1. Introduction
Cervical cancer is the second most common malignancy among women with more than half a million new cases reported annually. It can be detected and prevented at its early stage by cytology screening test. Over the last six decades, cervical smear of Pap strains has played an important role in controlling cervical cancer epidemic by detecting pre-cancerous changes and providing guidance for early treatments. It was estimated that a Pap smear test can reduce mortality rate by 70% or more [
1]. Cytological slides of vaginal smear of Pap strains are microscopically examined at 400× magnification. With such magnification, cytologists must examine thousands of field-of-views (FOVs) per slide and, thereby, are limited to investigate a limited number of samples per day. Despite the seemingly simplicity in detection of abnormal cervical cells, experienced physicians are required for an accurate diagnosis and there is a lack of such healthcare resources in many developing countries. Moreover, even with experienced pathologists, it is a tedious task to examine the Pap smear slides through microscope and the detection of cervical cancer cells can often be missed, due to the small size of the cervical intraepithelial neoplasia, overlapping clump of cells or masking by blood mucus and artifacts.
In the light of these challenges, different computer-aided diagnosis technologies have been proposed [
2,
3,
4,
5] to reduce the workload of pathologists and improve the efficiency and accuracy of cervical cancer detection. The computer-assisted screening for cervical cancer cells involves typically two steps: segmentation [
6] of cytoplasm and nucleus, and classification [
7]. After the entire image is segmented into nuclei and cytoplasm, these small pieces are classified into abnormal and normal ones, and finally used for the diagnosis of cervical cancer. The segmentation quality is, therefore, vital for the extraction of cell features and the consequential classification results. This has been a subject of extensive investigation [
6,
8,
9,
10,
11,
12,
13,
14,
15] in the last two decades. Cell overlapping is one of the difficult issues for accurate cell segmentation and effective methods to separate overlapping cells have been proposed [
11,
16,
17,
18]. After successful segmentation into individual cells, the performance for cell classification relies on the extraction of morphological, structural, and contextual features and optimal selection of classifiers [
19]. Due to the complicated characteristics of cervical cells and pathological subtleties associated with cervical cancer development, traditional computer vision methods can only provide limited generality and efficiency for segmentation and feature extraction.
With the advent of deep learning based artificial intelligence (AI) technology in recent years, particularly, the development of convolutional neural networks (CNNs) has demonstrated its potential for improving automated cervical cancer screening and diagnosis. Deep learning excels at recognizing patterns and high-level sematic features in large volumes of data, extracting relationships between complex features in the images, and identifying characteristics that is difficult be perceived by the human brain. AI software can process vast number of images rapidly and has already been used to assist clinicians in level classification of skin cancer [
20], breast cancer detection [
21,
22], end-to-end lung cancer screening [
23] and prediction of colorectal cancer outcomes [
24]. Significant progress has also been made in automated cervical cancer classification. With these classification methods we can leverage the deep CNNs for effective extraction of deep and high-level features [
3,
7,
25,
26,
27,
28,
29,
30,
31] of the cells to achieve more accurate cancer screening.
Deep learning technology has also led to breakthroughs in object detection, which refers to the task of both identifying the locations and categorizing object instances in images. Object detection is a key ability required by most computer and robot vision systems. The cutting-edge research in the field has been making amazing progress in many directions, such as automatic car-driving and object tracking. Current object detection models can be divided into two main categories, proposal-driven two-stage detectors and proposal-free single stage detectors. For the two-stage detectors, such as faster region-based CNN (FRCNN) series [
32,
33,
34,
35], region-based fully convolution network (R-FCN) [
36] and feature pyramid network (FPN) [
33], the detection process is completed in two steps. First the algorithms produce sparse candidate proposals, and then further refine the coordinates and classify these candidate proposals. The two-stage methods generally have higher accuracy at the cost of speed. In contrast, the one-stage detectors generate the object’s category labels and locations directly and have advantage in speed, such as YOLO [
37] for real-time object detection, single shot multibox detector (SSD) series [
38] and RetinaNet [
39]. More recently, a new category of anchor-free single-stage detectors has emerged [
40], which has completely abandoned the anchor mechanism.
The object detection methods have also begun to find applications for cervical cancer detection in cervical cytology images. At present, there are few network architectures designed specifically for the specific task of cervical cancer detection [
41,
42,
43,
44,
45,
46]. Xu et al. [
45] used the generic Faster RCNN for the detection of abnormal cells in cervical smear images scanned at 20× and showed that detection of various abnormal cells was feasible. Zhang et al. [
46] tested an R-FCN model for cervical cancer screening of liquid-based cytology (LBC) images. The performance evaluation was based on an interesting concept called hit degree which ignores the precise intersection-over-union (IOU) threshold, and a hit recall was counted if a ground truth box was hit by any detection result box. Accordingly, the precise positions of the cell nuclei are less important because the cells tend to form clumps in the Pap smear slides. Tan et al. [
44] also used Faster RCNN architecture for cervical cancer cell detection in ThinPrep cytologic test (TCT) images scanned with a seamless slider at up to 400× and achieved AUC of 0.67. More recently, Ma et al. [
42] proposed an improved Faster RCNN-FPN architecture for cervical cancer detection in cropped patches out of positive Pap smear images. They designed a lightweight booster consisting of a refinement and spatial-aware module, aimed to enhance feature details and spatial context information.
