Abstract
The marine sponge of the genus Geodia, Jaspis, Rhabdastrella, and Stelletta are characterized chemically by a variety of isomalabaricane triterpenes. This class of compounds drew spotlights in marine lead discovery due to their profound anti-proliferative properties. Further research on exploring its chemical diversity led to the identifications of two new isomalabaricane-type triterpenes rhabdastin H (1) and rhabdastin I (2). Their structures were unraveled using a series of spectroscopic approaches. These isolates were found to exhibit unique structural features with the only reported tetrahydrofuran functionality among all marine-derived isomalabaricanes. Both compounds 1 and 2 showed activities against K562 (IC50 11.7 and 9.8 μM) and Molt4 (IC50 16.5 and 11.0 μM) leukemic cells in MTT cell proliferative assay.
1. Introduction
Marine sponges continue to act as a fruitful source of bioactive and unusual metabolites. Among these metabolites are the sponge-derived isomalabaricanes. They were reported to exhibit potent anti-proliferative properties against a series of cancer cells, including colorectal carcinoma [1], ovarian carcinoma [2], acute promyelocytic leukemia [3], acute lymphoblastic leukemia [4], prostate carcinoma [3], gastric adenocarcinoma [3], ductal carcinoma [3], hepatocellular carcinoma [3], cervix carcinoma [3], papillomavirus-related endocervical adenocarcinoma [5], and malignant melanoma [4]. The isomalabaricane-type triterpenoids are characterized by an α-methyl group at C-8, owning to their trans-syn-trans tricyclic ring junction instead of trans-anti-trans of their isomers, malabaricanes [1]. Isomalabaricanes are only found in sponges and are considered as chemotaxonomic markers of Rhabdastrella sp. [1] Despite the significant difficulties in the isolation and characterization of isomalabaricanes due to light induced 13-E/Z isomerization [6], their significant cytotoxic activity prompted extensive research on this class of compounds aiming to find new drug leads.
The current study aimed to explore novel isomalabaricanes from marine sources. We isolated two unusual compounds with tetrahydrofuran functional group, the moiety that has been reported previously in malabaricane-type triterpenoids but without any cytotoxic assessments [7,8]. Then, the isolated compounds were evaluated against several cancer cell lines using MTT assay to further interpret their anti-proliferative properties.
