Survival and Virulence Potential of Drug-Resistant E. coli in Simulated Gut Conditions and Antibiotic Challenge
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Strains and Growth Conditions
2.2. Screening of Virulence Genes
2.3. Extraction and Profiling of Plasmid DNA
2.4. Conjugation Experiment
2.5. Growth Kinetics of MDR and Sensitive E. coli under In-Vitro Gut Conditions
2.6. Determination of Minimum Inhibitory Concentration (MIC) on Exposure to In-Vitro Gut Conditions
2.7. Effect of Simulated Gut Conditions and Ciprofloxacin Shock on the Expression of Different Virulence Genes
2.8. Bacterial Adhesion and Invasion Assay
2.9. Transcriptome Analysis of MDR Strain Grown in 3% Bile
2.9.1. Bile Exposure
2.9.2. Total RNA Isolation, cDNA Library Construction and Analysis
2.9.3. Gene Ontology and Cluster Analysis
2.9.4. Transcriptome Validation by qPCR
2.9.5. Validation of Transcriptome Data by Motility Test
2.9.6. Statistical Analysis
3. Results
3.1. Screening of Virulence Genes–Occurrence of traT Gene in Majority of the Isolates
3.2. Presence of Multiple Plasmids in MDR Strains
3.3. Trans-Conjugation Ablity of Quinolone Resitant Isolates
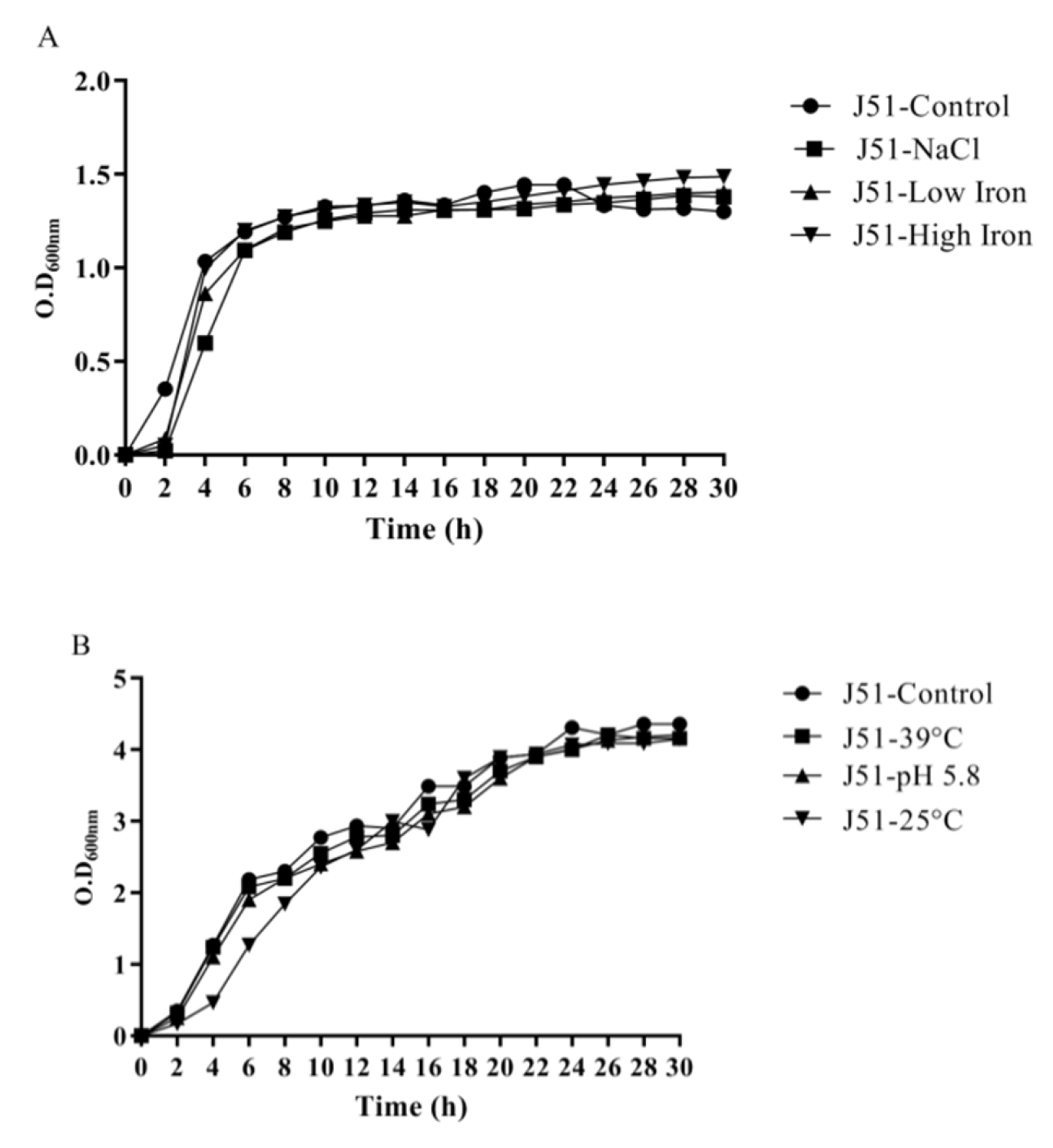
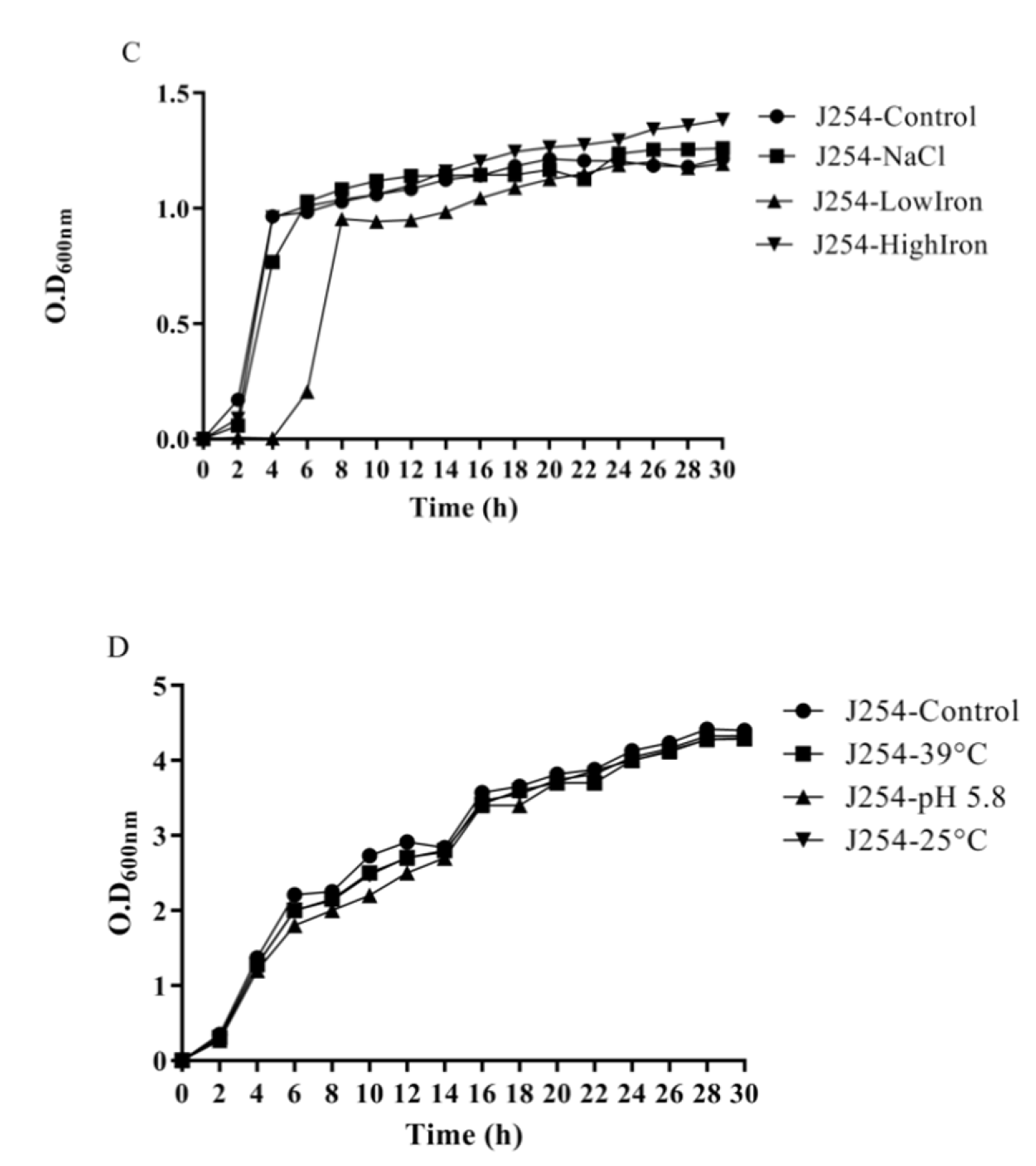
3.4. Reduction in Growth Kinetics of MDR and Sensitive E. coli upon Bile Challenge
3.5. Variation in Minimum Inhibitory Concentration (MIC) on Exposure to In-Vitro Gut Conditions
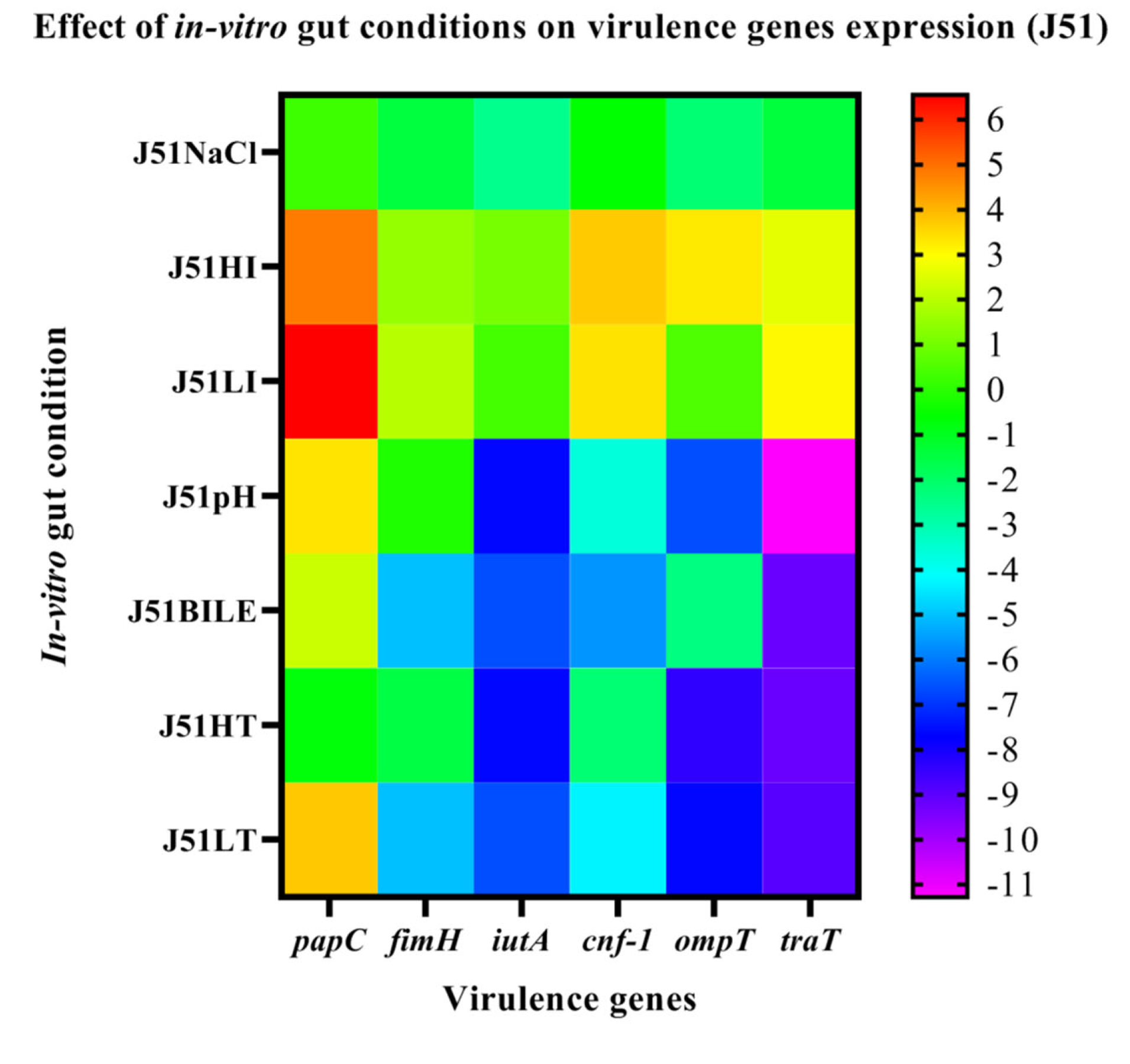
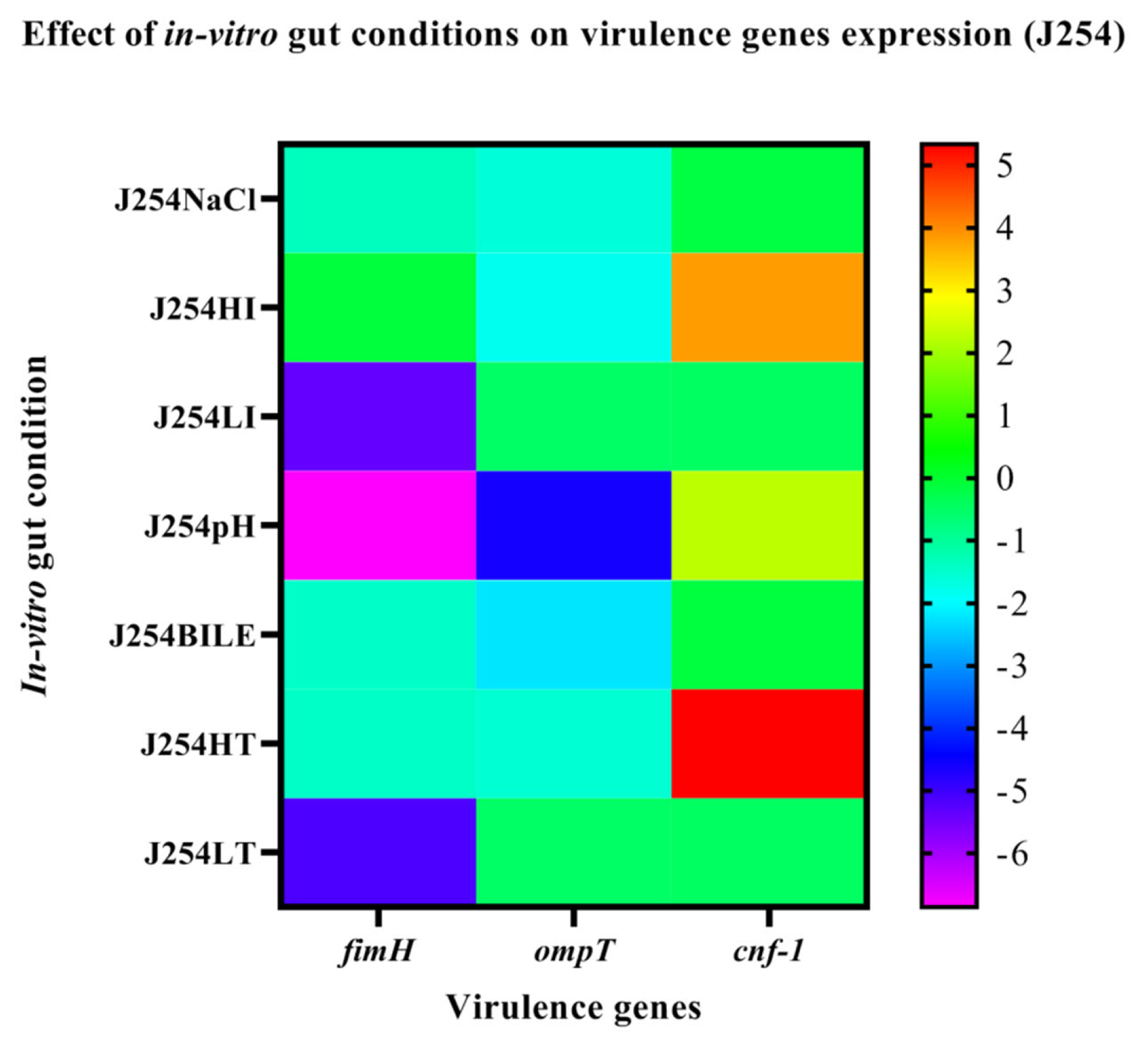
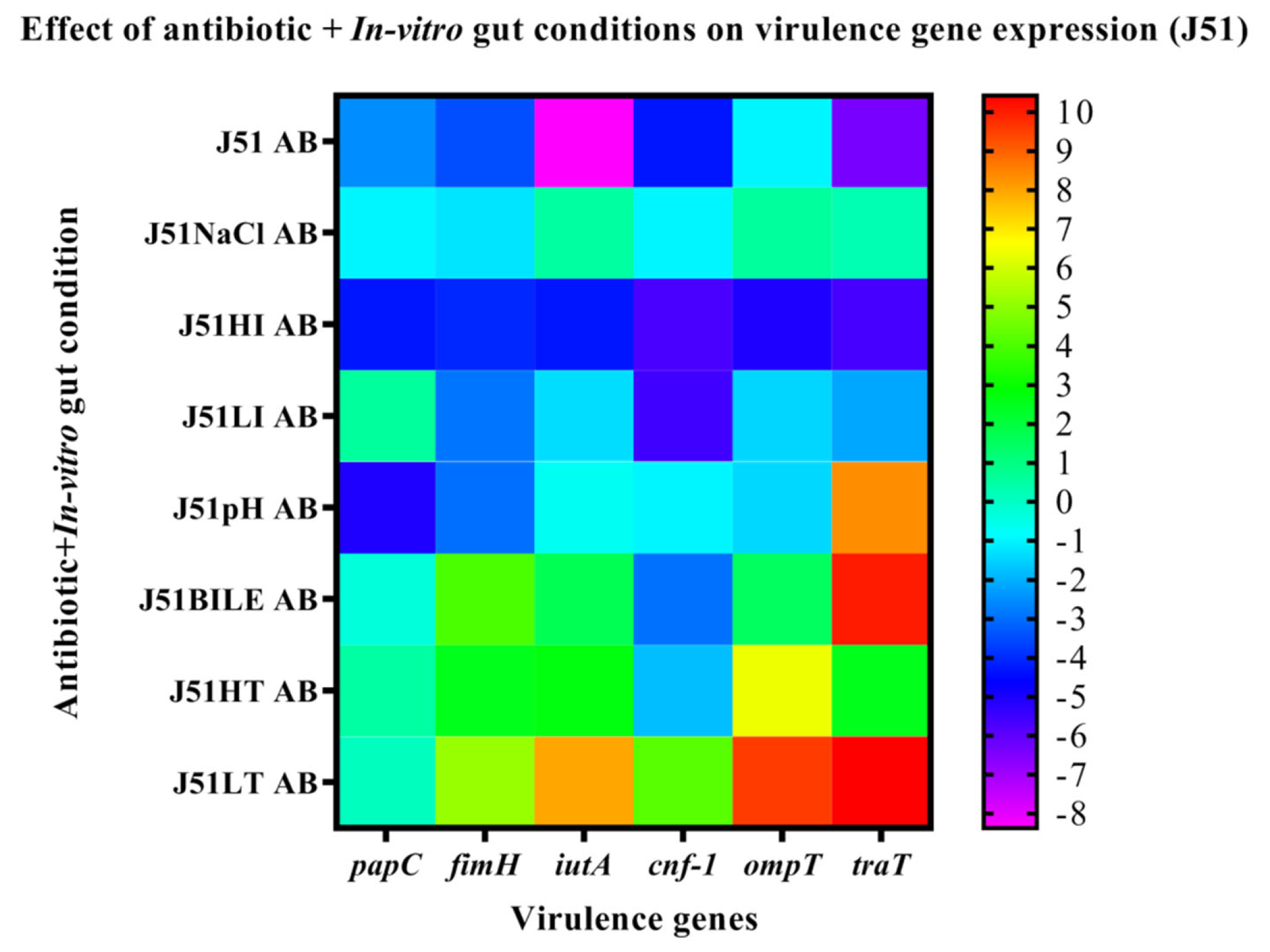
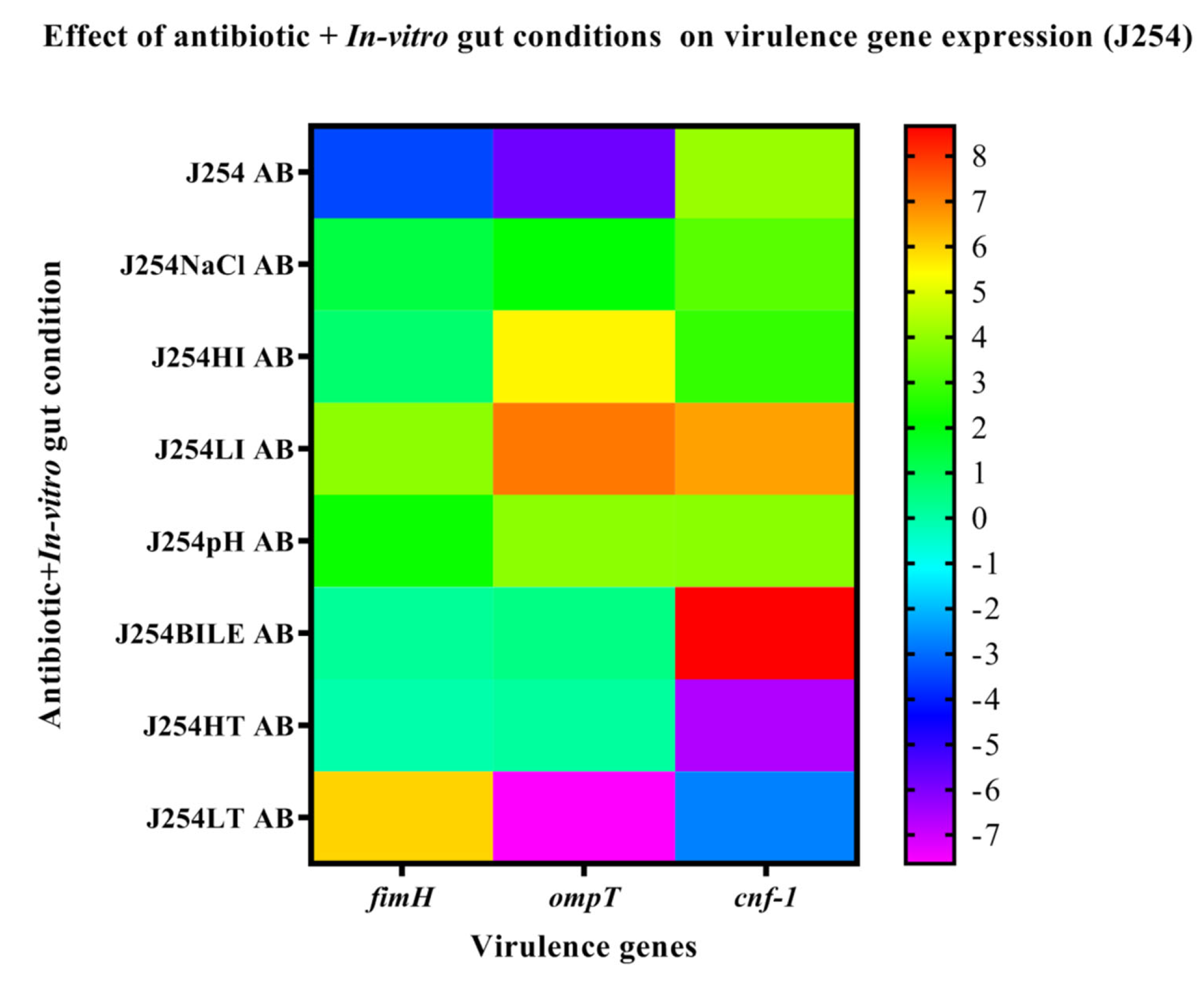
3.6. Effect of In-Vitro Gut Conditions and Ciprofloxacin Shock on the Expression of Virulence Genes
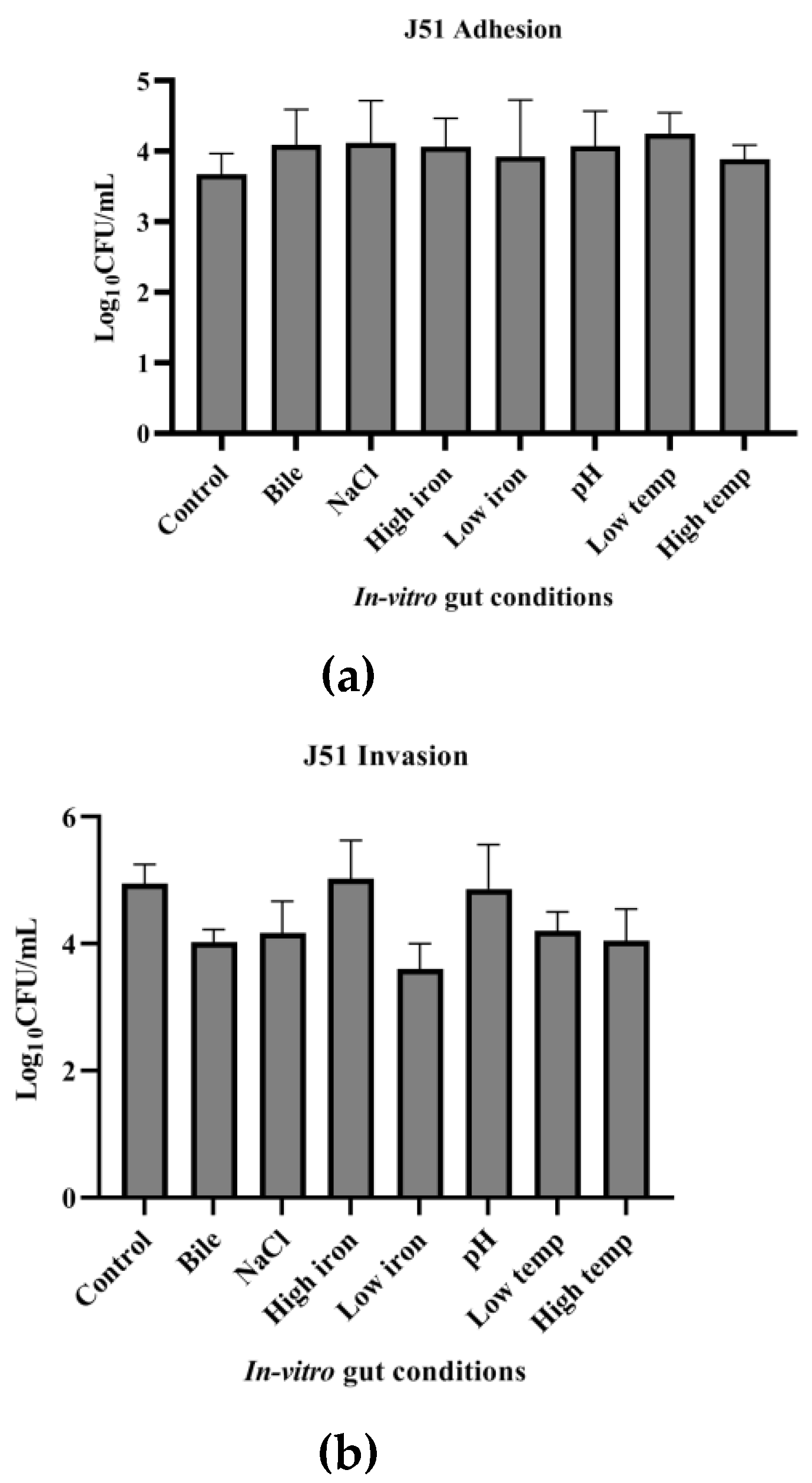
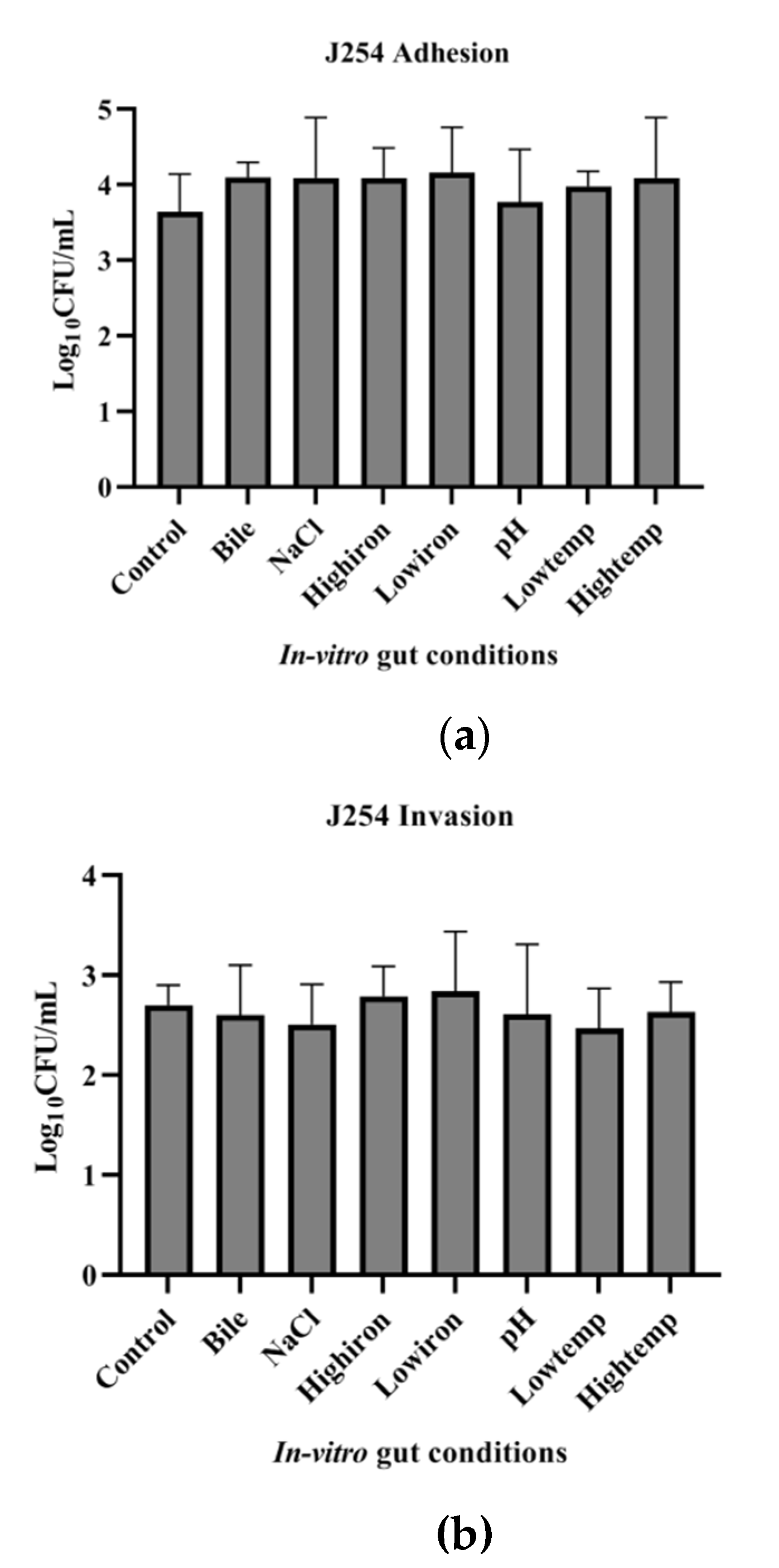
3.7. Adhesion and Invasion Potential of MDR and Sensitive Isolate
3.8. Functional Classification of Differentially Expressed Genes (DEGs) in MDR Isolate on Exposure to Bile and Combination of Bile and Ciprofloaxcin
