Transcriptomic Profiles in Zebrafish Liver Permit the Discrimination of Surface Water with Pollution Gradient and Different Discharges
Abstract
1. Introduction
2. Methods and Materials
2.1. Site Selection and Water Sampling
2.2. Fish Exposures
2.3. Microarray Analysis
2.4. Pathway Analysis
2.5. Clustering Analysis
2.6. Statistical Analysis
3. Results
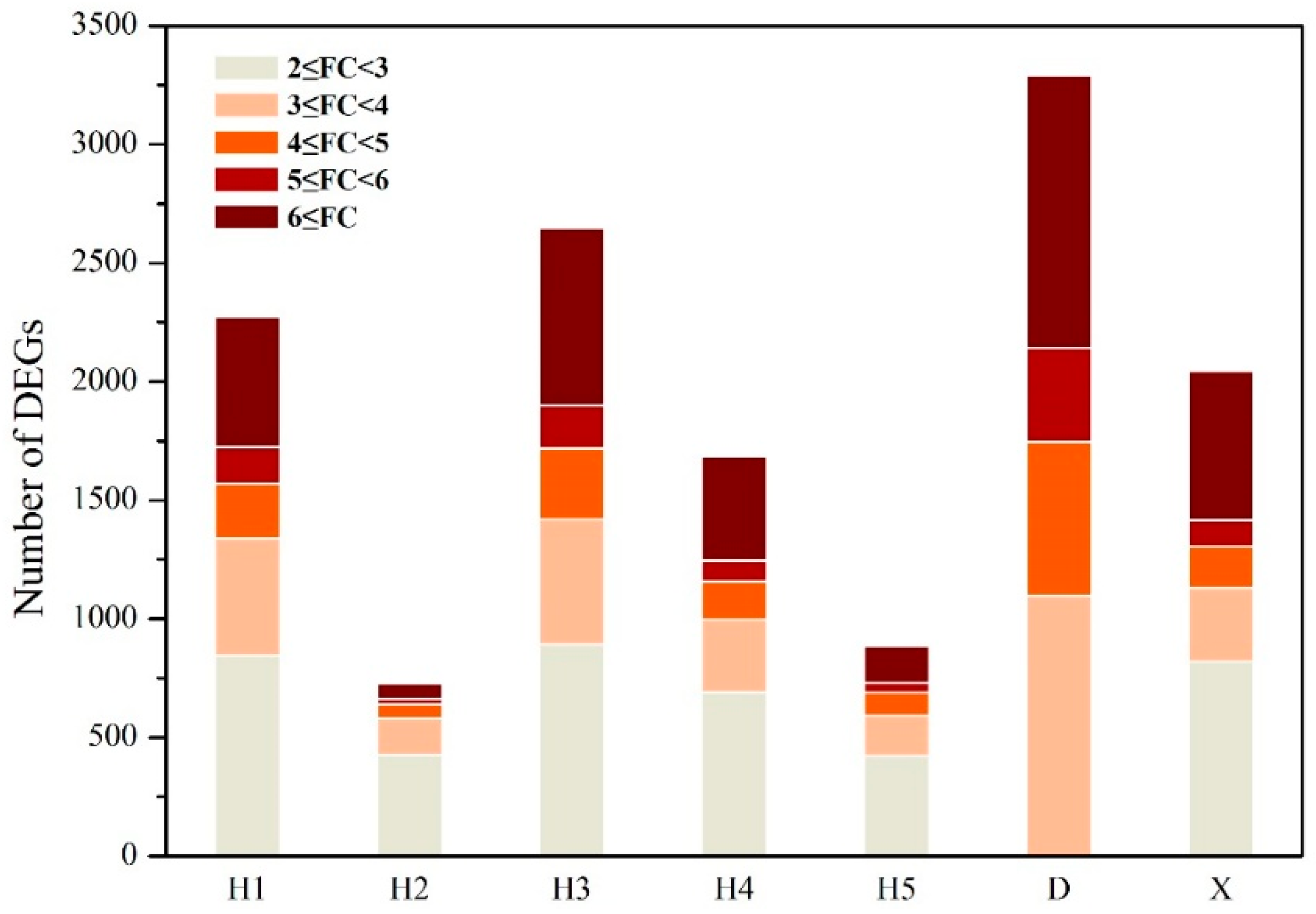
3.1. Differentially Expressed Genes Affected by River Water
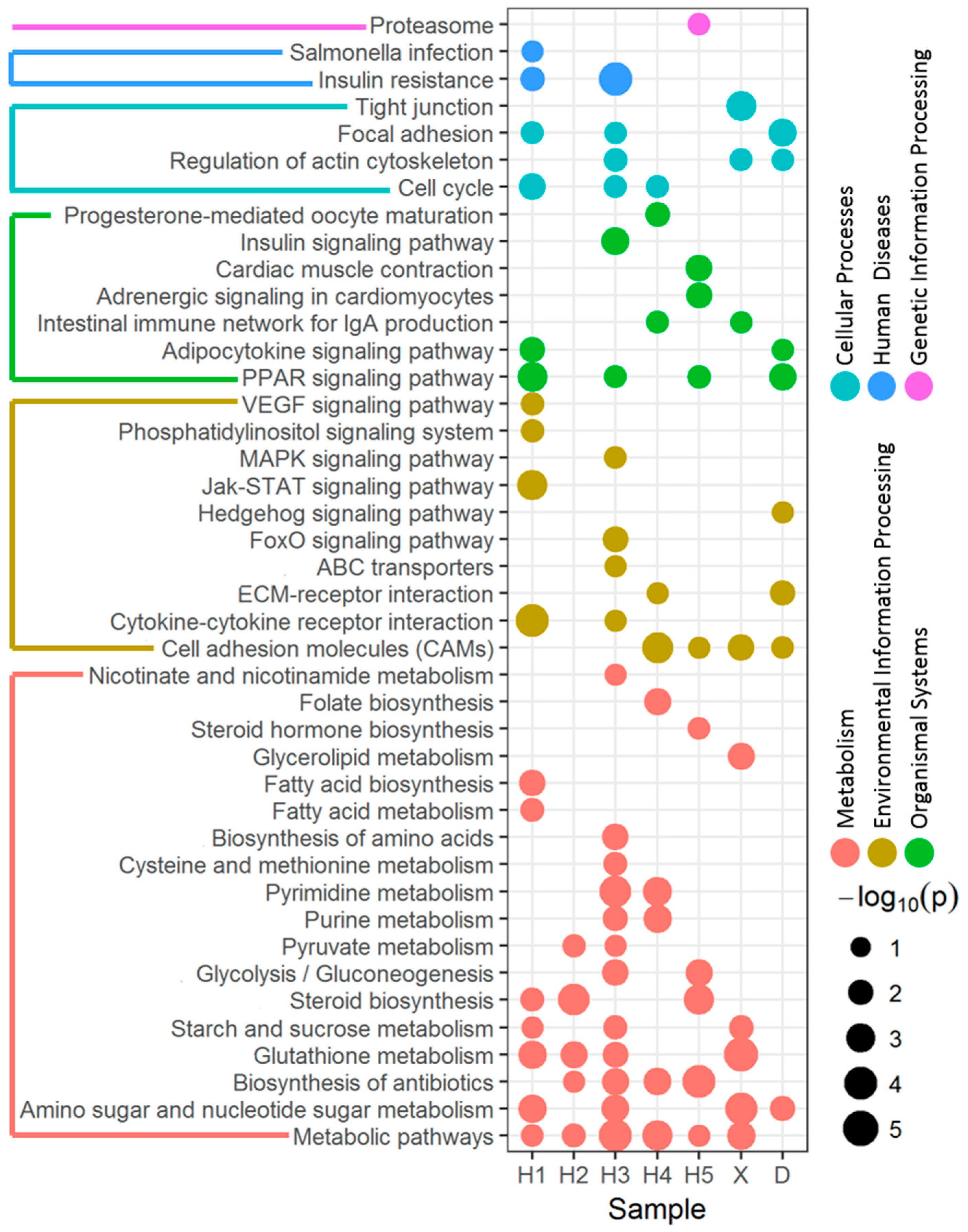
3.2. Enriched Pathway of DEGs Affected by River Water
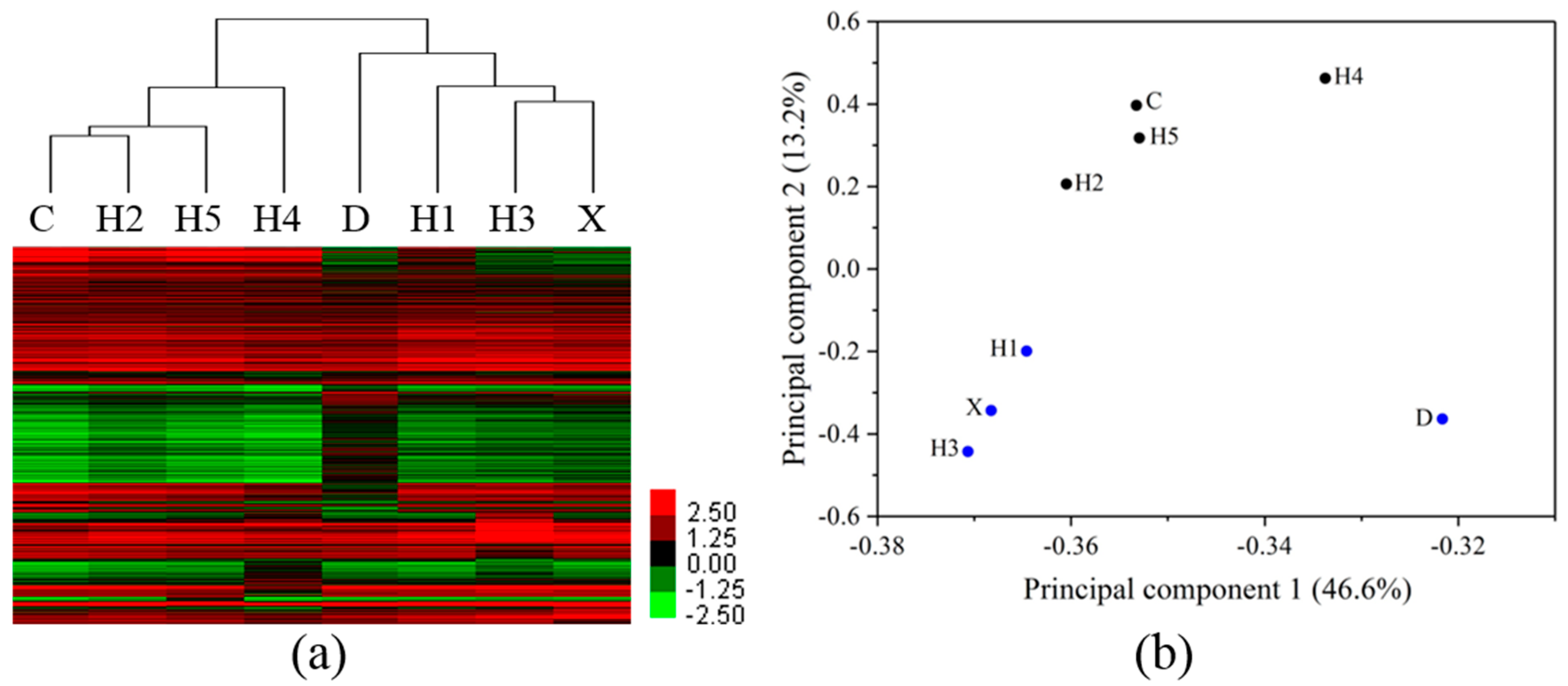
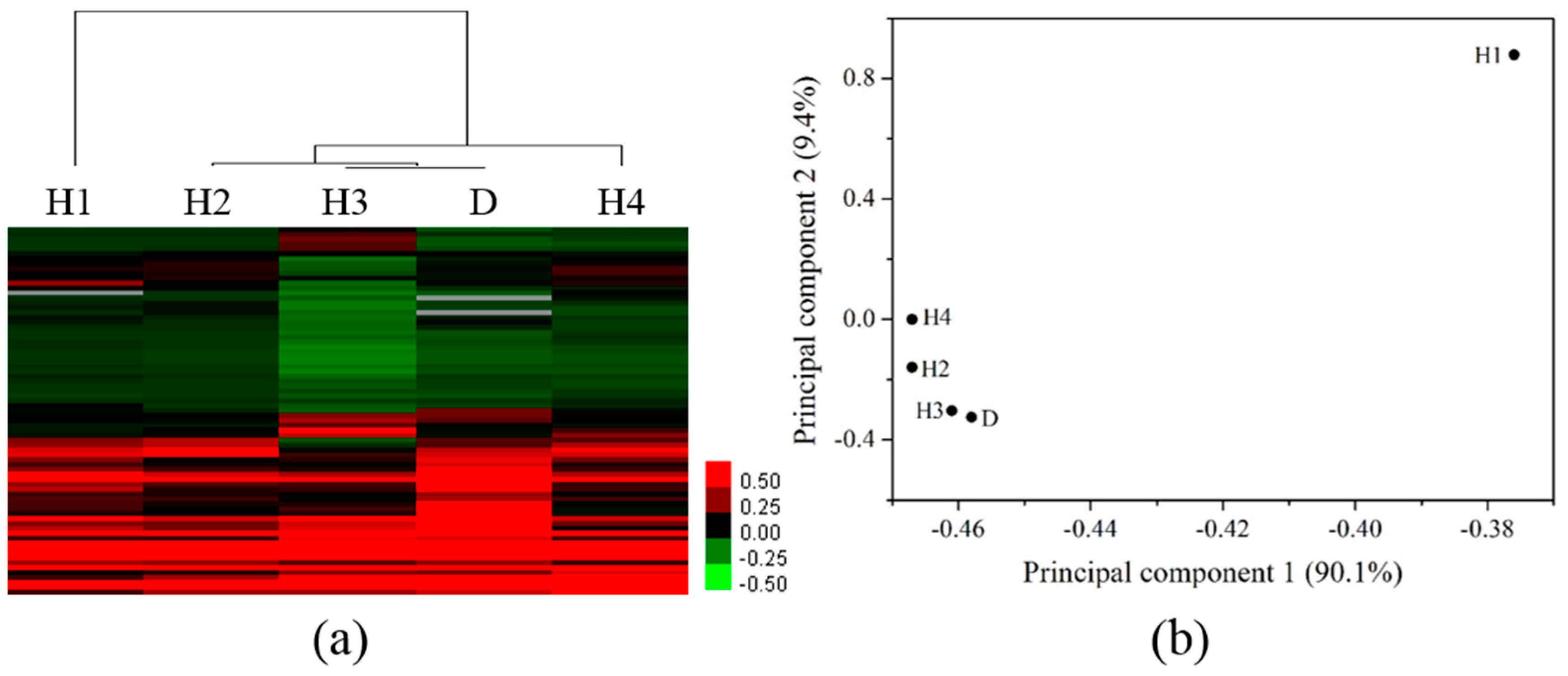
3.3. Differentiation of Gene Profiles for Pollution Gradients and Discharge Source
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Cwiertny, D.M.; Snyder, S.A.; Schlenk, D.; Kolodziej, E.P. Environmental designer drugs: When transformation may not eliminate risk. Environ. Sci. Technol. 2014, 48, 11737–11745. [Google Scholar] [CrossRef] [PubMed]
- Fenner, K.; Canonica, S.; Wackett, L.P.; Elsner, M. Evaluating pesticide degradation in the environment: Blind spots and emerging opportunities. Science 2013, 341, 752–758. [Google Scholar] [CrossRef] [PubMed]
- Qu, S.; Kolodziej, E.P.; Long, S.A.; Gloer, J.B.; Patterson, E.V.; Baltrusaitis, J.; Jones, G.D.; Benchetler, P.V.; Cole, E.A.; Kimbrough, K.C.; et al. Product-to-parent reversion of trenbolone: Unrecognized risks for endocrine disruption. Science 2013, 342, 347–351. [Google Scholar] [CrossRef] [PubMed]
- Neale, P.A.; Ait-Aissa, S.; Brack, W.; Creusot, N.; Denison, M.S.; Deutschmann, B.; Hilscherova, K.; Hollert, H.; Krauss, M.; Novak, J.; et al. Linking in vitro effects and detected organic micropollutants in surface water using mixture-toxicity modeling. Environ. Sci. Technol. 2015, 49, 14614–14624. [Google Scholar] [CrossRef] [PubMed]
- North, M.; Vulpe, C.D. Functional toxicogenomics: Mechanism-centered toxicology. Int. J. Mol. Sci. 2010, 11, 4796–4813. [Google Scholar] [CrossRef] [PubMed]
