Characterization of Chromosome-Mediated BlaOXA-894 in Shewanella xiamenensis Isolated from Pig Wastewater
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Isolate
2.2. Antibiotic Susceptibility Testing
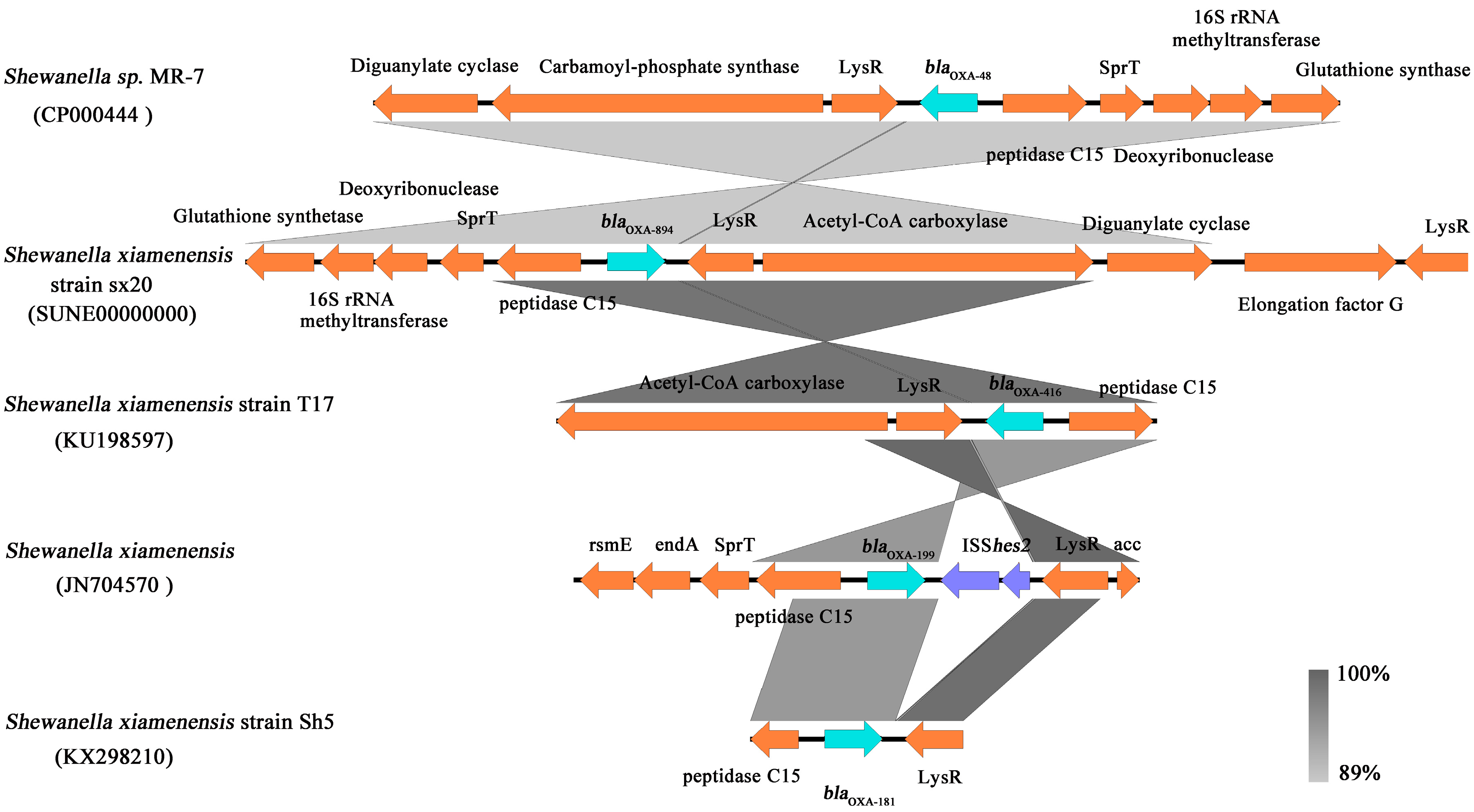
2.3. Whole-Genome Sequencing and Analysis
2.4. The Conjugation Assay
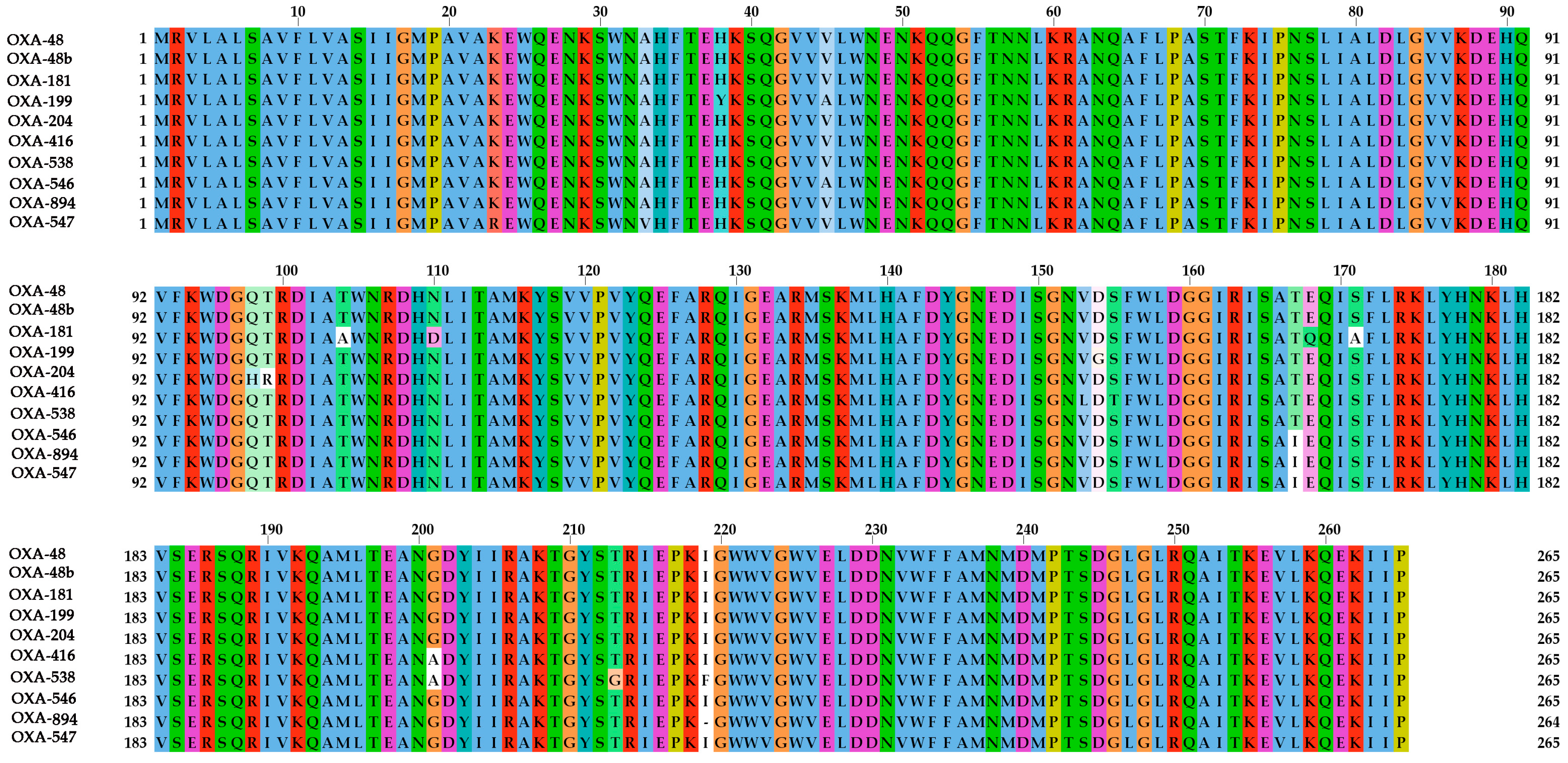
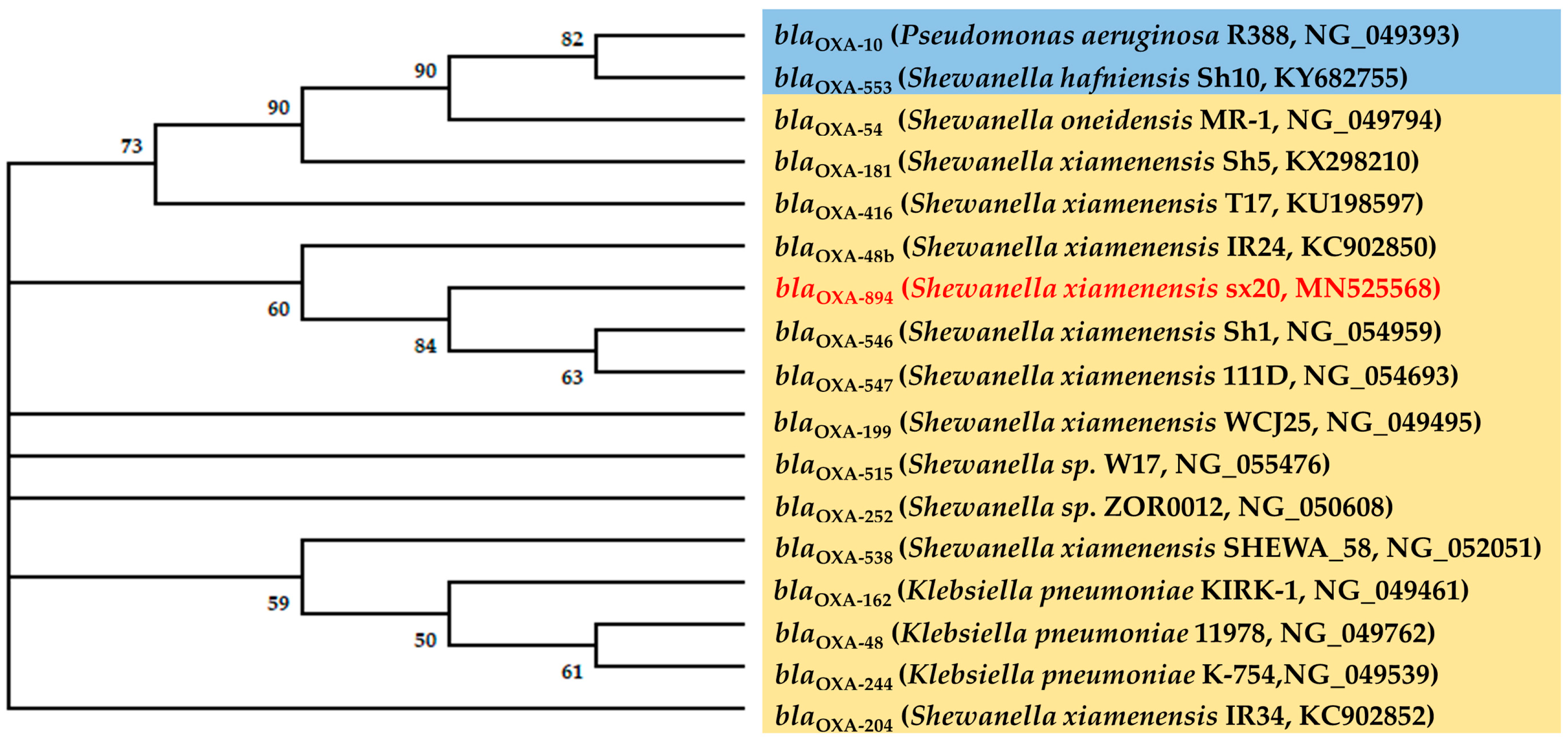
2.5. Phylogenetic Analysis of the BlaOXA-48-Like Genes
3. Results and Discussion
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Poirel, L.; Heritier, C.; Tolun, V.; Nordmann, P. Emergence of oxacillinase-mediated resistance to imipenem in Klebsiella pneumoniae. Antimicrob. Agents Chemother. 2004, 48, 15–22. [Google Scholar] [CrossRef] [PubMed]
- Poirel, L.; Potron, A.; Nordmann, P. OXA-48-like carbapenemases: The phantom menace. J. Antimicrob. Chemother. 2012, 67, 1597–1606. [Google Scholar] [CrossRef] [PubMed]
- Mairi, A.; Pantel, A.; Sotto, A.; Lavigne, J.P.; Touati, A. OXA-48-like carbapenemases producing Enterobacteriaceae in different niches. Eur. J. Clin. Microbiol. Infect. Dis. 2018, 37, 587–604. [Google Scholar] [CrossRef] [PubMed]
- Kasap, M.; Torol, S.; Kolayli, F.; Dundar, D.; Vahaboglu, H. OXA-162, a novel variant of OXA-48 displays extended hydrolytic activity towards imipenem, meropenem and doripenem. J. Enzym. Inhib. Med. Chem. 2013, 28, 990–996. [Google Scholar] [CrossRef] [PubMed]
- Potron, A.; Nordmann, P.; Lafeuille, E.; Al Maskari, Z.; Al Rashdi, F.; Poirel, L. Characterization of OXA-181, a carbapenem-hydrolyzing class D beta-lactamase from Klebsiella pneumoniae. Antimicrob. Agents Chemother. 2011, 55, 4896–4899. [Google Scholar] [CrossRef] [PubMed]
