Maternal Iron Deficiency Programs Offspring Cognition and Its Relationship with Gastrointestinal Microbiota and Metabolites
Abstract
1. Introduction
2. Materials and Methods
2.1. Animals and Ethics
2.2. Diet and Experimental Protocols
- Four-month-old male offspring of mothers receiving the control diet (sham control (SC group)) (n = 5).
- Four-month-old male offspring of mothers receiving the low-iron diet but the control diet through the pregnancy (ICC group) (n = 5).
- Four-month-old male offspring of mothers receiving the low-iron diet during pregnancy but the control diet during lactation (IDC group) (n = 5).
- Four-month-old male offspring from mothers continuously receiving the low-iron diet (IDD group) (n = 4).
2.3. Behavioral Evaluation
2.3.1. Days 1–4: Spatial Acquisition Phase
2.3.2. Day 5: Probe Trial Phase
2.4. Tissue Dissection and Collection
2.5. Quantitative Reverse-Transcription Polymerase Chain Reaction (qRT-PCR) Analysis
2.6. Western Blot (WB) Assay
2.7. Measurement of Plasma Acetate, Butyrate, and Propionate Concentrations
2.8. Gut Microbiota Profiling
2.9. Statistical Analyses
3. Results
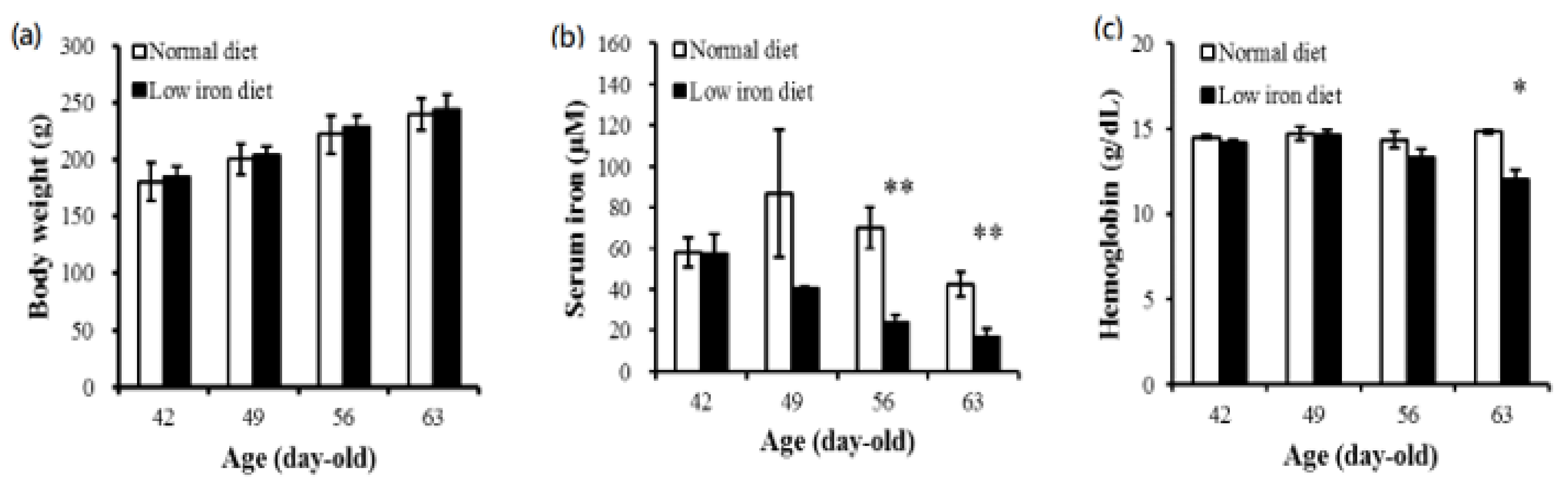
3.1. Maternal Body Weight and Serum Iron and Hemoglobin Concentrations
3.2. Adult Male Offspring Body Weights and Serum Iron and Hemoglobin Concentrations
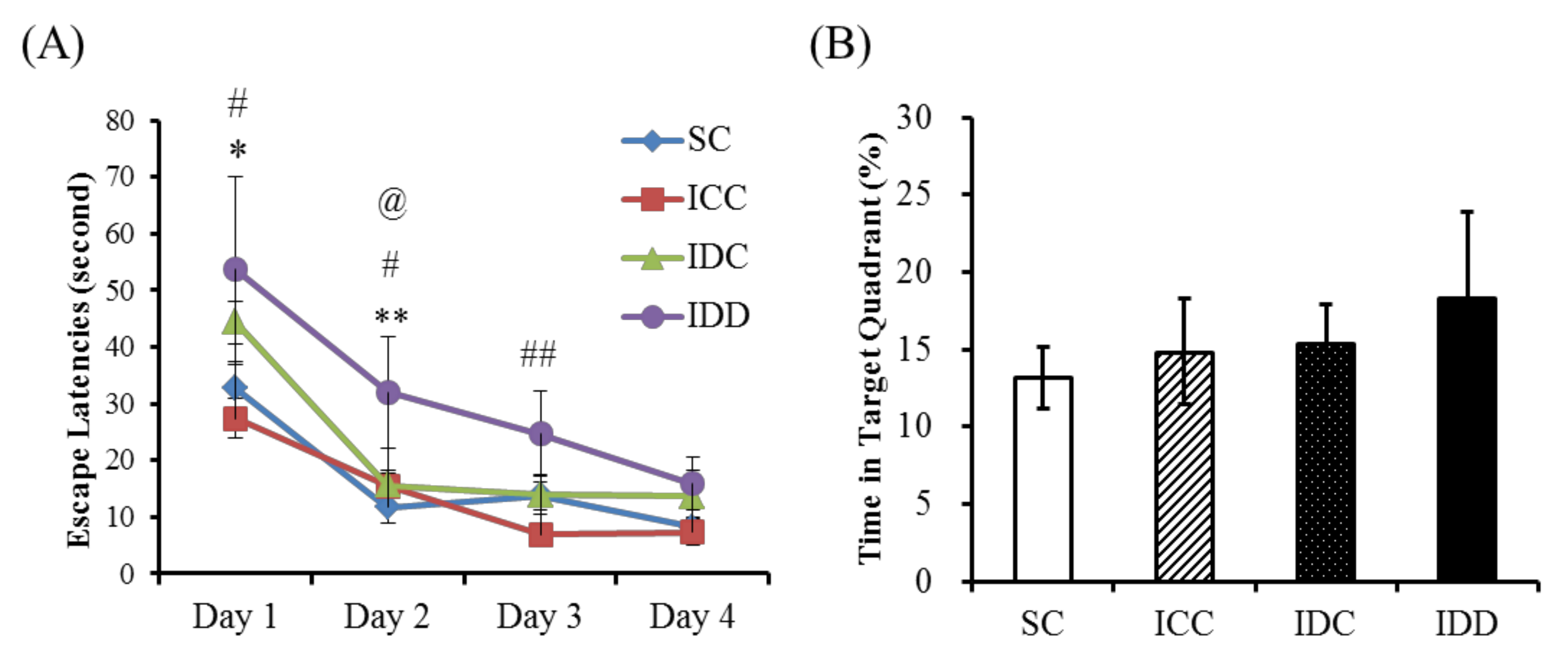
3.3. Morris Water Maze
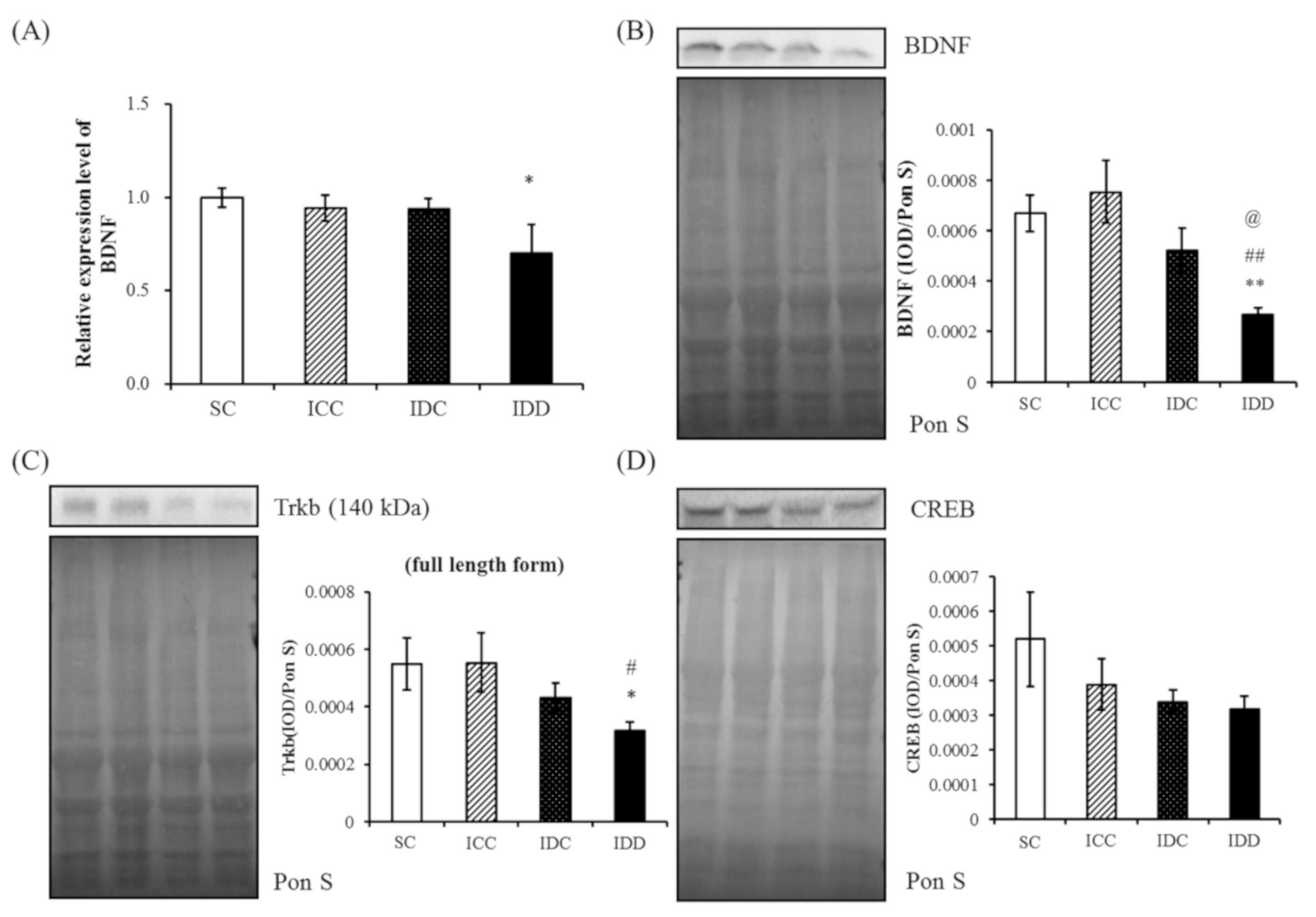
3.4. WB and qRT-PCR
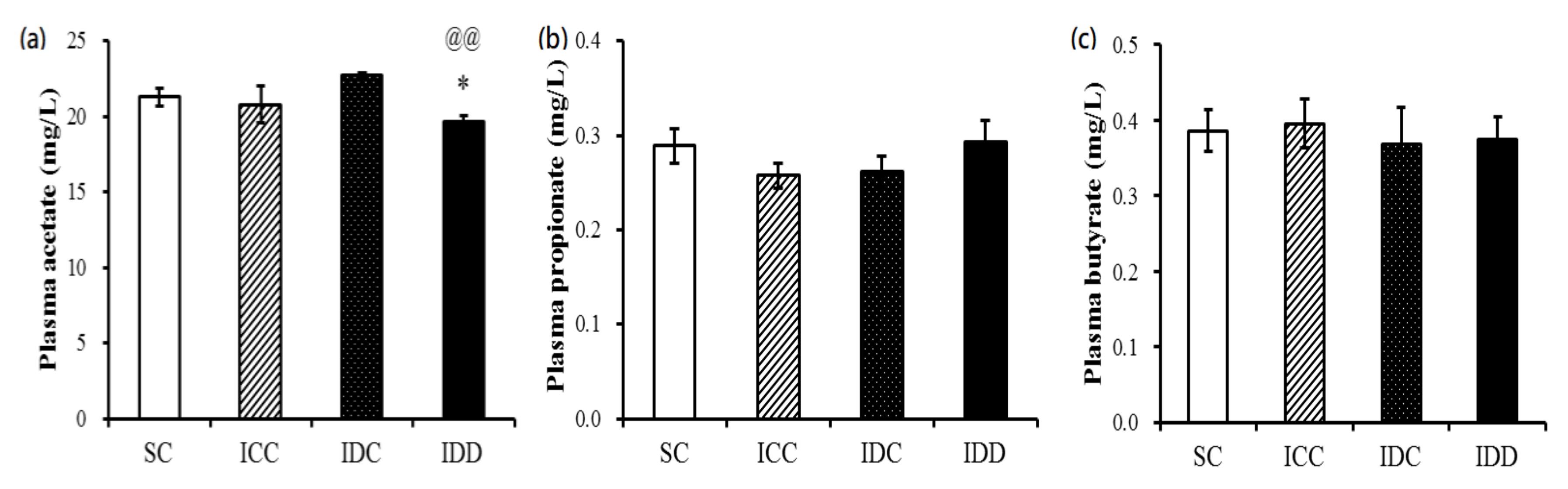
3.5. Short-Chain Fatty Acids (SCFAs)
3.6. Gut Microbiota
4. Discussion
4.1. BDNF Pathway
4.2. Neuronal and Metabolic Pathways between the Brain and Intestine
4.3. IDA and Microbiota
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Zimmermann, M.B.; Hurrell, R.F. Nutritional iron deficiency. Lancet 2007, 370, 511–520. [Google Scholar] [CrossRef]
- World Health Organization. WHO Guideline on Use of Ferritin Concentrations to Assess Iron Status in Individuals and Populations; Licence: CC BY-NC-SA 3; WHO: Geneva, Switzeland, 2020. [Google Scholar]