2. Results
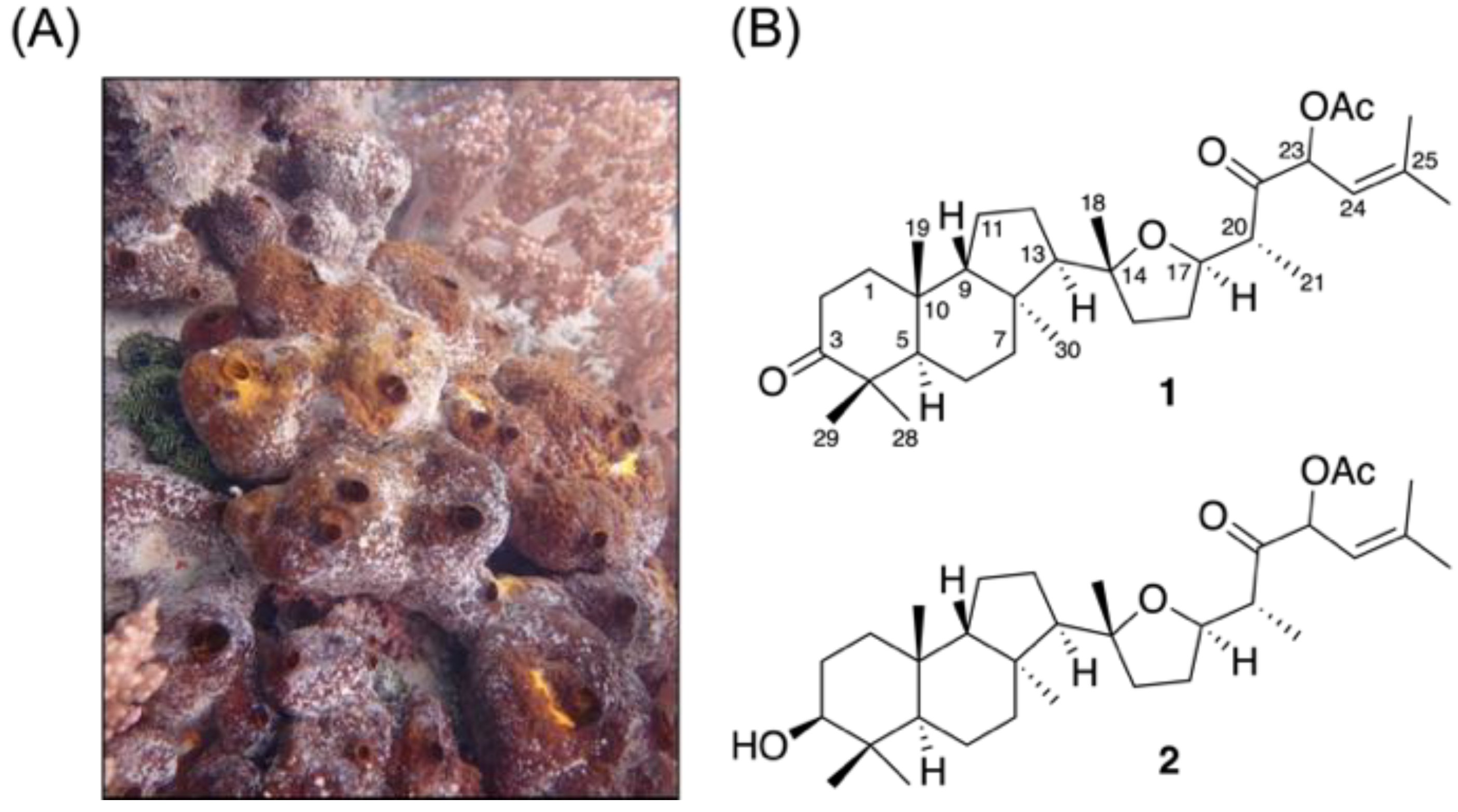
The freeze-dried specimen of the wild-type sponge Rhabdastrella sp. (Figure 1) was extracted with a 1:1 mixture of methanol (MeOH) and dichloromethane (CH2Cl2) (1:1) to provide the crude extract. The obtained crude extract was further fractionated and purified using normal and reversed-phase column chromatography yielding two isomalabaricanes, rhabdastin H (1) and rhabdastin I (2). These isolates demonstrated unique structural features with the presence of the first identified tetrahydrofuran moiety in this class of compounds.
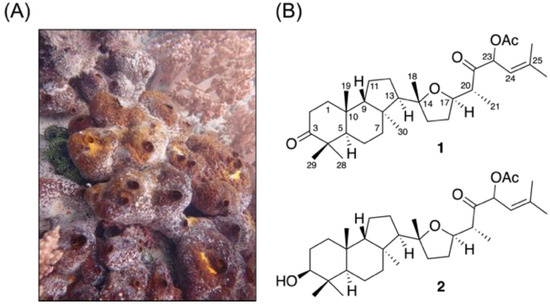
Figure 1.
(A) Aquatic ecology of the sponge Rhabdastrella sp. and (B) the isolated isomalabaricane triterpenes.
The molecular formula of 1 was suggested as C32H50O5 based on 13C NMR and HRESIMS data that showed a molecular ion peak at m/z 537.3539 [M + Na]+ implying eight degrees of unsaturation. The IR spectrum of 1 revealed the presence of carbonyl groups from absorptions at 1742, 1726, and 1703 cm−1. Its 13C NMR spectrum of 1 (Table 1), measured in CDCl3, showed the presence of thirty-two carbon signals, which were assigned by the assistance of DEPT data to nine methyl groups, eight sp3 methylenes, six sp3 methines (including two oxymathines), four sp3 quaternary carbons, one sp2 methine and four sp2 quaternary carbons (including two ketone carbonyl). The NMR signals at δC 169.8 (C) and 20.9 (CH3) and δH 2.12 (3H, s) and the IR absorption at 1742 cm−1 suggested the presence of an acetoxy group. Carbon signals of the eight methyl groups (δC 31.8, 29.3, 26.0, 24.5, 24.0, 19.3, 19.0 and 12.1), one oxygen-bearing methine carbon (δC 79.8), and one oxygenated quaternary carbon (δC 86.2), one trisubstituted carbon-carbon double bonds (δC 116.9, CH; 142.6, C), two ketones (δC 220.3 and 207.4) were also assigned. The resonances of one olefinic proton (δH 5.11, d, J = 10.0 Hz) and two oxygenated methines (δH 6.09, d, J = 10.0 Hz; δH 3.76, ddd, J = 14.0, 8.5, 4.5 Hz) were observed from the 1H NMR spectroscopic data of 1 (Table 1).

Table 1.
1H, 13C, 1H–1H COSY, and HMBC NMR data of 1.
