A Review of Toxins from Cnidaria
Abstract
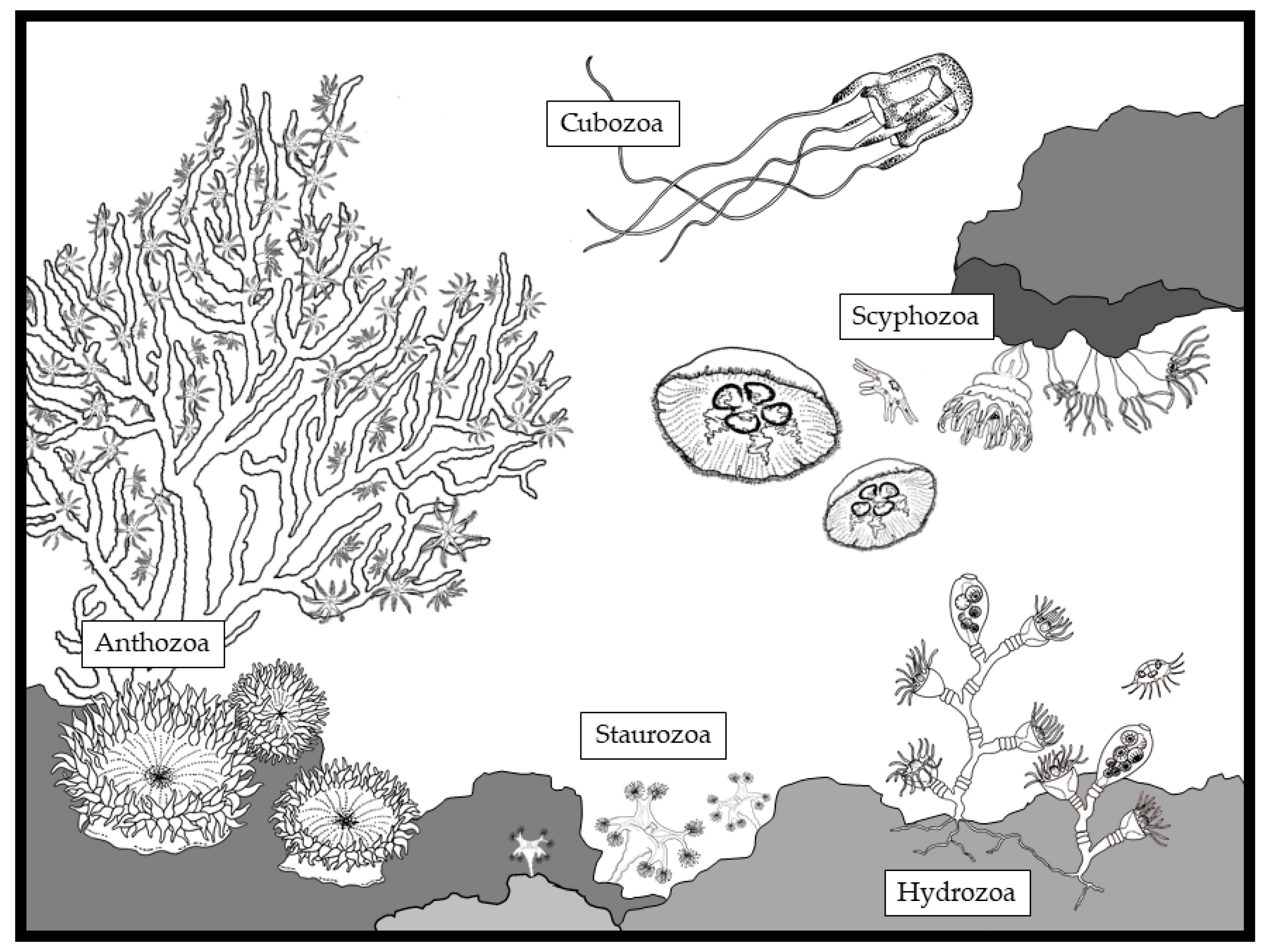
1. Introduction to the Phylum Cnidaria
2. Venoms of Cnidaria
2.1. Phospholipase A2
2.2. Metalloproteinases
2.3. Voltage-Gated Sodium and Potassium Channel Toxins
2.4. Kunitz-Type Proteinase Inhibitors
2.5. TRPV1 Channel Inhibitors
2.6. TRPA1 Channel Modulators
2.7. ASICs Channels Modulators
2.8. Beta-Defensin Like Alpha-Amylase Inhibitors
3. Omics Applied to Cnidaria to Identify Chemical Defences
4. Cnidaria Bioprospecting
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Arai, M.N. A Functional Biology of Scyphozoa; Chapman & Hall: London, UK, 1997; p. 315. [Google Scholar]
- Technau, U.; Steele, R.E. Evolutionary crossroads in developmental biology. Cnidaria. Dev. 2012, 138, 1447. [Google Scholar] [CrossRef] [PubMed]
- Mariscal, R.N. Nematocysts. In Coelenterate Biology; Lenhoff, K., Muscatine, L., Marian, E., Davis, L., Eds.; University of Hawaii Press: Honolulu, Hawaii, 1974; pp. 129–178. [Google Scholar]
- Östman, C. A guideline to nematocyst nomenclature and classification, and some notes on the systematic value of nematocysts. Sci. Mar. 2000, 64, 31–46. [Google Scholar] [CrossRef]
- Mariscal, R.N.; Conklin, E.J.; Bigger, C.H. The ptychocyst, a major new category of cnida used in tube construction by a cerianthid anemone. Biol. Bull. 1977, 152, 392–405. [Google Scholar] [CrossRef]
- Larson, R.J.; Fautin, D.G. Stauromedusae of the genus Manania (=Thaumatoscyphus) (Cnidaria, Scyphozoa) in the Northeast Pacific, including descriptions of new species Manania gwilliami and Manania handi. Can. J. Zool. 1989, 67, 1543–1549. [Google Scholar] [CrossRef]
- Ames, C.L.; Klompen, A.M.L.; Badhiwala, K.; Muffett, K.; Reft, A.J.; Kumar, M.; Janssen, J.D.; Schultzhaus, J.N.; Field, L.D.; Muroski, M.E.; et al. Cassiosomes are stinging-cell structures in the mucus of the upside-down jellyfish Cassiopea xamachana. Commun. Biol. 2020, 3, 67. [Google Scholar] [CrossRef]
- Fautin, D. Structural diversity, systematics, and evolution of cnidae. Toxicon 2009, 54, 1054–1064. [Google Scholar] [CrossRef] [PubMed]
- Moran, Y.; Genikhovich, G.; Gordon, D.; Wienkoop, S.; Zenkert, C.; Ôzbek, S.; Technau, U.; Gurevitz, M. Neurotoxin localization to ectodermal gland cells uncovers an alternative mechanism of venom delivery in sea anemones. Proc. Rotal Soc. Lond. (Part B) 2012, 279, 1351–1358. [Google Scholar] [CrossRef]
- Jouiaei, M.; Yanagihara, A.; Madio, B.; Nevalainen, T.J.; Alewood, P.F.; Fry, B. Ancient venom systems: A review on Cnidaria toxins. Toxins 2015, 7, 2251–2271. [Google Scholar] [CrossRef]
- Frazão, B.; Vasconcelos, V.; Antunes, A. Sea anemone (Cnidaria, Anthozoa, Actiniaria) toxins: An overview. Mar. Drugs 2012, 10, 1812–1851. [Google Scholar] [CrossRef]
- Mariottini, G.L. Hemolytic venoms from marine cnidarian jellyfish—An overview. J. Venom Res. 2014, 5, 22. [Google Scholar]
- Mariottini, G.L.; Pane, L. Mediterranean jellyfish venoms: A review on scyphomedusae. Mar. Drugs 2010, 8, 1122–1152. [Google Scholar] [CrossRef] [PubMed]
- Mariottini, G.L.; Pane, L. Cytotoxic and cytolytic cnidarian venoms. A review on health implications and possible therapeutic applications. Toxins 2014, 6, 108–151. [Google Scholar] [CrossRef] [PubMed]
- Nevalainen, T.J.; Peuravuori, H.J.; Quinn, R.J.; Llewellyn, L.E.; Benzie, J.A.H.; Fenner, P.J.; Winkel, K.D. Phospholipase A2 in Cnidaria. Comp. Biochem. Physiol. Part B Biochem. Mol. Biol. 2004, 139, 731–735. [Google Scholar] [CrossRef] [PubMed]
- Moran, Y.; Gordon, D.; Gurevitz, M. Sea anemone toxins affecting voltage-gated sodium channels—Molecular and evolutionary features. Toxicon 2009, 54, 1089–1101. [Google Scholar] [CrossRef]
- Merquiol, L.; Romano, G.; Ianora, A.; D’Ambra, I. Biotechnological applications of scyphomedusae. Mar. Drugs 2019, 17, 604. [Google Scholar] [CrossRef]
- Brinkman, D.L.; Konstantakopoulos, N.; McInerney, B.V.; Mulvenna, J.; Seymour, J.E.; Isbister, G.K.; Hodgson, W.C. Chironex fleckeri (box jellyfish) venom proteins: Expansion of a cnidarian toxin family that elicits variable cytolytic and cardiovascular effects. J. Biol. Chem. 2014, 289, 4798–4812. [Google Scholar] [CrossRef]
- Glasser, E.; Rachamim, T.; Aharonovich, D.; Sher, D. Hydra actinoporin-like toxin-1, an unusual hemolysin from the nematocyst venom of Hydra magnipapillata which belongs to an extended gene family. Toxicon 2014, 91, 103–113. [Google Scholar] [CrossRef]
- Yap, W.Y.; Tan, K.J.S.X.; Hwang, J.S. Expansion of Hydra actinoporin-like toxin (HALT) gene family: Expression divergence and functional convergence evolved through gene duplication. Toxicon 2019, 170, 10–20. [Google Scholar] [CrossRef]
- Zhang, M.; Fishman, Y.; Sher, D.; Zlotkin, E. Hydralysin, a novel animal group-selective paralytic and cytolytic protein from a noncnidocystic origin in Hydra. Biochemistry 2003, 42, 8939–8944. [Google Scholar] [CrossRef]
- Radwan, F.F.Y.; Aboul-Dahab, H.M. Milleporin-1, a new phospholipase A2 active protein from the fire coral Millepora platyphylla nematocysts. Comp. Biochem. Physiol. C Toxicol. Pharmacol. 2004, 139, 267–272. [Google Scholar] [CrossRef]
- Haddad, V., Jr.; Zara, F.; Marangoni, S.; Toyama, D.; Souza, A.; Oliveira, S.; Toyama, M. Identification of two novel cytolysins from the hydrozoan Olindias sambaquiensis (Cnidaria). J. Venom. Anim. Toxins Incl. Trop. Dis. 2014, 20, 10. [Google Scholar]
- Weston, A.J.; Chung, R.; Dunlap, W.C.; Morandini, A.C.; Marques, A.C.; Moura-da-Silva, A.M.; Ward, M.; Padilla, G.; da Silva, L.F.; Andreakis, N.; et al. Proteomic characterisation of toxins isolated from nematocysts of the South Atlantic jellyfish Olindias sambaquiensis. Toxicon 2013, 71, 11–17. [Google Scholar] [CrossRef] [PubMed]
- Stillway, L.W.; Lane, C.E. Phospholipase in the nematocyst toxin of Physalia physalis. Toxicon 1971, 9, 193–195. [Google Scholar] [CrossRef]
- Burnett, J.W.; Calton, G.G. Venomous pelagic coelenterates chemistry toxicology immunology and treatment of their stings. Toxicon 1987, 25, 581–602. [Google Scholar] [CrossRef]
- Werb, Z.; Banda, M.J.; McKerrow, J.H.; Sandhaus, R.A. Elastases and elastin degradation. J. Investig. Dermatol. 1982, 79, 154–159. [Google Scholar] [CrossRef]
- Dìaz Garcìa, C.; Fuentes-Silva, D.; Sanchez-Soto, C.; Domìnguez Pàrez, D.; Garcìa-Delgado, N.; Varela, C.; Mendoza-Hernàndez, G.; Rodrìguez-Romero, A.; Castañeda, O.; Hiriart, M. Toxins from Physalia physalis (Cnidaria) raise the intracellular Ca2+ of beta-cells and promotei Insulin secretion. Curr. Med. Chem. 2012, 19, 5414–5423. [Google Scholar] [CrossRef] [PubMed]
- Mas, R.; Menéndez, R.; Garateix, A.; Garcia, M.; Cha, M. Effects of a high molecular weight toxin from Physalia physalis on glutamate responses. Neuroscience 1989, 33, 269–273. [Google Scholar] [CrossRef]
- Menéndez, R.; Mas, R.; Garateix, A.; Garcia, M.; Chavez, M. Effects of a high molecular weight polypeptidic toxin from Physalia physalis (Portuguese man-of-war) on cholinergic responses. Comp. Biochem. Physiol. Part C Comp. Pharmacol. 1990, 95, 63–69. [Google Scholar]
- Tamkun, M.M.; Hessinger, D.A. Isolation and partial characterization of a hemolytic and toxic protein from the nematocyst venom of the Portuguese man-of-war, Physalia physalis. Biochim. Biophys. Acta (BBA) Protein Struct. 1981, 667, 87–98. [Google Scholar] [CrossRef]
- Neeman, I.; Calton, G.J.; Burnett, J.W. Purification and characterization of the endonuclease present in Physalia physalis venom. Comp. Biochem. Physiol. Part B Comp. Biochem. 1980, 67, 155–158. [Google Scholar] [CrossRef]
- Cormier, S.M. Physalia venom mediates histamine release from mast cells. J. Exp. Zool. 1981, 218, 117–120. [Google Scholar] [CrossRef] [PubMed]
- Radwan, F.F.Y.; Burnett, J.W.; Bloom, D.A.; Coliano, T.; Eldefrawi, M.E.; Erderly, H.; Aurelian, L.; Torres, M.; Heimer-de la Cotera, E.P. A comparison of the toxinological characteristics of two Cassiopea and Aurelia species. Toxicon 2001, 39, 245–257. [Google Scholar] [CrossRef]
