Screening for Colorectal Cancer Leading into a New Decade: The “Roaring ‘20s” for Epigenetic Biomarkers?
Abstract
1. Introduction
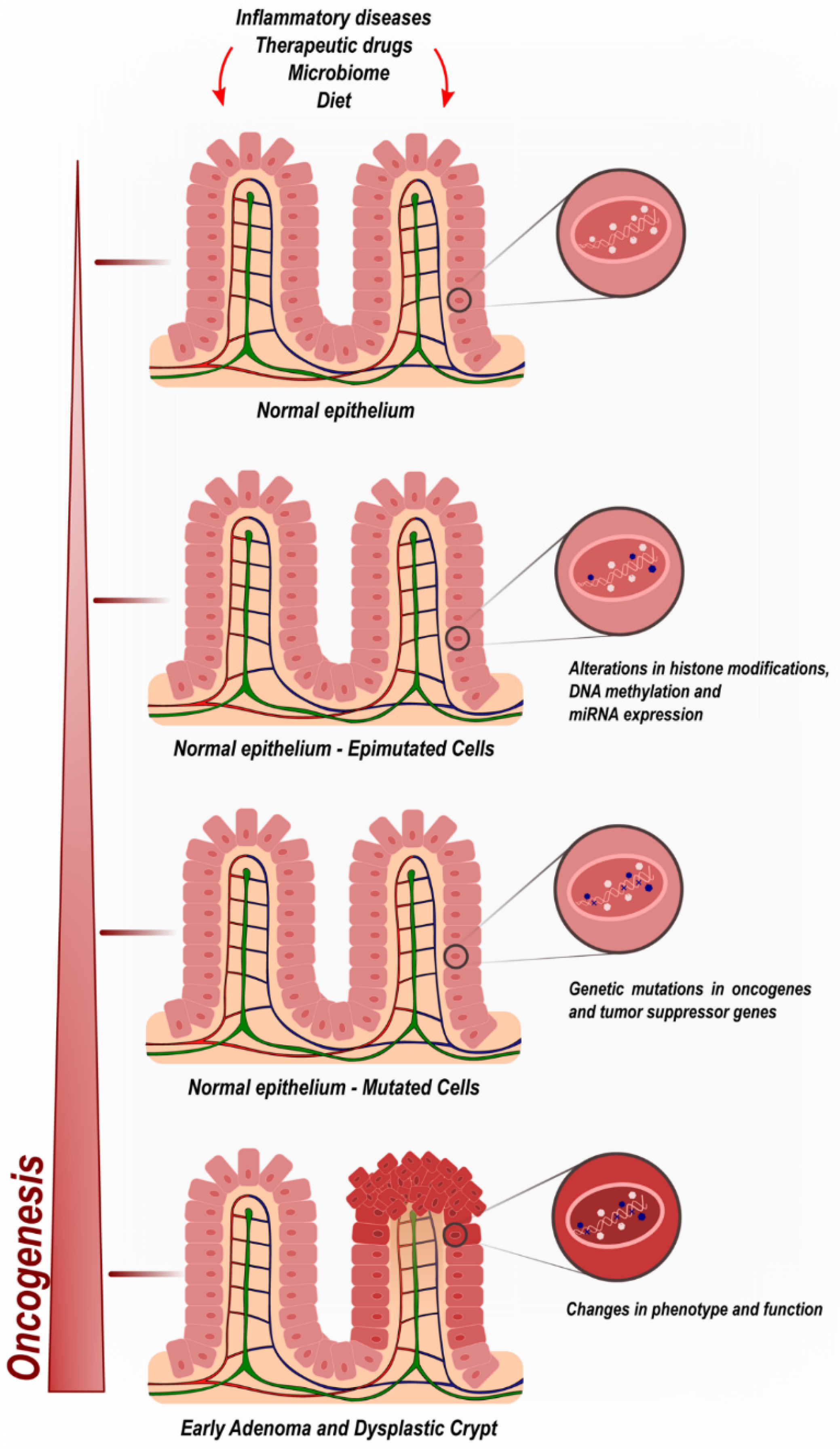
1.1. Genetic and Molecular Pathogenesis of CRC
1.2. Epigenetic Mechanisms in CRC—DNA Methylation
1.3. Epigenetic Mechanisms in CRC—MicroRNA Regulation
1.4. Epigenetic Mechanisms in CRC—Histone Modifications
2. Objectives
3. Materials and Methods
4. Techniques Used for Screening and Diagnosis of CRC
4.1. Digital Rectal Examination
4.2. Sigmoidoscopy
4.3. Colonoscopy
4.4. CT Colonography
4.5. Colon Capsule Endoscopy
4.6. Double Contrast Barium Enema
5. Biomarkers Used for Screening and Diagnosis of CRC
5.1. Fecal Screening Tests
5.1.1. Fecal Occult-Blood Tests—gFOBT and FIT
5.1.2. Fecal DNA Testing
5.1.3. Methylated Vimentin
5.2. Blood Screening Tests
6. Novel Biomarkers in CRC—An Epigenetic-Based Approach
6.1. DNA Methylation—Stool-Based Biomarkers
6.2. DNA Methylation—Blood-Based Biomarkers
6.3. Micro RNAs—Stool-Based Biomarkers
6.4. Micro RNAs—Blood-Based Biomarkers
6.5. Histone Modifications
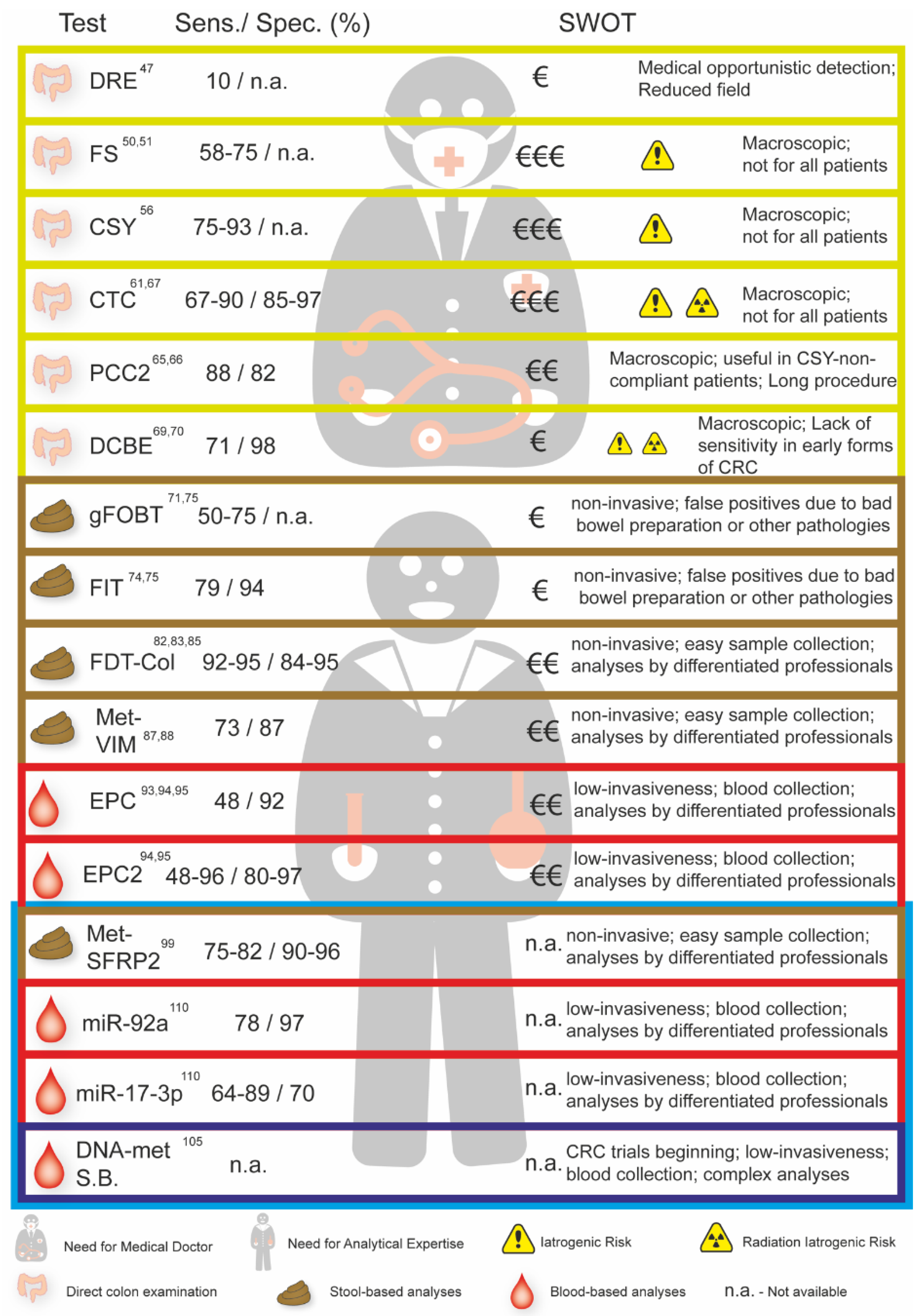
7. Discussion and Conclusions
- (1)
- It only detects CRC that developed with the bleeding lesions. However, bleeding can be sporadic, and many CRC lesions might progress without bleeding;
- (2)
- It detects lesions in a histological cancer phase, sometimes too late for an effective treatment;
- (3)
- It requires bowel preparation;
- (4)
- It presents many false positives due to peptic ulcers or bowel hemangiomas;
- (5)
- It presents many false positives due to the wide use of antiplatelet, anticoagulant, or non-steroid anti-inflammatory drugs;
- (6)
- In specific populations, it has low compliance due to religious or social reasons.
8. Future Directions
Author Contributions
Funding
Conflicts of Interest
References