- Potron, A.; Nordmann, P.; Poirel, L. Characterization of OXA-204, a carbapenem-hydrolyzing class D beta-lactamase from Klebsiella pneumoniae. Antimicrob. Agents Chemother. 2013, 57, 633–636. [Google Scholar] [CrossRef]
- Potron, A.; Rondinaud, E.; Poirel, L.; Belmonte, O.; Boyer, S.; Camiade, S.; Nordmann, P. Genetic and biochemical characterisation of OXA-232, a carbapenem-hydrolysing class D beta-lactamase from Enterobacteriaceae. Int. J. Antimicrob. Agents 2013, 41, 325–329. [Google Scholar] [CrossRef]
- Guo, L.; An, J.; Ma, Y.; Ye, L.; Luo, Y.; Tao, C.; Yang, J. Nosocomial Outbreak of OXA-48-Producing Klebsiella pneumoniae in a Chinese Hospital: Clonal Transmission of ST147 and ST383. PLoS ONE 2016, 11, e0160754. [Google Scholar] [CrossRef]
- Huang, J.; Sun, B.; Zhang, X. Shewanella xiamenensis sp. nov., isolated from coastal sea sediment. Int. J. Syst. Evol. Microbiol. 2010, 60, 1585–1589. [Google Scholar] [CrossRef]
- Huang, Y.; Cheng, J.; Wu, Z. Genomic and phylogenetic characterization of Shewanella xiamenensis isolated from giant grouper (Epinephelus lanceolatus) in Taiwan. Zoonoses Public Health 2019, 66. [Google Scholar] [CrossRef]
- Zong, Z. Nosocomial peripancreatic infection associated with Shewanella xiamenensis. J. Med. Microbiol. 2011, 60, 1387–1390. [Google Scholar] [CrossRef] [PubMed]
- Tacao, M.; Correia, A.; Henriques, I. Environmental Shewanella xiamenensis strains that carry blaOXA-48 or blaOXA-204 genes: Additional proof for blaOXA-48-like gene origin. Antimicrob. Agents Chemother. 2013, 57, 6399–6400. [Google Scholar] [CrossRef] [PubMed]