- Bayazit, V. Cytotoxic effects of some animal and vegetable extracts and some chemicals on adenohypophyse carcinoma, kidney adenocarcinoma and skin carcinoma cells. J. Med. Sci. 2004, 4, 1–10. [Google Scholar]
- Burnett, J.W.; Calton, G.J.; Larsen, J.B. Significant envenomation by Aurelia aurita, the moon jellyfish. Toxicon 1988, 26, 215–217. [Google Scholar] [CrossRef]
- Lee, H.; Jung, E.-S.; Kang, C.; Yoon, W.D.; Kim, J.-S.; Kim, E. Scyphozoan jellyfish venom metalloproteinases and their role in the cytotoxicity. Toxicon 2011, 58, 277–284. [Google Scholar] [CrossRef]
- Ovchinnikova, T.V.; Balandin, S.V.; Aleshina, G.M.; Tagaev, A.A.; Leonova, Y.F.; Krasnodembsky, E.D.; Men’shenin, A.V.; Kokryakov, V.N. Aurelin, a novel antimicrobial peptide from jellyfish Aurelia aurita with structural features of defensins and channel-blocking toxins. Biochem. Biophys. Res. Commun. 2006, 348, 514–523. [Google Scholar] [CrossRef]
- Kokelj, F.; Del Negro, P.; Tubaro, A. Dermossicità da Chrysaora hysoscella. G. Ital. Dermatol. Venereol. 1989, 124, 297–298. [Google Scholar] [PubMed]
- Walker, M.J.A. Pharmacological and biochemical properties of a toxin containing material from the jellyfish, Cyanea capillata. Toxicon 1977, 15, 3–14. [Google Scholar] [CrossRef]
- Lassen, S.; Helmholz, H.; Ruhnau, C.; Prange, A. A novel proteinaceous cytotoxin from the northern Scyphozoa Cyanea capillata (L.) with structural homology to cubozoan haemolysins. Toxicon 2011, 57, 721–729. [Google Scholar] [CrossRef]
- Lassen, S.; Wiebring, A.; Helmholz, H.; Ruhnau, C.; Prange, A. Isolation of a Nav channel blocking polypeptide from Cyanea capillata medusae—A neurotoxin contained in fishing tentacle isorhizas. Toxicon 2012, 59, 610–616. [Google Scholar] [CrossRef]
- Helmholz, H.; Ruhnau, C.; Schütt, C.; Prange, A. Comparative study on the cell toxicity and enzymatic activity of two northern scyphozoan species Cyanea capillata (L.) and Cyanea lamarckii (Péron & Léslieur). Toxicon 2007, 50, 53–64. [Google Scholar] [CrossRef] [PubMed]
- Kang, C.; Munawir, A.; Cha, M.; Sohn, E.-T.; Lee, H.; Kim, J.-S.; Yoon, W.D.; Lim, D.; Kim, E. Cytotoxicity and hemolytic activity of jellyfish Nemopilema nomurai (Scyphozoa: Rhizostomeae) venom. Comp. Biochem. Physiol. Part C Toxicol. Pharmacol. 2009, 150, 85–90. [Google Scholar] [CrossRef] [PubMed]
- Quadrifoglio, F.; Avian, M.; Del Negro, P.; Princi, T.; Scuka, M.; Gavinelli, E.; Rottini Sandrini, L. Nematocisti e tossine di Pelagia noctiluca (Forsskål). Nova Thalass. 1986, 8, 155–162. [Google Scholar]
- Salleo, A.; Calabrese, L.; Barra, D.; La Spada, G. Characterization of protein components of the capsule fluid ad of the capsule wall of the nematocysts of Pelagia noctiluca. Nova Thalass. 1986, 8, 119–122. [Google Scholar]
- Scarpa, C.; Kokelj, F.; Del Negro, P.; Tubaro, A. Valutazione dell’effetto irritante sulla cute umana di una preparazione di nematocisti di Pelagia noctiluca. Ann. It. Derm. Clin. Sper. 1987, 41, 337–341. [Google Scholar]
- Mariottini, G.L.; Giacco, E.; Pane, L. The mauve stinger Pelagia noctiluca (Forsskal, 1775). Distribution, ecology, toxicity and epidemiology of stings. A review. Mar. Drugs 2008, 6, 496–513. [Google Scholar] [CrossRef]
- Morabito, R.; Marino, A.; La Spada, G.; Pane, L.; Mariottini, G.L. The venom and the toxicity of Pelagia noctiluca (Cnidaria: Scyphozoa). A review of three decades of research in Italian laboratories and future perspectives. J. Biol. Res. 2015, 88. [Google Scholar] [CrossRef]
- Carneiro, R.F.V.; Nascimento, N.R.F.d.; Costa, P.P.C.; Gomes, V.M.; de Souza, A.J.F.; de Oliveira, S.C.B.; dos Santos Diz Filho, E.B.; Zara, F.J.; Fonteles, M.C.; de Oliveira Toyama, D. The extract of the jellyfish Phyllorhiza punctata promotes neurotoxic effects. J. Appl. Toxicol. 2011, 31, 720–729. [Google Scholar] [CrossRef]
- Rastogi, A.; Sarkar, A.; Chakrabarty, D. Partial purification and identification of a metalloproteinase with anticoagulant activity from Rhizostoma pulmo (Barrel Jellyfish). Toxicon 2017, 132, 29–39. [Google Scholar] [CrossRef]
- Cariello, L.; Romano, G.; Spagnuolo, A.; Zanetti, L. Isolation and partial characterization of Rhizolysin, a high molecular weight protein with hemolytic activity, from the jellyfish Rhizostoma pulmo. Toxicon 1988, 26, 1057–1065. [Google Scholar] [CrossRef]
- Allavena, A.; Mariottini, G.L.; Carli, A.M.; Contini, S.; Martelli, A. In vitro evaluation of the cytotoxic, hemolytic and clastogenic activities of Rhizostoma pulmo toxin(s). Toxicon 1998, 36, 933–936. [Google Scholar] [CrossRef]
- Li, C.; Yu, H.; Liu, S.; Xing, R.; Guo, Z.; Li, P. Factors affecting the protease activity of venom from jellyfish Rhopilema esculentum Kishinouye. BioOrg. Med. Chem. Lett. 2005, 15, 5370–5374. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.; Li, C.; Li, R.; Xing, R.; Liu, S.; Li, P. Factors influencing hemolytic activity of venom from the jellyfish Rhopilema esculentum Kishinouye. Food Chem. Toxicol. 2007, 45, 1173–1178. [Google Scholar] [CrossRef] [PubMed]
- Gusmani, L.; Avian, M.; Galil, B.; Patriarca, P.; Rottini, G. Biologically active polypeptides in the venom of the jellyfish Rhopilema nomadica. Toxicon 1997, 35, 637–648. [Google Scholar] [CrossRef]
- Yoffe, B.; Baruchin, A.M. Mediterranean jellyfish (Rhopilema nomadica) sting. Burns 2004, 5, 503–504. [Google Scholar] [CrossRef]
- Badré, S. Bioactive toxins from stinging jellyfish. Toxicon 2014, 91, 114–125. [Google Scholar] [CrossRef]
- Li, R.; Yu, H.; Xue, W.; Yue, Y.; Liu, S.; Xing, R.; Li, P. Jellyfish venomics and venom gland transcriptomics analysis of Stomolophus meleagris to reveal the toxins associated with sting. J. Proteom. 2014, 106, 17–29. [Google Scholar] [CrossRef]
- Chung, J.J.; Ratnapala, L.A.; Cooke, I.M.; Yanagihara, A.A. Partial purification and characterization of a hemolysin (CAH1) from Hawaiian box jellyfish (Carybdea alata) venom. Toxicon 2001, 39, 981–990. [Google Scholar] [CrossRef]
- Nagai, H.; Takuwa, K.; Nakao, M.; Ito, E.; Miyake, M.; Noda, M.; Nakajima, T. Novel proteinaceous toxins from the box jellyfish (sea wasp) Carybdea rastoni. Biochem. Biophys. Res. Commun. 2000, 275, 582–588. [Google Scholar] [CrossRef]
- Rottini, G.; Gusmani, L.; Parovel, E.; Avian, M.; Patriarca, P. Purification and properties of a cytolytic toxin in venom of the jellyfish Carybdea marsupialis. Toxicon 1995, 33, 315–326. [Google Scholar] [CrossRef]
- Sánchez-Rodríguez, J.; Torrens, E.; Segura-Puertas, L. Partial purification and characterization of a novel neurotoxin and three cytolysins from box jellyfish (Carybdea marsupialis) nematocyst venom. Arch. Toxicol. 2006, 80, 163–168. [Google Scholar] [CrossRef] [PubMed]
- Azuma, H.; Sekizaki, S.; Satoh, A.; Nakajima, T. Platelet aggregation caused by Carybdea rastonii toxins (CrTX-I, II, and III) obtained from a jellyfish, ++stonii. Proc. Soc. Exp. Biol. Med. 1986, 182, 34–42. [Google Scholar] [CrossRef] [PubMed]
- Avila Soria, G. Molecular Characterization of Carukia barnesi and Malo kingi, Cnidaria; Cubozoa; Carybdeidae. Ph.D. Thesis, James Cook University, Townsville, Australia, 2009. [Google Scholar]
- Jouiaei, M.; Casewell, N.; Yanagihara, A.; Nouwens, A.; Cribb, B.; Whitehead, D.; Jackson, T.; Ali, S.A.; Wagstaff, S.; Koludarov, I.; et al. Firing the sting: Chemically induced discharge of cnidae reveals novel proteins and peptides from box jellyfish (Chironex fleckeri) venom. Toxins 2015, 7, 936–950. [Google Scholar] [CrossRef] [PubMed]
- Brinkman, D.L.; Burnell, J.N. Biochemical and molecular characterisation of cubozoan protein toxins. Toxicon 2009, 54, 1162–1173. [Google Scholar] [CrossRef]