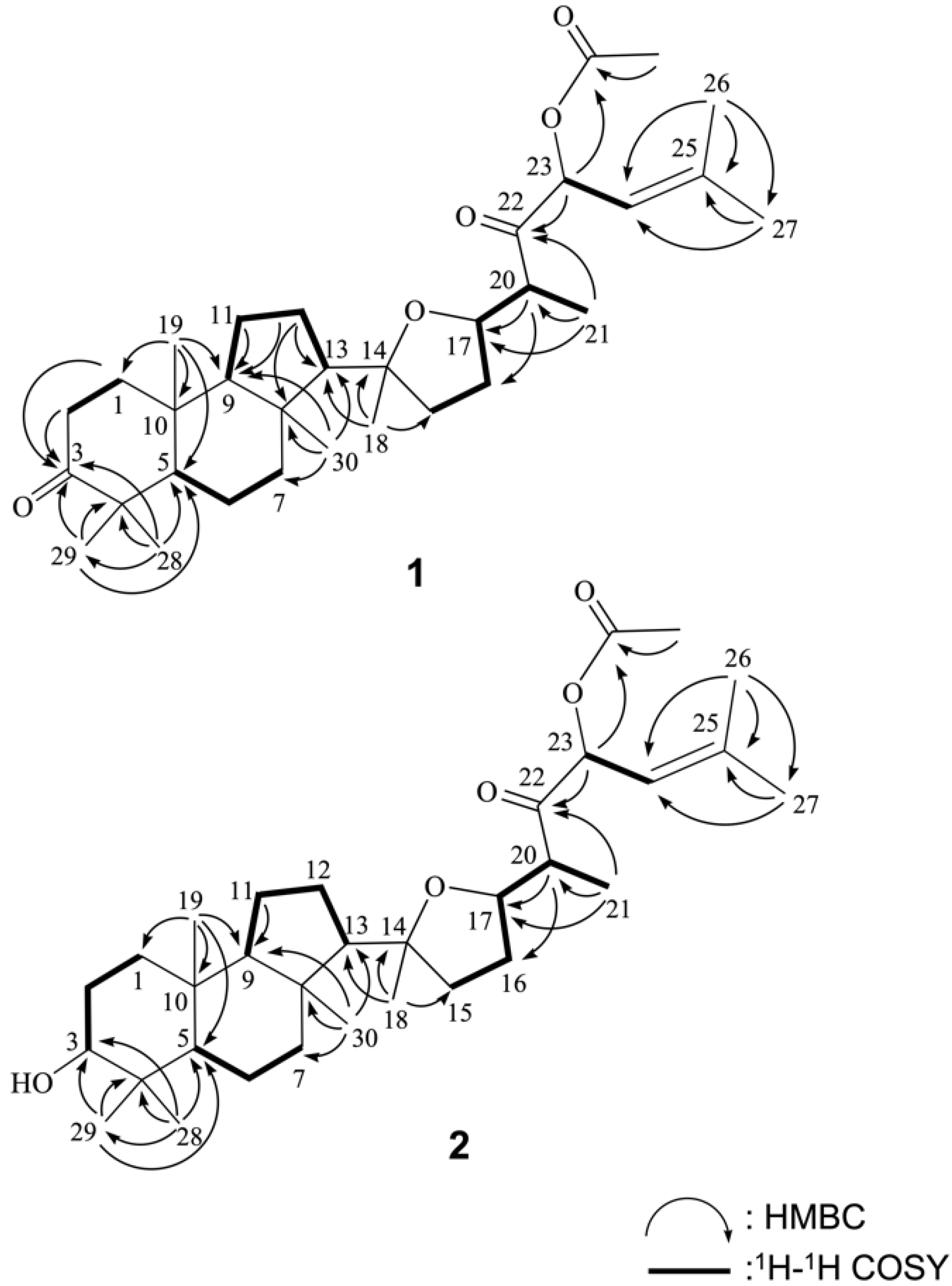
Based on the above results and with the assistance of 1H–1H COSY and HMBC experiments (Figure 2), the planar structure of 1 was determined. To establish the proton sequences in 1, the 1H–1H COSY spectrum analysis established five proton sequences from H2-1 to H2-2, H-5 to H2-7, and H2-11 to H2-12, H2-15 to H-21, and H-23 to H-24. These data, together with the HMBC correlations (Figure 2) from H2-1 and H2-2 to C-3, H2-11 and H2-12 to C-9, H2-12 to C-8 and C-13, H3-19 to C-1, C-5, C-9 and C-10, H3-28 and H3-29 to C-3, C-4 and C-5 and H3-30 to C-7, C-8, C-9 and C-13 established the connectivity within the 6-membered (A), 6-membered (B), and 5-membered (C) rings.
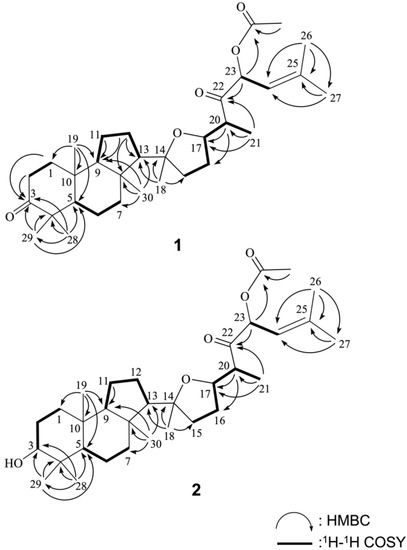
Figure 2.
