Wastewater Sequencing—An Innovative Method for Variant Monitoring of SARS-CoV-2 in Populations
Abstract
:1. Introduction
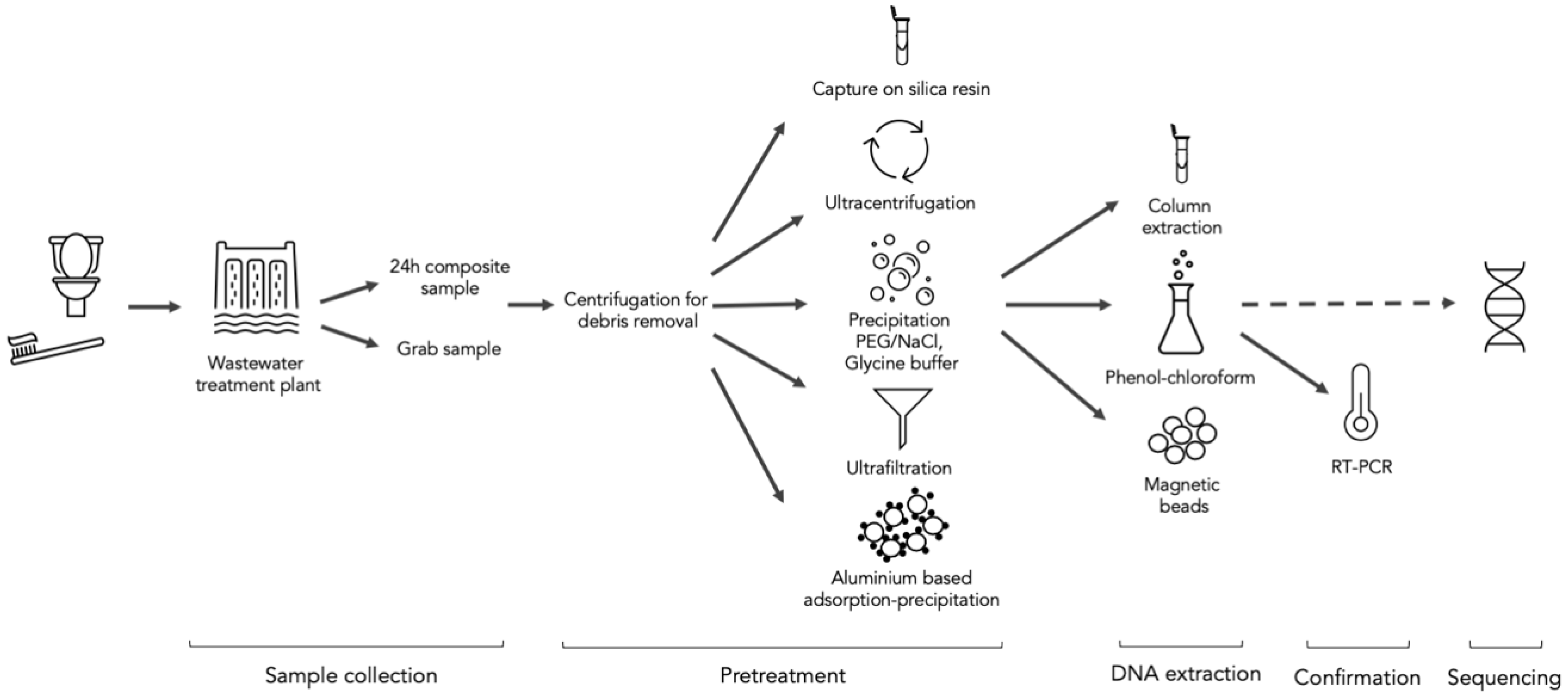
2. Methodology
2.1. Samples and Sampling Methods
2.2. Pretreatment of Wastewater Samples
2.3. Extraction of RNA
2.4. Confirmation by RT-qPCR
2.5. Sequencing Technology
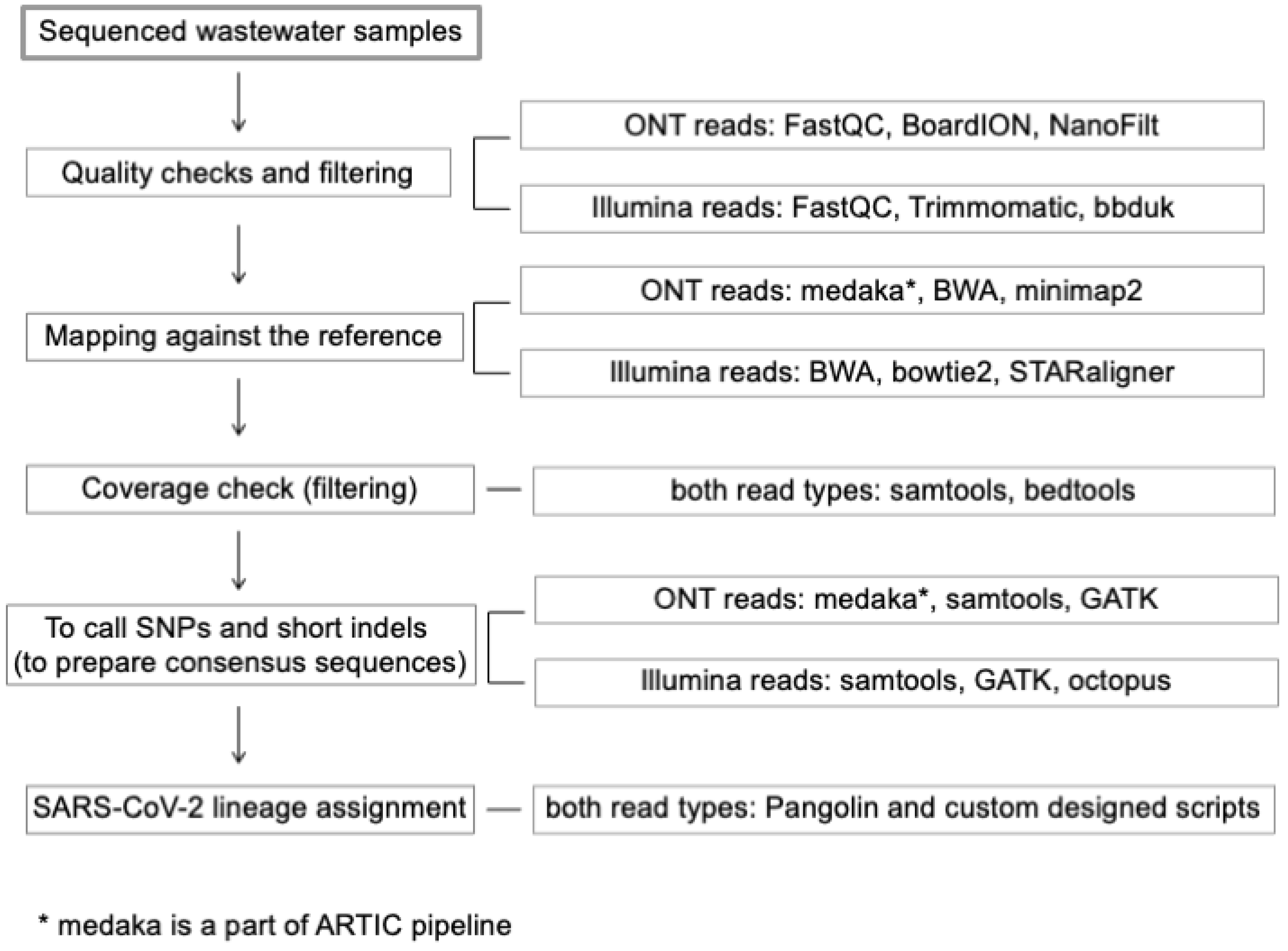
2.6. Data Analysis and Variant Identification
3. Discussion
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Fineberg, H.V. Pandemic Preparedness and Response—Lessons from the H1N1 Influenza of 2009. N. Engl. J. Med. 2014, 370, 1335–1342. [Google Scholar] [CrossRef] [Green Version]
- Wang, W.; Xu, Y.; Gao, R.; Lu, R.; Han, K.; Wu, G.; Tan, W. Detection of SARS-CoV-2 in Different Types of Clinical Specimens. JAMA 2020, 323, 1843–1844. [Google Scholar] [CrossRef] [Green Version]
- Zhang, M.; Feng, C.; Zhang, X.; Hu, S.; Zhang, Y.; Min, M.; Liu, B.; Ying, X.; Liu, Y. Susceptibility Factors of Stomach for SARS-CoV-2 and Treatment Implication of Mucosal Protective Agent in COVID-19. Front. Med. 2021, 7, 597967. [Google Scholar] [CrossRef]
- Chen, Y.; Chen, L.; Deng, Q.; Zhang, G.; Wu, K.; Ni, L.; Yang, Y.; Liu, B.; Wang, W.; Wei, C.; et al. The presence of SARS-CoV-2 RNA in the feces of COVID-19 patients. J. Med. Virol. 2020, 92, 833–840. [Google Scholar] [CrossRef] [Green Version]
- Amoah, I.D.; Kumari, S.; Bux, F. Coronaviruses in wastewater processes: Source, fate and potential risks. Environ. Int. 2020, 143, 105962. [Google Scholar] [CrossRef]
- Foladori, P.; Cutrupi, F.; Segata, N.; Manara, S.; Pinto, F.; Malpei, F.; Bruni, L.; La Rosa, G. SARS-CoV-2 from faeces to wastewater treatment: What do we know? A review. Sci. Total Environ. 2020, 743, 140444. [Google Scholar] [CrossRef]
- Larsen, D.A.; Wigginton, K.R. Tracking COVID-19 with wastewater. Nat. Biotechnol. 2020, 38, 1151–1153. [Google Scholar] [CrossRef]
- Daughton, C. The international imperative to rapidly and inexpensively monitor community-wide COVID-19 infection status and trends. Sci. Total Environ. 2020, 726, 138149. [Google Scholar] [CrossRef]
- Haramoto, E.; Malla, B.; Thakali, O.; Kitajima, M. First environmental surveillance for the presence of SARS-CoV-2 RNA in wastewater and river water in Japan. Sci. Total Environ. 2020, 737, 140405. [Google Scholar] [CrossRef]
- Hasan, S.W.; Ibrahim, Y.; Daou, M.; Kannout, H.; Jan, N.; Lopes, A.; Alsafar, H.; Yousef, A.F. Detection and quantification of SARS-CoV-2 RNA in wastewater and treated effluents: Surveillance of COVID-19 epidemic in the United Arab Emirates. Sci. Total Environ. 2021, 764, 142929. [Google Scholar] [CrossRef]
- Izquierdo-Lara, R.; Elsinga, G.; Heijnen, L.; Oude Munnink, B.B.; Schapendonk, C.M.E.; Nieuwenhuijse, D.; Kon, M.; Lu, L.; Aarestrup, F.M.; Lycett, S.; et al. Monitoring SARS-CoV-2 circulation and diversity through community wastewater sequencing. Public Glob. Health 2020, 27, 1405–1415. [Google Scholar] [CrossRef]
- Mlejnkova, H.; Sovova, K.; Vasickova, P.; Ocenaskova, V.; Jasikova, L.; Juranova, E. Preliminary Study of Sars-Cov-2 Occurrence in Wastewater in the Czech Republic. Int. J. Environ. Res. Public Health 2020, 17, 5508. [Google Scholar] [CrossRef]
- Randazzo, W.; Truchado, P.; Cuevas-Ferrando, E.; Simón, P.; Allende, A.; Sánchez, G. SARS-CoV-2 RNA in wastewater anticipated COVID-19 occurrence in a low prevalence area. Water Res. 2020, 181, 115942. [Google Scholar] [CrossRef]
- Westhaus, S.; Weber, F.-A.; Schiwy, S.; Linnemann, V.; Brinkmann, M.; Widera, M.; Greve, C.; Janke, A.; Hollert, H.; Wintgens, T.; et al. Detection of SARS-CoV-2 in raw and treated wastewater in Germany–Suitability for COVID-19 surveillance and potential transmission risks. Sci. Total Environ. 2021, 751, 141750. [Google Scholar] [CrossRef]
- Kitajima, M.; Ahmed, W.; Bibby, K.; Carducci, A.; Gerba, C.P.; Hamilton, K.A.; Haramoto, E.; Rose, J.B. SARS-CoV-2 in wastewater: State of the knowledge and research needs. Sci. Total Environ. 2020, 739, 139076. [Google Scholar] [CrossRef]
