Impact of Metformin on Cancer Biomarkers in Non-Diabetic Cancer Patients: A Systematic Review and Meta-Analysis of Clinical Trials
Abstract
1. Introduction
2. Methodology
2.1. Search Strategy
- 1.
- Metformin (metformin, dimethylbiguanidine, dimethylguanylguanidine, glucophage, metformin hydrochloride, and metformin HCl);
- 2.
- Cancer and metabolic biomarkers.
2.2. Inclusion Criteria and Selection Study
- (1).
- The research was based on clinical trial;
- (2).
- Non-diabetic patients with breast or endometrial cancers were selected as participants for the intervention and control groups;
- (3).
- Metformin was administrated for the intervention group;
- (4).
- The endpoint of each study was cancer biomarkers;
- (5).
- The language of articles had to be English.
2.3. Exclusion Criteria
- (1).
- Studies with unclear methodology and unclear results were not considered;
- (2).
- Duplicate studies were excluded;
- (3).
- Animals, cohort, case-report, case-control, letters, erratum, and conference papers were excluded.
2.4. Data Extraction
2.5. Quality Assessment
2.6. Statistical Analysis
3. Results
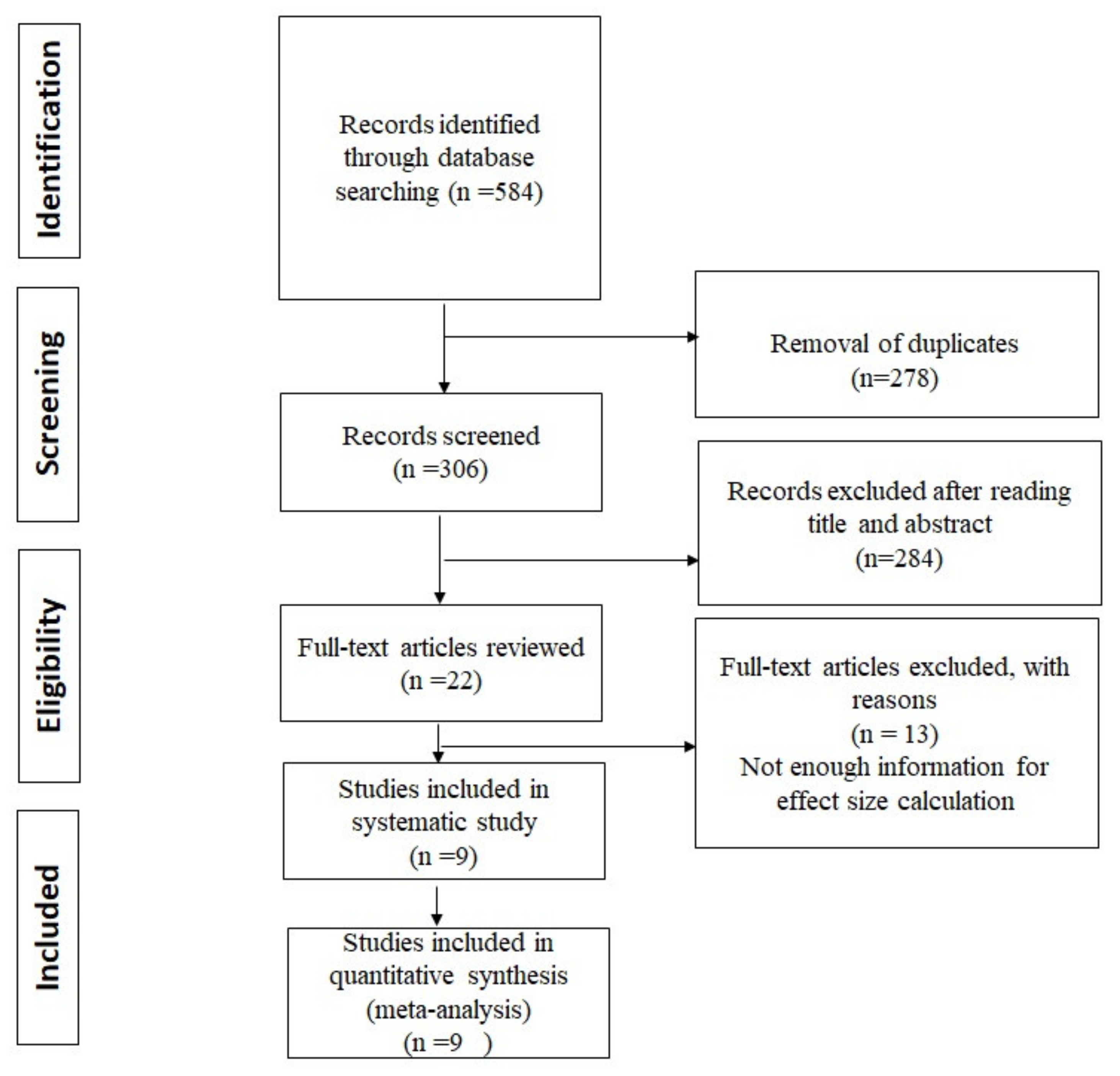
3.1. Study Selection
3.2. Features of the Study
3.3. Systematic Findings
3.4. Risk of Bias Assessment
3.5. Results of the Meta-Analysis
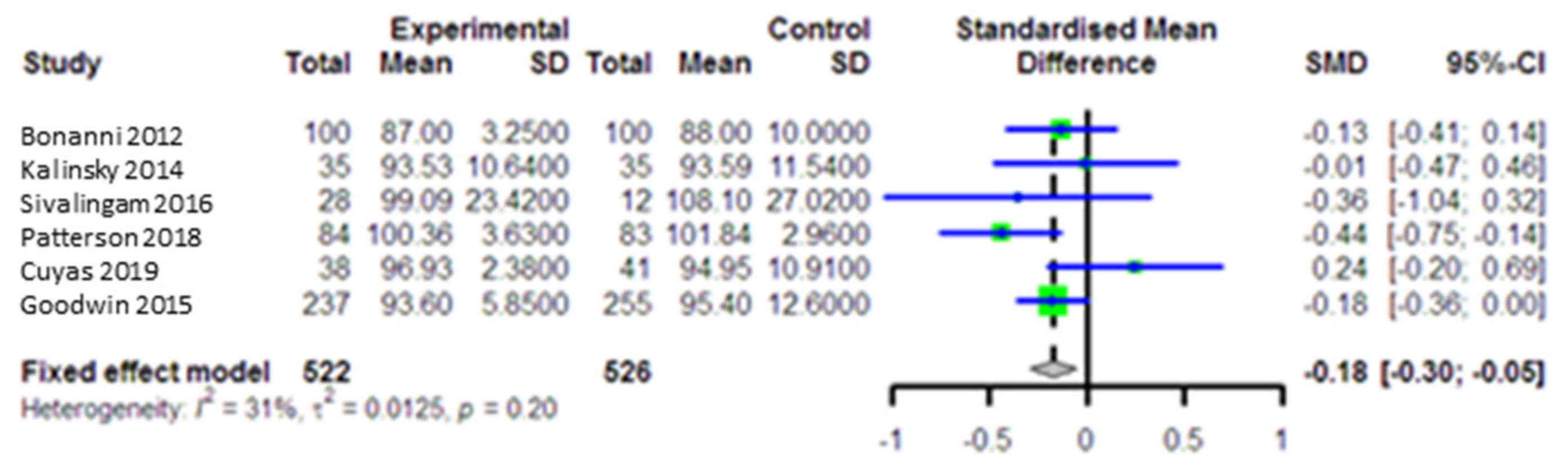
3.5.1. Impact of Metformin on Insulin Levels
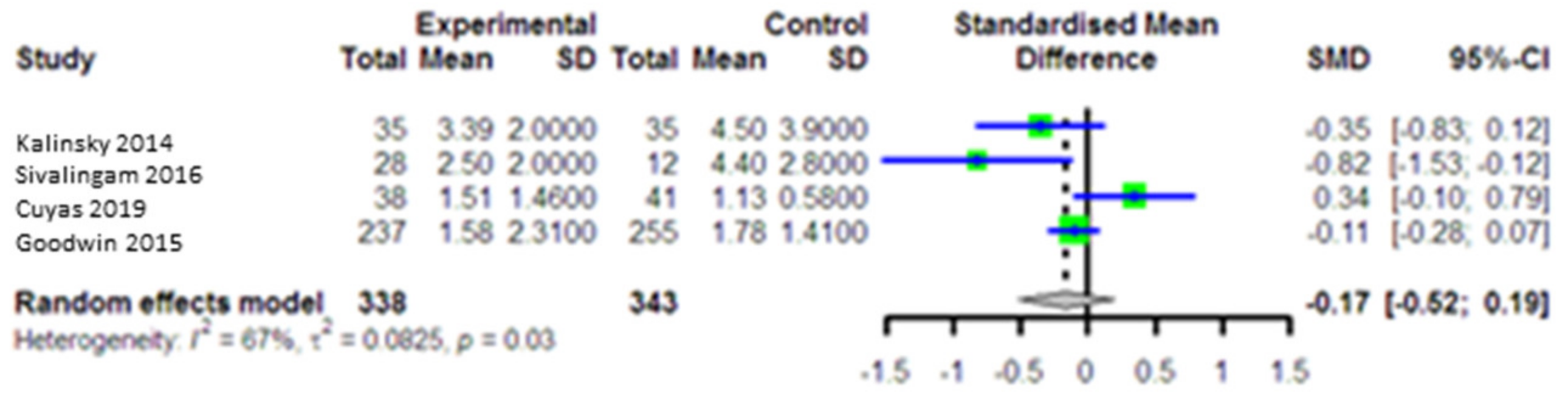
3.5.2. Impact of Metformin on Fasting Blood Glucose (FBS)
3.5.3. Impact of Metformin on HOMA-IR
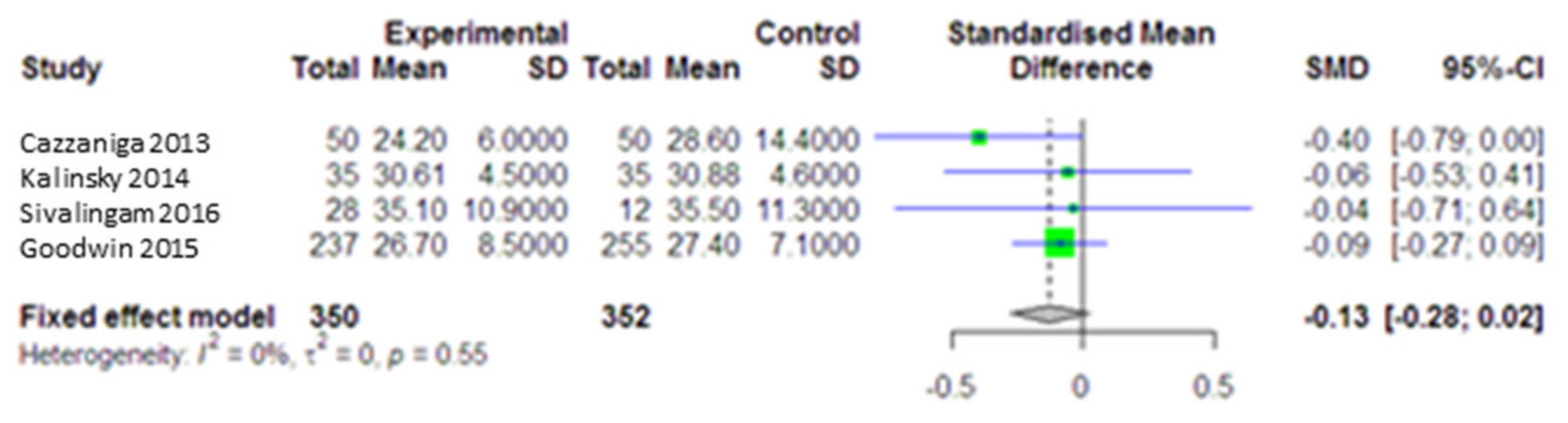
3.5.4. Impact of Metformin on Body Mass Index (BMI)
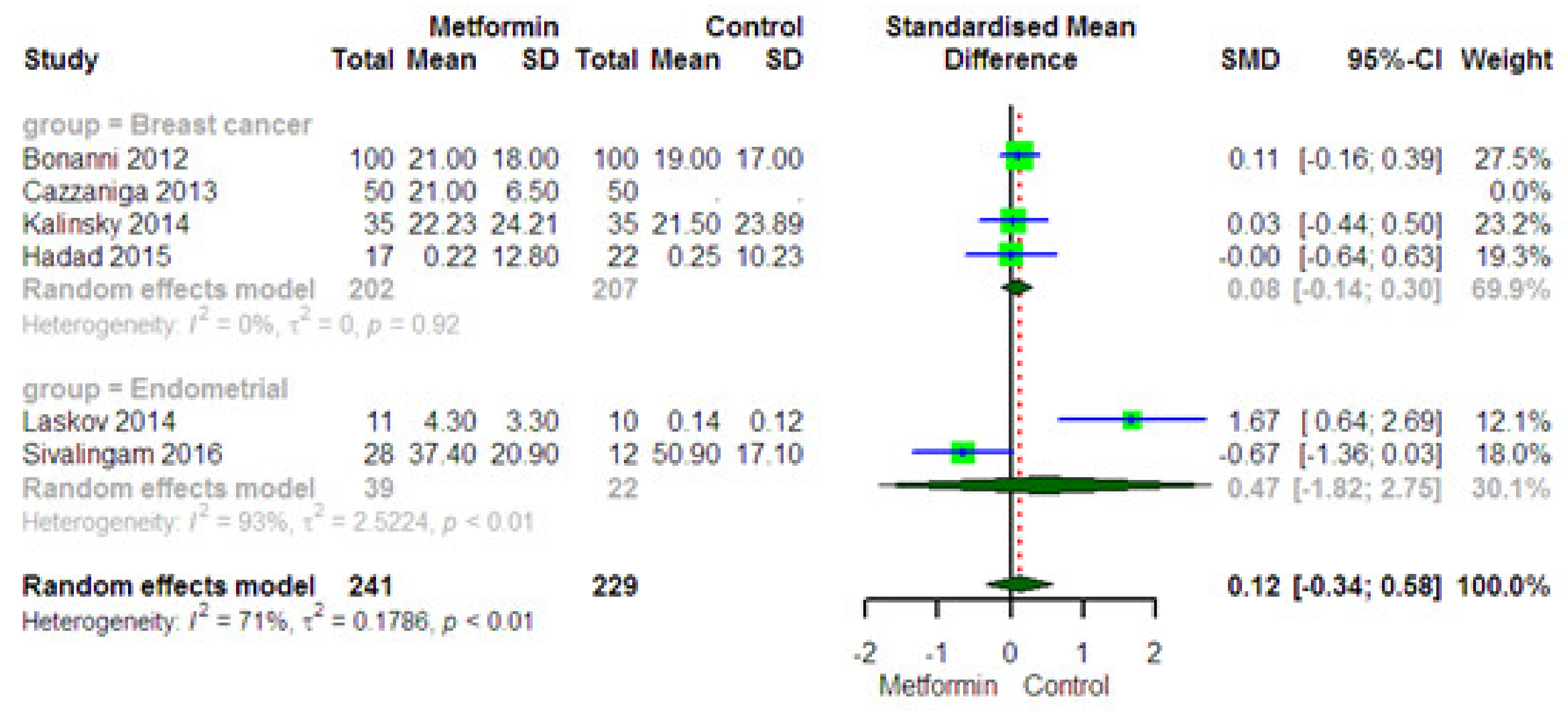
3.5.5. Impact of Metformin on the Ki-67 Level in Plasma
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed]
- Sana, M.; Malik, H.J. Current and emerging breast cancer biomarkers. J. Cancer Res. Ther. 2015, 11, 508–513. [Google Scholar] [CrossRef] [PubMed]
