Triterpene Glycosides from the Far Eastern Sea Cucumber Psolus chitonoides: Chemical Structures and Cytotoxicities of Chitonoidosides E1, F, G, and H
Abstract
:1. Introduction
2. Results and Discussion
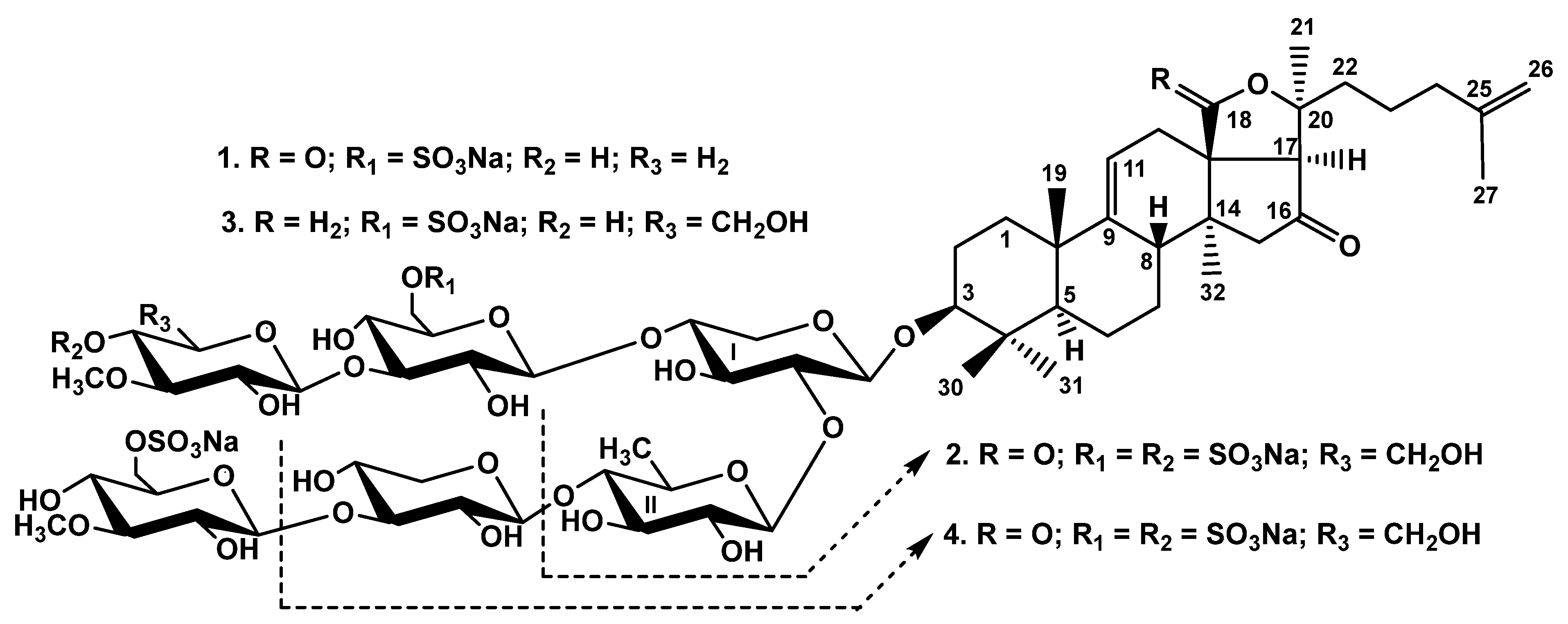
2.1. Structural Elucidation of the Glycosides
2.2. Bioactivity of the Glycosides
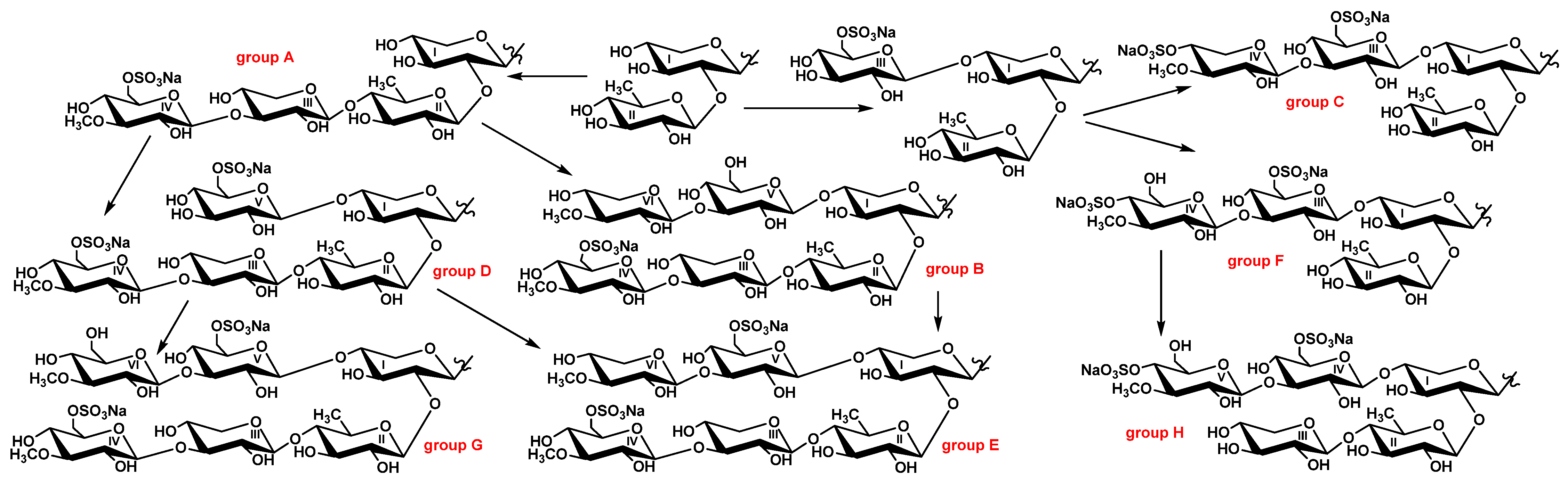
2.3. Metabolic Network of Carbohydrate Chains of Chitonoidosides of the Groups A–H
3. Materials and Methods
3.1. General Experimental Procedures
3.2. Animals and Cells
3.3. Extraction and Isolation
3.3.1. Chitonoidoside E1 (1)
3.3.2. Chitonoidoside F (2)
3.3.3. Chitonoidoside G (3)
3.3.4. Chitonoidoside H (4)
3.4. Cytotoxic Activity (MTT Assay Applied for HeLa Cells)
3.5. Cytotoxic Activity (MTS Assay Applied for DLD-1, THP-1 and HL-60 Cells)
3.6. Hemolytic Activity
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Aminin, D.L.; Menchinskaya, E.S.; Pislyagin, E.A.; Silchenko, A.S.; Avilov, S.A.; Kalinin, V.I. Sea cucumber triterpene glycosides as anticancer agents. In Studies in Natural Product Chemistry; Rahman, A.U., Ed.; Elsevier B.V.: Amsterdam, The Netherlands, 2016; Volume 49, pp. 55–105. [Google Scholar]
- Khotimchenko, Y. Pharmacological potential of sea cucumbers. Int. J. Mol. Sci. 2020, 19, 1342. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Menchinskaya, E.S.; Gorpenchenko, T.Y.; Silchenko, A.S.; Avilov, S.A.; Aminin, D.L. Modulation of doxorubicin intracellular accumulation and anticancer activity by triterpene glycoside cucumarioside A2-2. Mar. Drugs 2019, 17, 597. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, Y.-C.; Xue, C.-H.; Zhang, T.T.; Wang, Y.-M. Saponins from sea cucumber and their biological activities. Agric. Food Chem. 2018, 66, 7222–7237. [Google Scholar] [CrossRef] [PubMed]
- Gomes, A.R.; Freitas, A.C.; Duarte, A.C.; Rocha-Santos, T.A.P. Echinoderms: A review of bioactive compounds with potential health effects. In Studies in Natural Products Chemistry; Rahman, A.U., Ed.; Elsevier B.V.: Amsterdam, The Netherlands, 2016; Volume 49, pp. 1–54. [Google Scholar]
- Aminin, D.; Pisliagin, E.; Astashev, M.; Es’kov, A.; Kozhemyako, V.; Avilov, S.; Zelepuga, E.; Yurchenko, E.; Kaluzhskiy, L.; Kozlovskaya, E.; et al. Glycosides from edible sea cucumbers stimulate macrophages via purinergic receptors. Sci. Rep. 2016, 6, 39683. