SARS-CoV-2: An Overview of Virus Genetics, Transmission, and Immunopathogenesis
Abstract
1. Introduction
2. SARS-CoV-2: Virus Genetics and Evolution
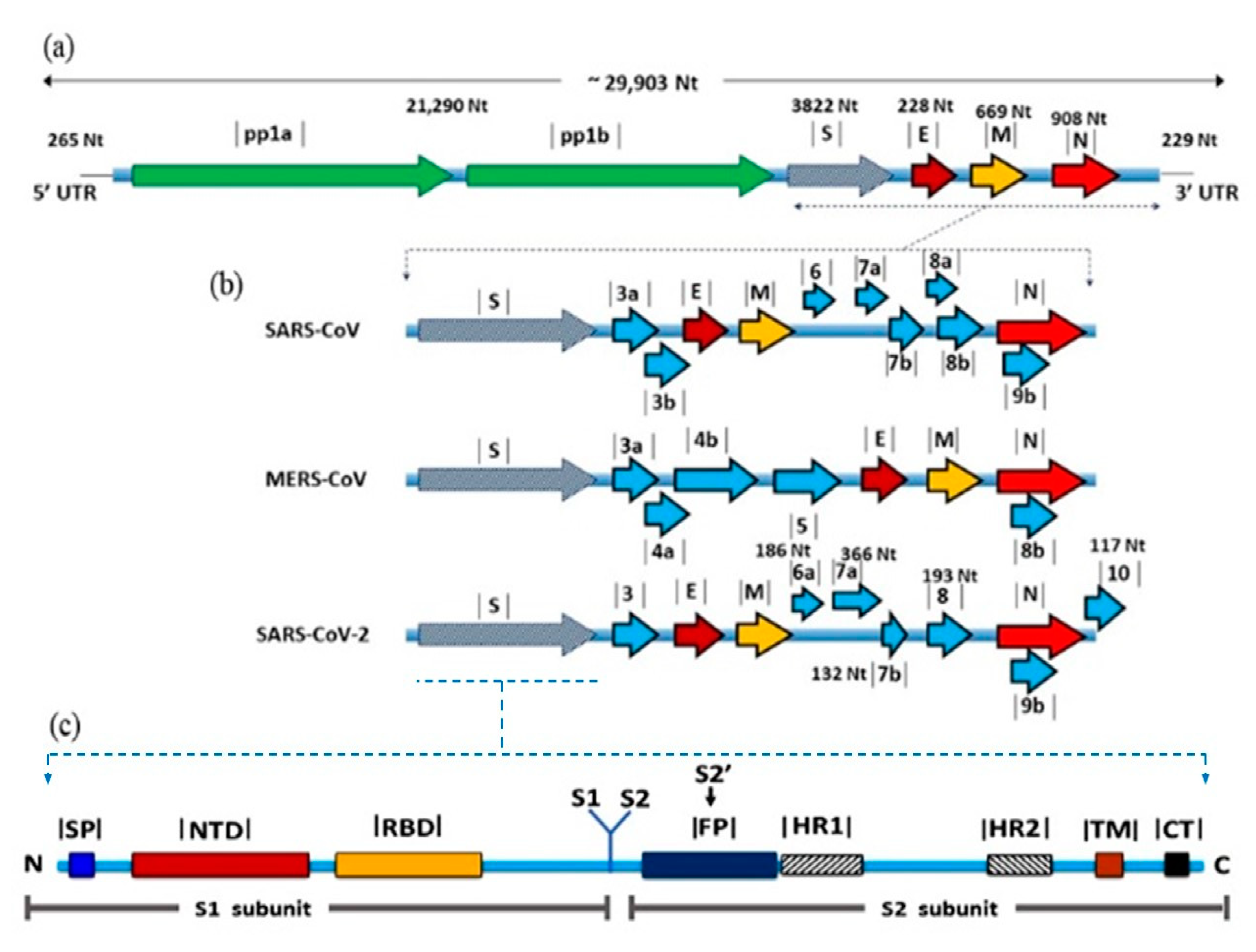
2.1. Taxonomy and Genome Organization
2.2. Origin and Evolution
3. S Protein: Role in Virus Evolution and Cross-Species Transmission
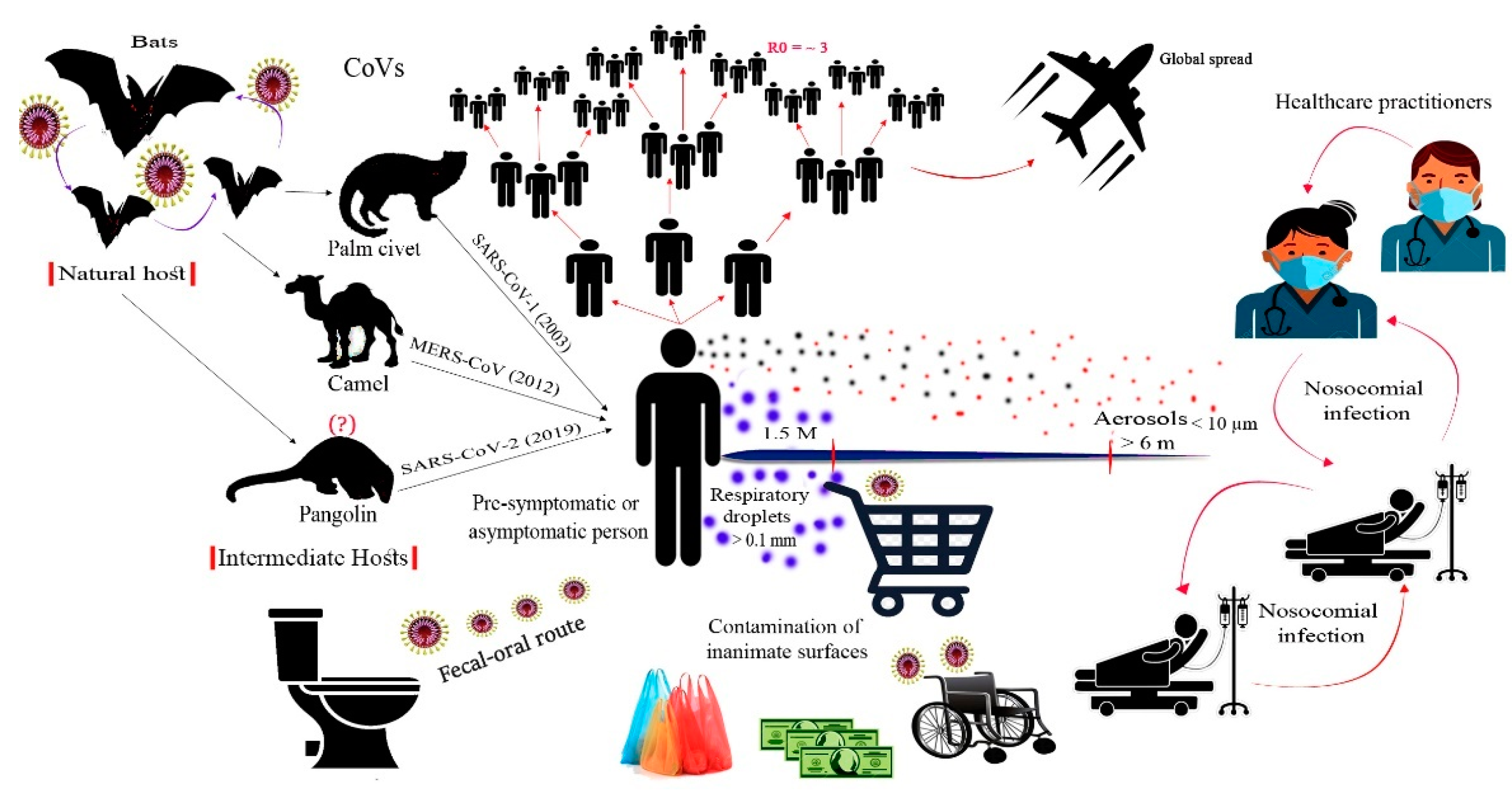
4. Transmission of SARS-CoV-2
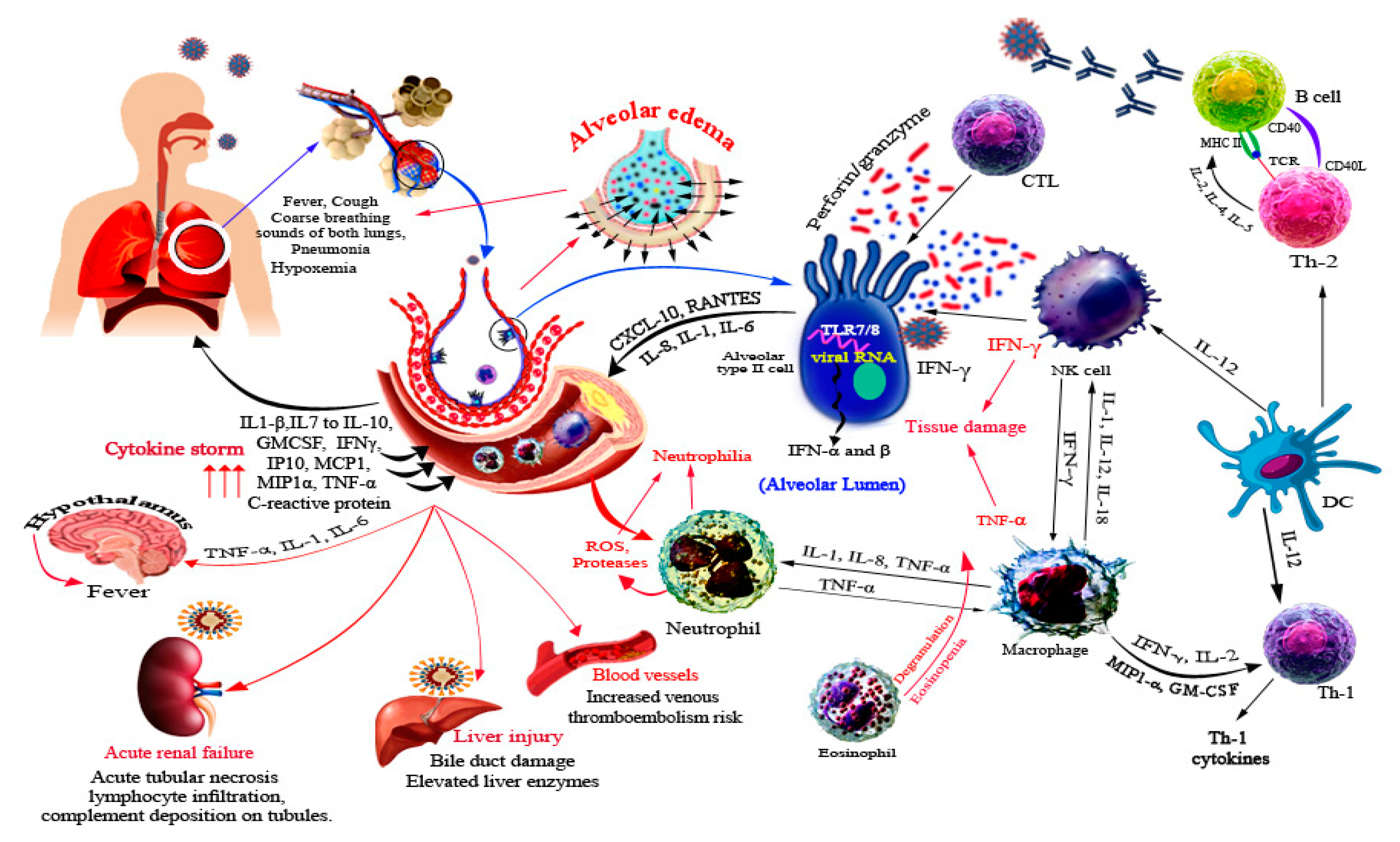
5. Immune Response against SARS-CoV-2
6. SARS-CoV-2 Related Organ Pathogenesis
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Weiss, S.R.; Navas-Martin, S. Coronavirus Pathogenesis and the Emerging Pathogen Severe Acute Respiratory Syndrome Coronavirus. Microbiol. Mol. Biol. Rev. 2005, 69, 635–664. [Google Scholar] [CrossRef]
- Assiri, A.; Al-Tawfiq, J.A.; Al-Rabeeah, A.A.; Al-Rabiah, F.A.; Al-Hajjar, S.; Al-Barrak, A.; Memish, Z.A. Epidemiological, demographic, and clinical characteristics of 47 cases of Middle East respiratory syndrome coronavirus disease from Saudi Arabia: A descriptive study. Lancet Infect. Dis. 2013, 13, 752–761. [Google Scholar] [CrossRef]
- Shang, J.; Ye, G.; Shi, K.; Wan, Y.; Luo, C.; Aihara, H.; Li, F. Structural basis of receptor recognition by SARS-CoV-2. Nature 2020, 581, 221–224. [Google Scholar] [CrossRef]
- COVID-19 situation reports. Available online: https://www.who.int/emergencies/diseases/novel-coronavirus-2019/situation-reports (accessed on 5 January 2020).
- Rossi, G.A.; Sacco, O.; Mancino, E.; Cristiani, L.; Midulla, F. Differences and similarities between SARS-CoV and SARS-CoV-2: Spike receptor-binding domain recognition and host cell infection with support of cellular serine proteases. Infection 2020, 48, 665–669. [Google Scholar] [CrossRef] [PubMed]
- Yadav, T.; Saxena, S.K. Transmission Cycle of SARS-CoV and SARS-CoV-2. In Coronavirus Disease 2019 (COVID-19); Springer: Singapore, 2020; pp. 33–42. [Google Scholar] [CrossRef]
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef]
- Ayittey, F.K.; Ayittey, M.K.; Chiwero, N.B.; Kamasah, J.S.; Dzuvor, C. Economic impacts of Wuhan 2019-nCoV on China and the world. J. Med. Virol. 2020, 92, 473–475. [Google Scholar] [CrossRef] [PubMed]
- Song, Z.; Xu, Y.; Bao, L.; Zhang, L.; Yu, P.; Qu, Y.; Zhu, H.; Zhao, W.; Han, Y.; Qin, C. From SARS to MERS, Thrusting Coronaviruses into the Spotlight. Viruses 2019, 11, 59. [Google Scholar] [CrossRef] [PubMed]
- Sawicki, S.G. Coronavirus Genome Replication. In Viral Genome Replication; Springer: Boston, MA, USA, 2008; pp. 25–39. [Google Scholar] [CrossRef]
- Li, Y.; Li, H.; Fan, R.; Wen, B.; Zhang, J.; Cao, X.; Wang, C.; Song, Z.; Li, S.; Li, X.; et al. Coronavirus Infections in the Central Nervous System and Respiratory Tract Show Distinct Features in Hospitalized Children. Intervirology 2016, 59, 163–169. [Google Scholar] [CrossRef]
- Tang, Q.; Song, Y.; Shi, M.; Cheng, Y.; Zhang, W.; Xia, X.-Q. Inferring the hosts of coronavirus using dual statistical models based on nucleotide composition. Sci. Rep. 2015, 5, 17155. [Google Scholar] [CrossRef] [PubMed]