- Nagai, H.; Takuwa-Kuroda, K.; Nakao, M.; Oshiro, N.; Iwanaga, S.; Nakajima, T. A novel protein toxin from the deadly box jellyfish (sea wasp, Habu-kurage) Chiropsalmus quadrigatus. Biosci. Biotechnol. Biochem. 2002, 66, 97–102. [Google Scholar] [CrossRef]
- Ramasamy, S.; Isbister, G.K.; Seymour, J.E.; Hodgson, W.C. The in vitro effects of two chirodropid (Chironex fleckeri and Chiropsalmus sp.) venoms: Efficacy of box jellyfish antivenom. Toxicon 2003, 41, 703–711. [Google Scholar] [CrossRef]
- Lin, X.-Y.; Ishida, M.; Nagashima, Y.; Kazuo, S. A polypeptide toxin in the sea anemone Actinia equina homologous with other sea anemone sodium channel toxins: Isolation and amino acid sequence. Toxicon 1996, 34, 57–65. [Google Scholar] [CrossRef]
- Shiomi, K.; Ishikawa, M.; Yamanaka, H.; Kikuchi, T. Isolation and properties of four serine protease inhibitors in the sea anemone Actinia equina. Nippon Suisan Gakkaishi 1989, 55, 1235–1241. [Google Scholar] [CrossRef]
- Maček, P.; Lebez, D. Isolation and characterization of three lethal and hemolytic toxins from the sea anemone Actinia equina L. Toxicon 1988, 26, 441–451. [Google Scholar] [CrossRef]
- Anderluh, G.; Križaj, I.; Štrukelj, B.; Gubenšek, F.; Maček, P.; Pungerčar, J. Equinatoxins, pore-forming proteins from the sea anemone Actinia equina, belong to a multigene family. Toxicon 1999, 37, 1391–1401. [Google Scholar] [CrossRef]
- Pungercar, J.; Anderluh, G.; Maček, P.; Franc, G.; Štrukelj, B. Sequence analysis of the cDNA encoding the precursor of equinatoxin V, a newly discovered hemolysin from the sea anemone Actinia equina. Biochim. Biophys. Acta (BBA) Protein Struct. Mol. Enzymol. 1997, 1341, 105–107. [Google Scholar] [CrossRef]
- Honma, T.; Minagawa, S.; Nagai, H.; Ishida, M.; Nagashima, Y.; Shiomi, K. Novel peptide toxins from acrorhagi, aggressive organs of the sea anemone Actinia equina. Toxicon 2005, 46, 768–774. [Google Scholar] [CrossRef] [PubMed]
- Bellomio, A.; Morante, K.; Barlič, A.; Gutiérrez-Aguirre, I.; Viguera, A.R.; Gonzàlez-Mañas, J.M. Purification, cloning and characterization of fragaceatoxin C, a novel actinoporin from the sea anemone Actinia fragacea. Toxicon 2009, 54, 869–880. [Google Scholar] [CrossRef] [PubMed]
- Norton, R.S.; Bobek, G.; Ivanov, J.O.; Thomson, M.; Fiala-Beer, E.; Moritz, R.L.; Simpson, R.J. Purification and characterisation of proteins with cardiac stimulatory and haemolytic activity from the anemone Actinia tenebrosa. Toxicon 1990, 28, 29–41. [Google Scholar] [CrossRef]
- Diochot, S.; Schweitz, H.; Béress, L.; Lazdunski, M. Sea anemone peptides with a specific blocking activity against the fast inactivating potassium channel Kv3.4. J. Biol. Chem. 1998, 273, 6744–6749. [Google Scholar] [CrossRef]
- Uechi, G.-I.; Toma, H.; Arakawa, T.; Sato, Y. Molecular characterization on the genome structure of hemolysin toxin isoforms isolated from sea anemone Actineria villosa and Phyllodiscus semoni. Toxicon 2010, 56, 1470–1476. [Google Scholar] [CrossRef]
- Oshiro, N.; Kobayashi, C.; Iwanaga, S.; Nozaki, M.; Namikoshi, M.; Spring, J.R.; Nagai, H. A new membrane-attack complex/perforin (MACPF) domain lethal toxin from the nematocyst venom of the Okinawan sea anemone Actineria villosa. Toxicon 2004, 43, 225–228. [Google Scholar] [CrossRef]
- Uechi, G.-I.; Toma, H.; Arakawa, T.; Sato, Y. Characterization of a novel proteinous toxin from sea anemone Actineria villosa. Protein J. 2011, 30, 422. [Google Scholar] [CrossRef]
- Talvinen, K.A.; Nevalainen, T.J. Cloning of a novel phospholipase A2 from the cnidarian Adamsia carciniopados. Comp. Biochem. Physiol. Part B Biochem. Mol. Biol. 2002, 132, 571–578. [Google Scholar] [CrossRef]
- Shiomi, K.; Qian, W.-H.; Lin, X.-Y.; Shimakura, K.; Nagashima, Y.; Ishida, M. Novel polypeptide toxins with crab lethality from the sea anemone Anemonia erythraea. Biochim. Biophys. Acta (BBA) Gen. Subj. 1997, 1335, 191–198. [Google Scholar] [CrossRef]
- Hasegawa, Y.; Honma, T.; Nagai, H.; Ishida, M.; Nagashima, Y.; Shiomi, K. Isolation and cDNA cloning of a potassium channel peptide toxin from the sea anemone Anemonia erythraea. Toxicon 2006, 48, 536–542. [Google Scholar] [CrossRef] [PubMed]
- Wunderer, G.; Eulitz, M. Amino-acid sequence of toxin I from Anemonia sulcata. Eur. J. Biochem. 1978, 89, 11–17. [Google Scholar] [CrossRef] [PubMed]
- Béress, L.; Béress, R.; Wunderer, G. Isolation and characterisation of three polypeptides with neurotoxic activity from Anemonia sulcata. Febs Lett. 1975, 50, 311–314. [Google Scholar] [CrossRef]
- Scheffler, J.-J.; Tsugita, A.; Linden, G.; Schweitz, H.; Lazdunski, M. The amino acid sequence of toxin V from Anemonia sulcata. Biochem. Biophys. Res. Commun. 1982, 107, 272–278. [Google Scholar] [CrossRef]
- Wunderer, G.; Bèress, L.; Machleidt, W.; Fritz, H. Broad-specificity inhibitors from sea anemones. In Methods in Enzymology; Academic Press: London, UK, 1976; Volume 45, pp. 881–888. [Google Scholar]
- Schweitz, H.; Bruhn, T.; Guillemare, E.; Moinier, D.; Lancelin, J.M.; Béress, L.; Lazdunski, M. Kalicludines and kaliseptine. Two different classes of sea anemone toxins for voltage sensitive K+ channels. J. Biol. Chem. 1995, 270, 25121–25126. [Google Scholar] [CrossRef]
- Kohno, Y.; Satoh, H.; Iguchi, A.; Nagai, H. Characterization of a new hemolytic protein toxin from the sea anemone Anthopleura asiatica. Fish. Sci. 2009, 75, 1049–1054. [Google Scholar] [CrossRef]
- Norton, T.R. Cardiotonic polypeptides from Anthopleura xanthogrammica (Brandt) and A. elegantissima (Brandt). Fed. Proc. 1981, 40, 21–25. [Google Scholar]
- Bruhn, T.; Schaller, C.; Schulze, C.; Sanchez-Rodriguez, J.; Dannmeier, C.; Ravens, U.; Heubach, J.F.; Eckhardt, K.; Schmidtmayer, J.; Schmidt, H.; et al. Isolation and characterisation of five neurotoxic and cardiotoxic polypeptides from the sea anemone Anthopleura elegantissima. Toxicon 2001, 39, 693–702. [Google Scholar] [CrossRef]
- Diochot, S.; Baron, A.; Rash, L.D.; Deval, E.; Escoubas, P.; Scarzello, S.; Salinas, M.; Lazdunski, M. A new sea anemone peptide, APETx2, inhibits ASIC3, a major acid-sensitive channel in sensory neurons. Embo J. 2004, 23, 1516–1525. [Google Scholar] [CrossRef]
- Sunahara, S.; Muramoto, K.; Tenma, K.; Kamiya, H. Amino acid sequence of two sea anemone toxins from Anthopleura fuscoviridis. Toxicon 1987, 25, 211–219. [Google Scholar] [CrossRef]
- Kelso, G.J.; Blumenthal, K.M. Identification and characterization of novel sodium channel toxins from the sea anemone Anthopleura xanthogrammica. Toxicon 1998, 36, 41–51. [Google Scholar] [CrossRef]
- Minagawa, S.; Ishida, M.; Shimakura, K.; Nagashima, Y.; Shiomi, K. Isolation and amino acid sequences of two kunitz-type protease inhibitors from the sea anemone Anthopleura aff. xanthogrammica. Comp. Biochem. Physiol. Part B Biochem. Mol. Biol. 1997, 118, 381–386. [Google Scholar] [CrossRef]
- Wang, L.; Ou, J.; Peng, L.; Zhong, X.; Du, J.; Liu, Y.; Huang, Y.; Liu, W.; Zhang, Y.; Dong, M.; et al. Functional expression and characterization of four novel neurotoxins from sea anemone Anthopleura sp. Biochem. Biophys. Res. Commun. 2004, 313, 163–170. [Google Scholar] [CrossRef] [PubMed]
- Malpezzi, E.L.A.; de Freitas, J.C.; Muramoto, K.; Kamiya, H. Characterization of peptides in sea anemone venom collected by a novel procedure. Toxicon 1993, 31, 853–864. [Google Scholar] [CrossRef]
- Oliveira, J.S.; Zaharenko, A.J.; Ferreira, W.A.; Konno, K.; Shida, C.u.S.; Richardson, M.; Lùcio, A.D.; Beirão, P.S.L.; de Freitas, J.C. BcIV, a new paralyzing peptide obtained from the venom of the sea anemone Bunodosoma caissarum. A comparison with the Na+ channel toxin BcIII. Biochim. Biophys. Acta (BBA) Proteins Proteom. 2006, 1764, 1592–1600. [Google Scholar] [CrossRef]
- Martins, R.D.; Alves, R.S.; Martins, A.M.C.; Barbosa, P.S.F.; Evangelista, J.S.A.M.; Evangelista, J.J.F.; Ximenes, R.M.; Toyama, M.H.; Toyama, D.O.; Souza, A.J.F.; et al. Purification and characterization of the biological effects of phospholipase A2 from sea anemone Bunodosoma caissarum. Toxicon 2009, 54, 413–420. [Google Scholar] [CrossRef]