- Polo, D.; Quintela-Baluja, M.; Corbishley, A.; Jones, D.L.; Singer, A.C.; Graham, D.W.; Romalde, J.L. Making waves: Wastewater-based epidemiology for COVID-19–approaches and challenges for surveillance and prediction. Water Res. 2020, 186, 116404. [Google Scholar] [CrossRef]
- Thompson, J.R.; Nancharaiah, Y.V.; Gu, X.; Lee, W.L.; Rajal, V.B.; Haines, M.B.; Girones, R.; Ng, L.C.; Alm, E.J.; Wuertz, S. Making waves: Wastewater surveillance of SARS-CoV-2 for population-based health management. Water Res. 2020, 184, 116181. [Google Scholar] [CrossRef]
- Singh, L.; Anyaneji, U.J.; Ndifon, W.; Turok, N.; Mattison, S.A.; Lessells, R.; Sinayskiy, I.; San, E.J.; Tegally, H.; Barnett, S.; et al. Implementation of an efficient SARS-CoV-2 specimen pooling strategy for high throughput diagnostic testing. Sci. Rep. 2021, 11, 17793. [Google Scholar] [CrossRef]
- Crits-Christoph, A.; Kantor, R.S.; Olm, M.R.; Whitney, O.N.; Al-Shayeb, B.; Lou, Y.C.; Flamholz, A.; Kennedy, L.C.; Greenwald, H.; Hinkle, A.; et al. Genome Sequencing of Sewage Detects Regionally Prevalent SARS-CoV-2 Variants. MBio 2021, 12, 9. [Google Scholar] [CrossRef]
- Jahn, K.; Dreifuss, D.; Topolsky, I.; Kull, A.; Ganesanandamoorthy, P.; Fernandez-Cassi, X.; Bänziger, C.; Stachler, E.; Fuhrmann, L.; Jablonski, K.P.; et al. Detection of SARS-CoV-2 variants in Switzerland by genomic analysis of wastewater samples. Infect. Dis. (Except. HIV/AIDS) 2021, 801, 1499757. [Google Scholar] [CrossRef]
- Pechlivanis, N.; Tsagiopoulou, M.; Maniou, M.C.; Togkousidis, A.; Mouchtaropoulou, E.; Chassalevris, T.; Chaintoutis, S.; Dovas, C.; Petala, M.; Kostoglou, M.; et al. Detecting SARS-CoV-2 lineages and mutational load in municipal wastewater; a use-case in the metropolitan area of Thessaloniki, Greece. Epidemiology 2022, 12, 2659. [Google Scholar] [CrossRef]
- Landgraff, C.; Wang, L.Y.R.; Buchanan, C.; Wells, M.; Schonfeld, J.; Bessonov, K.; Ali, J.; Robert, E.; Nadon, C. Metagenomic sequencing of municipal wastewater provides a near-complete SARS-CoV-2 genome sequence identified as the B.1.1.7 variant of concern from a Canadian municipality concurrent with an outbreak. Public Glob. Health 2021. [Google Scholar] [CrossRef]
- Wilton, T.; Bujaki, E.; Klapsa, D.; Fritzsche, M.; Mate, R.; Martin, J. Rapid increase of SARS-CoV-2 variant B.1.1.7 detected in sewage samples from England between October 2020 and January 2021. Msystems 2021, 6, e00353-21. [Google Scholar] [CrossRef]
- Bi, C.; Ramos-Mandujano, G.; Tian, Y.; Hala, S.; Xu, J.; Mfarrej, S.; Esteban, C.R.; Delicado, E.N.; Alofi, F.S.; Khogeer, A.; et al. Simultaneous Detection and Mutation Surveillance of SARS-CoV-2 and co-infections of multiple respiratory viruses by Rapid field-deployable sequencing. Med 2021, 2, 689–700. [Google Scholar] [CrossRef]
- Pérez-Cataluña, A.; Cuevas-Ferrando, E.; Randazzo, W.; Falcó, I.; Allende, A.; Sánchez, G. Comparing analytical methods to detect SARS-CoV-2 in wastewater. Sci. Total Environ. 2021, 758, 143870. [Google Scholar] [CrossRef]
- Smyth, D.S.; Trujillo, M.; Cheung, K.; Gao, A.; Hoxie, I.; Kannoly, S.; Kubota, N.; Markman, M.; San, K.M.; Sompanya, G.; et al. Detection of Mutations Associated with Variants of Concern Via High. Throughput Sequencing of SARS-CoV-2 Isolated from NYC Wastewater. Infect. Dis. (Except. HIV/AIDS) 2021, 848, 157740. [Google Scholar] [CrossRef]
- Fontenele, R.S.; Kraberger, S.; Hadfield, J.; Driver, E.M.; Bowes, D.; Holland, L.A.; Faleye, T.O.C.; Adhikari, S.; Kumar, R.; Inchausti, R.; et al. High-throughput sequencing of SARS-CoV-2 in wastewater provides insights into circulating variants. Water Res. 2021, 205, 117710. [Google Scholar] [CrossRef]
- Fuqua, J.L.; Rouchka, E.C.; Waigel, S.; Sokoloski, K.; Chung, D.; Zacharias, W.; Zhang, M.; Chariker, J.; Talley, D.; Santisteban, I.; et al. A Rapid Assessment of Wastewater for Genomic Surveillance of SARS-CoV-2 Variants at Sewershed Scale in Louisville, KY. Public Glob. Health 2021. [Google Scholar] [CrossRef]