- Chan, J.F.; Kok, K.H.; Zhu, Z.; Chu, H.; To, K.K.; Yuan, S.; Yuen, K.Y. Genomic characterization of the 2019 novel human-pathogenic coronavirus isolated from a patient with atypical pneumonia after visiting Wuhan. Emerg. Microbes Infect. 2020, 9, 221–236. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Li, X.; Li, T.; Zhang, S.; Wang, L.; Wu, X.; Liu, J. The genetic sequence, origin, and diagnosis of SARS-CoV-2. Eur. J. Clin. Microbiol. Infect. Dis. 2020, 39, 1629–1635. [Google Scholar] [CrossRef]
- Haan, C.A.M.D.; Rottier, P.J.M. Hosting the severe acute respiratory syndrome coronavirus: Specific cell factors required for infection. Cell Microbiol. 2006, 8, 1211–1218. [Google Scholar] [CrossRef]
- Lu, R.; Zhao, X.; Li, J.; Niu, P.; Yang, B.; Wu, H.; Wang, W.; Song, H.; Huang, B.; Zhu, N.; et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: Implications for virus origins and receptor binding. Lancet 2020, 395, 565–574. [Google Scholar] [CrossRef]
- Wu, A.; Peng, Y.; Huang, B.; Ding, X.; Wang, X.; Niu, P.; Meng, J.; Zhu, Z.; Zhang, Z.; Wang, J.; et al. Genome Composition and Divergence of the Novel Coronavirus (2019-nCoV) Originating in China. Cell Host Microbe 2020, 27, 325–328. [Google Scholar] [CrossRef] [PubMed]
- Paraskevis, D.; Kostaki, E.G.; Magiorkinis, G.; Panayiotakopoulos, G.; Sourvinos, G.; Tsiodras, S. Full-genome evolutionary analysis of the novel corona virus (2019-nCoV) rejects the hypothesis of emergence as a result of a recent recombination event. Infect. Genet. Evol. J. Mol. Epidemiol. Evol. Genet. Infect. Dis. 2020, 79, 104212. [Google Scholar] [CrossRef]
- Rehman, S.U.; Shafique, L.; Ihsan, A.; Liu, Q. Evolutionary Trajectory for the Emergence of Novel Coronavirus SARS-CoV-2. Pathogens 2020, 9, 240. [Google Scholar] [CrossRef]
- Guan, Y.; Zheng, B.J.; He, Y.Q.; Liu, X.L.; Zhuang, Z.X.; Cheung, C.L.; Luo, S.W.; Li, P.H.; Zhang, L.J.; Guan, Y.J.; et al. Isolation and characterization of viruses related to the SARS coronavirus from animals in southern China. Science 2003, 302, 276–278. [Google Scholar] [CrossRef]
- Haagmans, B.L.; Al Dhahiry, S.H.; Reusken, C.B.; Raj, V.S.; Galiano, M.; Myers, R.; Godeke, G.J.; Jonges, M.; Farag, E.; Diab, A.; et al. Middle East respiratory syndrome coronavirus in dromedary camels: An outbreak investigation. Lancet Infect. Dis. 2014, 14, 140–145. [Google Scholar] [CrossRef]
- Xiao, K.; Zhai, J.; Feng, Y.; Zhou, N.; Zhang, X.; Zou, J.; Li, N.; Guo, Y.; Li, X.; Shen, X.; et al. Isolation of SARS-CoV-2-related coronavirus from Malayan pangolins. Nature 2020, 583, 286–289. [Google Scholar] [CrossRef]
- Zhang, T.; Wu, Q.; Zhang, Z. Probable Pangolin Origin of SARS-CoV-2 Associated with the COVID-19 Outbreak. Curr. Biol. 2020, 30, 1346–1351. [Google Scholar] [CrossRef]
- Shi, J.; Wen, Z.; Zhong, G.; Yang, H.; Wang, C.; Huang, B.; Liu, R.; He, X.; Shuai, L.; Sun, Z.; et al. Susceptibility of ferrets, cats, dogs, and other domesticated animals to SARS–coronavirus 2. Science 2020, 368, 1016–1020. [Google Scholar] [CrossRef]
- Mandary, M.B.; Masomian, M.; Poh, C.L. Impact of RNA Virus Evolution on Quasispecies Formation and Virulence. Int. J. Mol. Sci. 2019, 20, 4657. [Google Scholar] [CrossRef]
