Marine Sulfated Polysaccharides as Promising Antiviral Agents: A Comprehensive Report and Modeling Study Focusing on SARS CoV-2
Abstract
:1. Introduction
2. Results and Discussion
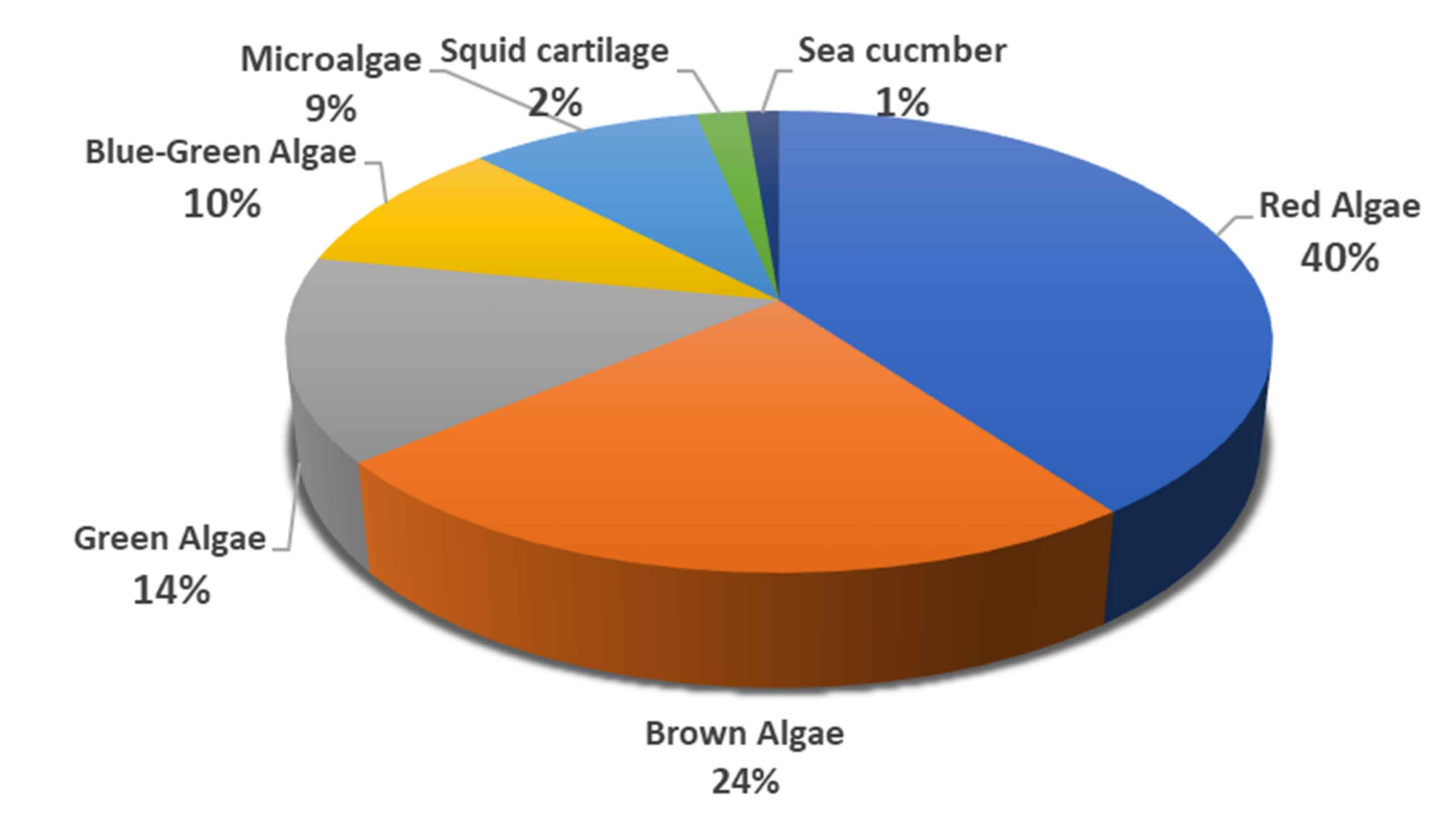
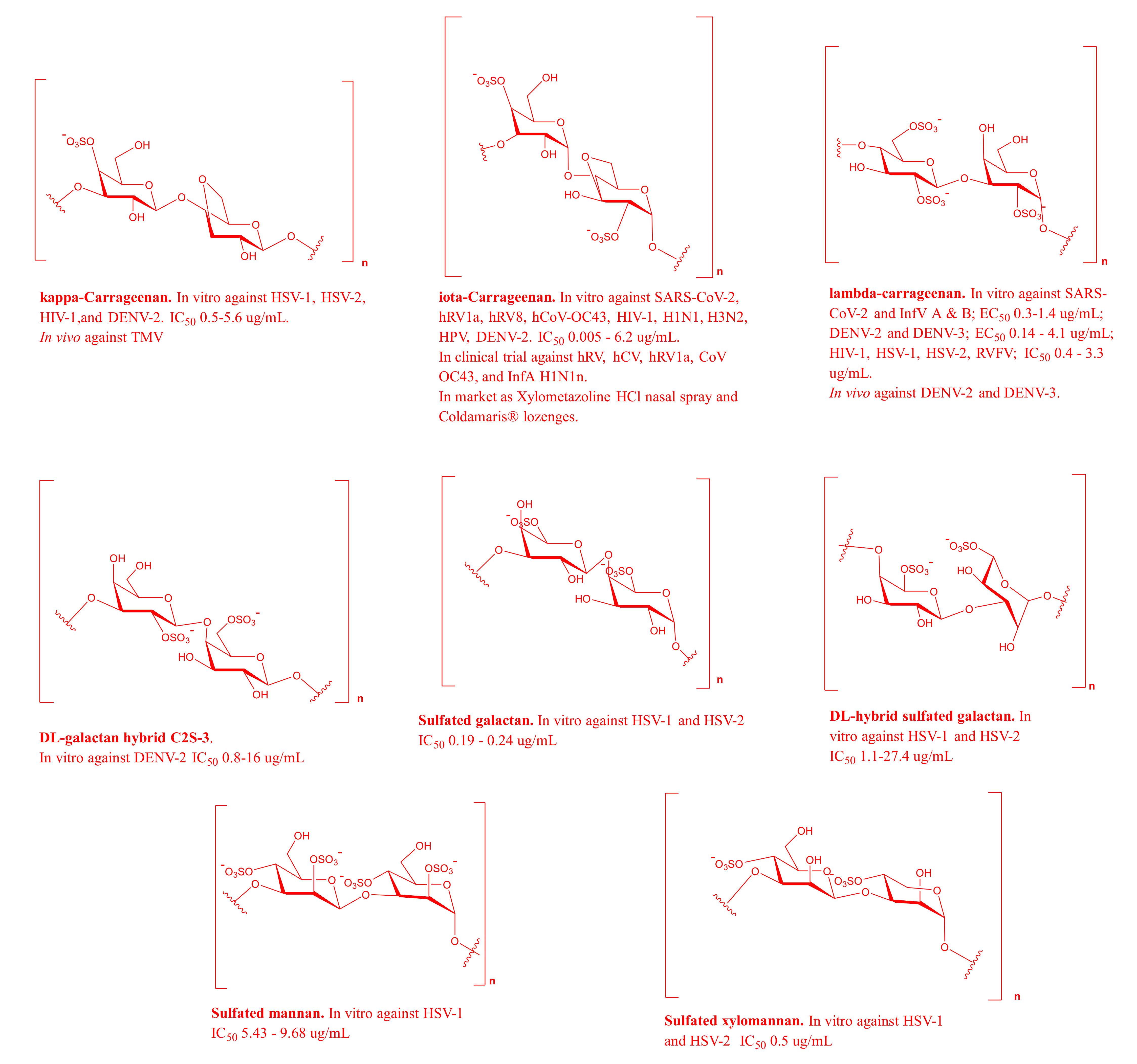
2.1. MSPs from Red Algae
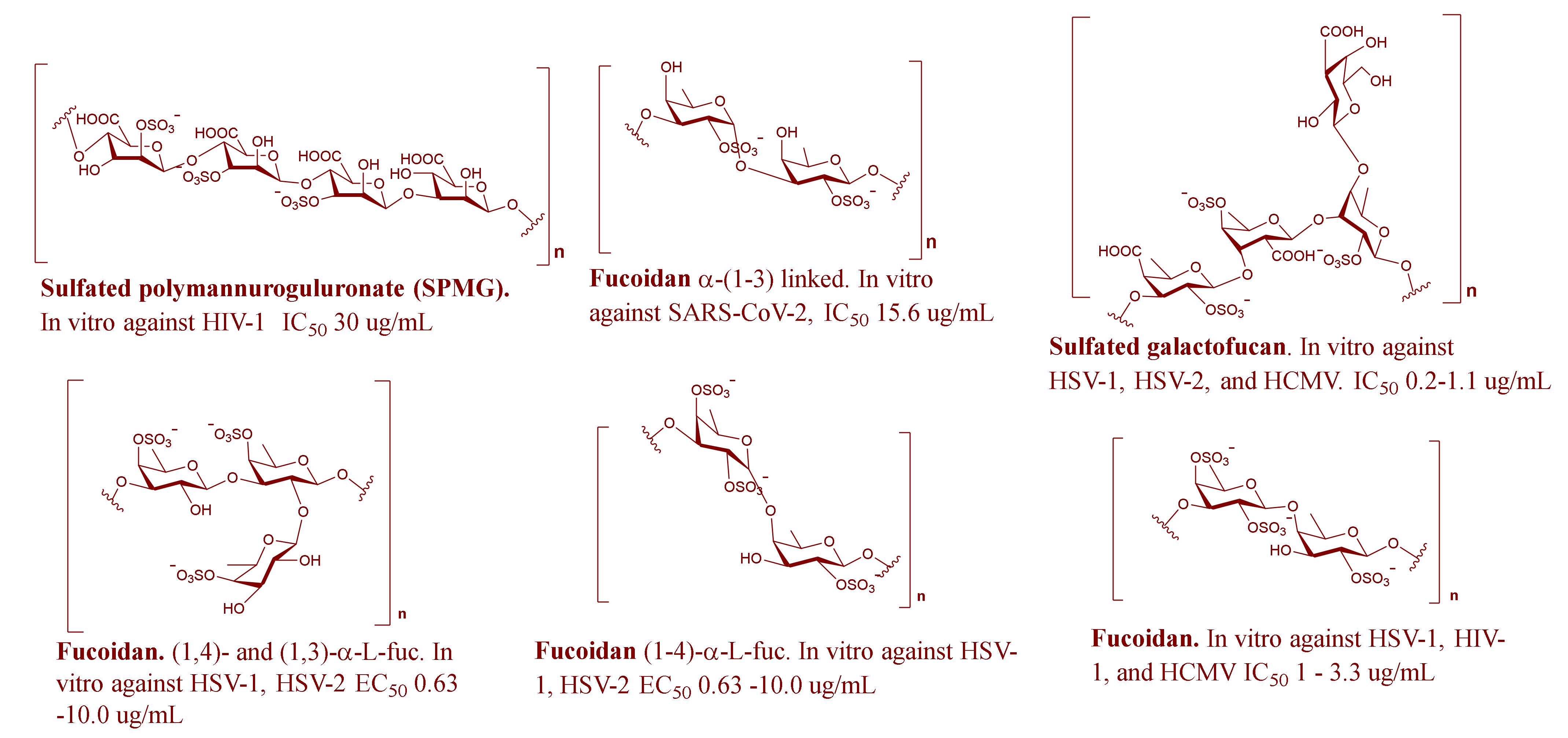
2.2. MSPs from Brown Algae
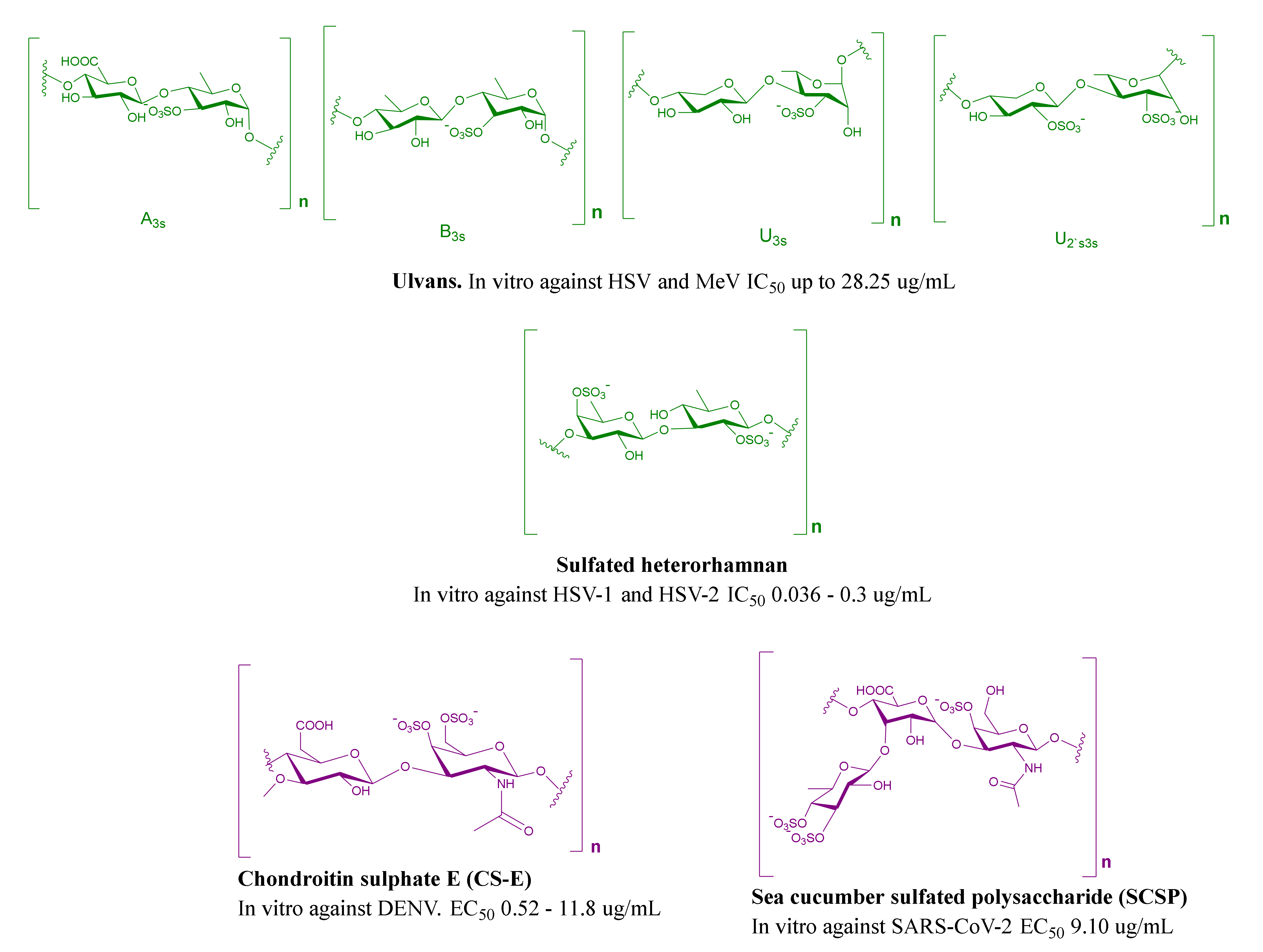
2.3. MSPs from Green Algae
2.4. MSPs from Miscellaneous Marine Sources
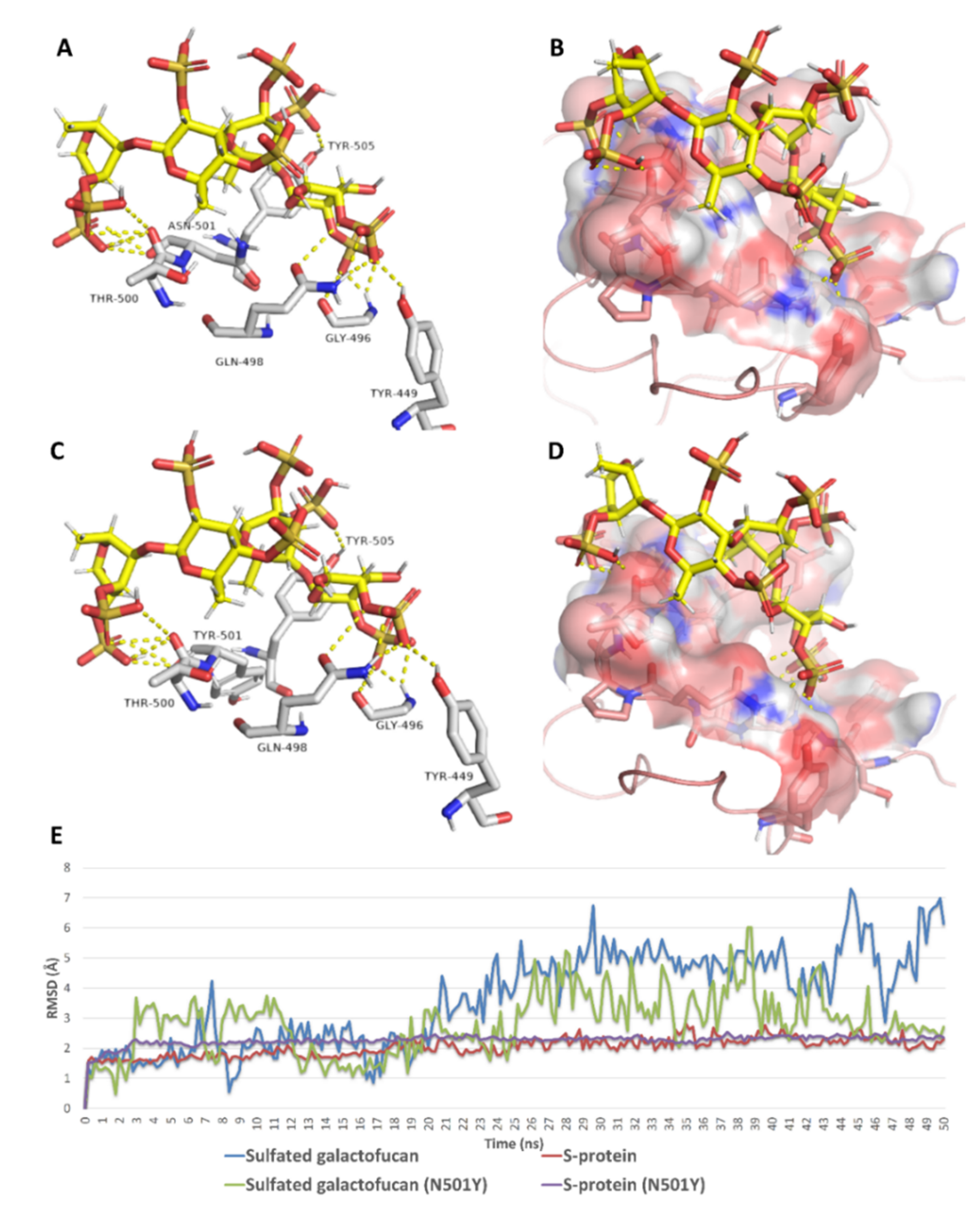
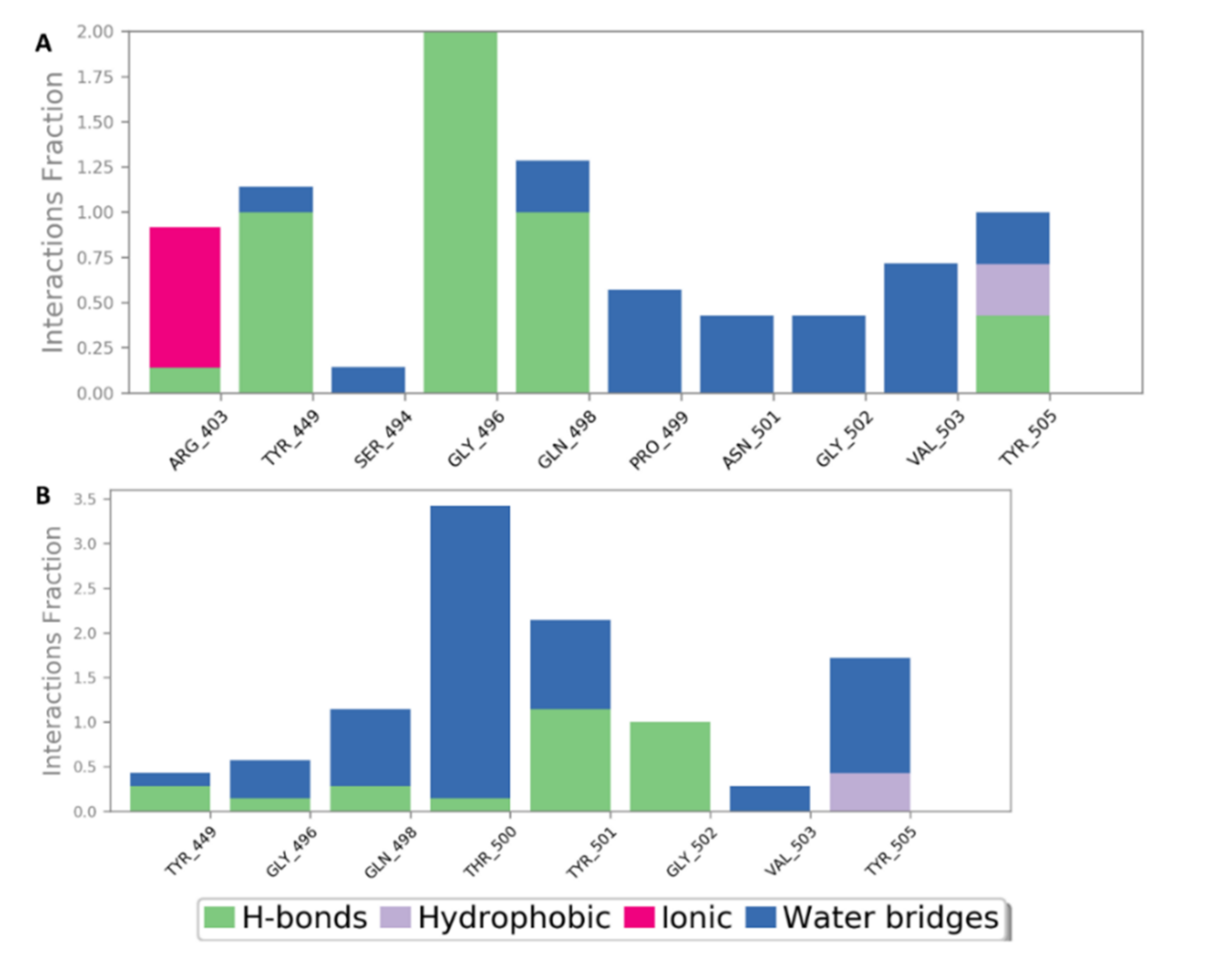
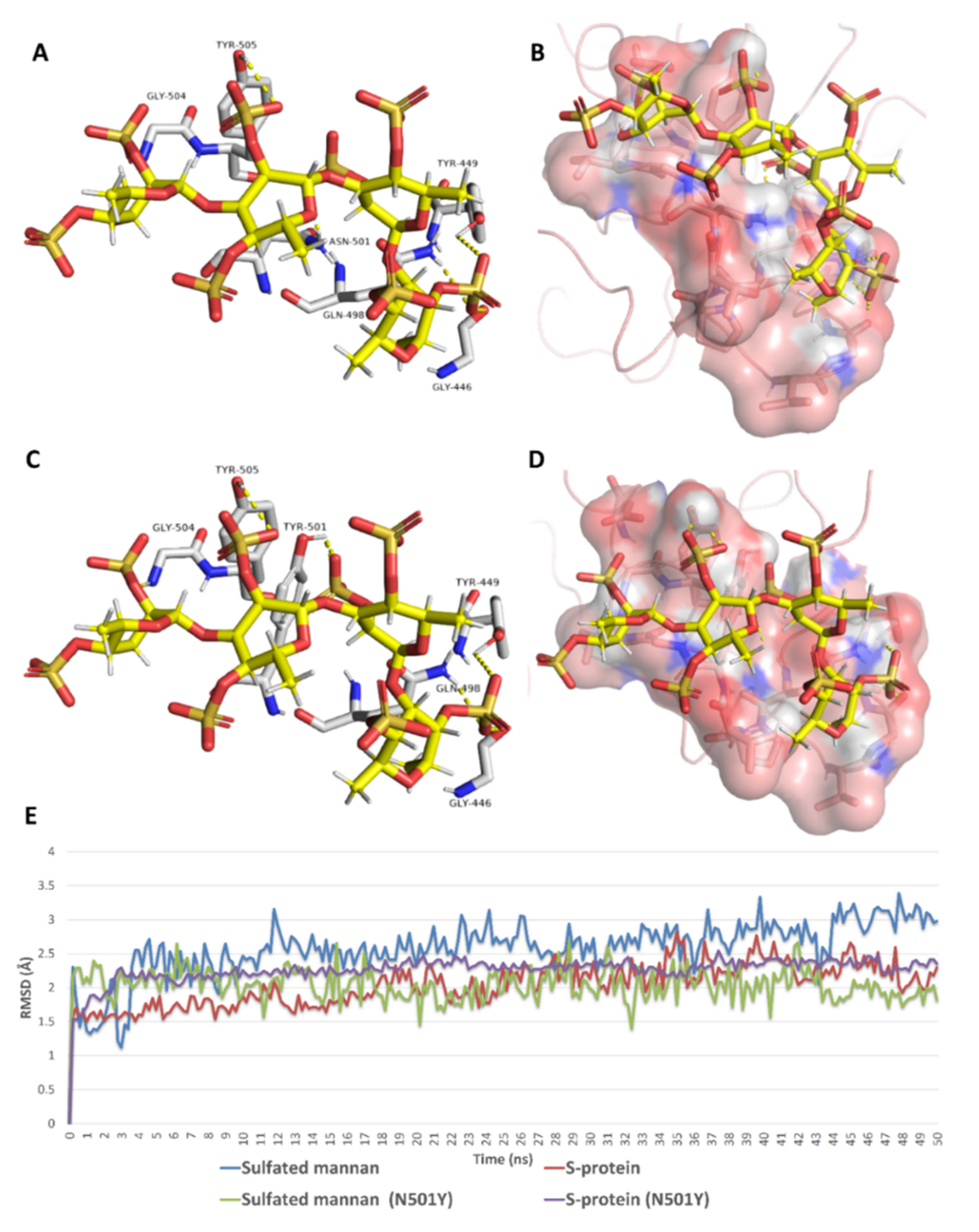
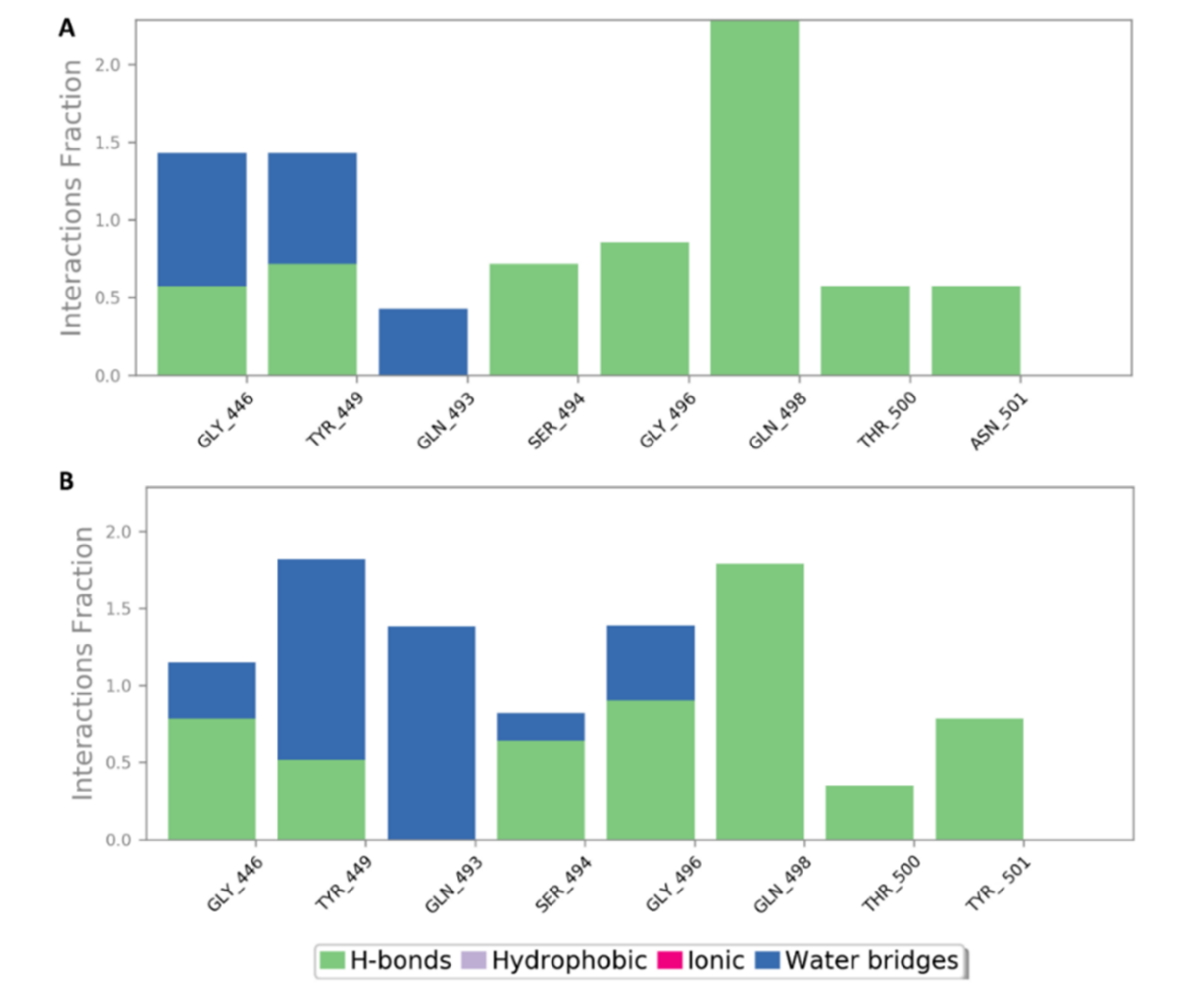
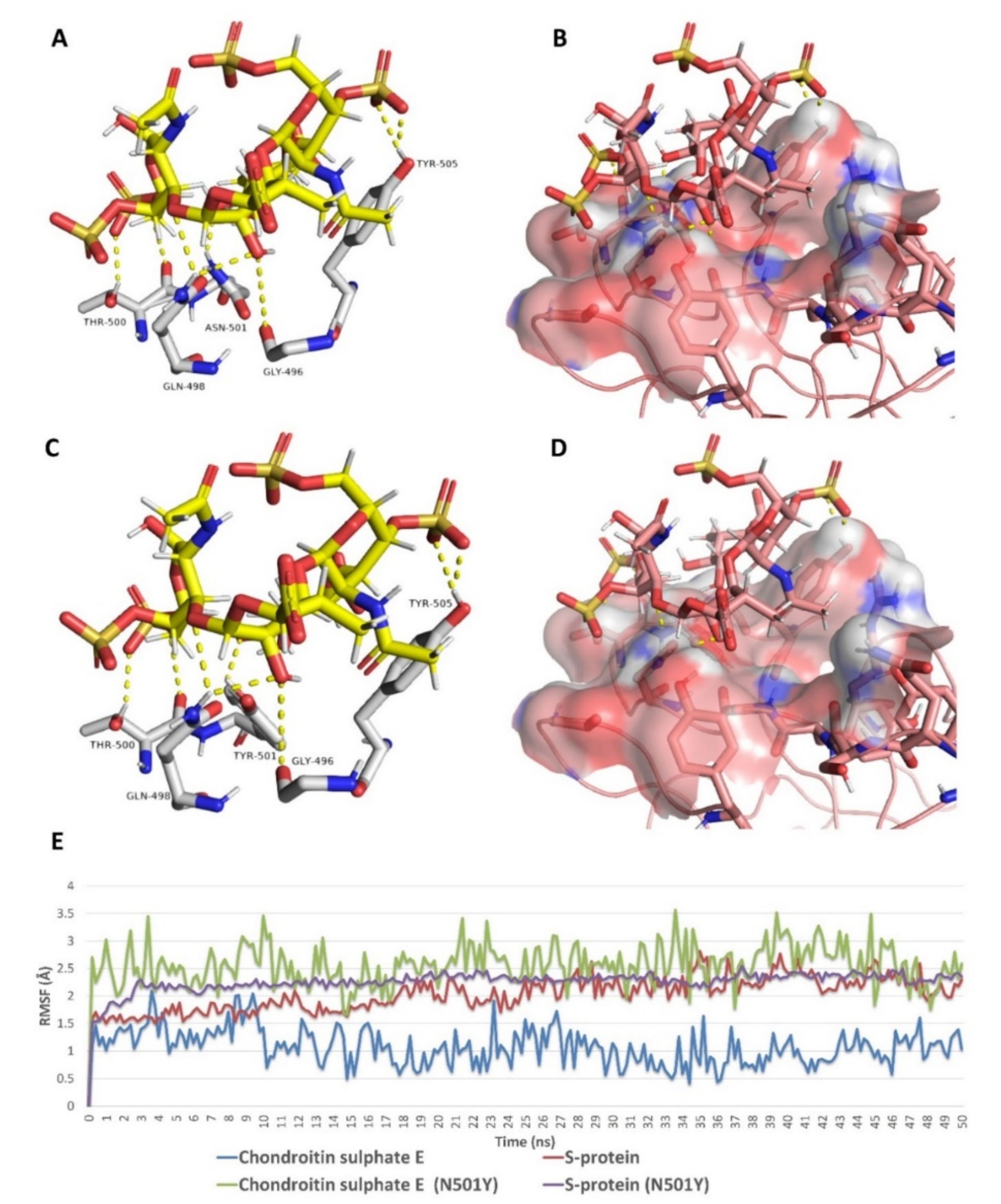
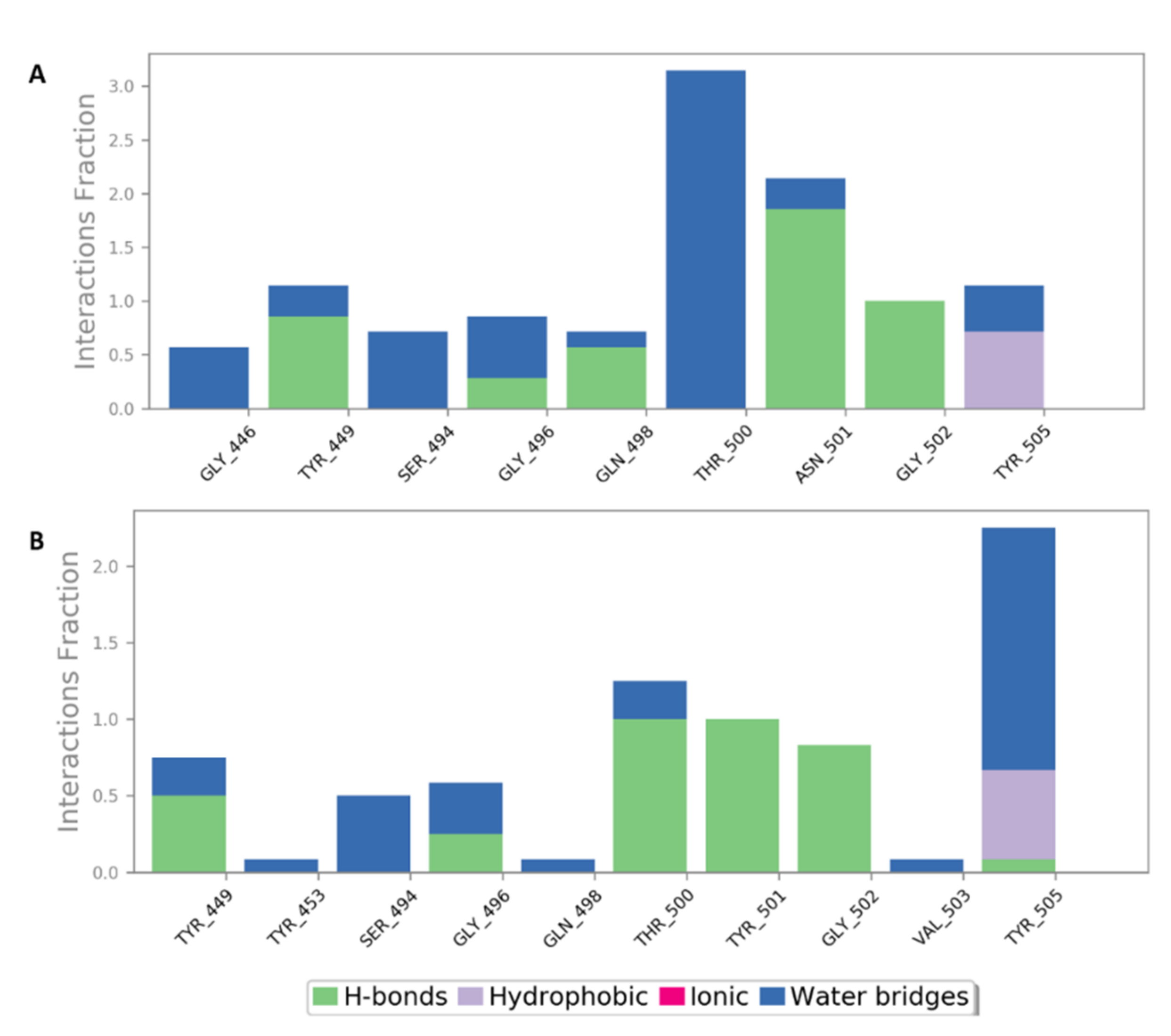
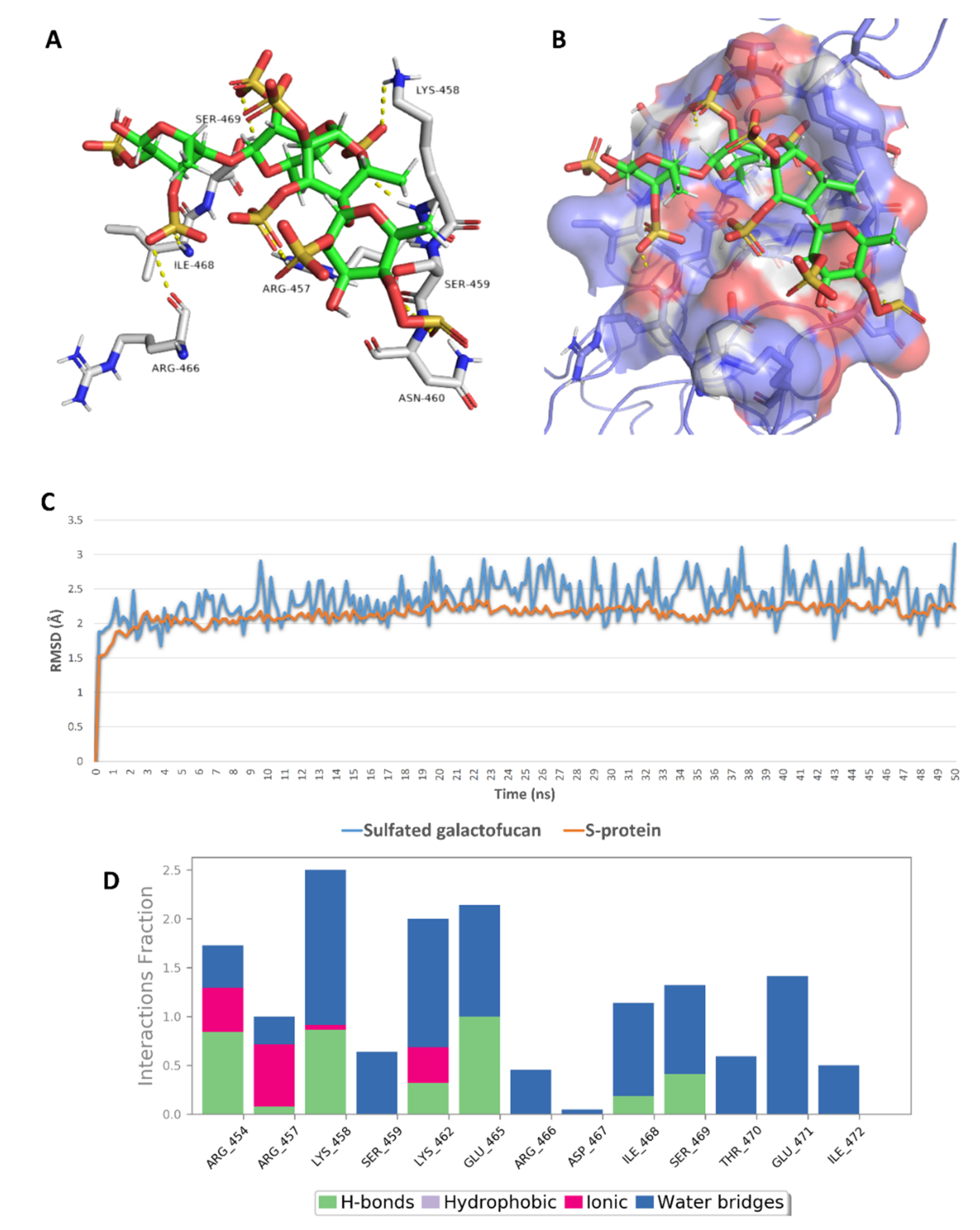
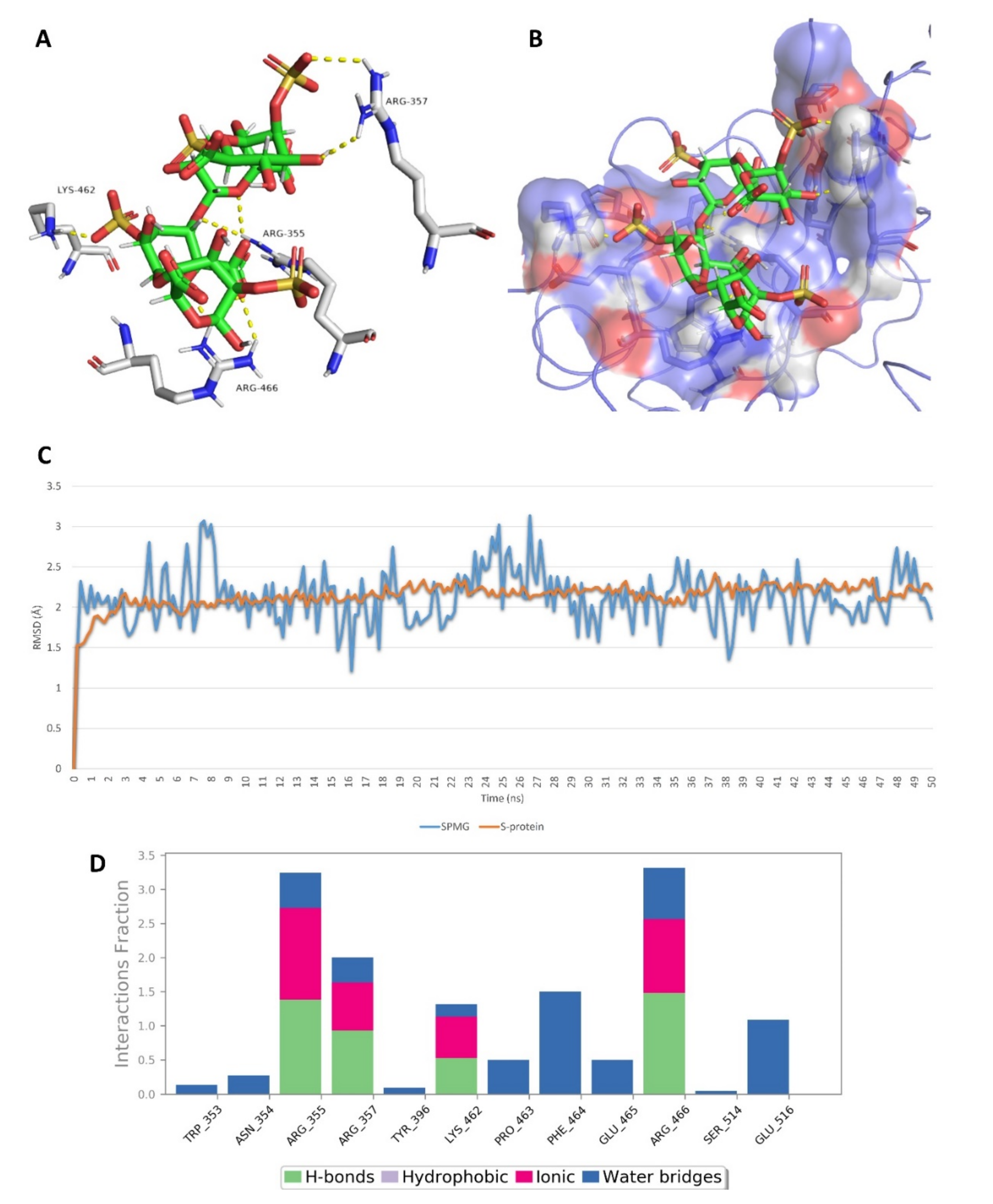
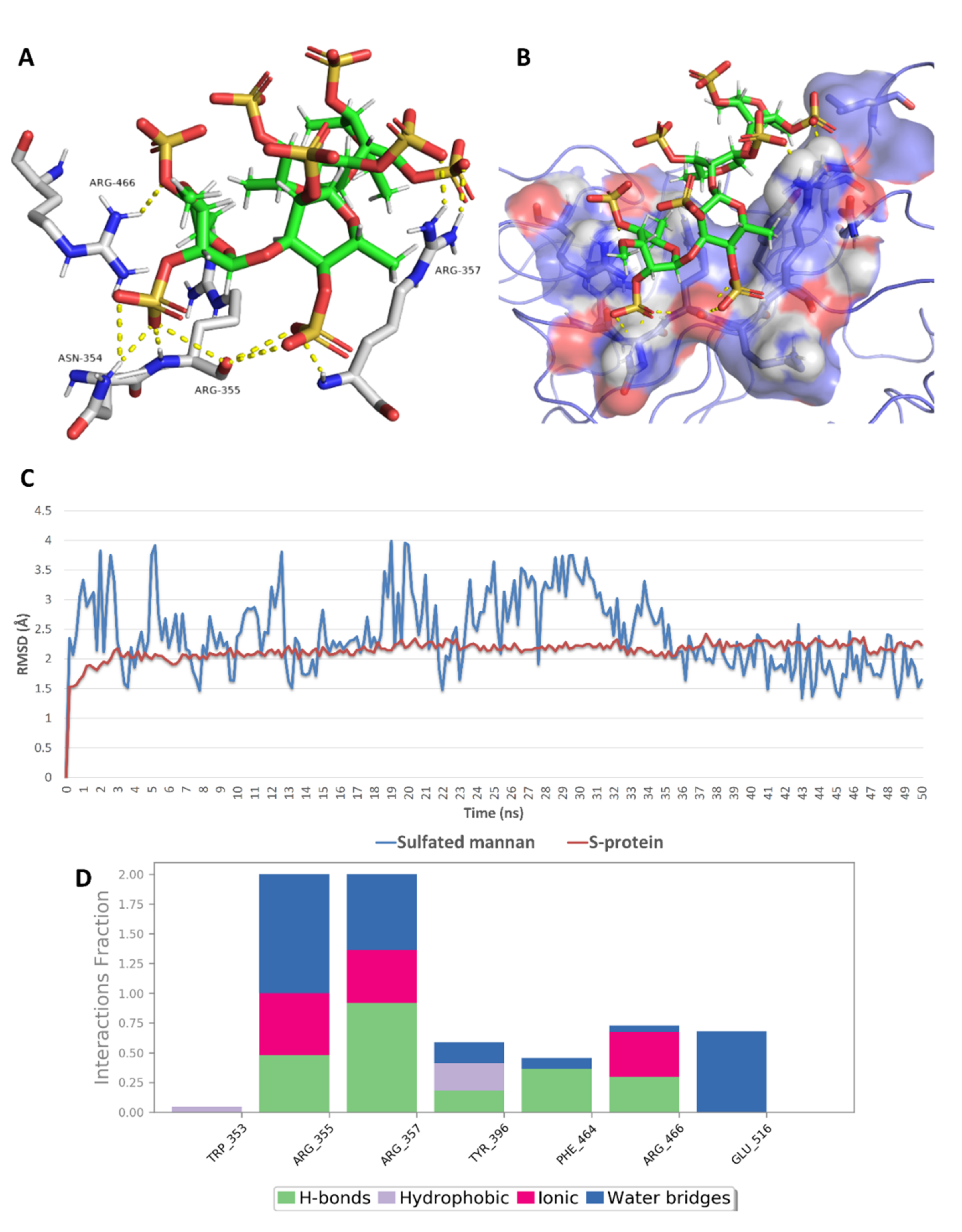
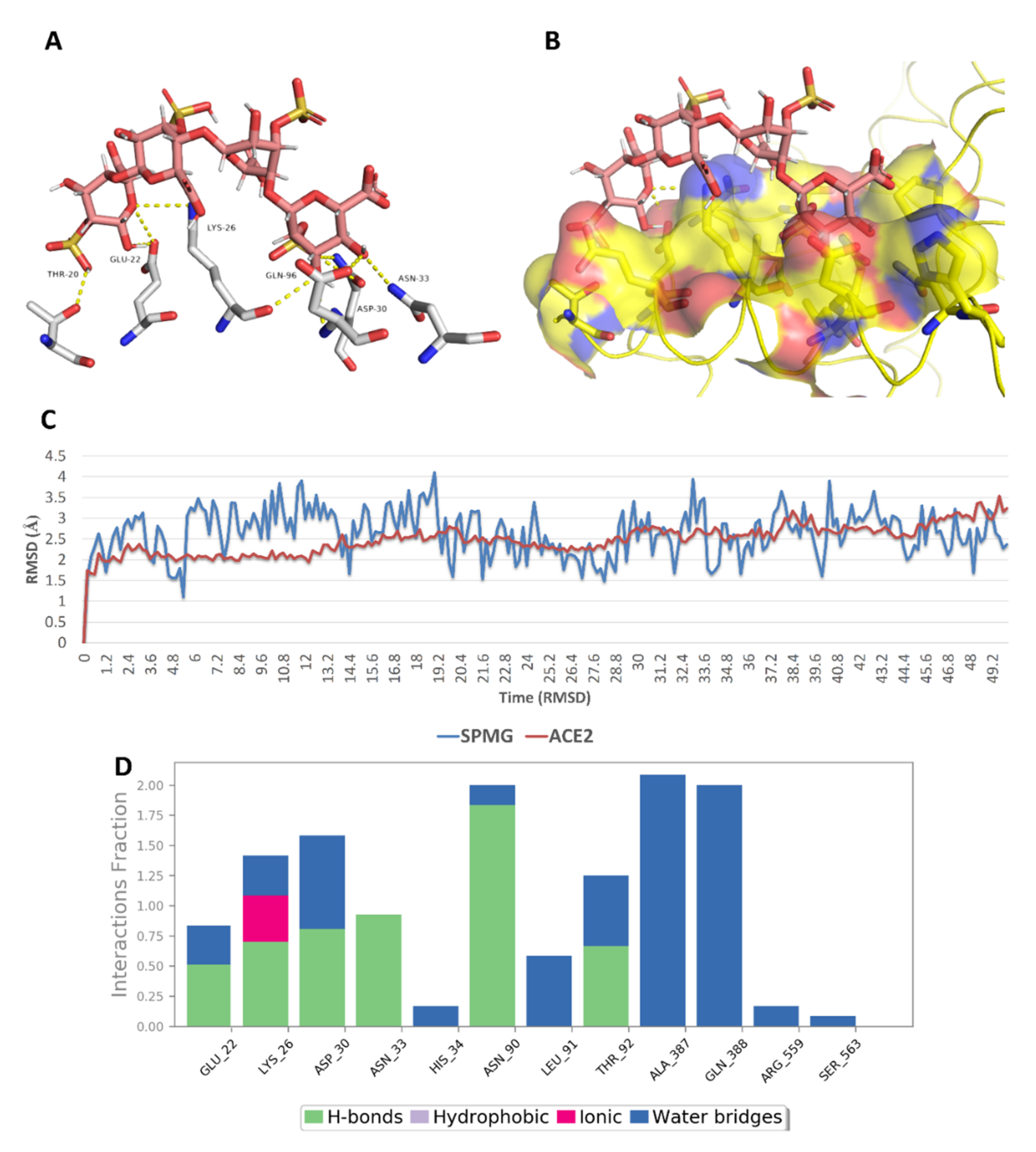
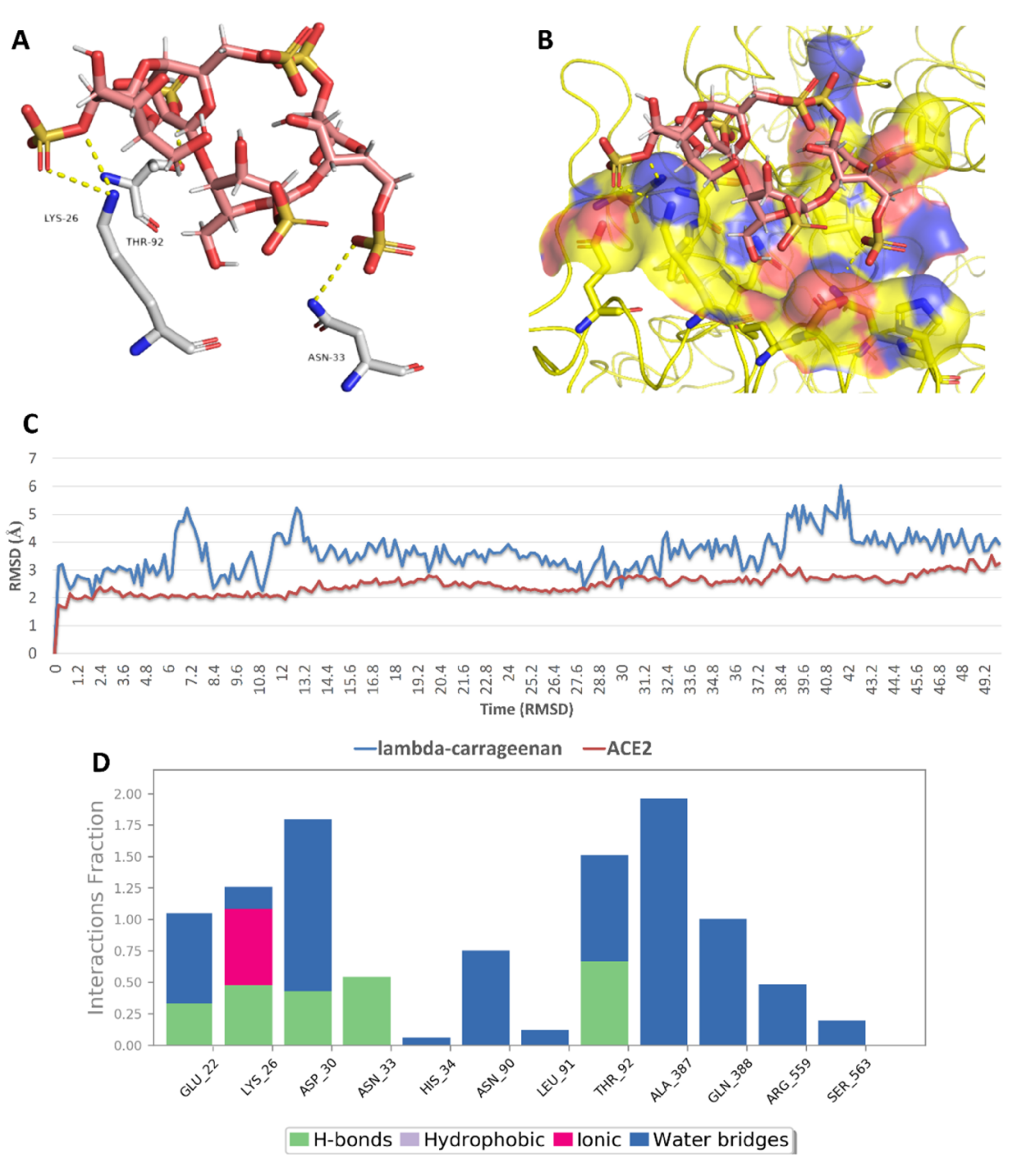
2.5. In Silico Investigation of MSPs against SARS CoV-2
2.6. Structure–Activity Relationship
3. Methodology
3.1. Databases Used
3.2. Molecular Modeling
3.2.1. Molecular Docking
3.2.2. Molecular Dynamic Simulation
3.2.3. Binding Free Energy Calculations
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Chen, L.; Huang, G. The antiviral activity of polysaccharides and their derivatives. Int. J. Biol. Macromol. 2018, 115, 77–82. [Google Scholar] [CrossRef] [PubMed]
- Lee, S. Chapter 4–Strategic Design of Delivery Systems for Nutraceuticals. In Nanotechnology Applications in Food; Oprea, A.E., Grumezescu, A.M., Eds.; Academic Press: Cambridge, MA, USA, 2017; pp. 65–86. [Google Scholar] [CrossRef]
- Ngo, D.-H.; Kim, S.-K. Sulfated polysaccharides as bioactive agents from marine algae. Int. J. Biol. Macromol. 2013, 62, 70–75. [Google Scholar] [CrossRef]
- Hans, N.; Malik, A.; Naik, S. Antiviral activity of sulfated polysaccharides from marine algae and its application in combating COVID-19: Mini review. Bioresour. Technol. Rep. 2021, 13, 100623. [Google Scholar] [CrossRef] [PubMed]
- Muthukumar, J.; Chidambaram, R.; Sukumaran, S. Sulfated polysaccharides and its commercial applications in food industries—A review. J. Food Sci. Technol. 2020. [Google Scholar] [CrossRef]
- Koenighofer, M.; Lion, T.; Bodenteich, A.; Prieschl-Grassauer, E.; Grassauer, A.; Unger, H.; Mueller, C.A.; Fazekas, T. Carrageenan nasal spray in virus confirmed common cold: Individual patient data analysis of two randomized controlled trials. Multidiscip. Respir. Med. 2014, 9, 57. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Graf, C.; Bernkop-Schnürch, A.; Egyed, A.; Koller, C.; Prieschl-Grassauer, E.; Morokutti-Kurz, M. Development of a nasal spray containing xylometazoline hydrochloride and iota-carrageenan for the symptomatic relief of nasal congestion caused by rhinitis and sinusitis. Int. J. Gen. Med. 2018, 11, 275. [Google Scholar] [CrossRef] [Green Version]
- Morokutti-Kurz, M.; Graf, C.; Prieschl-Grassauer, E. Amylmetacresol/2, 4-dichlorobenzyl alcohol, hexylresorcinol, or carrageenan lozenges as active treatments for sore throat. Int. J. Gen. Med. 2017, 10, 53. [Google Scholar] [CrossRef] [Green Version]
- Ekins, S.; Mestres, J.; Testa, B. In silico pharmacology for drug discovery: Methods for virtual ligand screening and profiling. Br. J. Pharmacol. 2007, 152, 9–20. [Google Scholar] [CrossRef] [Green Version]
- Sliwoski, G.; Kothiwale, S.; Meiler, J.; Lowe, E.W. Computational methods in drug discovery. Pharmacol. Rev. 2014, 66, 334–395. [Google Scholar] [CrossRef] [Green Version]
- Makhouri, F.R.; Ghasemi, J.B. In Silico Studies in Drug Research Against Neurodegenerative Diseases. Curr. Neuropharmacol. 2018, 16, 664–725. [Google Scholar] [CrossRef] [PubMed]
- Phillips, M.A. Has Molecular Docking Ever Brought Us a Medicine? IntechOpen: London, UK, 2018. [Google Scholar] [CrossRef] [Green Version]
- Fadda, E.; Woods, R.J. Molecular simulations of carbohydrates and protein–carbohydrate interactions: Motivation, issues and prospects. Drug Discov. Today 2010, 15, 596–609. [Google Scholar] [CrossRef] [PubMed]
- Hemmingson, J.A.; Falshaw, R.; Furneaux, R.H.; Thompson, K. Structure and Antiviral Activity of the Galactofucan Sulfates Extracted from UndariaPinnatifida (Phaeophyta). J. Appl. Phycol. 2006, 18, 185. [Google Scholar] [CrossRef]
- Vishchuk, O.S.; Ermakova, S.P.; Zvyagintseva, T.N. Sulfated polysaccharides from brown seaweeds Saccharina japonica and Undaria pinnatifida: Isolation, structural characteristics, and antitumor activity. Carbohydr. Res. 2011, 346, 2769–2776. [Google Scholar] [CrossRef]
- Mandal, P.; Mateu, C.G.; Chattopadhyay, K.; Pujol, C.A.; Damonte, E.B.; Ray, B. Structural Features and Antiviral Activity of Sulphated Fucans from the Brown Seaweed Cystoseira Indica. Antivir. Chem. Chemother. 2007, 18, 153–162. [Google Scholar] [CrossRef] [Green Version]
- Kwon, P.S.; Oh, H.; Kwon, S.-J.; Jin, W.; Zhang, F.; Fraser, K.; Hong, J.J.; Linhardt, R.J.; Dordick, J.S. Sulfated polysaccharides effectively inhibit SARS-CoV-2 in vitro. Cell Discov. 2020, 6, 50. [Google Scholar] [CrossRef]
- Song, S.; Peng, H.; Wang, Q.; Liu, Z.; Dong, X.; Wen, C.; Ai, C.; Zhang, Y.; Wang, Z.; Zhu, B. Inhibitory activities of marine sulfated polysaccharides against SARS-CoV-2. Food Funct. 2020, 11, 7415–7420. [Google Scholar] [CrossRef]
- Usoltseva, R.V.; Anastyuk, S.D.; Ishina, I.A.; Isakov, V.V.; Zvyagintseva, T.N.; Thinh, P.D.; Zadorozhny, P.A.; Dmitrenok, P.S.; Ermakova, S.P. Structural characteristics and anticancer activity in vitro of fucoidan from brown alga Padina boryana. Carbohydr. Polym. 2018, 184, 260–268. [Google Scholar] [CrossRef] [PubMed]
- Miao, B.; Geng, M.; Li, J.; Li, F.; Chen, H.; Guan, H.; Ding, J. Sulfated polymannuroguluronate, a novel anti-acquired immune deficiency syndrome (AIDS) drug candidate, targeting CD4 in lymphocytes. Biochem. Pharmacol. 2004, 68, 641–649. [Google Scholar] [CrossRef]