3.8.1. Identification of Significantly Expressed Genes Using Hierarchical Clustering
3.8.2. Downregulation of Various Metabolic Pathways on Exposure to Bile and Combination of Bile and Ciprofloxacin
3.9. Transcriptome Validation of Differentially Expressed Genes by qPCR
Decreased Motility on Exposure to Bile and Combination of Bile and Ciprofloxacin
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- De Kraker, M.E.A.; Stewardson, A.J.; Harbarth, S. Will 10 Million People Die a Year due to Antimicrobial Resistance by 2050? PLoS Med. 2016, 13, e1002184. [Google Scholar] [CrossRef] [PubMed]
- Murray, C.J.; Ikuta, K.S.; Sharara, F.; Swetschinski, L.; Aguilar, G.R.; Gray, A.; Han, C.; Bisignano, C.; Rao, P.; Wool, E.; et al. Global burden of bacterial antimicrobial resistance in 2019: A systematic analysis. Lancet 2022, 399, 629–655. [Google Scholar] [CrossRef]
- Dong, G.; Li, J.; Chen, L.; Bi, W.; Zhang, X.; Liu, H.; Zhi, X.; Zhou, T.; Cao, J. Effects of sub-minimum inhibitory concentrations of ciprofloxacin on biofilm formation and virulence factors of Escherichia coli. Braz. J. Infect. Dis. 2019, 23, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Spector, M.P.; Kenyon, W.J. Resistance and survival strategies of Salmonella enterica to environmental stresses. Food Res. Int. 2012, 45, 455–481. [Google Scholar] [CrossRef]
- Gonzales-Siles, L.; Sjöling, A. The different ecological niches of enterotoxigenic Escherichia coli. Environ. Microbiol. 2016, 18, 741–751. [Google Scholar] [CrossRef] [PubMed]
- Yoon, H.; Park, B.-Y.; Oh, M.-H.; Choi, K.-H.; Yoon, Y. Effect of NaCl on Heat Resistance, Antibiotic Susceptibility, and Caco-2 Cell Invasion of Salmonella. BioMed Res. Int. 2013, 2013, 274096. [Google Scholar] [CrossRef]
- Dichtl, S.; Demetz, E.; Haschka, D.; Tymoszuk, P.; Petzer, V.; Nairz, M.; Seifert, M.; Hoffmann, A.; Brigo, N.; Würzner, R.; et al. Dopamine Is a Siderophore-Like Iron Chelator That Promotes Salmonella enterica Serovar Typhimurium Virulence in Mice. mBio 2019, 10, e02624-18. [Google Scholar] [CrossRef]