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dyshlovoy, S.A.; Madanchi, R.; Hauschild, J.; Otte, K.; Alsdorf, W.H.; Schumacher, U.; Kalinin, V.I.; Silchenko, A.S.; Avilov, S.A.; Honecker, F.; et al. The marine triterpene glycoside frondoside A induces p53-independent apoptosis and inhibits autophagy in urothelial carcinoma cells. BMC Cancer 2017, 17, 93. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yun, S.-H.; Sim, E.-H.; Han, S.-H.; Kim, T.-R.; Ju, M.-H.; Han, J.-Y.; Jeong, J.-S.; Kim, S.-H.; Silchenko, A.S.; Stonik, V.A.; et al. In vitro and in vivo anti-leukemic effects of cladoloside C2 are mediated by activation of Fas/ceramide syntase 6/P38 kinase/C-Jun NH2-terminal kinase/caspase-8. Oncotarget 2018, 9, 495–511. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Omran, N.E.; Salem, H.K.; Eissa, S.H.; Kabbash, A.M.; Kandeil, M.A.; Salem, M.A. Chemotaxonomic study of the most abundant Egyptian sea-cucumbers using ultra-performance liquid chromatography (UPLC) coupled to high-resolution mass spectrometry (HRMS). Chemoecology 2020, 30, 35–48. [Google Scholar] [CrossRef]
- Kalinin, V.I.; Silchenko, A.S.; Avilov, S.A. Taxonomic significance and ecological role of triterpene glycosides from holothurians. Biol. Bull. 2016, 43, 532–540. [Google Scholar] [CrossRef]
- Kalinin, V.I.; Avilov, S.A.; Silchenko, A.S.; Stonik, V.A. Triterpene Glycosides of Sea Cucumbers (Holothuroidea, Echinodermata) as Taxonomic Markers. Nat. Prod. Commun. 2015, 10, 21–26. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Silchenko, A.S.; Kalinovsky, A.I.; Avilov, S.A.; Andrijaschenko, P.V.; Popov, R.S.; Dmitrenok, P.S.; Chingizova, E.A.; Kalinin, V.I. Triterpene Glycosides from the Far Eastern Sea Cucumber Thyonidium (=Duasmodactyla) kurilensis (Levin): The Structures, Cytotoxicities, and Biogenesis of Kurilosides A3, D1, G, H, I, I1, J, K, and K1. Mar. Drugs 2021, 19, 187. [Google Scholar] [CrossRef] [PubMed]
- Silchenko, A.S.; Kalinovsky, A.I.; Avilov, S.A.; Kalinin, V.I.; Andrijaschenko, P.V.; Dmitrenok, P.S.; Popov, R.S.; Chingizova, E.A.; Ermakova, S.P.; Malyarenko, O.S. Structures and bioactivities of six new triterpene glycosides, psolusosides E, F, G, H, H1 and I and the corrected structure of psolusoside B from the sea cucumber Psolus fabricii. Mar. Drugs 2019, 17, 358. [Google Scholar] [CrossRef] [Green Version]
- Zelepuga, E.A.; Silchenko, A.S.; Avilov, S.A.; Kalinin, V.I. Structure-activity relationships of holothuroid’s triterpene glycosides and some in silico insights obtained by molecular dynamics study on the mechanisms of their membranolytic action. Mar. Drugs 2021, 19, 604. [Google Scholar] [CrossRef] [PubMed]