- Ferlay, J.; Colombet, M.; Soerjomataram, I.; Mathers, C.; Parkin, D.M.; Piñeros, M.; Znaor, A.; Bray, F. Estimating the global cancer incidence and mortality in 2018: GLOBOCAN sources and methods. Int. J. Cancer 2018, 144, 1941–1953. [Google Scholar] [CrossRef] [PubMed]
- Siegel, R.L.; Miller, K.D.; Fedewa, S.A.; Ahnen, D.J.; Meester, R.G.S.; Barzi, A.; Jemal, A. Colorectal cancer statistics, 2017. CA Cancer J. Clin. 2017, 67, 177–193. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.-M.; Qian, J.-C.; Cai, Z.; Tang, T.; Wang, P.; Zhang, K.-H.; Deng, Z.-L.; Cai, J.-P. DNA alterations of microsatellite DNA, p53, APC and K-ras in Chinese colorectal cancer patients. Eur. J. Clin. Investig. 2012, 42, 751–759. [Google Scholar] [CrossRef] [PubMed]
- Church, J. Molecular genetics of colorectal cancer. Semin. Colon Rectal Surg. 2016, 27, 172–175. [Google Scholar] [CrossRef]
- Vogelstein, B.; Fearon, E.R.; Hamilton, S.R.; Kern, S.E.; Preisinger, A.C.; Leppert, M.; Smits, A.M.; Bos, J.L. Genetic alterations during colorectal-tumor development. N. Engl. J. Med. 1988, 319, 525–532. [Google Scholar] [CrossRef] [PubMed]
- Pino, M.S.; Chung, D.C. The chromosomal instability pathway in colon cancer. Gastroenterology 2010, 138, 2059–2072. [Google Scholar] [CrossRef]
- Lengauer, C.; Kinzler, K.W.; Vogelstein, B. Genetic instability in colorectal cancers. Nat. Cell Biol. 1997, 386, 623–627. [Google Scholar] [CrossRef]
- Grady, W.M.; Yu, M.; Markowitz, S.D. Epigenetic alterations in the gastrointestinal tract: Current and emerging use for biomarkers of cancer. Gastroenterology 2021, 160, 690–709. [Google Scholar] [CrossRef]
- Half, E.; Bercovich, D.; Rozen, P. Familial adenomatous polyposis. Orphanet J. Rare Dis. 2009, 4, 22. [Google Scholar] [CrossRef]
- Boland, C.R.; Goel, A. Microsatellite instability in colorectal cancer. Gastroenterology 2010, 138, 2073–2087. [Google Scholar] [CrossRef]
- TNM Classification. Available online: https://www.ncbi.nlm.nih.gov/books/NBK553187/ (accessed on 20 April 2021).