- Meiyu, G.; Fuchuan, L.; Xianliang, X.; Jing, L.; Zuowei, Y.; Huashi, G. The potential molecular targets of marine sulfated polymannuroguluronate interfering with HIV-1 entry: Interaction between SPMG and HIV-1 rgp120 and CD4 molecule. Antivir. Res. 2003, 59, 127–135. [Google Scholar] [CrossRef]
- Liu, H.; Geng, M.; Xin, X.; Li, F.; Zhang, Z.; Li, J.; Ding, J. Multiple and multivalent interactions of novel anti-AIDS drug candidates, sulfated polymannuronate (SPMG)-derived oligosaccharides, with gp120 and their anti-HIV activities. Glycobiology 2004, 15, 501–510. [Google Scholar] [CrossRef]
- Ponce, N.M.A.; Pujol, C.A.; Damonte, E.B.; Flores, M.L.; Stortz, C.A. Fucoidans from the brown seaweed Adenocystis utricularis: Extraction methods, antiviral activity and structural studies. Carbohydr. Res. 2003, 338, 153–165. [Google Scholar] [CrossRef]
- Ale, M.T.; Mikkelsen, J.D.; Meyer, A.S. Important Determinants for Fucoidan Bioactivity: A Critical Review of Structure-Function Relations and Extraction Methods for Fucose-Containing Sulfated Polysaccharides from Brown Seaweeds. Mar. Drugs 2011, 9, 2106–2130. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Adhikari, U.; Mateu, C.G.; Chattopadhyay, K.; Pujol, C.A.; Damonte, E.B.; Ray, B. Structure and antiviral activity of sulfated fucans from Stoechospermum marginatum. Phytochemistry 2006, 67, 2474–2482. [Google Scholar] [CrossRef] [PubMed]
- Shen, P.; Yin, Z.; Qu, G.; Wang, C. 11-fucoidan and its health benefits. In Bioactive Seaweeds for Food Applications; Qin, Y., Ed.; Academic Press: Cambridge, MA, USA, 2018; pp. 223–238. [Google Scholar]
- Hoshino, T.; Hayashi, T.; Hayashi, K.; Hamada, J.; Lee, J.-B.; Sankawa, U. An antivirally active sulfated polysaccharide from Sargassum horneri (TURNER) C. AGARDH. Biol. Pharm. Bull. 1998, 21, 730–734. [Google Scholar] [CrossRef] [Green Version]
- Wang, W.; Wu, J.; Zhang, X.; Hao, C.; Zhao, X.; Jiao, G.; Shan, X.; Tai, W.; Yu, G. Inhibition of Influenza A Virus Infection by Fucoidan Targeting Viral Neuraminidase and Cellular EGFR Pathway. Sci. Rep. 2017, 7, 40760. [Google Scholar] [CrossRef] [PubMed]
- Zhu, W.; Ooi, V.E.C.; Chan, P.K.S.; Ang, P.O., Jr. Isolation and characterization of a sulfated polysaccharide from the brown alga Sargassum patens and determination of its anti-herpes activity. Biochem. Cell Biol. 2003, 81, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Feldman, S.C.; Reynaldi, S.; Stortz, C.A.; Cerezo, A.S.; Damonte, E.B. Antiviral properties of fucoidan fractions from Leathesia difformis. Phytomedicine 1999, 6, 335–340. [Google Scholar] [CrossRef]
- Krylova, N.V.; Ermakova, S.P.; Lavrov, V.F.; Leneva, I.A.; Kompanets, G.G.; Iunikhina, O.V.; Nosik, M.N.; Ebralidze, L.K.; Falynskova, I.N.; Silchenko, A.S.; et al. The Comparative Analysis of Antiviral Activity of Native and Modified Fucoidans from Brown Algae Fucus evanescens In Vitro and In Vivo. Mar. Drugs 2020, 18, 224. [Google Scholar] [CrossRef] [Green Version]
- Lee, J.-B.; Hayashi, K.; Hashimoto, M.; Nakano, T.; Hayashi, T. Novel Antiviral Fucoidan from Sporophyll of Undaria pinnatifida (Mekabu). Chem. Pharm. Bull. 2004, 52, 1091–1094. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bandyopadhyay, S.S.; Navid, M.H.; Ghosh, T.; Schnitzler, P.; Ray, B. Structural features and in vitro antiviral activities of sulfated polysaccharides from Sphacelaria indica. Phytochemistry 2011, 72, 276–283. [Google Scholar] [CrossRef]
- Thuy, T.T.T.; Ly, B.M.; Van, T.T.T.; Van Quang, N.; Tu, H.C.; Zheng, Y.; Seguin-Devaux, C.; Mi, B.; Ai, U. Anti-HIV activity of fucoidans from three brown seaweed species. Carbohydr. Polym. 2015, 115, 122–128. [Google Scholar] [CrossRef]
- Elizondo-Gonzalez, R.; Cruz-Suarez, L.E.; Ricque-Marie, D.; Mendoza-Gamboa, E.; Rodriguez-Padilla, C.; Trejo-Avila, L.M. In vitro characterization of the antiviral activity of fucoidan from Cladosiphon okamuranus against Newcastle Disease Virus. Virol. J. 2012, 9, 307. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hidari, K.I.P.J.; Takahashi, N.; Arihara, M.; Nagaoka, M.; Morita, K.; Suzuki, T. Structure and anti-dengue virus activity of sulfated polysaccharide from a marine alga. Biochem. Biophys. Res. Commun. 2008, 376, 91–95. [Google Scholar] [CrossRef] [PubMed]
- Richards, C.; Williams, N.A.; Fitton, J.H.; Stringer, D.N.; Karpiniec, S.S.; Park, A.Y. Oral Fucoidan Attenuates Lung Pathology and Clinical Signs in a Severe Influenza a Mouse Model. Mar. Drugs 2020, 18, 246. [Google Scholar] [CrossRef]
- Synytsya, A.; Bleha, R.; Synytsya, A.; Pohl, R.; Hayashi, K.; Yoshinaga, K.; Nakano, T.; Hayashi, T. Mekabu fucoidan: Structural complexity and defensive effects against avian influenza A viruses. Carbohydr. Polym. 2014, 111, 633–644. [Google Scholar] [CrossRef]
- Talarico, L.B.; Zibetti, R.G.M.; Faria, P.C.S.; Scolaro, L.A.; Duarte, M.E.R.; Noseda, M.D.; Pujol, C.A.; Damonte, E.B. Anti-herpes simplex virus activity of sulfated galactans from the red seaweeds Gymnogongrus griffithsiae and Cryptonemia crenulata. Int. J. Biol. Macromol. 2004, 34, 63–71. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, T.; Pujol, C.A.; Damonte, E.B.; Sinha, S.; Ray, B. Sulfated Xylomannans from the Red Seaweed Sebdenia Polydactyla: Structural Features, Chemical Modification and Antiviral Activity. Antivir. Chem. Chemother. 2009, 19, 235–242. [Google Scholar] [CrossRef] [Green Version]
- Morokutti-Kurz, M.; Fröba, M.; Graf, P.; Große, M.; Grassauer, A.; Auth, J.; Schubert, U.; Prieschl-Grassauer, E. Iota-carrageenan neutralizes SARS-CoV-2 and inhibits viral replication in vitro. PLoS ONE 2021, 16, e0237480. [Google Scholar] [CrossRef] [PubMed]
- Hongu, T.; Phillips, G.O.; Takigami, M. 7-Polymer fibers for health and nutrition. In New Millennium Fibers; Hongu, T., Phillips, G.O., Takigami, M., Eds.; Woodhead Publishing: Sawston, UK, 2005; pp. 218–246. [Google Scholar]