As discussed above, the automated detection of cervical cancer cells in the cytology images is a multi-task process involving feature extraction, cell localization and classification. Although extensive research efforts in the past have made significant progress in automated classification and detection of the abnormal cervical cells, some bottle neck issues still remain open. Particularly relevant concerns include the following: (1) the abnormalities in the cancerous cells are subtle and complex; (2) Cells in the cervical Pap smear images exist in different sizes and their geometric shapes can be obscured due to clump formation and overlapping with artifacts; (3) The ROI pooling/aligning processes in the object detection algorithms tend to enhance the local receptive fields and lose global context information.
Aimed at mitigating these potential limiting factors for more accurate detection of abnormal cells in cervical cytology images, in this study, we introduced two functional extensions for one of the top-performing object detectors, the Faster RCNN-FPN framework: One is the infusion of deformable convolution network (DCN) [
47,
48,
49,
50,
51] in the last three stages of the bottom-up pathway of the FPN and the other is adding a global context aware (GCA) module [
52] alongside the RPN. A unique advantage of the Faster RCNN-FPN architecture is its inherent multi-scale nature in feature extraction and can facilitate the cell detection of different sizes. Infusion of the deformable convolution layers into the later three stages of the FPN structure can explicitly learn the geometric offset information of the objects associated with shape transformations. We added a GCA module into the Faster RCNN-FPN to strengthen the spatial correlation between the background and foreground. In the GCA module, the feature pyramid and attention strategies are used for global feature extraction and feature refinement, respectively. We leveraged the extracted global features as attention maps for contextual modulation to improve the cell detection performance. Moreover, we utilized various conventional computer-vision techniques for imaging process to preprocess the negative image samples and tag some of the representative cells aimed to enhance the subtle feature differences between the negative and positive cancerous cells. We conducted extensive experiments using the dataset from the “Digital Human Body” (DHB) Vision Challenge (tianchi.aliyun.com/competition/entrance/231757/information (accessed on 1 February 2021)) with the proposed deformable and global context aware (DGCA) Faster RCNN-FPN architecture (hereon abbreviated as DGCA-RCNN). Besides the exhaustive test, we evaluated the performance of the proposed DGCA-RCNN framework and conducted systematic performance comparison with five other state-of-the-art object detectors including ATSS [
53], RetinaNet [
39], Faster RCNN-FPN [
34], double-head [
54], and Cascade RCNN [
32]. We also carried out ablation analysis of the extensions in the DGCA-RCNN framework to assess their functional importance for the detector’s performance.
2. Materials and Methods
2.1. The Proposed Framework for Cervical Cancer Cell Detection