- Gunasekera, T.S.; Csonka, L.N.; Paliy, O. Genome-Wide Transcriptional Responses of Escherichia coli K-12 to Continuous Osmotic and Heat Stresses. J. Bacteriol. 2008, 190, 3712–3720. [Google Scholar] [CrossRef]
- Aditya, V.; Kotian, A.; Saikrishnan, S.; Rohit, A.; Mithoor, D.; Karunasagar, I.; Deekshit, V.K. Effect of ciprofloxacin and in vitro gut conditions on biofilm of Escherichia coli isolated from clinical and environmental sources. J. Appl. Microbiol. 2022, 132, 964–977. [Google Scholar] [CrossRef]
- Poirel, L.; Madec, J.-Y.; Lupo, A.; Schink, A.-K.; Kieffer, N.; Nordmann, P.; Schwarz, S. Antimicrobial Resistance in Escherichia coli. Microbiol. Spectr. 2018, 6. [Google Scholar] [CrossRef]
- Hamner, S.; McInnerney, K.; Williamson, K.; Franklin, M.J.; Ford, T. Bile Salts Affect Expression of Escherichia coli O157:H7 Genes for Virulence and Iron Acquisition, and Promote Growth under Iron Limiting Conditions. PLoS ONE 2013, 8, e74647. [Google Scholar] [CrossRef] [PubMed]
- Valat, C.; Hirchaud, E.; Drapeau, A.; Touzain, F.; Boisseson, C.; Haenni, M.; Blanchard, Y.; Madec, J.; de Boisseson, C. Overall changes in the transcriptome of Escherichia coli O26:H11 induced by a subinhibitory concentration of ciprofloxacin. J. Appl. Microbiol. 2020, 129, 1577–1588. [Google Scholar] [CrossRef] [PubMed]
- Akshay, S.D.; Anupama, K.P.; Deekshit, V.K.; Rohit, A.; Maiti, B. Effect of sub-minimum inhibitory concentration of ceftriaxone on the expression of outer membrane proteins in Salmonella enterica serovar Typhi. World J. Microbiol. Biotechnol. 2022, 38, 190. [Google Scholar] [CrossRef] [PubMed]
- Scheiring, J.; Andreoli, S.P.; Zimmerhackl, L.B. Treatment and outcome of Shiga-toxin-associated hemolytic uremic syndrome (HUS). Pediatr. Nephrol. 2008, 23, 1749–1760. [Google Scholar] [CrossRef] [PubMed]