- Rouquie, D.; Heneweer, M.; Botham, J.; Ketelslegers, H.; Markell, L.; Pfister, T.; Steiling, W.; Strauss, V.; Hennes, C. Contribution of new technologies to characterization and prediction of adverse effects. Crit. Rev. Toxicol. 2015, 45, 172–183. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Kemadjou, J.R.; Zinsmeister, C.; Bauer, M.; Legradi, J.; Muller, F.; Pankratz, M.; Jakel, J.; Strahle, U. Transcriptional profiling reveals barcode-like toxicogenomic responses in the zebrafish embryo. Genome Biol. 2007, 8, R227. [Google Scholar] [CrossRef] [PubMed]
- Vidal-Dorsch, D.E.; Colli-Dula, R.C.; Bay, S.M.; Greenstein, D.J.; Wiborg, L.; Petschauer, D.; Denslow, N.D. Gene expression of fathead minnows (pimephales promelas) exposed to two types of treated municipal wastewater effluents. Environ. Sci. Technol. 2013, 47, 11268–11277. [Google Scholar] [CrossRef] [PubMed]
- Ings, J.S.; Servos, M.R.; Vijayan, M.M. Hepatic transcriptomics and protein expression in rainbow trout exposed to municipal wastewater effluent. Environ. Sci. Technol. 2011, 45, 2368–2376. [Google Scholar] [CrossRef] [PubMed]
- Perkins, E.J.; Habib, T.; Escalon, B.L.; Cavallin, J.E.; Thomas, L.; Weberg, M.; Hughes, M.N.; Jensen, K.M.; Kahl, M.D.; Villeneuve, D.L.; et al. Prioritization of contaminants of emerging concern in wastewater treatment plant discharges using chemical: Gene interactions in caged fish. Environ. Sci. Technol. 2017, 51, 8701–8712. [Google Scholar] [CrossRef] [PubMed]
- Costigan, S.L.; Werner, J.; Ouellet, J.D.; Hill, L.G.; Law, R.D. Expression profiling and gene ontology analysis in fathead minnow (pimephales promelas) liver following exposure to pulp and paper mill effluents. Aquat. Toxicol. 2012, 122–123, 44–55. [Google Scholar] [CrossRef] [PubMed]
- Martinovic-Weigelt, D.; Mehinto, A.C.; Ankley, G.T.; Denslow, N.D.; Barber, L.B.; Lee, K.E.; King, R.J.; Schoenfuss, H.L.; Schroeder, A.L.; Villeneuve, D.L. Transcriptomic effects-based monitoring for endocrine active chemicals: Assessing relative contribution of treated wastewater to downstream pollution. Environ. Sci. Technol. 2014, 48, 2385–2394. [Google Scholar] [CrossRef] [PubMed]