- Silchenko, A.S.; Kalinovsky, A.I.; Avilov, S.A.; Andrijaschenko, P.V.; Popov, R.S.; Dmitrenok, P.S.; Chingizova, E.A.; Kalinin, V.I. Unusual Structures and Cytotoxicities of Chitonoidosides A, A1, B, C, D, and E, Six Triterpene Glycosides from the Far Eastern Sea Cucumber Psolus chitonoides. Mar. Drugs 2021, 19, 449. [Google Scholar] [CrossRef] [PubMed]
- Maltsev, I.I.; Stonik, V.A.; Kalinovsky, A.I.; Elyakov, G.B. Triterpene glycosides from sea cucumber Stichopus japonicus Selenka. Comp. Biochem. Physiol. 1984, 78B, 421–426. [Google Scholar] [CrossRef]
- Silchenko, A.S.; Kalinovsky, A.I.; Avilov, S.A.; Andrijaschenko, P.V.; Popov, R.S.; Dmitrenok, P.S.; Chingizova, E.A.; Ermakova, S.P.; Malyarenko, O.S.; Dautov, S.S.; et al. Structures and bioactivities of quadrangularisosides A, A1, B, B1, B2, C, C1, D, D1–D4, and E from the sea cucumber Colochirus quadrangularis: The first discovery of the glycosides, sulfated by C-4 of the terminal 3-O-methylglucose residue. Synergetic effect on colony formation of tumor HT-29 cells of these glycosides with radioactive irradiation. Mar. Drugs 2020, 18, 394. [Google Scholar]
- Silchenko, A.S.; Kalinovsky, A.I.; Avilov, S.A.; Andrijaschenko, P.V.; Popov, R.S.; Dmitrenok, P.S.; Chingizova, E.A.; Kalinin, V.I. Kurilosides A1, A2, C1, D, E and F-triterpene glycosides from the Far Eastern sea cucumber Thyonidium (=Duasmodactyla) kurilensis (Levin): Structures with unusual non-holostane aglycones and cytotoxicities. Mar. Drugs 2020, 18, 551. [Google Scholar] [CrossRef] [PubMed]
- Aminin, D.L.; Pislyagin, E.A.; Menchinskaya, E.S.; Silchenko, A.S.; Avilov, S.A.; Kalinin, V.I. Immunomodulatory and anticancer activity of sea cucumber triterpene glycosides. In Studies in Natural Products Chemistry; Rahman, A.U., Ed.; Elsevier Science, B.V.: Amsterdam, The Netherlands, 2014; Volume 41, pp. 75–94. [Google Scholar]
- Kim, C.G.; Kwak, J.-Y. Anti-cancer effects of triterpene glycosides, frondoside A and cucumarioside A2-2 isolated from sea cucumbers. In Handbook of Anticancer Drugs from Marine Origin; Kim, S.K., Ed.; Springer International Publishing: Cham, Switzerland, 2015; pp. 673–682. [Google Scholar]
- Careaga, V.P.; Maier, M.S. Cytotoxic triterpene glycosides from sea cucumbers. In Handbook of Anticancer Drugs from Marine Origin; Kim, S.K., Ed.; Springer International Publishing: Cham, Switzerland, 2015; pp. 515–528. [Google Scholar]
