Screening of Marine Bioactive Antimicrobial Compounds for Plant Pathogens
Abstract
1. Introduction
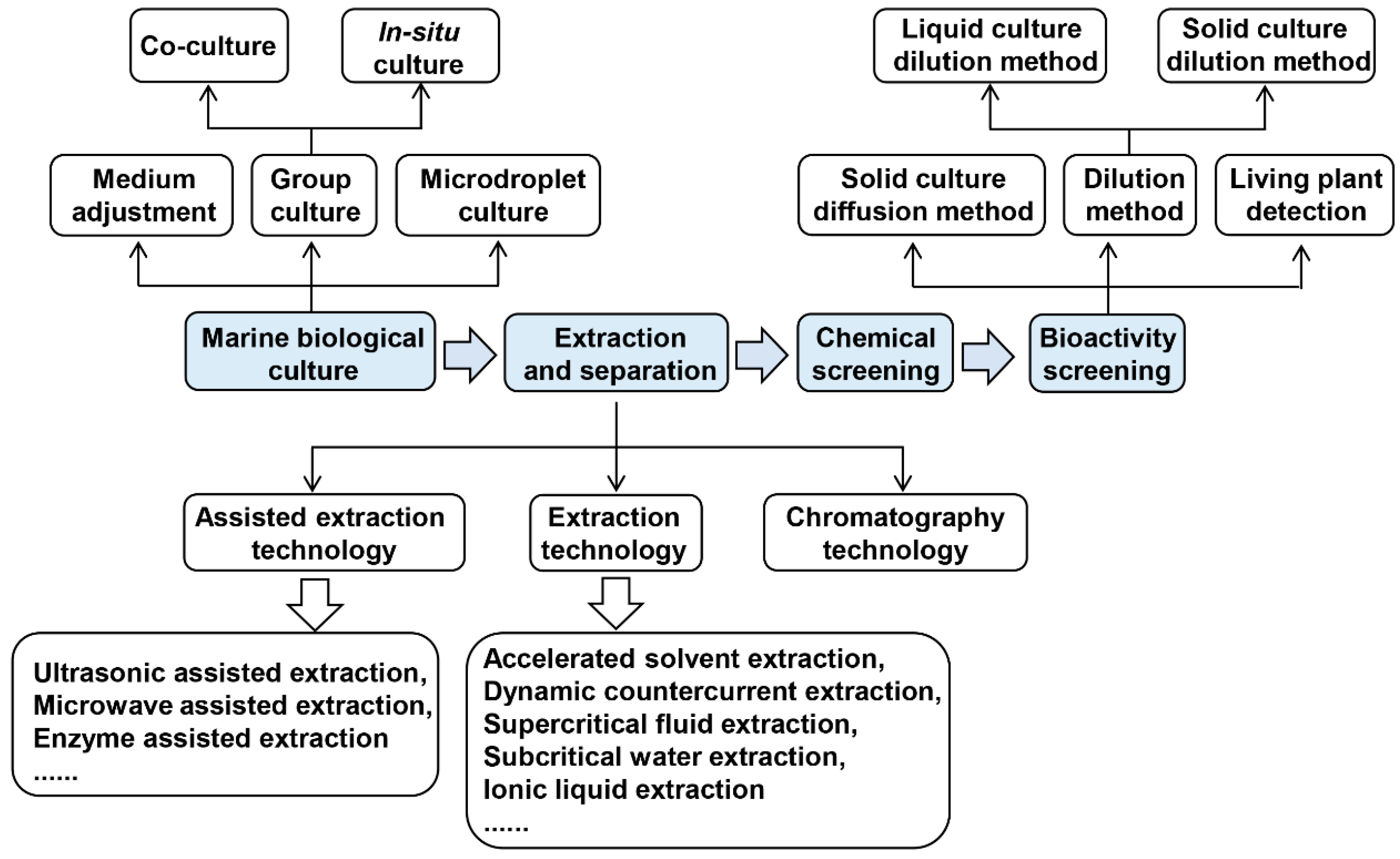
2. Methodologies of Screening Marine Antimicrobial Substances
2.1. Cultivation of Marine Organisms
2.1.1. Medium Optimization
2.1.2. Co-Cultivation
2.1.3. In Situ Cultivation
2.1.4. Microdroplets Cultivation
2.2. Extraction and Separation of Marine Natural Products
2.2.1. Auxiliary Extraction Technique
Ultrasound-Assisted Extraction (UAE)
Microwave-Assisted Extraction (MAE)
Enzyme-Assisted Extraction (EAE)
2.2.2. Extraction Technique
Accelerated Solvent Extraction (ASE)
Dynamic Countercurrent Extraction (DCE)
Supercritical Fluid Extraction (SFE)
Subcritical Water Extraction (SWE)
Ionic Liquid Extraction (ILE)
2.2.3. Chromatography Technique
Separation Based on Distribution Ratio of the Substance
Separation based on Adsorption of the Substance
Separation Based on Molecular Size of the Substance
2.3. Chemical Screening
2.4. Biological Activity Screening
2.4.1. Solid Culture Diffusion Method
2.4.2. Dilution Method
Liquid Culture Dilution Method
Solid Culture Dilution Method
2.4.3. Living Plant Detection
2.5. Active Substance Mechanism Research
2.5.1. Microscopic Observation
2.5.2. Dyeing Methods
2.5.3. Omics Methods
2.5.4. Molecular Docking Methods
3. Active Substances Derived from Different Marine Sources
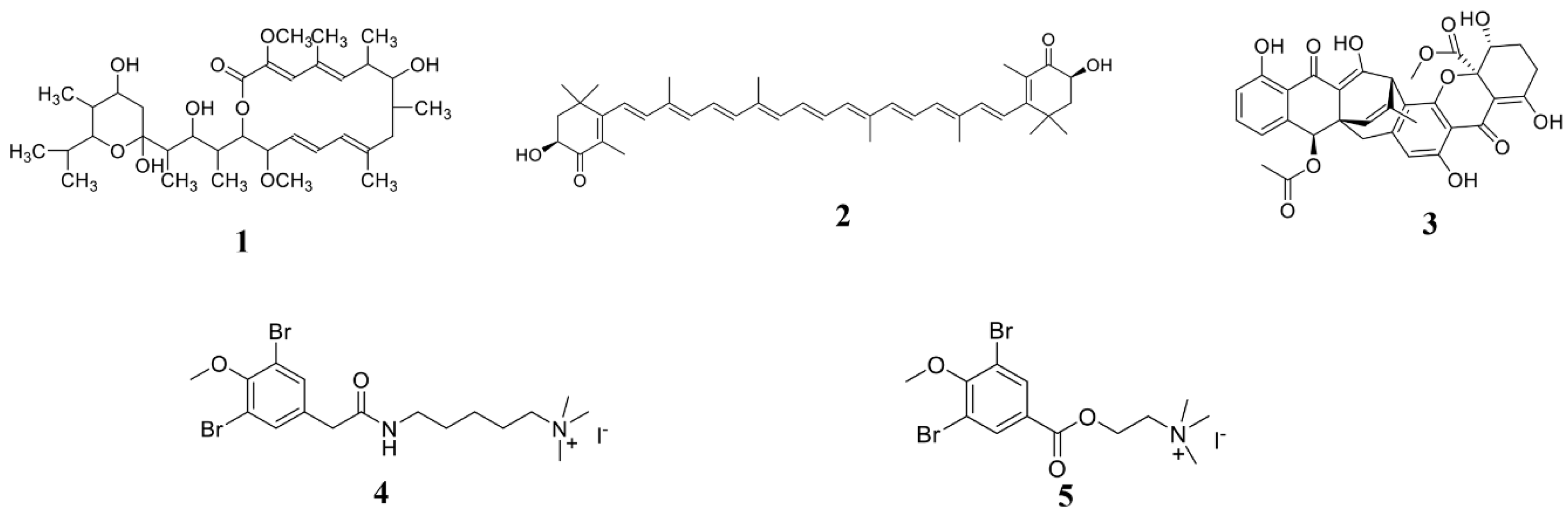
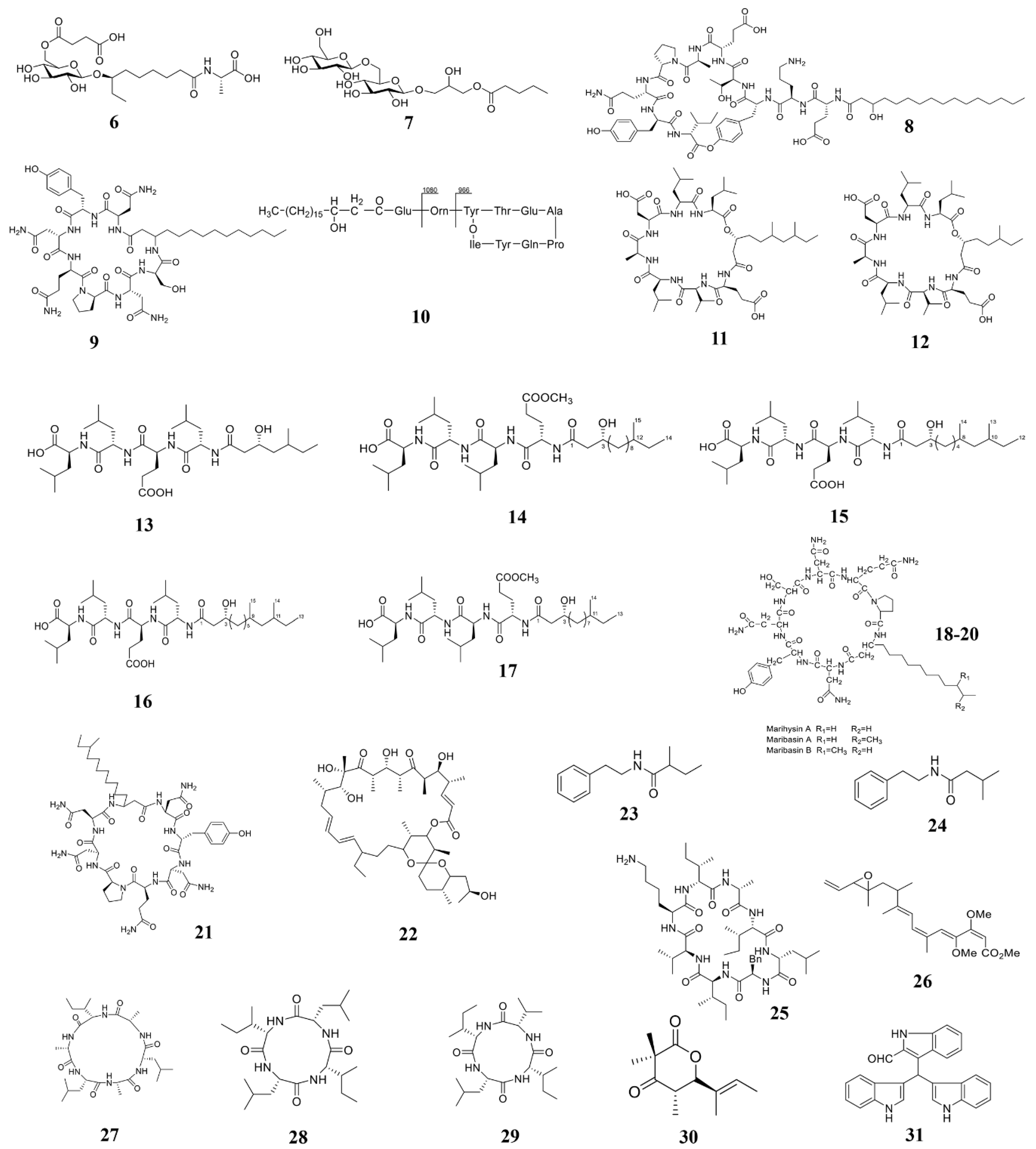
3.1. Marine Bacterium-Derived Anti-Microbial Compounds
3.1.1. Active Substances from Marine Bacillus
3.1.2. Active Substances from Marine Streptomyces
3.1.3. Active substances from other marine bacteria
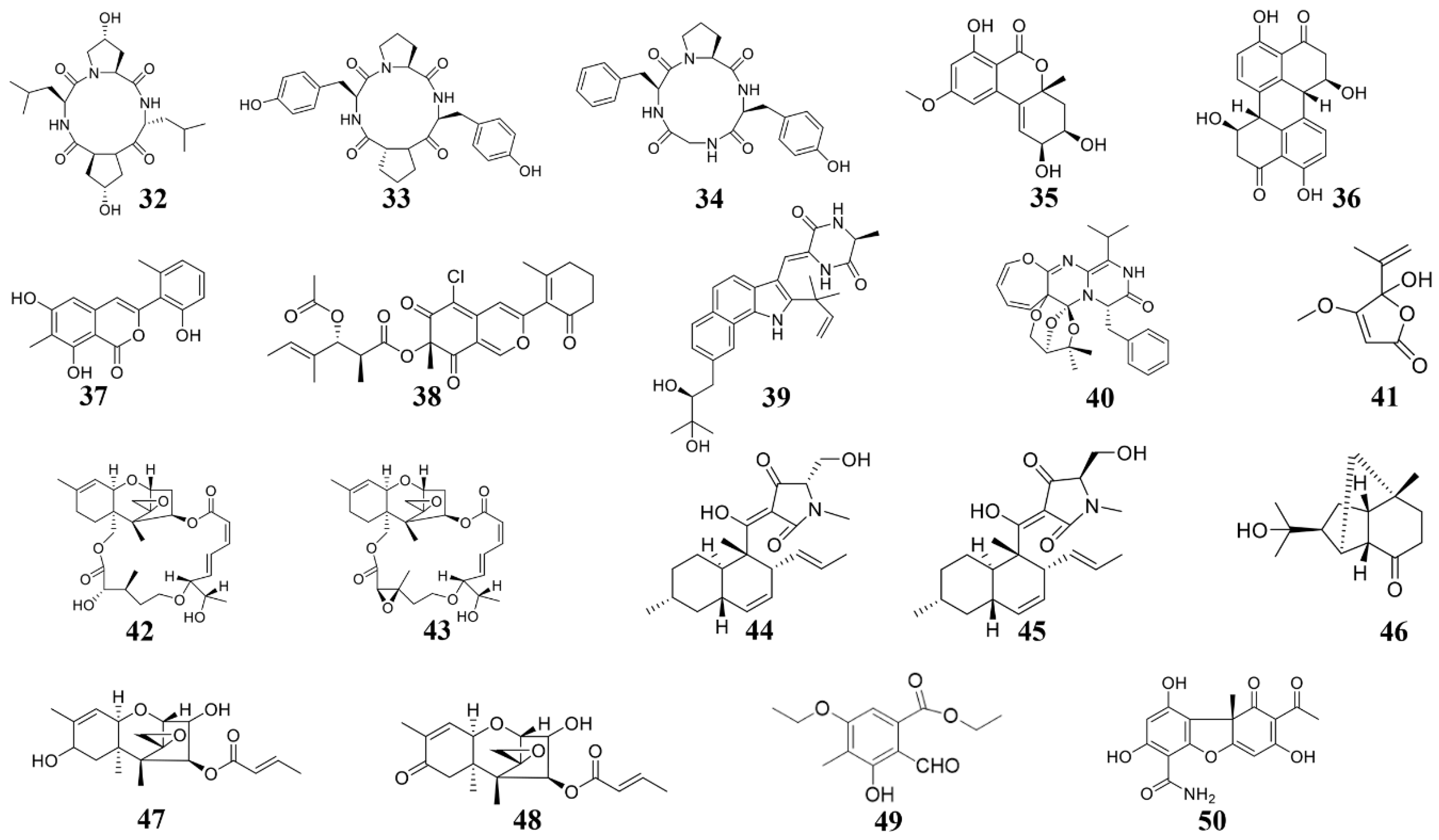
3.2. Marine Fungus-Derived Anti-Microbial Compounds
3.2.1. Active Substances from Marine Alternaria
3.2.2. Active Substances from Marine Pleosporales
3.2.3. Active Substances from Marine Parasitic Fungus
3.2.4. Active Substances from Other Marine Fungus
3.3. Sponge-Derived Anti-Microbial Compounds
3.4. Seaweed-Derived Anti-Microbial Compounds
3.5. Marine Animal-Derived Anti-Microbial Compounds
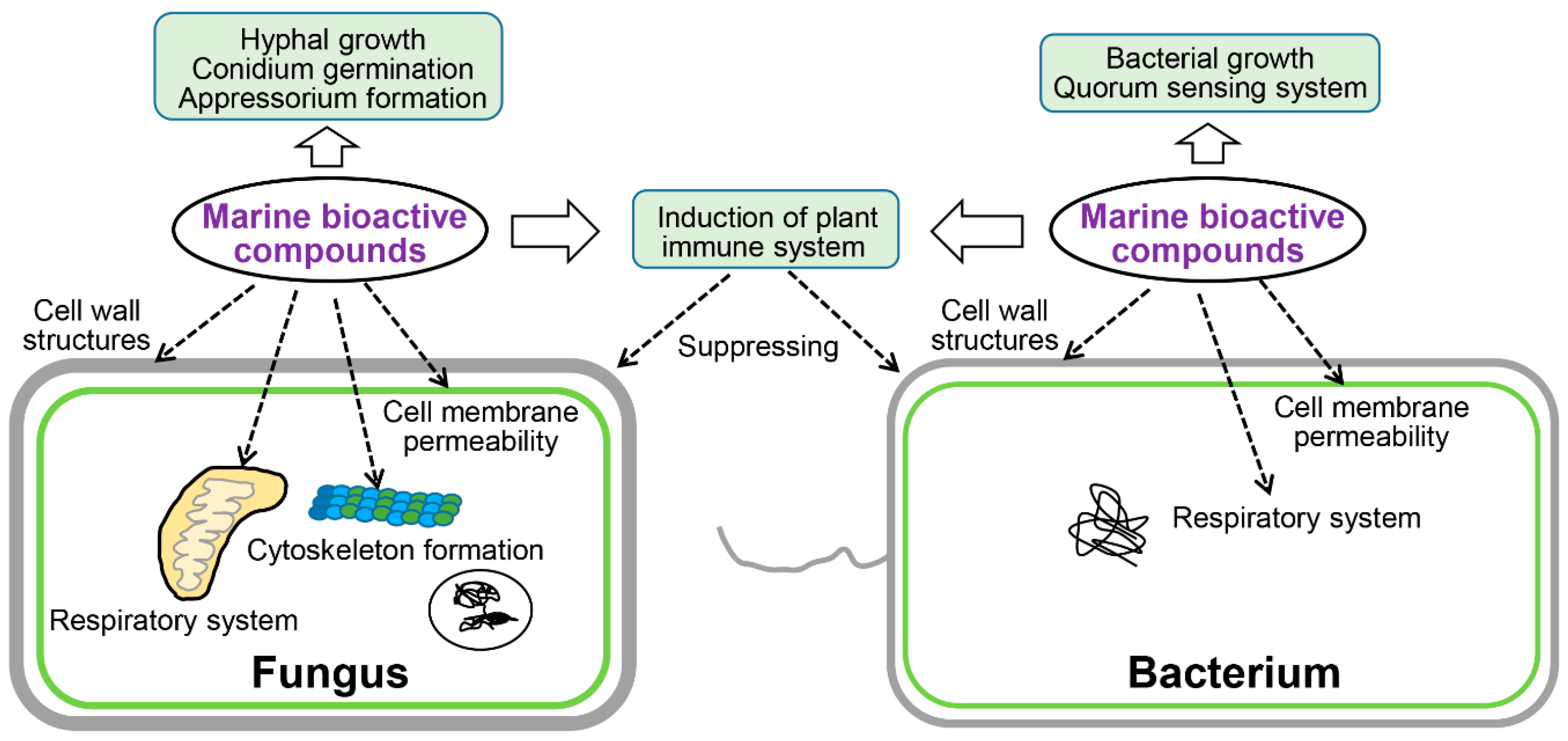
4. Microbicidal Mechanisms of Marine-Derived Bioactive Substances
4.1. Affect Cell Wall Structures
4.2. Affect Cell Membrane Permeability
4.3. Affect Fatty Acid Metabolism
4.4. Affect Respiratory System
4.5. Affect Cytoskeleton Formation
4.6. Affect Bacterial QS System
4.7. Induction of the Plant Immune System
5. Potential of Large-Scale Application of Marine Natural Products in Agricultural Production
5.1. Chemical Synthesis Methods
5.2. Synthetic Biology Methods
6. Prospects and Challenges
Author Contributions
Funding
Conflicts of Interest
References
- Jomo, K.S.; Chowdhury, A. COVID-19 Pandemic Recession and Recovery. Development (Rome) 2020, 63, 1–12. [Google Scholar]
- Addrah, M.E.; Zhang, Y.; Zhang, J.; Liu, L.; Zhou, H.; Chen, W.; Zhao, J. Fungicide treatments to control seed-borne fungi of sunflower seeds. Pathogens 2019, 9, 29. [Google Scholar] [CrossRef] [PubMed]
- Akinsanya, B.; Olaleru, F.; Samuel, O.B.; Akeredolu, E.; Isibor, P.O.; Adeniran, O.S.; Saliu, J.K.; Akhiromen, D.I. Bioaccumulation of organochlorine pesticides, Procamallanus sp. (Baylis, 1923) infections, and microbial colonization in African snakehead fish sampled from Lekki Lagoon, Lagos, Nigeria. Braz. J. Biol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Cheng, X.; Man, X.; Wang, Z.; Liang, L.; Zhang, F.; Wang, Z.; Liu, P.; Lei, B.; Hao, J.; Liu, X. Fungicide SYP-14288 Inducing multidrug resistance in Rhizoctonia solani. Plant Dis. 2020, 104, 2563–2570. [Google Scholar] [CrossRef] [PubMed]
- Gerber, P.F.; Gould, N.; McGahan, E. Potential contaminants and hazards in alternative chicken bedding materials and proposed guidance levels: A review. Poult. Sci 2020, 99, 6664–6684. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Hu, J.; Li, W.; Gao, Y.; Tian, Y. Changes of embryonic development, locomotor activity, and metabolomics in zebrafish co-exposed to chlorpyrifos and deltamethrin. J. Appl. Toxicol. 2020. [Google Scholar] [CrossRef]
- Santana, M.S.; Sandrini-Neto, L.; Di Domenico, M.; Prodocimo, M.M. Pesticide effects on fish cholinesterase variability and mean activity: A meta-analytic review. Sci. Total Environ. 2020, 757, 143829. [Google Scholar] [CrossRef]
- Aiello, D.; Vitale, A.; Perrone, G.; Tessitori, M.; Polizzi, G. Can biological control agents reduce multiple fungal infections causing decline of milkwort in ornamental nursery? Plants (Basel) 2020, 9, 1682. [Google Scholar] [CrossRef]
- Zhang, H.; Xie, W.; Hou, F.; Hu, J.; Yao, Z.; Zhao, Q.; Zhang, D. Response of microbial community to the lysis of Phaeocystis globosa induced by a biological algicide, prodigiosin. Environ. Pollut. 2020, 265, 115047. [Google Scholar] [CrossRef]
- Escribano-Viana, R.; Lopez-Alfaro, I.; Lopez, R.; Santamaria, P.; Gutierrez, A.R.; Gonzalez-Arenzana, L. Impact of chemical and biological fungicides applied to grapevine on grape biofilm, must, and wine microbial diversity. Front. Microbiol 2018, 9, 59. [Google Scholar] [CrossRef]
- Cordova, L.G.; Dalla Lana, F.; Paul, P.A.; Peres, N.A. A quantitative synthesis of the efficacy and profitability of conventional and biological fungicides for botrytis fruit rot management on strawberry in Florida. Plant Dis. 2019, 103, 2505–2511. [Google Scholar] [CrossRef] [PubMed]
- Cardoso, J.; Nakayama, D.G.; Sousa, E.; Pinto, E. Marine-derived compounds and prospects for their antifungal application. Molecules 2020, 25, 5856. [Google Scholar] [CrossRef] [PubMed]
- Khrunyk, Y.; Lach, S.; Petrenko, I.; Ehrlich, H. Progress in modern marine biomaterials research. Mar. Drugs 2020, 18, 589. [Google Scholar] [CrossRef] [PubMed]
- Stonik, V.A.; Makarieva, T.N.; Shubina, L.K. Antibiotics from marine bacteria. Biochemistry (Mosc) 2020, 85, 1362–1373. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Guo, J.; Wang, Z.; Liu, Y.; Song, H.; Wang, Q. Marine natural products for drug discovery: First discovery of kealiinines A-C and their derivatives as novel antiviral and antiphytopathogenic fungus agents. J. Agric. Food Chem. 2018, 66, 7310–7318. [Google Scholar] [CrossRef]