- Zimmerhackl, L.B. E. coli, Antibiotics, and the Hemolytic–Uremic Syndrome. N. Engl. J. Med. 2000, 342, 1990–1991. [Google Scholar] [CrossRef]
- Safdar, N.; Said, A.; Gangnon, R.E.; Maki, D.G. Risk of Hemolytic Uremic Syndrome After Antibiotic Treatment of Escherichia coli O157:H7 Enteritis. JAMA 2002, 288, 996–1001. [Google Scholar] [CrossRef]
- Sambrook, J.; Fritsch, E.F.; Maniatis, T. Molecular Cloning: A Laboratory Manual, 2nd ed.; Lanssen, K.A., Ed.; Cold Spring Harbor Laboratory Press: Long Island, NY, USA, 1989. [Google Scholar]
- Shrestha, U.T.; Shrestha, S.; Adhikari, N.; Rijal, K.R.; Shrestha, B.; Adhikari, B.; Banjara, M.R.; Ghimire, P. Plasmid Profiling and Occurrence of β-Lactamase Enzymes in Multidrug-Resistant Uropathogenic Escherichia coli in Kathmandu, Nepal. Infect. Drug Resist. 2020, 13, 1905–1917. [Google Scholar] [CrossRef]
- Wang, A.; Yang, Y.; Lu, Q.; Wang, Y.; Chen, Y.; Deng, L.; Ding, H.; Deng, Q.; Zhang, H.; Wang, C.; et al. Presence of qnr gene in Escherichia coli and Klebsiella pneumoniae resistant to ciprofloxacin isolated from pediatric patients in China. BMC Infect. Dis. 2008, 8, 68. [Google Scholar] [CrossRef]
- Poole, N.M.; Green, S.I.; Rajan, A.; Vela, L.E.; Zeng, X.-L.; Estes, M.K.; Maresso, A.W. Role for FimH in Extraintestinal Pathogenic Escherichia coli Invasion and Translocation through the Intestinal Epithelium. Infect. Immun. 2017, 85, 11. [Google Scholar] [CrossRef]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 2009, 4, 44–57. [Google Scholar] [CrossRef]
- Kanehisa, M.; Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef] [PubMed]
- Rezatofighi, S.E.; Mirzarazi, M.; Salehi, M. Virulence genes and phylogenetic groups of uropathogenic Escherichia coli isolates from patients with urinary tract infection and uninfected control subjects: A case-control study. BMC Infect. Dis. 2021, 21, 361. [Google Scholar] [CrossRef] [PubMed]