- Orts, D.J.B.; Moran, Y.; Cologna, C.T.; Peigneur, S.; Madio, B.; Praher, D.; Quinton, L.; De Pauw, E.; Bicudo, J.E.P.W.; Tytgat, J.; et al. BcsTx3 is a founder of a novel sea anemone toxin family of potassium channel blocker. Febs J. 2013, 280, 4839–4852. [Google Scholar] [CrossRef]
- Loret, E.; Menendez, R.; Mansuelle, P.; Sampieri, F.; Rochat, H. Positively charged amino acid residues located similarly in sea anemone and scorpion toxins. J. Biol. Chem. 1994, 269, 16785–16788. [Google Scholar]
- Aneiros, A.; Garcìa, I.; Martìnez, J.R.; Harvey, A.L.; Anderson, A.J.; Marshall, D.L.; Engström, Å.; Hellman, U.; Karlsson, E. A potassium channel toxin from the secretion of the sea anemone Bunodosoma granulifera. isolation, amino acid sequence and biological activity. Biochim. Biophys. Acta (BBA) Gen. Subj. 1993, 1157, 86–92. [Google Scholar] [CrossRef]
- Santana, A.N.C.; Leite, A.B.; França, M.S.F.; França, L.; Vale, O.C.; Cunha, R.B.; Ricart, C.A.O.; Sousa, M.V.; Carvalho, K.M. Partial sequence and toxic effects of granulitoxin, a neurotoxic peptide from the sea anemone Bunodosoma granulifera. Braz. J. Med Biol. Res. 1998, 31, 1335–1338. [Google Scholar] [CrossRef]
- Zaharenko, A.J.; Ferreira, W.A.; Oliveira, J.S.; Richardson, M.; Pimenta, D.C.; Konno, K.; Portaro, F.C.V.; de Freitas, J.C. Proteomics of the neurotoxic fraction from the sea anemone Bunodosoma cangicum venom: Novel peptides belonging to new classes of toxins. Comp. Biochem. Physiol. Part D Genom. Proteom. 2008, 3, 219–225. [Google Scholar] [CrossRef] [PubMed]
- Zaharenko, A.J.; Ferreira, W.A.; de Oliveira, J.S.; Konno, K.; Richardson, M.; Schiavon, E.; Wanke, E.; de Freitas, J.C. Revisiting cangitoxin, a sea anemone peptide: Purification and characterization of cangitoxins II and III from the venom of Bunodosoma cangicum. Toxicon 2008, 51, 1303–1307. [Google Scholar] [CrossRef] [PubMed]
- Cariello, L.; De Santis, A.; Fiore, F.; Piccoli, R.; Spagnuolo, A.; Zanetti, L.; Parente, A. Calitoxin, a neurotoxic peptide from the sea anemone Calliactis parasitica: Amino-acid sequence and electrophysiological properties. Biochemistry 1989, 28, 2484–2489. [Google Scholar] [CrossRef] [PubMed]
- Spagnuolo, A.; Zanetti, L.; Cariello, L.; Piccoli, R. Isolation and characterization of two genes encoding calitoxins, neurotoxic peptides from Calliactis parasitica (Cnidaria). Gene 1994, 138, 187–191. [Google Scholar] [CrossRef]
- Ständker, L.; Béress, L.; Garateix, A.; Christ, T.; Ravens, U.; Salceda, E.; Soto, E.; John, H.; Forssmann, W.-G.; Aneiros, A. A new toxin from the sea anemone Condylactis gigantea with effect on sodium channel inactivation. Toxicon 2006, 48, 211–220. [Google Scholar] [CrossRef] [PubMed]
- Shiomi, K.; Lin, X.-Y.; Nagashima, Y.; Ishida, M. Isolation and aminoacid sequences of polypeptide toxins in the Caribbean Sea anemone Condylactis passiflora. Fish. Sci. 1995, 61, 1016–1021. [Google Scholar] [CrossRef]
- Maeda, M.; Honma, T.; Shiomi, K. Isolation and cDNA cloning of type 2 sodium channel peptide toxins from three species of sea anemones (Cryptodendrum adhaesivum, Heterodactyla hemprichii and Thalassianthus aster) belonging to the family Thalassianthidae. Comp. Biochem. Physiol. Part B Biochem. Mol. Biol. 2010, 157, 389–393. [Google Scholar] [CrossRef]
- Ishida, M.; Yokoyama, A.; Shimakura, K.; Nagashima, Y.; Shiomi, K. Halcurin, a polypeptide toxin from the sea anemone Halcurias sp., with a structural resemblance to type 1 and 2 toxins. Toxicon 1997, 35, 537–544. [Google Scholar] [CrossRef]
- Zykova, T.A.; Kozlovskaia, E.P. Amino acid sequence of a neurotoxin from the anemone Radianthus macrodactylus. Bioorganicheskaia Khimiia 1989, 15, 1301–1306. [Google Scholar]
- Zykova, T.A.; Kozlovskaia, E.P.; Eliakov, G.B. Amino acid sequence of neurotoxin II from the sea anemone Radianthus macrodactylus. Bioorganicheskaia Khimiia 1988, 14, 878–882. [Google Scholar]
- Zykova, T.A.; Kozlovskaia, E.P. Disulfide bonds in neurotoxin-III from the sea anenome Radianthus macrodactylus. Bioorganicheskaia Khimiia 1989, 15, 904–907. [Google Scholar] [PubMed]
- Zykova, T.A.; Kozlovskaia, E.P.; Eliakov, G.B. Amino acid sequence of neurotoxins IV and V from the sea anemone Radianthus macrodactylus. Bioorganicheskaia Khimiia 1988, 14, 1489–1494. [Google Scholar] [PubMed]
- Shiomi, K.; Lin, X.-Y.; Nagashima, Y.; Ishida, M. Isolation and amino acid sequence of a polypeptide toxin from the sea anemone Radianthus crispus. Fish. Sci. 1996, 62, 629–633. [Google Scholar] [CrossRef]
- Zykova, T.A.; Vinokurov, L.M.; Markova, L.F.; Kozlovskaya, E.P.; Elyakov, G.B. Amino-acid sequence of trypsin inhibitor IV from Radianthus macrodactylus. Bioorg. Khim. 1985, 11, 293–301. [Google Scholar]
- Il’ina, A.; Lipkin, A.; Barsova, E.; Issaeva, M.; Leychenko, E.; Guzev, K.; Monastyrnaya, M.; Lukyanov, S.; Kozlovskaya, E. Amino acid sequence of RTX-A’s isoform actinoporin from the sea anemone, Radianthus macrodactylus. Toxicon 2006, 47, 517–520. [Google Scholar] [CrossRef]
- Klyshko, E.V.; Issaeva, M.P.; Monastyrnaya, M.M.; Il’yna, A.P.; Guzev, K.V.; Vakorina, T.I.; Dmitrenok, P.S.; Zykova, T.A.; Kozlovskaya, E.P. Isolation, properties and partial amino acid sequence of a new actinoporin from the sea anemone Radianthus macrodactylus. Toxicon 2004, 44, 315–324. [Google Scholar] [CrossRef]
- Kvetkina, A.; Leychenko, E.; Chausova, V.; Zelepuga, E.; Chernysheva, N.; Guzev, K.; Pislyagin, E.; Yurchenko, E.; Menchinskaya, E.; Aminin, D.; et al. A new multigene HCIQ subfamily from the sea anemone Heteractis crispa encodes Kunitz-peptides exhibiting neuroprotective activity against 6-hydroxydopamine. Sci. Rep. 2020, 10, 4205. [Google Scholar] [CrossRef]
- Kozlov, S.A.; Osmakov, D.I.; Andreev, Y.A.; Koshelev, S.G.; Gladkikh, I.N.; Monastyrnaya, M.M.; Kozlovskaya, E.P.; Grishin, E.V. A sea anemone polypeptide toxin inhibiting the ASIC3 acid-sensitive channel. Russ. J. BioOrg. Chem. 2012, 38, 578–583. [Google Scholar] [CrossRef]
- Kalina, R.; Gladkikh, I.; Dmitrenok, P.; Chernikov, O.; Koshelev, S.; Kvetkina, A.; Kozlov, S.; Kozlovskaya, E.; Monastyrnaya, M. New APETx-like peptides from sea anemone Heteractis crispa modulate ASIC1a channels. Peptides 2018, 104, 41–49. [Google Scholar] [CrossRef]
- Gladkikh, I.N.; Kvetkina, A.N.; Kostina, E.E.; Kalina, R.S.; Grebnev, B.B.; Koshelev, S.G.; Kozlov, S.A.; Monastyrnaya, M.M.; Kozlovskaya, E.P. Peptide modulators of ASIC channels of the sea anemone Urticina aff. coriacea (Cuvier, 1798) from the Sea of Okhotsk. Russ. J. Mar. Biol. 2018, 44, 458–464. [Google Scholar] [CrossRef]
- Gladkikh, I.; Monastyrnaya, M.; Zelepuga, E.; Sintsova, O.; Tabakmakher, V.; Gnedenko, O.; Ivanov, A.; Hua, K.-F.; Kozlovskaya, E. New Kunitz-Type HCRG polypeptides from the sea anemone Heteractis crispa. Marine Drugs 2015, 13, 6038–6063. [Google Scholar] [CrossRef] [PubMed]
- Monastyrnaya, M.; Peigneur, S.; Zelepuga, E.; Sintsova, O.; Gladkikh, I.; Leychenko, E.; Isaeva, M.; Tytgat, J.; Kozlovskaya, E. Kunitz-type peptide HCRG21 from the sea anemone Heteractis crispa is a full antagonist of the TRPV1 receptor. Mar. Drugs 2016, 14, 229. [Google Scholar] [CrossRef] [PubMed]
- Andreev, Y.A.; Kozlov, S.A.; Koshelev, S.G.; Ivanova, E.A.; Monastyrnaya, M.M.; Kozlovskaya, E.P.; Grishin, E.V. Analgesic compound from sea anemone Heteractis crispa is the first polypeptide inhibitor of vanilloid receptor 1 (TRPV1). J. Biol. Chem. 2008, 283, 23914–23921. [Google Scholar] [PubMed]
- Andreev, Y.A.; Kozlov, S.A.; Korolkova, Y.V.; Dyachenko, I.A.; Bondarenko, D.A.; Skobtsov, D.I.; Murashev, A.N.; Kotova, P.D.; Rogachevskaja, O.A.; Kabanova, N.V.; et al. Polypeptide modulators of TRPV1 produce analgesia without hyperthermia. Mar. Drugs 2013, 11, 5100–5115. [Google Scholar] [CrossRef] [PubMed]