Selected 1H-1H COSY and HMBC correlations of 1 and 2.
Ring A of 1 was found to possess one ketone at C-3 and two methyl groups (C-28 and C-29), one methyl group (C-19), and one methyl groups (C-30) attached at C-4, C-8, and C-10, respectively. The key HMBC correlations suggested the connection of H3-18 to C-13, C-14, and C-15, H-20 to C-16 and C-17; H3-21 to C-17, C-20, and C-22; H-23 to C-22; H3-26 and H3-27 to C-24 and C-25. Thus, the side chain C-20 to C-27 was found to possess one double bond at C-24/C-25, one ketone group at C-22, two methyl groups at C-25, and one methyl group at C-20. One acetoxy group at C-23 was confirmed by the HMBC correlation between an oxymethine proton resonating at δH 6.09 (H-23) and the protons of an acetate methyl (δH 2.12) to the ester carbonyl carbon at δC 169.8. An ether linkage was proposed between C-14 and C-17 forming a tetrahydrofuran ring based on the degrees of unsaturation and molecular formula. Based on the above analysis, the gross structure of 1 was established unambiguously and named rhabdastin H following up the previous investigation of cytotoxic isomalabaricanes with oxygenated and olefinic side chains from the sponge Rhabdastrella globostellata [9].
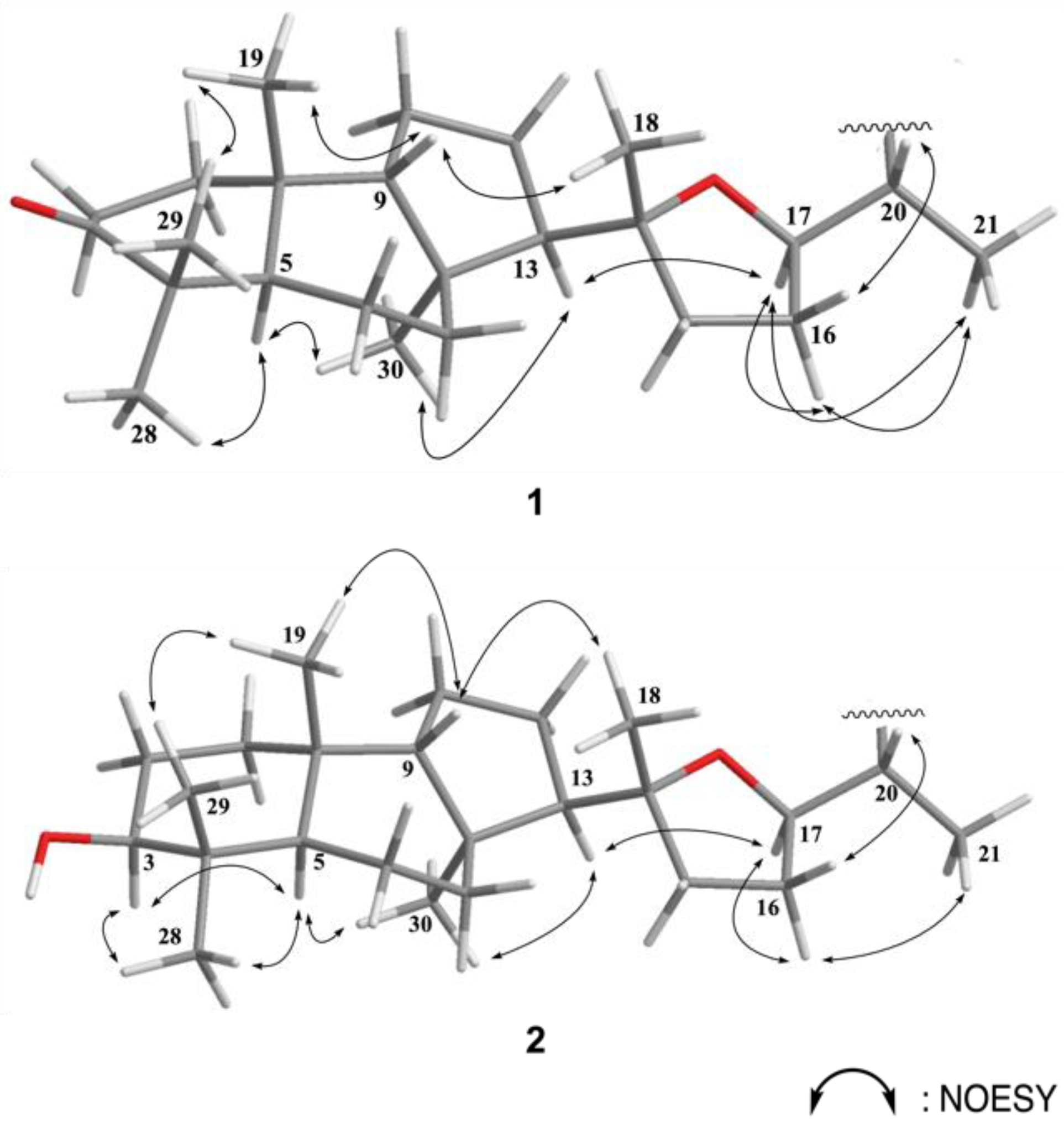
The relative configurations of the eight chiral centers at C-5, C-8, C-9, C-10, C-13, C-14, C-17, and C-20 in 1 were elucidated by the following NOE analysis (Figure 3). It was found that H3-19 (δH 0.79, s) showed NOE interaction with H-9 (δH 1.44, m) and H-9 with H3-18.