- Uma, B.; Prabhakar, K.; Rajendran, S.; Kavitha, K.; Sarayu, Y.L. Antibiotic sensitivity and plasmid profiles ofEscherichiacoliisolated from pediatric diarrhea. J. Glob. Infect. Dis. 2009, 1, 107–110. [Google Scholar] [CrossRef] [PubMed]
- Kotian, A.; Aditya, V.; Jazeela, K.; Karunasagar, I.; Karunasagar, I.; Deekshit, V.K. Effect of bile on growth and biofilm formation of non-typhoidal salmonella serovars isolated from seafood and poultry. Res. Microbiol. 2020, 171, 165–173. [Google Scholar] [CrossRef]
- Prouty, A.; Brodsky, I.; Manos, J.; Belas, R.; Falkow, S.; Gunn, J. Transcriptional regulation of Salmonella enterica serovar Typhimurium genes by bile. FEMS Immunol. Med. Microbiol. 2004, 41, 177–185. [Google Scholar] [CrossRef][Green Version]
- Papezova, K.; Gregorova, D.; Jonuschies, J.; Rychlik, I. Ordered expression of virulence genes inSalmonella enterica serovar typhimurium. Folia Microbiol. 2007, 52, 107–114. [Google Scholar] [CrossRef]
- Chen, T.; Dong, G.; Zhang, S.; Zhang, X.; Zhao, Y.; Cao, J.; Zhou, T.; Wu, Q. Effects of iron on the growth, biofilm formation and virulence of Klebsiella pneumoniae causing liver abscess. BMC Microbiol. 2020, 20, 36. [Google Scholar] [CrossRef]
- Cassat, J.E.; Skaar, E.P. Iron in Infection and Immunity. Cell Host Microbe 2013, 13, 509–519. [Google Scholar] [CrossRef]
- Fekri, K.; Khoshdel, A.; Rasoulynezhad, M.; Kheiri, S.; Malekpour, A.; Zamanzad, B. In vitro effect of iron chelators on the growth of Escherichia coli, Staphylococcus epidermidis, Staphylococcus aureus, Yersinia enterocolitica, and Pseudomonas aeruginosa Strains. J. Shahrekord Univ. Med. Sci. 2019, 21, 244–249. [Google Scholar] [CrossRef]
- Li, F.; Xiong, X.-S.; Yang, Y.-Y.; Wang, J.-J.; Wang, M.-M.; Tang, J.-W.; Liu, Q.-H.; Wang, L.; Gu, B. Effects of NaCl Concentrations on Growth Patterns, Phenotypes Associated with Virulence, and Energy Metabolism in Escherichia coli BW25113. Front. Microbiol. 2021, 12, 705326. [Google Scholar] [CrossRef]
