Canadian Consensus Recommendations on the Management of KRAS G12C-Mutated NSCLC
Abstract
1. Introduction
2. KRAS Mutations: Biology and Signaling Pathways
3. Epidemiology, Clinical Features, and Prognostic Implications
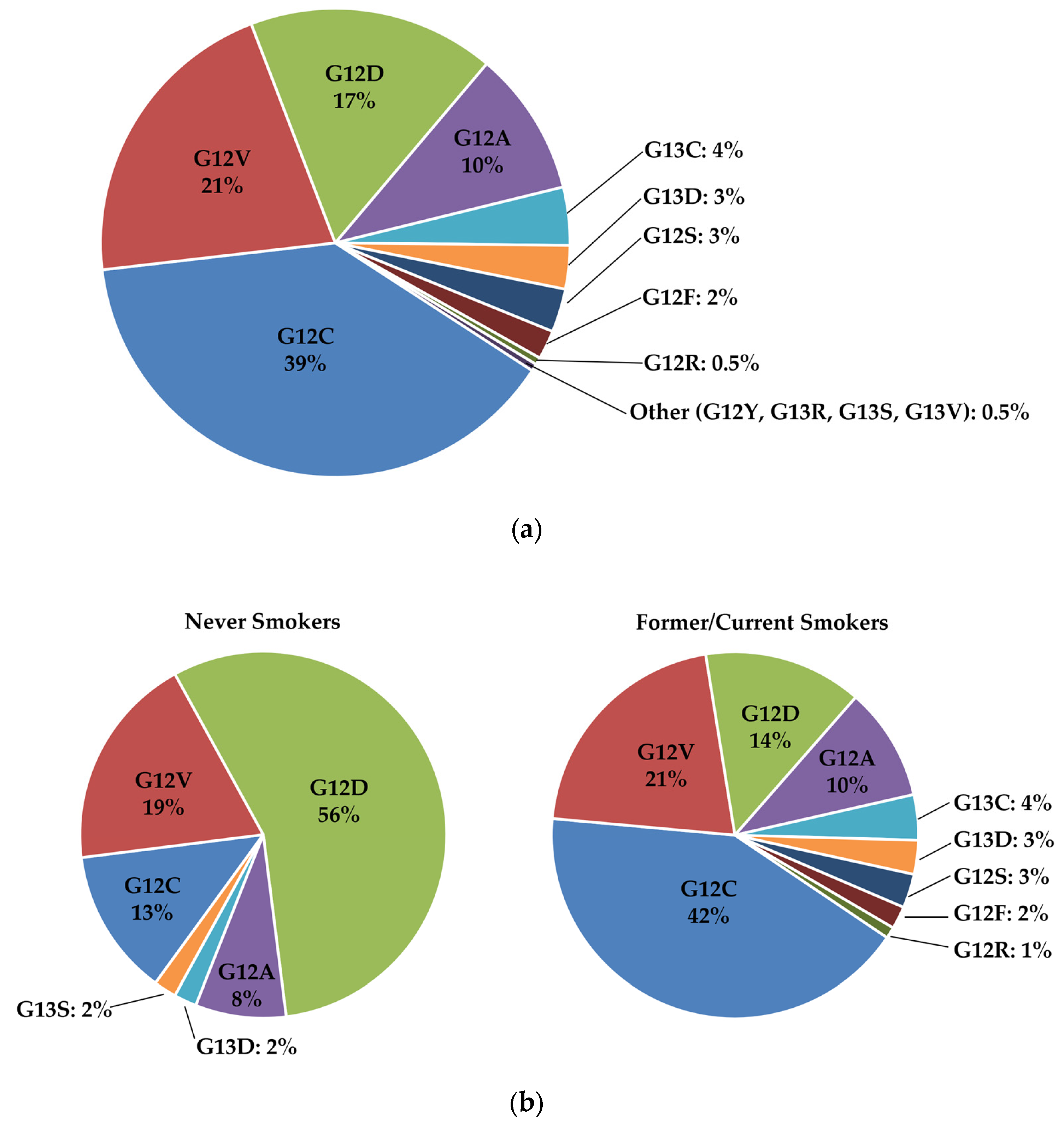
3.1. Overview
3.2. Co-Mutations and Their Impact on Treatment Outcomes
4. Identifying Patients with KRAS G12C
4.1. Testing Strategies
4.2. Testing for KRAS G12C
4.3. Should Liquid Biopsy Be Integrated into KRAS Testing in Patients with NSCLC?
4.4. TAT and Reporting of Biomarker Test Results
4.5. Recommendations
- Testing for KRAS G12C should be performed as part of a comprehensive panel that includes current standard-of-care biomarkers as summarized by international guidelines. Single-gene strategies for KRAS are not recommended.
- Although KRAS G12C status is currently required for patients with stage III/IV NSCLC, KRAS G12C could be included in the NGS panel and reflex biomarker testing for KRAS G12C could be initiated by the pathologist at the time of initial diagnosis in all patients diagnosed with stage IB-IV non-squamous NSCLC, if NGS is performed.
- The panel recognizes the lower frequency of KRAS G12C and other actionable alterations in squamous cell carcinoma and, in light of current evidence, suggests broad molecular testing in patients with advanced disease in which systemic therapy is being considered if resources are available and there is no significant additional burden on the molecular pathology laboratory.
- TP53, STK11, and KEAP1 are evolving biomarkers that may have implications in patients with KRAS mutations; therefore, molecular laboratories could consider preparing to include these alterations in NGS panels as they may become relevant for therapeutic decision making in the near future as more supporting data becomes available.
- Liquid biopsy should be considered when tissue biopsy is unavailable or inadequate for molecular testing, when invasive procedures for tissue procurement are contraindicated, or when urgent treatment decisions are required and delays are expected with tissue testing.
- Liquid biopsy results should be interpreted by qualified professionals in a patient-specific context. Providers should be aware of the limitations and use caution when interpreting standalone liquid biopsy results, as liquid biopsy is not a diagnosis in and of itself but rather augments our understanding of a histologically or cytologically diagnosed malignancy.
- Negative liquid biopsy results do not mean the absence of the target and tissue testing is recommended if liquid biopsy does not detect a mutation indicating the presence of tumour-derived DNA.
- The maximum acceptable TAT (from the acquisition of tissue to the oncologist having the report) for all biomarkers should not exceed 21 calendar days. Accelerated testing should be available for certain specific situations.
- Biomarker test results should be compiled and ideally reported in a single comprehensive biomarker report by the pathologist that includes information on PD-L1 expression.
5. Therapeutic Approaches for Patients with Advanced NSCLC Harbouring KRAS Mutations
5.1. Questions
- What is the preferred first-line therapy for treatment-naïve patients with KRAS mutations?
- What are the preferred subsequent lines of therapy for patients with KRAS mutations?