- Kalinin, V.I.; Silchenko, A.S.; Avilov, S.A.; Stonik, V.A. Non-holostane aglycones of sea cucumber triterpene glycosides. Structure, biosynthesis, evolution. Steroids 2019, 147, 42–51. [Google Scholar] [CrossRef] [PubMed]
- Kalinin, V.I.; Silchenko, A.S.; Avilov, S.A.; Stonik, V.A. Progress in the studies of triterpene glycosides from sea cucumbers (Holothuroidea, Echinodermata) Between 2017 and 2021. Nat. Prod. Commun. 2021, 16. [Google Scholar] [CrossRef]
| Atom | δC Mult. a–c | δH Mult. (J in Hz) d | HMBC | ROESY |
|---|---|---|---|---|
| Xyl1 (1→C-3) | ||||
| 1 | 104.8 CH | 4.67 d (7.1) | C: 3 | H-3; H-3, 5 Xyl1 |
| 2 | 82.2 CH | 3.96 m | C: 1 Qui2 | H-1 Qui2 |
| 3 | 75.1 CH | 4.16 m | H-1, 5 Xyl1 | |
| 4 | 77.9 CH | 4.16 m | H-1 Glc5 | |
| 5 | 63.5 CH2 | 4.38 dd (4.3; 12.2) | C: 3 Xyl1 | |
| 3.64 m | H-1 Xyl1 | |||
| Qui2 (1→2Xyl1) | ||||
| 1 | 104.6 CH | 5.03 d (8.0) | C: 2 Xyl1 | H-2 Xyl1; H-3, 5 Qui2 |
| 2 | 75.7 CH | 3.87 t (8.0) | C: 1, 3 Qui2 | |
| 3 | 74.8 CH | 3.98 t (9.3) | C: 2 Qui2 | H-1 Qui2 |
| 4 | 85.6 CH | 3.49 t (9.3) | C: 1 Xyl3; 3, 5 Qui2 | H-1 Xyl3 |
| 5 | 71.4 CH | 3.67 dd (6.2; 9.3) | H-1 Qui2 | |
| 6 | 17.8 CH3 | 1.61 d (6.2) | C: 4, 5 Qui2 | H-4 Qui2 |
| Xyl3 (1→4Qui2) | ||||
| 1 | 104.5 CH | 4.75 d (7.7) | C: 4 Qui2 | H-4 Qui2; H-3, 5 Xyl3 |
| 2 | 73.2 CH | 3.84 t (8.5) | C: 1 Xyl3 | |
| 3 | 87.1 CH | 4.04 t (8.5) | C: 1 MeGlc4; 2, 4 Xyl3 | H-1 MeGlc4 |
| 4 | 68.7 CH | 3.90 m | ||
| 5 | 65.7 CH2 | 4.13 dd (4.3; 11.1) | C: 4 Xyl3 | |
| 3.60 t (11.1) | C: 1, 3 Xyl3 | H-1 Xyl3 | ||
| MeGlc4 (1→3Xyl3) | ||||
| 1 | 104.4 CH | 5.11 d (7.9) | C: 3 Xyl3 | H-3 Xyl3; H-3, 5 MeGlc4 |
| 2 | 74.3 CH | 3.80 t (7.9) | C: 1 MeGlc4 | |
| 3 | 86.4 CH | 3.64 t (9.1) | C: 2, 4 MeGlc4, OMe | H-1, 5 MeGlc4 |
| 4 | 69.7 CH | 3.96 t (9.1) | ||
| 5 | 75.3 CH | 4.03 m | H-1, 3 MeGlc4 | |
| 6 | 67.1 CH2 | 4.97 dd (3.0; 11.5) | C: 5 MeGlc4 | |
| 4.71 dd (6.1; 11.5) | C: 5 MeGlc4 | |||
| OMe | 60.5 CH3 | 3.76 s | C: 3 MeGlc4 | |
| Glc5 (1→4Xyl1) | ||||
| 1 | 102.3 CH | 4.90 d (7.9) | C: 4 Xyl1 | H-4 Xyl1; H-3, 5 Glc5 |
| 2 | 73.2 CH | 3.84 t (9.1) | C: 1, 3 Glc5 | |
| 3 | 86.0 CH | 4.07 t (9.1) | C: 1 MeXyl6; 2 Glc5 | H-1 MeXyl6; H-1 Glc5 |
| 4 | 68.9 CH | 3.88 t (9.1) | ||
| 5 | 75.5 CH | 4.05 m | H-1 Glc5 | |
| 6 | 67.1 CH2 | 4.97 dd (1.9; 11.5) | C: 4, 5 Glc5 | |
| 4.70 dd (5.5; 11.5) | ||||
| MeXyl6 (1→3Glc5) | ||||
| 1 | 105.2 CH | 5.07 d (7.9) | C: 3 Glc5 | H-3 Glc5; H-3,5 MeXyl6 |
| 2 | 74.2 CH | 3.79 t (8.5) | C: 1 MeXyl6 | |