- Morales, D.R.; Morris, A.D. Metformin in cancer treatment and prevention. Annu. Rev. Med. 2015, 66, 17–29. [Google Scholar] [CrossRef] [PubMed]
- Vissers, P.A.; Cardwell, C.R.; van de Poll-Franse, L.V.; Young, I.S.; Pouwer, F.; Murray, L.J. The association between glucose-lowering drug use and mortality among breast cancer patients with type 2 diabetes. Breast Cancer Res. Treat. 2015, 150, 427–437. [Google Scholar] [CrossRef] [PubMed]
- Della Cort, C.M.; Ciaramella, V.; Di Maurog, C.; Castellonen, M.D.; Papaccio, F.; Fasano, M.; Sasso, F.C.; Martinelli, E.; Troiani, T.; De Vita, F.; et al. Metformin increases antitumor activity of MEK inhibitors through GLI1 downregulation in LKB1 positive human NSCLC cancer cells. Oncotarget 2016, 7, 4265–4278. [Google Scholar] [CrossRef] [PubMed]
- Morgillo, F.; Fasano, M.; Della Corte, C.M.; Sasso, F.C.; Papaccio, F.; Viscardi, G.; Esposito, G.; Di Liello, R.; Normanno, N.; Capuano, A.; et al. Results of the safety run-in part of the METAL (METformin in Advanced Lung cancer) study: A multicentre, open-label phase I–II study of metformin with erlotinib in second-line therapy of patients with stage IV non-small-cell lung cancer. ESMO Open 2017, 2, e000132. [Google Scholar] [CrossRef]
- Cantrell, L.A.; Zhou, C.; Mendivil, A.; Malloy, K.M.; Gehrig, P.A.; Bae-Jump, V.L. Metformin is a potent inhibitor of endometrial cancer cell proliferation—Implications for a novel treatment strategy. Gynecol. Oncol. 2010, 116, 92–98. [Google Scholar] [CrossRef] [PubMed]
- Sarfstein, R.; Friedman, Y.; Attias-Geva, Z.; Fishman, A.; Bruchim, I.; Werner, H. Metformin downregulates the insulin/IGF-I signaling pathway and inhibits different uterine serous carcinoma (USC) cells proliferation and migration in p53-dependent or-independent manners. PLoS ONE 2013, 8, e61537. [Google Scholar] [CrossRef]