- Potron, A.; Poirel, L.; Nordmann, P. Origin of OXA-181, an emerging carbapenem-hydrolyzing oxacillinase, as a chromosomal gene in Shewanella xiamenensis. Antimicrob. Agents Chemother. 2011, 55, 4405–4407. [Google Scholar] [CrossRef] [PubMed]
- Tacao, M.; Araujo, S.; Vendas, M.; Alves, A.; Henriques, I. Shewanella species as the origin of blaOXA-48 genes: Insights into gene diversity, associated phenotypes and possible transfer mechanisms. Int. J. Antimicrob. Agents 2018, 51, 340–348. [Google Scholar] [CrossRef] [PubMed]
- Zong, Z. Discovery of blaOXA-199, a chromosome-based blaOXA-48-like variant, in Shewanella xiamenensis. PLoS ONE 2012, 7, e48280. [Google Scholar] [CrossRef]
- Tafoukt, R.; Leangapichart, T.; Hadjadj, L.; Bakour, S.; Diene, S.M.; Rolain, J.M.; Touati, A. Characterisation of blaOXA-538, a new variant of blaOXA-48, in Shewanella xiamenensis isolated from river water in Algeria. J. Glob. Antimicrob. Resist. 2018, 13, 70–73. [Google Scholar] [CrossRef] [PubMed]
- Poirel, L.; Walsh, T.R.; Cuvillier, V.; Nordmann, P. Multiplex PCR for detection of acquired carbapenemase genes. Diagn. Microbiol. Infect. Dis. 2011, 70, 119–123. [Google Scholar] [CrossRef]
- Pasteran, F.; Tijet, N.; Melano, R.G. Simplified Protocol for Carba NP Test for Enhanced Detection of Carbapenemase Producers Directly from Bacterial Cultures. J. Clin. Microbiol. 2015, 53, 3908–3911. [Google Scholar] [CrossRef]
- Waterhouse, A.; Procter, J.; Martin, D. Jalview Version 2-a multiple sequence alignment editor and analysis workbench. Bioinformatics 2009, 25, 1189–1191. [Google Scholar] [CrossRef]