- Holland, L.A.; Kaelin, E.A.; Maqsood, R.; Estifanos, B.; Wu, L.; Varsani, A.; Halden, R.U.; Hogue, B.G.; Scotch, M.; Lim, E.S.; et al. An 81 nucleotide deletion in SARS-CoV-2 ORF7a identified from sentinel surveillance in Arizona (Jan–Mar 2020). J. Virol. 2020. [Google Scholar] [CrossRef]
- Taylor, J.K.; Coleman, C.M.; Postel, S.; Sisk, J.M.; Bernbaum, J.G.; Venkataraman, T.; Sundberg, E.J.; Frieman, M.B. Severe Acute Respiratory Syndrome Coronavirus ORF7a Inhibits Bone Marrow Stromal Antigen 2 Virion Tethering through a Novel Mechanism of Glycosylation Interference. J. Virol. 2015, 89, 11820–11833. [Google Scholar] [CrossRef]
- Tan, Y.; Tan, T.; Lee, M.; Tham, P.; Gunalan, V.; Druce, J.; Birch, C.; Catton, M.; Fu, N.; Yu, N.; et al. Induction of Apoptosis by the Severe Acute Respiratory Syndrome Coronavirus 7a Protein Is Dependent on Its Interaction with the Bcl-XL Protein. J. Virol. 2007, 81, 6346–6355. [Google Scholar] [CrossRef] [PubMed]
- Su, Y.; Anderson, D.; Young, B.E.; Zhu, F.; Linster, M.; Kalimuddin, S.; Low, J.; Yan, Z.; Jayakumar, J.; Sun, L.; et al. Discovery of a 382-nt deletion during the early evolution of SARS-CoV-2. bioRxiv 2020. [Google Scholar] [CrossRef]
- Muth, D.; Corman, V.M.; Roth, H.; Binger, T.; Dijkman, R.; Theresa, L.; Florian, G.G.; Balboni, A.; Battilani, M.; Rihtarič, D.; et al. Attenuation of replication by a 29 nucleotide deletion in SARS-coronavirus acquired during the early stages of human-to-human transmission. Sci. Rep. 2018, 8, 15177. [Google Scholar] [CrossRef]
- Young, B.E.; Fong, S.W.; Chan, Y.H.; Mak, T.; Ang, L.W.; Anderson, D.E.; Lee, C.Y.; Amrun, N.; Lee, B.; Goh, Y.G.; et al. Effects of a major deletion in the SARS-CoV-2 genome on the severity of infection and the inflammatory response: An observational cohort study. Lancet 2020, 396, 603–611. [Google Scholar] [CrossRef]
- Jewell, B.L. Monitoring differences between the SARS-CoV-2 B.1.1.7 variant and other lineages. Lancet 2021, 6, E267–E268. [Google Scholar]
- Hoffmann, M.; Arora, P.; Groß, R.; Seidel, A.; Hörnich, B.F.; Hahn, A.S.; Krüger, N.; Graichen, L.; Hofmann-Winkler, H.; Kempf, A.; et al. SARS-CoV-2 variants B.1.351 and P.1 escape from neutralizing antibodies. Cell 2021, 184, 2384–2393.e2312. [Google Scholar] [CrossRef] [PubMed]
- Burki, T. Understanding variants of SARS-CoV-2. Lancet 2021, 397, 462. [Google Scholar] [CrossRef]
- Karim, S.S.A. Vaccines and SARS-CoV-2 variants: The urgent need for a correlate of protection. Lancet 2021, 397, 1263–1264. [Google Scholar] [CrossRef]
- Bian, L.; Gao, F.; Zhang, J.; He, Q.; Mao, Q.; Xu, M.; Liang, Z. Effects of SARS-CoV-2 variants on vaccine efficacy and response strategies. Expert Rev. Vaccines 2021. [Google Scholar] [CrossRef] [PubMed]
- Vijgen, L.; Keyaerts, E.; Lemey, P.; Maes, P.; Van, K.; Nauwynck, H.; Pensaert, M.; Van Ranst, M. Evolutionary history of the closely related group 2 coronaviruses: Porcine hemagglutinating encephalomyelitis virus, bovine coronavirus, and human coronavirus OC43. J. Virol. 2006, 80, 7270–7274. [Google Scholar] [CrossRef] [PubMed]
- Alekseev, K.P.; Vlasova, A.N.; Jung, K.; Hasoksuz, M.; Zhang, X.; Halpin, R.; Wang, S.; Ghedin, E.; Spiro, D.; Saif, L.J.; et al. Bovine-Like Coronaviruses Isolated from Four Species of Captive Wild Ruminants Are Homologous to Bovine Coronaviruses, Based on Complete Genomic Sequences. J. Virol. 2008, 82, 12422–12431. [Google Scholar] [CrossRef] [PubMed]
