scholar] [CrossRef]
- Hale, M.B.; Blankenship, R.E.; Fuller, R.C. Menaquinone is the sole quinone in the facultatively aerobic green photosynthetic bacterium Chloroflexus aurantiacus. BBA - Bioenerg. 1983, 723, 376–382. [Google Scholar] [CrossRef]
- Kawasaki, A.; Dennis, P.G.; Forstner, C.; Raghavendra, A.K.H.; Mathesius, U.; Richardson, A.E.; Delhaize, E.; Gilliham, M.; Watt, M.; Ryan, P.R. Manipulating exudate composition from root apices shapes the microbiome throughout the root system. Plant Physiol. 2021, 187, 2279. [Google Scholar] [CrossRef]
- Dagorn, A.; Chapalain, A.; Mijouin, L.; Hillion, M.; Duclairoir-Poc, C.; Chevalier, S.; Taupin, L.; Orange, N.; Feuilloley, M.G.J. Effect of GABA, a bacterial metabolite, on pseudomonas fluorescens surface properties and Cytotoxicity. Int. J. Mol. Sci. 2013, 14, 12186–12204. [Google Scholar] [CrossRef]
- Ramesh, S.A.; Tyerman, S.D.; Gilliham, M.; Xu, B. γ-Aminobutyric acid (GABA) signalling in plants. Cell. Mol. Life Sci. 2017, 74, 1577–1603. [Google Scholar] [CrossRef]
- Li, L.; Dou, N.; Zhang, H.; Wu, C. The versatile GABA in plants. Plant Signal. Behav. 2021, 16, 1–22. [Google Scholar] [CrossRef]
- Wang, P.; Lopes, L.D.; Lopez-Guerrero, M.G.; van Dijk, K.; Alvarez, S.; Riethoven, J.-J.; Schachtman, D.P. Natural variation in root exudation of GABA and DIMBOA impacts the maize root endosphere and rhizosphere microbiomes. J. Exp. Bot. 2022, 73, 5052–5066. [Google Scholar] [CrossRef]
- Nascimento, F.X.; Hernández, A.G.; Glick, B.R.; Rossi, M.J. Plant growth-promoting activities and genomic analysis of the stress-resistant Bacillus megaterium STB1, a bacterium of agricultural and biotechnological interest. Biotechnol. Rep. 2020, 25, 1–9. [Google Scholar] [CrossRef]
- Lamont, J.R.; Wilkins, O.; Bywater-Ekegärd, M.; Smith, D.L. From yogurt to yield: Potential applications of lactic acid bacteria in plant production. Soil Biol. Biochem. 2017, 111, 1–9. [Google Scholar] [CrossRef]
- Raman, J.; Kim, J.S.; Choi, K.R.; Eun, H.; Yang, D.; Ko, Y.J.; Kim, S.J. Application of Lactic Acid Bacteria (LAB) in Sustainable Agriculture: Advantages and Limitations. Int. J. Mol. Sci. 2022, 23, 7784. [Google Scholar] [CrossRef]
- Somers, E.; Amke, A.; Croonenborghs, A.; van Overbeek, L.S.; Vanderleyden, J. Lactic acid bacteria in organic agricultural soils. In Proceedings of the Rhizosphere 2, Montpellier, France, 26–31 August 2007; pp. 26–31. [Google Scholar]
- Weber, M.; Fuchs, T.M. Metabolism in the Niche: A Large-Scale Genome-Based Survey Reveals Inositol Utilization To Be Widespread among Soil, Commensal, and Pathogenic Bacteria. Microbiol. Spectr. 2022, 10, e0201322. [Google Scholar] [CrossRef]
- Caspi, R.; Billington, R.; Keseler, I.M.; Kothari, A.; Krummenacker, M.; Midford, P.E.; Ong, W.K.; Paley, S.; Subhraveti, P.; Karp, P.D. The MetaCyc database of metabolic pathways and enzymes-a 2019 update. Nucleic Acids Res. 2020, 48, D453–D455. [Google Scholar] [CrossRef]
- Yoshida, K.I.; Yamaguchi, M.; Morinaga, T.; Kinehara, M.; Ikeuchi, M.; Ashida, H.; Fujita, Y. myo-inositol catabolism in Bacillus subtilis. J. Biol. Chem. 2008, 283, 10415–10424. [Google Scholar] [CrossRef]
- Pini, F.; East, A.K.; Appia-Ayme, C.; Tomek, J.; Karunakaran, R.; Mendoza-Suárez, M.; Edwards, A.; Terpolilli, J.J.; Roworth, J.; Downie, J.A.; et al. Bacterial Biosensors for in Vivo Spatiotemporal Mapping of Root Secretion. Plant Physiol. 2017, 174, 1289–1306. [Google Scholar] [CrossRef] [PubMed]
- Vílchez, J.I.; Yang, Y.; He, D.; Zi, H.; Peng, L.; Lv, S.; Kaushal, R.; Wang, W.; Huang, W.; Liu, R.; et al. DNA demethylases are required for myo-inositol-mediated mutualism between plants and beneficial rhizobacteria. Nat. Plants 2020, 6, 983–995. [Google Scholar] [CrossRef]