- Bansal, S.; Jonsson, C.B.; Taylor, S.L.; Figueroa, J.M.; Dugour, A.V.; Palacios, C.; César Vega, J. Iota-carrageenan and Xylitol inhibit SARS-CoV-2 in cell culture. bioRxiv 2020. [Google Scholar] [CrossRef]
- Jang, Y.; Shin, H.; Lee, M.K.; Kwon, O.S.; Shin, J.S.; Kim, Y.; Kim, M. Antiviral activity of lambda-carrageenan against influenza viruses in mice and severe acute respiratory syndrome coronavirus 2 in vitro. bioRxiv 2020. [Google Scholar] [CrossRef]
- Carlucci, M.J.; Ciancia, M.; Matulewicz, M.C.; Cerezo, A.S.; Damonte, E.B. Antiherpetic activity and mode of action of natural carrageenans of diverse structural types. Antivir. Res. 1999, 43, 93–102. [Google Scholar] [CrossRef]
- Chattopadhyay, K.; Ghosh, T.; Pujol, C.A.; Carlucci, M.J.; Damonte, E.B.; Ray, B. Polysaccharides from Gracilaria corticata: Sulfation, chemical characterization and anti-HSV activities. Int. J. Biol. Macromol. 2008, 43, 346–351. [Google Scholar] [CrossRef] [PubMed]
- Pujol, C.A.; Estevez, J.M.; Carlucci, M.J.; Ciancia, M.; Cerezo, A.S.; Damonte, E.B. Novel DL-Galactan Hybrids from the Red Seaweed Gymnogongrus Torulosus are Potent Inhibitors of Herpes Simplex Virus and Dengue Virus. Antivir. Chem. Chemother. 2002, 13, 83–89. [Google Scholar] [CrossRef] [Green Version]
- Kang, H.-K.; Seo, C.; Park, Y. The Effects of Marine Carbohydrates and Glycosylated Compounds on Human Health. Int. J. Mol. Sci. 2015, 16, 6018–6056. [Google Scholar] [CrossRef] [Green Version]
- Talarico, L.B.; Pujol, C.A.; Zibetti, R.G.M.; Faría, P.C.S.; Noseda, M.D.; Duarte, M.E.R.; Damonte, E.B. The antiviral activity of sulfated polysaccharides against dengue virus is dependent on virus serotype and host cell. Antivir. Res. 2005, 66, 103–110. [Google Scholar] [CrossRef] [PubMed]
- Cáceres, P.J.; Carlucci, M.a.J.; Damonte, E.B.; Matsuhiro, B.; Zúñiga, E.A. Carrageenans from chilean samples of Stenogramme interrupta (Phyllophoraceae): Structural analysis and biological activity. Phytochemistry 2000, 53, 81–86. [Google Scholar] [CrossRef]
- Mazumder, S.; Ghosal, P.K.; Pujol, C.A.; Carlucci, M.a.J.; Damonte, E.B.; Ray, B. Isolation, chemical investigation and antiviral activity of polysaccharides from Gracilaria corticata (Gracilariaceae, Rhodophyta). Int. J. Biol. Macromol. 2002, 31, 87–95. [Google Scholar] [CrossRef]
- Pereira, M.G.; Benevides, N.M.B.; Melo, M.R.S.; Valente, A.P.; Melo, F.R.; Mourão, P.A.S. Structure and anticoagulant activity of a sulfated galactan from the red alga, Gelidium crinale. Is there a specific structural requirement for the anticoagulant action? Carbohydr. Res. 2005, 340, 2015–2023. [Google Scholar] [CrossRef]
- Witvrouw, M.; Este, J.A.; Mateu, M.Q.; Reymen, D.; Andrei, G.; Snoeck, R.; Ikeda, S.; Pauwels, R.; Bianchini, N.V.; Desmyter, J.; et al. Activity of a Sulfated Polysaccharide Extracted from the Red Seaweed Aghardhiella Tenera against Human Immunodeficiency Virus and Other Enveloped Viruses. Antivir. Chem. Chemother. 1994, 5, 297–303. [Google Scholar] [CrossRef] [Green Version]
- Pérez Recalde, M.; Noseda, M.D.; Pujol, C.A.; Carlucci, M.J.; Matulewicz, M.C. Sulfated mannans from the red seaweed Nemalion helminthoides of the South Atlantic. Phytochemistry 2009, 70, 1062–1068. [Google Scholar] [CrossRef]
- Kolender, A.A.; Pujol, C.A.; Damonte, E.B.; Matulewicz, M.C.; Cerezo, A.S. The system of sulfated α-(1→ 3)-linked D-mannans from the red seaweed Nothogenia fastigiata: Structures, antiherpetic and anticoagulant properties. Carbohydr. Res. 1997, 304, 53–60. [Google Scholar] [CrossRef]
- Erra-Balsells, R.; Kolender, A.A.; Matulewicz, M.a.C.; Nonami, H.; Cerezo, A.S. Matrix-assisted ultraviolet laser-desorption ionization time-of-flight mass spectrometry of sulfated mannans from the red seaweed Nothogenia fastigiata. Carbohydr. Res. 2000, 329, 157–167. [Google Scholar] [CrossRef]
- Mandal, P.; Pujol, C.A.; Carlucci, M.J.; Chattopadhyay, K.; Damonte, E.B.; Ray, B. Anti-herpetic activity of a sulfated xylomannan from Scinaia hatei. Phytochemistry 2008, 69, 2193–2199. [Google Scholar] [CrossRef]
- Damonte, E.B.; Matulewicz, M.C.; Cerezob, A.S.; Coto, C.E. Herpes simplex Virus-Inhibitory Sulfated Xylogalactans from the Red Seaweed Nothogenia fastigiata. Chemotherapy 1996, 42, 57–64. [Google Scholar] [CrossRef]
- Navarro, D.A.; Stortz, C.A. The system of xylogalactans from the red seaweed Jania rubens (Corallinales, Rhodophyta). Carbohydr. Res. 2008, 343, 2613–2622. [Google Scholar] [CrossRef]
- Talarico, L.B.; Damonte, E.B. Interference in dengue virus adsorption and uncoating by carrageenans. Virology 2007, 363, 473–485. [Google Scholar] [CrossRef] [Green Version]
- Hemilä, H.; Chalker, E. Carrageenan nasal spray may double the rate of recovery from coronavirus and influenza virus infections: Re-analysis of randomized trial data. Res. Sq. 2020. [Google Scholar] [CrossRef]
- Leibbrandt, A.; Meier, C.; König-Schuster, M.; Weinmüllner, R.; Kalthoff, D.; Pflugfelder, B.; Graf, P.; Frank-Gehrke, B.; Beer, M.; Fazekas, T. Iota-carrageenan is a potent inhibitor of influenza A virus infection. PLoS ONE 2010, 5, e14320. [Google Scholar] [CrossRef] [PubMed]
- Grassauer, A.; Weinmuellner, R.; Meier, C.; Pretsch, A.; Prieschl-Grassauer, E.; Unger, H. Iota-Carrageenan is a potent inhibitor of rhinovirus infection. Virol. J. 2008, 5, 107. [Google Scholar] [CrossRef] [Green Version]
- Buck, C.B.; Thompson, C.D.; Roberts, J.N.; Müller, M.; Lowy, D.R.; Schiller, J.T. Carrageenan is a potent inhibitor of papillomavirus infection. PLoS Pathog 2006, 2, e69. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gomaa, H.H.; Elshoubaky, G.A. Antiviral activity of sulfated polysaccharides carrageenan from some marine seaweeds. Int. J. Curr. Pharm. Rev. Res. 2016, 7, 34–42. [Google Scholar]
- Malagoli, B.G.; Cardozo, F.T.G.S.; Gomes, J.H.S.; Ferraz, V.P.; Simões, C.M.O.; Braga, F.C. Chemical characterization and antiherpes activity of sulfated polysaccharides from Lithothamnion muelleri. Int. J. Biol. Macromol. 2014, 66, 332–337. [Google Scholar] [CrossRef] [PubMed]
