Whole Genome Sequencing of a Vietnamese Family from a Dioxin Contamination Hotspot Reveals Novel Variants in the Son with Undiagnosed Intellectual Disability
Abstract
1. Introduction
2. Materials and Methods
2.1. Subjects
2.2. Methods
3. Results
3.1. Clinical Assessment and Dioxin Level in the Father’s Serum
3.2. Whole Genome Sequencing and Identification of Variants
3.3. De Novo Variants
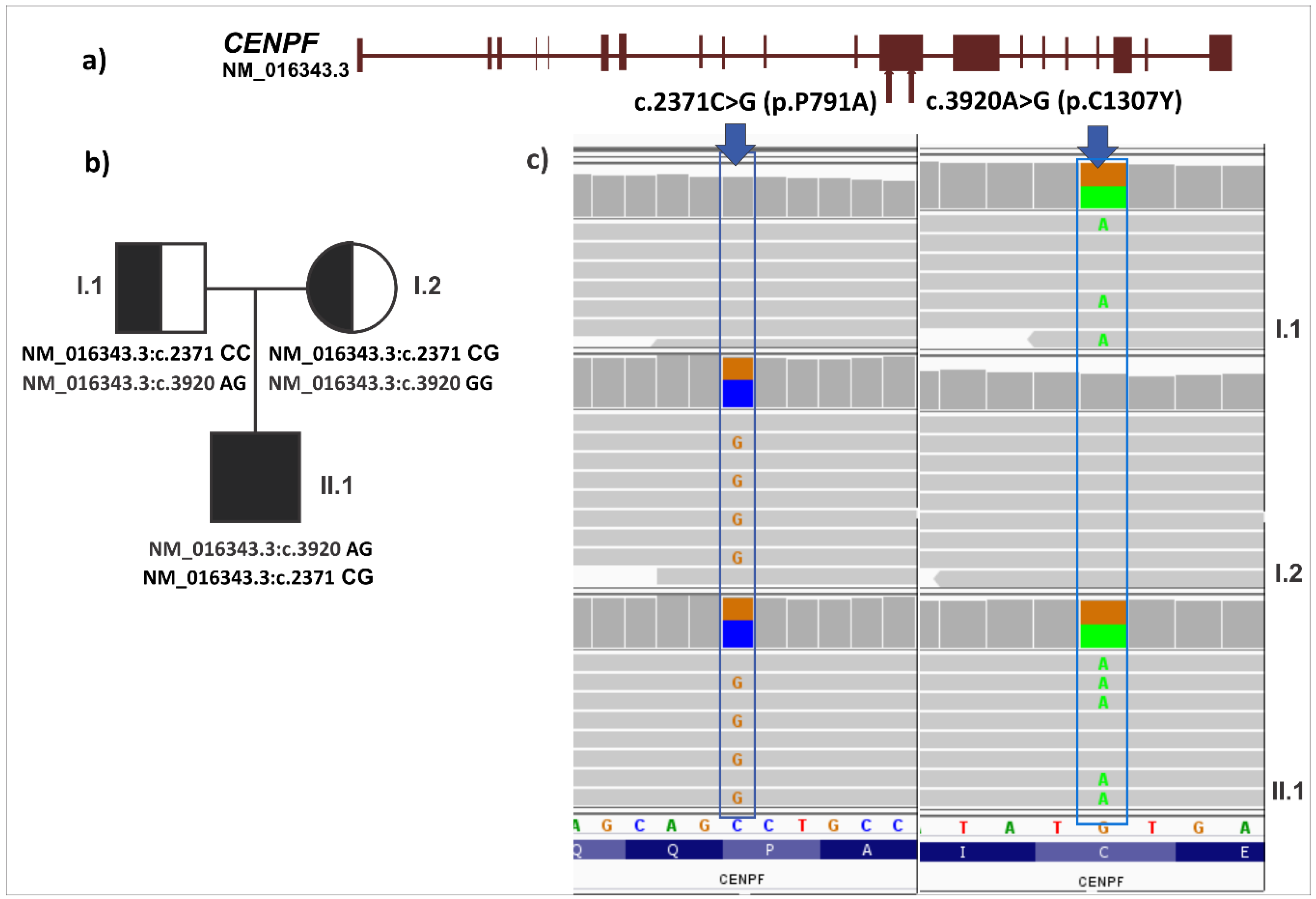
3.4. Analysis of Germline Variations under Recessive Model
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Stellman, J.M.; Stellman, S.D.; Christian, R.; Weber, T.; Tomasallo, C. The extent and patterns of usage of Agent Orange and other herbicides in Vietnam. Nature 2003, 422, 681–687. [Google Scholar] [CrossRef] [PubMed]
- Schecter, A.; Dai, L.C.; Thuy, L.T.; Quynh, H.T.; Minh, D.Q.; Cau, H.D.; Phiet, P.H.; Nguyen, N.T.; Constable, J.D.; Baughman, R.; et al. Agent Orange and the Vietnamese: The persistence of elevated dioxin levels in human tissues. Am. J. Public Health 1995, 85, 516–522. [Google Scholar] [CrossRef] [PubMed]
- Dwernychuk, L.W.; Cau, H.D.; Hatfield, C.T.; Boivin, T.G.; Hung, T.M.; Dung, P.T.; Thai, N.D. Dioxin reservoirs in southern Viet Nam—A legacy of Agent Orange. Chemosphere 2002, 47, 117–137. [Google Scholar] [CrossRef]
- Schecter, A.; Papke, O.; Tung, K.C.; Joseph, J.; Harris, T.R.; Dahlgren, J. Polybrominated diphenyl ether flame retardants in the US population: Current levels, temporal trends, and comparison with dioxins, dibenzofurans, and polychlorinated biphenyls. J. Occup. Environ. Med. 2005, 47, 199–211. [Google Scholar] [CrossRef] [PubMed]
- Schecter, A.; Cao Dai, L.; Päpke, O.; Prange, J.; Constable, J.D.; Matsuda, M.; Duc Thao, V.; Piskac, A.L. Recent dioxin contamination from agent orange in residents of a southern vietnam city. J. Occup. Environ. Med. 2001, 43, 435–443. [Google Scholar] [CrossRef] [PubMed]
- Tuyet-Hanh, T.T.; Vu-Anh, L.; Ngoc-Bich, N.; Tenkate, T. Environmental health risk assessment of dioxin exposure through foods in a dioxin hot spot-Bien Hoa City, Vietnam. Int. J. Environ. Res. Public Health 2010, 7, 2395–2406. [Google Scholar] [CrossRef] [PubMed]
- Huyen, D.T.; Igarashi, T.; Shiraiwa, T. Vertical distribution of dioxins in soil of Bien Hoa airbase, Vietnam. Springerplus 2015, 4, 300. [Google Scholar] [CrossRef] [PubMed]
- Manh, H.D.; Kido, T.; Tai, P.T.; Okamoto, R.; Honma, S.; Liang, S.X.; Anh le, T.; Maruzeni, S.; Nghi, T.N.; Nishijo, M.; et al. Levels of polychlorinated dibenzodioxins and polychlorinated dibenzofurans in breast milk samples from three dioxin-contaminated hotspots of Vietnam. Sci. Total Environ. 2015, 511, 416–422. [Google Scholar] [CrossRef] [PubMed]