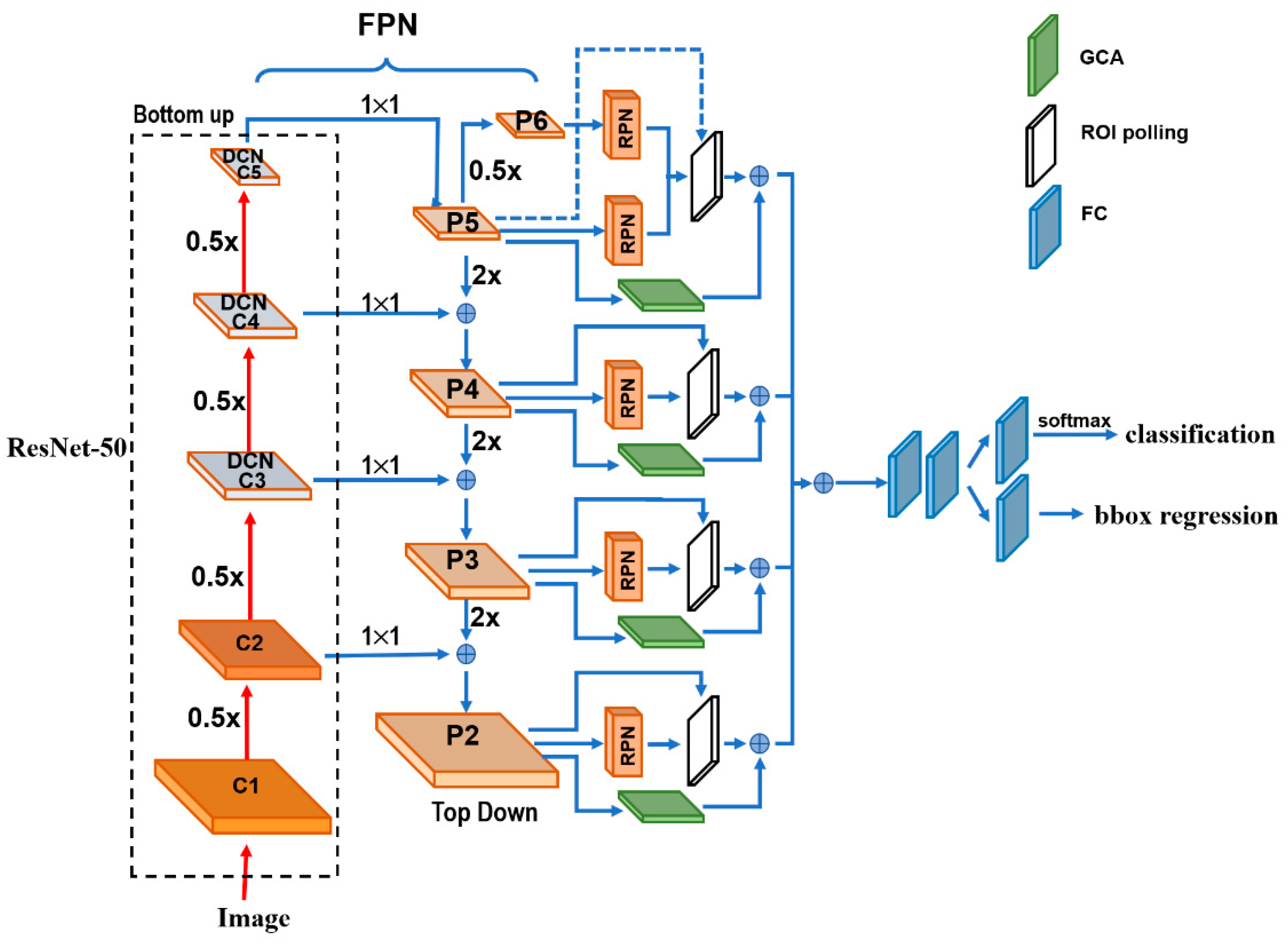
Figure 1 depicts the architecture details of the proposed DGCA-RCNN framework. It is based on the faster RCNN-FPN architecture consisting of image input, feature pyramid network (FPN) for feature extraction, feature map generation with region proposal network (RPN), classifier, and bonding box regressor. As shown in
Figure 1, the FPN consists of a bottom-up and top-down pathways. The bottom-up pathway is a deep convolutional network hierarchy for feature extraction with successively decreasing spatial dimension. As more high-level features are extracted, the semantic value for each layer increases. The bottom layers (C1) are not included for object detection because of their limited semantic value and large dimension. They are in high resolution, require too much memory and can result in significant slowdown. The framework only uses the upper layers for detection and, therefore, performs much worse for smaller objects. However, FPN also provides a top-down pathway to construct higher resolution layers from a semantic rich layer using up-samplings. While the reconstructed layers are semantically strong, locations are not precise after all the down- and up-samplings. Therefore, lateral connections between the reconstructed layers and the corresponding feature maps are utilized to generate pixelwise addition to improve the localization precision. They also act as skip connections to facilitate training.
In this study, we used ResNet-50 to construct the bottom-up pathway. It composes of 5 convolution modules (C1–C5) and each has multiple convolution layers. From C1 to C5 the spatial dimension is reduced by half at each stage. We apply a 1 × 1 convolution filter to reduce the C5 channel depth to 256-d (P5). This becomes the first feature map layer used for object prediction. Along the top-down route, the previous layer is up-sampled by a factor of 2 using the nearest neighbor up-sampling method. The up-sampled P5 was added with the 1 × 1 filtered C4 pixel-wise to generate P4. The same process is repeated to generate P3 and P2. In this way, the FPN network utilizes its inherent multi-scale pyramidal hierarchy of deep convolutional networks to construct feature pyramids and make independent predictions at different levels (P2, P3, P4, P5 and P6) for multi-scale object detection. The RPN network loops through the different prediction levels and makes full use of the feature maps of different scales at 32 × 32, 64 × 64, 128 × 128, 256 × 256 and 512 × 512, respectively. Furthermore, three aspect ratios of 1:1, 1:2 and 2:1 are utilized at each level. The RPN is trainable from end-to-end to generate simultaneously bounding-box proposals and objectness scores at each position. These regions may contain target objects and are sent to the subsequent classifier network to produce the final classification and refined anchor positions. Since the same classifier/regressor model is shared among the region proposals from the feature map pyramid of different levels, the region of interest (ROI) pooling layer uses max pooling to convert each ROI from the RPN into a feature map of fixed size at 7 × 7.