- Mandal, P.; Pujol, C.A.; Damonte, E.B.; Ghosh, T.; Ray, B. Xylans from Scinaia hatei: Structural features, sulfation and anti-HSV activity. Int. J. Biol. Macromol. 2010, 46, 173–178. [Google Scholar] [CrossRef] [PubMed]
- Nor Rashid, N.; Yusof, R.; Rothan, H.A. Antiviral and virucidal activities of sulphated polysaccharides against Japanese encephalitis virus. Trop. Biomed. 2020, 37, 713–721. [Google Scholar] [CrossRef] [PubMed]
- Reunov, A.; Nagorskaya, V.; Lapshina, L.; Yermak, I.; Barabanova, A. Effect of κ/ß-Carrageenan from red alga Tichocarpus crinitus (Tichocarpaceae) on infection of detached tobacco leaves with tobacco mosaic virus. J. Plant Dis. Prot. 2004, 111, 165–172. [Google Scholar] [CrossRef]
- Piccini, L.E.; Carro, A.C.; Quintana, V.M.; Damonte, E.B. Antibody-independent and dependent infection of human myeloid cells with dengue virus is inhibited by carrageenan. Virus Res. 2020, 290, 198150. [Google Scholar] [CrossRef] [PubMed]
- De S.F-Ticher, P.C.; Talarico, L.B.; Noseda, M.D.; Pita, B.; Guimarães, S.M.; Damonte, E.B.; Duarte, M.E.R. Chemical structure and antiviral activity of carrageenans from Meristiella gelidium against herpes simplex and dengue virus. Carbohydr. Polym. 2006, 63, 459–465. [Google Scholar] [CrossRef]
- Girond, S.; Crance, J.M.; Van Cuyck-Gandre, H.; Renaudet, J.; Deloince, R. Antiviral activity of carrageenan on hepatitis A virus replication in cell culture. Res. Virol. 1991, 142, 261–270. [Google Scholar] [CrossRef]
- Bouhlal, R.; Haslin, C.; Chermann, J.-C.; Colliec-Jouault, S.; Sinquin, C.; Simon, G.; Cerantola, S.; Riadi, H.; Bourgougnon, N. Antiviral Activities of Sulfated Polysaccharides Isolated from Sphaerococcus coronopifolius (Rhodophytha, Gigartinales) and Boergeseniella thuyoides (Rhodophyta, Ceramiales). Mar. Drugs 2011, 9, 1187–1209. [Google Scholar] [CrossRef]
- Lopes, N.; Ray, S.; Espada, S.F.; Bomfim, W.A.; Ray, B.; Faccin-Galhardi, L.C.; Linhares, R.E.C.; Nozawa, C. Green seaweed Enteromorpha compressa (Chlorophyta, Ulvaceae) derived sulphated polysaccharides inhibit herpes simplex virus. Int. J. Biol. Macromol. 2017, 102, 605–612. [Google Scholar] [CrossRef]
- Robic, A.; Rondeau-Mouro, C.; Sassi, J.F.; Lerat, Y.; Lahaye, M. Structure and interactions of ulvan in the cell wall of the marine green algae Ulva rotundata (Ulvales, Chlorophyceae). Carbohydr. Polym. 2009, 77, 206–216. [Google Scholar] [CrossRef]
- Wang, S.; Wang, W.; Hao, C.; Yunjia, Y.; Qin, L.; He, M.; Mao, W. Antiviral activity against enterovirus 71 of sulfated rhamnan isolated from the green alga Monostroma latissimum. Carbohydr. Polym. 2018, 200, 43–53. [Google Scholar] [CrossRef] [PubMed]
- Morán-Santibañez, K.; Cruz-Suárez, L.E.; Ricque-Marie, D.; Robledo, D.; Freile-Pelegrín, Y.; Peña-Hernández, M.A.; Rodríguez-Padilla, C.; Trejo-Avila, L.M. Synergistic Effects of Sulfated Polysaccharides from Mexican Seaweeds against Measles Virus. BioMed Res. Int. 2016, 2016, 8502123. [Google Scholar] [CrossRef] [Green Version]
- Aguilar-Briseño, J.A.; Cruz-Suarez, L.E.; Sassi, J.-F.; Ricque-Marie, D.; Zapata-Benavides, P.; Mendoza-Gamboa, E.; Rodríguez-Padilla, C.; Trejo-Avila, L.M. Sulphated Polysaccharides from Ulva clathrata and Cladosiphon okamuranus Seaweeds both Inhibit Viral Attachment/Entry and Cell-Cell Fusion, in NDV Infection. Mar. Drugs 2015, 13, 697–712. [Google Scholar] [CrossRef] [PubMed]
- Hardouin, K.; Bedoux, G.; Burlot, A.-S.; Donnay-Moreno, C.; Bergé, J.-P.; Nyvall-Collén, P.; Bourgougnon, N. Enzyme-assisted extraction (EAE) for the production of antiviral and antioxidant extracts from the green seaweed Ulva armoricana (Ulvales, Ulvophyceae). Algal Res. 2016, 16, 233–239. [Google Scholar] [CrossRef] [Green Version]
- Terasawa, M.; Hayashi, K.; Lee, J.-B.; Nishiura, K.; Matsuda, K.; Hayashi, T.; Kawahara, T. Anti-Influenza A Virus Activity of Rhamnan Sulfate from Green Algae Monostroma nitidum in Mice with Normal and Compromised Immunity. Mar. Drugs 2020, 18, 254. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.-B.; Koizumi, S.; Hayashi, K.; Hayashi, T. Structure of rhamnan sulfate from the green alga Monostroma nitidum and its anti-herpetic effect. Carbohydr. Polym. 2010, 81, 572–577. [Google Scholar] [CrossRef]
- Cassolato, J.E.F.; Noseda, M.D.; Pujol, C.A.; Pellizzari, F.M.; Damonte, E.B.; Duarte, M.E.R. Chemical structure and antiviral activity of the sulfated heterorhamnan isolated from the green seaweed Gayralia oxysperma. Carbohydr. Res. 2008, 343, 3085–3095. [Google Scholar] [CrossRef] [PubMed]
- Ohta, Y.; Lee, J.-B.; Hayashi, K.; Hayashi, T. Isolation of Sulfated Galactan from Codium fragile and Its Antiviral Effect. Biol. Pharm. Bull. 2009, 32, 892–898. [Google Scholar] [CrossRef] [Green Version]
- Lee, J.B.; Hayashi, K.; Maeda, M.; Hayashi, T. Antiherpetic activities of sulfated polysaccharides from green algae. Planta Med 2004, 70, 813–817. [Google Scholar] [CrossRef]
- Chiu, Y.-H.; Chan, Y.-L.; Li, T.-L.; Wu, C.-J. Inhibition of Japanese Encephalitis Virus Infection by the Sulfated Polysaccharide Extracts from Ulva lactuca. Mar. Biotechnol. 2012, 14, 468–478. [Google Scholar] [CrossRef]
- Song, L.; Chen, X.; Liu, X.; Zhang, F.; Hu, L.; Yue, Y.; Li, K.; Li, P. Characterization and Comparison of the Structural Features, Immune-Modulatory and Anti-Avian Influenza Virus Activities Conferred by Three Algal Sulfated Polysaccharides. Mar. Drugs 2015, 14, 4. [Google Scholar] [CrossRef]
- Pujol, C.A.; Ray, S.; Ray, B.; Damonte, E.B. Antiviral activity against dengue virus of diverse classes of algal sulfated polysaccharides. Int. J. Biol. Macromol. 2012, 51, 412–416. [Google Scholar] [CrossRef] [PubMed]