- Barreca, M.; Spano, V.; Montalbano, A.; Cueto, M.; Diaz Marrero, A.R.; Deniz, I.; Erdogan, A.; Lukic Bilela, L.; Moulin, C.; Taffin-de-Givenchy, E.; et al. Marine anticancer agents: An overview with a particular focus on their chemical classes. Mar. Drugs 2020, 18, 619. [Google Scholar] [CrossRef]
- Bracegirdle, J.; Keyzers, R.A. Marine-derived polyaromatic butenolides—isolation, synthesis and biological evaluations. Curr. Pharm. Des. 2020, 26, 4351–4361. [Google Scholar] [CrossRef]
- Santos, J.D.; Vitorino, I.; Reyes, F.; Vicente, F.; Lage, O.M. From ocean to medicine: Pharmaceutical applications of metabolites from marine bacteria. Antibiotics (Basel) 2020, 9, 455. [Google Scholar] [CrossRef]
- Teng, Y.F.; Xu, L.; Wei, M.Y.; Wang, C.Y.; Gu, Y.C.; Shao, C.L. Recent progresses in marine microbial-derived antiviral natural products. Arch. Pharm. Res. 2020, 43, 1215–1229. [Google Scholar] [CrossRef] [PubMed]
- Bunyapaiboonsri, T.; Yoiprommarat, S.; Suntivich, R.; Preedanon, S.; Komwijit, S.; Teerawatananond, T.; Sakayaroj, J. A cyclic lipodepsipeptide, a spirolactone, and a chromanone from the marine fungus Verruculina enalia (Kohlm.) Kohlm. & Volkm.-Kohlm. BCC 22226. Tetrahedron 2020, 76, 131497. [Google Scholar]
- Du, F.Y.; Ju, G.L.; Xiao, L.; Zhou, Y.M.; Wu, X. Sesquiterpenes and cyclodepsipeptides from marine-derived fungus Trichoderma longibrachiatum and their antagonistic activities against soil-borne pathogens. Mar. Drugs 2020, 18, 165. [Google Scholar] [CrossRef] [PubMed]
- Huang, R.-H.; Lin, W.; Zhang, P.; Liu, J.-Y.; Wang, D.; Li, Y.-Q.; Wang, X.-Q.; Zhang, C.-S.; Li, W.; Zhao, D.-L. Anti-phytopathogenic bacterial metabolites from the seaweed-derived fungus Aspergillus sp. D40. Front. Mar. Sci. 2020, 7, 7. [Google Scholar] [CrossRef]
- Betancur, L.A.; Forero, A.M.; Vinchira-Villarraga, D.M.; Cardenas, J.D.; Romero-Otero, A.; Chagas, F.O.; Pupo, M.T.; Castellanos, L.; Ramos, F.A. NMR-based metabolic profiling to follow the production of anti-phytopathogenic compounds in the culture of the marine strain Streptomyces sp. PNM-9. Microbiol. Res. 2020, 239, 126507. [Google Scholar] [CrossRef]
- Chakraborty, M.; Mahmud, N.U.; Gupta, D.R.; Tareq, F.S.; Shin, H.J.; Islam, T. Inhibitory effects of linear lipopeptides from a marine Bacillus subtilis on the wheat blast fungus Magnaporthe oryzae triticum. Front. Microbiol. 2020, 11, 665. [Google Scholar] [CrossRef] [PubMed]
- Fenical, W.B. Marine microbial natural products: The evolution of a new field of science. J. Antibiot. (Tokyo) 2020, 73, 481–487. [Google Scholar] [CrossRef] [PubMed]
- Berdy, B.; Spoering, A.L.; Ling, L.L.; Epstein, S.S. In situ cultivation of previously uncultivable microorganisms using the ichip. Nat. Protoc. 2017, 12, 2232–2242. [Google Scholar] [CrossRef]
- Nichols, D.; Cahoon, N.; Trakhtenberg, E.M.; Pham, L.; Mehta, A.; Belanger, A.; Kanigan, T.; Lewis, K.; Epstein, S.S. Use of ichip for high-throughput in situ cultivation of "uncultivable" microbial species. Appl. Environ. Microbiol. 2010, 76, 2445–2450. [Google Scholar] [CrossRef]
- Sherpa, R.T.; Reese, C.J.; Montazeri Aliabadi, H. Application of ichip to grow “uncultivable” microorganisms and its impact on antibiotic discovery. J. Pharm. Pharm. Sci. 2015, 18, 303–315. [Google Scholar] [CrossRef]
- Shleeva, M.O.; Kudykina, Y.K.; Vostroknutova, G.N.; Suzina, N.E.; Mulyukin, A.L.; Kaprelyants, A.S. Dormant ovoid cells of Mycobacterium tuberculosis are formed in response to gradual external acidification. Tuberculosis (Edinb) 2011, 91, 146–154. [Google Scholar] [CrossRef]
- Tanner, R.S. Manual of Environmental Microbiology, 3rd ed.; American Society of Microbiology: Edinburgh, UK, 2007. [Google Scholar]
- Schleifer, K.H. Classification of Bacteria and Archaea: Past, present and future. Syst. Appl. Microbiol. 2009, 32, 533–542. [Google Scholar] [CrossRef]
- Webster, J.; Weber, R. Introduction to Fungi; Cambridge University Press: Cambridge, UK, 2007. [Google Scholar]
- Tripp, H.J.; Kitner, J.B.; Schwalbach, M.S.; Dacey, J.W.; Wilhelm, L.J.; Giovannoni, S.J. SAR11 marine bacteria require exogenous reduced sulphur for growth. Nature 2008, 452, 741–744. [Google Scholar] [CrossRef] [PubMed]
- Sizova, M.; Hohmann, T.; Hazen, A.; Paster, B.; Halem, S.; Murphy, C.; Panikov, N.; Epstein, S. New approaches for isolation of previously uncultivated oral bacteria. Appl. Environ. Microb. 2012, 78, 194–203. [Google Scholar] [CrossRef] [PubMed]
- Puspita, I.D.; Kamagata, Y.; Tanaka, M.; Asano, K.; Nakatsu, C.H. Are uncultivated bacteria really uncultivable? Microbes. Environ. 2012, 27, 356–366. [Google Scholar] [CrossRef]
- Santini, J.M.; Kappler, U.; Ward, S.A.; Honeychurch, M.J.; vanden Hoven, R.N.; Bernhardt, P.V. The NT-26 cytochrome c552 and its role in arsenite oxidation. Biochim. Biophys. Acta 2007, 1767, 189–196. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Alain, K.; Querellou, J. Cultivating the uncultured: Limits, advances and future challenges. Extremophiles 2009, 13, 583–594. [Google Scholar] [CrossRef]
- Kwak, M.J.; Kong, H.G.; Choi, K.; Kwon, S.K.; Song, J.Y.; Lee, J.; Lee, P.A.; Choi, S.Y.; Seo, M.; Lee, H.J.; et al. Rhizosphere microbiome structure alters to enable wilt resistance in tomato. Nat. Biotechnol. 2018, 36, 1100–1109. [Google Scholar] [CrossRef] [PubMed]
- Nadell, C.D.; Xavier, J.B.; Foster, K.R. The sociobiology of biofilms. FEMS Microbiol. Rev. 2009, 33, 206–224. [Google Scholar] [CrossRef] [PubMed]
- Crocetti, G.R.; Banfield, J.F.; Keller, J.; Bond, P.L.; Blackall, L.L. Glycogen-accumulating organisms in laboratory-scale and full-scale wastewater treatment processes. Microbiology (Reading) 2002, 148, 3353–3364. [Google Scholar] [CrossRef]