- Nghi, T.N.; Nishijo, M.; Manh, H.D.; Tai, P.T.; Van Luong, H.; Anh, T.H.; Thao, P.N.; Trung, N.V.; Waseda, T.; Nakagawa, H.; et al. Dioxins and nonortho PCBs in breast milk of vietnamese mothers living in the largest hot spot of dioxin contamination. Environ. Sci. Technol. 2015, 49, 5732–5742. [Google Scholar] [CrossRef] [PubMed]
- National Academies of Sciences, Engineering, and Medicine, Institute of Medicine, Board on the Health of Select Populations, Committee to Review the Health Effects in Vietnam Veterans of Exposure to Herbicides (Tenth Biennial Update). National academies of sciences, engineering, and medicine. In Veterans and Agent Orange: Update 2014; National Academies Press: Washington, DC, USA, 2016. [Google Scholar]
- Ton, N.D.; Nakagawa, H.; Ha, N.H.; Duong, N.T.; Nhung, V.P.; Hien, L.T.T.; Hue, H.T.T.; Hoang, N.H.; Wong, J.H.; Nakano, K.; et al. Whole genome sequencing and mutation rate analysis of trios with paternal dioxin exposure. Hum. Mutat. 2018, 39, 1384–1392. [Google Scholar] [CrossRef] [PubMed]
- Pavlowsky, A.; Chelly, J.; Billuart, P. Emerging major synaptic signaling pathways involved in intellectual disability. Mol. Psychiatry 2012, 17, 682–693. [Google Scholar] [CrossRef] [PubMed]
- Vissers, L.E.; Gilissen, C.; Veltman, J.A. Genetic studies in intellectual disability and related disorders. Nat. Rev. Genet. 2016, 17, 9–18. [Google Scholar] [CrossRef] [PubMed]
- Iossifov, I.; O’Roak, B.J.; Sanders, S.J.; Ronemus, M.; Krumm, N.; Levy, D.; Stessman, H.A.; Witherspoon, K.T.; Vives, L.; Patterson, K.E.; et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature 2014, 515, 216–221. [Google Scholar] [CrossRef] [PubMed]
- The Deciphering Developmental Disorders Study. Large-scale discovery of novel genetic causes of developmental disorders. Nature 2015, 519, 223–228. [Google Scholar] [CrossRef] [PubMed]
- Rovelet-Lecrux, A.; Charbonnier, C.; Wallon, D.; Nicolas, G.; Seaman, M.N.; Pottier, C.; Breusegem, S.Y.; Mathur, P.P.; Jenardhanan, P.; Le Guennec, K.; et al. De novo deleterious genetic variations target a biological network centered on Abeta peptide in early-onset Alzheimer disease. Mol. Psychiatry 2015, 20, 1046–1056. [Google Scholar] [CrossRef] [PubMed]
- Gilissen, C.; Hehir-Kwa, J.Y.; Thung, D.T.; van de Vorst, M.; van Bon, B.W.; Willemsen, M.H.; Kwint, M.; Janssen, I.M.; Hoischen, A.; Schenck, A.; et al. Genome sequencing identifies major causes of severe intellectual disability. Nature 2014, 511, 344–347. [Google Scholar] [CrossRef] [PubMed]
- Agha, Z.; Iqbal, Z.; Azam, M.; Ayub, H.; Vissers, L.E.; Gilissen, C.; Ali, S.H.; Riaz, M.; Veltman, J.A.; Pfundt, R.; et al. Exome sequencing identifies three novel candidate genes implicated in intellectual disability. PLoS ONE 2014, 9, e112687. [Google Scholar] [CrossRef] [PubMed]