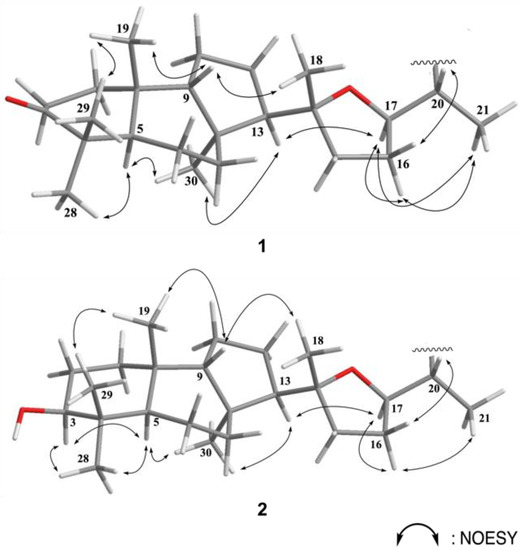
Figure 3.
Selected NOESY correlations for 1 and 2.
Since all naturally occurring isomalabaricanes displayed that H-5 is trans to Me-19, we assumed the β-orientation of H3-19. Thus, H3-18 and H-9 were suggested to be positioned on the β-face. One of the methyl groups (H3-29) at C-4, which showed a NOE correlation with H3-19, was β-oriented and the other one (H3-28) was α-oriented. The NOE correlations observed between H3-28/H-5, H-5/H3-30, H3-30/H-13, H-13/H-17, H-17/H3-21, and H-17/H-16 suggested α-orientation of H-5, H-13, H-17, H3-21, and H3-30. One of the methylene protons at C-16 (δH 1.97), which showed a NOE correlation with H-17 and H3-21, was assigned as H-16α and the other one (δH 1.58) as H-16β. Moreover, the detection of large proton coupling constants at H-17 (J = 13.5 Hz) and H-20 (J = 13.5 Hz) further supported their trans conformation in between. The observed NOE correlation between H-20 and H-16β suggested a β-orientation of H-20.
The HR-ESI-MS of rhabdastin I (2) showed a molecular ion peak at (m/z 539.3696 [M + Na]+) and the molecular formula C32H52O5 was suggested based on the HRESIMS and 13C NMR data. The IR spectrum of 2 showed the absorption of carbonyl groups (1745 and 1725 cm−1) and a hydroxy group (3436 cm−1). The 1H and 13C NMR spectroscopic data of 2 (Table 2) and 1 (Table 1) indicated similarity in structure except that the ketocarbonyl carbon (C-3) in 1 was replaced by a hydroxy group-bearing methine carbon in 2. In the 13C NMR spectrum, the signal at δc 220.3 was replaced by a signal at δc 79.5, and in the 1H NMR spectrum, a signal at δH 3.24 (dd, J = 11.5, 6.5 Hz) was attributed to a hydroxy-bearing methine at C-3. H-3 showed an NOE correlation with H-5 (δH 1.54, m) suggesting a β-orientation of the hydroxy group at C-3. Therefore, 2 was identified as the 2S-hydroxy derivative of 1.

Table 2.
1H, 13C, 1H–1H COSY, and HMBC NMR data of 2.