- Berninger, J.P.; Martinovic-Weigelt, D.; Garcia-Reyero, N.; Escalon, L.; Perkins, E.J.; Ankley, G.T.; Villeneuve, D.L. Using transcriptomic tools to evaluate biological effects across effluent gradients at a diverse set of study sites in minnesota, USA. Environ. Sci. Technol. 2014, 48, 2404–2412. [Google Scholar] [CrossRef] [PubMed]
- Sellin Jeffries, M.K.; Mehinto, A.C.; Carter, B.J.; Denslow, N.D.; Kolok, A.S. Taking microarrays to the field: Differential hepatic gene expression of caged fathead minnows from nebraska watersheds. Environ. Sci. Technol. 2012, 46, 1877–1885. [Google Scholar] [CrossRef] [PubMed]
- Falciani, F.; Diab, A.M.; Sabine, V.; Williams, T.D.; Ortega, F.; George, S.G.; Chipman, J.K. Hepatic transcriptomic profiles of european flounder (Platichthys flesus) from field sites and computational approaches to predict site from stress gene responses following exposure to model toxicants. Aquat. Toxicol. 2008, 90, 92–101. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.; He, M.; Liu, X.; Guo, W.; Liu, S. Contamination and ecological risk assessment of toxic trace elements in the Xi river, an urban river of Shenyang city, China. Environ. Monit. Assess. 2013, 185, 4321–4332. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.; He, M.; Liu, X.; Guo, W.; Liu, S. Distribution and contamination assessment of toxic trace elements in sediment of the daliao river system, China. Environ. Earth Sci. 2013, 70, 3163–3173. [Google Scholar] [CrossRef]
- Li, B.; Liu, R.; Gao, H.; Tan, R.; Zeng, P.; Song, Y. Spatial distribution and ecological risk assessment of phthalic acid esters and phenols in surface sediment from urban rivers in northeast China. Environ. Pollut. 2016, 219, 409–415. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; He, L.; Lu, Y.; Su, J.; Song, H.; Zeng, X.; Yu, Z. Distribution, source, and ecological risk assessment of polycyclic aromatic hydrocarbons (PAHs) in surface sediments from the hun river, northeast China. Environ. Monit. Assess. 2015, 187, 290. [Google Scholar] [CrossRef] [PubMed]