- Kang, G.H. Four molecular subtypes of colorectal cancer and their precursor lesions. Arch. Pathol. Lab. Med. 2011, 135, 698–703. [Google Scholar] [CrossRef] [PubMed]
- Mojarad, E.N.; Kuppen, P.J.K.; Aghdaei, H.A.; Zali, M.R. The CpG island methylator phenotype (CIMP) in colorectal cancer. Gastroenterol. Hepatol. Bed Bench 2013, 6, 120–128. [Google Scholar]
- Kalady, M.F.; DeJulius, K.L.; Sanchez, J.A.; Jarrar, A.; Liu, X.; Manilich, E.; Skacel, M.; Church, J.M. BRAF mutations in colorectal cancer are associated with distinct clinical characteristics and worse prognosis. Dis. Colon Rectum 2012, 55, 128–133. [Google Scholar] [CrossRef]
- Mojarad, E.N.; Farahani, R.K.; Haghighi, M.M.; Aghdaei, H.A.; Kuppen, P.; Zali, M. Clinical implications of BRAF mutation test in colorectal cancer. Gastroenterol. Hepatol. Bed Bench 2013, 6, 6–13. [Google Scholar]
- Lao, V.V.; Grady, W.M. Epigenetics and colorectal cancer. Nat. Rev. Gastroenterol. Hepatol. 2011, 8, 686–700. [Google Scholar] [CrossRef] [PubMed]
- Pancione, M.; Remo, A.; Colantuoni, V. Genetic and epigenetic events generate multiple pathways in colorectal cancer progression. Pathol. Res. Int. 2012, 2012, 1–11. [Google Scholar] [CrossRef]
- Jones, P.A. Functions of DNA methylation: Islands, start sites, gene bodies and beyond. Nat. Rev. Genet. 2012, 13, 484–492. [Google Scholar] [CrossRef]
- Yang, Y.; Chu, F.-H.; Xu, W.-R.; Sun, J.-Q.; Sun, X.; Ma, X.-M.; Yu, M.-W.; Yang, G.-W.; Wang, X.-M. Identification of regulatory role of DNA methylation in colon cancer gene expression via systematic bioinformatics analysis. Medicine 2017, 96, e8487. [Google Scholar] [CrossRef]
- Castelo-Branco, P.; Choufani, S.; Mack, S.C.; Gallagher, D.; Zhang, C.; Lipman, T.; Zhukova, N.; Walker, E.J.; Martin, D.; Merino, D.; et al. Methylation of the TERT promoter and risk stratification of childhood brain tumours: An integrative genomic and molecular study. Lancet Oncol. 2013, 14, 534–542. [Google Scholar] [CrossRef]
- Kerachian, M.A.; Javadmanesh, A.; Azghandi, M.; Shariatpanahi, A.M.; Yassi, M.; Davodly, E.S.; Talebi, A.; Khadangi, F.; Soltani, G.; Hayatbakhsh, A.; et al. Crosstalk between DNA methylation and gene expression in colorectal cancer, a potential plasma biomarker for tracing this tumor. Sci. Rep. 2020, 10, 1–13. [Google Scholar] [CrossRef]
- Milicic, A.; Harrison, L.-A.; Goodlad, R.; Hardy, R.G.; Nicholson, A.M.; Presz, M.; Sieber, O.; Santander, S.; Pringle, J.H.; Mandir, N.; et al. Ectopic expression of P-Cadherin correlates with promoter hypomethylation early in colorectal carcinogenesis and enhanced intestinal crypt fission in vivo. Cancer Res. 2008, 68, 7760–7768. [Google Scholar] [CrossRef]
- Strubberg, A.M.; Madison, B.B. MicroRNAs in the etiology of colorectal cancer: Pathways and clinical implications. Dis. Model. Mech. 2017, 10, 197–214. [Google Scholar] [CrossRef]
- Chi, Y.; Zhou, D. MicroRNAs in colorectal carcinoma--from pathogenesis to therapy. J. Exp. Clin. Cancer Res. 2016, 35, 43. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, H.; Takatsuka, S.; Akashi, H.; Yamamoto, E.; Nojima, M.; Maruyama, R.; Kai, M.; Yamano, H.-O.; Sasaki, Y.; Tokino, T.; et al. Genome-Wide profiling of chromatin signatures reveals epigenetic regulation of MicroRNA genes in colorectal cancer. Cancer Res. 2011, 71, 5646–5658. [Google Scholar] [CrossRef]
- Schetter, A.J.; Okayama, H.; Harris, C.C. The role of MicroRNAs in colorectal cancer. Cancer J. 2012, 18, 244–252. [Google Scholar] [CrossRef] [PubMed]
- Jung, G.; Hernández-Illán, E.; Moreira, L.; Balaguer, F.; Goel, A. Epigenetics of colorectal cancer: Biomarker and therapeutic potential. Nat. Rev. Gastroenterol. Hepatol. 2020, 17, 111–130. [Google Scholar] [CrossRef] [PubMed]
- Tan, M.; Luo, H.; Lee, S.; Jin, F.; Yang, J.S.; Montellier, E.; Buchou, T.; Cheng, Z.; Rousseaux, S.; Rajagopal, N.; et al. Identification of 67 histone marks and histone lysine crotonylation as a new type of histone modification. Cell 2011, 146, 1016–1028. [Google Scholar] [CrossRef]
- Arnaudo, A.M.; Garcia, B.A. Proteomic characterization of novel histone post-translational modifications. Epigenetics Chromatin 2013, 6, 24. [Google Scholar] [CrossRef]
- Bishnupuri, K.S.; Mishra, M.K. Epigenetics of colorectal cancer. Epigenetic Adv. Cancer 2016, 97–121. [Google Scholar] [CrossRef]
- Migheli, F.; Migliore, L. Epigenetics of colorectal cancer. Clin. Genet. 2012, 81, 312–318. [Google Scholar] [CrossRef]