- Hayashi, T.; Hayashi, K.; Maeda, M.; Kojima, I. Calcium Spirulan, an Inhibitor of Enveloped Virus Replication, from a Blue-Green Alga Spirulina platensis. J. Nat. Prod. 1996, 59, 83–87. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.-B.; Srisomporn, P.; Hayashi, K.; Tanaka, T.; Sankawa, U.; Hayashi, T. Effects of Structural Modification of Calcium Spirulan, a Sulfated Polysaccharide from Spirulina Platensis, on Antiviral Activity. Chem. Pharm. Bull. 2001, 49, 108–110. [Google Scholar] [CrossRef] [Green Version]
- Mader, J.; Gallo, A.; Schommartz, T.; Handke, W.; Nagel, C.-H.; Günther, P.; Brune, W.; Reich, K. Calcium spirulan derived from Spirulina platensis inhibits herpes simplex virus 1 attachment to human keratinocytes and protects against herpes labialis. J. Allergy Clin. Immunol. 2016, 137, 197–203. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ogura, F.; Hayashi, K.; Lee, J.-B.; Kanekiyo, K.; Hayashi, T. Evaluation of an Edible Blue-Green Alga, Aphanothece sacrum, for Its Inhibitory Effect on Replication of Herpes Simplex Virus Type 2 and Influenza Virus Type A. Biosci. Biotechnol. Biochem. 2010, 74, 1687–1690. [Google Scholar] [CrossRef] [Green Version]
- Hernández-Corona, A.; Nieves, I.; Meckes, M.; Chamorro, G.; Barron, B.L. Antiviral activity of Spirulina maxima against herpes simplex virus type 2. Antivir. Res. 2002, 56, 279–285. [Google Scholar] [CrossRef]
- Huleihel, M.; Ishanu, V.; Tal, J.; Arad, S. Antiviral effect of red microalgal polysaccharides on Herpes simplex and Varicella zoster viruses. J. Appl. Phycol. 2001, 13, 127–134. [Google Scholar] [CrossRef]
- Lee, J.-B.; Hayashi, K.; Hirata, M.; Kuroda, E.; Suzuki, E.; Kubo, Y.; Hayashi, T. Antiviral Sulfated Polysaccharide from Navicula directa, a Diatom Collected from Deep-Sea Water in Toyama Bay. Biol. Pharm. Bull. 2006, 29, 2135–2139. [Google Scholar] [CrossRef] [Green Version]
- Kim, M.; Yim, J.H.; Kim, S.-Y.; Kim, H.S.; Lee, W.G.; Kim, S.J.; Kang, P.-S.; Lee, C.-K. In vitro inhibition of influenza A virus infection by marine microalga-derived sulfated polysaccharide p-KG03. Antivir. Res. 2012, 93, 253–259. [Google Scholar] [CrossRef]
- Yim, J.H.; Kim, S.J.; Ahn, S.H.; Lee, C.K.; Rhie, K.T.; Lee, H.K. Antiviral Effects of Sulfated Exopolysaccharide from the Marine Microalga Gyrodinium impudicum Strain KG03. Mar. Biotechnol. 2004, 6, 17–25. [Google Scholar] [CrossRef]
- Hasui, M.; Matsuda, M.; Okutani, K.; Shigeta, S. In vitro antiviral activities of sulfated polysaccharides from a marine microalga (Cochlodinium polykrikoides) against human immunodeficiency virus and other enveloped viruses. Int. J. Biol. Macromol. 1995, 17, 293–297. [Google Scholar] [CrossRef]
- Huang, N.; Wu, M.-Y.; Zheng, C.-B.; Zhu, L.; Zhao, J.-H.; Zheng, Y.-T. The depolymerized fucosylated chondroitin sulfate from sea cucumber potently inhibits HIV replication via interfering with virus entry. Carbohydr. Res. 2013, 380, 64–69. [Google Scholar] [CrossRef] [PubMed]
- Kato, D.; Era, S.; Watanabe, I.; Arihara, M.; Sugiura, N.; Kimata, K.; Suzuki, Y.; Morita, K.; Hidari, K.I.P.J.; Suzuki, T. Antiviral activity of chondroitin sulphate E targeting dengue virus envelope protein. Antivir. Res. 2010, 88, 236–243. [Google Scholar] [CrossRef] [PubMed]
- Bergefall, K.; Trybala, E.; Johansson, M.; Uyama, T.; Naito, S.; Yamada, S.; Kitagawa, H.; Sugahara, K.; Bergström, T. Chondroitin sulfate characterized by the E-disaccharide unit is a potent inhibitor of herpes simplex virus infectivity and provides the virus binding sites on gro2C cells. J. Biol. Chem. 2005, 280, 32193–32199. [Google Scholar] [CrossRef] [Green Version]
- Perino, A.; Consiglio, P.; Maranto, M.; De Franciscis, P.; Marci, R.; Restivo, V.; Manzone, M.; Capra, G.; Cucinella, G.; Calagna, G. Impact of a new carrageenan-based vaginal microbicide in a female population with genital HPV-infection: First experimental results. Eur. Rev. Med. Pharm. Sci. 2019, 23, 6744–6752. [Google Scholar] [CrossRef]
- Damonte, E.B.; Matulewicz, M.C.; Cerezo, A.S. Sulfated seaweed polysaccharides as antiviral agents. Curr. Med. Chem. 2004, 11, 2399–2419. [Google Scholar] [CrossRef] [PubMed]
- Clausen, T.M.; Sandoval, D.R.; Spliid, C.B.; Pihl, J.; Perrett, H.R.; Painter, C.D.; Narayanan, A.; Majowicz, S.A.; Kwong, E.M.; McVicar, R.N.; et al. SARS-CoV-2 Infection Depends on Cellular Heparan Sulfate and ACE2. Cell 2020, 183, 1043–1057. [Google Scholar] [CrossRef]
- Walls, A.C.; Park, Y.J.; Tortorici, M.A.; Wall, A.; McGuire, A.T.; Veesler, D. Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein. Cell 2020, 181, 281–292. [Google Scholar] [CrossRef]
- Mottarella, S.E.; Beglov, D.; Beglova, N.; Nugent, M.A.; Kozakov, D.; Vajda, S. Docking server for the identification of heparin binding sites on proteins. J. Chem. Inf. Model 2014, 54, 2068–2078. [Google Scholar] [CrossRef]
- Singh, J.; Samal, J.; Kumar, V.; Sharma, J.; Agrawal, U.; Ehtesham, N.Z.; Sundar, D.; Rahman, S.A.; Hira, S.; Hasnain, S.E. Structure-Function Analyses of New SARS-CoV-2 Variants B.1.1.7, B.1.351 and B.1.1.28.1: Clinical, Diagnostic, Therapeutic and Public Health Implications. Viruses 2021, 13, 439. [Google Scholar] [CrossRef]
- Witvrouw, M.; De Clercq, E. Sulfated polysaccharides extracted from sea algae as potential antiviral drugs. Gen. Pharm. 1997, 29, 497–511. [Google Scholar] [CrossRef]
- Mukherjee, S.; Ghosh, K.; Hahn, F.; Wangen, C.; Strojan, H.; Müller, R.; Anand, N.; Ali, I.; Bera, K.; Ray, B.; et al. Chemically sulfated polysaccharides from natural sources: Assessment of extraction-sulfation efficiencies, structural features and antiviral activities. Int. J. Biol. Macromol. 2019, 136, 521–530. [Google Scholar] [CrossRef] [PubMed]
- Carlucci, M.J.; Scolaro, L.A.; Errea, M.I.; Matulewicz, M.C.; Damonte, E.B. Antiviral activity of natural sulphated galactans on herpes virus multiplication in cell culture. Planta Med. 1997, 63, 429–432. [Google Scholar] [CrossRef]
- Rodríguez, M.C.; Merino, E.R.; Pujol, C.A.; Damonte, E.B.; Cerezo, A.S.; Matulewicz, M.C. Galactans from cystocarpic plants of the red seaweed Callophyllis variegata (Kallymeniaceae, Gigartinales). Carbohydr. Res. 2005, 340, 2742–2751. [Google Scholar] [CrossRef] [PubMed]