2.2. Extensions to the Faster RCNN-FPN Architecture
The regular grids are used in the input feature maps for the convolution operations and CNNs are limited in modeling geometric transformations. Introducing deformable convolutional layers can add 2D offsets to the regular-grid sampling locations in the standard convolution and can facilitate the detection of the geometrically transformed objects without additional supervision [
55]. Considering the geometrical variations of diploids in the cervical smear images, we replaced the last convolution layers in C3, C4 and C5 of the ResNet-50 [
56] backbone by deformable convolutional layers in the proposed framework. To optimize the performance of the network and the efficiency of limited computation resource, the deformable convolution layers are usually implemented in C3–C5 of the FPN. The rationale is that the size of the feature maps in C1 and C2 are relatively large, computationally demanding, and not very sensitive to deformation effect. With the extension we can obtain the field offsets to model the various geometrical deformations of the cells on top of their ordinary feature maps. During training, both the convolutional kernels for generating the output features and the field offsets can be learned simultaneously. Therefore, we expect to improve the generality of the model in the aspects of scalability, geometry transformation and cell deformation.
As discussed above, ROI Pooling/Aligning is an indispensable process for the Faster RCNN-FPN architecture. It is used to rescale the object proposals cropped from the feature pyramid to generate a fixed-size feature map. However, these cropped feature maps of dominant local receptive fields possess very weak global context information. To alleviate this, we infused an off-the-shelf Global Context Aware (GCA) module to strengthen the spatial correlation between the background and foreground by fusing the global context information through attention mechanism. We leveraged the extracted global features as attention maps for contextual modulation to improve the cell detection. In the end, the GCA module also utilizes a max pooling to convert the global context features at different stages in the top-down route of FPN into the fixed size of 7 × 7 and merges them pixel-wise with the ROIs from the ROI pooling layer. The reference information from the background may help decide the spatial and category relationship between the targets and global contexts.
2.3. The Loss Functions
The loss function of the detection algorithm can be divided into two parts, one is the RPN loss, and the other is the loss of the detection network. To evaluate the RPN loss, we used the IOU metric which is defined as the ratio between the area of overlap and the area encompassed jointly by both the predicted bounding box and the ground-truth bounding box. Depending on the associated IOU each anchor is assigned a binary class label. If the IOU for a given anchor is >0.7 or <0.3, positive or negative label will be signed to the anchor, respectively. The detection model is optimized for a loss combining the two tasks: classification and localization. Therefore, the loss (
L) function sums up the cost of classification (
Lcls) and bounding box prediction (
Lbox):
where
Lcls is the log loss function over two classes (positive and negative), as we can easily translate a multi-class classification into a binary classification by predicting a sample being a target object versus not.
The bounding box loss
Lbox should measure the difference between
and
using a robust loss function. The smooth
L1 loss is adopted here as usual, and it is claimed to be less sensitive to outliers.
Other symbols’ definitions in the above equations are provided in
Table 1.
2.4. The Dataset and Preprocessing Pipeline
For the study we used the cervical cytology image dataset from “Digital Human Body” (DHB) Vision Challenge-Intelligent Diagnosis of Cervical Cancer Risk provided by the Alibaba Cloud TianChi Company (
https://tianchi.aliyun.com/competition/entrance/231757/information?scm=20140722.184.2.173 (accessed on 1 February 2021)). The original images were in KFB format and acquired with digital scanning under 20× magnification. Typical file size is in the range of 300–400 MB. The dataset contains 500 positive and 300 negative whole slide images (WSI) of 40,000 × 40,000. The positive WSIs have marked ROIs where the positions of 5414 lesions with abnormal squamous epithelial cells were labeled. The coordinate and size information for the ROIs were provided in an associated list document in json format. The abnormal squamous epithelial cells include mainly four types: atypical squamous cells (ASC-US) which cannot be clearly defined, low-grade intraepithelial lesions (LSIL), atypical squamous cells (ASC-H) which tend to have high intraepithelial cells, and high intraepithelial lesions (HIS). It is notable that there is no guarantee that there is no abnormal squamous epithelial cell outside the marked ROIs for the positive WSIs.