- Vijgen, L.; Keyaerts, E.; Moës, E.; Thoelen, I.; Wollants, E.; Lemey, P.; Vandamme, A.M.; Van Ranst, M. Complete genomic sequence of human coronavirus OC43: Molecular clock analysis suggests a relatively recent zoonotic coronavirus transmission event. J. Virol. 2005, 79, 1595–1604. [Google Scholar] [CrossRef] [PubMed]
- Pfefferle, S.; Oppong, S.; Drexler, J.F.; Gloza-Rausch, F.; Ipsen, A.; Seebens, A.; Müller, M.A.; Annan, A.; Vallo, P.; Adu-Sarkodie, Y.; et al. Distant Relatives of Severe Acute Respiratory Syndrome Coronavirus and Close Relatives of Human Coronavirus 229E in Bats, Ghana. Emerg. Infect. Dis. 2009, 15, 1377–1384. [Google Scholar] [CrossRef] [PubMed]
- Li, F. Structure, Function, and Evolution of Coronavirus Spike Proteins. Annu. Rev. Virol. 2016, 3, 237–261. [Google Scholar] [CrossRef] [PubMed]
- Wong, S.K.; Li, W.; Moore, M.J.; Choe, H.; Farzan, M. A 193-amino acid fragment of the SARS coronavirus S protein efficiently binds angiotensin-converting enzyme 2. J. Biol. Chem. 2004, 279, 3197–3201. [Google Scholar] [CrossRef]
- Du, L.; Zhao, G.; Kou, Z.; Ma, C.; Sun, S.; Poon, V.K.; Lu, L.; Wang, L.; Debnath, A.K.; Zheng, B.J.; et al. Identification of a receptor-binding domain in the S protein of the novel human coronavirus Middle East respiratory syndrome coronavirus as an essential target for vaccine development. J. Virol. 2013, 87, 9939–9942. [Google Scholar] [CrossRef]
- Hofmann, H.; Pyrc, K.; van der Hoek, L.; Geier, M.; Berkhout, B.; Pöhlmann, S. Human coronavirus NL63 employs the severe acute respiratory syndrome coronavirus receptor for cellular entry. Proc. Natl. Acad. Sci. USA 2005, 102, 7988–7993. [Google Scholar] [CrossRef] [PubMed]
- Hofmann, H.; Simmons, G.; Rennekamp, A.J.; Chaipan, C.; Gramberg, T.; Heck, E.; Geier, M.; Wegele, A.; Marzi, A.; Bates, P.; et al. Highly Conserved Regions within the Spike Proteins of Human Coronaviruses 229E and NL63 Determine Recognition of Their Respective Cellular Receptors. J. Virol. 2006, 80, 8639–8652. [Google Scholar] [CrossRef] [PubMed]
- Delmas, B.; Gelfi, J.; L’Haridon, R.; Vogelt, L.K.; Sjostromt, H.; Norent, O.; Laude, H. Aminopeptidase N is a major receptor for the enteropathogenic coronavirus TGEV. Nature 1992, 357, 417–420. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Tang, J.; Ma, Y.; Liang, X.; Yang, Y.; Peng, G.; Qi, Q.; Jiang, S.; Li, J.; Du, L.; et al. Receptor usage and cell entry of porcine epidemic diarrhea coronavirus. J. Virol. 2015, 89, 6121–6125. [Google Scholar] [CrossRef] [PubMed]
- Delmas, B.; Gelfi, J.; Sjöström, H.; Noren, O.; Laude, H. Further Characterization of Aminopeptidase-N as a Receptor for Coronaviruses. In Coronaviruses: Molecular Biology and Virus-Host Interactions; Laude, H., Vautherot, J.-F., Eds.; Springer: Boston, MA, USA, 1993; pp. 293–298. [Google Scholar]
- Raj, V.S.; Mou, H.; Smits, S.L.; Dekkers, D.H.; Müller, M.A.; Dijkman, R.; Muth, D.; Demmers, A.; Zaki, A.; Fouchier, R.M.; et al. Dipeptidyl peptidase 4 is a functional receptor for the emerging human coronavirus-EMC. Nature 2013, 495, 251–254. [Google Scholar] [CrossRef]