- Jiao, G.; Yu, G.; Zhang, J.; Ewart, H.S. Chemical structures and bioactivities of sulfated polysaccharides from marine algae. Mar. Drugs 2011, 9, 196–223. [Google Scholar] [CrossRef] [Green Version]
- Sapay, N.; Nurisso, A.; Imberty, A. Simulation of carbohydrates, from molecular docking to dynamics in water. Methods Mol. Biol. 2013, 924, 469–483. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Zhang, Y.; Wu, L.; Niu, S.; Song, C.; Zhang, Z.; Lu, G.; Qiao, C.; Hu, Y.; Yuen, K.-Y.; et al. Structural and Functional Basis of SARS-CoV-2 Entry by Using Human ACE2. Cell 2020, 181, 894–904. [Google Scholar] [CrossRef] [PubMed]
- Towler, P.; Staker, B.; Prasad, S.G.; Menon, S.; Tang, J.; Parsons, T.; Ryan, D.; Fisher, M.; Williams, D.; Dales, N.A.; et al. ACE2 X-ray structures reveal a large hinge-bending motion important for inhibitor binding and catalysis. J. Biol. Chem. 2004, 279, 17996–18007. [Google Scholar] [CrossRef] [Green Version]
- Seeliger, D.; De Groot, B.L. Ligand docking and binding site analysis with PyMOL and Autodock/Vina. J. Comput. Aided Mol. Des. 2010, 24, 417–422. [Google Scholar] [CrossRef] [Green Version]
- Bowers, K.J.; Chow, D.E.; Xu, H.; Dror, R.O.; Eastwood, M.P.; Gregersen, B.A.; Klepeis, J.L.; Kolossvary, I.; Moraes, M.A.; Sacerdoti, F.D. Scalable algorithms for molecular dynamics simulations on commodity clusters. In Proceedings of the SC’06: 2006 ACM/IEEE Conference on Supercomputing, Tampa, FL, USA, 11–17 November 2006; p. 43. [Google Scholar]
- Horner-Miller, B. CellSs: A programming model for the cell BE architecture. In Proceedings of the 2006 ACM/IEEE Conference on Supercomputing, Tampa, FL, USA, 11–17 November 2006; Association for Computing Machinery: New York, NY, USA, 2006. [Google Scholar]
- Schrodinger Maestro; LLC: New York, NY, USA, 2009; Available online: https://www.schrodinger.com/products/maestro (accessed on 18 July 2021).
- Jo, S.; Kim, T.; Iyer, V.G.; Im, W. CHARMM-GUI: A web-based graphical user interface for CHARMM. J. Comput. Chem. 2008, 29, 1859–1865. [Google Scholar] [CrossRef] [PubMed]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD: Visual molecular dynamics. J. Mol. Graph. 1996, 14, 33–38. [Google Scholar] [CrossRef]
- Kim, S.; Oshima, H.; Zhang, H.; Kern, N.R.; Re, S.; Lee, J.; Roux, B.; Sugita, Y.; Jiang, W.; Im, W. CHARMM-GUI Free Energy Calculator for Absolute and Relative Ligand Solvation and Binding Free Energy Simulations. J. Chem. Theory Comput. 2020, 16, 7207–7218. [Google Scholar] [CrossRef] [PubMed]
| Sources | Compounds | Activity | Efficacy | Mode of Action | References |
|---|---|---|---|---|---|
| Brown Algae | |||||
| Undaria pinnatifid | Sulfated galactofucan | HSV-1, HSV-2, and HCMV | IC50 = 0.2–1.1 μg/mL | Inhibit virus entry and host cell binding | [14,15] |
| Cystoseira indica | Sulfated Fucans | HSV-1 and HSV-2 | IC50 = 0.50–2.8 μg/mL | Inhibit adsorption | [16] |
| Saccharina japonica | Fucoidan: RPI-27 and RPI-28 in a complex | SARS-CoV-2 | EC50 = 8.3 ± 4.6 μg/mL | S-protein binding | [17] |
| Padina boryana | Fucoidan: α-(1→3)-linked | SARS-CoV-2 | IC50 = 15.6 μg/mL | S-protein binding | [18,19] |
| Laminaria japonica | Sulfated polymannuroguluronate (SPMG) | HIV-1 | IC50 = 30 μg/mL | Inhibit the virus entry and replication | [20] |
| Laminaria japonica | Sulfated polymannuroguluronate (SPMG) | HIV-1 | 100 μg/mL | Inhibit HIV-1 entry by suppressing rgp120 binding to sCD4 | [21] |
| Laminaria japonica | Sulfated polymannuronate (SPMG)-derived oligosaccharides | HIV-1 | ID50 = 5.3 nM | Inhibit HIV-1 entry by suppressing rgp120 binding to sCD4 | [22] |
| Adenocystis utricularis | Sulfated galactofucan | HSV-1 and HSV-2 | IC50 = 1.25–2.16 μg/mL | Unknown | [23,24] |
| Stoechospermum marginatum | Sulfated fucan: (1→4)- and (1→3)-linked-α-L-fucopyranosyl residues | HSV-1 and HSV-2 | EC50 = 0.63–10.0 μg/mL | Inhibit binding of the virus to the host cell receptor | [25,26] |
| Sargassum horneri | Fucoidan | HSV-1, HIV-1, and HCMV | IC50 = 1–3.3 μg/mL | Unknown | [27] |
| Kjellmaniella crassifolia | Fucoidan KW | Inf A virus (H1N1, H3N2) | IC50 < 6.5 μg/mL | Suppress the replication | [28] |
| Sargassum patens | Galactofucan | HSV-1 and HSV-2 | EC50 = 1.3–5.5 μg/mL | Inhibit adsorption | [29] |
| Leathesia difformis | Fucoidan | HSV-1, HSV-2, and HCMV | IC50 = 0.5–1.9 μg/mL | Inhibit protein synthesis and adsorption | [30] |
| Fucus evanescens | Fucoidan | HSV-2 | 10 mg/kg/day for 5 days | Unknown | [31] |
| Undaria pinnatifida | Fucoidan | HSV-1, HSV-2, and HCMV | IC50 = 1.5–2.6 μg/mL | Inhibit entry and binding | [32] |
| Sphacelaria indica | Xylogalactofucan and alginic acid | HSV-1 | IC50 = 0.6–10 μg/mL | Inhibit adsorption | [33] |
| Sargassum mcclurei, Sargassum polycystum, and Turbinara ornata | Fucoidan | HIV-1 | IC50 = 0.33–0.7 μg/mL | Inhibit entry | [34] |
| Scinaia hatei | Sulfated xylomannan | HSV-1 and HSV-2 | IC50 = 0.5–4.6 μg/mL | Inhibit replication and virus binding | [16] |
| Cladosiphon okamuranus | Fucoidan | NDV | IC50 = 0.75 ± 1.6 μg/mL | Inhibit viral-induced syncytia formation | [35] |
| Cladosiphon okamuranus | Fucoidan | DENV-2 | Structure-based analysis: fucoidan interacts directly with envelope glycoprotein (EGP) on DEN2 | Inhibit binding | [36] |
| Undaria pinnatifida- derived fucoidan (UPF) | Fucoidan | InfAV (H1N1, PR8) | 7.0 mg/day for 7 days | [37] | |
| Undaria pinnatifida | Fucoidan | Avian InfAV | 5 mg/day for 14 days | Decrease replication | [38] |
| Red Algae | |||||
| Gymnogongrus griffithsiae and Cryptonemia crenulata | Kappa/iota/nu Carrageenan and DL-galactan hybrid | HSV-1 and HSV-2 | IC50 = 0.5–5.6 μg/mL | Inhibit adsorption | [39] |
| Sebdenia polydactyla | Sulfated xylomannans | HSV-1 | IC50 = 0.35–2.8 μg/mL | Inhibit virus attachment to the host cell | [40] |