- Sun, H.; Li, F.; Zhang, T.; Zhang, X.; He, N.; Song, Q.; Zhao, L.; Sun, L.; Sun, T. Perfluorinated compounds in surface waters and wwtps in shenyang, china: Mass flows and source analysis. Water Res. 2011, 45, 4483–4490. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Hu, X.; Liu, R.; Zeng, P.; Song, Y. Occurrence and distribution of phthalic acid esters and phenols in hun river watersheds. Environ. Earth Sci. 2015, 73, 5095–5106. [Google Scholar] [CrossRef]
- Dharmadhikari, D.M.; Vanerkar, A.P.; Barhate, N.M. Chemical oxygen demand using closed microwave digestion system. Environ. Sci. Technol. 2005, 39, 6198–6201. [Google Scholar] [CrossRef] [PubMed]
- State Environmental Protection Administration. Standard Methods for the Examination of Water and Wastewater, 4th ed.; China Environmental Science Publish Press: Beijing, China, 2002.
- Porreca, I.; D’Angelo, F.; Gentilcore, D.; Carchia, E.; Amoresano, A.; Affuso, A.; Ceccarelli, M.; De Luca, P.; Esposito, L.; Guadagno, F.M.; et al. Cross-species toxicogenomic analyses and phenotypic anchoring in response to groundwater low-level pollution. BMC Genom. 2014, 15, 1067. [Google Scholar] [CrossRef] [PubMed]
- Pan, Y.X.; Luo, Z.; Zhuo, M.Q.; Wei, C.C.; Chen, G.H.; Song, Y.F. Oxidative stress and mitochondrial dysfunction mediated cd-induced hepatic lipid accumulation in zebrafish danio rerio. Aquat. Toxicol. 2018, 199, 12–20. [Google Scholar] [CrossRef] [PubMed]
- Huang da, W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using david bioinformatics resources. Nat. Protoc. 2009, 4, 44–57. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Goto, S.; Sato, Y.; Furumichi, M.; Tanabe, M. Kegg for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res. 2012, 40, D109–D114. [Google Scholar] [CrossRef] [PubMed]
- De Hoon, M.J.L.; Imoto, S.; Nolan, J.; Miyano, S. Open source clustering software. Bioinformatics 2004, 20, 1453–1454. [Google Scholar] [CrossRef] [PubMed]
- Lam, S.H.; Winata, C.L.; Tong, Y.; Korzh, S.; Lim, W.S.; Korzh, V.; Spitsbergen, J.; Mathavan, S.; Miller, L.D.; Liu, E.T.; et al. Transcriptome kinetics of arsenic-induced adaptive response in Zebrafish liver. Physiol. Genom. 2006, 27, 351–361. [Google Scholar] [CrossRef] [PubMed]