- Chiurazzi, P.; Pirozzi, F. Advances in understanding—Genetic basis of intellectual disability. F1000Research 2016, 5. [Google Scholar] [CrossRef] [PubMed]
- Schecter, A.; Pavuk, M.; Papke, O.; Malisch, R. The use of potassium dichromate and ethyl alcohol as blood preservatives for analysis of organochlorine contaminants. Chemosphere 2004, 57, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef] [PubMed]
- Fujimoto, A.; Totoki, Y.; Abe, T.; Boroevich, K.A.; Hosoda, F.; Nguyen, H.H.; Aoki, M.; Hosono, N.; Kubo, M.; Miya, F.; et al. Whole-genome sequencing of liver cancers identifies etiological influences on mutation patterns and recurrent mutations in chromatin regulators. Nat. Genet 2012, 44, 760–764. [Google Scholar] [CrossRef] [PubMed]
- Andersson, R.; Bruder, C.E.; Piotrowski, A.; Menzel, U.; Nord, H.; Sandgren, J.; Hvidsten, T.R.; Diaz de Stahl, T.; Dumanski, J.P.; Komorowski, J. A segmental maximum a posteriori approach to genome-wide copy number profiling. Bioinformatics 2008, 24, 751–758. [Google Scholar] [CrossRef] [PubMed]
- Adzhubei, I.A.; Schmidt, S.; Peshkin, L.; Ramensky, V.E.; Gerasimova, A.; Bork, P.; Kondrashov, A.S.; Sunyaev, S.R. A method and server for predicting damaging missense mutations. Nat. Methods 2010, 7, 248–249. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.; Sims, G.E.; Murphy, S.; Miller, J.R.; Chan, A.P. Predicting the functional effect of amino acid substitutions and indels. PLoS ONE 2012, 7, e46688. [Google Scholar] [CrossRef] [PubMed]
- Kumar, P.; Henikoff, S.; Ng, P.C. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat. Protoc. 2009, 4, 1073–1081. [Google Scholar] [CrossRef] [PubMed]
- The 1000 Genomes Project Consortium; Auton, A.; Brooks, L.D.; Durbin, R.M.; Garrison, E.P.; Kang, H.M.; Korbel, J.O.; Marchini, J.L.; McCarthy, S.; McVean, G.A.; et al. A global reference for human genetic variation. Nature 2015, 526, 68–74. [Google Scholar] [PubMed]
- Schecter, A.; Pavuk, M.; Constable, J.D.; Dai le, C.; Papke, O. A follow-up: High level of dioxin contamination in Vietnamese from agent orange, three decades after the end of spraying. J. Occup. Environ. Med. 2002, 44, 218–220. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.H.; Yuen, R.K.; Jin, X.; Wang, M.; Chen, N.; Wu, X.; Ju, J.; Mei, J.; Shi, Y.; He, M.; et al. Detection of clinically relevant genetic variants in autism spectrum disorder by whole-genome sequencing. Am. J. Hum. Genet. 2013, 93, 249–263. [Google Scholar] [CrossRef] [PubMed]
- Kong, A.; Frigge, M.L.; Masson, G.; Besenbacher, S.; Sulem, P.; Magnusson, G.; Gudjonsson, S.A.; Sigurdsson, A.; Jonasdottir, A.; Jonasdottir, A.; et al. Rate of de novo mutations and the importance of father’s age to disease risk. Nature 2012, 488, 471–475. [Google Scholar] [CrossRef] [PubMed]