- Prado, T.; Fumian, T.M.; Mannarino, C.F.; Resende, P.C.; Motta, F.C.; Eppinghaus, A.L.F.; Miagostovich, M.P. Wastewater-based epidemiology as a useful tool to track SARS-CoV-2 and support public health policies at municipal level in Brazil. Water Res. 2021, 191, 116810. [Google Scholar] [CrossRef]
- Ahmed, W.; Tscharke, B.; Bertsch, P.M.; Bibby, K.; Bivins, A.; Choi, P.; Clarke, L.; Dwyer, J.; Edson, J.; Nguyen, T.M.H.; et al. SARS-CoV-2 RNA monitoring in wastewater as a potential early warning system for COVID-19 transmission in the community: A temporal case study. Sci. Total Environ. 2021, 761, 144216. [Google Scholar] [CrossRef]
- Gregory, D.A.; Wieberg, C.G.; Wenzel, J.; Lin, C.-H.; Johnson, M.C. Monitoring SARS-CoV-2 Populations in Wastewater by Amplicon Sequencing and Using the Novel Program SAM Refiner. Viruses 2021, 13, 1647. [Google Scholar] [CrossRef]
- Avgeris, M.; Adamopoulos, P.G.; Galani, A.; Xagorari, M.; Gourgiotis, D.; Trougakos, I.P.; Voulgaris, N.; Dimopoulos, M.-A.; Thomaidis, N.S.; Scorilas, A. Novel Nested-Seq Approach for SARS-CoV-2 Real-Time Epidemiology and In-Depth Mutational Profiling in Wastewater. Int. J. Mol. Sci. 2021, 22, 8498. [Google Scholar] [CrossRef]
- Hillary, L.S.; Farkas, K.; Maher, K.H.; Lucaci, A.; Thorpe, J.; Distaso, M.A.; Gaze, W.H.; Paterson, S.; Burke, T.; Connor, T.R.; et al. Monitoring SARS-CoV-2 in municipal wastewater to evaluate the success of lockdown measures for controlling COVID-19 in the UK. Water Res. 2021, 200, 117214. [Google Scholar] [CrossRef]
- Sharif, S.; Ikram, A.; Khurshid, A.; Salman, M.; Mehmood, N.; Arshad, Y.; Ahmed, J.; Safdar, R.M.; Rehman, L.; Mujtaba, G.; et al. Detection of SARs-CoV-2 in wastewater using the existing environmental surveillance network: A potential supplementary system for monitoring COVID-19 transmission. PLoS ONE 2021, 16, e0249568. [Google Scholar] [CrossRef]
- Fongaro, G.; Stoco, P.H.; Souza, D.S.M.; Grisard, E.C.; Magri, M.E.; Rogovski, P.; Schörner, M.A.; Barazzetti, F.H.; Christoff, A.P.; de Oliveira, L.F.V.; et al. The presence of SARS-CoV-2 RNA in human sewage in Santa Catarina, Brazil, November 2019. Sci. Total Environ. 2021, 778, 146198. [Google Scholar] [CrossRef]
- Spurbeck, R.R.; Minard-Smith, A.; Catlin, L. Feasibility of neighborhood and building scale wastewater-based genomic epidemiology for pathogen surveillance. Sci. Total Environ. 2021, 789, 147829. [Google Scholar] [CrossRef]
- Mondal, S.; Feirer, N.; Brockman, M.; Preston, M.A.; Teter, S.J.; Ma, D.; Goueli, S.A.; Moorji, S.; Saul, B.; Cali, J.J. A direct capture method for purification and detection of viral nucleic acid enables epidemiological surveillance of SARS-CoV-2. Sci. Total Environ. 2021, 795, 148834. [Google Scholar] [CrossRef]
- Bar-Or, I.; Weil, M.; Indenbaum, V.; Bucris, E.; Bar-Ilan, D.; Elul, M.; Levi, N.; Aguvaev, I.; Cohen, Z.; Shirazi, R.; et al. Detection of SARS-CoV-2 variants by genomic analysis of wastewater samples in Israel. Sci. Total Environ. 2021, 789, 148002. [Google Scholar] [CrossRef]
- Martínez-Puchol, S.; Itarte, M.; Rusiñol, M.; Forés, E.; Mejías-Molina, C.; Andrés, C.; Antón, A.; Quer, J.; Abril, J.F.; Girones, R.; et al. Exploring the diversity of coronavirus in sewage during COVID-19 pandemic: Don’t miss the forest for the trees. Sci. Total Environ. 2021, 800, 149562. [Google Scholar] [CrossRef]
- Rubio-Acero, R.; Beyerl, J.; Muenchhoff, M.; Roth, M.S.; Castelletti, N.; Paunovic, I.; Radon, K.; Springer, B.; Nagel, C.; Boehm, B.; et al. Spatially resolved qualified sewage spot sampling to track SARS-CoV-2 dynamics in Munich—One year of experience. Sci. Total Environ. 2021, 797, 149031. [Google Scholar] [CrossRef]
- Carcereny, A.; Martínez-Velázquez, A.; Bosch, A.; Allende, A.; Truchado, P.; Cascales, J.; Romalde, J.L.; Lois, M.; Polo, D.; Sánchez, G.; et al. Monitoring Emergence of the SARS-CoV-2 B.1.1.7 Variant through the Spanish National SARS-CoV-2 Wastewater Surveillance System (VATar COVID-19). Environ. Sci. Technol. 2021, 55, 11756–11766. [Google Scholar] [CrossRef] [PubMed]