- Ung, C.Y.; Lam, S.H.; Zhang, X.; Li, H.; Zhang, L.; Li, B.; Gong, Z. Inverted expression profiles of sex-biased genes in response to toxicant perturbations and diseases. PLoS ONE 2013, 8. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2017. [Google Scholar]
- Sanderson, J.T. The steroid hormone biosynthesis pathway as a target for endocrine-disrupting chemicals. Toxicol. Sci. 2006, 94, 3–21. [Google Scholar] [CrossRef] [PubMed]
- Hasenbein, M.; Werner, I.; Deanovic, L.A.; Geist, J.; Fritsch, E.B.; Javidmehr, A.; Foe, C.; Fangue, N.A.; Connon, R.E. Transcriptomic profiling permits the identification of pollutant sources and effects in ambient water samples. Sci. Total Environ. 2014, 468–469, 688–698. [Google Scholar] [CrossRef] [PubMed]
- Hara-Yamamura, H.; Nakashima, K.; Hoque, A.; Miyoshi, T.; Kimura, K.; Watanabe, Y.; Okabe, S. Evaluation of whole wastewater effluent impacts on HepG2 using DNA microarray-based transcriptome analysis. Environ. Sci. Technol. 2013, 47, 5425–5432. [Google Scholar] [CrossRef] [PubMed]
- Uren-Webster, T.M.; Lewis, C.; Filby, A.L.; Paull, G.C.; Santos, E.M. Mechanisms of toxicity of di(2-ethylhexyl) phthalate on the reproductive health of male zebrafish. Aquat. Toxicol. 2010, 99, 360–369. [Google Scholar] [CrossRef] [PubMed]
- Zheng, B.; Liu, R.; Liu, Y.; Jin, F.; An, L. Phenolic endocrine-disrupting chemicals and intersex in wild crucian carp from Hun River, China. Chemosphere 2015, 120, 743–749. [Google Scholar] [CrossRef] [PubMed]
- Song, S.; An, L.; Zheng, B.; Zhao, Y.; Li, Z.; Chen, H.; Zhao, X.; Liu, D. Vitellogenin mrna expression in wild crucian carp from Hun River. Asian J. Ecotoxicol. 2013, 8, 121–129. [Google Scholar]
- Jin, S.Q.; Antinore, M.J.; Lung, F.T.; Dong, X.; Zhao, H.C.; Fan, F.Y.; Colchagie, A.B.; Blanck, P.; Roller, P.P.; Fornace, A.J.; et al. The GADD45 inhibition of Cdc2 kinase correlates with GADD45-mediated growth suppression. J. Biol. Chem. 2000, 275, 16602–16608. [Google Scholar] [CrossRef] [PubMed]