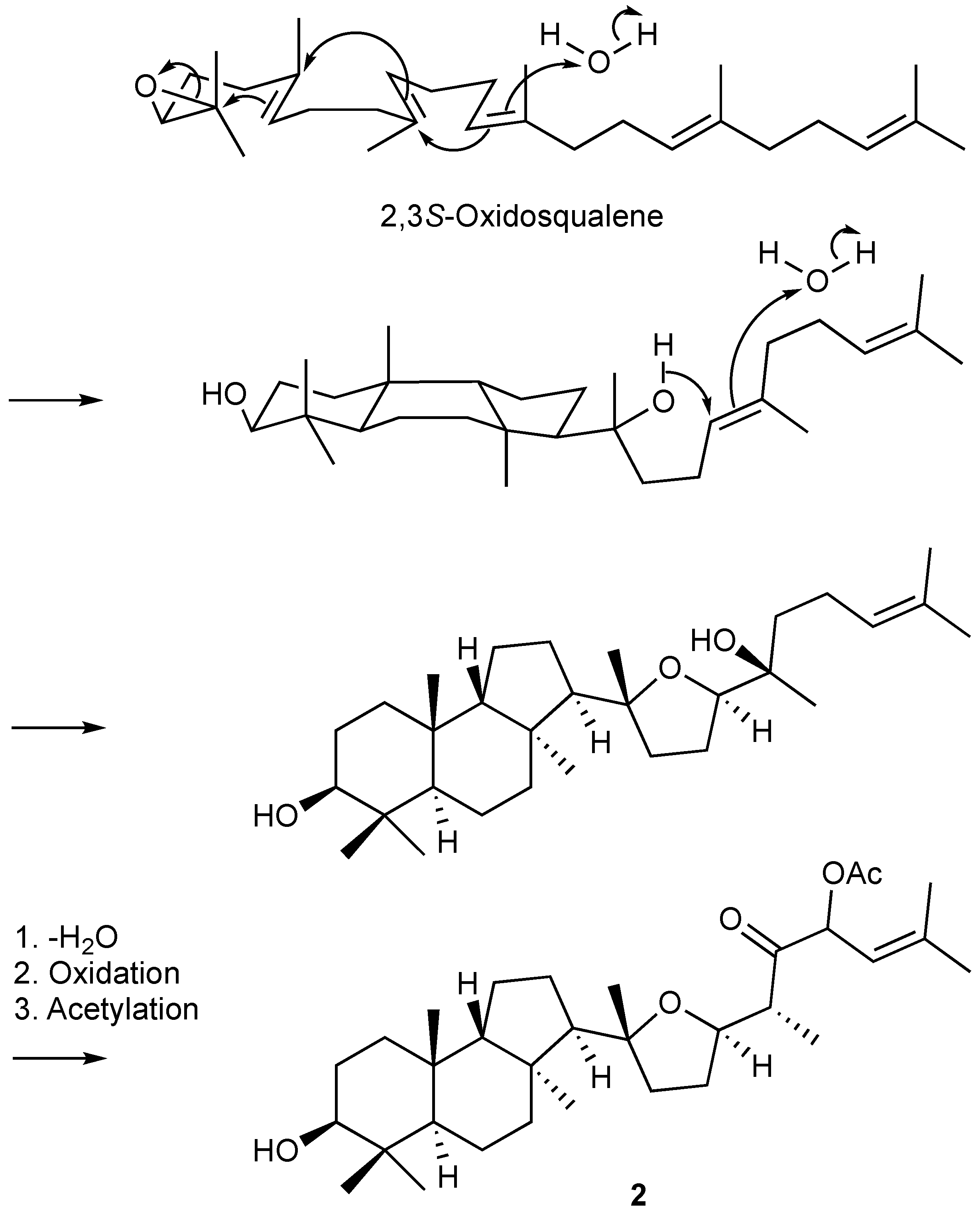
The plausible biosynthesis route of the obtained isomalabaricanes presented in Figure 4. The main isomalabaricane 6/6/5 carbocyclic skeleton might be derived from 2,3S-oxidosqualene through hydroxylation and electrocyclizations. The attached tetrahydrofuran moiety and the subsequent side chain were suggested to go through a hydroxylation, an electrocyclization, dehydration, oxidation, and acetylation, forming the first identified isomalabaricane with tetrahydrofuran functionality.
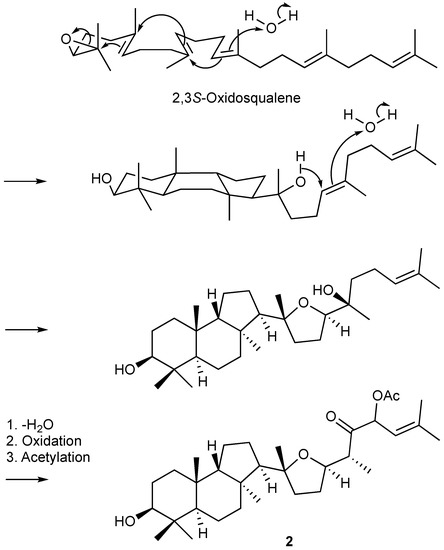
Figure 4.
The plausible biosynthetic pathway of 2.