| 3 | 86.6 CH | 3.57 t (9.1) | C: 2, 4 MeXyl6; OMe | H-1 MeXyl6; OMe |
| 4 | 69.9 CH | 3.98 t (9.1) | C: 3, 5 MeXyl6 | |
| 5 | 66.4 CH2 | 4.14 dd (5.5; 12.1) | C: 1, 3, 4 MeXyl6 | |
| 3.61 t (11.5) | H-1 MeXyl6 | |||
| OMe | 60.5 CH3 | 3.79 s | C: 3 MeXyl6 | H-3 MeXyl6 |
| Atom | δC Mult. a–c | δH Mult. (J in Hz) d | HMBC | ROESY |
|---|---|---|---|---|
| Xyl1 (1→C-3) | ||||
| 1 | 104.8 CH | 4.67 d (7.6) | C: 3 | H-3; H-3, 5 Xyl1 |
| 2 | 82.1 CH | 3.98 t (8.3) | C: 1 Qui2; 3 Xyl1 | H-1 Qui2 |
| 3 | 75.1 CH | 4.17 t (8.3) | C: 4 Xyl1 | |
| 4 | 78.0 CH | 4.16 m | H-1 Xyl1; H-1 Glc3 | |
| 5 | 63.5 CH2 | 4.38 d (10.2) | ||
| 3.63 m | H-1, 3 Xyl1 | |||
| Qui2 (1→2Xyl1) | ||||
| 1 | 104.9 CH | 5.06 d (8.1) | C: 2 Xyl1 | H-2 Xyl1; H-3, 5 Qui2 |
| 2 | 76.2 CH | 3.88 t (9.1) | C: 1 Qui2 | H-4 Qui2 |
| 3 | 76.8 CH | 4.06 t (9.1) | C: 2, 4 Qui2 | H-1, 5 Qui2 |
| 4 | 76.2 CH | 3.58 t (9.1) | C: 3, 5 Qui2 | H-2 Qui2 |
| 5 | 72.8 CH | 3.70 dd (6.3; 9.9) | C: 4 Qui2 | H-1, 3 Qui2 |
| 6 | 18.2 CH3 | 1.53 d (6.3) | C: 4, 5 Qui2 | |
| Glc3 (1→4Xyl1) | ||||
| 1 | 102.2 CH | 4.90 d (8.0) | C: 4 Xyl1 | H-4 Xyl1; H-3, 5 Glc3 |
| 2 | 73.3 CH | 3.83 t (9.2) | C: 1 Glc3 | |
| 3 | 86.0 CH | 4.17 t (9.2) | C: 2, 4 Glc3; 1 MeGlc4 | H-1 MeGlc4; H-1 Glc3 |
| 4 | 68.9 CH | 3.87 t (9.2) | C: 5 Glc3 | |
| 5 | 75.1 CH | 4.04 m | H-1 Glc3 | |
| 6 | 67.2 CH2 | 4.94 d (10.3) | ||
| 4.68 m | ||||
| MeGlc4 (1→3Glc3) | ||||
| 1 | 104.3 CH | 5.18 d (8.0) | C: 3 Glc3 | H-3 Glc3; H-3, 5 MeGlc4 |
| 2 | 74.0 CH | 3.86 t (8.7) | C: 1, 3 MeGlc4 | H-4 MeGlc4 |
| 3 | 85.3 CH | 3.71 t (8.7) | C: 4, 5 MeGlc4, OMe | H-1, 5 MeGlc4; Ome |
| 4 | 76.3 CH | 4.88 t (9.6) | C: 3, 5 MeGlc4 | H-2 MeGlc4 |
| 5 | 76.4 CH | 3.86 t (8.7) | H-1 MeGlc4 | |
| 6 | 61.2 CH2 | 4.51 d (11.4) | ||
| 4.33 dd (4.4; 11.4) | ||||
| OMe | 60.7 CH3 | 3.93 s | C: 3 MeGlc4 |
| Position | δC Mult. a | δH Mult. (J in Hz) b | HMBC | ROESY |
|---|---|---|---|---|
| 1 | 35.9 CH2 | 1.60 m | H-11 | |
| 1.28 m | ||||
| 2 | 26.7 CH2 | 2.08 m | ||
| 1.83 m | H-19, H-30 | |||
| 3 | 88.7 CH | 3.12 dd (4.2; 11.8) | C: 1 Xyl1 | H-5, H-31, H1-Xyl1 |
| 4 | 39.6 C | |||
| 5 | 52.7 CH | 0.75 brd (12.0) | C: 10, 19 | H-3, H-31 |
| 6 | 20.9 CH2 | 1.57 m | H-31 | |
| 1.35 m | H-19, H-30 | |||
| 7 | 28.7 CH2 | 1.57 m | ||
| 1.17 m | ||||
| 8 | 40.9 CH | 2.31 m | H-18, H-19 | |
| 9 | 150.9 C | |||
| 10 | 39.5 C | |||
| 11 | 114.7 CH | 5.30 m | H-1 | |
| 12 | 33.8 CH2 | 2.38 m | H-32 | |
| 2.25 m | H-21 | |||