- Dominguez-Sola, D.; Ying, C.Y.; Grandori, C.; Ruggiero, L.; Chen, B.; Li, M.; Galloway, D.A.; Gu, W.; Gautier, J.; Dalla-Favera, R. Non-transcriptional control of DNA replication by c-Myc. Nature 2007, 448, 445–451. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, V.T.; Fuse, Y.; Tamaoki, J.; Akiyama, S.-I.; Muratani, M.; Tamaru, Y.; Kobayashi, M. Conservation of the Nrf2-mediated gene regulation of proteasome subunits and glucose metabolism in Zebrafish. Oxidative Med. Cell. Longev. 2016, 2016, 5720574. [Google Scholar] [CrossRef] [PubMed]
- Hosiner, D.; Gerber, S.; Lichtenberg-Frate, H.; Glaser, W.; Schueller, C.; Klipp, E. Impact of acute metal stress in saccharomyces cerevisiae. PLoS ONE 2014, 9. [Google Scholar] [CrossRef] [PubMed]
- Tiveron, M.-C.; Cremer, H. CXCL12/CXCR4 signalling in neuronal cell migration. Curr. Opin. Neurobiol. 2008, 18, 237–244. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Sun, B.; Wu, H.; Nie, P. Effects of pure microcystin-LR on the transcription of immune related genes and heat shock proteins in larval stage of zebrafish (Danio rerio). Aquaculture 2009, 289, 154–160. [Google Scholar] [CrossRef]
- Williams, J.H.; Farag, A.M.; Stansbury, M.A.; Young, P.A.; Bergman, H.L.; Petersen, N.S. Accumulation of hsp70 in juvenile and adult rainbow trout gill exposed to metal-contaminated water and/or diet. Environ. Toxicol. Chem. 1996, 15, 1324–1328. [Google Scholar] [CrossRef]
- Vijayan, M.M.; Pereira, C.; Kruzynski, G.; Iwama, G.K. Sublethal concentrations of contaminant induce the expression of hepatic heat shock protein 70 in two salmonids. Aquat. Toxicol. 1998, 40, 101–108. [Google Scholar] [CrossRef]
- Duan, J.C.; Yu, Y.; Li, Y.; Li, Y.B.; Liu, H.C.; Jing, L.; Yang, M.; Wang, J.; Li, C.Q.; Sun, Z.W. Low-dose exposure of silica nanoparticles induces cardiac dysfunction via neutrophil-mediated inflammation and cardiac contraction in zebrafish embryos. Nanotoxicology 2016, 10, 575–585. [Google Scholar] [CrossRef] [PubMed]
- Howe, K.; Clark, M.D.; Torroja, C.F.; Torrance, J.; Berthelot, C.; Muffato, M.; Collins, J.E.; Humphray, S.; McLaren, K.; Matthews, L.; et al. The zebrafish reference genome sequence and its relationship to the human genome. Nature 2013, 496, 498–503. [Google Scholar] [CrossRef] [PubMed]
