Antibacterial Activity of Marine and Black Band Disease Cyanobacteria against Coral-Associated Bacteria
Abstract
:1. Introduction
2. Results
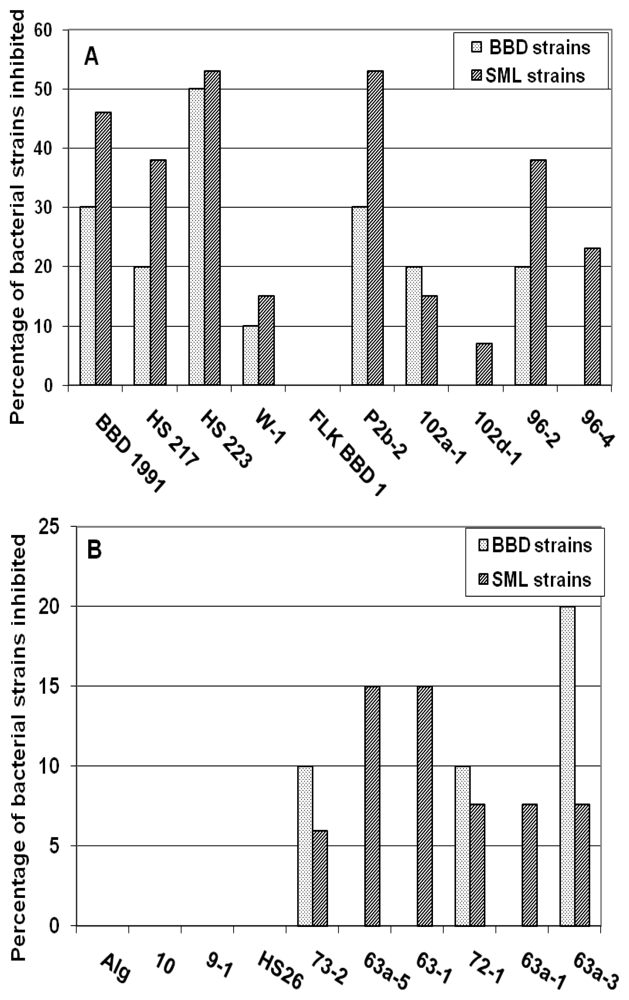
2.1. Inhibition of BBD and SML Bacteria during Co-Cultivation with Cyanobacteria
2.2. Interactions between Individual Bacterial and Cyanobacterial Strains
2.3. Activity of Cyanobacterial Extracts
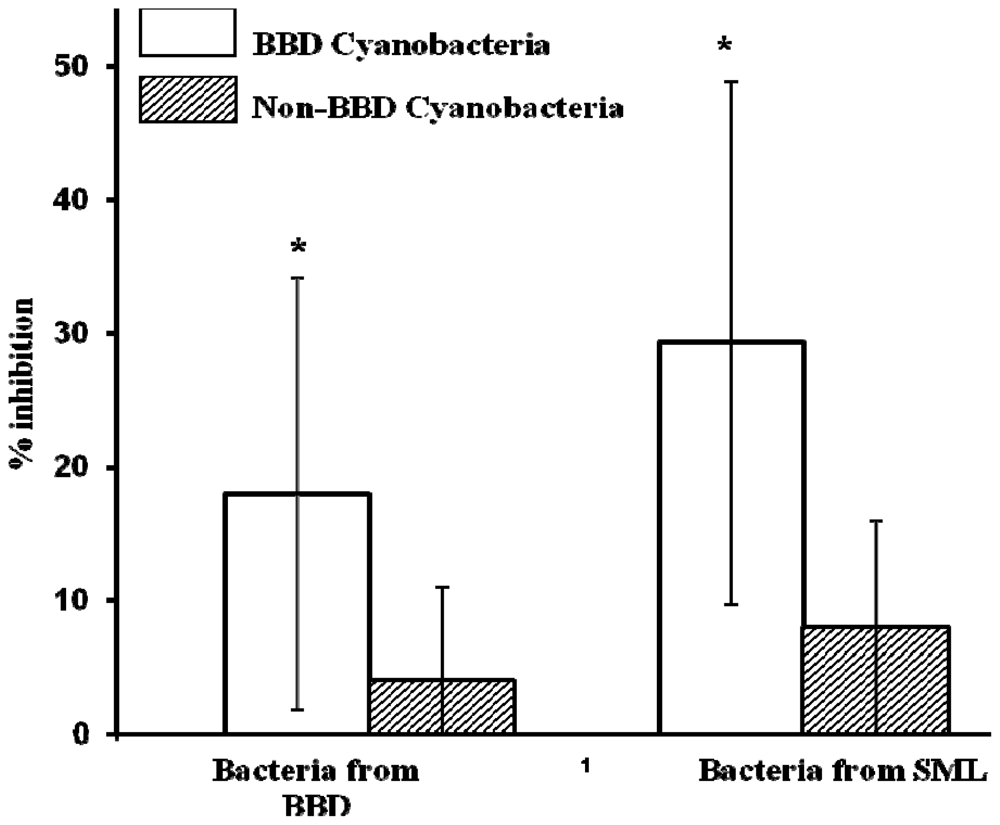
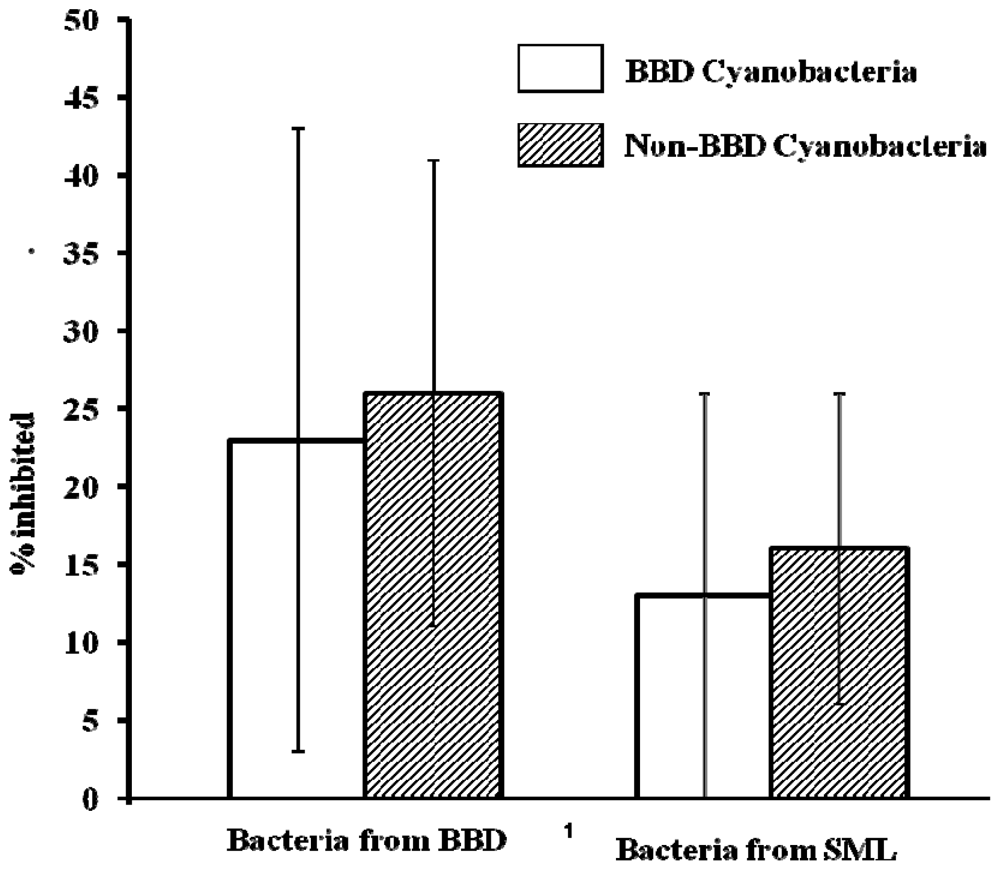
2.4. Activity of Extracts vs. Co-Cultivation with SML and BBD Bacteria
2.5. Antibacterial Activity against Known Coral Pathogens
3. Discussion
4. Experimental Section
4.1. Source of Cyanobacterial and Bacterial Isolates
4.2. Field Sampling
4.3. Co-Cultivation of Cyanobacteria and Heterotrophic Bacteria
4.4. Preparation of Cyanobacterial Extracts
4.5. Activity of Cyanobacterial Extracts
4.6. Statistical Analysis
5. Conclusions
Acknowledgements
- Samples Availability: Available from the authors.