- Kozlov, S.A.; Andreev, Y.A.; Murashev, A.N.; Skobtsov, D.I.; D′yachenko, I.A.; Grishin, E.V. New polypeptide components from the Heteractis crispa sea anemone with analgesic activity. Russ. J. Bioorganic Chem. 2009, 35, 711–719. [Google Scholar] [CrossRef]
- Tabakmakher, V.; Sintsova, O.; Krivoshapko, O.; Zelepuga, E.; Monastyrnaya, M.; Kozlovskaya, E.P. Analgesic effect of novel Kunitz-type polypeptides of the sea anemone Heteractis crispa. Dokl. Biochem. Biophys. 2015, 461, 80–83. [Google Scholar] [CrossRef]
- Sintsova, O.V.; Monastyrnaya, M.M.; Pislyagin, E.A.; Menchinskaya, E.S.; Leychenko, E.V.; Aminin, D.L.; Kozlovskaya, E.P. Anti-inflammatory activity of a polypeptide from the Heteractis crispa sea anemone. Russ. J. BioOrg. Chem. 2015, 41, 590–596. [Google Scholar] [CrossRef]
- Sintsova, O.; Pislyagin, E.; Gladkikh, I.; Monastyrnaya, M.; Menchinskaya, E.; Leychenko, E.; Aminin, D.; Kozlovskaya, E. Kunitz-type peptides of the sea anemone Heteractis crispa: Potential anti-inflammatory compounds. Russ. J. BioOrg. Chem. 2017, 43, 91–97. [Google Scholar] [CrossRef]
- Khoo, K.S.; Kam, W.K.; Khoo, H.E.; Gopalakrishnakone, P.; Chung, M.C.M. Purification and partial characterization of two cytolysins from a tropical sea anemone, Heteractis magnifica. Toxicon 1993, 31, 1567–1579. [Google Scholar] [CrossRef]
- Sintsova, O.; Gladkikh, I.; Chausova, V.; Monastyrnaya, M.; Anastyuk, S.; Chernikov, O.; Yurchenko, E.; Aminin, D.; Isaeva, M.; Leychenko, E.; et al. Peptide fingerprinting of the sea anemone Heteractis magnifica mucus revealed neurotoxins, Kunitz-type proteinase inhibitors and a new β-defensin α-amylase inhibitor. J. Proteom. 2018, 173, 12–21. [Google Scholar] [CrossRef]
- Wang, Y.; Chua, K.L.; Khoo, H.E. A new cytolysin from the sea anemone, Heteractis magnifica: Isolation, cDNA cloning and functional expression1. Biochim. Biophys. Acta (BBA) Protein Struct. Mol. Enzymol. 2000, 1478, 9–18. [Google Scholar] [CrossRef]
- Gendeh, G.S.; Young, L.C.; de Medeiros, C.L.C.; Jeyaseelan, K.; Harvey, A.L.; Chung, M.C.M. A new potassium channel toxin from the sea anemone Heteractis magnifica: Isolation, cDNA cloning, and functional expression. Biochemistry 1997, 36, 11461–11471. [Google Scholar] [CrossRef] [PubMed]
- Schweitz, H.; Bidard, J.N.; Frelin, C.; Pauron, D.; Vijverberg, H.P.M.; Mahasneh, D.M.; Lazdunski, M.; Vilbois, F.; Tsugita, A. Purification, sequence, and pharmacological properties of sea anemone toxins from Radianthus paumotensis. A new class of sea anemone toxins acting on the sodium channel. Biochemistry 1985, 24, 3554–3561. [Google Scholar] [CrossRef] [PubMed]
- Kvetkina, A.N.; Leychenko, E.V.; Yurchenko, E.A.; Pislyagin, E.A.; Peigneur, S.; Tytgat, Y.; Isaeva, M.P.; Aminin, D.L.; Kozlovskaya, E.P. A new Iq-peptide of the Kunitz Type from the Heteractis magnifica sea anemone exhibits neuroprotective activity in a model of Alzheimer’s disease. Russ. J. BioOrg. Chem. 2018, 44, 416–423. [Google Scholar] [CrossRef]
- Sintsova, O.; Gladkikh, I.; Kalinovskii, A.; Zelepuga, E.; Monastyrnaya, M.; Kim, N.; Shevchenko, L.; Peigneur, S.; Tytgat, J.; Kozlovskaya, E.; et al. Magnificamide, a neta-defensin-like peptide from the mucus of the sea anemone Heteractis magnifica, Is a strong inhibitor of mammalian alpha-amylases. Mar. Drugs 2019, 17, 542. [Google Scholar] [CrossRef]
- Krebs, H.C.; Habermehl, G.G. Isolierung und strukturaufklärung eines hämolytisch aktiven peptids aus der seeanemone Metridium senile. Naturwissenschaften 1987, 74, 395–396. [Google Scholar] [CrossRef]
- Logashina, Y.A.; Mosharova, I.V.; Korolkova, Y.V.; Shelukhina, I.V.; Dyachenko, I.A.; Palikov, V.A.; Palikova, Y.A.; Murashev, A.N.; Kozlov, S.A.; Stensvåg, K.; et al. Peptide from sea anemone Metridium senile affects transient receptor potential ankyrin-repeat 1 (TRPA1) function and produces analgesic effect. J. Biol. Chem. 2017, 292, 2992–3004. [Google Scholar] [CrossRef]
- Moran, Y.; Weinberger, H.; Sullivan, J.C.; Reitzel, A.M.; Finnerty, J.R.; Gurevitz, M. Concerted evolution of sea anemone neurotoxin genes Is revealed through analysis of the Nematostella vectensis genome. Mol. Biol. Evol. 2008, 25, 737–747. [Google Scholar] [CrossRef]
- Sachkova, M.Y.; Singer, S.A.; Macrander, J.; Reitzel, A.M.; Peigneur, S.; Tytgat, J.; Moran, Y. The birth and death of toxins with distinct functions: A case study in the sea anemone Nematostella. Mol. Biol. Evol. 2019, 36, 2001–2012. [Google Scholar] [CrossRef]
- Moran, Y.; Praher, D.; Schlesinger, A.; Ayalon, A.; Tal, Y.; Technau, U. Analysis of soluble protein contents from the nematocysts of a model sea anemone sheds light on venom evolution. Mar. Biotechnol. 2013, 15, 329–339. [Google Scholar] [CrossRef]
- Il’ina, A.P.; Monastyrnaya, M.M.; Isaeva, M.P.; Guzev, K.V.; Rasskazov, V.A.; Kozlovskaya, E.P. Primary structures of actinoporins from sea anemone Oulactis orientalis. Russ. J. BioOrg. Chem. 2005, 31, 320–324. [Google Scholar] [CrossRef]
- Nishida, S.; Fujita, S.; Warashina, A.; Satake, M.; Tamiya, N. Amino acid sequence of a sea anemone toxin from Parasicyonis actinostoloides. Febs Lett. 1985, 150, 171–173. [Google Scholar] [CrossRef] [PubMed]
- Mizuno, M.; Nozaki, M.; Morine, N.; Suzuki, N.; Nishikawa, K.; Morgan, B.P.; Matsuo, S. A protein toxin from the sea anemone Phyllodiscus semoni targets the kidney and causes a severe renal injury with predominant glomerular endothelial damage. Am. J. Pathol. 2007, 171, 402–414. [Google Scholar] [CrossRef] [PubMed]
- Nagai, H.; Oshiro, N.; Takuwa-Kuroda, K.; Iwanaga, S.; Nozaki, M.; Nakajima, T. Novel proteinaceous toxins from the nematocyst venom of the Okinawan sea anemone Phyllodiscus semoni Kwietniewski. Biochem. Biophys. Res. Commun. 2002, 294, 760–763. [Google Scholar] [CrossRef]
- Rodrìguez, A.A.; Salceda, E.; Garateix, A.G.; Zaharenko, A.J.; Peigneur, S.; Lòpez, O.; Pons, T.; Richardson, M.; Dìaz, M.; Hernàndez, Y.; et al. A novel sea anemone peptide that inhibits acid-sensing ion channels. Peptides 2014, 53, 3–12. [Google Scholar] [CrossRef]
- Jiang, X.-Y.; Yang, W.-l.; Chen, H.-P.; Tu, H.-B.; Wu, W.-Y.; Wei, J.-W.; Wang, J.; Liu, W.-H.; Xu, A.-L. Cloning and characterization of an acidic cytolysin cDNA from sea anemone Sagartia rosea. Toxicon 2002, 40, 1563–1569. [Google Scholar] [CrossRef]
- Kem, W.R.; Parten, B.; Pennington, M.W.; Price, D.A.; Dunn, B.M. Isolation, characterization, and amino acid sequence of a polypeptide neurotoxin occurring in the sea anemone Stichodactyla helianthus. Biochemistry 1989, 28, 3483–3489. [Google Scholar] [CrossRef]
- Tysoe, C.; Williams, L.K.; Keyzers, R.; Nguyen, N.T.; Tarling, C.; Wicki, J.; Goddard-Borger, E.D.; Aguda, A.H.; Perry, S.; Foster, L.J.; et al. Potent human alpha-amylase inhibition by the beta-defensin-like protein Helianthamide. Acs. Cent. Sci. 2016, 2, 154–161. [Google Scholar] [CrossRef]
- Delfìn, J.; Martìnez, I.; Antuch, W.; Morera, V.; Gonzàlez, Y.; Rodìguez, R.; Màquez, M.; Saroyàn, A.; Larionova, N.; Dìaz, J.; et al. Purification, characterization and immobilization of proteinase inhibitors from Stichodactyla helianthus. Toxicon 1996, 34, 1367–1376. [Google Scholar] [CrossRef]
- Castañeda, O.; Sotolongo, V.; Amor, A.M.; Stðcklin, R.; Anderson, A.J.; Harvey, A.L.; Engstrðm, Å.; Wernstedt, C.; Karlsson, E. Characterization of a potassium channel toxin from the Caribbean sea anemone Stichodactyla helianthus. Toxicon 1995, 33, 603–613. [Google Scholar] [CrossRef]
- Huerta, V.; Morera, V.; Guanche, Y.; Chinea, G.; Gonzàlez, L.J.; Betancourt, L.; Martìnez, D.; Alvarez, C.; Lanio, M.E.; Besada, V. Primary structure of two cytolysin isoforms from Stichodactyla helianthus differing in their hemolytic activity. Toxicon 2001, 39, 1253–1256. [Google Scholar] [CrossRef]
- Shiomi, K.; Honma, T.; Ide, M.; Nagashima, Y.; Ishida, M.; Chino, M. An epidermal growth factor-like toxin and two sodium channel toxins from the sea anemone Stichodactyla gigantea. Toxicon 2003, 41, 229–236. [Google Scholar] [CrossRef]