- Arulkumaran, S.; Regan, L.; Papageorghiou, A.; Monga, A. Oxford Desk Reference: Obstetrics and Gynaecology; OUP: Oxford, UK, 2011; pp. 180–181. [Google Scholar]
- Recommendations to prevent and control iron deficiency in the United States. Centers for Disease Control and Prevention. MMWR. Recomm. Rep. 1998, 47. Available online: http://www.ncbi.nlm.nih.gov/pubmed/9563847 (accessed on 1 July 2020).
- Gambling, L.; Danzeisen, R.; Fosset, C.; Andersen, H.S.; Dunford, S.; Srai, S.K.S.; McArdle, H.J. Iron and copper interactions in development and the effect on pregnancy outcome. J. Nutr. 2003, 133, 1554S–1556S. [Google Scholar] [CrossRef] [PubMed]
- Tamura, T.; Goldenberg, R.L.; Hou, J.; Johnston, K.E.; Cliver, S.P.; Ramey, S.L.; Nelson, K.G. Cord serum ferritin concentrations and mental and psychomotor development of children at five years of age. J. Pediatr. 2002, 140, 165–170. [Google Scholar] [CrossRef] [PubMed]
- Rao, R.; Tkáč, I.; Schmidt, A.T.; Georgieff, M.K. Fetal and neonatal iron deficiency causes volume loss and alters the neurochemical profile of the adult rat hippocampus. Nutr. Neurosci. 2011, 14, 59–65. [Google Scholar] [CrossRef] [PubMed]
- Unger, E.L.; Hurst, A.R.; Georgieff, M.K.; Schallert, T.; Rao, R.B.; Connor, J.R.; Kaciroti, N.; Lozoff, B.; Felt, B. Behavior and Monoamine Deficits in Prenatal and Perinatal Iron Deficiency Are Not Corrected by Early Postnatal Moderate-Iron or High-Iron Diets in Rats12. J. Nutr. 2012, 142, 2040–2049. [Google Scholar] [CrossRef] [PubMed]
- Mu, J.-S.; Li, W.-P.; Yao, Z.-B.; Zhou, X.F. Deprivation of endogenous brain-derived neurotrophic factor results in impairment of spatial learning and memory in adult rats. Brain Res. 1999, 835, 259–265. [Google Scholar] [CrossRef]
- Tran, P.V.; Carlson, E.S.; Fretham, S.J.B.; Georgieff, M.K. Early-Life Iron Deficiency Anemia Alters Neurotrophic Factor Expression and Hippocampal Neuron Differentiation in Male Rats1–3. J. Nutr. 2008, 138, 2495–2501. [Google Scholar] [CrossRef]