5.2. ICIs in Patients with KRAS Mutations
5.3. Recommendations: First-Line Treatment of Advanced KRAS G12C-Mutated NSCLC
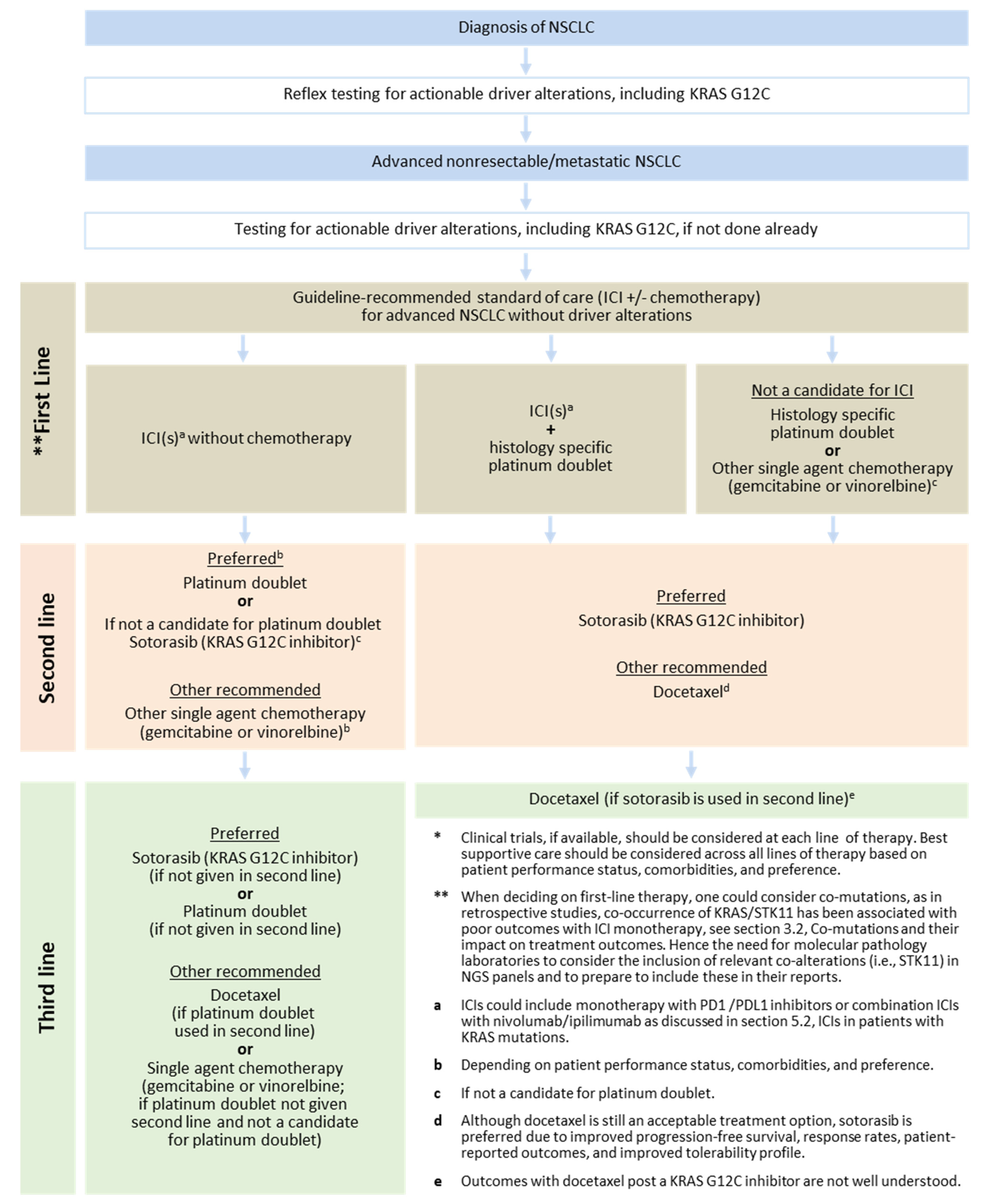
- Patients with advanced KRAS G12C-mutated NSCLC that are eligible for treatment should be offered guideline-recommended standard of care for advanced NSCLC without driver alterations (ICI(s) +/− chemotherapy), Figure 2.
- The impact of co-mutations, including TP53, STK11, and KEAP1, is currently under investigation and the presence of these mutations should not impact first-line treatment decisions at present.
5.4. Direct Inhibition of KRAS
5.4.1. Overview
5.4.2. Sotorasib
5.4.3. Adagrasib
5.5. Recommendations: Second- and Subsequent-Line Treatment of Advanced KRASs G12C-Mutated NSCLC
- For patients with locally advanced or metastatic KRAS G12C-mutated NSCLC who have progressed on ICI(s) and/or chemotherapy, the panel recommends sotorasib over docetaxel based on improved progression-free survival, response rates, patient-reported outcomes, and improved tolerability profile.
- It is reasonable to offer sotorasib as second-line therapy to patients who progressed on only having either monotherapy ICI that are not candidates for platinum doublet or patients with a contraindication to ICIs that received chemotherapy in the first-line setting
- Given that sotorasib is approved by Health Canada and has phase 3 trial data, the panel recommends the use of sotorasib in the current treatment algorithm of advanced KRAS G12C mutated NSCLC. However, the panel recognizes that the use of adagrasib or a clinical trial with another KRAS G12C inhibitor (monotherapy or in combination) is an acceptable treatment option in this line of therapy.
- Sotorasib and adagrasib have a similar mechanism of action; it is not recommended to switch between these agents at the time of progression.
5.6. What Are the Treatment Options for Metastatic NSCLC Patients with KRAS G12C and Brain Metastases?
5.6.1. Overview
5.6.2. Recommendations
- KRAS G12C inhibitors, sotorasib and adagrasib, appear to have activity in brain metastases, but there is insufficient evidence to recommend one agent over the other in patients with brain metastases.
- Currently, there is insufficient evidence to make recommendations regarding the sequencing of KRAS G12C inhibitors and radiation in patients with brain metastases.
- Due to uncertainties related to potential interaction with radiation therapy, the panel does not recommend concurrent use of KRAS G12C inhibitors with stereotactic radiosurgery (SRS) or whole-brain radiation therapy (WBRT).
5.7. What Are the Common Toxicities Associated with KRAS G12C Inhibitors and How Should They Be Managed?
5.7.1. Overview
5.7.2. Recommendations
- As KRAS G12C inhibitors can cause hepatotoxicity, regular monitoring of liver function (ALT, AST, and total bilirubin) is recommended. Clinicians should follow the drug’s product monograph for dose reductions, interruptions, or permanent discontinuations.
- Corticosteroids are recommended in patients presenting with grade 2–4 hepatotoxicity.
- The expert panel agreed that a low dose of prednisone might be considered for preventing another occurrence of hepatotoxicity upon re-challenge with a KRAS inhibitor.
- Clinicians should be aware of an increased risk of AEs, in particular hepatotoxicity, when initiating sotorasib post-ICI.
- Sequencing of KRAS G12C inhibitors, ICI, and the need for and duration of washout period between these agents is yet to be determined. Physicians should consider and weigh the risk of hepatotoxicity vs. the risk of disease progression.
- Caution is required in patients initiated on a KRAS G12C inhibitor within 3 months post-ICI. These patients may require more frequent monitoring of liver function tests.
5.8. What Are the Potential Strategies to Overcome Treatment Resistance in Patients with KRAS G12C?
5.8.1. Overview
5.8.2. Recommendations
- The panel strongly encourages the enrolment of patients with KRAS G12C mutated NSCLC into clinical trials with KRAS inhibitors.
- Currently repeated biopsy (liquid or tissue) in patients progressing on or after KRAS G12C inhibitors is not recommended outside clinical trial settings.
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Siegel, R.L.; Miller, K.D.; Fuchs, H.E.; Jemal, A. Cancer Statistics, 2021. CA Cancer J. Clin. 2021, 71, 7–33. [Google Scholar] [CrossRef] [PubMed]
- Veluswamy, R.; Mack, P.C.; Houldsworth, J.; Elkhouly, E.; Hirsch, F.R. KRAS G12C-Mutant Non-Small Cell Lung Cancer: Biology, Developmental Therapeutics, and Molecular Testing. J. Mol. Diagn. 2021, 23, 507–520. [Google Scholar] [CrossRef] [PubMed]
- Amgen Canada Lumakras Product Monograph (September 2021; Submission Control Number: 248435). Available online: https://www.amgen.ca/-/media/Themes/CorporateAffairs/amgen-ca/amgen-ca/documents/products/en/lumakras_pm.pdf (accessed on 4 May 2023).