- Withman, B.; Gunasekera, T.S.; Beesetty, P.; Agans, R.; Paliy, O. Transcriptional Responses of Uropathogenic Escherichia coli to Increased Environmental Osmolality Caused by Salt or Urea. Infect. Immun. 2013, 81, 80–89. [Google Scholar] [CrossRef] [PubMed]
- Keerthirathne, T.P.; Ross, K.; Fallowfield, H.; Whiley, H. A Review of Temperature, pH, and Other Factors that Influence the Survival of Salmonella in Mayonnaise and Other Raw Egg Products. Pathogens 2016, 5, 63. [Google Scholar] [CrossRef]
- Noor, R.; Islam, Z.; Munshi, S.K.; Rahman, F. Influence of temperature on Escherichia coli growth in different culture media. J. Pure Appl. Microbiol. 2013, 7, 899–904. [Google Scholar]
- Derakhshan, S.; Ahmadi, S.; Ahmadi, E.; Nasseri, S.; Aghaei, A. Characterization of Escherichia coli isolated from urinary tract infection and association between virulence expression and antimicrobial susceptibility. BMC Microbiol. 2022, 22, 89. [Google Scholar] [CrossRef] [PubMed]
- Pope, L.M.; Reed, K.E.; Payne, S.M. Increased protein secretion and adherence to HeLa cells by Shigella spp. following growth in the presence of bile salts. Infect. Immun. 1995, 63, 3642–3648. [Google Scholar] [CrossRef]
- Kortman, G.A.M.; Boleij, A.; Swinkels, D.W.; Tjalsma, H. Iron Availability Increases the Pathogenic Potential of Salmonella Typhimurium and Other Enteric Pathogens at the Intestinal Epithelial Interface. PLoS ONE 2012, 7, e29968. [Google Scholar] [CrossRef] [PubMed]
- Deekshit, V.; Kumar, B.; Rai, P.; Karunasagar, I. Differential expression of virulence genes and role of gyrA mutations in quinolone resistant and susceptible strains of Salmonella Weltevreden and Newport isolated from seafood. J. Appl. Microbiol. 2015, 119, 970–980. [Google Scholar] [CrossRef] [PubMed]
- Prieto, A.I.; Ramos-Morales, F.; Casadesús, J. Bile-Induced DNA Damage in Salmonella enterica. Genetics 2004, 168, 1787–1794. [Google Scholar] [CrossRef]
- Kwan, B.W.; Lord, D.M.; Peti, W.; Page, R.; Benedik, M.J.; Wood, T.K. The MqsR/MqsA toxin/antitoxin system protects Escherichia coli during bile acid stress. Environ. Microbiol. 2015, 17, 3168–3181. [Google Scholar] [CrossRef]