To further clarify the anti-proliferative potential of the isolated compounds, four cancer cell lines (DLD-1: colorectal adenocarcinoma; T-47D: ductal carcinoma; Molt4: acute lymphoblastic leukemia; K562: chronic myelogenous leukemia) were used for MTT screening (Table 3). Compounds 1 and 2 were found to exhibit anti-proliferative activities against Molt4 and K562 leukemic cells with the IC50 value ranging from 9.81 to 16.54 μM.

Table 3.
Anti-proliferative activities of compounds 1 and 2.
3. Materials and Methods
3.1. General Experimental Procedures
Optical rotation spectra were recorded on a JASCO P-1010 polarimeter (JASCO, Tokyo, Japan). UV spectra were analyzed using JASCO UV-530 ultraviolet spectrophotometers. IR spectra were obtained on a Fourier-transform IR spectrophotometer Varian Digilab FTS 1000 (Varian Inc., Palo Alto, CA, USA). NMR spectra were obtained on a JEOL ECZ 400S or an ECZ 600R NMR (JEOL, Tokyo, Japan). HRESIMS data were collected on a Bruker APEX II instrument (Bruker Daltonik, Bremen, Germany). TLC was performed on Kieselgel 60 F254 (0.25 mm, Merck, Darmstadt, Germany) and/or RP-18 F254 (0.25 mm) coated plates and then visualized by spraying with 50% H2SO4 and heating on a hot plate. Silica gel 60 (Merck, 40−63 μm and 63−200 μm) were used for column chromatography. A Rheodyne 7725 injection port, a Hitachi L-2455 Photodiode Array Detector, and a Hitachi L-7100 pump (Hitachi, Tokyo, Japan), as well as a column Supelco Ascentis® C-18 Cat #: 581343-U, were applied for HPLC chromatography. All methods were carried out following the relevant guidelines and regulations.
3.2. Animal Material
The specimen of the wild-type sponge Rhabdastrella sp. was collected by scuba diving at a depth about 3–5 m from Kenting, Pingtung, Taiwan in December 2017. The voucher specimen was deposited at −20 °C at the National Museum of Marine Biology and Aquarium, Taiwan (specimen No. 2017-1221-SP). Taxonomic identification was performed by Dr. Hsing-Hui Li using 18S DNA sequence and morphology determination.
3.3. Extraction and Isolation
Rhabdastrella sp. (500 g fresh weight) was collected and freeze-dried. The freeze-dried material (75 g, dry weight) was minced and extracted three times with a 1:1 mixture of methanol (MeOH) and dichloromethane (CH2Cl2). The crude extract was evaporated under reduced pressure to afford a residue (8 g), and the residue was subjected to a normal phase column chromatography on silica gel (70–230 mesh), using n-hexane, increasing polarity of n-hexane:EtOAc mixtures, and acetone to yield 13 fractions: L1 (eluted by n-hexane), L2 (eluted by n-hexane:EtOAc, 100:1), L3 (50:1), L4 (30:1), L5 (20:1), L6 (10:1), L7 (5:1), L8 (3:1), L9 (2:1), L10 (1:1), L11 (1:2), L12 (eluted by EtOAc) and L13 (eluted by acetone). L10 was further separated with silica gel (n-hexane:acetone 6:1) using normal phase HPLC to afford ten subfractions (L10-1 to L10-10). Subfraction L10-4 was then subjected to a reversed-phase HPLC (RP-HPLC) (MeOH:H2O, 85:15), yielding 1 (3.1 mg). Similarly, the subfraction L10-6 was purified by RP-HPLC (MeOH:H2O, 80:20) to provide 2 (8.5 mg).
Rhabdastin H (1): Colorless oil; −47.1 (c 0.03, CHCl3); IR (ATR, CHCl3) νmax 1742 and 1726 cm–1; 1H NMR data, see Table 1; HRESIMS m/z 537.3539 [M + Na]+ (calcd. 537.3550, see supplementary materials).
Rhabdastin I (2): Colorless oil; −137.5 (c 0.02, CHCl3); IR (ATR, CHCl3) νmax 3436, 1745, and 1725 cm–1; 1H NMR data, see Table 2; ESIMS m/z 539.3696 [M + Na]+ (calcd. 539.3707, see supplementary materials).
3.4. MTT Cell Proliferation Assay
MTT assay was used to examine the cellular proliferation of DLD-1 (colorectal adenocarcinoma), T-47D (ductal carcinoma), Molt4 (acute lymphoblastic leukemia), and K562 (chronic myelogenous leukemia) after 1 and 2 treatments. American Type Culture Collection (ATCC, Manassas, VA, USA) was the source for all cell lines. Briefly, cells at 1 × 105 cells/mL were seeded at 96-well plates (150 μL/well) and incubated with several concentrations of compounds 1 and 2 for 24 h. After adding 50 μL MTT solution (1 mg/mL in PBS), the culture was incubated at 37 °C for 4 h, following which 200 μL DMSO was added to dissolve the formazan. The plate was then read on an ELISA microplate reader at 595 nm absorbance.