- Williams, R.K.; Jiang, G.S.; Holmes, K.V. Receptor for mouse hepatitis virus is a member of the carcinoembryonic antigen family of glycoproteins. Proc. Natl. Acad. Sci. USA 1991, 88, 5533–5536. [Google Scholar] [CrossRef] [PubMed]
- Schultze, B.; Gross, H.J.; Brossmer, R.; Herrler, G. The S protein of bovine coronavirus is a hemagglutinin recognizing 9-O-acetylated sialic acid as a receptor determinant. J. Virol. 1991, 65, 6232–6237. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Moore, M.J.; Vasilieva, N.; Sui, J.; Wong, S.K.; Berne, M.A.; Somasundaran, M.; Sullivan, J.L.; Luzuriaga, K.; Greenough, T.C.; et al. Angiotensin-converting enzyme 2 is a functional receptor for the SARS coronavirus. Nature 2003, 426, 450–454. [Google Scholar] [CrossRef]
- Walls, A.C.; Park, Y.-J.; Tortorici, M.A.; Wall, A.; McGuire, A.T.; Veesler, D. Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein. Cell 2020, 181, 281–292.e6. [Google Scholar] [CrossRef] [PubMed]
- Schultze, B.; Cavanagh, D.; Herrler, G. Neuraminidase treatment of avian infectious bronchitis coronavirus reveals a hemagglutinating activity that is dependent on sialic acid-containing receptors on erythrocytes. Virology 1992, 189, 792–794. [Google Scholar] [CrossRef]
- Li, F. Receptor recognition and cross-species infections of SARS coronavirus. Antiviral. Res. 2013, 100, 246–254. [Google Scholar] [CrossRef]
- Braun, E.; Sauter, D. Furin-mediated protein processing in infectious diseases and cancer. Clin. Transl. Immunol. 2019, 8, e1073. [Google Scholar] [CrossRef] [PubMed]
- Claas, E.C.; Osterhaus, A.D.; van Beek, R.; Jong, J.C.; Rimmelzwaan, G.F.; Senne, D.A.; Krauss, S.; Shortridge, K.F.; Webster, R.G. Human influenza A H5N1 virus related to a highly pathogenic avian influenza virus. Lancet Lond Engl. 1998, 351, 472–477. [Google Scholar] [CrossRef]
- Izaguirre, G. The Proteolytic Regulation of Virus Cell Entry by Furin and Other Proprotein Convertases. Viruses 2019, 11, 837. [Google Scholar] [CrossRef]
- Yamada, Y.; Liu, D.X. Proteolytic activation of the spike protein at a novel RRRR/S motif is implicated in furin-dependent entry, syncytium formation, and infectivity of coronavirus infectious bronchitis virus in cultured cells. J. Virol. 2009, 83, 8744–8758. [Google Scholar] [CrossRef]
- Cheng, J.; Zhao, Y.; Xu, G.; Zhang, K.; Jia, W.; Sun, Y.; Zhao, J.; Xue, J.; Hu, Y.; Zhang, G. The S2 Subunit of QX-type Infectious Bronchitis Coronavirus Spike Protein Is an Essential Determinant of Neurotropism. Viruses 2019, 11, 972. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Zhang, Y.; Wu, L.; Niu, C.; Song, Z.; Zhang, G.; Lu, C.; Qiao, Y.; Hu, K.; Yuen, Y.; et al. Structural and Functional Basis of SARS-CoV-2 Entry by Using Human ACE2. Cell 2020, 181, 894–904.e9. [Google Scholar] [CrossRef] [PubMed]
- Shang, J.; Wan, Y.; Luo, C.; Ye, G.; Geng, Q.; Auerbach, A.; Li, F. Cell entry mechanisms of SARS-CoV-2. Proc. Natl. Acad. Sci. USA 2020, 117, 11727–11734. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Y.; Cao, D.; Zhang, Y.; Ma, J.; Qi, J.; Wang, Q.; Lu, G.; Wu, Y.; Yan, J.; Shi, Y.; et al. Cryo-EM structures of MERS-CoV and SARS-CoV spike glycoproteins reveal the dynamic receptor binding domains. Nat. Commun. 2017, 8, 15092. [Google Scholar] [CrossRef]