We utilized a library package provided by Ningbo Jiangfeng Konfoong Bioinformation Tech Co., Ltd (
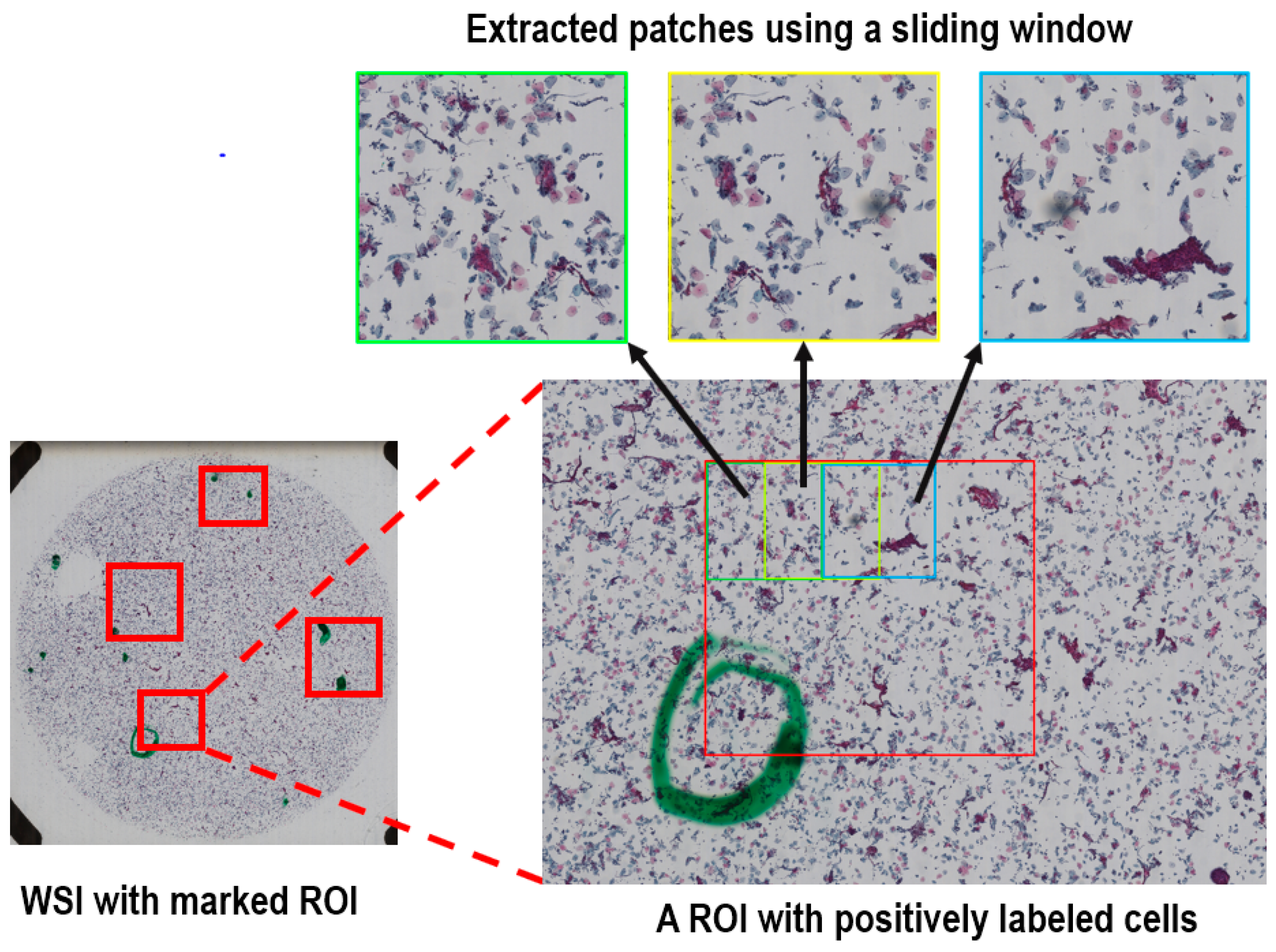
https://www.exporthub.com/eh-ningbo-konfoong-bioinformation-tech-co-ltd/ (accessed on 1 February 2021)) to convert from KFB to PNG format which has 3-channel RGB and 8-bit depth in each channel. Due to the limited GPU memory, the WSIs cannot be fed into the model directly and were cropped into smaller patches. As illustrated in
Figure 2, for the training dataset we cropped all the marked ROIs in each positive WSI using a sliding window method to extract systematically patches of 2000 × 2000 out of each ROI. The overlap of the sliding window was 50% in both directions. On average, about 20 patches were generated per positive sample.
For the negative samples without any specified ROI and the test dataset of the positive samples, we used the same sliding window method to crop the entire region within the radius of 20,000 voxels from the center of each WSI, because a statistical analysis of the marked ROIs for the positive samples showed that all positively labeled cells were within this range. There are rarely any cells at the edge of the images.
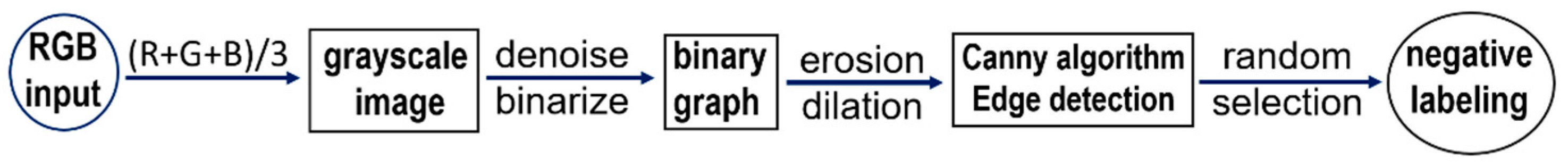
To enhance the features in the negative image samples, we applied computer-vision techniques to label some of the negative cells. The negative tagging can improve the performance of the model and further reduce the false positive rate. As schematically illustrated in
Figure 3, the procedure to label the negative cells included the following steps: Firstly, the RGB images were color averaged to generate grayscale images which were denoised using a series of filters. Then corrosion and dilation operations were applied to the binarized images before edge detection using the canny algorithm. Finally, for each of the preprocessed image patch a few cells of appropriate sizes were selected in a pseudo-random fashion and labeled as negative cells. The selection of the negative cells was carried by using a random number generator to attain an even distribution for the labeled negative cells after ranking the edge-detected negative cells according to their areas. For quality assurance the labeled negative samples were manually inspected to verify the exclusion of artifacts and background. An example set of the labeled cells (both negative and positive) are provided in
Figure 4 in magnified display. The total number of the extracted image patches with negatively labeled cells was controlled to be the same as that for the positively labeled image patches (see
Table 2).