References
- Weil, E. Coral reef diseases in the wider Caribbean. In Coral Health and Disease; Rosenberg, E, Loya, Y, Eds.; Springer-Verlag: Berlin, Germany, 2004; pp. 35–68. [Google Scholar]
- Sutherland, K; Porter, J; Torres, C. Disease and immunity in Caribbean and Indo-Pacific zooxanthellate corals. Mar. Ecol. Prog. Ser 2004, 266, 273–302. [Google Scholar]
- Dinsdale, E. Abundance of black band disease on corals from one location on the Great Barrier Reef: a comparison with abundance in the Caribbean Region. Proceedings of the 9th International Coral Reef Symposium, Bali, Indonesia, 23–27 October 2000; 2, pp. 1238–1244.
- Al-Moghrabi, M. Unusual black band disease (BBD) outbreak in the northern tip of the Gulf of Aquaba (Jordan). Coral Reefs 2001, 19, 330–331. [Google Scholar]
- Rützler, K; Santavy, D; Antonius, A. The black band disease of Atlantic reef corals. III. Distribution, ecology and development. PSZNI Mar. Ecol 1983, 4, 329–358. [Google Scholar]
- Ducklow, HW; Mitchell, R. Observations on natural and artificially diseased tropical corals: A scanning electron microscope study. Microb. Ecol 1979, 5, 215–223. [Google Scholar]
- Cooney, RP; Pantos, O; Le Tissier, MDA; Barer, MR; O’Donell, A; Bythell, JC. Characterization of the bacterial consortium associated with black band disease in coral using molecular microbiological techniques. Environ. Microbiol 2002, 4, 401–413. [Google Scholar]
- Frias-Lopez, J; Zerkle, A; Bonheyo, G; Fouke, B. Partitioning of bacterial communities between seawater and healthy, black band diseased, and dead coral surfaces. Appl. Environ. Microbiol 2002, 68, 2214–2228. [Google Scholar]
- Frias-Lopez, J; Klaus, JS; Boneyo, GT; Fouke, BW. Bacterial community associated with black band disease in corals. Appl. Environ. Microbiol 2004, 70, 5955–5962. [Google Scholar]
- Sekar, R; Mills, DK; Remily, ER; Voss, JD; Richardson, LL. Microbial communities in the surface mucopolysaccharide layer and the black band microbial mat of black band diseased Siderastrea siderea. Appl. Environ. Microbiol 2006, 72, 5963–5973. [Google Scholar]
- Reshef, L; Koren, O; Loya, Y; Zilber-Rosenberg, I; Rosenberg, E. The coral probiotic hypothesis. Environ. Microbiol 2006, 8, 2068–2073. [Google Scholar]
- Rohwer, F; Breitbart, M; Jara, J; Azam, F; Knowlton, N. Diversity of bacteria associated with the Caribbean coral Montastraea franksi. Coral Reefs 2001, 20, 85–91. [Google Scholar]
- Rohwer, F; Segritan, V; Azam, F; Knowlton, N. Diversity and distribution of coral-associated bacteria. Mar. Ecol. Prog. Ser 2002, 243, 1–10. [Google Scholar]
- Klaus, JS; Frias-Lopez, J; Bonheyo, GT; Heikoop, JM; Fouke, BW. Bacterial communities inhabiting the healthy tissues of two Caribbean reef corals: interspecific and spatial variation. Coral Reefs 2005, 24, 129–137. [Google Scholar]
- Bourne, DG; Munn, CB. Diversity of bacteria associated with the coral Pocillopora damicornis from the Great Barrier Reef. Environ. Microbiol 2005, 7, 1162–1174. [Google Scholar]
- Ritchie, KB. Regulation of microbial populations by coral surface mucus and mucus-associated bacteria. Mar. Ecol. Prog. Ser 2006, 322, 1–14. [Google Scholar]