| 13 | 56.3 C | |||
| 14 | 40.3 C | |||
| 15 | 50.5 CH2 | 2.47 d (15.9) | C: 14, 16, 32 | H-18 |
| 2.19 d (15.9) | C: 13, 16 | H-32 | ||
| 16 | 218.1 C | |||
| 17 | 63.8 CH | 2.35 s | C: 12, 13, 16, 18, 20, 21 | H-12, H-21, H-22, H-32 |
| 18 | 73.8 CH2 | 4.02 m | ||
| 3.65 d (9.1) | C: 12, 20 | |||
| 19 | 22.2 CH3 | 0.97 s | C: 1, 5, 9, 10 | H-1, H-2, H-6, H-8, H-18 |
| 20 | 86.6 C | |||
| 21 | 26.1 CH3 | 1.32 s | C: 17, 20, 22 | H-12, H-17, H-18, H-22 |
| 22 | 37.8 CH2 | 1.70 m | ||
| 1.57 m | H-21, H-24 | |||
| 23 | 22.7 CH2 | 1.69 m | ||
| 1.56 m | ||||
| 24 | 38.2 CH2 | 1.95 m | C: 23 | H-21 |
| 25 | 146.0 C | |||
| 26 | 110.2 CH2 | 4.72 brs | C: 24, 27 | |
| 4.71 brs | C: 24, 27 | |||
| 27 | 22.2 CH3 | 1.65 s | C: 24, 25, 26 | H-24 |
| 30 | 16.5 CH3 | 0.97 s | C: 3, 4, 5, 31 | H-31 |
| 31 | 27.9 CH3 | 1.13 s | C: 3, 4, 5, 30 | H-1, H-3, H-5, H-30 |
| 32 | 21.4 CH3 | 0.78 s | C: 13, 14, 15 | H-15, H-17 |
| Atom | δC Mult. a–c | δH Mult. (J in Hz) d | HMBC | ROESY |
|---|---|---|---|---|
| Xyl1 (1→C-3) | ||||
| 1 | 104.8 CH | 4.66 d (6.9) | C: 3 | H-3; H-3, 5 Xyl1 |
| 2 | 82.1 CH | 3.97 t (8.8) | C: 1 Qui2; 1, 3 Xyl1 | H-1 Qui2 |
| 3 | 75.1 CH | 4.16 t (8.8) | C: 4 Xyl1 | |
| 4 | 77.8 CH | 4.16 m | ||
| 5 | 63.5 CH2 | 4.37 dd (4.1; 11.8) | ||
| 3.62 m | H-1 Xyl1 | |||
| Qui2 (1→2Xyl1) | ||||
| 1 | 104.5 CH | 5.04 d (7.3) | C: 2 Xyl1 | H-2 Xyl1; H-3, 5 Qui2 |
| 2 | 75.7 CH | 3.87 t (9.0) | C: 1, 3 Qui2 | H-4 Qui2 |
| 3 | 74.8 CH | 3.98 t (9.0) | C: 2, 4 Qui2 | H-5 Qui2 |
| 4 | 85.6 CH | 3.49 t (9.0) | C: 1 Xyl3; 3, 5 Qui2 | H-1 Xyl3; H-2 Qui2 |
| 5 | 71.4 CH | 3.68 dd (6.2; 9.0) | H-1 Qui2 | |
| 6 | 17.8 CH3 | 1.62 d (6.2) | C: 4, 5 Qui2 | H-4, 5 Qui2 |
| Xyl3 (1→4Qui2) | ||||
| 1 | 104.4 CH | 4.75 d (7.7) | C: 4 Qui2 | H-4 Qui2; H-3, 5 Xyl3 |
| 2 | 73.2 CH | 3.84 t (8.3) | C: 1, 3 Xyl3 | |
| 3 | 87.0 CH | 4.04 t (8.3) | C: 1 MeGlc4; 2, 4 Xyl3 | H-1 MeGlc4; H-1 Xyl3 |
| 4 | 68.8 CH | 3.89 m | C: 5 Xyl3 | |
| 5 | 65.7 CH2 | 4.12 dd (5.3; 11.2) | ||
| 3.59 d (11.2) | C: 1 Xyl3 | H-1 Xyl3 | ||
| MeGlc4 (1→3Xyl3) | ||||
| 1 | 104.6 CH | 5.12 d (7.9) | C: 3 Xyl3 | H-3 Xyl3; H-3, 5 MeGlc4 |
| 2 | 74.3 CH | 3.80 t (8.5) | C: 1 MeGlc4 | |
| 3 | 86.4 CH | 3.64 t (8.5) | C: 4 MeGlc4, OMe | H-1 MeGlc4; OMe |
| 4 | 69.9 CH | 3.96 t (8.5) | C: 3, 5, 6 MeGlc4 | H-2, 6 MeGlc4 |
| 5 | 75.5 CH | 4.03 m | H-1, 3 MeGlc4 | |
| 6 | 67.1 CH2 | 4.97 d (10.7) | ||
| 4.71 dd (5.6; 11.3) | C: 5 MeGlc4 | |||
| OMe | 60.5 CH3 | 3.76 s | C: 3 MeGlc4 | |