- Nichols, D.; Lewis, K.; Orjala, J.; Mo, S.; Ortenberg, R.; O’Connor, P.; Zhao, C.; Vouros, P.; Kaeberlein, T.; Epstein, S.S. Short peptide induces an “uncultivable” microorganism to grow in vitro. Appl. Environ. Microbiol. 2008, 74, 4889–4897. [Google Scholar] [CrossRef]
- Tanaka, Y.; Tamaki, H.; Tanaka, K.; Tozawa, E.; Matsuzawa, H.; Toyama, T.; Kamagata, Y.; Mori, K. “Duckweed-microbe co-cultivation method” for isolating a wide variety of microbes including taxonomically novel microbes. Microbes. Environ. 2018, 33, 402–406. [Google Scholar] [CrossRef]
- Kaeberlein, T.; Lewis, K.; Epstein, S.S. Isolating “uncultivable” microorganisms in pure culture in a simulated natural environment. Science 2002, 296, 1127–1129. [Google Scholar] [CrossRef] [PubMed]
- Jung, D.; Seo, E.Y.; Epstein, S.S.; Joung, Y.; Han, J.; Parfenova, V.V.; Belykh, O.I.; Gladkikh, A.S.; Ahn, T.S. Application of a new cultivation technology, I-tip, for studying microbial diversity in freshwater sponges of Lake Baikal, Russia. FEMS Microbiol. Ecol. 2014, 90, 417–423. [Google Scholar] [CrossRef] [PubMed]
- Zengler, K.; Walcher, M.; Clark, G.; Haller, I.; Toledo, G.; Holland, T.; Mathur, E.J.; Woodnutt, G.; Short, J.M.; Keller, M. High-throughput cultivation of microorganisms using microcapsules. Methods Enzymol. 2005, 397, 124–130. [Google Scholar] [PubMed]
- Jiang, C.Y.; Dong, L.; Zhao, J.K.; Hu, X.; Shen, C.; Qiao, Y.; Zhang, X.; Wang, Y.; Ismagilov, R.F.; Liu, S.J.; et al. High-throughput single-cell cultivation on microfluidic streak plates. Appl. Environ. Microbiol. 2016, 82, 2210–2218. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Cao, J.; Yan, X.; Cheng, W.; Wang, X.; Yang, R.; Guo, Y. Discovery of natural product-based fungicides (ii): Semisynthesis and biological activity of sarisan attached 3-phenylisoxazolines as antifungal agents. Chem. Biodivers. 2020. [Google Scholar] [CrossRef]
- Aliano-Gonzalez, M.J.; Jarillo, J.A.; Carrera, C.; Ferreiro-Gonzalez, M.; Alvarez, J.A.; Palma, M.; Ayuso, J.; Barbero, G.F.; Espada-Bellido, E. Optimization of a novel method based on ultrasound-assisted extraction for the quantification of anthocyanins and total phenolic compounds in blueberry samples (Vaccinium corymbosum L.). Foods 2020, 9, 1763. [Google Scholar] [CrossRef]
- Quiroz, J.Q.; Torres, A.C.; Ramirez, L.M.; Garcia, M.S.; Gomez, G.C.; Rojas, J. Optimization of the microwave-assisted extraction process of bioactive compounds from annatto seeds (Bixa orellana L.). Antioxidants (Basel) 2019, 8, 37. [Google Scholar] [CrossRef]
- Alboofetileh, M.; Rezaei, M.; Tabarsa, M.; You, S. Bioactivities of Nizamuddinia zanardinii sulfated polysaccharides extracted by enzyme, ultrasound and enzyme-ultrasound methods. J. Food Sci. Technol. 2019, 56, 1212–1220. [Google Scholar] [CrossRef]
- Amarante, S.J.; Catarino, M.D.; Marcal, C.; Silva, A.M.S.; Ferreira, R.; Cardoso, S.M. Microwave-assisted extraction of phlorotannins from Fucus vesiculosus. Mar. Drugs 2020, 18, 559. [Google Scholar] [CrossRef]
- Sebastian, J.; Rouissi, T.; Brar, S.K.; Hegde, K.; Verma, M. Microwave-assisted extraction of chitosan from Rhizopus oryzae NRRL 1526 biomass. Carbohydr. Polym. 2019, 219, 431–440. [Google Scholar] [CrossRef]
- Dominguez-Rodriguez, G.; Marina, M.L.; Plaza, M. Enzyme-assisted extraction of bioactive non-extractable polyphenols from sweet cherry (Prunus avium L.) pomace. Food Chem. 2021, 339, 128086. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, T.T.; Mikkelsen, M.D.; Tran, V.H.N.; Trang, V.T.D.; Rhein-Knudsen, N.; Holck, J.; Rasin, A.B.; Cao, H.T.T.; Van, T.T.T.; Meyer, A.S. Enzyme-assisted fucoidan extraction from brown macroalgae Fucus distichus subsp. evanescens and Saccharina latissima. Mar. Drugs 2020, 18, 296. [Google Scholar] [CrossRef] [PubMed]
- Habeebullah, S.F.K.; Alagarsamy, S.; Sattari, Z.; Al-Haddad, S.; Fakhraldeen, S.; Al-Ghunaim, A.; Al-Yamani, F. Enzyme-assisted extraction of bioactive compounds from brown seaweeds and characterization. J. Appl. Phycol. 2019, 1–15. [Google Scholar] [CrossRef]
- Olech, M.; Lyko, L.; Nowak, R. Influence of accelerated solvent extraction conditions on the LC-ESI-MS/MS polyphenolic profile, triterpenoid content, and antioxidant and anti-lipoxygenase activity of Rhododendron luteum sweet leaves. Antioxidants (Basel) 2020, 9, 822. [Google Scholar] [CrossRef] [PubMed]
- Johnson, T.A.; Sohn, J.; Vaske, Y.M.; White, K.N.; Cohen, T.L.; Vervoort, H.C.; Tenney, K.; Valeriote, F.A.; Bjeldanes, L.F.; Crews, P. Myxobacteria versus sponge-derived alkaloids: The bengamide family identified as potent immune modulating agents by scrutiny of LC-MS/ELSD libraries. Bioorg. Med. Chem. 2012, 20, 4348–4355. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Liu, C.; Pan, Y.; Qi, Y.; Li, Y.; Li, S. Ultrasound-assisted dynamic extraction coupled with parallel countercurrent chromatography for simultaneous extraction, purification, and isolation of phytochemicals: Application to isoflavones from red clover. Anal. Bioanal. Chem. 2015, 407, 4597–4606. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Y.; He, X.; Wang, T.; Zhang, X.; Li, Z.; Xu, X.; Zhang, W.; Yan, X.; Li, S.; He, S. Efficient preparation of bafilomycin a1 from marine streptomyces lohii fermentation using three-phase extraction and high-speed counter-current chromatography. Mar. Drugs 2020, 18, 332. [Google Scholar] [CrossRef]
- He, X.; Ding, L.; Yi, M.; Xu, J.; Zhou, X.; Zhang, W.; He, S. Separation of five diketopiperazines from the marine fungus Alternaria alternate HK-25 by high-speed counter-current chromatography. J. Sep. Sci. 2019, 42, 2510–2516. [Google Scholar] [CrossRef]
- Bader, C.D.; Neuber, M.; Panter, F.; Krug, D.; Muller, R. Supercritical fluid extraction enhances discovery of secondary metabolites from myxobacteria. Anal. Chem. 2020, 92, 15403–15411. [Google Scholar] [CrossRef]