- Zhao, Y.; Fang, X.; Wang, Y.; Zhang, J.; Jiang, S.; Liu, Z.; Ma, Z.; Xu, L.; Li, E.; Zhang, K. Comprehensive analysis for histone acetylation of human colon cancer cells treated with a novel HDAC inhibitor. Curr. Pharm. Des. 2014, 20, 1866–1873. [Google Scholar] [CrossRef]
- Vaiopoulos, A.G.; Athanasoula, K.C.; Papavassiliou, A.G. Epigenetic modifications in colorectal cancer: Molecular insights and therapeutic challenges. Biochim. Biophys. Acta (BBA)-Mol. Basis Dis. 2014, 1842, 971–980. [Google Scholar] [CrossRef]
- Suzuki, H.; Gabrielson, E.; Chen, W.; Anbazhagan, R.; Van Engeland, M.; Weijenberg, M.P.; Herman, J.G.; Baylin, S.B. A genomic screen for genes upregulated by demethylation and histone deacetylase inhibition in human colorectal cancer. Nat. Genet. 2002, 31, 141–149. [Google Scholar] [CrossRef]
- Stypula-Cyrus, Y.; Damania, D.; Kunte, D.P.; Cruz, M.D.; Subramanian, H.; Roy, H.K.; Backman, V. HDAC Up-Regulation in early colon field carcinogenesis is involved in cell tumorigenicity through regulation of chromatin structure. PLoS ONE 2013, 8, e64600. [Google Scholar] [CrossRef] [PubMed]
- Rex, D.K.; Boland, R.C.; Dominitz, J.A.; Giardiello, F.M.; Johnson, D.A.; Kaltenbach, T.; Levin, T.R.; Lieberman, D.; Robertson, D.J. Colorectal cancer screening: Recommendations for physicians and patients from the U.S. Multi-Society task force on colorectal cancer. Am. J. Gastroenterol. 2017, 112, 1016–1030. [Google Scholar] [CrossRef] [PubMed]
- Hillner, B.E.; Siegel, B.A.; Shields, A.F.; Liu, D.; Gareen, I.; Hanna, L.; Ba, S.H.S.; Coleman, R.E. The impact of positron emission tomography (PET) on expected management during cancer treatment. Cancer 2008, 115, 410–418. [Google Scholar] [CrossRef] [PubMed]
- Eddy, D.M. Screening for colorectal cancer. Ann. Intern. Med. 1990, 113, 373–384. [Google Scholar] [CrossRef]
- Carethers, J.M.; Jung, B.H. Genetics and genetic biomarkers in sporadic colorectal cancer. Gastroenterology 2015, 149, 1177–1190.e3. [Google Scholar] [CrossRef] [PubMed]
- Rawson, J.B.; Bapat, B. Epigenetic biomarkers in colorectal cancer diagnostics. Expert Rev. Mol. Diagn. 2012, 12, 499–509. [Google Scholar] [CrossRef]
- Chen, E.; Xu, X.; Liu, T. Hereditary nonpolyposis colorectal cancer and cancer syndromes: Recent basic and clinical discoveries. J. Oncol. 2018, 2018, 1–11. [Google Scholar] [CrossRef]
- Winawer, S. Colorectal cancer screening and surveillance: Clinical guidelines and rationale—Update based on new evidence. Gastroenterology 2003, 124, 544–560. [Google Scholar] [CrossRef]
- Vernon, S.W. Participation in colorectal cancer screening: A review. J. Natl. Cancer Inst. 1997, 89, 1406–1422. [Google Scholar] [CrossRef] [PubMed]
- US Preventive Services Task Force; Bibbins-Domingo, K.; Grossman, D.C.; Curry, S.J.; Davidson, K.W.; Epling, J.W., Jr.; García, F.A.R.; Gillman, M.W.; Harper, D.M.; Kemper, A.R.; et al. Screening for colorectal cancer. JAMA 2016, 315, 2564–2575. [Google Scholar] [CrossRef] [PubMed]
- Ventura, L.; Mantellini, P.; Grazzini, G.; Castiglione, G.; Buzzoni, C.; Rubeca, T.; Sacchettini, C.; Paci, E.; Zappa, M. The impact of immunochemical faecal occult blood testing on colorectal cancer incidence. Dig. Liver Dis. 2014, 46, 82–86. [Google Scholar] [CrossRef]
- Jacob, B.J.; Moineddin, R.; Sutradhar, R.; Baxter, N.N.; Urbach, D.R. Effect of colonoscopy on colorectal cancer incidence and mortality: An instrumental variable analysis. Gastrointest. Endosc. 2012, 76, 355–364.e1. [Google Scholar] [CrossRef] [PubMed]
- Pignone, M.; Rich, M.; Teutsch, S.M.; Berg, A.O.; Lohr, K.N. Screening for colorectal cancer in adults at average risk: A summary of the evidence for the U.S. Preventive Services Task Force. Ann. Intern. Med. 2002, 137, 132–141. [Google Scholar] [CrossRef]
- Nadeau, M.; Walaszek, A.; Perdue, D.G.; Rhodes, K.L.; Haverkamp, D.; Forster, J. Influences and Practices in Colorectal Cancer Screening Among Health Care Providers Serving Northern Plains American Indians. Prev. Chronic. Dis. 2016, 13, E167. [Google Scholar] [CrossRef]
- Lin, J.S.; Piper, M.A.; Perdue, L.A.; Rutter, C.M.; Webber, E.M.; O’Connor, E.; Smith, N.; Whitlock, E.P. Screening for colorectal cancer. JAMA 2016, 315, 2576–2594. [Google Scholar] [CrossRef]
- Holme, Ø.; Bretthauer, M.; Fretheim, A.; Odgaard-Jensen, J.; Hoff, G.; Odgaard-Jensen, J. Flexible sigmoidoscopy versus faecal occult blood testing for colorectal cancer screening in asymptomatic individuals. Cochrane Database Syst. Rev. 2013. [Google Scholar] [CrossRef]