- Skoulidis, F.; Li, B.T.; Dy, G.K.; Price, T.J.; Falchook, G.S.; Wolf, J.; Italiano, A.; Schuler, M.; Borghaei, H.; Barlesi, F.; et al. Sotorasib for Lung Cancers with KRAS p.G12C Mutation. N. Engl. J. Med. 2021, 384, 2371–2381. [Google Scholar] [CrossRef] [PubMed]
- de Langen, A.J.; Johnson, M.L.; Mazieres, J.; Dingemans, A.C.; Mountzios, G.; Pless, M.; Wolf, J.; Schuler, M.; Lena, H.; Skoulidis, F.; et al. Sotorasib versus docetaxel for previously treated non-small-cell lung cancer with KRASG12C mutation: A randomised, open-label, phase 3 trial. Lancet 2023, 401, 733–746. [Google Scholar] [CrossRef]
- Mok, T.S.K.; Lawler, W.E.; Shum, M.K.; Shaker, R.; Dakhil, S.R.; Spira, A.I.; Barlesi, F.; Reck, M.; Garassino, M.C.; Spigel, D.R.; et al. KRYSTAL-12: A randomized phase 3 study of adagrasib (MRTX849) versus docetaxel in patients (pts) with previously treated non-small-cell lung cancer (NSCLC) with KRASG12C mutation. J. Clin. Oncol. 2021, 39, TPS9129. [Google Scholar] [CrossRef]
- Garrido, P.; Olmedo, M.E.; Gómez, A.; Paz Ares, L.; López-Ríos, F.; Rosa-Rosa, J.M.; Palacios, J. Treating KRAS-mutant NSCLC: Latest evidence and clinical consequences. Ther. Adv. Med. Oncol. 2017, 9, 589–597. [Google Scholar] [CrossRef]
- Corral de la Fuente, E.; Olmedo Garcia, M.E.; Gomez Rueda, A.; Lage, Y.; Garrido, P. Targeting KRAS in Non-Small Cell Lung Cancer. Front. Oncol. 2022, 11, 792635. [Google Scholar] [CrossRef]
- Hobbs, G.A.; Der, C.J.; Rossman, K.L. RAS isoforms and mutations in cancer at a glance. J. Cell Sci. 2016, 129, 1287–1292. [Google Scholar] [CrossRef]
- Prior, I.A.; Lewis, P.D.; Mattos, C. A comprehensive survey of Ras mutations in cancer. Cancer Res. 2012, 72, 2457–2467. [Google Scholar] [CrossRef]
- Kato, S.; Fujiwara, Y.; Hong, D.S. Targeting KRAS: Crossroads of Signaling and Immune Inhibition. J. Immunother. Precis Oncol. 2022, 5, 68–78. [Google Scholar] [CrossRef]
- Moore, A.R.; Rosenberg, S.C.; McCormick, F.; Malek, S. RAS-targeted therapies. Nat. Rev. Drug Discov. 2021, 19, 533–552. [Google Scholar] [CrossRef] [PubMed]
- Ye, J.; Lin, M.; Zhang, C.; Zhu, X.; Li, S.; Liu, H.; Yin, J.; Yu, H.; Zhu, K. Tissue gene mutation profiles in patients with colorectal cancer and their clinical implications. Biomed. Rep. 2020, 13, 43–48. [Google Scholar] [CrossRef]
- Cox, A.D.; Fesik, S.W.; Kimmelman, A.C.; Luo, J.; Der, C.J. Drugging the undruggable RAS: Mission possible? Nat. Rev. Drug Discov. 2014, 13, 828–851. [Google Scholar] [CrossRef] [PubMed]
- Ritu, K.; Kumar, P.; Singh, A.; Nupur, K.; Spalgias, S.; Mrigpuri, P.; Rajkumar. Untangling the KRAS mutated lung cancer subsets and its therapeutic implications. Mol. Biomed. 2021, 2, 40. [Google Scholar] [CrossRef] [PubMed]
- Friedlaender, A.; Drilon, A.; Weiss, G.J.; Banna, G.L.; Addeo, A. KRAS as a druggable target in NSCLC: Rising like a phoenix after decades of development failures. Cancer Treat. Rev. 2020, 85, 101978. [Google Scholar] [CrossRef]
- Yu, H.A.; Sima, C.S.; Shen, R.; Kass, S.; Gainor, J.; Shaw, A.; Hames, M.; Iams, W.; Aston, J.; Lovly, C.M.; et al. Prognostic impact of KRAS mutation subtypes in 677 patients with metastatic lung adenocarcinomas. J. Thorac. Oncol. 2015, 10, 431–437. [Google Scholar] [CrossRef]
- Salgia, R.; Pharaon, R.; Mambetsariev, I.; Nam, A.; Sattler, M. The improbable targeted therapy: KRAS as an emerging target in non-small cell lung cancer (NSCLC). Cell Rep. Med. 2021, 2, 100186. [Google Scholar] [CrossRef]
- Jordan, E.J.; Kim, H.R.; Arcila, M.E.; Barron, D.; Chakravarty, D.; Gao, J.; Chang, M.T.; Ni, A.; Kundra, R.; Jonsson, P.; et al. Prospective Comprehensive Molecular Characterization of Lung Adenocarcinomas for Efficient Patient Matching to Approved and Emerging Therapies. Cancer Discov. 2017, 7, 596–609. [Google Scholar] [CrossRef]
- Nusrat, M.; Roszik, J.; Holla, J.; Hong, M.G.; Coker, O.; Johnson, B.; Ahnert, J.R.; Janku, F.; Kopetz, S.; Shaw, K.R.; et al. Therapeutic vulnerabilities among KRAS G12C mutant (mut) advanced cancers based on co-alteration (co-alt) patterns. J. Clin. Oncol. 2020, 38 (Suppl. S15), 3625. [Google Scholar] [CrossRef]
- Roberts, P.J.; Stinchcombe, T.E.; Der, C.J.; Socinski, M.A. Personalized medicine in non-small-cell lung cancer: Is KRAS a useful marker in selecting patients for epidermal growth factor receptor-targeted therapy? J. Clin. Oncol. 2010, 28, 4769–4777. [Google Scholar] [CrossRef]
- Aran, V.; Zalis, M.; Montella, T.; de Sousa, C.A.M.; Ferrari, B.L.; Gil Ferreira, C. Evaluation of KRAS Concomitant Mutations in Advanced Lung Adenocarcinoma Patients. Medicina 2021, 57, 1039. [Google Scholar] [CrossRef] [PubMed]
- Garcia, B.N.C.; van Kempen, L.C.; Kuijpers, C.C.H.J.; Schuuring, E.; Willems, S.M.; van der Wekken, A.J. Prevalence of KRAS p.(G12C) in stage IV NSCLC patients in the Netherlands; a nation-wide retrospective cohort study. Lung Cancer 2022, 167, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Scheffler, M.; Ihle, M.A.; Hein, R.; Merkelbach-Bruse, S.; Scheel, A.H.; Siemanowski, J.; Brägelmann, J.; Kron, A.; Abedpour, N.; Ueckeroth, F.; et al. K-ras Mutation Subtypes in NSCLC and Associated Co-occuring Mutations in Other Oncogenic Pathways. J. Thorac. Oncol. 2019, 14, 606–616. [Google Scholar] [CrossRef] [PubMed]