- Rios, G.; Lacoux, C.; Leclercq, V.; Diamant, A.; Lebrigand, K.; Lazuka, A.; Soyeux, E.; Lacroix, S.; Fassy, J.; Couesnon, A.; et al. Monitoring SARS-CoV-2 variants alterations in Nice neighborhoods by wastewater nanopore sequencing. Lancet Reg. Health Eur. 2021, 10, 100202. [Google Scholar] [CrossRef] [PubMed]
- Swift, C.L.; Isanovic, M.; Correa Velez, K.E.; Norman, R.S. Community-level SARS-CoV-2 sequence diversity revealed by wastewater sampling. Sci. Total Environ. 2021, 801, 149691. [Google Scholar] [CrossRef]
- Wurtz, N.; Revol, O.; Jardot, P.; Giraud-Gatineau, A.; Houhamdi, L.; Soumagnac, C.; Annessi, A.; Lacoste, A.; Colson, P.; Aherfi, S.; et al. Monitoring the Circulation of SARS-CoV-2 Variants by Genomic Analysis of Wastewater in Marseille, South-East France. Pathogens 2021, 10, 1042. [Google Scholar] [CrossRef] [PubMed]
- Ai, Y.; Davis, A.; Jones, D.; Lemeshow, S.; Tu, H.; He, F.; Ru, P.; Pan, X.; Bohrerova, Z.; Lee, J. Wastewater SARS-CoV-2 monitoring as a community-level COVID-19 trend tracker and variants in Ohio, United States. Sci. Total Environ. 2021, 801, 149757. [Google Scholar] [CrossRef] [PubMed]
- Dharmadhikari, T.; Rajput, V.; Yadav, R.; Boargaonkar, R.; Patil, D.; Kale, S.; Kamble, S.P.; Dastager, S.G.; Dharne, M.S. High throughput sequencing based direct detection of SARS-CoV-2 fragments in wastewater of Pune, West India. Sci. Total Environ. 2021, 807, 151038. [Google Scholar] [CrossRef]
- Rouchka, E.C.; Chariker, J.H.; Saurabh, K.; Waigel, S.; Zacharias, W.; Zhang, M.; Talley, D.; Santisteban, I.; Puccio, M.; Moyer, S.; et al. The Rapid Assessment of Aggregated Wastewater Samples for Genomic Surveillance of SARS-CoV-2 on a City-Wide Scale. Pathogens 2021, 10, 1271. [Google Scholar] [CrossRef]
- Vo, V.; Tillett, R.L.; Chang, C.-L.; Gerrity, D.; Betancourt, W.Q.; Oh, E.C. SARS-CoV-2 variant detection at a university dormitory using wastewater genomic tools. Sci. Total Environ. 2022, 805, 149930. [Google Scholar] [CrossRef] [PubMed]
- Cornman, R.S.; McKenna, J.E.; Fike, J.; Oyler-McCance, S.J.; Johnson, R. An experimental comparison of composite and grab sampling of stream water for metagenetic analysis of environmental DNA. PeerJ 2018, 6, e5871. [Google Scholar] [CrossRef] [Green Version]
- Medema, G.; Been, F.; Heijnen, L.; Petterson, S. Implementation of environmental surveillance for SARS-CoV-2 virus to support public health decisions: Opportunities and challenges. Curr. Opin. Environ. Sci. Health 2020, 17, 49–71. [Google Scholar] [CrossRef]
- Michael-Kordatou, I.; Karaolia, P.; Fatta-Kassinos, D. Sewage analysis as a tool for the COVID-19 pandemic response and management: The urgent need for optimised protocols for SARS-CoV-2 detection and quantification. J. Environ. Chem. Eng. 2020, 8, 104306. [Google Scholar] [CrossRef]
- Spurbeck, R.R.; Minard-Smith, A.T.; Catlin, L.A. Applicability of Neighborhood and Building Scale Wastewater-Based Genomic Epidemiology to Track the SARS-CoV-2 Pandemic and other Pathogens. Epidemiology 2021, 800, 149180. [Google Scholar] [CrossRef]
- Hokajärvi, A.-M.; Rytkönen, A.; Tiwari, A.; Kauppinen, A.; Oikarinen, S.; Lehto, K.-M.; Kankaanpää, A.; Gunnar, T.; Al-Hello, H.; Blomqvist, S.; et al. The detection and stability of the SARS-CoV-2 RNA biomarkers in wastewater influent in Helsinki, Finland. Sci. Total Environ. 2021, 770, 145274. [Google Scholar] [CrossRef]
- Dumke, R.; de la Cruz Barron, M.; Oertel, R.; Helm, B.; Kallies, R.; Berendonk, T.U.; Dalpke, A. Evaluation of Two Methods to Concentrate SARS-CoV-2 from Untreated Wastewater. Pathogens 2021, 10, 195. [Google Scholar] [CrossRef]
- Farkas, K.; Hillary, L.S.; Malham, S.K.; McDonald, J.E.; Jones, D.L. Wastewater and public health: The potential of wastewater surveillance for monitoring COVID-19. Curr. Opin. Environ. Sci. Health 2020, 17, 14–20. [Google Scholar] [CrossRef]
- Krebs, S.; Fischaleck, M.; Blum, H. A simple and loss-free method to remove TRIzol contaminations from minute RNA samples. Anal. Biochem. 2009, 387, 136–138. [Google Scholar] [CrossRef]