- Ara, K.M.; Jowkarderis, M.; Raofie, F. Optimization of supercritical fluid extraction of essential oils and fatty acids from flixweed (Descurainia Sophia L.) seed using response surface methodology and central composite design. J. Food Sci. Technol. 2015, 52, 4450–4458. [Google Scholar] [CrossRef]
- Messina, C.M.; Manuguerra, S.; Renda, G.; Santulli, A. Biotechnological applications for the sustainable use of marine by-products: In vitro antioxidant and pro-apoptotic effects of astaxanthin extracted with supercritical co2 from Parapeneus longirostris. Mar. Biotechnol. (NY) 2019, 21, 565–576. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Li, Y.; Li, Y.; Ye, D.; Yuan, L.; Sun, Y.; Han, D.; Hu, Q. Solid matrix-supported supercritical CO(2) enhances extraction of gamma-linolenic acid from the Cyanobacterium arthrospira (spirulina) platensis and bioactivity evaluation of the molecule in zebrafish. Mar. Drugs 2019, 17, 203. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Jing, J.; Wang, J.; Xu, J.; Hu, Z. Extraction of low methoxyl pectin from fresh sunflower heads by subcritical water extraction. ACS Omega 2020, 5, 15095–15104. [Google Scholar] [CrossRef] [PubMed]
- Pangestuti, R.; Getachew, A.T.; Siahaan, E.A.; Chun, B.S. E3S Web of Conferences; Les Ulis Cedex A: Les Ulis, France, 2020; p. 03013. [Google Scholar]
- Alboofetileh, M.; Rezaei, M.; Tabarsa, M.; You, S.; Mariatti, F.; Cravotto, G. Subcritical water extraction as an efficient technique to isolate biologically-active fucoidans from Nizamuddinia zanardinii. Int. J. Biol. Macromol. 2019, 128, 244–253. [Google Scholar] [CrossRef] [PubMed]
- Okamura, H.; Hirayama, N. Recent progress in ionic liquid extraction for the separation of rare earth elements. Anal. Sci. 2020, 37, 119–130. [Google Scholar] [CrossRef]
- Liu, Y.H.; Alimujiang, A.; Wang, X.; Luo, S.W.; Balamurugan, S.; Yang, W.D.; Liu, J.S.; Zhang, L.; Li, H.Y. Ethanol induced jasmonate pathway promotes astaxanthin hyperaccumulation in Haematococcus pluvialis. Bioresour. Technol. 2019, 289, 121720. [Google Scholar] [CrossRef]
- Wong, L.L.; Natarajan, G.; Boleij, M.; Thi, S.S.; Winnerdy, F.R.; Mugunthan, S.; Lu, Y.; Lee, J.M.; Lin, Y.; van Loosdrecht, M.; et al. Extracellular protein isolation from the matrix of anammox biofilm using ionic liquid extraction. Appl. Microbiol. Biotechnol. 2020, 104, 3643–3654. [Google Scholar] [CrossRef]
- Al-Rubaye, A.F.; Hameed, I.H.; Kadhim, M.J. A review: Uses of gas chromatography-Mass Spectrometry (GC-MS) technique for analysis of bioactive natural compounds of some plants. Int. J. Tox. Pharm. Res. 2017, 9, 81–85. [Google Scholar] [CrossRef]
- Gong, Y.; Huang, X.Y.; Pei, D.; Duan, W.D.; Zhang, X.; Sun, X.; Di, D.L. The applicability of high-speed counter current chromatography to the separation of natural antioxidants. J. Chromatogr. A 2020, 1623, 461150. [Google Scholar] [CrossRef]
- Chen, B.; Liu, Z.; Zhang, Y.; Li, W.; Sun, Y.; Wang, Y.; Wang, Y.; Sun, Y. Application of high-speed counter-current chromatography and HPLC to separate and purify of three polyacetylenes from Platycodon grandiflorum. J. Sep. Sci. 2018, 41, 789–796. [Google Scholar] [CrossRef]
- Wu, H.; Liu, J.; Duan, N.; Han, R.; Zhang, X.; Leng, X.; Liu, W.; Han, L.; Li, X.; Xing, S. Isolation and purification of an antibiotic polyketide jbir-99 from the marine fungus Meyerozyma guilliermondii by high-speed counter-current chromatography. Preprints 2019, 11, 20–28. [Google Scholar]
- Xia, M.; Liu, C.; Gao, L.; Lu, Y. One-step preparative separation of phytosterols from edible brown seaweed Sargassum horneri by high-speed countercurrent chromatography. Mar. Drugs 2019, 17, 691. [Google Scholar] [CrossRef] [PubMed]
- Vreeker, G.C.; Wuhrer, M. Reversed-phase separation methods for glycan analysis. Anal. Bioanal. Chem. 2017, 409, 359–378. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Sun, C. Fengycins, cyclic lipopeptides from marine bacillus subtilis strains, kill the plant-pathogenic fungus Magnaporthe grisea by inducing reactive oxygen species production and chromatin condensation. Appl. Environ. Microbiol. 2018, 84. [Google Scholar] [CrossRef]
- Ma, Z.; Hu, J. Plipastatin A1 produced by a marine sediment-derived Bacillus amyloliquefaciens SH-B74 contributes to the control of gray mold disease in tomato. 3 Biotech 2018, 8, 125. [Google Scholar] [CrossRef]
- Xu, S.; Lu, X.; Li, M.; Wang, J.; Li, H.; He, X.; Feng, J.; Wu, J.; Chen, K.; Ouyang, P. Separation of 5-aminovalerate from its bioconversion liquid by macroporous adsorption resin: Mechanism and dynamic separation. J. Chem. Technol. Biot. 2020, 95, 686–693. [Google Scholar] [CrossRef]
- Chen, J.; Liu, H.; Xia, Z.; Zhao, X.; Wu, Y.; An, M. Purification and structural analysis of the effective anti-TMV compound ε-poly-L-lysine produced by Streptomyces ahygroscopicus. Molecules 2019, 24, 1156. [Google Scholar] [CrossRef]
- He, Y.; Ding, Y.; Wu, Q.; Chen, M.; Zhao, S.E.; Zhang, J.; Wei, X.; Zhang, Y.; Bai, J.; Mo, S. Identification of the potential biological preservative tetramycin a-producing strain and enhancing its production. Front. Microbiol. 2020, 10, 2925. [Google Scholar] [CrossRef]
- Ma, R.; Zhou, R.; Tong, R.; Shi, S.; Chen, X. At-line hyphenation of high-speed countercurrent chromatography with Sephadex LH-20 column chromatography for bioassay-guided separation of antioxidants from vine tea (Ampelopsis grossedentata). J. Chromatogr. B 2017, 1040, 112–117. [Google Scholar] [CrossRef]
- Du, F.-Y.; Li, X.-M.; Sun, Z.-C.; Meng, L.-H.; Wang, B.-G. Secondary metabolites with agricultural antagonistic potentials from Beauveria felina, a marine-derived entomopathogenic fungus. J. Agr. Food Chem. 2020, 68, 14824–14831. [Google Scholar] [CrossRef]
- Huang, R.-H.; Gou, J.-Y.; Zhao, D.-L.; Wang, D.; Liu, J.; Ma, G.-Y.; Li, Y.-Q.; Zhang, C.-S. Phytotoxicity and anti-phytopathogenic activities of marine-derived fungi and their secondary metabolites. RSC Adv. 2018, 8, 37573–37580. [Google Scholar] [CrossRef]