- Whitlock, E.P.; Lin, J.S.; Liles, E.; Beil, T.L.; Fu, R. Screening for colorectal cancer: A targeted, updated systematic review for the U.S. preventive services task force. Ann. Intern. Med. 2008, 149, 638–658. [Google Scholar] [CrossRef]
- Swartz, A.W.; Eberth, J.; Josey, M.M.J.; Strayer, S.M. Reanalysis of All-Cause mortality in the U.S. Preventive services task force 2016 evidence report on colorectal cancer screening. Ann. Intern. Med. 2017, 167, 602. [Google Scholar] [CrossRef]
- Wallace, M.B. New strategies to improve polypectomy during colonoscopy. Gastroenterol. Hepatol. 2017, 13, 1–12. [Google Scholar]
- D’Andrea, E.; Ahnen, D.J.; Sussman, D.A.; Najafzadeh, M. Quantifying the impact of adherence to screening strategies on colorectal cancer incidence and mortality. Cancer Med. 2020, 9, 824–836. [Google Scholar] [CrossRef]
- Ko, C.W. Colonoscopy risks: What is known and what are the next steps? Gastroenterology 2018, 154, 473–475. [Google Scholar] [CrossRef]
- Frazier, A.L.; Colditz, G.; Fuchs, C.S.; Kuntz, K.M. Cost-effectiveness of screening for colorectal cancer in the general population. JAMA 2000, 284, 1954–1961. [Google Scholar] [CrossRef] [PubMed]
- US Preventive Services Task Force. Screening for colorectal cancer: Recommendation and rationale. Ann. Intern. Med. 2002, 137, 129–131. [Google Scholar] [CrossRef]
- Kim, S.Y.; Kim, H.-S.; Park, H.J. Adverse events related to colonoscopy: Global trends and future challenges. World J. Gastroenterol. 2019, 25, 190–204. [Google Scholar] [CrossRef] [PubMed]
- Hong, N.; Park, S.H. CT colonography in the diagnosis and management of colorectal cancer: Emphasis on pre- and post-surgical evaluation. World J. Gastroenterol. 2014, 20, 2014–2022. [Google Scholar] [CrossRef]
- Screening for Colorectal Cancer: Strategies in Patients at Average Risk-UpToDate. Available online: https://www.uptodate.com/contents/screening-for-colorectal-cancer-strategies-in-patients-at-average-risk (accessed on 27 March 2018).
- De Gonzalez, A.B.; Kim, K.P.; Yee, J. CT Colonography: Perforation rates and potential radiation risks. Gastrointest. Endosc. Clin. 2010, 20, 279–291. [Google Scholar] [CrossRef] [PubMed]
- Halligan, S.; Wooldrage, K.; Dadswell, E.; Kralj-Hans, I.; Von Wagner, C.; Edwards, R.; Yao, G.; Kay, C.; Burling, D.; Faiz, O.; et al. Computed tomographic colonography versus barium enema for diagnosis of colorectal cancer or large polyps in symptomatic patients (SIGGAR): A multicentre randomised trial. Lancet 2013, 381, 1185–1193. [Google Scholar] [CrossRef]
- Ijspeert, J.E.G.; Nolthenius, C.J.T.; Kuipers, E.J.; van Leerdam, M.E.; Nio, C.Y.; Thomeer, M.G.J.; Biermann, K.; van de Vijver, M.; Dekker, E.; Stoker, J. CT-Colonography vs. colonoscopy for detection of High-Risk sessile serrated polyps. Am. J. Gastroenterol. 2016, 111, 516–522. [Google Scholar] [CrossRef] [PubMed]
- Stoop, E.M.; de Haan, M.C.; de Wijkerslooth, T.R.; Bossuyt, P.M.; van Ballegooijen, M.; Nio, C.Y.; van de Vijver, M.; Biermann, K.; Thomeer, M.; van Leerdam, M.E.; et al. Participation and yield of colonoscopy versus non-cathartic CT colonography in population-based screening for colorectal cancer: A randomised controlled trial. Lancet Oncol. 2012, 13, 55–64. [Google Scholar] [CrossRef]
- Rex, D.K.; Adler, S.N.; Aisenberg, J.; Burch, W.C.; Carretero, C.; Chowers, Y.; Fein, S.A.; Fern, S.E.; Sainz, I.F.-U.; Fich, A.; et al. Accuracy of capsule colonoscopy in detecting colorectal polyps in a screening population. Gastroenterology 2015, 148, 948–957.e2. [Google Scholar] [CrossRef] [PubMed]
- Spada, C.; Hassan, C.; Munoz-Navas, M.; Neuhaus, H.; Deviere, J.; Fockens, P.; Coron, E.; Gay, G.; Toth, E.; Riccioni, M.E.; et al. Second-generation colon capsule endoscopy compared with colonoscopy. Gastrointest. Endosc. 2011, 74, 581–589.e1. [Google Scholar] [CrossRef]
- El Zoghbi, M.; Cummings, L.C. New era of colorectal cancer screening. World J. Gastrointest. Endosc. 2016, 8, 252–258. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-urien, I.; Carretero, C.; Gay, G.; Delvaux, M.; Lapalus, M.G.; Ponchon, T.; Neuhaus, H.; Philipper, M.; Costamagna, G.; Riccioni, M.E.; et al. Capsule endoscopy for the detection of polyps and cancer. NEJM 2009, 361, 264–270. [Google Scholar]
- Glick, S. Double-Contrast barium enema for colorectal cancer screening. Am. J. Roentgenol. 2000, 174, 1529–1537. [Google Scholar] [CrossRef]