- Erental, A.; Sharon, I.; Engelberg-Kulka, H. Two Programmed Cell Death Systems in Escherichia coli: An Apoptotic-Like Death Is Inhibited by the mazEF-Mediated Death Pathway. PLOS Biol. 2012, 10, e1001281. [Google Scholar] [CrossRef]
- Sun, R.; Zhao, X.; Meng, Q.; Huang, P.; Zhao, Q.; Liu, X.; Zhang, W.; Zhang, F.; Fu, Y. Genome-Wide Screening and Characterization of Genes Involved in Response to High Dose of Ciprofloxacin in Escherichia coli. Microb. Drug Resist. 2022, 28, 501–510. [Google Scholar] [CrossRef] [PubMed]
- Shu, Q.; Crick, S.L.; Pinkner, J.S.; Ford, B.; Hultgren, S.J.; Frieden, C. The E. coli CsgB nucleator of curli assembles to β-sheet oligomers that alter the CsgA fibrillization mechanism. Proc. Natl. Acad. Sci. USA 2012, 109, 6502–6507. [Google Scholar] [CrossRef] [PubMed]
- Dörr, T.; Vulić, M.; Lewis, K. Ciprofloxacin Causes Persister Formation by Inducing the TisB toxin in Escherichia coli. PLOS Biol. 2010, 8, e1000317. [Google Scholar] [CrossRef] [PubMed]
- Gurnev, P.A.; Ortenberg, R.; Dörr, T.; Lewis, K.; Bezrukov, S.M. Persister-promoting bacterial toxin TisB produces anion-selective pores in planar lipid bilayers. FEBS Lett. 2012, 586, 2529–2534. [Google Scholar] [CrossRef] [PubMed]
- Cameron, D.; Shan, Y.; Zalis, E.A.; Isabella, V.; Lewis, K. A Genetic Determinant of Persister Cell Formation in Bacterial Pathogens. J. Bacteriol. 2018, 200, 17. [Google Scholar] [CrossRef]



| Functional Annotation Cluster | Enrichment Score | Gene Count |
|---|---|---|
| C vs. B | ||
| Annotation Cluster 1 | 3.27 | |
| Transposition | 10 | |
| DNA Recombination | 16 | |
| Annotation Cluster 2 | 2.71 | |
| Bacterial type flagellum-dependent cell motility | 11 | |
| Bacterial type flagellum dependent swarming facility | 8 | |
| Annotation Cluster 3 | 2.31 | |
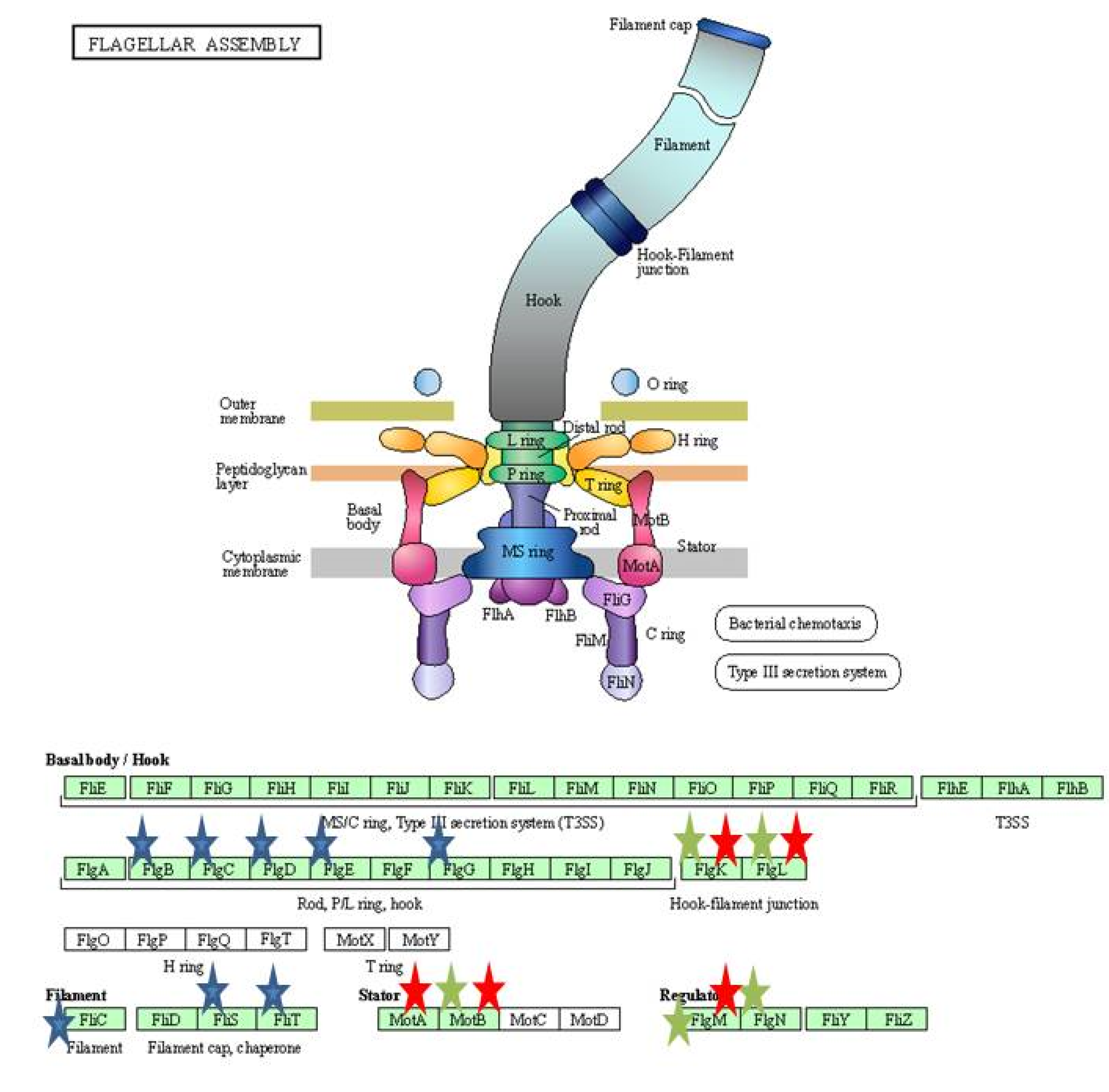
| Flagellar assembly | 14 | |