| Glc5 (1→4Xyl1) | ||||
| 1 | 102.2 CH | 4.88 d (7.9) | C: 4 Xyl1 | H-4 Xyl1; H-3, 5 Glc5 |
| 2 | 73.2 CH | 3.84 t (9.0) | C: 1, 3 Glc5 | |
| 3 | 86.0 CH | 4.16 t (9.0) | C: 1 MeGlc6; 2 Glc5 | H-1 MeGlc6; H-1 Glc5 |
| 4 | 69.0 CH | 3.89 t (9.0) | C: 3 Glc5 | |
| 5 | 75.5 CH | 4.02 m | H-1 Glc5 | |
| 6 | 67.1 CH2 | 4.93 d (10.7) | ||
| 4.68 dd (6.2; 11.3) | ||||
| MeGlc6 (1→3Glc5) | ||||
| 1 | 104.4 CH | 5.18 d (7.5) | C: 3 Glc5 | H-3 Glc5; H-3, 5 MeGlc6 |
| 2 | 74.5 CH | 3.84 t (8.8) | C: 1 MeGlc6 | |
| 3 | 86.8 CH | 3.66 t (8.8) | C: 2, 4 MeGlc6, OMe | H-1 MeGlc6 |
| 4 | 70.3 CH | 3.89 m | C: 5 MeGlc6 | H-6 MeGlc6 |
| 5 | 77.5 CH | 3.89 m | H-1 MeGlc6 | |
| 6 | 61.7 CH2 | 4.34 dd (2.2; 11.7) | ||
| 4.05 dd (5.1; 11.7) | C: 4 MeGlc6 | |||
| OMe | 60.6 CH3 | 3.80 s | C: 3 MeGlc6 |
| Atom | δC Mult. a–c | δH Mult. (J in Hz) d | HMBC | ROESY |
|---|---|---|---|---|
| Xyl1 (1→C-3) | ||||
| 1 | 104.6 CH | 4.66 d (6.9) | C: 3 | H-3; H-3, 5 Xyl1 |
| 2 | 81.9 CH | 3.97 m | C: 1 Qui2; 1 Xyl1 | H-1 Qui2 |
| 3 | 75.1 CH | 4.16 m | C: 4 Xyl1 | H-1 Xyl1 |
| 4 | 78.0 CH | 4.15 m | H-1 Glc4 | |
| 5 | 63.5 CH2 | 4.38 m | C: 3 Xyl1 | |
| 3.62 m | H-1 Xyl1 | |||
| Qui2 (1→2Xyl1) | ||||
| 1 | 104.4 CH | 5.07 d (6.9) | C: 2 Xyl1 | H-2 Xyl1; H-5 Qui2 |
| 2 | 75.7 CH | 3.87 t (9.2) | C: 1, 3 Qui2 | H-4 Qui2 |
| 3 | 74.8 CH | 4.00 t (9.2) | C: 2, 4 Qui2 | H-1 Qui2 |
| 4 | 85.6 CH | 3.47 t (9.2) | C: 1 Xyl3; 3, 5 Qui2 | H-1 Xyl3; H-2 Qui2 |
| 5 | 71.4 CH | 3.70 dd (6.2; 9.2) | H-1, 3 Qui2 | |
| 6 | 17.8 CH3 | 1.61 d (6.2) | C: 4, 5 Qui2 | H-4, 5 Qui2 |
| Xyl3 (1→4Qui2) | ||||
| 1 | 104.8 CH | 4.70 d (6.6) | C: 4 Qui2 | H-4 Qui2; H-3, 5 Xyl3 |
| 2 | 73.3 CH | 3.82 t (8.4) | C: 1, 3 Xyl3 | |
| 3 | 77.2 CH | 4.05 t (8.4) | C: 2, 4 Xyl3 | |
| 4 | 70.1 CH | 4.03 m | C: 3 Xyl3 | |
| 5 | 66.5 CH2 | 4.14 brd (11.2) | C: 3 Xyl3 | |
| 3.59 t (11.2) | C: 1, 3, 4 Xyl3 | H-1 Xyl3 | ||
| Glc4 (1→4Xyl1) | ||||
| 1 | 102.3 CH | 4.89 d (8.1) | C: 4 Xyl1 | H-4 Xyl1; H-3, 5 Glc4 |
| 2 | 74.0 CH | 3.82 t (9.1) | C: 1 Glc4 | |
| 3 | 86.0 CH | 4.16 t (9.1) | C: 1 MeGlc5 | H-1 MeGlc5; H-1 Glc4 |
| 4 | 69.0 CH | 3.86 t (9.1) | C: 3 Glc4 | |
| 5 | 75.2 CH | 4.03 m | H-1 Glc4 | |
| 6 | 67.2 CH2 | 4.96 d (12.2) | ||
| 4.69 m | ||||
| MeGlc5 (1→3Glc4) | ||||
| 1 | 104.3 CH | 5.18 d (8.1) | C: 3 Glc4 | H-3 Glc4; H-3, 5 MeGlc5 |
| 2 | 74.1 CH | 3.86 t (9.1) | C: 1 MeGlc5 | H-4 MeGlc5 |
| 3 | 85.2 CH | 3.71 t (9.1) | C: 4 MeGlc5; OMe | H-1 MeGlc5 |
| 4 | 76.2 CH | 4.88 t (9.1) | C: 3, 5 MeGlc5 | H-6 MeGlc5 |