- Honma, T.; Kawahata, S.; Ishida, M.; Nagai, H.; Nagashima, Y.; Shiomi, K. Novel peptide toxins from the sea anemone Stichodactyla haddoni. Peptides 2008, 29, 536–544. [Google Scholar] [CrossRef] [PubMed]
- Razpotnik, A.; Križaj, I.; Kem, W.R.; Maček, P.; Turk, T. A new cytolytic protein from the sea anemone Urticina crassicornis that binds to cholesterol- and sphingomyelin-rich membranes. Toxicon 2009, 53, 762–769. [Google Scholar] [CrossRef]
- Logashina, Y.A.; Solstad, R.G.; Mineev, K.S.; Korolkova, Y.V.; Mosharova, I.V.; Dyachenko, I.A.; Palikov, V.A.; Palikova, Y.A.; Murashev, A.N.; Arseniev, A.S.; et al. New disulfide-stabilized fold provides sea anemone peptide to exhibit both antimicrobial and TRPA1 potentiating properties. Toxins 2017, 9, 154. [Google Scholar] [CrossRef]
- Osmakov, D.I.; Kozlov, S.A.; Andreev, Y.A.; Koshelev, S.G.; Sanamyan, N.P.; Sanamyan, K.E.; Dyachenko, I.A.; Bondarenko, D.A.; Murashev, A.N.; Mineev, K.S.; et al. Sea anemone peptide with uncommon Y-hairpin structure inhibits acid-sensing ion channel 3 (ASIC3) and reveals analgesic activity. J. Biol. Chem. 2013, 288, 23116–23127. [Google Scholar] [CrossRef]
- Cline, E.I.; Wiebe, L.I.; Young, J.D.; Samuel, J. Toxic effects of the novel protein UPI from the sea anemone Urticina piscivora. Pharmacol. Res. 1995, 32, 309–314. [Google Scholar] [CrossRef]
- Murakami, M.; Kudo, I. Phospholipase A2. J. Biochem. 2002, 131, 285–292. [Google Scholar] [CrossRef]
- Romero, L.; Marcussi, S.; Marchi-Salvador, D.; Silva, F.; Fuly, A.; Stabeli, R.; Da Silva, S.; González, J.; del Monte-MartÃnez, A.; Soares, A. Enzymatic and structural characterization of a basic phospholipase A2 from the sea anemone Condylactis gigantea. Biochimie 2010, 92, 1063–1071. [Google Scholar] [CrossRef]
- Fry, B.G.; Roelants, K.; Champagne, D.E.; Scheib, H.; Tyndall, J.D.A.; King, G.F.; Nevalainen, T.J.; Norman, J.A.; Lewis, R.J.; Norton, R.S.; et al. The toxicogenomic multiverse: Convergent recruitment of proteins into animal venoms. Annu. Rev. Genom. Hum. Genet. 2009, 10, 483–511. [Google Scholar] [CrossRef]
- Undheim, E.A.B.; King, G.F. On the venom system of centipedes (Chilopoda), a neglected group of venomous animals. Toxicon 2011, 57, 512–524. [Google Scholar] [CrossRef] [PubMed]
- Fox, J.W.; Serrano, S.M.T. Structural considerations of the snake venom metalloproteinases, key members of the M12 reprolysin family of metalloproteinases. Toxicon 2005, 45, 969–985. [Google Scholar] [CrossRef] [PubMed]
- Messerli, S.M.; Greenberg, R.M. Cnidarian toxins acting on voltage-gate ion channels. Mar. Drugs 2006, 4, 70–81. [Google Scholar] [CrossRef]
- Smith, J.J.; Blumenthal, K.M. Site-3 sea anemone toxins: Molecular probes of gating mechanisms in voltage-dependent sodium channels. Toxicon 2007, 49, 159–170. [Google Scholar] [CrossRef] [PubMed]
- Kalina, R.S.; Peigneur, S.; Zelepuga, E.A.; Dmitrenok, P.S.; Kvetkina, A.N.; Kim, N.Y.; Leychenko, E.V.; Tytgat, J.; Kozlovskaya, E.P.; Monastyrnaya, M.M.; et al. New insights into the Type II toxins from the sea anemone Heteractis crispa. Toxins 2020, 12, 44. [Google Scholar] [CrossRef] [PubMed]
- Liao, Q.; Feng, Y.; Yang, B.; Lee, S.M.-Y. Cnidarian peptide neurotoxins: A new source of various ion channel modulators or blockers against central nervous systems disease. Drug Discov. Today 2019, 24, 189–197. [Google Scholar] [CrossRef] [PubMed]
- Honma, T.; Shiomi, K. Peptide toxins in sea anemones: Structural and functional aspects. Mar. Biotechnol. 2006, 8, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Cuypers, E.; Yanagihara, A.; Karlsson, E.; Tytgat, J. Jellyfish and other cnidarian envenomations cause pain by affecting TRPV1 channels. Febs Lett. 2006, 580, 5728–5732. [Google Scholar] [CrossRef]
- Levine, J.D.; Alessandri-Haber, N. TRP channels: Targets for the relief of pain. Biochim. Biophys. Acta (BBA) Mol. Basis Dis. 2007, 1772, 989–1003. [Google Scholar] [CrossRef]
- Fernandes, E.S.; Fernandes, M.A.; Keeble, J.E. The functions of TRPA1 and TRPV1: Moving away from sensory nerves. Br. J. Pharmacol. 2012, 166, 510–521. [Google Scholar] [CrossRef]
- Tysoe, C.; Withers, S.G. Structural dissection of Helianthamide reveals the basis of Its potent inhibition of human pancreaticalpha-amylase. Biochemistry 2018, 57, 5384–5387. [Google Scholar] [CrossRef] [PubMed]
- Vogg, M.C.; Beccari, L.; Iglesias Ollé, L.; Rampon, C.; Vriz, S.; Perruchoud, C.; Wenger, Y.; Galliot, B. An evolutionarily-conserved Wnt3/β-catenin/Sp5 feedback loop restricts head organizer activity in Hydra. Nat. Commun. 2019, 10, 312–315. [Google Scholar] [CrossRef] [PubMed]
- Chapman, J.A.; Kirkness, E.F.; Simakov, O.; Hampson, S.E.; Mitros, T.; Weinmaier, T.; Rattei, T.; Balasubramanian, P.G.; Borman, J.; Busam, D.; et al. The dynamic genome of Hydra. Nature 2010, 464, 592–596. [Google Scholar] [CrossRef] [PubMed]
- Ohdera, A.; Ames, C.L.; Dikow, R.B.; Kayal, E.; Chiodin, M.; Busby, B.; La, S.; Pirro, S.; Collins, A.G.; Medina, M.N.; et al. Box, stalked, and upside-down? Draft genomes from diverse jellyfish (Cnidaria, Acraspeda) lineages: Alatina alata (Cubozoa), Calvadosia cruxmelitensis (Staurozoa), and Cassiopea xamachana (Scyphozoa). Gigascience 2019, 8, 1–15. [Google Scholar] [CrossRef]
- Wilding, C.S.; Fletcher, N.; Smith, E.K.; Prentis, P.; Weedall, G.D.; Stewart, Z. The genome of the sea anemone Actinia equina (L.): Meiotic toolkit genes and the question of sexual reproduction. Mar. Genom. 2020, 53, 100753. [Google Scholar] [CrossRef]
- Surm, J.M.; Stewart, Z.K.; Papanicolaou, A.; Pavasovic, A.; Prentis, P.J. The draft genome of Actinia tenebrosa reveals insights into toxin evolution. Ecol. Evol. 2019, 9, 11314–11328. [Google Scholar] [CrossRef]
- Shinzato, C.; Shoguchi, E.; Kawashima, T.; Hamada, M.; Hisata, K.; Tanaka, M.; Fujie, M.; Fujiwara, M.; Koyanagi, R.; Ikuta, T.; et al. Using the Acropora digitifera genome to understand coral responses to environmental change. Nature 2011, 476, 320–323. [Google Scholar] [CrossRef]
- Ying, H.; Hayward, D.C.; Cooke, I.; Wang, W.; Moya, A.; Siemering, K.R.; Sprungala, S.; Ball, E.E.; Forêt, S.; Miller, D.J. The whole-genome sequence of the coral Acropora millepora. Genome Biol. Evol. 2019, 11, 1374–1379. [Google Scholar] [CrossRef]
- Jeon, Y.; Park, S.G.; Lee, N.; Weber, J.A.; Kim, H.-S.; Hwang, S.-J.; Woo, S.; Kim, H.-M.; Bhak, Y.; Jeon, S.; et al. The draft genome of an octocoral, Dendronephthya gigantea. Genome Biol. Evol. 2019, 11, 949–953. [Google Scholar] [CrossRef]
- Baumgarten, S.; Simakov, O.; Esherick, L.Y.; Liew, Y.J.; Lehnert, E.M.; Michell, C.T.; Li, Y.; Hambleton, E.A.; Guse, A.; Oates, M.E.; et al. The genome of Aiptasia, a sea anemone model for coral symbiosis. Proc. Natl. Acad. Sci. USA 2015, 112, 11893–11898. [Google Scholar] [CrossRef]
- Helmkampf, M.; Bellinger, M.R.; Geib, S.M.; Sim, S.B.; Takabayashi, M. Draft genome of the rice coral Montipora capitata obtained fromlLinked-read sequencing. Genome Biol. Evol. 2019, 11, 2045–2054. [Google Scholar] [CrossRef] [PubMed]
- Putnam, N.H.; Srivastava, M.; Hellsten, U.; Dirks, B.; Chapman, J.; Salamov, A.; Terry, A.; Shapiro, H.; Lindquist, E.; Kapitonov, V.V.; et al. Sea anemone genome reveals ancestral eumetazoan gene repertoire and genomic organization. Science 2007, 317, 86–94. [Google Scholar] [CrossRef] [PubMed]
- Prada, C.; Kamel, B.; Budd, A.; Woodley, C.; Schmutz, J.; Grimwood, J.; Iglesias-Prieto, R.; Pandolfi, J.; Levitan, D.; Johnson, K.; et al. Empty niches after extinctions increase population sizes of modern corals. Curr. Biol. 2016, 26. [Google Scholar] [CrossRef] [PubMed]
- Cunning, R.; Bay, R.A.; Gillette, P.; Baker, A.C.; Traylor-Knowles, N. Comparative analysis of the Pocillopora damicornis genome highlights role of immune system in coral evolution. Sci. Rep. 2018, 8, 16134. [Google Scholar] [CrossRef]