| C vs. BAB | ||
| Annotation Cluster 1 | 3.9 | |
| Biosynthesis of secondary metabolites | 62 | |
| Carbon metabolism | 31 | |
| Pyruvate metabolism | 19 | |
| Citric acid cycle | 12 | |
| Annotation Cluster 2 | 3.55 | |
| DNA recombination | 19 | |
| Transposition | 15 | |
| Annotation Cluster 3 | 2.82 | |
| Glycolysis/Gluconeogenesis | 15 | |
| Annotation Cluster 4 | 2.6 | |
| NADP binding | 20 | |
| B vs. BAB | ||
| Annotation Cluster 1 | 2.8 | |
| Metabolic pathways | 12 | |
| Annotation Cluster 2 | 2.17 | |
| ATP binding | 10 | |
| Response to heat | 6 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Aditya, V.; Kotian, A.; Sanil, A.; Thaseena, P.M.-a.; Karunasagar, I.; Deekshit, V.K. Survival and Virulence Potential of Drug-Resistant E. coli in Simulated Gut Conditions and Antibiotic Challenge. Int. J. Environ. Res. Public Health 2022, 19, 12805. https://doi.org/10.3390/ijerph191912805
Aditya V, Kotian A, Sanil A, Thaseena PM-a, Karunasagar I, Deekshit VK. Survival and Virulence Potential of Drug-Resistant E. coli in Simulated Gut Conditions and Antibiotic Challenge. International Journal of Environmental Research and Public Health. 2022; 19(19):12805. https://doi.org/10.3390/ijerph191912805
Chicago/Turabian StyleAditya, Vankadari, Akshatha Kotian, Anisha Sanil, Poidal Mohammed-ali Thaseena, Indrani Karunasagar, and Vijaya Kumar Deekshit. 2022. "Survival and Virulence Potential of Drug-Resistant E. coli in Simulated Gut Conditions and Antibiotic Challenge" International Journal of Environmental Research and Public Health 19, no. 19: 12805. https://doi.org/10.3390/ijerph191912805
APA StyleAditya, V., Kotian, A., Sanil, A., Thaseena, P. M.-a., Karunasagar, I., & Deekshit, V. K. (2022). Survival and Virulence Potential of Drug-Resistant E. coli in Simulated Gut Conditions and Antibiotic Challenge. International Journal of Environmental Research and Public Health, 19(19), 12805. https://doi.org/10.3390/ijerph191912805