- Harder, T; Lau, SCK; Dobretsov, S; Fang, TK; Quian, P-Y. A distinctive epibiotic bacterial community on the soft coral Dendronephtya sp. and antibacterial activity of coral tissue extracts suggest a chemical mechanism against bacterial epibiosis. FEMS Microbiol. Ecol 2003, 43, 337–347. [Google Scholar]
- Koh, EGL. Do scleractinian corals engage in chemical warfare against microbes? J. Chem. Ecol 1997, 23, 379–398. [Google Scholar]
- Gochfield, DJ; Aeby, GS. Antibacterial chemical defenses in Hawaiian corals provide possible protection from disease. Mar. Ecol. Prog. Ser 2008, 362, 119–128. [Google Scholar]
- Kim, K. Antimicrobial activity in gorgonian corals (Coelentrate, Octocorallia). Coral Reefs 1994, 13, 75–80. [Google Scholar]
- Rypien, KL; Ward, JR; Azam, F. Antagonistic interactions among coral-associated bacteria. Environ. Microbiol 2010, 12, 28–39. [Google Scholar]
- Kelman, D; Kashman, Y; Rosenberg, E; Kushmaro, A; Loya, Y. Antimicrobial activity of Red Sea corals. Mar. Biol 2006, 149, 357–363. [Google Scholar]
- Østensvik, Ø; Skulberg, OM; Underdal, OM; Hormazabal, V. Antibacterial properties of extracts from selected planktonic freshwater cyanobacteria—a comparative study of bacterial bioassays. J. Appl. Microbiol 1998, 84, 1117–1124. [Google Scholar]
- Jaki, B; Orjala, J; Sticher, O. A novel extracellular diterpenoid with antibacterial activity from the cyanobacterium Nostoc commune. J. Nat. Prod 1999, 62, 502–503. [Google Scholar]
- Skulberg, OM. Microalgae as a source of bioactive molecules—experience from cyanophyte research. J. Appl. Phycol 2000, 12, 341–348. [Google Scholar]
- Martins, RF; Ramos, MF; Herfindal, L; Sousa, JA; Skærven, K; Vasconcelos, VM. Antimicrobial and cytotoxic assessment of marine cyanobacteria—Synechocystis and Synechococcus. Mar. Drugs 2008, 6, 1–11. [Google Scholar]
- Morrow, KM; Paul, VJ; Liles, MR; Chadwick, NE. Allelochemicals produced by Caribbean macroalgae and cyanobacteria have species-specific effects on reef coral microorganisms. Coral Reefs 2011, 30, 309–320. [Google Scholar]
- Kreitlow, S; Mundt, S; Lindequist, U. Cyanobacteria—a potential source of new biologically active substances. J. Biotechnol 1999, 70, 61–63. [Google Scholar]
- Miller, AW; Richardson, LL. A meta-analysis of 16S rRNA gene clone libraries from the polymicrobial black band disease of corals. FEMS Microbiol. Ecol 2011, 75, 231–241. [Google Scholar]
- Nissimov, JRE; Munn, CB. Antimicrobial properties of resident coral mucus bacteria of Oculina patagonica. FEMS Microbiol. Lett 2009, 292, 210–215. [Google Scholar]
- Rosenberg, E; Kushmaro, A; Kramarsky-Winter, E; Banin, E; Loya, Y. The role of microorganisms in coral bleaching. ISME J 2009, 3, 139–146. [Google Scholar]
- Richardson, LL; Sekar, R; Myers, JL; Gantar, M; Remily, ER; Kaczmarsky, LT; Voss, JD; Boyer, GL; Zimba, PV. The presence of the cyanobacterial toxin microcystin in black band disease of corals. FEMS Microbiol. Ecol 2007, 272, 182–187. [Google Scholar]