- Arbour, K.C.; Jordan, E.; Kim, H.R.; Dienstag, J.; Yu, H.A.; Sanchez-Vega, F.; Lito, P.; Berger, M.; Solit, D.B.; Hellmann, M.; et al. Effects of Co-Occurring Genomic Alterations on Outcomes in Patients With KRAS-Mutant Non-Small Cell Lung Cancer. Clin. Cancer Res. 2018, 24, 334–340. [Google Scholar] [CrossRef] [PubMed]
- Boudeau, J.; Sapkota, G.; Alessi, D.R. LKB1, a protein kinase regulating cell proliferation and polarity. FEBS Lett. 2003, 546, 159–165. [Google Scholar] [CrossRef] [PubMed]
- Sitthideatphaiboon, P.; Galan-Cobo, A.; Negrao, M.V.; Qu, X.; Poteete, A.; Zhang, F.; Liu, D.D.; Lewis, W.E.; Kemp, H.N.; Lewis, J.; et al. STK11/LKB1 Mutations in NSCLC Are Associated with KEAP1/NRF2-Dependent Radiotherapy Resistance Targetable by Glutaminase Inhibition. Clin. Cancer Res. 2021, 27, 1720–1733. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, A.; Kang, M.I.; Okawa, H.; Ohtsuji, M.; Zenke, Y.; Chiba, T.; Igarashi, K.; Yamamoto, M. Oxidative stress sensor Keap1 functions as an adaptor for Cul3-based E3 ligase to regulate proteasomal degradation of Nrf2. Mol. Cell Biol. 2004, 24, 7130–7139. [Google Scholar] [CrossRef] [PubMed]
- Ferrer, I.; Zugazagoitia, J.; Herbertz, S.; John, W.; Paz-Ares, L.; Schmid-Bindert, G. KRAS-Mutant Non-Small Cell Lung Cancer: From Biology to Therapy. Lung Cancer 2018, 124, 53–64. [Google Scholar] [CrossRef]
- Skoulidis, F.; Heymach, J.V. Co-occurring genomic alterations in non-small-cell lung cancer biology and therapy. Nat. Rev. Cancer 2019, 19, 495–509. [Google Scholar] [CrossRef]
- Romero, R.; Sayin, V.I.; Davidson, S.M.; Bauer, M.R.; Singh, S.X.; LeBoeuf, S.E.; Karakousi, T.R.; Ellis, D.C.; Bhutkar, A.; Sánchez-Rivera, F.; et al. Keap1 loss promotes Kras-driven lung cancer and results in dependence on glutaminolysis. Nat. Med. 2017, 23, 1362–1368. [Google Scholar] [CrossRef]
- Zhao, J.; Han, Y.; Li, J.; Chai, R.; Bai, C. Prognostic value of KRAS/TP53/PIK3CA in non-small cell lung cancer. Oncol. Lett. 2019, 17, 3233–3240. [Google Scholar] [CrossRef]
- Kansanen, E.; Kuosmanen, S.M.; Leinonen, H.; Levonen, A.L. The Keap1-Nrf2 pathway: Mechanisms of activation and dysregulation in cancer. Redox. Biol. 2013, 1, 45–49. [Google Scholar] [CrossRef]
- Facchinetti, F.; Bluthgen, M.V.; Tergemina-Clain, G.; Faivre, L.; Pignon, J.P.; Planchard, D.; Remon, J.; Soria, J.C.; Lacroix, L.; Besse, B. LKB1/STK11 mutations in non-small cell lung cancer patients: Descriptive analysis and prognostic value. Lung Cancer 2017, 112, 62–68. [Google Scholar] [CrossRef]
- Koyama, S.; Akbay, E.A.; Li, Y.Y.; Aref, A.R.; Skoulidis, F.; Herter-Sprie, G.S.; Buczkowski, K.A.; Liu, Y.; Awad, M.M.; Denning, W.L.; et al. STK11/LKB1 Deficiency Promotes Neutrophil Recruitment and Proinflammatory Cytokine Production to Suppress T-cell Activity in the Lung Tumor Microenvironment. Cancer Res. 2016, 76, 999–1008. [Google Scholar] [CrossRef]
- Cekani, E.; Epistolio, S.; Dazio, G.; Cefalì, M.; Wannesson, L.; Frattini, M.; Froesch, P. Molecular Biology and Therapeutic Perspectives for K-Ras Mutant Non-Small Cell Lung Cancers. Cancers 2022, 14, 4103. [Google Scholar] [CrossRef]
- Jia, Y.; Jiang, T.; Li, X.; Zhao, C.; Zhang, L.; Zhao, S.; Liu, X.; Qiao, M.; Luo, J.; Shi, J.; et al. Characterization of distinct types of KRAS mutation and its impact on first-line platinum-based chemotherapy in Chinese patients with advanced non-small cell lung cancer. Oncol. Lett. 2017, 14, 6525–6532. [Google Scholar] [CrossRef]
- Ma, Z.; Zhang, Y.; Deng, C.; Fu, F.; Deng, L.; Li, Y.; Chen, H. The prognostic value of Kirsten rat sarcoma viral oncogene homolog mutations in resected lung adenocarcinoma differs according to clinical features. J. Thorac. Cardiovasc. Surg. 2022, 163, e73–e85. [Google Scholar] [CrossRef]
- Si, X.; Pan, R.; Ma, S.; Li, L.; Liang, L.; Zhang, P.; Chu, Y.; Wang, H.; Wang, M.; Zhang, X.; et al. Genomic characteristics of driver genes in Chinese patients with non-small cell lung cancer. Thorac. Cancer 2021, 12, 357–363. [Google Scholar] [CrossRef]
- Das, B.R.; Bhaumik, S.; Ahmad, F.; Mandsaurwala, A.; Satam, H. Molecular spectrum of somatic EGFR and KRAS gene mutations in non small cell lung carcinoma: Determination of frequency, distribution pattern and identification of novel variations in Indian patients. Pathol. Oncol. Res. 2015, 21, 675–687. [Google Scholar] [CrossRef]
- Fathi, Z.; Mousavi, S.A.J.; Roudi, R.; Ghazi, F. Distribution of KRAS, DDR2, and TP53 gene mutations in lung cancer: An analysis of Iranian patients. PLoS ONE 2018, 26, e0200633. [Google Scholar] [CrossRef]
- Cavagna, R.; Escremim de Paula, F.; Sant’Anna, D.; Santana, I.; da Silva, V.D.; da Silva, E.C.A.; Bacchi, C.E.; Miziara, J.E.; Dias, J.M.; De Marchi, P.; et al. Frequency of KRAS p.Gly12Cys Mutation in Brazilian Patients With Lung Cancer. JCO Glob. Oncol. 2021, 7, 639–645. [Google Scholar] [CrossRef]
- Wing, S.E.; Jankowska, M.M.; Zou, X.; Sosa, E.; Yang, J.A.; Benmarhnia, T.; Neuhausen, S.L.; Nelson, R.; Salgia, R.; Gray, S.W.; et al. Neighborhood disadvantage is associated with KRAS-mutated non-small cell lung cancer risk. J. Cancer Res. Clin. Oncol. 2022. [Google Scholar] [CrossRef]
- Canadian Partnership against Cancer Lung Cancer and Equity. A Focus on Income and Geography. Available online: https://s22457.pcdn.co/wp-content/uploads/2020/11/Lung-cancer-and-equity-report-EN.pdf (accessed on 16 February 2023).