- Zakikhani, M.; Dowling, R.; Fantus, I.G.; Sonenberg, N.; Pollak, M. Metformin is an AMP kinase-dependent growth inhibitor for breast cancer cells. Cancer Res. 2006, 66, 10269–10273. [Google Scholar] [CrossRef]
- Bonanni, B.; Puntoni, M.; Cazzaniga, M.; Pruneri, G.; Serrano, D.; Guerrieri-Gonzaga, A.; Gennari, A.; Trabacca, M.S.; Galimberti, V.; Veronesi, P.; et al. Dual effect of metformin on breast cancer proliferation in a randomized presurgical trial. J. Clin. Oncol. 2012, 30, 2593–2600. [Google Scholar] [CrossRef]
- Hadad, S.; Iwamoto, T.; Jordan, L.; Purdie, C.; Bray, S.; Baker, L.; Jellema, G.; Deharo, S.; Hardie, D.G.; Pusztai, L.; et al. Evidence for biological effects of metformin in operable breast cancer: A pre-operative, window-of-opportunity, randomized trial. Breast Cancer Res. Treat. 2011, 128, 783–794. [Google Scholar] [CrossRef] [PubMed]
- Joshua, A.; Zannella, V.; Downes, M.; Bowes, B.; Hersey, K.; Koritzinsky, M.; Schwab, M.; Hofmann, U.; Evans, A.; van der Kwast, T.; et al. A pilot ‘window of opportunity’ neoadjuvant study of metformin in localised prostate cancer. Prostate Cancer Prostatic Dis. 2014, 17, 252–258. [Google Scholar] [CrossRef] [PubMed]
- Laskov, I.; Drudi, L.; Beauchamp, M.-C.; Yasmeen, A.; Ferenczy, A.; Pollak, M.; Gotlieb, W.H. Anti-diabetic doses of metformin decrease proliferation markers in tumors of patients with endometrial cancer. Gynecol. Oncol. 2014, 134, 607–614. [Google Scholar] [CrossRef]
- Mitsuhashi, A.; Kiyokawa, T.; Sato, Y.; Shozu, M. Effects of metformin on endometrial cancer cell growth in vivo: A preoperative prospective trial. Cancer 2014, 120, 2986–2995. [Google Scholar] [CrossRef] [PubMed]
- Schuler, K.M.; Rambally, B.S.; DiFurio, M.J.; Sampey, B.P.; Gehrig, P.A.; Makowski, L.; Bae-Jump, V. Antiproliferative and metabolic effects of metformin in a preoperative window clinical trial for endometrial cancer. Cancer Med. 2015, 4, 161–173. [Google Scholar] [CrossRef]