- Her, N.; Amy, G.; Foss, D.; Cho, J.; Yoon, Y.; Kosenka, P. Optimization of method for detecting and characterizing NOM by HPLC− size exclusion chromatography with UV and on-line DOC detection. Environ. Sci. Technol. 2002, 36, 1069–1076. [Google Scholar] [CrossRef] [PubMed]
- Sherma, J.; Fried, B. Handbook of Thin-Layer Chromatography; CRC press: Boca Raton, FL, USA, 2003. [Google Scholar]
- De Hoffmann, E. Mass spectrometry. In Kirk-Othmer Encyclopedia of Chemical Technology; Wiley-Interscience: Hoboken, NJ, USA, 2000. [Google Scholar]
- Hore, P.J. Nuclear Magnetic Resonance; Oxford University Press: New York, NY, USA, 2015. [Google Scholar]
- Schweder, T.; Lindequist, U.; Lalk, M. Screening for new metabolites from marine microorganisms. Adv. Biochem. Eng. Biotechnol. 2005, 96, 1–48. [Google Scholar] [PubMed]
- Heatley, N.G. A method for the assay of penicillin. Biochem. J. 1944, 38, 61–65. [Google Scholar] [CrossRef]
- Schalchli, H.; Hormazabal, E.; Rubilar, O.; Briceno, G.; Mutis, A.; Zocolo, G.J.; Diez, M.C. Production of ligninolytic enzymes and some diffusible antifungal compounds by white-rot fungi using potato solid wastes as the sole nutrient source. J. Appl. Microbiol. 2017, 123, 886–895. [Google Scholar] [CrossRef]
- Zhao, Z.; Wang, Q.; Wang, K.; Brian, K.; Liu, C.; Gu, Y. Study of the antifungal activity of Bacillus vallismortis ZZ185 in vitro and identification of its antifungal components. Bioresour. Technol. 2010, 101, 292–297. [Google Scholar] [CrossRef]
- Bonev, B.; Hooper, J.; Parisot, J. Principles of assessing bacterial susceptibility to antibiotics using the agar diffusion method. J. Antimicrob. Chemother. 2008, 61, 1295–1301. [Google Scholar] [CrossRef]
- Afshari, M.A.; Shams-Ghahfarokhi, M.; Razzaghi-Abyaneh, M. Antifungal susceptibility and virulence factors of clinically isolated dermatophytes in Tehran, Iran. Iran. J. Microbiol. 2016, 8, 36–46. [Google Scholar]
- Monteiro, M.C.; de la Cruz, M.; Cantizani, J.; Moreno, C.; Tormo, J.R.; Mellado, E.; De Lucas, J.R.; Asensio, F.; Valiante, V.; Brakhage, A.A.; et al. A new approach to drug discovery: High-throughput screening of microbial natural extracts against Aspergillus fumigatus using resazurin. J. Biomol. Screen 2012, 17, 542–549. [Google Scholar] [CrossRef]
- Li, C.; Wang, J.; Luo, C.; Ding, W.; Cox, D.G. A new cyclopeptide with antifungal activity from the co-culture broth of two marine mangrove fungi. Nat. Prod. Res. 2014, 28, 616–621. [Google Scholar] [CrossRef]
- Huang, D.; Li, X.; Sun, L.; Xiu, Z.; Nishino, N. Synthesis, evaluation and molecular modeling of cyclic tetrapeptide histone deacetylase inhibitors as anticancer agents. J. Pept. Sci. 2012, 18, 242–251. [Google Scholar] [CrossRef] [PubMed]
- Benhadj, M.; Gacemi-Kirane, D.; Toussaint, M.; Hotel, L.; Bontemps, C.; Duval, R.E.; Aigle, B.; Leblond, P. Diversity and antimicrobial activities of Streptomyces isolates from Fetzara Lake, north eastern Algeria. Ann. Biol. Clin. (Paris) 2018, 76, 81–95. [Google Scholar] [CrossRef] [PubMed]
- Cao, F.; Meng, Z.H.; Mu, X.; Yue, Y.F.; Zhu, H.J. Absolute configuration of bioactive azaphilones from the marine-derived fungus Pleosporales sp. CF09-1. J. Nat. Prod. 2019, 82, 386–392. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Kong, Q.; Qin, C.; Chen, Y.; Chen, Y.; Lv, R.; Zhou, G. Identification of lipopeptides in Bacillus megaterium by two-step ultrafiltration and LC-ESI-MS/MS. AMB Express 2016, 6, 79. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.-J.; Sah, R.L.; Doong, J.-Y.H.; Grodzinsky, A.J. Fluorometric assay of DNA in cartilage explants using Hoechst 33258. Anal. Biochem. 1988, 174, 168–176. [Google Scholar] [CrossRef]
- Reers, M.; Smiley, S.T.; Mottola-Hartshorn, C.; Chen, A.; Lin, M.; Chen, L.B. Methods in Enzymology; Elsevier: Amsterdam, The Netherlands, 1995; Volume 260. [Google Scholar]
- Morris, G.M.; Lim-Wilby, M. Molecular Modeling of Proteins; Springer: Berlin/Heidelberg, Germany, 2008. [Google Scholar]
- Zhang, M.; Ding, X.; Kang, J.; Gao, Y.; Wang, Z.; Wang, Q. Marine natural product for pesticide candidate: Pulmonarin alkaloids as novel antiviral and anti-phytopathogenic-fungus agents. J. Agric. Food Chem. 2020, 68, 11350–11357. [Google Scholar] [CrossRef] [PubMed]
- Tareq, F.S.; Lee, H.S.; Lee, Y.J.; Lee, J.S.; Shin, H.J. Ieodoglucomide C and ieodoglycolipid, new glycolipids from a marine-derived bacterium Bacillus licheniformis 09IDYM23. Lipids 2015, 50, 513–519. [Google Scholar] [CrossRef]
- Ma, Z.; Zhang, S.; Sun, K.; Hu, J. Identification and characterization of a cyclic lipopeptide iturin A from a marine-derived Bacillus velezensis 11-5 as a fungicidal agent to Magnaporthe oryzae in rice. J. Plant Dis. Protect. 2019, 127, 15–24. [Google Scholar] [CrossRef]
- Tareq, F.S.; Hasan, C.M.; Lee, H.-S.; Lee, Y.-J.; Lee, J.S.; Surovy, M.Z.; Islam, M.T.; Shin, H.J. Gageopeptins A and B, new inhibitors of zoospore motility of the phytopathogen Phytophthora capsici from a marine-derived bacterium Bacillus sp. 109GGC020. Bioorg. Med. Chem. Lett. 2015, 25, 3325–3329. [Google Scholar] [CrossRef]
- Gu, K.-B.; Zhang, D.-J.; Guan, C.; Xu, J.-H.; Li, S.-L.; Shen, G.-M.; Luo, Y.-C.; Li, Y.-G. Safe antifungal lipopeptides derived from Bacillus marinus B-9987 against grey mold caused by Botrytis cinerea. J. Integr. Agric. 2017, 16, 1999–2008. [Google Scholar] [CrossRef][Green Version]
- Ma, Z.; Wang, N.; Hu, J.; Wang, S. Isolation and characterization of a new iturinic lipopeptide, mojavensin A produced by a marine-derived bacterium Bacillus mojavensis B0621A. J. Antibiot. 2012, 65, 317–322. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharya, S.; Das, A.; Samadder, S.; Rajan, S.S. Biosynthesis and characterization of a thermostable, alkali-tolerant chitinase from Bacillus pumilus JUBCH08 displaying antagonism against phytopathogenic Fusarium oxysporum. 3 Biotech 2016, 6, 87. [Google Scholar] [CrossRef] [PubMed]