- Gantar, M; Sekar, R; Richardson, LL. Cyanotoxins from black band disease of corals and from other coral reef environments. Microb. Ecol 2009, 58, 856–864. [Google Scholar]
- Stanić, D; Oehrle, S; Gantar, M; Richardson, LL. Microcystin production and ecological physiology of Caribbean black band disease cyanobacteria. Environ. Microbiol 2011, 13, 900–910. [Google Scholar]
- Richardson, LL; Miller, AW; Broderick, E; Kaczmarsky, L; Gantar, M; Stanić, D; Sekar, R. Sulfide, microcystin, and the etiology of black band disease. Dis. Aquat. Org 2009, 87, 79–90. [Google Scholar]
- Richardson, LL; Kuta, KG. Ecological physiology of the black band disease cyanobacterium Phormidium corallyticum. FEMS Microbiol. Ecol 2003, 43, 287–298. [Google Scholar]
- Myers, JL; Sekar, R; Richardson, LL. Molecular detection and ecological significance of the cyanobacterial genera Geitlerinema and Leptolyngbya in black band disease of corals. Appl. Environ. Microbiol 2007, 73, 5173–5182. [Google Scholar]
- Myers, JL; Richardson, LL. Adaptation of cyanobacteria to the sulfide-rich microenvironment of black band disease of coral. FEMS Microbiol. Ecol 2009, 67, 242–251. [Google Scholar]
- Sato, Y; Willis, BL; Bourne, DG. Successional changes in bacterial communities during the development of black band disease on the reef coral, Montipora hispida. ISME J 2009, 4, 203–214. [Google Scholar]
- Voss, JD; Mills, DK; Myers, JL; Remily, ER; Richardson, LL. Black band disease microbial community variation on corals in three regions of the wider Caribbean. Microb. Ecol 2007, 54, 730–739. [Google Scholar]
- Denner, EBM; Smith, G; Busse, H-J; Schumann, P; Narzt, T; Polson, S; Lubitz, W; Richardson, LL. Aurantimonas coralicida gen. nov., sp. nov., the causative agent of white plague type II on Caribbean scleractinian corals. Int. J. Syst. Evol. Microbiol 2003, 53, 1115–1122. [Google Scholar]
- Patterson, K; Porter, J; Ritchie, K; Polson, S; Mueller, E; Peters, E; Santavy, D; Smith, G. The etiology of white pox, a lethal disease of the Caribbean elkhorn coral, Acropora palmata. Proc. Natl. Acad. Sci. USA 2002, 99, 8725–8730. [Google Scholar]
- Kushmaro, A; Banin, E; Stackebrandt, E; Rosenberg, E. Vibrio shiloi sp. nov: the causative agent of bleaching of the coral Oculina patagonica. Int. J. Syst. Evol. Microbiol 2001, 51, 1383–1388. [Google Scholar]
- Sekar, R; Kaczmarsky, LT; Richardson, LL. Microbial community composition of black band disease on the coral host Siderastrea siderea from three regions of the wider Caribbean. Mar. Ecol. Prog. Ser 2008, 362, 85–98. [Google Scholar]
- Anagnostidis, K; Komarek, J. Modern approach to the classification system of cyanophytes. 3-Oscillatoriales. Arch. Hydrobiol. Suppl 1988, 80, 327–472. [Google Scholar]
- Gantar, M; Berry, JP; Thomas, S; Wang, M; Rein, KS; Perez, R; Gawley, RE. Allelopathic activity among cyanobacteria and microalgae isolated from Florida freshwater habitats. FEMS Microbiol. Ecol 2008, 64, 55–64. [Google Scholar]
- Rippka, R; Deruelles, J; Waterbury, JB; Herdman, M; Stanier, RY. Generic assignments, strain histories and properties of pure cultures of cyanobacteria. J. Gen. Microbiol 1979, 111, 1–61. [Google Scholar]