- Conrad, D.F.; Keebler, J.E.; DePristo, M.A.; Lindsay, S.J.; Zhang, Y.; Casals, F.; Idaghdour, Y.; Hartl, C.L.; Torroja, C.; Garimella, K.V.; et al. Variation in genome-wide mutation rates within and between human families. Nat. Genet. 2011, 43, 712–714. [Google Scholar] [CrossRef] [PubMed]
- Neale, B.M.; Kou, Y.; Liu, L.; Ma’ayan, A.; Samocha, K.E.; Sabo, A.; Lin, C.F.; Stevens, C.; Wang, L.S.; Makarov, V.; et al. Patterns and rates of exonic de novo mutations in autism spectrum disorders. Nature 2012, 485, 242–245. [Google Scholar] [CrossRef] [PubMed]
- Michaelson, J.J.; Shi, Y.; Gujral, M.; Zheng, H.; Malhotra, D.; Jin, X.; Jian, M.; Liu, G.; Greer, D.; Bhandari, A.; et al. Whole-genome sequencing in autism identifies hot spots for de novo germline mutation. Cell 2012, 151, 1431–1442. [Google Scholar] [CrossRef] [PubMed]
- Wolvetang, E.W.; Bradfield, O.M.; Tymms, M.; Zavarsek, S.; Hatzistavrou, T.; Kola, I.; Hertzog, P.J. The chromosome 21 transcription factor ETS2 transactivates the beta-APP promoter: Implications for Down syndrome. Biochim. Biophys. Acta 2003, 1628, 105–110. [Google Scholar] [CrossRef]
- Chumakov, A.M.; Chen, D.L.; Chumakova, E.A.; Koeffler, H.P. Localization of the c-ets-2 transactivation domain. J. Virol. 1993, 67, 2421–2425. [Google Scholar] [PubMed]
- Yi, Z.; Li, Y.; Ma, W.; Li, D.; Zhu, C.; Luo, J.; Wang, Y.; Huang, X.; Yuan, W.; Liu, M.; et al. A novel KRAB zinc-finger protein, ZNF480, expresses in human heart and activates transcriptional activities of AP-1 and SRE. Biochem. Biophys. Res. Com. 2004, 320, 409–415. [Google Scholar] [CrossRef] [PubMed]
- Shen, E.H.; Overly, C.C.; Jones, A.R. The Allen Human Brain Atlas: Comprehensive gene expression mapping of the human brain. Trends Neurosci. 2012, 35, 711–714. [Google Scholar] [CrossRef] [PubMed]
- Piton, A.; Redin, C.; Mandel, J.L. XLID-causing mutations and associated genes challenged in light of data from large-scale human exome sequencing. Am. J. Hum. Genet. 2013, 93, 368–383. [Google Scholar] [CrossRef] [PubMed]
- Waters, A.M.; Asfahani, R.; Carroll, P.; Bicknell, L.; Lescai, F.; Bright, A.; Chanudet, E.; Brooks, A.; Christou-Savina, S.; Osman, G.; et al. The kinetochore protein, CENPF, is mutated in human ciliopathy and microcephaly phenotypes. J. Med. Genet. 2015, 52, 147–156. [Google Scholar] [CrossRef] [PubMed]

| Congener | TEQ (ppt) |
|---|---|
| Sample amount (g) | 15.236 |
| lipid content (%) | 0.177 |
| Sample weight (lipid) (g) | 0.027 |
| Values in pg/g (ppt), lipid based | |
| 2.3.7.8-Tetra-CDD (TCDD) | 87 |
| 1.2.3.7.8-Penta-CDD (PeCDD) | 12 |