- Correia, G.; Rodrigues, L.; Gameiro da Silva, M.; Gonçalves, T. Airborne route and bad use of ventilation systems as non-negligible factors in SARS-CoV-2 transmission. Med. Hypotheses 2020, 141, 109781. [Google Scholar] [CrossRef]
- Guo, Y.-R.; Cao, Q.-D.; Hong, Z.S.; Tan, Y.; Chen, S.; Jin, H.; Tan, K.; Wang, D.; Yan, Y. The origin, transmission and clinical therapies on coronavirus disease 2019 (COVID-19) outbreak-an update on the status. Mil. Med. Res. 2020, 7, 11. [Google Scholar] [CrossRef] [PubMed]
- Meselson, M. Droplets and Aerosols in the Transmission of SARS-CoV-2. N. Engl. J. Med. 2020, 382, 2063. [Google Scholar] [CrossRef] [PubMed]
- Xiao, S.; Li, Y.; Wong, T.; Hui, D.S.C. Role of fomites in SARS transmission during the largest hospital outbreak in Hong Kong. PLoS ONE 2017, 12. [Google Scholar] [CrossRef]
- Zhou, J.; Li, C.; Zhao, G.; Chu, H.; Wang, D.; Yan, H.H.; Poon, V.K.; Wen, L.; Wong, B.H.; Zhao, X.; et al. Human intestinal tract serves as an alternative infection route for Middle East respiratory syndrome coronavirus. Sci. Adv. 2017, 3, eaao4966. [Google Scholar] [CrossRef]
- Wang, W.; Xu, Y.; Gao, R.; Lu, R.; Han, K.; Wu, G.; Tan, W. Detection of SARS-CoV-2 in Different Types of Clinical Specimens. JAMA 2020, 323, 1843–1844. [Google Scholar] [CrossRef] [PubMed]
- Knowlton, S.D.; Boles, C.L.; Perencevich, E.N.; Diekema, D.J.; Nonnenmann, M.W.; CDC Epicenters Program. Bioaerosol concentrations generated from toilet flushing in a hospital-based patient care setting. Antimicrob. Resist. Infect. Control. 2018, 7, 16. [Google Scholar] [CrossRef] [PubMed]
- Zeng, H.; Xu, C.; Fan, J.; Tang, Y.; Deng, Q.; Zhang, W.; Long, X. Antibodies in Infants Born to Mothers With COVID-19 Pneumonia. JAMA 2020, 323, 1848–1849. [Google Scholar] [CrossRef]
- Zeng, L.; Xia, S.; Yuan, W.; Yan, K.; Xiao, F.; Shao, J.; Zhou, W. Neonatal Early-Onset Infection With SARS-CoV-2 in 33 Neonates Born to Mothers With COVID-19 in Wuhan, China. JAMA Pediatr. 2020. [Google Scholar] [CrossRef]
- Wei, W.E. Presymptomatic Transmission of SARS-CoV-2—Singapore, January 23–March 16, 2020. MMWR Morb. Mortal. Wkly. Rep. 2020, 69. [Google Scholar] [CrossRef] [PubMed]
- Fiorillo, L.; Cervino, G.; Matarese, M.; D’Amico, C.; Surace, G.; Paduano, V.; Fiorillo, M.T.; Moschella, A.; Bruna, A.; Romano, G.L.; et al. COVID-19 Surface Persistence: A Recent Data Summary and Its Importance for Medical and Dental Settings. Int. J. Environ. Res. Public Health 2020, 17, 3132. [Google Scholar] [CrossRef] [PubMed]
- Chin, A.W.H.; Chu, J.T.S.; Perera, M.R.A.; Hui, K.P.Y.; Yen, H.-L.; Chan, M.C.W.; Peiris, M.; Poon, L.L.M. Stability of SARS-CoV-2 in different environmental conditions. Lancet Microbe 2020, 1, e10. [Google Scholar] [CrossRef]
- Felsenstein, S.; Herbert, J.A.; McNamara, P.S.; Hedrich, C.M. COVID-19: Immunology and treatment options. Clin. Immunol. 2020, 215, 108448. [Google Scholar] [CrossRef]
- Prompetchara, E.; Ketloy, C.; Palaga, T. Immune responses in COVID-19 and potential vaccines: Lessons learned from SARS and MERS epidemic. Asian Pac. J. Allergy Immunol. 2020, 38, 1–9. [Google Scholar] [CrossRef]