- MacPhail, C.; Snow, S. Not All Canadian Cancer Patients Are Equal-Disparities in Public Cancer Drug Funding across Canada. Curr. Oncol. 2022, 29, 2064–2072. [Google Scholar] [CrossRef]
- Sun, J.M.; Hwang, D.W.; Ahn, J.S.; Ahn, M.J.; Park, K. Prognostic and predictive value of KRAS mutations in advanced non-small cell lung cancer. PLoS ONE 2013, 8, e64816. [Google Scholar] [CrossRef]
- Tsao, M.S.; Aviel-Ronen, S.; Ding, K.; Lau, D.; Liu, N.; Sakurada, A.; Whitehead, M.; Zhu, C.Q.; Livingston, R.; Johnson, D.H.; et al. Prognostic and predictive importance of p53 and RAS for adjuvant chemotherapy in non small-cell lung cancer. J. Clin. Oncol. 2007, 25, 5240–5247. [Google Scholar] [CrossRef]
- Chaft, J.E.; Rusch, V.; Ginsberg, M.S.; Paik, P.K.; Finley, D.J.; Kris, M.G.; Price, K.A.; Azzoli, C.G.; Fury, M.G.; Riely, G.J.; et al. Phase II trial of neoadjuvant bevacizumab plus chemotherapy and adjuvant bevacizumab in patients with resectable nonsquamous non-small-cell lung cancers. J. Thorac. Oncol. 2013, 8, 1084–1090. [Google Scholar] [CrossRef]
- Ghimessy, A.K.; Gellert, A.; Schlegl, E.; Hegedus, B.; Raso, E.; Barbai, T.; Timar, J.; Ostoros, G.; Megyesfalvi, Z.; Gieszer, B.; et al. KRAS Mutations Predict Response and Outcome in Advanced Lung Adenocarcinoma Patients Receiving First-Line Bevacizumab and Platinum-Based Chemotherapy. Cancers 2019, 11, 1514. [Google Scholar] [CrossRef]
- D’Incecco, A.; Andreozzi, M.; Ludovini, V.; Rossi, E.; Capodanno, A.; Landi, L.; Tibaldi, C.; Minuti, G.; Salvini, J.; Coppi, E.; et al. PD-1 and PD-L1 expression in molecularly selected non-small-cell lung cancer patients. Br. J. Cancer 2015, 112, 95–102. [Google Scholar] [CrossRef]
- Garassino, M.C.; Marabese, M.; Rusconi, P.; Rulli, E.; Martelli, O.; Farina, G.; Scanni, A.; Broggini, M. Different types of K-Ras mutations could affect drug sensitivity and tumour behaviour in non-small-cell lung cancer. Ann. Oncol. 2011, 22, 235–237. [Google Scholar] [CrossRef]
- Mazieres, J.; Drilon, A.; Lusque, A.; Mhanna, L.; Cortot, A.B.; Mezquita, L.; Thai, A.A.; Mascaux, C.; Couraud, S.; Veillon, R.; et al. Immune checkpoint inhibitors for patients with advanced lung cancer and oncogenic driver alterations: Results from the IMMUNOTARGET registry. Ann. Oncol. 2019, 30, 1321–1328. [Google Scholar] [CrossRef]
- Dong, Z.Y.; Zhong, W.Z.; Zhang, X.C.; Su, J.; Xie, Z.; Liu, S.Y.; Tu, H.Y.; Chen, H.J.; Sun, Y.L.; Zhou, Q.; et al. Potential Predictive Value of TP53 and KRAS Mutation Status for Response to PD-1 Blockade Immunotherapy in Lung Adenocarcinoma. Clin. Cancer Res. 2017, 23, 3012–3024. [Google Scholar] [CrossRef] [PubMed]
- Schoenfeld, A.J.; Rizvi, H.; Bandlamudi, C.; Sauter, J.L.; Travis, W.D.; Rekhtman, N.; Plodkowski, A.J.; Perez-Johnston, R.; Sawan, P.; Beras, A.; et al. Clinical and molecular correlates of PD-L1 expression in patients with lung adenocarcinomas. Ann. Oncol. 2020, 31, 599–608. [Google Scholar] [CrossRef] [PubMed]
- Lauko, A.; Kotecha, R.; Barnett, A.; Li, H.; Tatineni, V.; Ali, A.; Patil, P.; Mohammadi, A.M.; Chao, S.T.; Murphy, E.S.; et al. Impact of KRAS mutation status on the efficacy of immunotherapy in lung cancer brain metastases. Sci. Rep. 2021, 11, 18174, Erratum in Sci. Rep. 2022, 12, 1147. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Ricciuti, B.; Nguyen, T.; Li, X.; Rabin, M.S.; Awad, M.M.; Lin, X.; Johnson, B.E.; Christiani, D.C. Association between Smoking History and Tumor Mutation Burden in Advanced Non-Small Cell Lung Cancer. Cancer Res. 2021, 81, 2566–2573. [Google Scholar] [CrossRef]
- Kim, J.Y.; Kronbichler, A.; Eisenhut, M.; Hong, S.H.; van der Vliet, H.J.; Kang, J.; Shin, J.I.; Gamerith, G. Tumor Mutational Burden and Efficacy of Immune Checkpoint Inhibitors: A Systematic Review and Meta-Analysis. Cancers 2019, 11, 1798. [Google Scholar] [CrossRef]
- Skoulidis, F.; Goldberg, M.E.; Greenawalt, D.M.; Hellmann, M.D.; Awad, M.M.; Gainor, J.F.; Schrock, A.B.; Hartmaier, R.J.; Trabucco, S.E.; Gay, L.; et al. STK11/LKB1 Mutations and PD-1 Inhibitor Resistance in KRAS-Mutant Lung Adenocarcinoma. Cancer Discov. 2018, 8, 822–835. [Google Scholar] [CrossRef]
- Ricciuti, B.; Arbour, K.C.; Lin, J.J.; Vajdi, A.; Vokes, N.; Hong, L.; Zhang, J.; Tolstorukov, M.Y.; Li, Y.Y.; Spurr, L.F.; et al. Diminished Efficacy of Programmed Death-(Ligand)1 Inhibition in STK11- and KEAP1-Mutant Lung Adenocarcinoma Is Affected by KRAS Mutation Status. J. Thorac. Oncol. 2022, 17, 399–410. [Google Scholar] [CrossRef]
- Papillon-Cavanagh, S.; Doshi, P.; Dobrin, R.; Szustakowski, J.; Walsh, A.M. STK11 and KEAP1 mutations as prognostic biomarkers in an observational real-world lung adenocarcinoma cohort. ESMO Open 2020, 5, e000706, Erratum in ESMO Open 2020, 5, e000706corr1. [Google Scholar] [CrossRef]
- Di Federico, A.; De Giglio, A.; Parisi, C.; Gelsomino, F. STK11/LKB1 and KEAP1 mutations in non-small cell lung cancer: Prognostic rather than predictive? Eur. J. Cancer 2021, 157, 108–113. [Google Scholar] [CrossRef]
- Bontoux, C.; Hofman, V.; Brest, P.; Ilié, M.; Mograbi, B.; Hofman, P. Daily Practice Assessment of KRAS Status in NSCLC Patients: A New Challenge for the Thoracic Pathologist Is Right around the Corner. Cancers 2022, 14, 1628. [Google Scholar] [CrossRef]
- Bruno, R.; Fontanini, G. Next Generation Sequencing for Gene Fusion Analysis in Lung Cancer: A Literature Review. Diagnostics 2020, 10, 521. [Google Scholar] [CrossRef] [PubMed]
- Rolfo, C.; Mack, P.; Scagliotti, G.V.; Aggarwal, C.; Arcila, M.E.; Barlesi, F.; Bivona, T.; Diehn, M.; Dive, C.; Dziadziuszko, R.; et al. Liquid Biopsy for Advanced NSCLC: A Consensus Statement From the International Association for the Study of Lung Cancer. J. Thorac. Oncol. 2021, 16, 1647–1662. [Google Scholar] [CrossRef] [PubMed]
- Zill, O.A.; Banks, K.C.; Fairclough, S.R.; Mortimer, S.A.; Vowles, J.V.; Mokhtari, R.; Gandara, D.R.; Mack, P.C.; Odegaard, J.I.; Nagy, R.J.; et al. The Landscape of Actionable Genomic Alterations in Cell-Free Circulating Tumor DNA from 21,807 Advanced Cancer Patients. Clin. Cancer Res. 2018, 24, 3528–3538. [Google Scholar] [CrossRef]