- Miller, R.A.; Chu, Q.; Xie, J.; Foretz, M.; Viollet, B.; Birnbaum, M.J. Biguanides suppress hepatic glucagon signalling by decreasing production of cyclic AMP. Nature 2013, 494, 256–260. [Google Scholar] [CrossRef]
- Pernicova, I.; Korbonits, M. Metformin—Mode of action and clinical implications for diabetes and cancer. Nat. Rev. Endocrinol. 2014, 10, 143–156. [Google Scholar]
- Zheng, J.; Woo, S.-L.; Hu, X.; Botchlett, R.; Chen, L.; Huo, Y.; Wu, C. Metformin and metabolic diseases: A focus on hepatic aspects. Front. Med. 2015, 9, 173–186. [Google Scholar]
- Cazzaniga, M.; DeCensi, A.; Pruneri, G.; Puntoni, M.; Bottiglieri, L.; Varricchio, C.; Guerrieri-Gonzaga, A.; Gentilini, O.D.; Pagani, G.; Dell’Orto, P.; et al. The effect of metformin on apoptosis in a breast cancer presurgical trial. Br. J. Cancer 2013, 109, 2792–2797. [Google Scholar] [CrossRef]
- Kalinsky, K.; Crew, K.D.; Refice, S.; Xiao, T.; Wang, A.; Feldman, S.M.; Taback, B.; Ahmad, A.; Cremers, S.; Hibshoosh, H.; et al. Presurgical trial of metformin in overweight and obese patients with newly diagnosed breast cancer. Cancer Investig. 2014, 32, 150–157. [Google Scholar] [CrossRef]
- Patterson, R.E.; Marinac, C.R.; Sears, D.D.; Kerr, J.; Hartman, S.J.; Cadmus-Bertram, L.; Villaseñor, A.; Flatt, S.W.; Godbole, S.; Li, H.; et al. The effects of metformin and weight loss on biomarkers associated with breast cancer outcomes. JNCI J. Natl. Cancer Inst. 2018, 110, 1239–1247. [Google Scholar] [CrossRef] [PubMed]
- Hadad, S.M.; Coates, P.; Jordan, L.B.; Dowling, R.J.; Chang, M.C.; Done, S.J.; Purdie, C.A.; Goodwin, P.J.; Stambolic, V.; Moulder-Thompson, S.; et al. Evidence for biological effects of metformin in operable breast cancer: Biomarker analysis in a pre-operative window of opportunity randomized trial. Breast Cancer Res. Treat. 2015, 150, 149–155. [Google Scholar] [CrossRef] [PubMed]