| Euchema spinosum | Iota-carrageenan | SARS-CoV-2 | IC50 = 2.6 μg/mL | Inhibit replication | [41,42] |
| Euchema spinosum | Iota-carrageenan with Xylitol® nasal spray | SARS-CoV-2 | IC50 < 6.0 μg/mL | Inhibit SARS-CoV-2 in vitro | [42,43] |
| Chondrus crispus | lambda-carrageenan | SARS-CoV-2, and InfV A and B | EC50 = 0.3–1.4 μg/mL | Prevent the entry | [42,44] |
| Euchema spinosum | Iota-carrageenan containing Xylometazoline HCL | hRV1a, hRV8, and hCoV-OC43 | IC50 for hRV1a and hRV8 = 1.56–6.2 μg/mLMIC for hCoV-OC43 = 0.024 μg/mL | Prevent the entry | [7,42] |
| Gigartina skottsbergii | κ/ι /μ/ν-carrageenan | HIV-1 | IC50 = 0.4–3.3 μg/m | Prevent binding and replication of virions | [45] |
| Gymnogongrus torulosus | DL-hybrid sulfated galactan | HSV-1 and HSV-2 | IC50 = 1.1 to 27.4 μg/mL | Unknown | [46,47] |
| Cryptonemia crenulata | DL-galactan hybrid C2S-3 | DENV-2 | IC50 = 1 μg/mL | Block virus multiplication | [48,49] |
| Stenogramme interrupta | ξ- and λ-carrageenans | HSV-1 and HSV-2 | IC50 = 0.65–2.88 μg/mL | Inhibit binding and replication | [50] |
| Gracilaria corticata | Sulfated galactan | HSV-1 and HSV-2 | IC50 = 0.19–0.24 μg/mL | Inhibit virus attachment to the host cell | [51,52] |
| Aghardhiella tenera | Sulfated galactan | HIV-1 and HIV-2 | IC50 = 0.05–0.5 μg/mL | Inhibit cytopathic effect of HIV-1 and HIV-2 in MT-4 cells | [53] |
| Nemalion helminthoides | Sulfated mannan | HSV-1 | IC50 = 5.43–9.68 μg/mL | Unknown | [54] |
| Nothogenia fastigiata | Sulfated xylomannan | HSV-1 and HSV-2 | IC50 = 0.6–1.3 μg/mL | Prevent binding and replication | [55,56] |
| Scinaia hatei | Sulfated xylomannan | HSV | IC50 = 0.5 μg/mL | Inhibit replication | [57] |
| Nothogenia fastigiata | Sulfated xylogalactans | HSV-1 | EC50 = 15.0–32.6 μg/mL | Inhibit virus attachment to the host cell | [58,59] |
| Euchema spinosum Chondrus crispus | Lambda/Iota-carrageenan | DENV-2 and DENV-3 | EC50 = 0.14–4.1 μg/mL | Interference with virus adsorption | [42,60] |
| Euchema spinosum | Iota-carrageenan nasal spray | hRV, hCV, and InfA | 3 times per day for 7 days | Inhibit virus attachment to the host cell | [6,42] |
| Euchema spinosum | Iota-carrageenan containing lozenges (Coldamaris®) | hRV1a, CoV OC43, and InfA H1N1n | 10 mg daily | Prevent binding and inhibit replication | [8] |
| Euchema spinosum | Iota-carrageenan nasal spray | hRV1a, CoV OC43, and InfA H1N1n | Total of 1.0 mg daily | Prevent the virus from binding to cell surfaces or penetrating the cells | [42,61] |
| Euchema spinosum | Iota-carrageenan, nasal spray | H1N1 and H3N2 | IC50 = 0.04–0.2 μg/mL | Effectively inhibit virus adsorption to host cells and reduce replication | [42,62] |
| Euchema spinosum | Iota-Carrageenan | hRV | 104–107 TCID50/mL; 5–50 μg/mL | Prevent binding and/or the entry into the cells, inhibit replication | [42,63] |
| Euchema spinosum | Iota-Carrageenan | HPV | IC50 = 0.005 μg/mL | Blocking the initial interaction of capsids with cells and exert a postattachment inhibitory effect. | [64] |
| Acanthophora specifira | Lambda-carrageenan | HSV-1 and RVFV | IC50 = 75.8–80.5 μg/mL | Inhibit replication | [65] |
| Lithothamnion muelleri | Sulfated xylogalactans | HSV-1 and HSV-2 | EC50 = 49.64–125.79 μg/mL | Inhibit adsorption and penetration | [59,66] |
| Scinaia hatei | Sulfated xylan | HSV-1 and HSV-2 | IC50 = 0.22–1.37 μg/mL | Inhibit entry | [67] |
| Euchema spinosum | Iota-Carrageenan | SARS-COV-2 | IC50 ≥ 125 μg/mL | Unknown | [18,42] |
| Sigma (Sigma Aldrich) | Carrageenan | Japanese encephalitis virus (JEV) | EC50 = 15 µg/mL | Inhibit attachment and the cellular entry stages | [68] |
| Tichocarpus crinitus | κ/β-carrageenan | Tobacco Mosaic Virus (TMV) | 1 mg/mL | Significantly inhibit virus replication | [69] |
| Chondrus crispus | Lambda-carrageenan | DENV-2 and DENV-3 | >99% reduction in virus production at 20 μg/mL | Inhibit entry | [42,70] |
| Meristiella gelidium | Iota/kappa/nu-hybrid carrageenan | DENV-2 | IC50 = 0.14–1.6 μg/mL | Unknown | [71] |
| Iota/lambda/kappa-carrageenan | Hepatitis A Virus (HAV) | Ratio of CD50 to ED50 >400 | Inhibit replication | [72] | |
| Sphaerococcus coronopifolius and Boergeseniella thuyoides | Sulfated galactan | HIV-1 and HSV-1 | EC50 = 4.1–17.2 μg/mL | Inhibit adsorption and replication | [73] |
| Green Algae | |||||
| Enteromorpha compressa | Ulvan | HSV | IC50 = 28.25 μg/mL | Inhibit adsorption and replication | [74,75] |
| Monostroma latissimum | Sulfated rhamnan | EV71 | IC50 = 0.5 μg/mL | inhibit replication | [76] |
| Ulva intestinalis | Ulvan | Measles virus (MeV) | IC50 = 3.6 μg/mL | Reduce the formation of syncytia | [77] |
| Ulva clathrata | Ulvan | NDV | IC50 = 0.1 μg/mL | Inhibit cell-cell fusion via a direct effect on F0 protein | [78] |
| Ulva armoricana | Ulvan (enzymatic preparations) | HSV-1 | EC50 = 320.9–373.0 μg/mL | Unknown | [79] |
| Monostroma nitidum | Rhamnan Sulfate | HSV-1, HSV-2, HCMV, MeV, MuV, HCV, HCoV and HIV | EC50 = 0.77–8.30 μg/mL | Inhibit viral proliferation | [80] |
| Monostroma nitidum | Rhamnan Sulfate | HSV-2 | IC50 = 0.87 μg/mL | Suggested to inhibit adsorption or penetration | [81] |
| Gayralia oxysperma | Sulfated heterorhamnan | HSV-1 and HSV-2 | IC50 = 0.036–0.3 μg/mL | Inhibit multiplication | [82] |
| Codium fragile | Sulfated galactan | HSV-2 | IC50 = 4.7 μg/mL | Interference with the early steps such as virus adsorption to and penetration into host cells | [83] |
| Monostroma nitidum, C. okamurai, C. scapelliformis, Chaetomorpha crassa, C. spiralis, Codium adhaerens, C. fragile, Caulerpa brachypus and Caulerpa latum | Sulfated arabinoxylogalactan | HSV-1 | IC50 = 0.38–8.5 μg/mL | Inhibit binding, penetration, and the late stages of replication | [84] |
| Enteromorpha compressa | Sulfated heteroglycuronan (chemically modified) | HSV-1 | IC50 = 28.25 μg/mL | Inhibit replication | [74] |