- Sosna, J.; Sella, T.; Sy, O.; Lavin, P.T.; Eliahou, R.; Fraifeld, S.; Libson, E. Critical analysis of the performance of Double-Contrast barium enema for detecting colorectal polyps ≥ 6 mm in the era of CT colonography. Am. J. Roentgenol. 2008, 190, 374–385. [Google Scholar] [CrossRef]
- Allison, J.E.; Fraser, C.G.; Halloran, S.P.; Young, G. Population screening for colorectal cancer means getting FIT: The past, present, and future of colorectal cancer screening using the fecal immunochemical test for hemoglobin (FIT). Gut Liver 2014, 8, 117–130. [Google Scholar] [CrossRef]
- Allison, J.E.; Sakoda, L.C.; Levin, T.R.; Tucker, J.P.; Tekawa, I.S.; Cuff, T.; Pauly, M.P.; Shlager, L.; Palitz, A.M.; Zhao, W.K.; et al. Screening for colorectal neoplasms with new fecal occult blood tests: Update on performance characteristics. J. Natl. Cancer Inst. 2007, 99, 1462–1470. [Google Scholar] [CrossRef]
- Baldwin, L.-M.; Schneider, J.L.; Schwartz, M.; Rivelli, J.S.; Green, B.B.; Petrik, A.F.; Coronado, G.D. First-year implementation of mailed FIT colorectal cancer screening programs in two Medicaid/Medicare health insurance plans: Qualitative learnings from health plan quality improvement staff and leaders. BMC Health Serv. Res. 2020, 20, 132. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.K.; Liles, E.G.; Bent, S.; Levin, T.R.; Corley, D.A. Accuracy of fecal immunochemical tests for colorectal cancer. Ann. Intern. Med. 2014, 160, 171–181. [Google Scholar] [CrossRef] [PubMed]
- Zauber, A.G. The impact of screening on colorectal cancer mortality and incidence: Has it really made a difference? Dig. Dis. Sci. 2015, 60, 681–691. [Google Scholar] [CrossRef] [PubMed]
- Libby, G.; Fraser, C.G.; Carey, F.A.; Brewster, D.; Steele, R.J.C. Occult blood in faeces is associated with all-cause and non-colorectal cancer mortality. Gut 2018, 67, 2116–2123. [Google Scholar] [CrossRef]
- Geiger, T.M.; Ricciardi, R. Screening options and recommendations for colorectal cancer. Clin. Colon Rectal Surg. 2009, 22, 209–217. [Google Scholar] [CrossRef]
- Nadel, M.R.; Shapiro, J.A.; Klabunde, C.N.; Seeff, L.C.; Uhler, R.; Smith, R.A.; Ransohoff, D.F. A national survey of primary care physicians’ methods for screening for fecal occult blood. Ann. Intern. Med. 2005, 142, 86–94. [Google Scholar] [CrossRef]
- Griffith, C.D.M.; Saunders, J.H. False-negative results of Hemoccult test in colorectal cancer. BMJ 1981, 283, 795. [Google Scholar] [CrossRef][Green Version]
- Ahlquist, D.A. Multi-Target stool DNA test: A new high bar for noninvasive screening. Dig. Dis. Sci. 2015, 60, 623–633. [Google Scholar] [CrossRef]
- Brenner, H.; Werner, S.; Chen, H. Multitarget stool DNA testing for colorectal-cancer screening. N. Engl. J. Med. 2014, 371, 184–188. [Google Scholar] [CrossRef]
- Wang, X.; Wang, D.; Zhang, H.; Feng, M.; Wu, X. Genome-wide analysis of DNA methylation identifies two CpG sites for the early screening of colorectal cancer. Epigenomics 2020, 12, 37–52. [Google Scholar] [CrossRef]
- Ahlquist, D.A. Molecular detection of colorectal neoplasia. Gastroenterology 2010, 138, 2127–2139. [Google Scholar] [CrossRef] [PubMed]
- Jaffe, R.M.; Kasten, B.; Young, D.S.; MacLowry, J.D. False-Negative Stool Occult Blood Tests Caused by Ingestion of Ascorbic Acid (Vitamin C). Ann. Intern. Med. 1975, 83, 824–826. [Google Scholar] [CrossRef] [PubMed]
- Imperiale, T.F.; Ransohoff, D.F.; Itzkowitz, S.H.; Levin, T.R.; Lavin, P.; Lidgard, G.P.; Ahlquist, D.A.; Berger, B.M. Multitarget stool DNA testing for colorectal-cancer screening. N. Engl. J. Med. 2014, 370, 1287–1297. [Google Scholar] [CrossRef]
- Ladabaum, U.; Mannalithara, A. Comparative effectiveness and cost effectiveness of a multitarget stool DNA test to screen for colorectal neoplasia. Gastroenterology 2016, 151, 427–439.e6. [Google Scholar] [CrossRef]
- Farzanehfar, M.; Pakbaz, B.; Jabini, R.; Soltani, N.; Ayatollahi, H. Quantitative study of vimentin gene methylation in stool samples for colorectal cancer screening. J. Adv. Pharm. Technol. Res. 2019, 10, 121–125. [Google Scholar] [CrossRef]
- Itzkowitz, S.H.; Jandorf, L.; Brand, R.; Rabeneck, L.; Schroy, P.; Sontag, S.; Johnson, D.; Skoletsky, J.; Durkee, K.; Markowitz, S.; et al. Improved fecal DNA test for colorectal cancer screening. Clin. Gastroenterol. Hepatol. 2007, 5, 111–117. [Google Scholar] [CrossRef] [PubMed]
- Ned-Sykes, R.; Melillo, S.; Marrone, M. Fecal DNA testing for Colorectal Cancer Screening: The ColoSure™ test. PLoS Curr. 2011, 3, RRN1220. [Google Scholar] [CrossRef]
- Sellin, M.E.; Sandblad, L.; Stenmark, S.; Gullberg, M. Deciphering the rules governing assembly order of mammalian septin complexes. Mol. Biol. Cell 2011, 22, 3152–3164. [Google Scholar] [CrossRef]