- Sivalingam, V.; Kitson, S.; McVey, R.; Roberts, C.; Pemberton, P.; Gilmour, K.; Ali, S.; Renehan, A.G.; Kitchener, H.C.; Crosbie, E.J. Measuring the biological effect of presurgical metformin treatment in endometrial cancer. Br. J. Cancer 2016, 114, 281–289. [Google Scholar] [CrossRef]
- Goodwin, P.J.; Parulekar, W.R.; Gelmon, K.A.; Shepherd, L.E.; Ligibel, J.A.; Hershman, D.L.; Rastogi, P.; Mayer, I.A.; Hobday, T.J.; Lemieux, J.; et al. Effect of metformin vs placebo on weight and metabolic factors in NCIC CTG MA 32. J. Natl. Cancer Inst. 2015, 107, djv006. [Google Scholar] [CrossRef] [PubMed]
- Cuyàs, E.; Buxó, M.; Ferri Iglesias, M.J.; Verdura, S.; Pernas, S.; Dorca, J.; Álvarez, I.; Martínez, S.; Pérez-Garcia, J.M.; Batista-López, N.; et al. The C allele of ATM rs11212617 associates with higher pathological complete remission rate in breast cancer patients treated with neoadjuvant metformin. Front. Oncol. 2019, 9, 193. [Google Scholar] [CrossRef]
- Tang, G.H.; Satkunam, M.; Pond, G.R.; Steinberg, G.R.; Blandino, G.; Schünemann, H.J.; Muti, P. Association of metformin with breast cancer incidence and mortality in patients with type II diabetes: A GRADE-assessed systematic review and meta-analysis. Cancer Epidemiol. Prev. Biomark. 2018, 27, 627–635. [Google Scholar] [CrossRef]
- Rahmani, J.; Manzari, N.; Thompson, J.; Gudi, S.; Chhabra, M.; Naik, G.; Mousavi, S.M.; Varkaneh, H.K.; Clark, C.; Zhang, Y. The effect of metformin on biomarkers associated with breast cancer outcomes: A systematic review, meta-analysis, and dose-response of randomized clinical trials. Clin. Transl. Oncol. 2020, 22, 37–49. [Google Scholar] [CrossRef]