| Cyanobacteria | Bacterial strains inhibited or stimulated by co-cultivation | Bacterial strains inhibited by lipophilic extracts | |
|---|---|---|---|
| BBD Isolates | |||
| BBD 1991 | Geitlerinema | B3, B5, B6, S1, S3, S5, S7, S9, S11 | B1, B2, B5, B9, B10, P2, P2, S2, S8, S12 |
| HS 217 | Geitlerinema | B3, B6, S5, S6, S7, S8, S13 | P1 |
| HS 223 | Geitlerinema | B1, B3, B5, B6, B8, P3, S1, S3, S4, S6, S7, S9, S13 | B1, B2, B3, B5, B9, B10, P1, P2, S2, S3 |
| W-1 | Geitlerinema | B6, S6, S7 | B9, B10, S9, S10 |
| FLK BBD1 | Leptolyngbya | No effect | B1, B2, B5, B9, P1, P2, S3, S6, S8 |
| P2b-2 | Leptolyngbya | B1, B3, B6, S3, S4, S5, S6, S7, S11, S13 | B5, B9, S1, S4, S7, S9, S10 |
| 102a-1 | Leptolyngbya | B1, B6, S3, S7 | B4 |
| 102d-1 | Leptolyngbya | S2, S9, S13 | B4 |
| 96-2 | Leptolyngbya | B2, B6, S3, S6, S7, S8, S13 | No effect |
| 96-4 | Spirulina | S3, S4, S9 | B4, S2, S10 |
| Other marine isolates | |||
| Alg | Leptolyngbya | No effect | B2, B3, B4, B5, B9, P1, P2, S2, S4, S8, S12 |
| 10 | Leptolyngbya | No effect | B9, P2, S4 |
| 9-1 | Leptolyngbya | No effect | B5, B9, P1, S12 |
| HS 26 | Leptolyngbya | No effect | B5, S12 |
| 73-2 | Phormidium | B1, S2, S8, S9 | B1, B4, B5, P2, S1, S2, S4 |
| 63a-5 | Pseudanabaena | S6, S7 | B5, B10, P2, S4, S9, S10 |
| 63-1 | Pseudanabaena | S4, S5, S9 | No effect |
| 72-1 | Pseudanabaena | B3, S4 | B5, B9, B10, P2, S7, S9, S10 |
| 63a-1 | Synechococcus | B1, B2, S2, S4, S5, S9, S10 | B1, B2, B3, B5, B9, P1, P2, S2, S9 |
| 63a-3 | Synechococcus | B1, B2, S5, S9 | B5, B9, B10, S7, S9, S12 |
| Strain designation | GenBank closest relative | Accession No. of closest relative 1 | Strain abbreviation 2 |
|---|---|---|---|
| Strains from BBD | |||
| HS-216-1a | Vibrio harveyi | AY750576 | B1 |
| HS-216-3d | Bacillus megaterium | AJ17381 | B2 |
| HS-217-1a | Bacillus cereus | AY305275 | B3 |
| HS-217-1c | Photobacterium eurosenbergii | AJ842344 | B4 |
| HS-216-4f | Marinobacter sp. | AY196982 | B5 |
| HS-217-2g | Alteromonas sp. | AY626838 | B6 |
| HS-216-4g | Marinobacter aquaeolei | AJ000726 | B7 |
| HS-216-4i | Idiomarina sp. | AB167047 | B8 |
| HS-217-2d | Vibrio harveyi | AY750575 | B9 |
| HS-216-4a | Methylarcula sp. | AJ534208 | B10 |
| Known coral pathogens | |||
| Aurantimonas coralicida | N/A | P1 | |
| Serratia marcescens | N/A | P2 | |
| Vibrio shiloi | N/A | P3 | |
| Strains from healthy corals | |||
| 1-1 | Vibrio sp. | EU267634 | S1 |
| 1-2 | Alcanivorax sp. | EU781516 | S2 |
| 1-3 | Vibrionaceae | EF584057 | S3 |
| 1-7 | Vibrio sp. | EU267643 | S4 |
| 1-8a | Bacillus sp. | EU070391 | S5 |
| 1.8b | Bacillus sp. | FJ461465 | S6 |
| 1-9 | Vibrio harveyi | DQ995240 | S7 |
| 1-10 | Vibrio sp. | EU267643 | S8 |
| 1-11 | Vibrio sp. | FJ178079 | S9 |
| 1-12 | Vibrio sp. | EF100710 | S10 |
| 1-13 | Vibrio sp. | FJ457416 | S11 |
| 1-14 | Vibrio sp. | EU276991 | S12 |
| 1-16 | Vibrio sp. | EF584084 | S13 |
| Isolate | Closest relative | Similarity % | GenBank access. No. | Location/Source | Reference |
|---|---|---|---|---|---|
| BBD Isolates | |||||