- Berensmeier, S. Magnetic particles for the separation and purification of nucleic acids. Appl. Microbiol. Biotechnol. 2006, 73, 495–504. [Google Scholar] [CrossRef]
- CDC. Labs. Centers for Disease Control and Prevention. Available online: https://www.cdc.gov/coronavirus/2019-ncov/lab/rt-pcr-panel-primer-probes.html (accessed on 11 February 2020).
- Heikema, A.P.; Horst-Kreft, D.; Boers, S.A.; Jansen, R.; Hiltemann, S.D.; de Koning, W.; Kraaij, R.; de Ridder, M.A.J.; van Houten, C.B.; Bont, L.J.; et al. Comparison of Illumina versus Nanopore 16S rRNA Gene Sequencing of the Human Nasal Microbiota. Genes 2020, 11, 1105. [Google Scholar] [CrossRef]
- McNaughton, A.L.; Roberts, H.E.; Bonsall, D.; de Cesare, M.; Mokaya, J.; Lumley, S.F.; Golubchik, T.; Piazza, P.; Martin, J.B.; de Lara, C.; et al. Illumina and Nanopore methods for whole genome sequencing of hepatitis B virus (HBV). Sci. Rep. 2019, 9, 7081. [Google Scholar] [CrossRef] [Green Version]
- La Rosa, G.; Bonadonna, L.; Lucentini, L.; Kenmoe, S.; Suffredini, E. Coronavirus in water environments: Occurrence, persistence and concentration methods-A scoping review. Water Res. 2020, 179, 115899. [Google Scholar] [CrossRef]
- Schumann, V.-F.; de Castro Cuadrat, R.R.; Wyler, E.; Wurmus, R.; Deter, A.; Quedenau, C.; Dohmen, J.; Faxel, M.; Borodina, T.; Altmüller, J.; et al. COVID-19 infection dynamics revealed by SARS-CoV-2 wastewater sequencing analysis and deconvolution. medRxiv 2021. [Google Scholar] [CrossRef]
- Bull, R.A.; Adikari, T.N.; Ferguson, J.M.; Hammond, J.M.; Stevanovski, I.; Beukers, A.G.; Naing, Z.; Yeang, M.; Verich, A.; Gamaarachchi, H.; et al. Analytical validity of nanopore sequencing for rapid SARS-CoV-2 genome analysis. Nat. Commun. 2020, 11, 6272. [Google Scholar] [CrossRef]
- O’Toole, Á.; Scher, E.; Underwood, A.; Jackson, B.; Hill, V.; McCrone, J.T.; Colquhoun, R.; Ruis, C.; Abu-Dahab, K.; Taylor, B.; et al. Assignment of epidemiological lineages in an emerging pandemic using the pangolin tool. Virus Evol. 2021, 7, veab064. [Google Scholar] [CrossRef]
- La Rosa, G.; Mancini, P.; Bonanno Ferraro, G.; Veneri, C.; Iaconelli, M.; Lucentini, L.; Bonadonna, L.; Brusaferro, S.; Brandtner, D.; Fasanella, A.; et al. Rapid screening for SARS-CoV-2 variants of concern in clinical and environmental samples using nested RT-PCR assays targeting key mutations of the spike protein. Water Res. 2021, 197, 117104. [Google Scholar] [CrossRef]
- Hata, A.; Hara-Yamamura, H.; Meuchi, Y.; Imai, S.; Honda, R. Detection of SARS-CoV-2 in wastewater in Japan during a COVID-19 outbreak. Sci. Total Environ. 2021, 758, 143578. [Google Scholar] [CrossRef]
- Cook, L.; Starr, K.; Boonyaratanakornkit, J.; Hayden, R.; Sam, S.S.; Caliendo, A.M. Does Size Matter? Comparison of Extraction Yields for Different-Sized DNA Fragments by Seven Different Routine and Four New Circulating Cell-Free Extraction Methods. J. Clin. Microbiol. 2018, 56, e01061-18. [Google Scholar] [CrossRef] [Green Version]
- Kemp, S.; Collier, D.; Datir, R.; Ferreira, I.; Gayed, S.; Jahun, A.; Hosmillo, M.; Rees-Spear, C.; Mlcochova, P.; Lumb, I.U.; et al. Neutralising antibodies in Spike mediated SARS-CoV-2 adaptation. medRxiv 2020. [Google Scholar] [CrossRef]
- Panchal, D.; Prakash, O.; Bobde, P.; Pal, S. SARS-CoV-2: Sewage surveillance as an early warning system and challenges in developing countries. Environ. Sci. Pollut. Res. 2021, 28, 22221–22240. [Google Scholar] [CrossRef]
- Castrignanò, E.; Yang, Z.; Feil, E.J.; Bade, R.; Castiglioni, S.; Causanilles, A.; Gracia-Lor, E.; Hernandez, F.; Plósz, B.G.; Ramin, P.; et al. Enantiomeric profiling of quinolones and quinolones resistance gene qnrS in European wastewaters. Water Res. 2020, 175, 115653. [Google Scholar] [CrossRef] [PubMed]
- Xagoraraki, I.; O’Brien, E. Wastewater-Based Epidemiology for Early Detection of Viral Outbreaks. In Women in Water Quality; Springer: Berlin/Heidelberg, Germany, 2020; pp. 75–97. [Google Scholar] [CrossRef] [Green Version]