To assess the performance of the trained model, we picked 80 positive and negative samples as the test dataset in a pseudo-random fashion. The rest of the dataset including both the positive and negative cases were used to train the proposed DGCA-RCNN model. The test dataset was also cropped by using the same sliding window described above to crop systematically each WSI within the radius of 20,000 voxels. The details of training and test datasets are summarized in
Table 2.
We performed the standard data augmentation procedure by adding image rotations and adjustment of the overall brightness. Rotations were performed three times on each image with a step size of 90°. Rotation may slightly alter the image quality but should not change the abnormality or normality of the cells. The overall image brightness was changed by about ±20%. The slightly darkening or brightening the entire FOV should not affect the detection results. Changing the brightness of the images can replicate the clinical conditions of different image qualities and stain settings in preparing the slides. Zero-mean normalization to attain centralization of the input data was also implemented for each RGB channel. This can improve the generality of the model by reducing the interference from settings, such as the staining conditions and imaging equipment.
2.5. The Implementation and Training Procedure of the Models
We implement the DGCA-RCNN model using PyTorch (version 1.8.1). for the Python environment (version 3.5). The training and test of the model were carried out on a GPU server equipped with 4 NVIDIA TITAN X (Pascal) with 12 GB GPU memory. For comparison, based on the same backbone of ResNet50, we also implemented other five closely related the state-of-the-art object detection models developed in recent years including ATSS [
53], RetinaNet [
39], Cascade RCNN [
32], Double head [
54], and Faster RCNN-FPN [
57]. We compared these models by strictly ruling out all the implementation inconsistencies between them.
Cross-validation procedure was implemented by further splitting the training-set into three folds and the model converged after about 12,000 iterations. To avoid over-fitting and prevent model selection bias, the cross-validation procedure was applied with a nested five-folds outer loop. After obtaining a score for each parameter set using the inner loop, we trained the model using the best parameter set from the three-folds inner loop on the whole data assigned to the inner loop without further manipulation of the parameter grid. The final model was selected according to the performance from the outer loop. We used the alternating training strategy to train the model, the RPN was first trained to generate the initial region proposals. The weights of the shared convolutional layers were initialized based on the pre-trained model using ImageNet. The other weights of the RPN were initialized randomly. The generated region proposals by the RPN were then used to train the Faster RCNN model. The weights of the shared convolutional layers were then initialized with the tuned weights by the RPN. The other Faster R-CNN weights were initialized randomly. When the weights for Faster RCNN and the shared layers were tuned, the tuned weights in the shared layers were again used to train the RPN, and the process repeats iteratively until convergency. In the initial stage, we used a warm-up strategy to attain learning stability and in the final stage we adopted a scheme of multi-step decay to improve convergency. In the first 500 iterations, the learning rate was linearly increased to 0.005. Then, the learning rate was kept flat in the next 7500 iterations. The ramp-down epochs were completed in three steps by reducing the learning rate by a factor of 10 in each step.
We also conducted ablation experiments with the proposed model to analyze the performance contributions from the different components, particularly the deformable convolution layers, GCA model and labeling of the negative samples.
2.6. Evaluation Metrics to Assess the Performance of the Models
When the IOU between the predicted and the ground truth bounding box exceeds a specified threshold value, it is considered as a correct detection. Depending on the specified confidence threshold, the detection results can have four possible types of outcomes: true positive (TP), false positive (FP), true negative (TN) and false negative (FN). Based on the detection results, the performances of the tested models are assessed using the following metrics:
Precision (
P) is the model’s ability to identify only the relevant objects and is defined as the percentage of correct positive predictions among all positive detections:
Recall (
R) is the model’s ability to detect all the relevant cases (also known as sensitivity) and defined as the percentage of the true positives among all positive ground truth cases:
Both precision and recall metrics are important for object detection, it is, therefore, necessary to determine the precision–recall curve that shows the tradeoff between the precision and recall values for different thresholds. Furthermore, with the precision-recall curve the average precision (
AP) can be estimated, which is a widely used metric to access the accuracy of an object detector.