- Rolfo, C.; Mack, P.C.; Scagliotti, G.V.; Baas, P.; Barlesi, F.; Bivona, T.G.; Herbst, R.S.; Mok, T.S.; Peled, N.; Pirker, R.; et al. Liquid Biopsy for Advanced Non-Small Cell Lung Cancer (NSCLC): A Statement Paper from the IASLC. J. Thorac. Oncol. 2018, 13, 1248–1268. [Google Scholar] [CrossRef]
- Bauml, J.; Levy, B. Clonal Hematopoiesis: A New Layer in the Liquid Biopsy Story in Lung Cancer. Clin. Cancer Res. 2018, 24, 4352–4354. [Google Scholar] [CrossRef]
- Hu, Y.; Ulrich, B.C.; Supplee, J.; Kuang, Y.; Lizotte, P.H.; Feeney, N.B.; Guibert, N.M.; Awad, M.M.; Wong, K.K.; Jänne, P.A.; et al. False-Positive Plasma Genotyping Due to Clonal Hematopoiesis. Clin. Cancer Res. 2018, 24, 4437–4443. [Google Scholar] [CrossRef]
- Shlush, L.I. Age-related clonal hematopoiesis. Blood 2018, 131, 496–504. [Google Scholar] [CrossRef] [PubMed]
- Anglesio, M.S.; Papadopoulos, N.; Ayhan, A.; Nazeran, T.M.; Noë, M.; Horlings, H.M.; Lum, A.; Jones, S.; Senz, J.; Seckin, T.; et al. Cancer-Associated Mutations in Endometriosis without Cancer. N. Engl. J. Med. 2017, 376, 1835–1848. [Google Scholar] [CrossRef] [PubMed]
- Chang, J.C.; Montecalvo, J.; Borsu, L.; Lu, S.; Larsen, B.T.; Wallace, W.D.; Sae-Ow, W.; Mackinnon, A.C.; Kim, H.R.; Bowman, A.; et al. Bronchiolar Adenoma: Expansion of the Concept of Ciliated Muconodular Papillary Tumors With Proposal for Revised Terminology Based on Morphologic, Immunophenotypic, and Genomic Analysis of 25 Cases. Am. J. Surg. Pathol. 2018, 42, 1010–1026. [Google Scholar] [CrossRef] [PubMed]
- Sasaki, E.; Masago, K.; Fujita, S.; Hanai, N.; Yatabe, Y. Frequent KRAS and HRAS mutations in squamous cell papillomas of the head and neck. J. Pathol. Clin. Res. 2020, 6, 154–159. [Google Scholar] [CrossRef]
- Wittersheim, M.; Heydt, C.; Hoffmann, F.; Büttner, R. KRAS mutation in papillary fibroelastoma: A true cardiac neoplasm? J. Pathol. Clin. Res. 2017, 3, 100–104. [Google Scholar] [CrossRef]
- Leighl, N.B.; Page, R.D.; Raymond, V.M.; Daniel, D.B.; Divers, S.G.; Reckamp, K.L.; Villalona-Calero, M.A.; Dix, D.; Odegaard, J.I.; Lanman, R.B.; et al. Clinical Utility of Comprehensive Cell-free DNA Analysis to Identify Genomic Biomarkers in Patients with Newly Diagnosed Metastatic Non-small Cell Lung Cancer. Clin. Cancer Res. 2019, 25, 4691–4700. [Google Scholar] [CrossRef]
- Merker, J.D.; Oxnard, G.R.; Compton, C.; Diehn, M.; Hurley, P.; Lazar, A.J.; Lindeman, N.; Lockwood, C.M.; Rai, A.J.; Schilsky, R.L.; et al. Circulating Tumor DNA Analysis in Patients With Cancer: American Society of Clinical Oncology and College of American Pathologists Joint Review. J. Clin. Oncol. 2018, 36, 1631–1641. [Google Scholar] [CrossRef]
- Koessler, T.; Paradiso, V.; Piscuoglio, S.; Nienhold, R.; Ho, L.; Christinat, Y.; Terracciano, L.M.; Cathomas, G.; Wicki, A.; McKee, T.A.; et al. Reliability of liquid biopsy analysis: An inter-laboratory comparison of circulating tumor DNA extraction and sequencing with different platforms. Lab. Investig. 2020, 100, 1475–1484. [Google Scholar] [CrossRef] [PubMed]
- Lampignano, R.; Neumann, M.H.D.; Weber, S.; Kloten, V.; Herdean, A.; Voss, T.; Groelz, D.; Babayan, A.; Tibbesma, M.; Schlumpberger, M.; et al. Multicenter Evaluation of Circulating Cell-Free DNA Extraction and Downstream Analyses for the Development of Standardized (Pre)analytical Work Flows. Clin. Chem. 2020, 66, 149–160. [Google Scholar] [CrossRef] [PubMed]
- Lindeman, N.I.; Cagle, P.T.; Aisner, D.L.; Arcila, M.E.; Beasley, M.B.; Bernicker, E.H.; Colasacco, C.; Dacic, S.; Hirsch, F.R.; Kerr, K.; et al. Updated Molecular Testing Guideline for the Selection of Lung Cancer Patients for Treatment With Targeted Tyrosine Kinase Inhibitors: Guideline From the College of American Pathologists, the International Association for the Study of Lung Cancer, and the Association for Molecular Pathology. J. Mol. Diagn. 2018, 20, 129–159. [Google Scholar] [CrossRef]
- Cheema, P.K.; Gomes, M.; Banerji, S.; Joubert, P.; Leighl, N.B.; Melosky, B.; Sheffield, B.S.; Stockley, T.; Ionescu, D.N. Consensus recommendations for optimizing biomarker testing to identify and treat advanced EGFR-mutated non-small-cell lung cancer. Curr. Oncol. 2020, 27, 321–329. [Google Scholar] [CrossRef]
- Li, M.M.; Datto, M.; Duncavage, E.J.; Kulkarni, S.; Lindeman, N.I.; Roy, S.; Tsimberidou, A.M.; Vnencak-Jones, C.L.; Wolff, D.J.; Younes, A.; et al. Standards and Guidelines for the Interpretation and Reporting of Sequence Variants in Cancer: A Joint Consensus Recommendation of the Association for Molecular Pathology, American Society of Clinical Oncology, and College of American Pathologists. J. Mol. Diagn. 2017, 19, 4–23. [Google Scholar] [CrossRef] [PubMed]
- Cagle, P.T.; Sholl, L.M.; Lindeman, N.I.; Alsabeh, R.; Divaris, D.X.; Foulis, P.; Lee, G.; Neal, J.W.; Nowak, J.A.; Yu, P.P.; et al. Template for reporting results of biomarker testing of specimens from patients with non-small cell carcinoma of the lung. Arch. Pathol. Lab. Med. 2014, 138, 171–174. [Google Scholar] [CrossRef] [PubMed]
- Hanna, N.H.; Schneider, B.J.; Temin, S.; Baker, S., Jr.; Brahmer, J.; Ellis, P.M.; Gaspar, L.E.; Haddad, R.Y.; Hesketh, P.J.; Jain, D.; et al. Therapy for Stage IV Non-Small-Cell Lung Cancer Without Driver Alterations: ASCO and OH (CCO) Joint Guideline Update. J. Clin. Oncol. 2020, 38, 1608–1632. [Google Scholar] [CrossRef] [PubMed]
- Reck, M.; Rodríguez-Abreu, D.; Robinson, A.G.; Hui, R.; Csőszi, T.; Fülöp, A.; Gottfried, M.; Peled, N.; Tafreshi, A.; Cuffe, S.; et al. Pembrolizumab versus Chemotherapy for PD-L1-Positive Non-Small-Cell Lung Cancer. N. Engl. J. Med. 2016, 375, 1823–1833. [Google Scholar] [CrossRef] [PubMed]
- Gandhi, L.; Rodríguez-Abreu, D.; Gadgeel, S.; Esteban, E.; Felip, E.; De Angelis, F.; Domine, M.; Clingan, P.; Hochmair, M.J.; Powell, S.F.; et al. Pembrolizumab plus Chemotherapy in Metastatic Non-Small-Cell Lung Cancer. N. Engl. J. Med. 2018, 378, 2078–2092. [Google Scholar] [CrossRef] [PubMed]
- Paz-Ares, L.; Luft, A.; Vicente, D.; Tafreshi, A.; Gümüş, M.; Mazières, J.; Hermes, B.; Çay Şenler, F.; Csőszi, T.; Fülöp, A.; et al. Pembrolizumab plus Chemotherapy for Squamous Non-Small-Cell Lung Cancer. N. Engl. J. Med. 2018, 379, 2040–2051. [Google Scholar] [CrossRef] [PubMed]