| 1.2.3.4.7.8-Hexa-CDD | 9.0 |
| 1.2.3.6.7.8-Hexa-CDD | 33 |
| 1.2.3.7.8.9-Hexa-CDD | 8.2 |
| 1.2.3.4.6.7.8-Hepta-CDD | 35 |
| OCDD | 385 |
| 2.3.7.8-Tetra-CDF | 3.2 |
| 1.2.3.7.8-Penta-CDF | 2.8 |
| 2.3.4.7.8-Penta-CDF | 9.6 |
| 1.2.3.4.7.8-Hexa-CDF | 27 |
| 1.2.3.6.7.8-Hexa-CDF | 15 |
| 1.2.3.7.8.9-Hexa-CDF | n.d. (3) |
| 2.3.4.6.7.8-Hexa-CDF | 8.4 |
| 1.2.3.4.6.7.8-Hepta-CDF | 18 |
| 1.2.3.4.7.8.9-Hepta-CDF | n.d. (4) |
| OCDF | n.d. (13) |
| 3,3’,4,4’-TCB (77) | n.d. (690) |
| 3,4,4’,5-TCB (81) | n.d. (29) |
| 3,3’,4,4’,5-PeCB (126) | 123 |
| 3,3’,4,4’,5,5’-HxCB (169) | 61 |
| 2,3,3’,4,4’-PeCB (105) | 5005 |
| 2,3,4,4’,5-PeCB (114) | 742 |
| 2,3’,4,4’,5-PeCB (118) | 22,592 |
| 2’,3,4,4’,5-PeCB (123) | 244 |
| 2,3,3’,4,4’,5-HxCB (156) | 5873 |
| 2,3,3’,4,4’,5’-HxCB (157) | 1579 |
| 2,3’,4,4’,5,5’-HxCB (167) | 2883 |
| 2,3,3’,4,4’,5,5’-HpCB (189) | 938 |
| Total PCDDs/PCDFs | 654 |
| TEQ (World Health Organization, WHO) based on PCDD/F | 115 |
| Type | Father | Mother | Proband | |
|---|---|---|---|---|
| WGS deep coverage (x) | 32.2 | 31.8 | 31.5 | |
| SNV | 1,461,494 | 1,419,542 | 1,388,686 | |
| Shared with dbSNP v138 | 1,438,017 | 1,396,439 | 1,366,410 | |
| Shared with 1000G | 6619 | 6963 | 6526 | |
| Novel variants | 16,858 | 16,140 | 15,750 | |
| Intronic | 13,555 | 13,068 | 12,751 | |
| Exonic | 243 | 239 | 204 | |
| 5’UTR | 54 | 46 | 38 | |
| 3’UTR | 309 | 289 | 259 | |
| ncRNA_intronic | 2551 | 2355 | 2379 | |
| ncRNA_exonic | 145 | 140 | 117 | |
| Splicing site | 1 | 3 | 2 | |
| Indel | 185,588 | 180,389 | 174,440 | |
| Shared with dbSNP v138 | 91,552 | 89,948 | 87,532 | |
| Shared with 1000G | 5048 | 4942 | 4560 | |
| Novel variants | 88,988 | 85,499 | 82,348 | |
| Intronic | 73,626 | 70,493 | 67,997 | |
| Exonic | 63 | 56 | 45 | |
| 5’UTR | 129 | 114 | 109 | |
| 3’UTR | 1850 | 1807 | 1690 | |
| ncRNA_intronic | 12,750 | 12,495 | 12,000 | |
| ncRNA_exonic | 502 | 479 | 448 | |
| Splicing site | 68 | 55 | 59 | |
| Total (SNV + Indel) | 1,647,082 | 1,599,931 | 1,563,126 |
| Type of Variant | Chromosomal Position | Transcript Level Position | Gene | Ref | Variant | AA Change | Provean Prediction | SIFT Prediction | Polyphen-2 Prediction | |||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Prediction | Score | Prediction | Score | Prediction | Score | |||||||
| de novo | NC_000021.8:g.40182013 | NM_001256295.1:c.47 | ETS2 | CC | CT | T22I | Not detected | Not detected | Not detected | Not detected | benign | 0.055 |
| de novo | NC_000019.9:g.52826007 | NM_144684.2:c.1504 | ZNF480 | CC | CT | R502W | Deleterious | −2.74 | Tolerated | 0.237 | probably damaging | 0.999 |
| compound heterozygosity | NC_000001.10:g.214814052 | NM_016343.3:c.2371 | CENPF | CC | CG | P791A | Neutral | −0.24 | Tolerated | 0.597 | benign | 0.001 |