- Hill, A.; Mesens, N.; Steemans, M.; Xu, J.J.; Aleo, M.D. Comparisons between in vitro whole cell imaging and in vivo zebrafish-based approaches for identifying potential human hepatotoxicants earlier in pharmaceutical development. Drug Metab. Rev. 2012, 44, 127–140. [Google Scholar] [CrossRef] [PubMed]
- Traver, D.; Herbomel, P.; Patton, E.E.; Murphey, R.D.; Yoder, J.A.; Litman, G.W.; Catic, A.; Amemiya, C.T.; Zon, L.I.; Trede, N.S. The zebrafish as a model organism to study development of the immune system. Adv. Immunol. 2003, 81, 254–330. [Google Scholar]
- Marin-Juez, R.; Jong-Raadsen, S.; Yang, S.; Spaink, H.P. Hyperinsulinemia induces insulin resistance and immune suppression via Ptpn6/Shp1 in zebrafish. J. Endocrinol. 2014, 222, 229–241. [Google Scholar] [CrossRef] [PubMed]
- Oehlers, S.H.B.; Flores, M.V.; Hall, C.J.; O’Toole, R.; Swift, S.; Crosier, K.E.; Crosier, P.S. Expression of zebrafish cxcl8 (interleukin-8) and its receptors during development and in response to immune stimulation. Dev. Comp. Immunol. 2010, 34, 352–359. [Google Scholar] [CrossRef] [PubMed]
- Staples, A.; Adams, W.J.; Parkerton, T.F.; Gorsuch, J.W.; Biddinger, G.R.; Reinert, K.H. Aquatic toxicity of eighteen phthalate esters. Environ. Toxicol. Chem. 1997, 16, 875–891. [Google Scholar] [CrossRef]
- Wang, L.; Ying, G.G.; Zhao, J.L.; Liu, S.; Yang, B.; Zhou, L.J.; Tao, R.; Su, H.C. Assessing estrogenic activity in surface water and sediment of the liao river system in northeast china using combined chemical and biological tools. Environ. Pollut. 2011, 159, 148–156. [Google Scholar] [CrossRef] [PubMed]
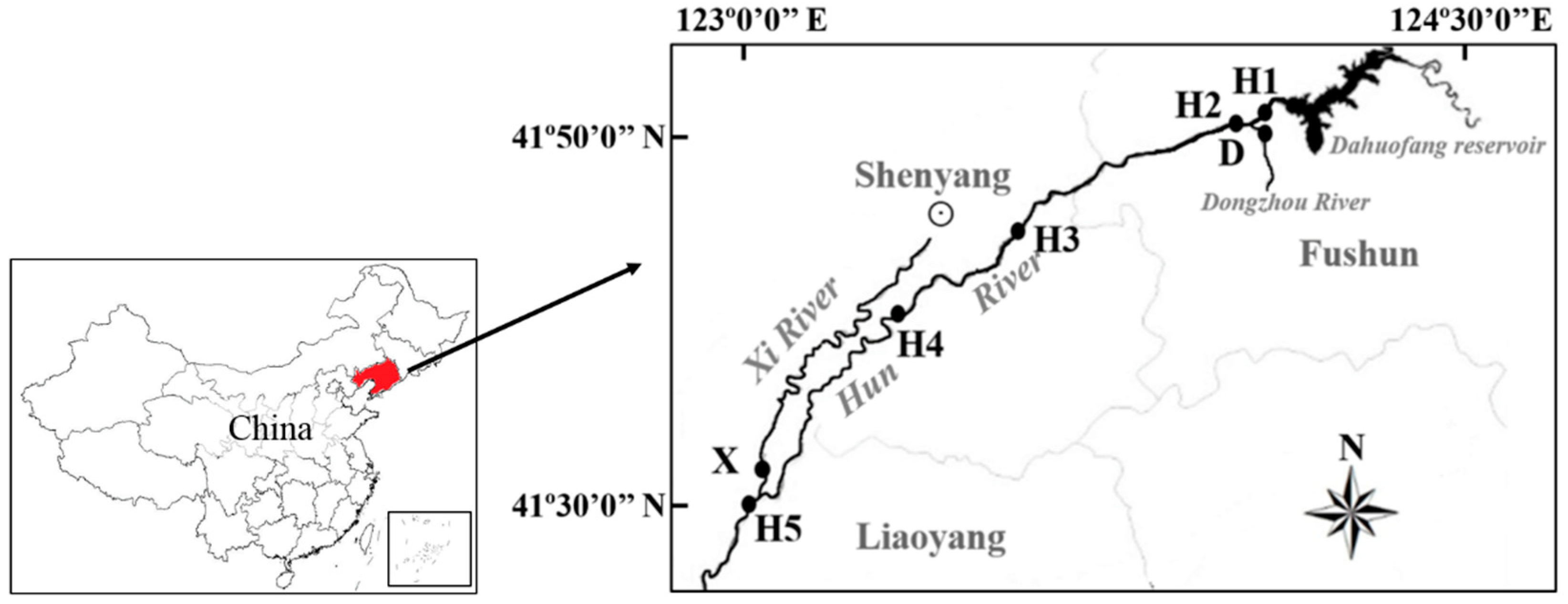
| Site | Longitude (East) | Latitude (North) | pH | Dissolved Oxygen (mg/L) | Chemical Oxygen Demand (mg/L) | Ammonia Nitrogen (mg/L) |
|---|---|---|---|---|---|---|
| H1 | 124°05.627 | 41°53.426 | 8.41 | 7.89 | 7 | 0.1 |
| H2 | 123°58.214 | 41°52.640 | 7.06 | 8.01 | 5 | 0.1 |
| H3 | 123°34.452 | 41°48.704 | 7.01 | 5.03 | 9 | 2.3 |
| H4 | 123°18.311 | 41°42.900 | 7.91 | 9.57 | 13 | 2.7 |
| H5 | 122°57.322 | 41°29.674 | 7.94 | 7.45 | 2 | 0.1 |
| X | 122°59.705 | 41°31.554 | 7.75 | 6.30 | 10 | 0.1 |
| D | 124°02.061 | 41°51.695 | 7.32 | 7.53 | 20 | 0.03 |