AP computes the average precision value for recall value over 0 to 1. The general definition for
AP is to estimate the area under the precision-recall curve. That is
In practical calculation of
AP, the precision–recall curve,
P(R), is usually interpolated into multiple discrete points. For a set of queries (
n), the mean average precision (
mAP) is computed as follows:
where
k is the query index in the set and
APk is the average precision for a given query
k. In this study, we adopted the latest MS Common Objects in Context (COCO) method and a 101-point interpolated
AP definition was used in the calculation for three different IOU thresholds. Besides the metrics outlined above, other metrics, such as specificity, sensitivity and area under curve (AUC) of the receiver operating characteristics (ROC) are also commonly used along for evaluating detection models. The ROC is a graph of the relationship between the true-positive rate (sensitivity is also known as recall) and the false-positive rate (1-specificity), which measures the effects of varying decision thresholds and accounts for all possible combinations of various correct and incorrect detections. Both
mAP and AUC for the test set were computed for the following three IOU thresholds: 0.1, 0.3 and 0.5. Here the ROC calculation was based on the multiple cancer cells in the image.
3. Results
The number of parameters and frame per second (FPS) for the different models are summarized in
Table 3.
Compared with the closely related model Faster RCNN-FPN, the proposed DGCA-RCNN model does not introduce significant overhead in terms of model complexity and computation burden. The number of parameters and time efficiency as indicated by FPS are quite comparable.