| Ulva lactuca | Ulva sulfated polysaccharide extracts | JEV | 0.75 mg/day for 7 days | The survival rate significantly increased | [85] |
| Ulva Pertusa | Ulvan | Avian InfAV | 50 mg twice (INJ) immunizations dose | Enhance AIV-specific antibody production and improve the humoral immunity level | [86] |
| Caulerpa racemosa | Sulfated heteropolysaccharide | DENV-2 | IC50 = 0.6 μg/mL | Interfere with virus multiplication | [87] |
| Blue-Green Algae | |||||
| Spirulina platensis | Calcium Spirulan (Ca-SP) | HSV-1, HCMV, MeV, MuV, InfA, HIV-1 | EC50 = 0.92–23 μg/mL | Inhibit replication and penetration | [88] |
| Spirulina platensis | Calcium spirulan, Sodium spirulan, and potassium spirulan | HSV-1 | IC50 = 0.46–0.88 μg/mL | Inhibit replication | [89] |
| Spirulina platensis | Spirularin® HS “Ocean Pharma (Topical cream containing Ca-SP in combination with SPME) | HSV-1 | 15 mg/g twice daily for 3 weeks(10 mg/g SPME) | Significantly prevent herpes labialis exacerbation | [90] |
| Aphanothece sacrum | Sulfated polysaccharides (ASWPH) | HSV-2 and Inf A (H1N1) | IC50 = 0.32–1.2 μg/mL | Inhibit adsorption | [91] |
| Spirulina maxima | Hot water extract (HWE) | HSV-1, HSV-2, HCMV, and pseudorabies virus (PRV) | ED50 = 0.069–0.333 mg/mL | Inhibit adsorption and penetration | [92] |
| Microalgae | |||||
| Red; Porphyridium sp. | cell-wall sulphated polysaccharide | HSV-1, HSV-2 and Varicella Zoster Virus (VZV) | IC50 = 1 µg/mL | Prevent adsorption and/or inhibit the production of new viral particles | [93] |
| Diatom; Navicula directa | Naviculan | HSV-1, HSV-2, HIV, and Inf A | IC50 = 7.4–170 µg/mL | Inhibit binding and penetration. Inhibit cell–cell fusion between HIV gp160- and CD4. | [94] |
| Gyrodinium impudium | P-KG03 | InfA H1N1 and H3N2 | EC50 = 0.19–0.48 μg/mL | Inhibit replication and entry | [95] |
| Gyrodinium impudicum | P-KG03 | EMCV | EC50 = 26.9 µg/ml | Inhibit replication | [96] |
| Cochlodinium polykrikoides | Extracellular sulfated polysaccharides (A1 and A2) | HIV-1, HSV-1, Inf A, RSV-A, and RSV-B | EC50 = 1.1–4.52 µg/mL | Inhibit replication | [97] |
| Marine animal-derived sulfated polysaccharides | |||||
| Thelenota ananas | Sea cucumber Fucosylated chondroitin sulfate | HIV-1 | EC50 = 0.73–31.86 μg/mL | inhibit the entry and replication | [98] |
| Stichopus japonicus | Sea cucumber sulfated polysaccharide (SCSP) | SARS-COV-2 | EC50 = 9.10 μg/mL | Interact with the S glycoprotein | [18] |
| Squid cartilage | Chondroitin sulphate E (CS-E) | DENV | EC50 = 0.52–11.8 μg/mL | Inhibit the entry via targeting E protein | [99] |
| Squid cartilage | Chondroitin sulfate E (CS-E) | HSV-1 and HSV-2 | IC50 = 0.06 to 0.2 μg/mL | Inhibit binding | [100] |
| No. | Compound | Vina Score (kcal/mol) # | ΔG * | Average RMSD (Å) | Reported Activity Against SARS CoV-2 | |||
|---|---|---|---|---|---|---|---|---|
| Original Strain | Mutated Strain ** | Original Strain | Mutated Strain ** | Original Strain | Mutated Strain ** | |||
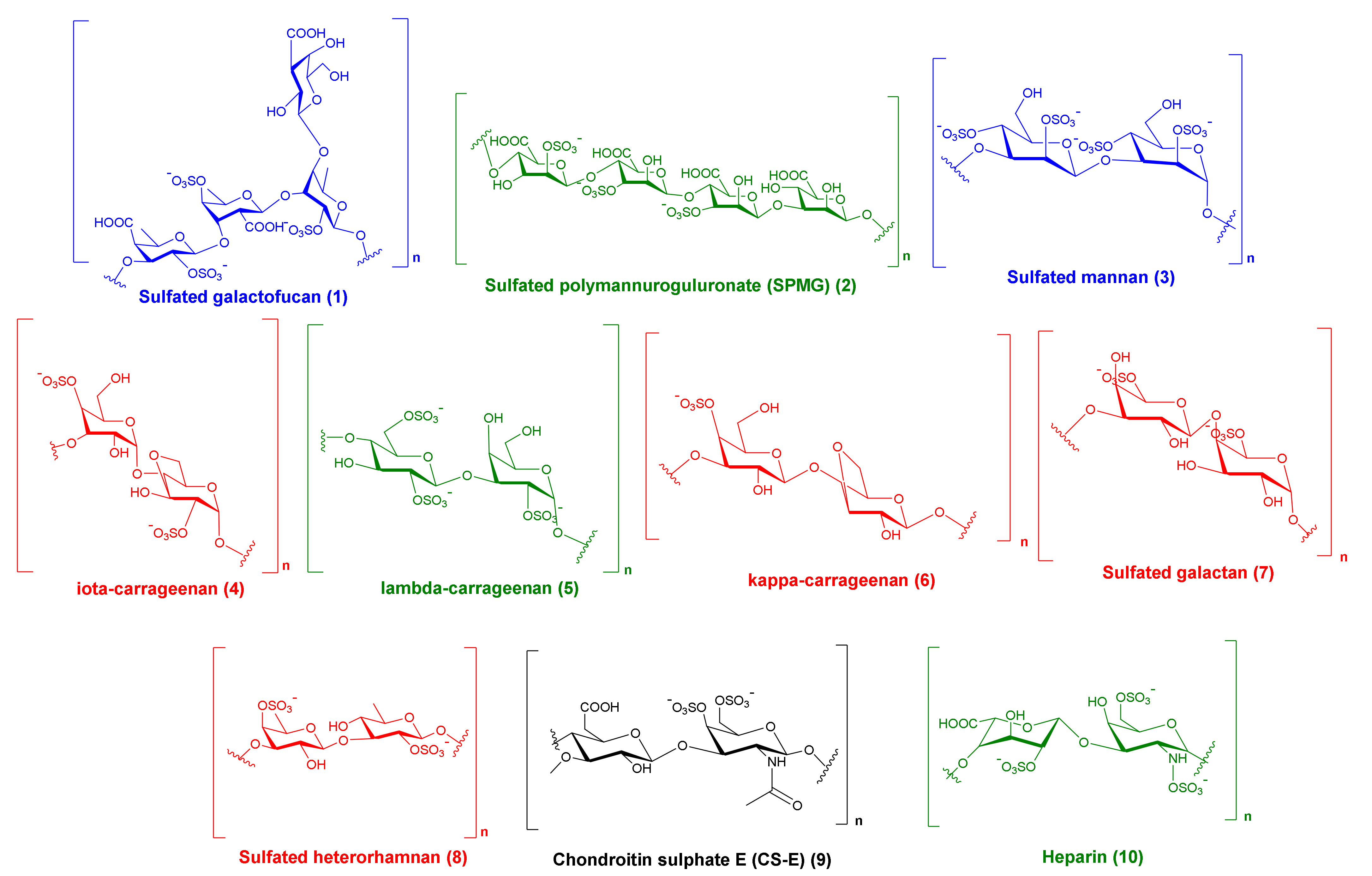
| 1 | Sulfated galactofucan: α- (1,3)- and (1,4)-α-L- (alternating) | −6.3 | −6.4 | −6.0 | −6.3 | 5.1 | 4.5 | No |
| 2 | Sulfated polymannuroguluronate (SPMG) | −7.5 | −6.4 | −7.3 | −5.9 | 2.6 | 4.1 | Yes |
| 3 | Sulfated mannan | −7.6 | −7.7 | −7.2 | −7.4 | 2.3 | 1.9 | No |
| 4 | iota-carrageenan | −6.0 | −6.0 | −0.8 | −0.9 | >15 *** | >15 *** | Yes |
| 5 | lambda-carrageenan | −7.0 | −7.0 | −5.4 | −5.8 | 4.6 | 4.3 | Yes |
| 6 | kappa-carrageenan | −6.6 | −6.4 | −1.3 | −1.9 | >15 *** | >15 *** | Yes |
| 7 | Sulfated galactan | −6.2 | −6.3 | −0.4 | −0.1 | >15 *** | >15 *** | No |
| 8 | Sulfated heterorhamnan | −6.1 | −6.2 | −0.1 | −0.7 | >15 *** | >15 *** | No |
| 9 | Chondroitin sulphate E (CS-E) | −7.6 | −7.5 | −7.1 | −6.1 | 1.1 | 2.8 | No |
| 10 | Heparin | −6.5 | −6.4 | −6.1 | −6.1 | 3.3 | 3.3 | Yes |
| No. | Compound | Vina Score (kcal/mol) | ΔG * | Average RMSD (Å) | Reported Activity Against SARS CoV-2 |
|---|---|---|---|---|---|
| 1 | Sulfated galactofucan: α- (1,3)- and (1,4)-α-L- (alternating) | −5.3 | −5.7 | 2.4 | No |
| 2 | Sulfated polymannuroguluronate (SPMG) | −5.5 | −5.9 | 2.1 | No |
| 3 | Sulfated mannan | −5.4 | −5.1 | 2.6 | No |
| 4 | iota-carrageenan | −5.0 | −1.7 | >15 ** | Yes |
| 5 | lambda-carrageenan | −5.5 | −5.4 | 2.5 | Yes |
| 10 | Heparin | −5.0 | −5.8 | 1.9 | Yes |
| No. | Compound | Vina Score (kcal/mol) | ΔG * | Average RMSD (Å) | Reported Activity Against SARS CoV-2 |
|---|---|---|---|---|---|
| 2 | Sulfated polymannuroguluronate (SPMG) | −5.9 | −5.4 | 2.8 | No |
| 5 | lambda -carrageenan | −5.5 | −5.0 | 3.1 | Yes |
| 10 | Heparin | −5.3 | −5.0 | 3.0 | Yes |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Salih, A.E.M.; Thissera, B.; Yaseen, M.; Hassane, A.S.I.; El-Seedi, H.R.; Sayed, A.M.; Rateb, M.E. Marine Sulfated Polysaccharides as Promising Antiviral Agents: A Comprehensive Report and Modeling Study Focusing on SARS CoV-2. Mar. Drugs 2021, 19, 406. https://doi.org/10.3390/md19080406
Salih AEM, Thissera B, Yaseen M, Hassane ASI, El-Seedi HR, Sayed AM, Rateb ME. Marine Sulfated Polysaccharides as Promising Antiviral Agents: A Comprehensive Report and Modeling Study Focusing on SARS CoV-2. Marine Drugs. 2021; 19(8):406. https://doi.org/10.3390/md19080406
Chicago/Turabian StyleSalih, Abdalla E. M., Bathini Thissera, Mohammed Yaseen, Ahmed S. I. Hassane, Hesham R. El-Seedi, Ahmed M. Sayed, and Mostafa E. Rateb. 2021. "Marine Sulfated Polysaccharides as Promising Antiviral Agents: A Comprehensive Report and Modeling Study Focusing on SARS CoV-2" Marine Drugs 19, no. 8: 406. https://doi.org/10.3390/md19080406
APA StyleSalih, A. E. M., Thissera, B., Yaseen, M., Hassane, A. S. I., El-Seedi, H. R., Sayed, A. M., & Rateb, M. E. (2021). Marine Sulfated Polysaccharides as Promising Antiviral Agents: A Comprehensive Report and Modeling Study Focusing on SARS CoV-2. Marine Drugs, 19(8), 406. https://doi.org/10.3390/md19080406