- Kim, M.S.; Froese, C.D.; Estey, M.P.; Trimble, W.S. SEPT9 occupies the terminal positions in septin octamers and mediates polymerization-dependent functions in abscission. J. Cell Biol. 2011, 195, 815–826. [Google Scholar] [CrossRef]
- Song, L.; Li, Y. Methylated Sept9 gene is a sensitive biomarker for all stages of colorectal cancer. Colorec Cancer 2015, 1, 7. [Google Scholar] [CrossRef]
- Church, T.R.; Wandell, M.; Lofton-Day, C.; Mongin, S.J.; Burger, M.; Payne, S.R.; Castaños-Vélez, E.; Blumenstein, B.A.; Rösch, T.; Osborn, N.K.; et al. Prospective evaluation of methylatedSEPT9in plasma for detection of asymptomatic colorectal cancer. Gut 2014, 63, 317–325. [Google Scholar] [CrossRef] [PubMed]
- Warren, J.D.; Xiong, W.; Bunker, A.M.; Vaughn, C.P.; Furtado, L.V.; Roberts, W.L.; Fang, J.C.; Samowitz, W.S.; Heichman, K.A. Septin 9 methylated DNA is a sensitive and specific blood test for colorectal cancer. BMC Med. 2011, 9, 133. [Google Scholar] [CrossRef] [PubMed]
- Constâncio, V.; Nunes, S.P.; Henrique, R.; Jerónimo, C. DNA Methylation-Based Testing in liquid biopsies as detection and prognostic biomarkers for the four major cancer types. Cells 2020, 9, 624. [Google Scholar] [CrossRef]
- Wu, Y.; Wan, X.; Jia, G.; Xu, Z.; Tao, Y.; Song, Z.; Du, T. Aberrantly methylated and expressed genes as prognostic epigenetic biomarkers for colon cancer. DNA Cell Biol. 2020, 39, 1961–1969. [Google Scholar] [CrossRef] [PubMed]
- Tham, C.; Chew, M.; Soong, R.; Lim, J.; Ang, M.; Tang, C.; Zhao, Y.; Ong, S.Y.K.; Liu, Y. Postoperative serum methylation levels ofTAC1andSEPT9are independent predictors of recurrence and survival of patients with colorectal cancer. Cancer 2014, 120, 3131–3141. [Google Scholar] [CrossRef]
- Jiang, D.; Xie, X.; Lu, Z.; Liu, L.; Qu, Y.; Wu, S.; Li, Y.; Li, G.; Wang, H.; Xu, G. Establishment of a colorectal cancer-related microrna-mrna regulatory network by microarray and bioinformatics. Front. Genet. 2020, 11, 560186. [Google Scholar] [CrossRef]
- Zhang, H.; Qi, J.; Wu, Y.-Q.; Zhang, P.; Jiang, J.; Wang, Q.-X.; Zhu, Y.-Q. Accuracy of early detection of colorectal tumours by stool methylation markers: A meta-analysis. World J. Gastroenterol. 2014, 20, 14040–14050. [Google Scholar] [CrossRef] [PubMed]
- Oh, T.J.; Oh, H.I.; Seo, Y.Y.; Jeong, D.; Kim, C.; Kang, H.W.; Han, Y.D.; Chung, H.C.; Kim, N.K.; An, S. Feasibility of quantifying SDC2 methylation in stool DNA for early detection of colorectal cancer. Clin. Epigenetics 2017, 9, 126. [Google Scholar] [CrossRef]
- Leon, S.A.; Shapiro, B.; Sklaroff, D.M.; Yaros, M.J. Free DNA in the serum of cancer patients and the effect of therapy. Cancer Res. 1977, 37, 646–650. [Google Scholar]
- Rasmussen, S.L.; Krarup, H.B.; Sunesen, K.G.; Pedersen, I.S.; Madsen, P.H.; Thorlacius-Ussing, O. Hypermethylated DNA as a biomarker for colorectal cancer: A systematic review. Color. Dis. 2016, 18, 549–561. [Google Scholar] [CrossRef]
- Bagheri, H.; Mosallaei, M.; Bagherpour, B.; Khosravi, S.; Salehi, A.R.; Salehi, R. TFPI2 and NDRG4 gene promoter methylation analysis in peripheral blood mononuclear cells are novel epigenetic noninvasive biomarkers for colorectal cancer diagnosis. J. Gene Med. 2020, 22, e3189. [Google Scholar] [CrossRef] [PubMed]
- Bin Lee, B.; Lee, E.J.; Jung, E.H.; Chun, H.-K.; Chang, D.K.; Song, S.Y.; Park, J.; Kim, D.-H. Aberrant Methylation of APC, MGMT, RASSF2A, and Wif-1 Genes in Plasma as a Biomarker for Early Detection of Colorectal Cancer. Clin. Cancer Res. 2009, 15, 6185–6191. [Google Scholar] [CrossRef]
- Chen, X.; Gole, J.; Gore, A.; He, Q.; Lu, M.; Min, J.; Yuan, Z.; Yang, X.; Jiang, Y.; Zhang, T.; et al. Non-invasive early detection of cancer four years before conventional diagnosis using a blood test. Nat. Commun. 2020, 11, 1–10. [Google Scholar] [CrossRef]
- Link, A.; Balaguer, F.; Shen, Y.; Nagasaka, T.; Lozano, J.J.; Boland, C.R.; Goel, A. Fecal MicroRNAs as novel biomarkers for colon cancer screening. Cancer Epidemiol. Biomark. Prev. 2010, 19, 1766–1774. [Google Scholar] [CrossRef]
- Wang, S.; Xiang, J.; Li, Z.; Lu, S.; Hu, J.; Gao, X.; Yu, L.; Wang, L.; Wang, J.; Wu, Y.; et al. A plasma microRNA panel for early detection of colorectal cancer. Int. J. Cancer 2015, 136, 152–161. [Google Scholar] [CrossRef] [PubMed]
- Müller, H.M.; Oberwalder, M.; Fiegl, H.; Morandell, M.; Goebel, G.; Zitt, M.; Mühlthaler, M.; Öfner, D.; Margreiter, R.; Widschwendter, M. Methylation changes in faecal DNA: A marker for colorectal cancer screening? Lancet 2004, 363, 1283–1285. [Google Scholar] [CrossRef]