- Ben Addi, A.; Lefort, A.; Hua, X.; Libert, F.; Communi, D.; Ledent, C.; Macours, P.; Tilley, S.L.; Boeynaems, J.M.; Robaye, B. Modulation of murine dendritic cell function by adenine nucleotides and adenosine: Involvement of the A(2B) receptor. Eur. J. Immunol. 2008, 38, 1610–1620. [Google Scholar] [CrossRef]
- De Wit, E.; Van Doremalen, N.; Falzarano, D.; Munster, V.J. SARS and MERS: Recent insights into emerging coronaviruses. Nat. Rev. Microbiol. 2016, 14, 523–534. [Google Scholar] [CrossRef]
- Zhao, Y.; Zhao, Z.; Wang, Y.; Zhou, Y.; Ma, Y.; Zuo, W. Single-cell RNA expression profiling of ACE2, the putative receptor of Wuhan 2019-nCov. bioRxiv 2020. [Google Scholar] [CrossRef]
- Zhang, B.; Zhou, X.; Qiu, Y.; Feng, F.; Feng, J.; Jia, Y.; Zhu, H.; Hu, K.; Liu, J.; Liu, Z.; et al. Clinical characteristics of 82 death cases with COVID-19. medRxiv. [CrossRef]
- Qin, C.; Zhou, L.; Hu, Z.; Zhang, S.; Yang, S.; Tao, Y.; Xie, C.; Ma, K.; Shang, K.; Wang, W.; et al. Dysregulation of Immune Response in Patients With Coronavirus 2019 (COVID-19) in Wuhan, China. Clin. Infect. Dis. 2020, 71. [Google Scholar] [CrossRef]
- Rokni, M.; Ghasemi, V.; Tavakoli, Z. Immune responses and pathogenesis of SARS-CoV-2 during an outbreak in Iran: Comparison with SARS and MERS. Rev. Med. Virol. 2020, 30, e2107. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.; Rhee, J.-W.; Cheng, P.; Waliany, S.; Chang, A.; Witteles, R.M.; Maecker, H.; Davis, M.M.; Nguyen, P.K.; Wu, S.M.; et al. Cardiovascular Complications in Patients with COVID-19: Consequences of Viral Toxicities and Host Immune Response. Curr. Cardiol. Rep. 2020, 22, 32. [Google Scholar] [CrossRef]
- Xu, Z.; Shi, L.; Wang, Y.; Zhang, J.; Huang, L.; Zhang, C.; Liu, S.; Zhao, P.; Liu, H.; Zhu, L.; et al. Pathological findings of COVID-19 associated with acute respiratory distress syndrome. Lancet Respir. Med. 2020, 8, 420–422. [Google Scholar] [CrossRef]
- Rojas, M.; Rodríguez, Y.; Monsalve, D.M.; Acosta-Ampudia, Y.; Camacho, B.; Gallo, J.E.; Rojas-Villarraga, A.; Ramírez-Santana, C.; Díaz-Coronado, J.C.; Manrique, R.; et al. Convalescent plasma in Covid-19: Possible mechanisms of action. Autoimmun. Rev. 2020, 19, 102554. [Google Scholar] [CrossRef] [PubMed]
- Wajnberg, A.; Mansour, M.; Leven, E.; Bouvier, N.M.; Patel, G.; Firpo, A.; Mendu, R.; Jhang, J.; Arinsburg, S.; Gitman, M.; et al. Humoral immune response and prolonged PCR positivity in a cohort of 1343 SARS-CoV 2 patients in the New York City region. medRxiv 2020. [Google Scholar] [CrossRef]
- Yuki, K.; Fujiogi, M.; Koutsogiannaki, S. COVID-19 pathophysiology: A review. Clin. Immunol. 2020, 215, 108427. [Google Scholar] [CrossRef]
- Fanelli, V.; Fiorentino, M.; Cantaluppi, V.; Gesualdo, E.L.; Stallone, G.; Ronco, C.; Castellano, G. Acute kidney injury in SARS-CoV-2 infected patients. Crit. Care 2020, 24, 155. [Google Scholar] [CrossRef]
- Li, H.; Liu, L.; Zhang, D.; Xu, J.; Dai, H.; Tang, N.; Su, X.; Calo, B. SARS-CoV-2 and viral sepsis: Observations and hypotheses. Lancet 2020, 395, 1517–1520. [Google Scholar] [CrossRef]