- Burcombe, R.; Wilson, G.D.; Dowsett, M.; Khan, I.; Richman, P.I.; Daley, F.; Detre, S.; Makris, A. Evaluation of Ki-67 proliferation and apoptotic index before, during and after neoadjuvant chemotherapy for primary breast cancer. Breast Cancer Res. 2006, 8, R31. [Google Scholar] [CrossRef]
- Dowsett, M.; Smith, I.E.; Ebbs, S.R.; Dixon, J.M.; Skene, A.; A’Hern, R.; Salter, J.; Detre, S.; Hills, M.; Walsh, G.; et al. Prognostic value of Ki67 expression after short-term presurgical endocrine therapy for primary breast cancer. J. Natl. Cancer Inst. 2007, 99, 167–170. [Google Scholar] [CrossRef]
- Dowsett, M.; Smith, I.E.; Ebbs, S.R.; Dixon, J.M.; Skene, A.; Griffith, C.; Boeddinghaus, I.; Salter, J.; Detre, S.; Hills, M.; et al. Short-term changes in Ki-67 during neoadjuvant treatment of primary breast cancer with anastrozole or tamoxifen alone or combined correlate with recurrence-free survival. Clin. Cancer Res. 2005, 11, 951s–958s. [Google Scholar]
- Liu, T.; Gao, H.; Yang, M.; Zhao, T.; Liu, Y.; Lou, G. Correlation of TNFAIP8 overexpression with the proliferation, metastasis, and disease-free survival in endometrial cancer. Tumor Biol. 2014, 35, 5805–5814. [Google Scholar] [CrossRef] [PubMed]
- Salvesen, H.B.; Iversen, O.E.; Akslen, L.A. Identification of high-risk patients by assessment of nuclear Ki-67 expression in a prospective study of endometrial carcinomas. Clin. Cancer Res. 1998, 4, 2779–2785. [Google Scholar] [PubMed]
- Salvesen, H.B.; Iversen, O.E.; Akslen, L.A. Prognostic significance of angiogenesis and Ki-67, p53, and p21 expression: A population-based endometrial carcinoma study. J. Clin. Oncol. 1999, 17, 1382–1390. [Google Scholar] [CrossRef] [PubMed]
- Stefansson, I.M.; Salvesen, H.B.; Akslen, L.A. Prognostic impact of alterations in P-cadherin expression and related cell adhesion markers in endometrial cancer. J. Clin. Oncol. 2004, 22, 1242–1252. [Google Scholar] [CrossRef] [PubMed]