| compound heterozygosity | NC_000001.10:g.214815601 | NM_016343.3:c.3920 | CENPF | GG | GA | C1307Y | Deleterious | −4.22 | Tolerated | 0.087 | probably damaging | 1 |
| compound heterozygosity | NC_000002.11:g.179514619 | XM_005246830.1:c.907 | TTN | GG | AG | P13274S | Deleterious | −3.53 | Tolerated | 0.354 | benign | 0.006 |
| compound heterozygosity | NC_000002.11:g.179578790 | NM_005246830.1:c.25647 | TTN | CC | CG | K8865N | Deleterious | −3.35 | Tolerated | 0.091 | probably damaging | 0.978 |
| homozygous deletion | NC_000001.10:g. 149040000_149195000del | NBPF25P | NC_000001.10:g. 149040000_149195000del (150 kb deletion) | |||||||||
| homozygous deletion | NC_000012.11:g.10580000_10590000del | KLRC1, KLRC2 | NC_000012.11:g.10580000_10590000del (10 kb deletion) | |||||||||
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nguyen, D.T.; Nguyen, H.H.; Nguyen, T.D.; Nguyen, T.T.H.; Nakano, K.; Maejima, K.; Sasaki-Oku, A.; Nguyen, V.B.; Nguyen, D.B.; Le, B.Q.; et al. Whole Genome Sequencing of a Vietnamese Family from a Dioxin Contamination Hotspot Reveals Novel Variants in the Son with Undiagnosed Intellectual Disability. Int. J. Environ. Res. Public Health 2018, 15, 2629. https://doi.org/10.3390/ijerph15122629
Nguyen DT, Nguyen HH, Nguyen TD, Nguyen TTH, Nakano K, Maejima K, Sasaki-Oku A, Nguyen VB, Nguyen DB, Le BQ, et al. Whole Genome Sequencing of a Vietnamese Family from a Dioxin Contamination Hotspot Reveals Novel Variants in the Son with Undiagnosed Intellectual Disability. International Journal of Environmental Research and Public Health. 2018; 15(12):2629. https://doi.org/10.3390/ijerph15122629
Chicago/Turabian StyleNguyen, Dang Ton, Hai Ha Nguyen, Thuy Duong Nguyen, Thi Thanh Hoa Nguyen, Kaoru Nakano, Kazuhiro Maejima, Aya Sasaki-Oku, Van Ba Nguyen, Duy Bac Nguyen, Bach Quang Le, and et al. 2018. "Whole Genome Sequencing of a Vietnamese Family from a Dioxin Contamination Hotspot Reveals Novel Variants in the Son with Undiagnosed Intellectual Disability" International Journal of Environmental Research and Public Health 15, no. 12: 2629. https://doi.org/10.3390/ijerph15122629
APA StyleNguyen, D. T., Nguyen, H. H., Nguyen, T. D., Nguyen, T. T. H., Nakano, K., Maejima, K., Sasaki-Oku, A., Nguyen, V. B., Nguyen, D. B., Le, B. Q., Wong, J. H., Tsunoda, T., Nakagawa, H., Fujimoto, A., & Nong, V. H. (2018). Whole Genome Sequencing of a Vietnamese Family from a Dioxin Contamination Hotspot Reveals Novel Variants in the Son with Undiagnosed Intellectual Disability. International Journal of Environmental Research and Public Health, 15(12), 2629. https://doi.org/10.3390/ijerph15122629