- Voolstra, C.R.; Li, Y.; Liew, Y.J.; Baumgarten, S.; Zoccola, D.; Flot, J.-F.; Tambutté, S.; Allemand, D.; Aranda, M. Comparative analysis of the genomes of Stylophora pistillata and Acropora digitifera provides evidence for extensive differences between species of corals. Sci. Rep. 2017, 7, 17583. [Google Scholar] [CrossRef]
- Yahalomi, D.; Haddas-Sasson, M.; Rubinstein, N.D.; Feldstein, T.; Diamant, A.; Huchon, D. The ìmultipartite mitochondrial genome of Enteromyxum leei (Myxozoa): Eight fast-evolving megacircles. Mol. Biol. Evol. 2017, 34, 1551–1556. [Google Scholar] [CrossRef]
- Chang, E.S.; Moran, N.; Nimrod, D.R.; Arik, D.; Hervé, P.; Dorothée, H.; Paulyn, C. Genomic insights into the evolutionary origin of Myxozoa within Cnidaria. Proc. Natl. Acad. Sci. USA 2015, 112, 14912–14917. [Google Scholar] [CrossRef]
- Yang, Y.; Xiong, J.; Zhou, Z.; Huo, F.; Miao, W.; Ran, C.; Liu, Y.; Zhang, J.; Feng, J.; Wang, M.; et al. The genome of the myxosporean Thelohanellus kitauei shows adaptations to nutrient acquisition within its fish host. Genome Biol. Evol. 2014, 6, 3182–3198. [Google Scholar] [CrossRef]
- Huang, C.; Morlighem, J.-Ã.t.R.L.; Zhou, H.; Lima, Ė.P.; Gomes, P.B.; Cai, J.; Lou, I.; Pérez, C.D.; Lee, S.M.; Ràdis-Baptista, G. The transcriptome of the zoanthid Protopalythoa variabilis (Cnidaria, Anthozoa) predicts a basal repertoire of toxin-like and venom-auxiliary polypeptides. Genome Biol. Evol. 2016, 8, 3045–3064. [Google Scholar] [CrossRef]
- Ames, C.L.; Ryan, J.F.; Bely, A.E.; Cartwright, P.; Collins, A.G. A new transcriptome and transcriptome profiling of adult and larval tissue in the box jellyfish Alatina alata: An emerging model for studying venom, vision and sex. BMC. Genom. 2016, 17, 650. [Google Scholar]
- Koch, T.; Grimmelikhuijzen, C. Global neuropeptide annotations from the genomes and transcriptomes of cubozoa, scyphozoa, staurozoa (Cnidaria: Medusozoa), and Octocorallia (Cnidaria: Anthozoa). Front. Endocrinol. 2019, 10, 831. [Google Scholar] [CrossRef] [PubMed]
- Ponce, D.; Brinkman, D.L.; Potriquet, J.; Mulvenna, J. Tentacle transcriptome and venom proteome of the Pacific sea nettle, Chrysaora fuscescens (Cnidaria: Scyphozoa). Toxins 2016, 8, 102. [Google Scholar] [CrossRef] [PubMed]
- Brinkman, D.L.; Aziz, A.; Loukas, A.; Potriquet, J.; Seymour, J.; Mulvenna, J. Venom proteome of the box jellyfish Chironex fleckeri. PLoS ONE 2012, 7, e47866. [Google Scholar] [CrossRef] [PubMed]
- Frazão, B.; Campos, A.; Osãrio, H.; Thomas, B.; Leandro, S.; Teixeira, A.; Vasconcelos, V.; Antunes, A. Analysis of Pelagia noctiluca proteome reveals a red fluorescent protein, a zinc metalloproteinase and a peroxiredoxin. Protein J. 2017, 36, 77–97. [Google Scholar] [CrossRef] [PubMed]
- Rachamim, T.; Morgenstern, D.; Aharonovich, D.; Brekhman, V.; Lotan, T.; Sher, D. The dynamically evolving nematocyst content of an anthozoan, a scyphozoan, and a hydrozoan. Mol. Biol. Evol. 2015, 32, 740–753. [Google Scholar] [CrossRef] [PubMed]
- Jaimes-Becerra, A.; Chung, R.; Morandini, A.C.; Weston, A.J.; Padilla, G.; Gacesa, R.; Ward, M.; Long, P.F.; Marques, A.C. Comparative proteomics reveals recruitment patterns of some protein families in the venoms of Cnidaria. Toxicon 2017, 137, 19–26. [Google Scholar] [CrossRef]
- Lohr, K.E.; Khattri, R.B.; Guingab-Cagmat, J.; Camp, E.F.; Merritt, M.E.; Garrett, T.J.; Patterson, J.T. Metabolomic profiles differ among unique genotypes of a threatened Caribbean coral. Sci. Rep. 2019, 9, 6067. [Google Scholar] [CrossRef]
- Turk, T.; Kem, W.R. The phylum Cnidaria and investigations of its toxins and venoms until 1990. Toxicon 2009, 54, 1031–1037. [Google Scholar] [CrossRef]
- Prentis, P.J.; Pavasovic, A.; Norton, R.S. Sea anemones: Quiet achievers in the field of peptide toxins. Toxins 2018, 10, 36. [Google Scholar] [CrossRef]
- Kawabata, T.; Lindsay, D.; Kitamura, M.; Konishi, S.; Nishikawa, J.; Nishida, S.; Kamio, M.; Nagai, H. Evaluation of the bioactivities of water-soluble extracts from twelve deep-sea jellyfish species. Fish. Sci. 2013, 79, 487–494. [Google Scholar] [CrossRef]
- Moritz, M.I.G.; Marostica, L.L.; Bianco, E.M.; Almeida, M.T.R.; Carraro, J.L.; Cabrera, G.M.; Palermo, J.A.; Simðes, C.M.O.; Schenkel, E.P. Polyoxygenated steroids from the octocoral Leptogorgia punicea and in vitro evaluation of their cytotoxic activity. Mar. Drugs 2014, 12, 5864–5880. [Google Scholar] [CrossRef]
- Ayed, Y. Evaluation of anti-proliferative and anti-inflammatory activities of Pelagia noctiluca venom in Lipopolysaccharide/Interferon-g stimulated RAW264.7 macrophages. Biomed. Pharmacother. 2016, 84, 1986–1991. [Google Scholar] [CrossRef] [PubMed]
- Silva, T.; de Andrade, P.; Paiva-Martins, F.; Valentäo, P.; Pereira, D. In vitro anti-Inflammatory and cytotoxic effects of aqueous extracts from the edible sea anemones Anemonia sulcata and Actinia equina. Int. J. Mol. Sci. 2017, 18, 653. [Google Scholar] [CrossRef] [PubMed]
- Mariottini, G.L.; Grice, I.D. Antimicrobials from Cnidarians. A new perspective for anti-infective therapy? Mar. Drugs 2016, 14, 48. [Google Scholar] [CrossRef] [PubMed]
- Coppola, D.; Oliviero, M.; Vitale, G.A.; Lauritano, C.; D’Ambra, I.; Iannace, S.; de Pascale, D. Marine collagen from alternative and sustainable sources: Extraction, processing and applications. Mar. Drugs 2020, 18, 214. [Google Scholar] [CrossRef]
- Lauritano, C.; Ferrante, M.I.; Rogato, A. Marine natural products from microalgae: An -omics overview. Mar. Drugs 2019, 17, 269. [Google Scholar] [CrossRef]
| Class | Life Stage | Cnida Type | Reference |
|---|---|---|---|
| Hydrozoa | Alternance polyp/medusa | Nematocyst | [3,4] |
| Scyphozoa | Alternance polyp/medusa | Nematocyst | [3,4] |
| Cubozoa | Alternance polyp/medusa | Nematocyst | [3,4] |
| Anthozoa | Polyp only | Nematocysts | [3,4] |
| Spirocyst (Zoantharia only) | [4] | ||
| Ptychocytes (Ceriantharia only) | [5] | ||
| Staurozoa | Alternance polyp/medusa | Nematocysts | [6] |
| Species | Compound | Molecular Mass (kDa) | Biological Activity | References |
|---|---|---|---|---|
| Hydrozoa | ||||
| Hydra magnipapillata | CqTX-A | ~40 | Cardiovascular, hemolytic | [18] |
| HALT-1 | N/A | Hemolytic, cytolysis | [19] | |
| HALT1–HALT7 | N/A | Cytolytic | [20] | |
| H. viridissima | Hydralysin | 27 | Neurotoxic, cytolytic, paralytic | [21] |
| Millepora platyphylla | Milleporin-1 | 30–34 | Cytolytic, hemolytic | [22] |
| Olindias sambaquiensis | Oshem1 | 3.013 | Cytolytic, hemolytic, myonecrotic, and cytotoxic | [23] |
| Oshem2 | 3.376 | Cytolytic | [23] | |
| Metalloproteinases | N/A | Cytolytic, neurotoxic | [24] | |
| Physalia physalis | Phospholipase A2 | N/A | N/A | [25] |
| Phospholipase B | N/A | N/A | [25] | |
| Collagenase | 25 | Cytotoxic, hemolytic | [26] | |
| Elastases | N/A | Musculotoxic, cytolytic, hemolytic | [27] | |
| PpV19.3 | 4.72 | Neurotoxic, cardiotoxic | [28] | |
| PpV9.4 | 0.55 | Hemolytic | [28] | |
| P3 | 85 | Neurotoxic | [29] | |
| P1 | 220 | Neurotoxic | [30] | |
| Physalitoxin | 220 | Hemolytic | [31] | |
| DNase | 75 | Cytolytic | [32] | |
| Histamine | N/A | N/A | [33] | |
| Tubularia larynx | Phospholipase A2 | N/A | Cytolytic, hemolytic | [15] |
| Scyphozoa | ||||
| Aurelia aurita | Phospholipase A2 | N/A | Cytolytic | [15,34] |
| Proteolytic enzymes | N/A | Hemolytic, neurotoxic, myotoxic, local skin irritation | [35] | |
| Tetramine and unidentified protein | N/A | Dermotoxic, temporary paralysis, edema | [36] | |
| Metalloproteinases | N/A | Gelatinolytic, caseinolytic, fibrinolytic | [37] | |
| Aurelin | 4.30 | Antimicrobial, neurotoxic (voltage-gated potassium channel inhibitor) | [38] | |
| Cassiopea andromeda | Phospholipase A2 | N/A | Hemolytic, dermonecrotic, local skin irritation | [34] |
| C. xamancha | Phospholipase A2 | N/A | Hemolytic, dermonecrotic, local skin irritation | [34] |