- Basu, S.; Kumar, D.; Anupurba, S.; Verma, A.; Kumar, A. Effect of maternal iron deficiency anemia on fetal neural development. J. Perinatol. 2017, 38, 233–239. [Google Scholar] [CrossRef]
- Bekinschtein, P.; Cammarota, M.; Igaz, L.M.; Bevilaqua, L.R.; Izquierdo, I.; Medina, J.H. Persistence of Long-Term Memory Storage Requires a Late Protein Synthesis- and BDNF- Dependent Phase in the Hippocampus. Neuron 2007, 53, 261–277. [Google Scholar] [CrossRef]
- Angoa-Pérez, M.; Anneken, J.H.; Kuhn, D.M. The role of brain-derived neurotrophic factor in the pathophysiology of psychiatric and neurological disorders. J. Psychiatry Psychiatr. Disord. 2017, 1, 252–269. [Google Scholar]
- Cryan, J.F.; Dinan, T.G. Mind-altering microorganisms: The impact of the gut microbiota on brain and behaviour. Nat. Rev. Neurosci. 2012, 13, 701–712. [Google Scholar] [CrossRef] [PubMed]
- Dostal, A.; Chassard, C.; Hilty, F.M.; Zimmermann, M.B.; Jaeggi, T.; Rossi, S.; Lacroix, C. Iron Depletion and Repletion with Ferrous Sulfate or Electrolytic Iron Modifies the Composition and Metabolic Activity of the Gut Microbiota in Rats. J. Nutr. 2011, 142, 271–277. [Google Scholar] [CrossRef] [PubMed]
- Woodman, A.G.; Care, A.S.; Mansour, Y.; Cherak, S.J.; Panahi, S.; Gragasin, F.S.; Bourque, S.L. Modest and Severe Maternal Iron Deficiency in Pregnancy are Associated with Fetal Anaemia and Organ-Specific Hypoxia in Rats. Sci. Rep. 2017, 7, 46573. [Google Scholar] [CrossRef] [PubMed]
- Lui, C.C.; Wang, J.-Y.; Tain, Y.-L.; Chen, Y.-C.; Chang, K.-A.; Lai, M.-C.; Huang, L.-T. Prenatal stress in rat causes long-term spatial memory deficit and hippocampus MRI abnormality: Differential effects of postweaning enriched environment. Neurochem. Int. 2011, 58, 434–441. [Google Scholar] [CrossRef] [PubMed]