- Skalova, A.; Chudejova, K.; Rotova, V. Molecular Characterization of OXA-48-Like-Producing Enterobacteriaceae in the Czech Republic and Evidence for Horizontal Transfer of pOXA-48-Like Plasmids. Antimicrob. Agents Chemother. 2017, 61. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef] [PubMed]
- Antonelli, A.; Di Palo, D.M.; Galano, A.; Becciani, S.; Montagnani, C.; Pecile, P.; Galli, L.; Rossolini, G.M. Intestinal carriage of Shewanella xiamenensis simulating carriage of OXA-48-producing Enterobacteriaceae. Diagn. Microbiol. Infect. Dis. 2015, 82, 1–3. [Google Scholar] [CrossRef] [PubMed]
- Yousfi, K.; Touati, A.; Lefebvre, B.; Fournier, E.; Cote, J.C.; Soualhine, H.; Walker, M.; Bougdour, D.; Tremblay, C.; Bekal, S. A Novel Plasmid, pSx1, Harboring a New Tn1696 Derivative from Extensively Drug-Resistant Shewanella xiamenensis Encoding OXA-416. Microb. Drug Resist. 2017, 23, 429–436. [Google Scholar] [CrossRef] [PubMed]
- Palaniyandi, S.; Mitra, A.; Herren, C.D.; Zhu, X.; Mukhopadhyay, S. LuxS contributes to virulence in avian pathogenic Escherichia coli O78:K80:H9. Vet. Microbiol. 2013, 166, 567–575. [Google Scholar] [CrossRef] [PubMed]

| Species | Year | blaOXA-48-Like Gene | Imipenem | Meropenem | Ertapenem | Reference |
|---|---|---|---|---|---|---|
| S. xiamenensis IR33 | 2013 | blaOXA-48 | 4 | 1 | 8 | [12] |
| S. xiamenensis Sh31 | 2018 | blaOXA-48 | 8 | 2 | 8 | [14] |
| S. xiamenensis IR24 | 2013 | blaOXA-48-like | 4 | 2 | 8 | [12] |
| S. xiamenensis S12 | 2011 | blaOXA-181 | 0.75 | 0.25 | 2 | [13] |
| S. xiamenensis Sh5 | 2018 | blaOXA-181 | 4 | 5 | 4 | [14] |
| S. xiamenensis AS69 | 2017 | blaOXA-181 | 0.75 | - | - | [16] |
| S. xiamenensis AS85 | 2017 | blaOXA-181 | 0.75 | - | - | [16] |
| S. xiamenensis AS100 | 2017 | blaOXA-199 | 0.5 | - | - | [16] |