- Castiglioni, S.; Salgueiro-González, N.; Bijlsma, L.; Celma, A.; Gracia-Lor, E.; Beldean-Galea, M.S.; Mackuľak, T.; Emke, E.; Heath, E.; Kasprzyk-Hordern, B.; et al. New psychoactive substances in several European populations assessed by wastewater-based epidemiology. Water Res. 2021, 195, 116983. [Google Scholar] [CrossRef]
- Ort, C.; van Nuijs, A.L.N.; Berset, J.-D.; Bijlsma, L.; Castiglioni, S.; Covaci, A.; de Voogt, P.; Emke, E.; Fatta-Kassinos, D.; Griffiths, P.; et al. Spatial differences and temporal changes in illicit drug use in Europe quantified by wastewater analysis. Addictions 2014, 109, 1338–1352. [Google Scholar] [CrossRef] [PubMed]
- Thomas, K.V.; Bijlsma, L.; Castiglioni, S.; Covaci, A.; Emke, E.; Grabic, R.; Hernández, F.; Karolak, S.; Kasprzyk-Hordern, B.; Lindberg, R.H.; et al. Comparing illicit drug use in 19 European cities through sewage analysis. Sci. Total Environ. 2012, 432, 432–439. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- NORMAN–SCORE Joint Initiative to Facilitate Data Comparison Between “SARS-CoV-2 in Sewage” Studies|NORMAN (n.d.). Available online: http://www.normandata.eu/?q=node/361 (accessed on 11 April 2021).
- Daughton, C.G. Wastewater surveillance for population-wide COVID-19: The present and future. Sci. Total Environ. 2020, 736, 139631. [Google Scholar] [CrossRef] [PubMed]
| Extraction | RT-qPCR Confirmation Genes | PCR Kit | Seq Technology | Primers | Ref. Genome | Reference |
|---|---|---|---|---|---|---|
| Magnetic extraction with NucliSENS easyMAG system (Biomerieux, Marcy-l’Étoile, France) | N1–N3 and E | Not specified | Illumina (San Diego, CA, USA)/ONT MinION (Oxford, UK) | 2 pools of 89,500 bp primers | 2 datasets—Dutch–Belgian full length SARS-CoV-2 genomes (8th of July 2020) and GISAID database (1st of July 2020) | [11] |
| QiaAmp Viral MiniKit (Qiagen, (Hilden, Germany)) | None | None | Illumina NovaSeq 6000 (San Diego, CA, USA) | ARTIC V3 primers | GISAID | [20] |
| AllPrep DNA/RNA Mini Kit (Qiagen (Hilden, Germany)) | N1 | TaqPath 1-Step RT-qPCR Master Mix or TaqMan Fast Virus 1-Step Master Mix (Thermo Fisher Scientific (Waltham, MA, USA)) | Illumina NextSeq 550 (San Diego, CA, USA) | Respiratory Virus Oligo Panel and Gut Microbiome | EPI_ISL_402124 and GISAID (23 August 2020) | [19] |
| Direct extraction of sample with silica columns (Zymo III-P (Irvine, CA, USA)) | None | |||||
| “Milk of Silica” method | None | |||||
| RNeasy PowerMicrobiome Kit+ Onestep PCR inhibitor removal kit (Zymo Research (Irvine, CA, USA)) following concentration step | N2 | Not specified | Illumina MiSeq (San Diego, CA, USA) | ARTIC V3 primers | NC_045512 | [21] |
| RNeasy Mini Kit (Qiagen, (Hilden, Germany)) | N1 and N2 | 2019-nCoV CDC RUO Kit, TaqPath 1-Step RT-qPCR Master Mix, CG (Thermo Fisher Scientific (Waltham, MA, USA)) | ONT MinION (Oxford, UK) | ARTIC V3 primers | MN908947.3 | [22] |
| RNeasy Mini Kit (Qiagen (Hilden, Germany) | ||||||
| High Pure Viral RNA kit (Roche (Basel, Switzerland) | RdRp, ORF8b, and specific primers for detection of B.1.1.7 variant | Not specified | Illumina MiSeq (San Diego, CA, USA) | Not specified | GISAID (1st of March 2021) | [23] |
| Direct-Zol RNA Miniprep kit (Zymoresearch (Irvine, CA, USA)) or Trizol Reagent (Invitrogen (Waltham, MA, USA)) | N genes | nCov CDC EUA Kit | ONT MinION (Oxford, UK) | 13 sets of primers covering N gene, S gene, ORF1ab, and ORF8 | NC_045512 | [24] |
| Maxwell RSC Pure Food GMO and Authentication Kit (Promega (Madison, WI, USA) | N1, E, and RdRp | RT-qPCR using One Step PrimeScriptTM RT-PCR Kit (Perfect Real Time) (Takara Bio (Kusacu, Japan)) | Illumina MiSeq (San Diego, CA, USA) | ARTIC V3 primers | MN908947.3 | [25] |
| TRIzol extraction | None | None | Illumina iSeq100 (San Diego, CA, USA) | Primers targeting RBD domain | MN908947 | [26] |
| RNaeasy MiniKit (Qiagen (Hilden, Germany)) | E | SuperScript III Platinum One-Step qRT-PCR Kit (Invitrogen (Waltham, MA, USA)) | Illumina HiSeq 2500 (San Diego, CA, USA) | Swift Nomalase Amplicon SARS-CoV-2 panel (SWIFT (Coralville, IA, USA)) | NC_045512.2 | [27] |