| BBD 1991 | Geitlerinema 1 | 99 | DQ151461 | Florida Keys, Algae Reef, BBD on Montastraea annularis | [36] |
| HS 217 | Geitlerinema | 99 | EF110974 | LSI Bahamas, Horseshoe Reef, BBD on Siderastrea siderea | [37] |
| HS 223 | Geitlerinema | 99 | DQ680351 | LSI Bahamas, Horseshoe Reef, BBD on Siderastrea siderea | [37] |
| W-1 | Geitlerinema | 99 | EF154084 | Florida Keys, Watson’s Reef, BBD on Siderastrea siderea | [37] |
| FLK BBD1 | Leptolyngbya | 98 | EF110975 | Florida Keys, South Carysfort, BBD on Montastraea annularis | [37] |
| P2b-2 | Leptolyngbya | 98 | EF372581 | Philippines, BBD on Porites lutea | [37] |
| 102a-1 | Leptolyngbya | 97 | EU743966 | Florida Keys, BBD on Dendrogyra cylindrus | [33] |
| 102d-1 | Leptolyngbya 2 | 97 | EU743968 | Florida Keys, BBD on Montastraea annularis | [33] |
| 96-2 | Leptolyngbya 3,4 | N/A | N/A | Florida Keys, BBD on Montastraea annularis | This work |
| 96-4 | Spirulina | 93 | EU743969 | Florida Keys, BBD on Montastraea annularis | [33] |
| Other marine isolates | |||||
| Alg | Leptolyngbya 3,4 | N/A | N/A | Florida Keys, Algae Reef, mat on sediment | [33] |
| 10 | Leptolyngbya | 99 | FJ232377 | Florida Keys, NN Dry Rocks, mat on Montastraea cavernosa | [33] |
| 9-1 | Leptolyngbya 3,4 | N/A | N/A | Florida Keys, Horseshoe reef, mat on Colpophyllia natans | This work |
| HS26 | Leptolyngbya | N/A | FJ232376 | Florida Keys, Horseshoe reef, mat on coral away from BBD | [33] |
| 73-2 | Phormidium | 97 | EU196366 | Gulf of Mexico, plankton | [38] |
| 63a-5 | Pseudanabaena | 97 | FJ026734 | Florida Keys, mat on sediment | [33] |
| 63-1 | Pseudanabaena 2 | 98 | EU110976 | Florida Keys, mat on sediment | [38] |
| 72-1 | Pseudanabaena | 98 | EU196365 | Florida Keys, NN dry rocks, mat on Montastraea cavernosa | [38] |
| 63a-1 | Synechococcus 2 | 98 | EU743972 | Florida Keys, mat on sediment | [33] |
| 63a-3 | Synechococcus | 98 | EU743971 | Florida Keys, mat on sediment | [33] |
© 2011 by the authors; licensee MDPI, Basel, Switzerland This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Gantar, M.; Kaczmarsky, L.T.; Stanić, D.; Miller, A.W.; Richardson, L.L. Antibacterial Activity of Marine and Black Band Disease Cyanobacteria against Coral-Associated Bacteria. Mar. Drugs 2011, 9, 2089-2105. https://doi.org/10.3390/md9102089
Gantar M, Kaczmarsky LT, Stanić D, Miller AW, Richardson LL. Antibacterial Activity of Marine and Black Band Disease Cyanobacteria against Coral-Associated Bacteria. Marine Drugs. 2011; 9(10):2089-2105. https://doi.org/10.3390/md9102089
Chicago/Turabian StyleGantar, Miroslav, Longin T. Kaczmarsky, Dina Stanić, Aaron W. Miller, and Laurie L. Richardson. 2011. "Antibacterial Activity of Marine and Black Band Disease Cyanobacteria against Coral-Associated Bacteria" Marine Drugs 9, no. 10: 2089-2105. https://doi.org/10.3390/md9102089
APA StyleGantar, M., Kaczmarsky, L. T., Stanić, D., Miller, A. W., & Richardson, L. L. (2011). Antibacterial Activity of Marine and Black Band Disease Cyanobacteria against Coral-Associated Bacteria. Marine Drugs, 9(10), 2089-2105. https://doi.org/10.3390/md9102089