The results for the test dataset indicate that the proposed DGCA-RCNN framework can improve the detection performance for cervical abnormal cells at all tested 3 IOU thresholds compared with the other state-of-the-art models based on the same backbone ResNet50. As shown in
Table 4,
Table 5 and
Table 6, the DGCA-RCNN model achieved the highest mAP and AUC at all IOU thresholds. Faster RCNN-FPN has the next best performance. Compared with the closely related model Faster RCNN-FPN, the mAP for the DGCA-RCNN model is boosted by 7–9% depending on the IOU threshold, while AUC is increased by 1–2%. As expected, the performance degrades with the increasing IOU thresholds and the corresponding mAP values are systematically reduced (see
Table 4). A similar trend has also been observed for the AUC results (
Table 5).
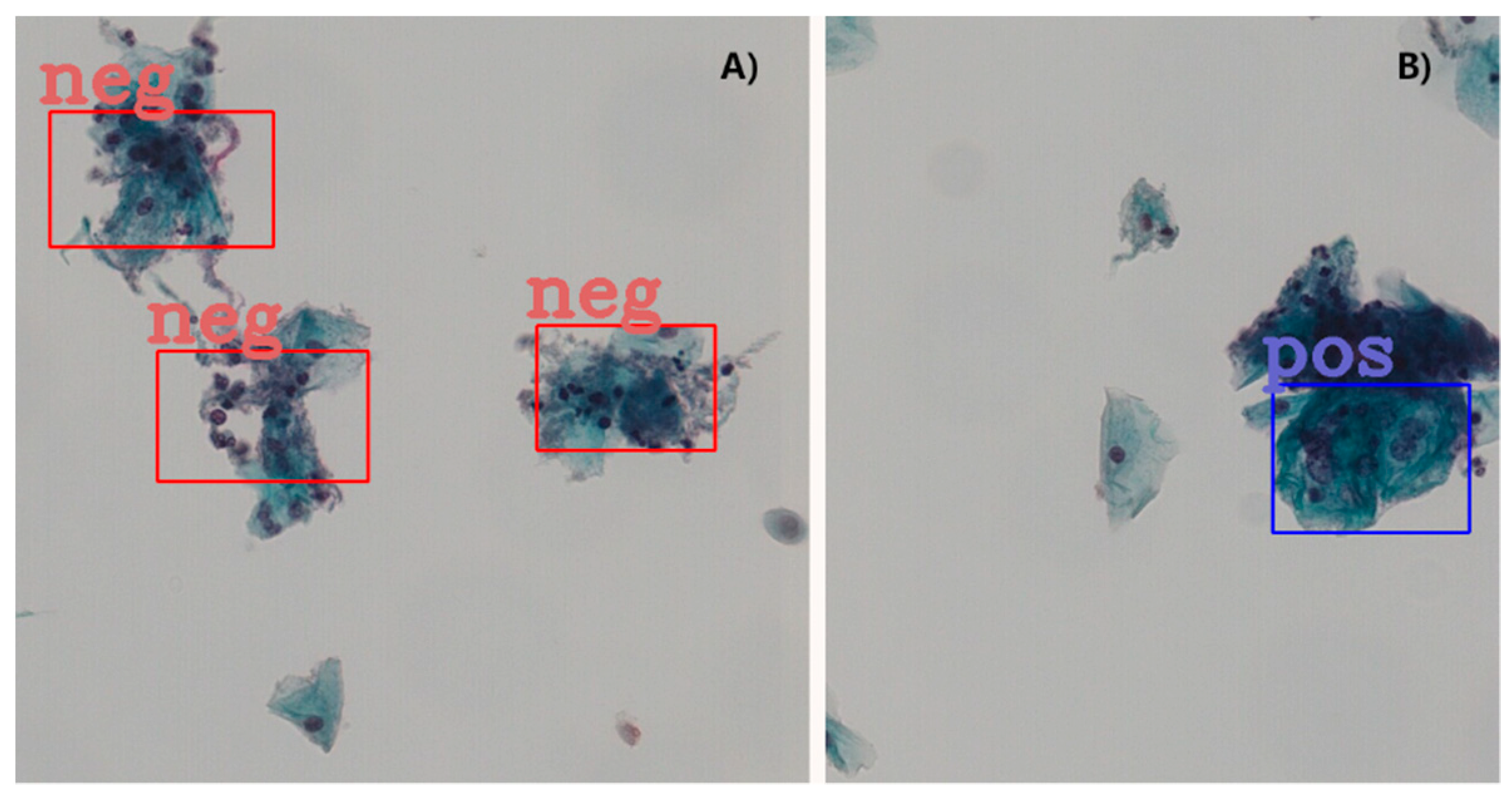
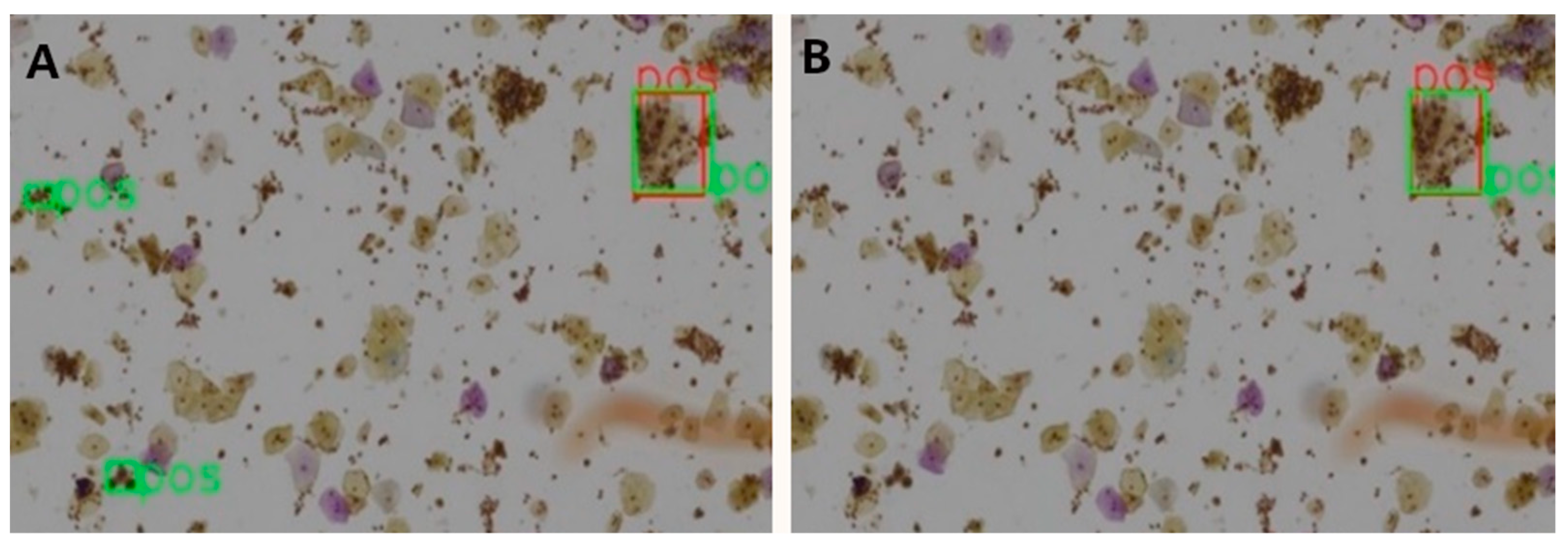
Table 7 shows the ablation results for the proposed model assessed at patch level for 3 different IOU thresholds. The mAP has been steadily boosted by adding the proposed extensions. Introducing deformable convolution layers improved mAP by 2.8–4.5%. The effect of introducing GCA module is quite inconsequential for mAP (0.1–0.6%), but nevertheless the effect is positive. The tagging of the negative image samples with computer vision techniques made significant improvement in mAP by 2.1–4.9%. To better understand this effect, we carefully compared the detection results with and without the tagging of the negative samples.
A clear observation is that the negative tagging reduces false positive detections.
Figure 5 shows such an example without (
Figure 5A) and with (
Figure 5B) the use of tagging the negative image samples. With the additional labeling information from the negative images, some subtle features of the negative cells appear to be enhanced and lead to reduced false positive detections, particularly in regions with clusters of nuclei.