4. Conclusions
The current study highlighted the discovery of the first isomalabaricane derivatives with tetrahydrofuran moiety. Although the identified functionalities did not lead to a dramatic increase in the anti-proliferative properties, the chemical diversity of this class of triterpenes was enriched by these compounds with such unique structures.
Supplementary Materials
The following are available online at https://www.mdpi.com/article/10.3390/md19040206/s1, ESIMS, HRESIMS, IR, 1D, 2D, and DEPT NMR spectra of 1–2.
Author Contributions
K.-H.L., W.-C.W. and J.-H.S. conceived and designed the experiments; Z.-H.H., K.-H.L. and J.-H.S. performed the sample collections, extraction, isolation, structures determination, and qualitative HPLC analysis; the pharmacological experiments were carried out by W.-C.W.; K.-H.L., W.-C.W. and J.-H.S. contributed reagents and analysis tools; B.-R.P., K.-H.L., M.E.-S., W.-C.W. and J.-H.S. participated in data interpretation, wrote the manuscript and revised the paper. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by the grant from the Ministry of Science and Technology, Taiwan (MOST 108-2320-B-291-001-MY3 and MOST 110-2320-B-038-013); Ministry of Education, Taiwan (DP2-110-21121-01-N-12-03); and from Taipei Medical University (TMU109-AE1-B15).
Institutional Review Board Statement
Not applicable.
Data Availability Statement
The data presented in this study are available on request from the corresponding author.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Tasdemir, D.; Mangalindan, G.C.; Concepcion, G.P.; Verbitski, S.M.; Rabindran, S.; Miranda, M.; Greenstein, M.; Hooper, J.N.; Harper, M.K.; Ireland, C.M. Bioactive isomalabaricane triterpenes from the marine sponge Rhabdastrella globostellata. J. Nat. Prod. 2002, 65, 210–214. [Google Scholar] [CrossRef] [PubMed]
- Clement, J.A.; Li, M.; Hecht, S.M.; Kingston, D.G. Bioactive isomalabaricane triterpenoids from Rhabdastrella globostellata that stabilize the binding of DNA polymerase beta to DNA. J. Nat. Prod. 2006, 69, 373–376. [Google Scholar] [CrossRef] [PubMed]
- Lv, F.; Deng, Z.; Li, J.; Fu, H.; van Soest, R.W.; Proksch, P.; Lin, W. Isomalabaricane-type compounds from the marine sponge Rhabdastrella aff. distincta. J. Nat. Prod. 2004, 67, 2033–2036. [Google Scholar] [CrossRef] [PubMed]
- Meragelman, K.M.; McKee, T.C.; Boyd, M.R. New cytotoxic isomalabaricane triterpenes from the sponge Jaspis species. J. Nat. Prod. 2001, 64, 389–392. [Google Scholar] [CrossRef] [PubMed]
- Aoki, S.; Sanagawa, M.; Watanabe, Y.; Setiawan, A.; Arai, M.; Kobayashi, M. Novel isomarabarican triterpenes, exhibiting selective anti-proliferative activity against vascular endothelial cells, from marine sponge Rhabdastrella globostellata. Bioorg. Med. Chem. 2007, 15, 4818–4828. [Google Scholar] [CrossRef] [PubMed]
- McCormick, J.L.; McKee, T.C.; Cardellina, J.H.; Leid, M.; Boyd, M.R. Cytotoxic triterpenes from a marine sponge, Stelletta sp. J. Nat. Prod. 1996, 59, 1047–1050. [Google Scholar] [CrossRef] [PubMed]
- Prabhakar, S.A.; Rajesh, K.G.; Adavi, R.V.B.; Raghuram, R.A.; Ravi, K.B.; Appa, R.V.N.A. New malabaricane triterpenes from the oleoresin of Ailanthus malabarica. Fitoterapia 2015, 100, 166–173. [Google Scholar]
- William, F.P.; Iain, C.P.; Ashok, G.B.; Sukh, D. The structure of malabricol. Tetrahedron Lett. 1979, 43, 4153–4154. [Google Scholar]
- Miyabi, H.; Kazuomi, T.; Toshiyuki, H.; Hiroaki, O.; Tatsuhiko, F.; Shin-ichi, A.; Yusuke, T.; Ryuji, I.; Masaharu, K.; Matsumi, D.; et al. Cytotoxic isomalabaricane derivatives and a monocyclic triterpene glycoside from the sponge Rhabdastrella globostellata. J. Nat. Prod. 2010, 73, 1512–1518. [Google Scholar]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).