| Chemicals | Concentration (μg/L) | ||||
|---|---|---|---|---|---|
| H1 | H2 | H3 | H4 | D | |
| Diisobutyl phthalate | 5.03 | 35.5 | 80.2 | 20.2 | 125 |
| Di(2-ethylhexyl) phthalate | 1.39 | 3.43 | 9.71 | 4.36 | 11.3 |
| 2,4-di-tert-butylphenol | 1.59 | 2.54 | 7.31 | 4.65 | 8.87 |
| Benzyl benzoate | 6.12 | 6.49 | 6.54 | 6.42 | 7.26 |
| m-Cresol | 1.91 | 1.23 | 1.51 | 1.42 | 5.67 |
| Dibutyl phthalate | 0.975 | 2.12 | 2.36 | 2.62 | 3.36 |
| 2-Methylpyridine | 1.7 | 1.42 | 1.05 | 1.04 | 2.55 |
| 2,6-di-tert-butyl-p-cresol | 0.558 | 1.018 | 2.22 | 0.865 | 1.92 |
| o-Cresol | 0.821 | 0.48 | 0.536 | 0.449 | 1.522 |
| 2-Methylnaphthalene | 0.568 | 0.471 | 0.787 | 0.512 | 1.201 |
| Dimethyl phthalate | 0.585 | 0.855 | 0.813 | 1.77 | 1.06 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, Z.; Liu, W.; Qu, Y.; Quan, X.; Zeng, P.; He, M.; Zhou, Y.; Liu, R. Transcriptomic Profiles in Zebrafish Liver Permit the Discrimination of Surface Water with Pollution Gradient and Different Discharges. Int. J. Environ. Res. Public Health 2018, 15, 1648. https://doi.org/10.3390/ijerph15081648
Zhang Z, Liu W, Qu Y, Quan X, Zeng P, He M, Zhou Y, Liu R. Transcriptomic Profiles in Zebrafish Liver Permit the Discrimination of Surface Water with Pollution Gradient and Different Discharges. International Journal of Environmental Research and Public Health. 2018; 15(8):1648. https://doi.org/10.3390/ijerph15081648
Chicago/Turabian StyleZhang, Zhou, Wei Liu, Yuanyuan Qu, Xie Quan, Ping Zeng, Mengchang He, Yanmei Zhou, and Ruixia Liu. 2018. "Transcriptomic Profiles in Zebrafish Liver Permit the Discrimination of Surface Water with Pollution Gradient and Different Discharges" International Journal of Environmental Research and Public Health 15, no. 8: 1648. https://doi.org/10.3390/ijerph15081648
APA StyleZhang, Z., Liu, W., Qu, Y., Quan, X., Zeng, P., He, M., Zhou, Y., & Liu, R. (2018). Transcriptomic Profiles in Zebrafish Liver Permit the Discrimination of Surface Water with Pollution Gradient and Different Discharges. International Journal of Environmental Research and Public Health, 15(8), 1648. https://doi.org/10.3390/ijerph15081648