- Sheen, J.-M.; Chen, Y.-C.; Hsu, M.-H.; Tain, Y.-L.; Yu, H.-R.; Huang, L.-T. Combined Intraperitoneal and Intrathecal Etanercept Reduce Increased Brain Tumor Necrosis Factor-Alpha and Asymmetric Dimethylarginine Levels and Rescues Spatial Deficits in Young Rats after Bile Duct Ligation. Front. Cell. Neurosci. 2016, 10. [Google Scholar] [CrossRef] [PubMed]
- Sheen, J.-M.; Yu, H.-R.; Tain, Y.-L.; Chen, Y.-C.; Hsu, M.-H.; Huang, L.-T. Young rats with increased circulatory asymmetric dimethylarginine exhibited spatial deficit and alterations in dorsal hippocampus brain-derived neurotrophic factor and asymmetric dimethylarginine: Effects of melatonin. Int. J. Dev. Neurosci. 2019, 78, 83–89. [Google Scholar] [CrossRef]
- Beard, J.L.; Connor, J.R. Iron status and neural functioning. Annu. Rev. Nutr. 2003, 23, 41–58. [Google Scholar] [CrossRef]
- Lozoff, B.; Georgieff, M.K. Iron Deficiency and Brain Development. Semin. Pediatr. Neurol. 2006, 13, 158–165. [Google Scholar] [CrossRef]
- Schmidt, A.T.; Waldow, K.J.; Grove, W.M.; Salinas, J.A.; Georgieff, M.K. Dissociating the long-term effects of fetal/neonatal iron deficiency on three types of learning in the rat. Behav. Neurosci. 2007, 121, 475–482. [Google Scholar] [CrossRef]
- Chang, S.; Zeng, L.; Brouwer, I.D.; Kok, F.J.; Yan, H. Effect of iron deficiency anemia in pregnancy on child mental development in rural China. Pediatrics 2013, 131, e755–e763. [Google Scholar] [CrossRef]
- Felt, B.T.; Lozoff, B. Brain iron and behavior of rats are not normalized by treatment of iron deficiency anemia during early development. J. Nutr. 1996, 126, 693–701. [Google Scholar] [CrossRef] [PubMed]
- Mihaila, C.; Schramm, J.C.; Strathmann, F.G.; Lee, D.L.; Gelein, R.M.; Luebke, A.E.; Mayer-Proschel, M. Identifying a window of vulnerability during fetal development in a maternal iron restriction model. PLoS ONE 2011, 6, e17483. [Google Scholar] [CrossRef] [PubMed]
- Ranade, S.C.; Nawaz, S.; Chakrabarti, A.; Gressens, P.; Mani, S. Spatial memory deficits in maternal iron deficiency paradigms are associated with altered glucocorticoid levels. Horm. Behav. 2013, 64, 26–36. [Google Scholar] [CrossRef] [PubMed]