| TRIzol and DNA-zol 96 MagBead RNA kit (Zymoresearch (Irvine, CA, USA)) | None | Taq 1-Step Multiplex Master Mix (Thermo Fisher (Waltham, MA, USA)) | Illumina (San Diego, CA, USA) | Swift Nomalase Amplicon SARS-CoV-2 panel (SWIFT (Coralville, IA, USA)) | NC_045512.2 | [28] |
| QiaAmp Viral MiniKit (Qiagen (Hilden, Germany)) | N2 | SuperScriptTM III PlatinumTM One-Step qRT-PCR Kit (Invitrogen (Waltham, MA, USA)) | Illumina MiSeq (San Diego, CA, USA) | Primers targeting whole SARS-CoV-2 genome | Not specified | [29] |
| RNeasy PowerMicrobiome Kit (Qiagen (Hilden, Germany)) | N1, N2, E | iTaq™ Universal Probes One-Step Reaction Mix (Bio-Rad Laboratories (Hercules, Clearwater, FL, USA) | Illumina MiSeq (San Diego, CA, USA) | Six N1 primers | NC_045512.2 | [30] |
| QiaAmp Viral MiniKit (Qiagen (Hilden, Germany)) | None | None | Illumina MiSeq (San Diego, CA, USA) | Primers covering 3 regions of S gene | NC_045512.2 | [31] |
| Water DNA/RNA magnetic bead kit (Idexx (Westbrook, ME, USA) | None | Novel nested and nested real-time PCR, dNTPs mix (Jena Bioscience GmbH (Jena, Germany)), Kapa Taq polymerase (Kapa Biosystems (Potters Bar, UK); for probe fluorescent-based real-time PCR Kapa Probe Fast Universal (2X) qPCR Master Mix (Kapa Biosystems (Potters Bar, UK) | Semi-conductor sequencing technology | In-house design primers for S gene and 5 specific regions | NC_045512.2 | [32] |
| Magnetic extraction with NucliSENS easyMAG system (Biomerieux (Marcy-l’Étoile, France)) | N1 and E | 1 × RNA Ultrasense Reaction Mix with 1 µL RNA Ultrasense Enzyme Mix (Invitrogen (Waltham, MA, USA) | Illumina MiSeq (San Diego, CA, USA) | ARTIC V3 primers | MN908947.3 | [33] |
| Spin star nucleic acid kit (BioservUK (Rotherham, UK)) | ORF1ab, N, and E | (Kit 1) Real-Time Fluorescent RT-PCR Kit for detecting 2019-nCoV by BGI China (Shenzhen, China) (IVD and CE marked; catalogue no. MFG030010), takes ORF 1ab gene as the target domain, (Kit 2) qRT-PCR for Novel Coronavirus (2019-nCoV) Nucleic Acid Diagnostic Kit (PCR-Flourescence, Probing, IVD marked) by Sansure Biotech (Sansure Biotech Inc China, ref no. S3102E, Changsha, China). The kit utilizes the novel coronavirus (2019-nCoV) ORF-1 gene and a conserved coding nucleocapsid protein N-gene as the target regions and finally (Kit 3) detection Kit for 2019 Novel Coronavirus RNA (PCR-Fluorescence Probing), for the E gene: Superscript III one step RT-PCR with platinum Taq Polymerase (Invitrogen (Waltham, MA, USA)) | ABI 3100 (Thermofisher (Waltham, MA, USA)) | ORF1a primer | Not specified | [34] |
| QiaAmp Viral MiniKit (Qiagen (Hilden, Germany)) | S, N1, and RdRP | OneStep qPCR Quantinova kit (Qiagen (Hilden, Germany)) | Illumina MiSeq (San Diego, CA, USA) | S and RdRP primers | NC_045512.2 and other coronaviruses | [35] |
| TRIzol extraction and QiaAmp Viral MiniKit (Qiagen (Hilden, Germany)) | N1 | 4× TaqMan Fast Viral One-Step Master Mix (Thermo Fisher (Waltham, MA, USA)) | Illumina MiSeq (San Diego, CA, USA) | Swift Nomalase Amplicon SARS-CoV-2 panel (SWIFT (Coralville, IA, USA)) | NC_045512.2 | [36] |
| Maxwell RSC Pure Food GMO and Authentication Kit (Promega (Madison, WI, USA)), Maxwell Enviro Wastewater kit (Promega (Madison, WI, USA)), Wizard Enviro Wastewater TNA kit (Promega (Madison, WI, USA)) | N1, N2, and E | SARS-CoV-2 RT-qPCR Detection Kit for Wastewater (Promega Corp. (Madison, WI, USA)) | Illumina MiniSeq (San Diego, CA, USA) | Swift Nomalase Amplicon SARS-CoV-2 Additional Genome Coverage panel (SWIFT (Coralville, IA, USA)) | NC_045512.2 | [37] |
| Magnetic extraction with NucliSENS easyMAG system (Biomerieux (Marcy-l’Étoile, France)) | E | SensiFast reaction mix (Bioline (London, UK)) | Illumina MiSeq (San Diego, CA, USA) | ARTIC V3 primers | NC_045512.2 | [38] |
| QiaAmp Viral MiniKit (Qiagen (Hilden, Germany))) | N1, N2 | Allplex™ 2019-nCoV Assay (Seegene (Wallnut Creek, USA) | Illumina MiSeq (San Diego, CA, USA) | VirCapSeq Enrichment kit (Roche (Basel, Switzerland)) | NC_045512.2 | [39] |
| AllPrep PowerViral DNA/RNA kit and Rneasy Power Microbiome kit (Qiagen (Hilden, Germany))) | N1 | One-step QuantiNova Multiplex RT-PCR Kit | Illumina HiSeq 1500 (San Diego, CA, USA) | ARTIC V2 primers | NC_045512.2 | [40] |