| S. xiamenensis IR34 | 2013 | blaOXA-204 | >32 | 8 | >32 | [12] |
| S. xiamenensis Sh33 | 2018 | blaOXA-204 | >32 | 8 | >32 | [14] |
| S. xiamenensis ZYW1 | 2019 | blaOXA-416 | 1 | - | - | [10] |
| S. xiamenensis DDP1 | 2013 | blaOXA-416 | 32 | 16 | >32 | [22] |
| S. xiamenensis T17 | 2017 | blaOXA-416 | 1 | 1 | 4 | [23] |
| S. xiamenensis AS58 | 2017 | blaOXA-538 | 3 | - | - | [16] |
| S. xiamenensis Sh1 | 2018 | blaOXA-546 | 1 | 0.5 | 3 | [14] |
| S. xiamenensis sx20 | 2019 | blaOXA-894 | 32 | 8 | 8 | This study |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zou, H.; Zhou, Z.; Xia, H.; Zhao, Q.; Li, X. Characterization of Chromosome-Mediated BlaOXA-894 in Shewanella xiamenensis Isolated from Pig Wastewater. Int. J. Environ. Res. Public Health 2019, 16, 3768. https://doi.org/10.3390/ijerph16193768
Zou H, Zhou Z, Xia H, Zhao Q, Li X. Characterization of Chromosome-Mediated BlaOXA-894 in Shewanella xiamenensis Isolated from Pig Wastewater. International Journal of Environmental Research and Public Health. 2019; 16(19):3768. https://doi.org/10.3390/ijerph16193768
Chicago/Turabian StyleZou, Huiyun, Ziyu Zhou, Huiyu Xia, Qian Zhao, and Xuewen Li. 2019. "Characterization of Chromosome-Mediated BlaOXA-894 in Shewanella xiamenensis Isolated from Pig Wastewater" International Journal of Environmental Research and Public Health 16, no. 19: 3768. https://doi.org/10.3390/ijerph16193768
APA StyleZou, H., Zhou, Z., Xia, H., Zhao, Q., & Li, X. (2019). Characterization of Chromosome-Mediated BlaOXA-894 in Shewanella xiamenensis Isolated from Pig Wastewater. International Journal of Environmental Research and Public Health, 16(19), 3768. https://doi.org/10.3390/ijerph16193768