- Bekinschtein, P.; Cammarota, M.; Katche, C.; Slipczuk, L.; Rossato, J.I.; Goldin, A.; Izquierdo, I.; Medina, J.H. BDNF is essential to promote persistence of long-term memory storage. Proc. Natl. Acad. Sci. USA 2008, 105, 2711–2716. [Google Scholar] [CrossRef]
- Chouthai, N.S.; Sampers, J.; Desai, N.; Smith, G.M. Changes in neurotrophin levels in umbilical cord blood from infants with different gestational ages and clinical conditions. Pediatr. Res. 2003, 53, 965–969. [Google Scholar] [CrossRef]
- Radka, S.F.; Hoist, P.A.; Fritsche, M.; Altar, C.A. Presence of brain-derived neurotrophic factor in brain and human and rat but not mouse serum detected by a sensitive and specific immunoassay. Brain Res. 1996, 709, 122–130. [Google Scholar] [CrossRef]
- Cunha, C.; Brambilla, R.; Thomas, K.L. A simple role for BDNF in learning and memory? Front. Mol. Neurosci. 2010, 3. [Google Scholar] [CrossRef]
- Estrada, J.; Contreras, I.; Pliego-Rivero, F.B.; Otero, G. Molecular mechanisms of cognitive impairment in iron deficiency: Alterations in brain-derived neurotrophic factor and Insulin-like growth factor expression and function in the central nervous system. Nutr. Neurosci. 2013, 17, 193–206. [Google Scholar] [CrossRef]
- Novotný, M.; Klimova, B.; Valis, M. Microbiome and cognitive impairment: Can any diets influence learning processes in a positive way? Front. Aging Neurosci. 2019, 11. [Google Scholar] [CrossRef]
- Fröhlich, E.E.; Farzi, A.; Mayerhofer, R.; Reichmann, F.; Jačan, A.; Wagner, B.; Zinser, E.; Bordag, N.; Magnes, C.; Fröhlich, E.; et al. Cognitive impairment by antibiotic-induced gut dysbiosis: Analysis of gut microbiota-brain communication. Brain Behav. Immun. 2016, 56, 140–155. [Google Scholar] [CrossRef]
- Macfarlane, G.T.; Macfarlane, S. Fermentation in the human large intestine. J. Clin. Gastroenterol. 2011, 45, S120–S127. [Google Scholar] [CrossRef] [PubMed]