- Turchinovich, A.; Weiz, L.; Langheinz, A.; Burwinkel, B. Characterization of extracellular circulating microRNA. Nucleic Acids Res. 2011, 39, 7223–7233. [Google Scholar] [CrossRef]
- Ng, E.K.-O.; Chong, W.W.S.; Jin, H.; Lam, E.K.Y.; Shin, V.Y.; Yu, J.; Poon, T.C.W.; Ng, S.S.M.; Sung, J.J.Y. Differential expression of microRNAs in plasma of patients with colorectal cancer: A potential marker for colorectal cancer screening. Gut 2009, 58, 1375–1381. [Google Scholar] [CrossRef]
- Zhang, H.; Zhu, M.; Shan, X.; Zhou, X.; Wang, T.; Zhang, J.; Tao, J.; Cheng, W.; Chen, G.; Li, J.; et al. A panel of seven-miRNA signature in plasma as potential biomarker for colorectal cancer diagnosis. Gene 2019, 687, 246–254. [Google Scholar] [CrossRef]
- Li, T.; Leong, M.H.; Harms, B.; Kennedy, G.; Chen, L. MicroRNA-21 as a potential colon and rectal cancer biomarker. World J. Gastroenterol. 2013, 19, 5615–5621. [Google Scholar] [CrossRef]
- Luo, X.; Stock, C.; Burwinkel, B.; Brenner, H. Identification and evaluation of plasma MicroRNAs for early detection of colorectal cancer. PLoS ONE 2013, 8, e62880. [Google Scholar] [CrossRef]
- Peng, Q.; Zhang, X.; Min, M.; Zou, L.; Shen, P.; Zhu, Y. The clinical role of microRNA-21 as a promising biomarker in the diagnosis and prognosis of colorectal cancer: A systematic review and meta-analysis. Oncotarget 2017, 8, 44893–44909. [Google Scholar] [CrossRef]
- James, J.; Riis, L.B.; Malham, M.; Høgdall, E.; Langholz, E.; Nielsen, B.S. MicroRNA biomarkers in IBD—Differential diagnosis and prediction of Colitis-Associated cancer. Int. J. Mol. Sci. 2020, 21, 7893. [Google Scholar] [CrossRef]
- Arroyo, J.; Chevillet, J.; Kroh, E.M.; Ruf, I.K.; Pritchard, C.C.; Gibson, D.F.; Mitchell, P.; Bennett, C.; Pogosova-Agadjanyan, E.L.; Stirewalt, D.L.; et al. Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc. Natl. Acad. Sci. USA 2011, 108, 5003–5008. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Paredes, M.; Esteller, M. Cancer epigenetics reaches mainstream oncology. Nat. Med. 2011, 17, 330–339. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Huang, S.-K.; Zhao, M.; Yang, M.; Zhong, J.-L.; Gu, Y.-Y.; Peng, H.; Che, Y.-Q.; Huang, C.-Z. Identification of a circulating microrna signature for colorectal cancer detection. PLoS ONE 2014, 9, e87451. [Google Scholar] [CrossRef]
- Baylin, S.B.; Jones, P.A. A decade of exploring the cancer epigenome—biological and translational implications. Nat. Rev. Cancer 2011, 11, 726–734. [Google Scholar] [CrossRef] [PubMed]
- Yoruker, E.E.; Holdenrieder, S.; Gezer, U. Potential of circulating nucleosome-associated histone modifications in cancer. Transl. Cancer Res. 2018, 7, S185–S191. [Google Scholar] [CrossRef]
- Rasmussen, L.; Christensen, I.J.; Herzog, M.; Micallef, J.; Nielsen, H.J. For the Danish Collaborative Group on Early Detection of Colorectal Cancer Circulating cell-free nucleosomes as biomarkers for early detection of colorectal cancer. Oncotarget 2017, 9, 10247–10258. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Almeida-Lousada, H.; Mestre, A.; Ramalhete, S.; Price, A.J.; de Mello, R.A.; Marreiros, A.D.; Neves, R.P.d.; Castelo-Branco, P. Screening for Colorectal Cancer Leading into a New Decade: The “Roaring ‘20s” for Epigenetic Biomarkers? Curr. Oncol. 2021, 28, 4874-4893. https://doi.org/10.3390/curroncol28060411
Almeida-Lousada H, Mestre A, Ramalhete S, Price AJ, de Mello RA, Marreiros AD, Neves RPd, Castelo-Branco P. Screening for Colorectal Cancer Leading into a New Decade: The “Roaring ‘20s” for Epigenetic Biomarkers? Current Oncology. 2021; 28(6):4874-4893. https://doi.org/10.3390/curroncol28060411
Chicago/Turabian StyleAlmeida-Lousada, Hélder, André Mestre, Sara Ramalhete, Aryeh J. Price, Ramon Andrade de Mello, Ana D. Marreiros, Ricardo Pires das Neves, and Pedro Castelo-Branco. 2021. "Screening for Colorectal Cancer Leading into a New Decade: The “Roaring ‘20s” for Epigenetic Biomarkers?" Current Oncology 28, no. 6: 4874-4893. https://doi.org/10.3390/curroncol28060411
APA StyleAlmeida-Lousada, H., Mestre, A., Ramalhete, S., Price, A. J., de Mello, R. A., Marreiros, A. D., Neves, R. P. d., & Castelo-Branco, P. (2021). Screening for Colorectal Cancer Leading into a New Decade: The “Roaring ‘20s” for Epigenetic Biomarkers? Current Oncology, 28(6), 4874-4893. https://doi.org/10.3390/curroncol28060411