| First Author Year Country | Study Design | Sample Size | Subjects | Age Range Duration | Intervention Dosage | Control Dosage | Result | Reference |
|---|---|---|---|---|---|---|---|---|
| Bonanni 2012 Italy | RCT | 200 | Operable Breast Cancer | 42–634 Weeks | 1700 mg/day | 1700 mg/day | Insulin (↔) FBS (↔) BMI (Reduced) Ki-67 (↔) | [10] |
| Cazzaniga 2013 Italy | RCT | 100 | Operable Breast Cancer | 45–624 weeks | 1700 mg/day | 1700 mg/day | HOMA-IR (↔) BMI (↔) Ki-67 (↔) | [19] |
| Laskov 2014 Canada | RCT | 21 | Operable Endometrial Cancer | 49–765 Weeks | 1500 mg/day | No Drug | Insulin (Reduced) BMI (↔) Ki-67 (Reduced) | [13] |
| Kalinsky 2014 USA | RCT | 70 | Operable Breast Cancer | 45.3–67.32 Weeks | 1500 mg/day | No Drug | Insulin (Reduced) FBS (↔) HOMA-IR (Reduced) BMI (Reduced) Ki-67 (↔) | [20] |
| Hadad 2015 UK | RCT | 39 | Operable Breast Cancer | - 2 Weeks | 500 mg/day, 2000 mg/day | No Drug | Insulin (↔) BMI (↔) Ki-67 (Reduced) | [22] |
| Goodwin 2015 Canada | RCT | 507 | Primary Breast Cancer | - 42 weeks | 850 mg/dsy | 850 mg/day | Insulin (Reduced) FBS (Reduced) HOMA-IR (Reduced) BMI (Reduced) Ki-67 (Reduced) | [24] |
| Sivalingam 2016 UK | RCT | 40 | Operable Endometrial Cancer | 54.7–774 Weeks | 1700 mg/day | No Drug | Insulin (↔) FBS (↔) HOMA-IR (↔) BMI (↔) Ki-67 (Reduced) | [23] |
| Patterson 2018 USA | RCT | 167 | Operable Breast Cancer | 55.7–70.524 Weeks | 500 mg/day, 1500 mg/day, 2000 mg/day | 500 mg/day, 2000 mg/day | Insulin (Reduced) FBS (↔) BMI (↔) | [21] |
| Cuyas 2019 Spain | RCT | 79 | Operable Breast Cancer | - 24 Weeks | 850 mg/day | 850 mg/day | Insulin (↔) FBS (↔) HOMA-IR (↔) | [25] |
| Study | Random Sequence | Allocation Concealment | Blinding | Incomplete Outcome Data | Selective Reporting | Other Bias | Reference |
|---|---|---|---|---|---|---|---|
| Hadad 2015 | L | L | L | H | U | L | [22] |
| Goodwin 2015 | L | L | L | U | L | U | [24] |