- Ibarra, A.; Heyck, M. Microbiota and memory: A symbiotic therapy to counter cognitive decline? Brain Circ. 2019, 5, 124–129. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, Y.; Sugahara, H.; Shimada, K.; Mitsuyama, E.; Kuhara, T.; Yasuoka, A.; Kondo, T.; Abe, K.; Xiao, J.-Z. Therapeutic potential of Bifidobacterium breve strain A1 for preventing cognitive impairment in Alzheimer’s disease. Sci. Rep. 2017, 7, 13510. [Google Scholar] [CrossRef] [PubMed]
- Hsiao, E.Y.; McBride, S.W.; Hsien, S.; Sharon, G.; Hyde, E.R.; McCue, T.; Codelli, J.A.; Chow, J.; Reisman, S.E.; Petrosino, J.F.; et al. Microbiota modulate behavioral and physiological abnormalities associated with neurodevelopmental disorders. Cell 2013, 155, 1451–1463. [Google Scholar] [CrossRef] [PubMed]
- Dinan, T.G.; Cryan, J.F. Gut instincts: Microbiota as a key regulator of brain development, ageing and neurodegeneration. J. Physiol. 2016, 595, 489–503. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Covington, A.; Pamer, E.G. The intestinal microbiota: Antibiotics, colonization resistance, and enteric pathogens. Immunol. Rev. 2017, 279, 90–105. [Google Scholar] [CrossRef]
- Sjödin, K.S.; Domellöf, M.; Lagerqvist, C.; Hernell, O.; Lönnerdal, B.; Szymlek-Gay, E.A.; Sjödin, A.; E West, C.; Lind, T. Administration of ferrous sulfate drops has significant effects on the gut microbiota of iron-sufficient infants: A randomised controlled study. Gut 2018, 68, 2095.1–2097. [Google Scholar] [CrossRef]
- Muleviciene, A.; D’Amico, F.; Turroni, S.; Candela, M.; Jankauskiene, A. Iron deficiency anemia-related gut microbiota dysbiosis in infants and young children: A pilot study. Acta Microbiol. Et Immunol. Hung. 2018, 65, 551–564. [Google Scholar] [CrossRef]
- Saji, N.; Murotani, K.; Hisada, T.; Tsuduki, T.; Sugimoto, T.; Kimura, A.; Niida, S.; Toba, K.; Sakurai, T. The relationship between the gut microbiome and mild cognitive impairment in patients without dementia: A cross-sectional study conducted in Japan. Sci. Rep. 2019, 9, 1–10. [Google Scholar] [CrossRef]