- Chen, X.; Schreiber, K.; Appel, J.; Makowka, A.; Fähnrich, B.; Roettger, M.; Hajirezaei, M.R.; Sönnichsen, F.D.; Schönheit, P.; Martin, W.F.; et al. The Entner-Doudoroff pathway is an overlooked glycolytic route in cyanobacteria and plants. Proc. Natl. Acad. Sci. USA 2016, 113, 5441–5446. [Google Scholar] [CrossRef] [PubMed]
- Fierer, N.; Ladau, J.; Clemente, J.C.; Leff, J.W.; Owens, S.M.; Pollard, K.S.; Knight, R.; Gilbert, J.A.; McCulley, R.L. Reconstructing the microbial diversity and function of pre-agricultural tallgrass prairie soils in the United States. Science 2013, 342, 621–624. [Google Scholar] [CrossRef] [PubMed]
| Phylum (%) | Season | ||
|---|---|---|---|
| Spring_BC | Summer_DC | Autumn_AC | |
| Proteobacteria | 28.96 | 30.04 | 28.49 |
| Acidobacteria | 16.91 | 12.41 | 15.29 |
| Actinobacteria | 6.26 | 14.37 | 11.25 |
| Planctomycetes | 9.50 | 10.76 | 10.71 |
| Bacteroidetes | 8.68 | 7.67 | 7.95 |
| Verrucomicrobia | 6.64 | 3.94 | 5.24 |
| Bacteria_unclassified | 5.75 | 4.89 | 4.74 |
| Firmicutes | 1.75 | 3.67 | 2.80 |
| Gemmatimonadetes | 2.83 | 2.30 | 2.79 |
| Chloroflexi | 1.21 | 1.68 | 1.21 |
| Cyanobacteria | - | 1.22 | 0.85 |
| Patescibacteria | 1.94 | - | 0.69 |
| Nitrospirae | 1.35 | - | 0.54 |
| Total % | 91.78 | 92.95 | 92.55 |
| Phylum | Class (%) | Season | ||
|---|---|---|---|---|
| Spring_BC | Summer_DC | Autumn_AC | ||
| Acidobacteria | Acidobacteriia | 2.27 | 1.77 | 2.13 |
| Blastocatellia (Subgroup_4) | 4.95 | 4.49 | 5.21 | |
| Holophagae | 1.20 | 0.37 | - | |
| Subgroup_6 | 9.29 | 6.02 | 7.95 | |
| Actinobacteria | Acidimicrobiia | 1.64 | 2.26 | 2.01 |
| Actinobacteria | 3.09 | 9.35 | 6.99 | |
| Rubrobacteria | - | - | 1.07 | |
| Thermoleophilia | 1.52 | 2.75 | 2.14 | |
| Bacteria_unclassified | Bacteria_unclassified | 5.75 | 4.89 | 4.74 |
| Bacteroidetes | Bacteroidia | 8.68 | 7.66 | 7.95 |
| Chloroflexi | Anaerolineae | 0.94 | 0.92 | 0.81 |
| Chloroflexia | 0.41 | 1.60 | 1.34 | |
| KD4-96 | 0.61 | 0.78 | 0.23 | |
| Cyanobacteria | Oxyphotobacteria | - | 1.22 | 0.85 |
| Firmicutes | Bacilli | 1.75 | 3.55 | 2.68 |
| Clostridia | - | 1.05 | 1.08 | |
| Gemmatimonadetes | Gemmatimonadetes | 2.83 | 2.30 | 2.79 |
| Nitrospirae | Nitrospira | 1.35 | - | 0.54 |
| Patescibacteria | Parcubacteria | 1.94 | - | 0.29 |
| Saccharimonadia | - | - | 1.22 | |
| Planctomycetes | Phycisphaerae | 5.03 | 7.00 | 6.71 |
| Planctomycetacia | 4.47 | 3.75 | 4.00 | |
| Proteobacteria | Alphaproteobacteria | 15.13 | 18.79 | 16.95 |
| Deltaproteobacteria | 3.55 | 2.71 | 2.71 | |
| Gammaproteobacteria | 10.28 | 8.54 | 8.83 | |
| Verrucomicrobia | Verrucomicrobiae | 6.64 | 3.94 | 5.24 |
| Total % | 93.32 | 95.71 | 96.46 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Stavridou, E.; Karamichali, I.; Lagiotis, G.; Patsea, E.; Osathanunkul, M.; Madesis, P. Seasonal Shifts in Soil Microbiome Structure Are Associated with the Cultivation of the Local Runner Bean Variety around the Lake Mikri Prespa. Biology 2022, 11, 1595. https://doi.org/10.3390/biology11111595
Stavridou E, Karamichali I, Lagiotis G, Patsea E, Osathanunkul M, Madesis P. Seasonal Shifts in Soil Microbiome Structure Are Associated with the Cultivation of the Local Runner Bean Variety around the Lake Mikri Prespa. Biology. 2022; 11(11):1595. https://doi.org/10.3390/biology11111595
Chicago/Turabian StyleStavridou, Evangelia, Ioanna Karamichali, Georgios Lagiotis, Elena Patsea, Maslin Osathanunkul, and Panagiotis Madesis. 2022. "Seasonal Shifts in Soil Microbiome Structure Are Associated with the Cultivation of the Local Runner Bean Variety around the Lake Mikri Prespa" Biology 11, no. 11: 1595. https://doi.org/10.3390/biology11111595
APA StyleStavridou, E., Karamichali, I., Lagiotis, G., Patsea, E., Osathanunkul, M., & Madesis, P. (2022). Seasonal Shifts in Soil Microbiome Structure Are Associated with the Cultivation of the Local Runner Bean Variety around the Lake Mikri Prespa. Biology, 11(11), 1595. https://doi.org/10.3390/biology11111595