| Sivalingam 2016 | H | L | U | L | H | U | [23] |
| Cazzaniga 2013 | L | U | L | H | U | L | [19] |
| Cuyas 2019 | U | H | L | U | L | L | [25] |
| Bonanni 2012 | L | U | L | L | U | H | [10] |
| Laskov 2014 | L | H | H | L | U | L | [13] |
| Kalinsky 2014 | H | U | L | U | L | L | [20] |
| Patterson 2018 | U | L | H | L | L | H | [21] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Farkhondeh, T.; Amirabadizadeh, A.; Aramjoo, H.; Llorens, S.; Roshanravan, B.; Saeedi, F.; Talebi, M.; Shakibaei, M.; Samarghandian, S. Impact of Metformin on Cancer Biomarkers in Non-Diabetic Cancer Patients: A Systematic Review and Meta-Analysis of Clinical Trials. Curr. Oncol. 2021, 28, 1412-1423. https://doi.org/10.3390/curroncol28020134
Farkhondeh T, Amirabadizadeh A, Aramjoo H, Llorens S, Roshanravan B, Saeedi F, Talebi M, Shakibaei M, Samarghandian S. Impact of Metformin on Cancer Biomarkers in Non-Diabetic Cancer Patients: A Systematic Review and Meta-Analysis of Clinical Trials. Current Oncology. 2021; 28(2):1412-1423. https://doi.org/10.3390/curroncol28020134
Chicago/Turabian StyleFarkhondeh, Tahereh, Alireza Amirabadizadeh, Hamed Aramjoo, Silvia Llorens, Babak Roshanravan, Farhad Saeedi, Marjan Talebi, Mehdi Shakibaei, and Saeed Samarghandian. 2021. "Impact of Metformin on Cancer Biomarkers in Non-Diabetic Cancer Patients: A Systematic Review and Meta-Analysis of Clinical Trials" Current Oncology 28, no. 2: 1412-1423. https://doi.org/10.3390/curroncol28020134
APA StyleFarkhondeh, T., Amirabadizadeh, A., Aramjoo, H., Llorens, S., Roshanravan, B., Saeedi, F., Talebi, M., Shakibaei, M., & Samarghandian, S. (2021). Impact of Metformin on Cancer Biomarkers in Non-Diabetic Cancer Patients: A Systematic Review and Meta-Analysis of Clinical Trials. Current Oncology, 28(2), 1412-1423. https://doi.org/10.3390/curroncol28020134