- Treangen, T.J.; Wagner, J.; Burns, M.P.; Villapol, S. Traumatic brain injury in mice induces acute bacterial dysbiosis within the fecal microbiome. Front. Immunol. 2018, 9. [Google Scholar] [CrossRef]
- Arciniegas, D.B.; Held, K.; Wagner, P. Cognitive impairment following traumatic brain injury. Curr. Treat. Options Neurol. 2002, 4, 43–57. [Google Scholar] [CrossRef] [PubMed]


© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hsieh, H.-Y.; Chen, Y.-C.; Hsu, M.-H.; Yu, H.-R.; Su, C.-H.; Tain, Y.-L.; Huang, L.-T.; Sheen, J.-M. Maternal Iron Deficiency Programs Offspring Cognition and Its Relationship with Gastrointestinal Microbiota and Metabolites. Int. J. Environ. Res. Public Health 2020, 17, 6070. https://doi.org/10.3390/ijerph17176070
Hsieh H-Y, Chen Y-C, Hsu M-H, Yu H-R, Su C-H, Tain Y-L, Huang L-T, Sheen J-M. Maternal Iron Deficiency Programs Offspring Cognition and Its Relationship with Gastrointestinal Microbiota and Metabolites. International Journal of Environmental Research and Public Health. 2020; 17(17):6070. https://doi.org/10.3390/ijerph17176070
Chicago/Turabian StyleHsieh, Hsin-Yi, Yu-Chieh Chen, Mei-Hsin Hsu, Hong-Ren Yu, Chung-Hao Su, You-Lin Tain, Li-Tung Huang, and Jiunn-Ming Sheen. 2020. "Maternal Iron Deficiency Programs Offspring Cognition and Its Relationship with Gastrointestinal Microbiota and Metabolites" International Journal of Environmental Research and Public Health 17, no. 17: 6070. https://doi.org/10.3390/ijerph17176070
APA StyleHsieh, H.-Y., Chen, Y.-C., Hsu, M.-H., Yu, H.-R., Su, C.-H., Tain, Y.-L., Huang, L.-T., & Sheen, J.-M. (2020). Maternal Iron Deficiency Programs Offspring Cognition and Its Relationship with Gastrointestinal Microbiota and Metabolites. International Journal of Environmental Research and Public Health, 17(17), 6070. https://doi.org/10.3390/ijerph17176070