| 5 | 76.4 CH | 3.84 t (9.1) | ||
| 6 | 61.7 CH2 | 4.50 d (12.2) | ||
| 4.33 dd (6.2; 12.2) | ||||
| OMe | 60.7 CH3 | 3.93 s | C: 3 MeGlc5 |
| Glycosides | ED50, µM, Erythrocytes | Cytotoxicity, ED50 µM | |||
|---|---|---|---|---|---|
| HeLa | DLD-1 | THP-1 | HL-60 | ||
| Chitonoidoside E | 0.45 ± 0.01 | 5.73 ± 0.10 | 8.93 ± 1.28 | 0.58 ± 0.07 | 5.73 ± 0.37 |
| Chitonoidoside E1 (1) | 0.64 ± 0.01 | 18.00 ± 0.59 | 34.12 ± 2.12 | 0.58 ± 0.04 | 9.97 ± 0.94 |
| Chitonoidoside F (2) | 1.17 ± 0.07 | 37.99 ± 1.36 | 71.11 ± 1.22 | 4.87 ± 0.47 | 41.23 ± 1.11 |
| Chitonoidoside G (3) | 0.65 ± 0.01 | 14.52 ± 1.08 | 12.44 ± 1.07 | 4.81 ± 0.46 | 8.23 ± 0.33 |
| Chitonoidoside H (4) | 0.89 ± 0.05 | 17.02 ± 1.18 | 36.62 ± 1.51 | 2.88 ± 0.23 | 8.13 ± 0.45 |
| Chitonoidoside A | 1.27 ± 0.03 | 39.48 ± 1.15 | 32.68 ± 2.56 | 2.93 ± 0.17 | 8.95 ± 0.35 |
| Cisplatin | - | 16.94 ± 0.25 | > 80.00 | 56.12 ± 3.91 | 8.58 ± 0.54 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Silchenko, A.S.; Kalinovsky, A.I.; Avilov, S.A.; Andrijaschenko, P.V.; Popov, R.S.; Chingizova, E.A.; Kalinin, V.I.; Dmitrenok, P.S. Triterpene Glycosides from the Far Eastern Sea Cucumber Psolus chitonoides: Chemical Structures and Cytotoxicities of Chitonoidosides E1, F, G, and H. Mar. Drugs 2021, 19, 696. https://doi.org/10.3390/md19120696
Silchenko AS, Kalinovsky AI, Avilov SA, Andrijaschenko PV, Popov RS, Chingizova EA, Kalinin VI, Dmitrenok PS. Triterpene Glycosides from the Far Eastern Sea Cucumber Psolus chitonoides: Chemical Structures and Cytotoxicities of Chitonoidosides E1, F, G, and H. Marine Drugs. 2021; 19(12):696. https://doi.org/10.3390/md19120696
Chicago/Turabian StyleSilchenko, Alexandra S., Anatoly I. Kalinovsky, Sergey A. Avilov, Pelageya V. Andrijaschenko, Roman S. Popov, Ekaterina A. Chingizova, Vladimir I. Kalinin, and Pavel S. Dmitrenok. 2021. "Triterpene Glycosides from the Far Eastern Sea Cucumber Psolus chitonoides: Chemical Structures and Cytotoxicities of Chitonoidosides E1, F, G, and H" Marine Drugs 19, no. 12: 696. https://doi.org/10.3390/md19120696
APA StyleSilchenko, A. S., Kalinovsky, A. I., Avilov, S. A., Andrijaschenko, P. V., Popov, R. S., Chingizova, E. A., Kalinin, V. I., & Dmitrenok, P. S. (2021). Triterpene Glycosides from the Far Eastern Sea Cucumber Psolus chitonoides: Chemical Structures and Cytotoxicities of Chitonoidosides E1, F, G, and H. Marine Drugs, 19(12), 696. https://doi.org/10.3390/md19120696