- Jianyou, L.; Jianrong, X.; Yongheng, C. Isolation and identification of two marine-derived Streptomyces from marine mud of coast and offshore Zhuhai, and bioactive potential for plant pathogenic fungi. Afr. J. Biotechnol. 2011, 10, 11855–11860. [Google Scholar]
- Buatong, J.; Rukachaisirikul, V.; Sangkanu, S.; Surup, F.; Phongpaichit, S. Antifungal metabolites from marine-derived Streptomyces sp. AMA49 against Pyricularia oryzae. J. Pure Appl. Microbio. 2019, 13, 653–665. [Google Scholar] [CrossRef]
- Pesic, A.; Baumann, H.I.; Kleinschmidt, K.; Ensle, P.; Wiese, J.; Süssmuth, R.D.; Imhoff, J.F. Champacyclin, a new cyclic octapeptide from Streptomyces strain C42 isolated from the Baltic Sea. Mar. Drugs 2013, 11, 4834–4857. [Google Scholar] [CrossRef]
- Fudou, R.; Iizuka, T.; Yamanaka, S. Haliangicin, a novel antifungal metabolite produced by a marine myxobacterium. 1. Fermentation and biological characteristics. J. Antibiot. (Tokyo) 2001, 54, 149–152. [Google Scholar] [CrossRef]
- Yang, L.; Tan, R.-X.; Wang, Q.; Huang, W.-Y.; Yin, Y.-X. Antifungal cyclopeptides from Halobacillus litoralis YS3106 of marine origin. Tetrahedron Lett. 2002, 43, 6545–6548. [Google Scholar] [CrossRef]
- Manwar, A.V.; Khandelwal, S.R.; Chaudhari, B.L.; Meyer, J.M.; Chincholkar, S.B. Siderophore production by a marine Pseudomonas aeruginosa and its antagonistic action against phytopathogenic fungi. Appl. Biochem. Biotechnol. 2004, 118, 243–251. [Google Scholar] [CrossRef]
- Tarman, K.; Palm, G.J.; Porzel, A.; Merzweiler, K.; Arnold, N.; Wessjohann, L.A.; Unterseher, M.; Lindequist, U. Helicascolide C, a new lactone from an Indonesian marine algicolous strain of Daldinia eschscholzii (Xylariaceae, Ascomycota). Phytochem. Lett. 2012, 5, 83–86. [Google Scholar] [CrossRef]
- Nair, V.; Schuhmann, I.; Anke, H.; Kelter, G.; Fiebig, H.H.; Helmke, E.; Laatsch, H. Marine bacteria, XLVII—Psychrotolerant bacteria from extreme antarctic habitats as producers of rare bis- and trisindole alkaloids. Planta Med. 2016, 82, 910–918. [Google Scholar] [CrossRef]
- Manavalan, T.; Manavalan, A.; Ramachandran, S.; Heese, K. Identification of a novel thermostable alkaline protease from Bacillus megaterium-TK1 for the detergent and leather industry. Biology (Basel) 2020, 9, 472. [Google Scholar] [CrossRef]
- Mohamed, O.G.; Khalil, Z.G.; Salim, A.A.; Cui, H.; Blumenthal, A.; Capon, R.J. Lincolnenins A-D: Isomeric bactericidal bianthracenes from Streptomyces lincolnensis. J. Org. Chem. 2020. [Google Scholar] [CrossRef] [PubMed]
- Vitale, G.A.; Coppola, D.; Palma Esposito, F.; Buonocore, C.; Ausuri, J.; Tortorella, E.; de Pascale, D. Antioxidant molecules from marine fungi: Methodologies and perspectives. Antioxidants (Basel) 2020, 9, 1183. [Google Scholar] [CrossRef]
- Huang, S.; Ding, W.; Li, C.; Cox, D.G. Two new cyclopeptides from the co-culture broth of two marine mangrove fungi and their antifungal activity. Pharmacogn. Mag. 2014, 10, 410–414. [Google Scholar] [PubMed]
- Zhao, D.L.; Wang, D.; Tian, X.Y.; Cao, F.; Li, Y.Q.; Zhang, C.S. Anti-phytopathogenic and cytotoxic activities of crude extracts and secondary metabolites of marine-derived fungi. Mar. Drugs 2018, 16, 36. [Google Scholar] [CrossRef]
- Cao, F.; Yang, J.K.; Liu, Y.F.; Zhu, H.J.; Wang, C.Y. Pleosporalone A, the first azaphilone characterized with aromatic A-ring from a marine-derived Pleosporales sp. fungus. Nat. Prod. Res. 2016, 30, 2448–2452. [Google Scholar] [CrossRef]
- Du, F.Y.; Li, X.; Li, X.M.; Zhu, L.W.; Wang, B.G. Indolediketopiperazine alkaloids from eurotium cristatum en-220, an endophytic fungus isolated from the marine alga Sargassum thunbergii. Mar. Drugs 2017, 15, 24. [Google Scholar] [CrossRef]
- Zhang, P.; Mandi, A.; Li, X.M.; Du, F.Y.; Wang, J.N.; Li, X.; Kurtan, T.; Wang, B.G. Varioxepine A, a 3H-oxepine-containing alkaloid with a new oxa-cage from the marine algal-derived endophytic fungus Paecilomyces variotii. Org. Lett. 2014, 16, 4834–4837. [Google Scholar] [CrossRef]
- Xie, L.W.; Jiang, S.M.; Zhu, H.H.; Sun, W.; Ouyang, Y.C.; Dai, S.K.; Li, X. Potential inhibitors against Sclerotinia sclerotiorum, produced by the fungus Myrothecium sp. associated with the marine sponge Axinella sp. Eur. J.Plant Pathol. 2008, 122, 571–578. [Google Scholar] [CrossRef]
- Zhao, D.; Han, X.; Wang, D.; Liu, M.; Gou, J.; Peng, Y.; Liu, J.; Li, Y.; Cao, F.; Zhang, C. Bioactive 3-decalinoyltetramic acids derivatives from a marine-derived strain of the fungus Fusarium equiseti D39. Front. Microbiol. 2019, 10, 1285. [Google Scholar] [CrossRef]
- Wang, J.; Ding, W.; Li, C.; Huang, S.; She, Z.; Lin, Y. A new polysubstituted benzaldehyde from the co-culture broth of two marine fungi (Strains Nos. E33 and K38). Chem. Nat. Comp. 2013, 49, 799–802. [Google Scholar] [CrossRef]
- Bonugli-Santos, R.C.; Dos Santos Vasconcelos, M.R.; Passarini, M.R.; Vieira, G.A.; Lopes, V.C.; Mainardi, P.H.; Dos Santos, J.A.; de Azevedo Duarte, L.; Otero, I.V.; da Silva Yoshida, A.M.; et al. Marine-derived fungi: Diversity of enzymes and biotechnological applications. Front. Microbiol. 2015, 6, 269. [Google Scholar] [CrossRef] [PubMed]
- Gaspar, H.; Feio, S.S.; Rodrigues, A.I.; Van Soest, R. Antifungal activity of (+)-curcuphenol, a metabolite from the marine sponge Didiscus oxeata. Mar. Drugs 2004, 2, 8–13. [Google Scholar] [CrossRef]
- Shin, D.S.; Lee, T.H.; Lee, H.S.; Shin, J.; Oh, K.B. Inhibition of infection of the rice blast fungus by halisulfate 1, an isocitrate lyase inhibitor. FEMS Microbiol. Lett. 2007, 272, 43–47. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.; Wang, G.; Xiao, L.; Xu, X.; Liu, X.; Xu, P.; Lin, X. Bis(2,3-dibromo-4,5-dihydroxybenzyl) ether, a marine algae derived bromophenol, inhibits the growth of Botrytis cinerea and interacts with DNA molecules. Mar. Drugs 2014, 12, 3838–3851. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.S.; Lee, T.H.; Lee, J.H.; Chae, C.S.; Chung, S.C.; Shin, D.S.; Shin, J.; Oh, K.B. Inhibition of the pathogenicity of Magnaporthe grisea by bromophenols, isocitrate lyase inhibitors, from the red alga Odonthalia corymbifera. J. Agric. Food Chem. 2007, 55, 6923–6928. [Google Scholar] [CrossRef]