- Mok, T.S.K.; Wu, Y.L.; Kudaba, I.; Kowalski, D.M.; Cho, B.C.; Turna, H.Z.; Castro, G., Jr.; Srimuninnimit, V.; Laktionov, K.K.; Bondarenko, I.; et al. Pembrolizumab versus chemotherapy for previously untreated, PD-L1-expressing, locally advanced or metastatic non-small-cell lung cancer (KEYNOTE-042): A randomised, open-label, controlled, phase 3 trial. Lancet 2019, 393, 1819–1830. [Google Scholar] [CrossRef] [PubMed]
- Hellmann, M.D.; Paz-Ares, L.; Bernabe Caro, R.; Zurawski, B.; Kim, S.W.; Carcereny Costa, E.; Park, K.; Alexandru, A.; Lupinacci, L.; de la Mora Jimenez, E.; et al. Nivolumab plus Ipilimumab in Advanced Non-Small-Cell Lung Cancer. N. Engl. J. Med. 2019, 381, 2020–2031. [Google Scholar] [CrossRef]
- Paz-Ares, L.; Ciuleanu, T.E.; Cobo, M.; Schenker, M.; Zurawski, B.; Menezes, J.; Richardet, E.; Bennouna, J.; Felip, E.; Juan-Vidal, O.; et al. First-line nivolumab plus ipilimumab combined with two cycles of chemotherapy in patients with non-small-cell lung cancer (CheckMate 9LA): An international, randomised, open-label, phase 3 trial. Lancet Oncol. 2021, 22, 198–211, Erratum in Lancet Oncol. 2021, 22, e92. [Google Scholar] [CrossRef]
- Herbst, R.S.; Giaccone, G.; de Marinis, F.; Reinmuth, N.; Vergnenegre, A.; Barrios, C.H.; Morise, M.; Felip, E.; Andric, Z.; Geater, S.; et al. Atezolizumab for First-Line Treatment of PD-L1-Selected Patients with NSCLC. N. Engl. J. Med. 2020, 383, 1328–1339. [Google Scholar] [CrossRef]
- Gogishvili, M.; Melkadze, T.; Makharadze, T.; Giorgadze, D.; Dvorkin, M.; Penkov, K.; Laktionov, K.; Nemsadze, G.; Nechaeva, M.; Rozhkova, I.; et al. Cemiplimab plus chemotherapy versus chemotherapy alone in non-small cell lung cancer: A randomized, controlled, double-blind phase 3 trial. Nat. Med. 2022, 28, 2374–2380. [Google Scholar] [CrossRef]
- Herbst, R.S.; Lopes, G.; Kowalski, D.M.; Kasahara, K.; Wu, Y.-L.; De Castro, G.; Cho, B.C.; Turna, H.Z.; Cristescu, R.; Aurora-Garg, D.; et al. LBA4 Association of KRAS mutational status with response to pembrolizumab monotherapy given as first-line therapy for PD-L1-positive advanced non-squamous NSCLC in Keynote-042. Ann. Oncol. 2019, 30, xi63–xi64. [Google Scholar] [CrossRef]
- Gadgeel, S.; Rodriguez-Abreu, D.; Felip, E.; Esteban, E.; Speranza, G.; Reck, M.; Hui, R.; Boyer, M.; Garon, E.B.; Horinouchi, H.; et al. LBA5—KRAS mutational status and efficacy in KEYNOTE-189: Pembrolizumab (pembro) plus chemotherapy (chemo) vs placebo plus chemo as first-line therapy for metastatic non-squamous NSCLC. Ann. Oncol. 2019, 30, xi64–xi65. [Google Scholar] [CrossRef]
- Nakajima, E.C.; Ren, Y.; Vallejo, J.J.; Akinboro, O.; Mishra-Kalyani, P.S.; Larkins, E.A.; Drezner, N.L.; Tang, S.; Pazdur, R.; Beaver, J.A.; et al. Outcomes of first-line immune checkpoint inhibitors with or without chemotherapy according to KRAS mutational status and PD-L1 expression in patients with advanced NSCLC: FDA pooled analysis. J. Clin. Oncol. 2022, 40 (Suppl. S16), 9001. [Google Scholar] [CrossRef]
- Jänne, P.A.; Riely, G.J.; Gadgeel, S.M.; Heist, R.S.; Ou, S.I.; Pacheco, J.M.; Johnson, M.L.; Sabari, J.K.; Leventakos, K.; Yau, E.; et al. Adagrasib in Non-Small-Cell Lung Cancer Harboring a KRASG12C Mutation. N. Engl. J. Med. 2022, 387, 120–131. [Google Scholar] [CrossRef] [PubMed]
- U.S. Food & Drug Administration: FDA Grants Accelerated Approval to Adagrasib for KRAS G12C-Mutated NSCLC. Available online: www.fda.gov/drugs/resources-information-approved-drugs/fda-grants-accelerated-approval-adagrasib-kras-g12c-mutated-nsclc (accessed on 4 May 2023).
- NCCN Clinical Practice Guidelines in Oncology (NCCN Guidelines®) Non-Small Cell Lung Cancer. Version 1. 2023. Available online: https://www.nccn.org/professionals/physician_gls/pdf/nscl.pdf (accessed on 4 May 2023).
- Purkey, H. Discovery of GDC-6036, a clinical stage treatment for KRAS G12C-positive cancers. Cancer Res. 2022, 82 (Suppl. S12), ND11. [Google Scholar] [CrossRef]
- Brachmann, S.M.; Weiss, A.; Guthy, D.A.; Beyer, K.; Voshol, J.; Maira, M.; Prahallad, A.; Porta, D.G.; Schnell, C.; Ostermann, N.; et al. JDQ443, a covalent irreversible inhibitor of KRAS G12C, exhibits a novel binding mode and demonstrates potent anti-tumor activity and favorable pharmacokinetic properties in preclinical models. Mol. Cancer Ther. 2021, 20 (Suppl. S12), P124. [Google Scholar] [CrossRef]
- Sacher, A.; Patel, M.R.; Miller, W.H.; Desai, J.; Garralda, E.; Bowyer, S.; Kim, T.W.; De Miguel, M.; Falcon, A.; Krebs, M.G.; et al. OA03.04 Phase I A Study to Evaluate GDC-6036 Monotherapy in Patients with Non-small Cell Lung Cancer (NSCLC) with KRAS G12C Mutation. J. Thor. Oncol. 2022, 17, S8–S9. [Google Scholar] [CrossRef]
- Tan, D.S.; Shimizu, T.; Solomon, B.; Heist, R.S.; Schuler, M.; De Miguel Luken, M.J.; Gazzah, A.; Wermke, M.; Dooms, C.; Loong, H.H.; et al. KontRASt-01: A phase Ib/II, dose-escalation study of JDQ443 in patients (pts) with advanced, KRAS G12C-mutated solid tumors. Cancer Res. 2022, 82, 12. [Google Scholar] [CrossRef]
- Ali, A.; Goffin, J.R.; Arnold, A.; Ellis, P.M. Survival of patients with non-small-cell lung cancer after a diagnosis of brain metastases. Curr. Oncol. 2013, 20, e300–e306. [Google Scholar] [CrossRef]
- Goldberg, S.B.; Schalper, K.A.; Gettinger, S.N.; Mahajan, A.; Herbst, R.S.; Chiang, A.C.; Lilenbaum, R.; Wilson, F.H.; Omay, S.B.; Yu, J.B.; et al. Pembrolizumab for management of patients with NSCLC and brain metastases: Long-term results and biomarker analysis from a non-randomised, open-label, phase 2 trial. Lancet Oncol. 2020, 21, 655–663. [Google Scholar] [CrossRef]
- Ramalingam, S.; Skoulidis, F.; Govindan, R.; Velcheti, V.; Li, B.; Besse, B.; Dy, G.; Kim, D.; Schuler, M.; Vincent, M.; et al. Efficacy of sotorasib in KRAS p.G12C-mutated NSCLC with stable brain metastases: A post-hoc analysis of CodeBreaK100. J. Thorac. Oncol. 2021, 16, S1123. [Google Scholar] [CrossRef]
- Chour, A.; Denis, J.; Lafitte, C.; Mascaux, C.; Zysman, M.; Lemaitre, A.; Swalduz, A.; Gounant, V.; Cortot, A.; Darrason, M.; et al. Sotorasib-induced liver and non-liver toxicity associated with sequential sotorasib following anti-PD(L)1 in KRASG12C mutant lung cancer. Ann. Oncol. 2022, 33, S49–S50. [Google Scholar] [CrossRef]