- Zhou, M.; Zhang, X.; Qu, J. Coronavirus disease 2019 (COVID-19): A clinical update. Front. Med. 2020, 14, 126–135. [Google Scholar] [CrossRef] [PubMed]
- Ng, W.H.; Tipih, T.; Makoah, N.A.; Vermeulen, J.G.; Goedhals, D.; Sempa, J.B.; Burt, F.J.; Taylor, A.; Mahalingam, S. Comorbidities in SARS-CoV-2 Patients: A Systematic Review and Meta-Analysis. mBio 2021, 12, 1. [Google Scholar] [CrossRef] [PubMed]
- Peng, Y.H.; Lin, Y.S.; Chen, C.H.; Tsai, K.Y.; Hung, Y.C.; Chen, H.J.; Liao, W.C.; Ho, W.C. Type 1 diabetes is associated with an increased risk of venous thromboembolism: A retrospective population-based cohort study. PLoS ONE 2020, 15, e0226997. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Xu, Y.Z.; Liu, B.; Wu, R.; Yang, Y.Y.; Xiao, X.Q.; Zhang, X. Pioglitazone upregulates angiotensin converting enzyme 2 expression in insulin-sensitive tissues in rats with high-fat diet-induced nonalcoholic steatohepatitis. Sci. World J. 2014, 2014, 603409. [Google Scholar] [CrossRef] [PubMed]
| Genus | Species | Receptor | Fusion Triggers | Reference |
|---|---|---|---|---|
| Alphacoronavirus | HCoV-NL63 | ACE2 | [44] | |
| HCoV-229E | Aminopeptidase N (APN) | [45] | ||
| TGEV | [46] | |||
| PEDV | [47] | |||
| PRCV | [48] [48] [48] | |||
| Feline-CoV | ||||
| Canine-CoV | ||||
| Betacoronavirus | MERS-CoV | Dipeptidyl peptidase 4 (DPP4) |
| [49] |
| MHV | Carcinoembryonic antigen-related cell adhesion molecule 1 |
| [50] | |
| Bovine-CoV | 5-N-acetyl-9-O-acetyl neuraminic acid, Neu 5,9 Ac2 | [51] | ||
| HCoV-OC43 | ||||
| HKU4 | Dipeptidyl peptidase 4 |
| ||
| SARS-CoV-1 | ACE2 |
| [52] | |
| SARS-CoV-2 | [53] | |||
| Gammacoronavirs | IBV | Sugar | [54] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Farrag, M.A.; Amer, H.M.; Bhat, R.; Hamed, M.E.; Aziz, I.M.; Mubarak, A.; Dawoud, T.M.; Almalki, S.G.; Alghofaili, F.; Alnemare, A.K.; et al. SARS-CoV-2: An Overview of Virus Genetics, Transmission, and Immunopathogenesis. Int. J. Environ. Res. Public Health 2021, 18, 6312. https://doi.org/10.3390/ijerph18126312
Farrag MA, Amer HM, Bhat R, Hamed ME, Aziz IM, Mubarak A, Dawoud TM, Almalki SG, Alghofaili F, Alnemare AK, et al. SARS-CoV-2: An Overview of Virus Genetics, Transmission, and Immunopathogenesis. International Journal of Environmental Research and Public Health. 2021; 18(12):6312. https://doi.org/10.3390/ijerph18126312
Chicago/Turabian StyleFarrag, Mohamed A., Haitham M. Amer, Rauf Bhat, Maaweya E. Hamed, Ibrahim M. Aziz, Ayman Mubarak, Turki M Dawoud, Sami G Almalki, Fayez Alghofaili, Ahmad K. Alnemare, and et al. 2021. "SARS-CoV-2: An Overview of Virus Genetics, Transmission, and Immunopathogenesis" International Journal of Environmental Research and Public Health 18, no. 12: 6312. https://doi.org/10.3390/ijerph18126312
APA StyleFarrag, M. A., Amer, H. M., Bhat, R., Hamed, M. E., Aziz, I. M., Mubarak, A., Dawoud, T. M., Almalki, S. G., Alghofaili, F., Alnemare, A. K., Al-Baradi, R. S., Alosaimi, B., & Alturaiki, W. (2021). SARS-CoV-2: An Overview of Virus Genetics, Transmission, and Immunopathogenesis. International Journal of Environmental Research and Public Health, 18(12), 6312. https://doi.org/10.3390/ijerph18126312