- de Felicio, R.; de Albuquerque, S.; Young, M.C.; Yokoya, N.S.; Debonsi, H.M. Trypanocidal, leishmanicidal and antifungal potential from marine red alga Bostrychia tenella J. Agardh (Rhodomelaceae, Ceramiales). J. Pharm. Biomed. Anal. 2010, 52, 763–769. [Google Scholar] [CrossRef]
- Guidara, M.; Yaich, H.; Amor, I.B.; Fakhfakh, J.; Gargouri, J.; Lassoued, S.; Blecker, C.; Richel, A.; Attia, H.; Garna, H. Effect of extraction procedures on the chemical structure, antitumor and anticoagulant properties of ulvan from Ulva lactuca of Tunisia coast. Carbohydr. Polym. 2021, 253, 117283. [Google Scholar] [CrossRef]
- Righini, H.; Baraldi, E.; Garcia Fernandez, Y.; Martel Quintana, A.; Roberti, R. Different antifungal activity of Anabaena sp., Ecklonia sp., and Jania sp. against Botrytis cinerea. Mar. Drugs 2019, 17, 299. [Google Scholar] [CrossRef]
- Scaglioni, P.T.; Pagnussatt, F.A.; Lemos, A.C.; Nicolli, C.P.; Del Ponte, E.M.; Badiale-Furlong, E. Nannochloropsis sp. and Spirulina sp. As a source of antifungal compounds to mitigate contamination by Fusarium graminearum species complex. Curr. Microbiol. 2019, 76, 930–938. [Google Scholar] [CrossRef]
- Scaglioni, P.T.; Scarpino, V.; Marinaccio, F.; Vanara, F.; Furlong, E.B.; Blandino, M. Impact of microalgal phenolic extracts on the control of Fusarium graminearum and deoxynivalenol contamination in wheat. World Mycotoxin J. 2019, 12, 367–378. [Google Scholar] [CrossRef]
- Lopez-Abarrategui, C.; Alba, A.; Silva, O.N.; Reyes-Acosta, O.; Vasconcelos, I.M.; Oliveira, J.T.; Migliolo, L.; Costa, M.P.; Costa, C.R.; Silva, M.R.; et al. Functional characterization of a synthetic hydrophilic antifungal peptide derived from the marine snail Cenchritis muricatus. Biochimie 2012, 94, 968–974. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.J.; Cai, J.J.; Cai, L.; Qu, H.D.; Yang, M.; Zhang, M. Cloning and expression of a hepcidin gene from a marine fish (Pseudosciaena crocea) and the antimicrobial activity of its synthetic peptide. Peptides 2009, 30, 638–646. [Google Scholar] [CrossRef] [PubMed]
- Dorr, T. Understanding tolerance to cell wall-active antibiotics. Ann. N. Y. Acad. Sci. 2020. [Google Scholar] [CrossRef] [PubMed]
- Lozoya-Perez, N.E.; Clavijo-Giraldo, D.M.; Martinez-Duncker, I.; Garcia-Carnero, L.C.; Lopez-Ramirez, L.A.; Nino-Vega, G.A.; Mora-Montes, H.M. Influences of the culturing media in the virulence and cell wall of Sporothrix schenckii, Sporothrix brasiliensis, and Sporothrix globosa. J. Fungi (Basel) 2020, 6, 323. [Google Scholar] [CrossRef]
- Lanver, D.; Mendoza-Mendoza, A.; Brachmann, A.; Kahmann, R. Sho1 and Msb2-related proteins regulate appressorium development in the smut fungus Ustilago maydis. Plant Cell 2010, 22, 2085–2101. [Google Scholar] [CrossRef]
- bin Yusof, M.T.; Kershaw, M.J.; Soanes, D.M.; Talbot, N.J. FAR1 and FAR2 regulate the expression of genes associated with lipid metabolism in the rice blast fungus Magnaporthe oryzae. PLoS ONE 2014, 9, e99760. [Google Scholar] [CrossRef]
- Deng, S.; Gu, Z.; Yang, N.; Li, L.; Yue, X.; Que, Y.; Sun, G.; Wang, Z.; Wang, J. Identification and characterization of the peroxin 1 gene MoPEX1 required for infection-related morphogenesis and pathogenicity in Magnaporthe oryzae. Sci. Rep. 2016, 6, 36292. [Google Scholar] [CrossRef]
- Oh, K.-B.; Lee, J.H.; Chung, S.-C.; Shin, J.; Shin, H.J.; Kim, H.-K.; Lee, H.-S. Antimicrobial activities of the bromophenols from the red alga Odonthalia corymbifera and some synthetic derivatives. Bioorg. Med. Chem. Lett. 2008, 18, 104–108. [Google Scholar] [CrossRef]
- Lian, N.; Wang, X.; Jing, Y.; Lin, J. Regulation of cytoskeleton-associated protein activities: Linking cellular signals to plant cytoskeletal function. J. Integr. Plant Biol. 2020. [Google Scholar] [CrossRef]
- Haque, M.; Islam, S.; Sheikh, M.A.; Dhingra, S.; Uwambaye, P.; Labricciosa, F.M.; Iskandar, K.; Charan, J.; Abukabda, A.B.; Jahan, D. Quorum sensing: A new prospect for the management of antimicrobial-resistant infectious diseases. Expert. Rev. Anti. Infect. Ther. 2020, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Soto-Aceves, M.P.; Cocotl-Yanez, M.; Servin-Gonzalez, L.; Soberon-Chavez, G. The Rhl quorum sensing system is at the top of the regulatory hierarchy under phosphate limiting conditions in Pseudomonas aeruginosa PAO1. J. Bacteriol. 2020. [Google Scholar] [CrossRef] [PubMed]
- de Almeida, C.L.; Falcao Hde, S.; Lima, G.R.; Montenegro Cde, A.; Lira, N.S.; de Athayde-Filho, P.F.; Rodrigues, L.C.; de Souza Mde, F.; Barbosa-Filho, J.M.; Batista, L.M. Bioactivities from marine algae of the genus Gracilaria. Int. J. Mol. Sci. 2011, 12, 4550–4573. [Google Scholar] [CrossRef] [PubMed]
- Ji, X.; Guo, J.; Liu, Y.; Lu, A.; Wang, Z.; Li, Y.; Yang, S.; Wang, Q. Marine-natural-product development: First discovery of nortopsentin alkaloids as novel antiviral, anti-phytopathogenic-fungus, and insecticidal agents. J. Agric. Food Chem. 2018, 66, 4062–4072. [Google Scholar] [CrossRef]
- Corey, E.J. The Logic of Chemical Synthesis; Wiley-Interscience: Hoboken, NJ, USA, 1991. [Google Scholar]
- Jessop, P.G.; Leitner, W. Chemical Synthesis Using Supercritical Fluids; John Wiley & Sons: Hoboken, NJ, USA, 2008. [Google Scholar]
- Ausländer, S.; Ausländer, D.; Fussenegger, M. Synthetic biology—The synthesis of biology. Angew. Chem. Int. Ed. 2017, 56, 6396–6419. [Google Scholar] [CrossRef]
- Sanchez-Muñoz, R.; Perez-Mata, E.; Almagro, L.; Cusido, R.M.; Bonfill, M.; Palazon, J.; Moyano, E. A novel hydroxylation step in the taxane biosynthetic pathway: A new approach to paclitaxel production by synthetic biology. Front. Bioeng. Biotech. 2020, 8, 410. [Google Scholar] [CrossRef]
- Paddon, C.J.; Westfall, P.J.; Pitera, D.J.; Benjamin, K.; Fisher, K.; McPhee, D.; Leavell, M.; Tai, A.; Main, A.; Eng, D. High-level semi-synthetic production of the potent antimalarial artemisinin. Nature 2013, 496, 528–532. [Google Scholar] [CrossRef]