- Schoenfeld, A.J.; Arbour, K.C.; Rizvi, H.; Iqbal, A.N.; Gadgeel, S.M.; Girshman, J.; Kris, M.G.; Riely, G.J.; Yu, H.A.; Hellmann, M.D. Severe immune-related adverse events are common with sequential PD-(L)1 blockade and osimertinib. Ann. Oncol. 2019, 30, 839–844. [Google Scholar] [CrossRef] [PubMed]
- Begum, P.; Goldin, R.D.; Possamai, L.A.; Popat, S. Severe Immune Checkpoint Inhibitor Hepatitis in KRAS G12C-Mutant NSCLC Potentially Triggered by Sotorasib: Case Report. JTO Clin. Res. Rep. 2021, 2, 100213. [Google Scholar] [CrossRef] [PubMed]
- Li, B.T.; Falchook, G.S.; Durm, G.A.; Burns, T.F.; Skoulidis, F.; Ramalingam, S.S.; Spira, A.; Bestvina, C.M.; Goldberg, S.B.; Veluswamy, R.; et al. CodeBreaK 100/101: First report of safety/efficacy of sotorasib in combination with pembrolizumab or atezolizumab in advanced KRAS p.G12C NSCLC. J. Thorac. Oncol. 2022, 17, S10–S11. [Google Scholar] [CrossRef]
- Zhao, Y.; Murciano-Goroff, Y.R.; Xue, J.Y.; Ang, A.; Lucas, J.; Mai, T.T.; Da Cruz Paula, A.F.; Saiki, A.Y.; Mohn, D.; Achanta, P.; et al. Diverse alterations associated with resistance to KRAS(G12C) inhibition. Nature 2021, 599, 679–683. [Google Scholar] [CrossRef]
- Blaquier, J.B.; Cardona, A.F.; Recondo, G. Resistance to KRASG12C Inhibitors in Non-Small Cell Lung Cancer. Front. Oncol. 2021, 11, 787585. [Google Scholar] [CrossRef]
- Dunnett-Kane, V.; Nicola, P.; Blackhall, F.; Lindsay, C. Mechanisms of Resistance to KRASG12C Inhibitors. Cancers 2021, 13, 151. [Google Scholar] [CrossRef]
- Li, B.T.; Velcheti, V.; Price, T.J.; Hong, D.S.; Fakih, M.; Kim, S.W.; Falchook, G.S.; Delord, J.-P.; Dy, G.K.; Ramalingam, S.S.; et al. Largest evaluation of acquired resistance to sotorasib in KRAS p.G12C-mutated non–small cell lung cancer (NSCLC) and colorectal cancer (CRC): Plasma biomarker analysis of CodeBreaK100. J. Clinl. Oncol. 2022, 40 (Suppl. S16), 102. [Google Scholar] [CrossRef]
- Zhang, J.; Zhang, J.; Liu, Q.; Fan, X.X.; Leung, E.L.; Yao, X.J.; Liu, L. Resistance looms for KRAS G12C inhibitors and rational tackling strategies. Pharmacol. Ther. 2022, 229, 108050. [Google Scholar] [CrossRef]
- Reita, D.; Pabst, L.; Pencreach, E.; Guérin, E.; Dano, L.; Rimelen, V.; Voegeli, A.C.; Vallat, L.; Mascaux, C.; Beau-Faller, M. Direct Targeting KRAS Mutation in Non-Small Cell Lung Cancer: Focus on Resistance. Cancers 2022, 14, 1321. [Google Scholar] [CrossRef]
- Awad, M.M.; Liu, S.; Rybkin, I.I.; Arbour, K.C.; Dilly, J.; Zhu, V.W.; Johnson, M.L.; Heist, R.S.; Patil, T.; Riely, G.J.; et al. Acquired Resistance to KRASG12C Inhibition in Cancer. N. Engl. J. Med. 2021, 384, 2382–2393. [Google Scholar] [CrossRef]
- Canon, J.; Rex, K.; Saiki, A.Y.; Mohr, C.; Cooke, K.; Bagal, D.; Gaida, K.; Holt, T.; Knutson, C.G.; Koppada, N.; et al. The clinical KRAS(G12C) inhibitor AMG 510 drives anti-tumour immunity. Nature 2019, 575, 217–223. [Google Scholar] [CrossRef] [PubMed]
| Trial | Krystal-1 [94] | CodeBreak100 [4] | CodeBreak 200 [5] |
|---|---|---|---|
| Phase | 2 | 2 | 3 |
| Drug | Adagrasib | Sotorasib | Sotorasib |
| N | 116 | 126 | 171 |
| ORR | 42.9% (95% CI 33.5–52.6) | 37.1% (95% CI 28.6–46.2) | 28.1% (95% CI 21.5–35.4) |
| DCR | 79.5% (95% CI 70.8–86.5%) | 80.6% (95% CI 72.6–87.2%) | 82.5% (95% CI 75.9–87.8) |
| TTR | 1.4 months (range 0.9–7.1 months) | 1.4 months (range 1.2–10.1 months) | 1.4 months (range 1.2–8.3 months) |
| DoR | 8.5 months (95% CI 6.2–13.8) | 11.1 months (95% CI 6.9-NE) | 8.6 months (95% CI 7.1–18.0) |
| PFS, median | 6.5 months (95% CI 4.7–8.4) | 6.8 months (95% CI 5.1–8.2) | 5.6 months (95% CI 4.3–7.8) |
| OS, median | 12.6 months (95% CI 9.2–19.2) | 12.5 months (95% 10-NE) | 10.6 months (95% CI 8.9–14.0) |
| Follow up, median | 12.9 months | 15.3 months | 17.7 months |
| Trial | Krystal-1 [94] | CodeBreak100 [4] | ||
|---|---|---|---|---|
| Drug | Adagrasib | Sotorasib | ||
| N | 116 | 126 | ||
| Any TRAE | Any Grade | Grade 3–4 | Any Grade | Grade 3–4 |
| 113 (97%) | 50 (43%) | 88 (70%) | 26 (21%) | |
| Most Frequent TRAEs | ||||
| Diarrhea | 73 (63%) | 1 (<1%) | 40 (32%) | 5 (4%) |
| Nausea | 72 (62%) | 5 (4%) | 24 (19%) | 0 |
| Fatigue | 47 (41%) | 5 (4%) | 14 (11%) | 0 |
| Vomiting | 55 (47%) | 1 (<1%) | 10 (8%) | 0 |
| ALT increased | 32 (28%) | 5 (4%) | 19 (15%) | 8 (6%) |
| AST increased | 29 (25%) | 4 (3%) | 19 (15%) | 7 (6%) |
| Decreased appetite | 28 (24%) | 4 (3%) | 16 (13%) | 0 |
| Blood alkaline phosphatase increase | 12 (10%) | 4 (4%) | 17 (14%) | 1 (1%) |
| Peripheral edema | 12 (10%) | 0 | 18 (14%) | 0 |
| Anemia | 21 (18%) | 6 (5%) | 18 (14%) | 1 (1%) |
| Severity | Sotorasib Action | Medical Management, Monitoring and Follow-Up |
|---|---|---|
| Grade 2–4 AST or ALT | First occurrence:
|
|
Second occurrence:
|
| |
Third occurrence:
|
| |
| AST or ALT > 3 ×ULN with totalbilirubin > 2 x ULN in the absence of alternative causes | Permanently discontinue |
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cheema, P.K.; Banerji, S.O.; Blais, N.; Chu, Q.S.-C.; Juergens, R.A.; Leighl, N.B.; Sacher, A.; Sheffield, B.S.; Snow, S.; Vincent, M.; et al. Canadian Consensus Recommendations on the Management of KRAS G12C-Mutated NSCLC. Curr. Oncol. 2023, 30, 6473-6496. https://doi.org/10.3390/curroncol30070476
Cheema PK, Banerji SO, Blais N, Chu QS-C, Juergens RA, Leighl NB, Sacher A, Sheffield BS, Snow S, Vincent M, et al. Canadian Consensus Recommendations on the Management of KRAS G12C-Mutated NSCLC. Current Oncology. 2023; 30(7):6473-6496. https://doi.org/10.3390/curroncol30070476
Chicago/Turabian StyleCheema, Parneet K., Shantanu O. Banerji, Normand Blais, Quincy S.-C. Chu, Rosalyn A. Juergens, Natasha B. Leighl, Adrian Sacher, Brandon S. Sheffield, Stephanie Snow, Mark Vincent, and et al. 2023. "Canadian Consensus Recommendations on the Management of KRAS G12C-Mutated NSCLC" Current Oncology 30, no. 7: 6473-6496. https://doi.org/10.3390/curroncol30070476
APA StyleCheema, P. K., Banerji, S. O., Blais, N., Chu, Q. S.-C., Juergens, R. A., Leighl, N. B., Sacher, A., Sheffield, B. S., Snow, S., Vincent, M., Wheatley-Price, P. F., Yip, S., & Melosky, B. L. (2023). Canadian Consensus Recommendations on the Management of KRAS G12C-Mutated NSCLC. Current Oncology, 30(7), 6473-6496. https://doi.org/10.3390/curroncol30070476