| Maxwell RSC Pure Food GMO and Authentication Kit (Promega (Madison, WI, USA))) and NucleoSpin RNA Virus kit (Macharey-Nagel (Duren, Germany)) | N1, S | N1: PrimeScript One-Step RT-PCR Kit (Takara Bio (Kusacu, Japan)) and 2019-nCoV RUO qPCR Probe Assay primer/probe mix (IDT, Integrated DNA Technologies (Newark, NJ, USA)); S:TaqMan RT-qPCR assay and RT-qPCR mastermix was prepared using the PrimeScript One-Step RT-PCR Kit (Takara Bio (Kusacu, Japan)) | Illumina MiSeq (San Diego, CA, USA) | ARTIC V3 primers (S gene only) | NC_045512.2 | [41] |
| AllPrep PowerViral DNA/RNA kit and Rneasy Power Microbiome kit (Qiagen (Hilden, Germany))) | N1 | qScript XLT One-Step RT-qPCR ToughMix (Quantabio (Beverly, MA, USA)) | ONT PromethION (Oxford, UK) | ARTIC V3 primers | MN908947 | [42] |
| AllPrep PowerViral DNA/RNA kit (Qiagen (Hilden, Germany))) | None | None | ONT GridION X5 (Oxford, UK) | ARTIC V3 primers | MN908947.3 | [43] |
| EZ1 Virus Mini Kit (Qiagen (Hilden, Germany))) | N genes | Superscript III Platinum One-step Quantitative RT-PCR systems with the ROX kit (Invitrogen (Waltham, MA, USA)) | Illumina NovaSeq 6000 (San Diego, CA, USA) | ARTIC V3 primers | NC_045512.2 | [44] |
| RNeasy PowerMicrobiome Kit (Qiagen (Hilden, Germany))) | N1, N2, and E | High-Capacity cDNA Reverse Transcription Kit (Applied Biosystems (Waltham, MA, USA)), ddPCR | Illumina NextSeq 550 (San Diego, CA, USA) | COVIDSeq kit (Illumina (San Diego, USA)) | Not specified | [45] |
| Rneasy Power Water Kit (Qiagen (Hilden, Germany))) | E, RdRP, N | TRUPCR SARS-CoV-2 RT qPCR kit (V-3.2) (3B BlackBio Biotech India Limited (Bhopal, India)) | ONT MinION (Oxford, UK) | ARTIC V3 primers | MN908947.3 | [46] |
| Trizol Reagent (Invitrogen (Waltham, MA, USA)) and Direct-Zol RNA Miniprep kit (Zymoresearch (Irvine, CA, USA)) and RNA cleanud up with Zymo RNA Clean & Concentrator (Zymoresearch (Irvine, CA, USA)) | N1 | Taq 1-Step Multiplex Master Mix (Thermo Fisher Scientific (Waltham, MA, USA)) | Illumina NextSeq 550 (San Diego, CA, USA) | Swift Nomalase Amplicon SARS-CoV-2 panel (SWIFT (Coralville, IA, USA)) | NC_045512.2 | [47] |
| QiaAmp Viral MiniKit (Qiagen (Hilden, Germany))) | None | None | Illumina NextSeq 500 (San Diego, CA, USA) | CleanPlex SARS-CoV-2 FLEX panel (Paragon Genomics) | NC_045512.2 | [48] |
| Institution | Target Genes |
|---|---|
| US CDC, Atlanta, GA, USA | N1, N2, N3 |
| Charité, Berlin, Germany | RdRp, E, N |
| China CDC, Beijing, China | ORF1ab and N |
| Institute Pasteur, Paris, France | Two targets in RdRp (IP2 and IP4) |
| National Institute of Infectious Diseases, Tokyo, Japan | Pancorona and multiple targets, Spike protein |
| HKU, Hong Kong, China | ORF1b-nsp14, N |
| National Institute of Health, Nonthaburi, Thailand | N |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tamáš, M.; Potocarova, A.; Konecna, B.; Klucar, Ľ.; Mackulak, T. Wastewater Sequencing—An Innovative Method for Variant Monitoring of SARS-CoV-2 in Populations. Int. J. Environ. Res. Public Health 2022, 19, 9749. https://doi.org/10.3390/ijerph19159749
Tamáš M, Potocarova A, Konecna B, Klucar Ľ, Mackulak T. Wastewater Sequencing—An Innovative Method for Variant Monitoring of SARS-CoV-2 in Populations. International Journal of Environmental Research and Public Health. 2022; 19(15):9749. https://doi.org/10.3390/ijerph19159749
Chicago/Turabian StyleTamáš, Michal, Alena Potocarova, Barbora Konecna, Ľubos Klucar, and Tomas Mackulak. 2022. "Wastewater Sequencing—An Innovative Method for Variant Monitoring of SARS-CoV-2 in Populations" International Journal of Environmental Research and Public Health 19, no. 15: 9749. https://doi.org/10.3390/ijerph19159749
APA StyleTamáš, M., Potocarova, A., Konecna, B., Klucar, Ľ., & Mackulak, T. (2022). Wastewater Sequencing—An Innovative Method for Variant Monitoring of SARS-CoV-2 in Populations. International Journal of Environmental Research and Public Health, 19(15), 9749. https://doi.org/10.3390/ijerph19159749