- Gomez-Escribano, J.P.; Alt, S.; Bibb, M.J. Next generation sequencing of actinobacteria for the discovery of novel natural products. Mar. Drugs 2016, 14, 78. [Google Scholar] [CrossRef]
- Liu, M.; Karuso, P.; Feng, Y.; Kellenberger, E.; Liu, F.; Wang, C.; Quinn, R.J. Is it time for artificial intelligence to predict the function of natural products based on 2D-structure. MedChemComm 2019, 10, 1667–1677. [Google Scholar] [CrossRef]
| Bacteria Sources | Substances | Activity to Pathogens | Year [Ref] |
|---|---|---|---|
| B. licheniformis 09IDYM23 | Ieodoglucomide C (6) and ieodoglycolipid (7) | Aspergillus niger, Botrytis cinerea, Colletotrichum acutatum, R. solani | 2015 [105] |
| B. amyloliquefaciens SH-B74 | Plipastatin A1 (8) | B. cinerea | 2018 [78] |
| B. velezensis 11-5 | Iturin A (9) | M. oryzae | 2019 [106] |
| B.s subtilis BS155 | Fengycin BS155 (10) | M. oryzae | 2018 [77] |
| Bacillus sp. 109GGC020 | Gageopeptins A (11) and B (12) | P. capsica, B. cinera, R. solani | 2015 [107] |
| B. subtilis 109GGC020 | Gageopeptides A–D (13–16) and gageotetrin B (17) | M. oryzae Triticum (MoT) | 2020 [24] |
| B. marinus B-9987 | Lipopeptides (18–20) | B. cinerea | 2017 [108] |
| B. mojavensis B0621A | Mojavensin A (21) | Valsa mali, F. verticillioides, F. oxysporum | 2012 [109] |
| B. pumilus JUBCH08 | Chitinase | F. oxysporum | 2016 [110] |
| S. roseobiolascens XAS585, S. roseofulvus XAS588 | Fermented broth (FBE) | inhibit mycelial development and conidial germination | 2011 [111] |
| Streptomyces sp. AMA49 | Oligomycin A | Pyricularia oryzae | 2019 [112] |
| Streptomyces sp. PNM-9 | 2-methyl-N-(2′-phenylethyl)-butanamide (23) 3-methyl-N-(2′-phenylethyl)-butanamide (24) | B. glumae | 2020 [23] |
| Streptomyces Strain C42 | Champacyclin (25) | Erwinia amylovora | 2013 [113] |
| Haliangium luteum AJ-13395 | Haliangicin (26) | a wide range of fungi | 2001 [114] |
| H. litoralis YS3106 | Halolitoralin (27–29) | moderate antifungal activity | 2002 [115] |
| Pseudomonas aeruginosa | Siderophores | A. niger, A. oryzae, A. flavus, F. oxysporum, Sclerotium rolfsii | 2004 [116] |
| Daldinia eschscholzii | Helicascolide C (30) | Cladosporium cucumerinum | 2012 [117] |
| Vibrio splendidus T262 | Trisindolal (31) | Phytophthora infestans, B. cinerea | 2016 [118] |
| Fungi Sources | Substances | Activity to Pathogens | Year [Ref] |
|---|---|---|---|
| Phomopsis sp. K38 and Alternaria sp. E33 | Cyclo-(L-leucyl-trans-4-hydroxy-L-prolyl-D-leucyl-trans-4-hydroxy-L-proline) (32) | G. graminis, F. graminearum, R. cerealis, H. sativum | 2014 [96] |
| Phomopsis sp. K38 and Alternaria sp. E33 | Cyclo (D-Pro-L-Tyr-L-Pro-L-Tyr) (33) and cyclo (Gly-L-Phe-L-Pro-L-Tyr) (34) | G. graminis, F. graminearum, R. cerealis, H. sativum | 2014 [122] |
| Alternaria sp. (P8) | Benzopyranone (35) | A. brassicicola | 2018 [84] |
| Fusarium equiseti (P18) and Alternaria sp. (P8) | Stemphyperylenol (36) | A. brassicicola, Pestallozzia theae | 2018 [123] |
| Pleosporales sp. CF09-1 | Pleosporalone A (37) | B. cinerea, Phytophthora capsica, R. oryzae | 2016 [124] |
| Pleosporales sp. CF09-1 | Pleosporalones B (38) | F. oxysporum, A. brassicicola | 2019 [99] |
| Eurotium cristatum EN-220 | Rubrumazine B (39) | M. oryzae | 2017 [125] |
| Paecilomyces variotii | Varioxepine A (40) | F. graminearum | 2014 [126] |
| Aspergillus sp. D40 | Penicillic acid (41) | Ralstonia solanacearum, a few other bacteria | 2020 [22] |
| Myrothecium sp. | Roridin A (42) and roridin D (43) | Sclerotinia sclerotiorum, M. oryzae | 2008 [127] |
| Fusarium equiseti D39 | Equisetin (44) and epi-equisetin (45) | P. syringae, R. cerealis | 2019 [128] |
| Trichoderma longibrachiatum | Sesquiterpenes (46–48) | B. cinerea, C. lagrnarium | 2020 [21] |
| Nos. K38 and E33 | Ethyl 5-ethoxy-2-formyl-3-hydroxy-4-methylbenzoate (49) | F. graminearum, R. solani, P. sojae, Gloeosporium musae | 2013 [129] |
| Verruculina enalia BCC 22226 | (-)-cercosporamide (50) | M. oryzae, C. acutatum | 2020 [20] |
| Sponge Sources | Substances | Activity to Pathogens | Year [Ref] |
|---|---|---|---|
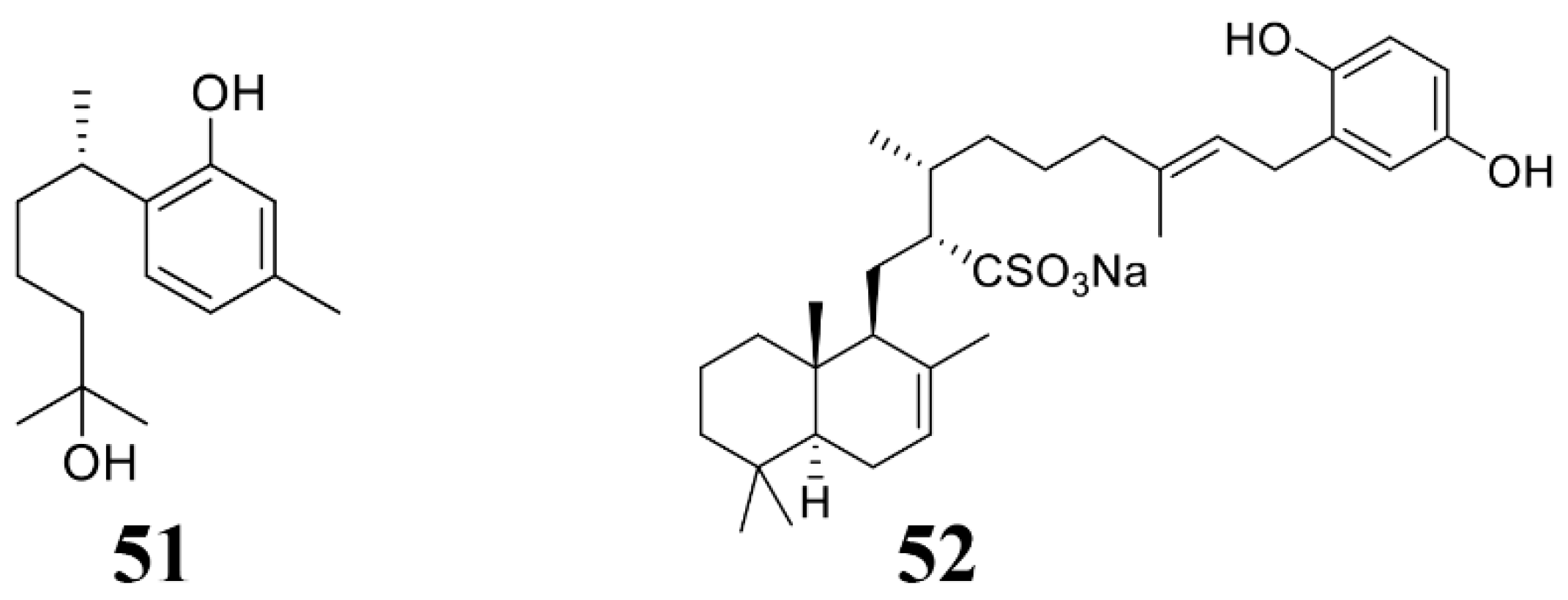
| Didiscus oxeata | (+)-curcuphenol (51) | C. cucumerinum, F. oxysporum, A. ramosa, A. niger, B. cinerea, P. expansum, R. oryzae, T. Harzianum, T. mentagrophytes, T. Koningii | 2004 [131] |
| Hippospongia spp. | Halisulfate 1 (52) | M. oryzae. | 2007 [132] |
| Seaweed Sources | Substances | Activity to Pathogens | Year [Ref] |
|---|---|---|---|
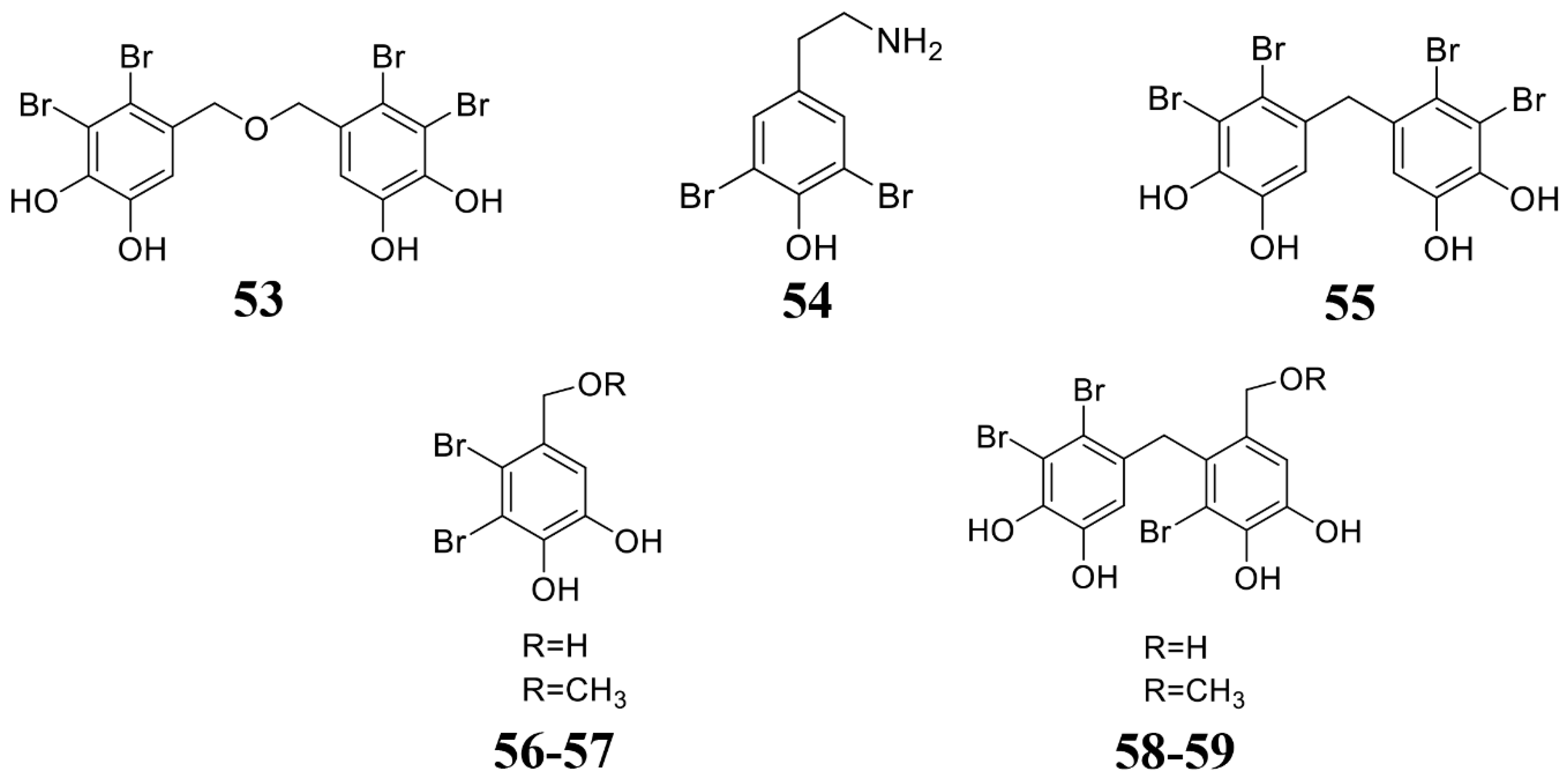
| Leathesia nana, Rhodomela confervoides, Rhodomela confervoides | Bis(2,3-dibromo-4,5-dihydroxybenzyl) ether (BDDE) (53) | B. cinerea. | 2014 [133] |
| Odonthalia corymbifera | Bromophenols (54–59) | M. oryzae. | 2007 [134] |
| Bostrychia tenella J. Agardh (Rhodomelaceae, Ceramiales) | n-hexane (BT-H) and dichloromethane (BT-D) | C. sphaerospermum, C. cladosporioides | 2010 [135] |
| Anabaena sp., Ecklonia sp., Jania sp. | Water extracts and polysaccharides | B. cinerea. | 2019 [137] |
| Nannochloropsis sp. and Spirulina sp. | Microalgal phenolic extracts (MPE) | F. graminearum | 2019 [139] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, X.; Zhao, H.; Chen, X. Screening of Marine Bioactive Antimicrobial Compounds for Plant Pathogens. Mar. Drugs 2021, 19, 69. https://doi.org/10.3390/md19020069
Li X, Zhao H, Chen X. Screening of Marine Bioactive Antimicrobial Compounds for Plant Pathogens. Marine Drugs. 2021; 19(2):69. https://doi.org/10.3390/md19020069
Chicago/Turabian StyleLi, Xiaohui, Hejing Zhao, and Xiaolin Chen. 2021. "Screening of Marine Bioactive Antimicrobial Compounds for Plant Pathogens" Marine Drugs 19, no. 2: 69. https://doi.org/10.3390/md19020069
APA StyleLi, X., Zhao, H., & Chen, X. (2021). Screening of Marine Bioactive Antimicrobial Compounds for Plant Pathogens. Marine Drugs, 19(2), 69. https://doi.org/10.3390/md19020069