| Chrysaora hysoscella | Cationic protein | N/A | Dermotoxic, cytotoxic | [39] |
| C. quinquecirrha | DNase | 110 | Dermonecrotic, cytotoxic | [26] |
| Acid protease | 120–150 | N/A | [26] | |
| Metallopeptidase | 100 | N/A | [26] | |
| Collagenase | N/A | N/A | [26] | |
| Cyanea capillata | Basic protein(s) | 70 | Cardiotoxic, dermonecrotic, musculotoxic | [26,40] |
| CcTX-1 | 31.173 | Cytotoxic | [41] | |
| CcNT | 8.22 | Neurotoxic | [42] | |
| Phospholipase A2 | N/A | Cytolytic, cytotoxic, hemolytic | [15,43] | |
| C. lamarckii | ClGP-1 | 27 | Cytotoxic | [43] |
| Phospholipase A2 | N/A | Cytolytic, cytotoxic, hemolytic | [43] | |
| C. nozakii | Metalloproteinases | N/A | Gelatinolytic, caseinolytic, fibrinolytic | [37] |
| Nemopilema nomurai | Metalloproteinases | 28–36 | Gelatinolytic, caseinolytic, fibrinolytic | [37] |
| 20–40/10–15 | Cytotoxic, hemolytic | [44] | ||
| Pelagia noctiluca | Proteinaceous macromolecules | 44–66 | Hemolytic, cytotoxic, dermonecrotic, hemolytic, local tissue damage | [45,46,47,48,49] |
| Phyllorhiza punctata | Phospholipase A2 | N/A | Neurotoxic | [50] |
| Rhizostoma pulmo | Rhizoprotease | 95 | Proteolytic, hemolytic | [51] |
| Rhizolysin | 260 | Hemolytic | [52] | |
| N/A | Cytotoxic, hemolytic | [53] | ||
| Rhopilema esculentum | Metalloproteinases | N/A | Gelatinolytic, caseinolytic, fibrinolytic | [37] |
| Hyaluronidase | 55–95 | Degradation of extracellular matrix components | [37] | |
| N/A | Proteolytic, cytotoxic, hemolytic | [54,55] | ||
| R. nomadica | Phospholipase A2 | N/A | Hemolytic | [56] |
| Serine protease | N/A | Local skin damage | [57] | |
| Rhopilema spp. | Phospholipase A2 | N/A | Hemolytic | [58] |
| Stomolophus meleagris | SmP90 | 90 | Radical scavenging | [59] |
| Phospholipase A2 | N/A | Cytotoxic, cytolytic, hemolytic, local tissue damage | [59] | |
| 218 toxins including C-lectin and metalloprotease | N/A | Voltage-gated potassium channel inhibitor | [59] | |
| Cubozoa | ||||
| Alatina moseri | CaTX-A | 43 | Hemolytic | [18] |
| Akatina alata | CAH1 | 42 | Hemolytic | [60] |
| CaTX-A | 43 | Hemolytic | [61] | |
| CaTX-B | 45 | Hemolytic | [61] | |
| C. marsupialis | Haemolysin | 102–107 | Hemolytic | [62] |
| CmHl1 | 139 | Cytolytic | [63] | |
| CmHl5 | 220 | Cytolytic | [63] | |
| CmH17 | 139 | Cytolytic | [63] | |
| CmNt | 120 | Neurotoxic | [63] | |
| C. rastonii | Phospholipase A2 | N/A | Cytolytic | [15] |
| CrTX-I | N/A | Hemolytic | [64] | |
| CrTX-II | N/A | Hemolytic | [64] | |
| CrTX-III | N/A | Hemolytic | [64] | |
| CrTX-A | 43 | Cutaneous inflammation of human skin | [61] | |
| CrTX-B | 46 | N/A | [61] | |
| Carukia barnesi | Phospholipase A2 | N/A | Cytolytic, hemolytic | [15] |
| CbTX-I | 21.67 | Neurotoxic | [65] | |
| CbTX-II | 18.16 | Neurotoxic | [65] | |
| Chironex fleckeri | Phospholipase A2 | N/A | Cytolytic, hemolytic | [15] |
| Metalloproteinases | 17–130 | N/A | [66] | |
| CfTX-1 | 43 | Cardiotoxic, cytotoxic, dermonecrotic, lethal | [67] | |
| CfTX-2 | 45 | Cardiotoxic, cytotoxic, dermonecrotic, lethal | [67] | |
| CfTX-A | 40 | Hemolytic | [18] | |
| CfTX-B | 42 | Hemolytic | [18] | |
| CfTX-Bt | 31.29 | N/A | [18] | |
| Chiropsalmus quadrigatus | CqTX-A | 44 | Hemolytic, neurotoxic, myotoxic | [58,68,69] |
| Malo kingi | MkTX-A; MkTX-B | 43–45 | Dermonecrotic, inflammatory | [65] |
| 43–45 | Dermonecrotic, inflammatory | [49] | ||
| Anthozoa (Hexacorallia) | ||||
| Acropora spp. | Phospholipase A2 | N/A | Catalytic | [15] |
| Actinia australis | Phospholipase A2 | N/A | Catalytic | [15] |
| Actinia equina | AeI | N/A | Lethal activity to crabs, Type I sodium channel toxin | [70] |
| AEPI-I, II, III and IV | 6.2–7 | Kunitz-type toxins | [71] | |
| Equinatoxin-I, II and III | 19 | Cytolytic and hemolytic | [72] | |
| Equinatoxin-IV | N/A | Hemolytic | [73] | |
| Equinatoxin-V | N/A | N/A | [74] | |
| Acrorhagin I and II | N/A | Lethal activity to crabs | [75] | |
| Acrorhagin Ia and IIa | N/A | N/A | [75] | |
| Actinia fragacea | Fragaceatoxin C | 20 | Lytic | [76] |
| Actinia tenebrosa | Tenebrosin-A, B and C | 19–20 | Hemolytic | [77] |
| Actinia villosa | Avt-I | 19 | Hemolytic | [78] |
| Avt-II | N/A | Cytolysis | [78,79] | |
| AvTX-60A | 60 | Fatal toxicity to mice | [80] | |
| Avt120 | 120 | Lethal activity to mice, cytotoxic | [81] | |
| Adamsia palliata | AcPLA2 | 13.5 | Phospholipase A2 catalytic activity | [82] |
| Adamsia carciniopados | Phospholipase A2 | N/A | Phospholipase A2 catalytic activity | [15] |
| Anemonia erythraea | AETX-I | 5 | Crab lethality, Type I sodium channel toxin | [83] |
| AETX-K | ~4 | Type I potassium channel toxin | [84] | |
| AETX-II | 6.5 | Crab lethality | [83] | |
| AETX-III | 6.6 | Crab lethality | [83] | |
| Anemonia sulcata | Toxin I | N/A | Type I sodium channel toxin | [85] |
| ATX-II | 5 | Cardiotoxic action, Type I sodium channel toxin | [62,85] | |
| Toxin III | 2.7 | N/A | [86] | |
| ATX-V | N/A | Type I sodium channel toxin | [87] | |
| SA5 II | N/A | Kunitz-type proteinase toxin | [88] | |
| Kalicludin 1, 2, 3 | ~6.7 | Type II potassium channel toxin, Kunitz inhibitors | [89] | |
| BDS-I and BDS-II | 4.7 | Inactivating Kv3.4 channel | [78] | |
| Kaliseptine | 3.8 | Potassium channel toxin | [89] | |
| Anthopleura asiatica | Bandaporin | 20 | Hemolytic | [90] |
| Anthopleura elegantissima | Anthopleurin-C | N/A | Cardiotoxic to rats, Type I sodium channel toxin | [91] |
| APE 1-1, 1-2, 2-1, 2-2, 3, 4 and 5-3 | 5 | Crab paralysis, Type I sodium channel toxin | [92] | |
| APET x2 | 4.6 | ASIC3 inhibition (acting on channel external side) | [93] | |
| Anthopleura fuscoviridis | AFT-I and II | N/A | Type I sodium channel toxins | [94] |
| Anthopleura xanthogrammica | Anthopleurin-A and B | N/A | Cardiotoxic to rats, Type I sodium channel toxin | [91] |
| Toxin PCR1-2, 2-1, 2-5, 2-10, 3-6, 3-7, 3-3,4 | N/A | Type I sodium channel toxin | [95] | |
| AXPI-I and II | N/A | Kunitz-type inhibitor | [96] | |
| Anthopleura spp. | Hk2a, Hk7a, Hk8a, Hk16a | ~5 | Rat heart stimulation, Type I sodium channel toxins | [97] |
| Bolecera tuedia | Phospolipase A2 | N/A | Phospholipase A2 catalytic activity | [15] |
| Bunodosoma caissarum | Bc-I, II and III | N/A | Toxic on crustacean nerves, Type I sodium channel toxin | [98] |
| Bc-IV | ~5 | Weak-paralyzing action in swimming crabs | [99] | |
| BcPLA(2)1 | ~5 | Renal function and induced insulin secretion in conditions of high glucose concentration | [100] | |
| BcsTx3 | 5.71 | Potassium channel inhibition | [101] | |
| Bunodosoma granulifera | Bg II and III | N/A | Neurotoxic on mice, Type I sodium channel toxins | [102] |
| Bg toxin | ~4 | Type I potassium channel toxin | [103] | |
| Granulitoxin (GRX) | ~5 | Severe neurotoxic effects such as circular movements, aggressive behavior, dyspnea, tonic-clonic convulsion and death in mice | [104] | |
| Bunodosoma cangicum | Cangitoxin | 5 | Type I sodium channel toxin | [105] |
| Cangitoxin II and III (CGTX-II and CGTX-III) | ~5 | Type I sodium channel toxin | [106] | |
| Bcg 25.96, 28.19, 30.24 | ~4 | Type I potassium channel toxin, Neurotoxic to crabs | [105] | |
| Bcg 31.16, 28.78, 25.52, 29.21 | 4–5 | May target different types of ion channels, Neurotoxic to crabs | [105] | |
| Bcg 21.75, 23.41, 21.00 | 3 | Possible new class of potassium channel blockers, Neurotoxic to crabs | [105] | |
| Calliactis parasitica | Calitoxin 1 | ~5 | Increased transmitter release, causing repetitive firing of the axons in neuromuscular preparation of crustaceans | [107] |
| Calitoxin 2 | N/A | N/A | [108] | |
| Condylactis gigantea | CgNa | ~5 | Type I sodium channel toxin | [109] |
| Condylactis passiflora | CpI, II and III | N/A | Lethal activity against crabs, Type I sodium channel toxin | [110] |
| Cryptodendrum adhaesivum | Ca I | N/A | Type II sodium channel toxin | [111] |
| Dendronephthya spp. | Phospholipase A2 | N/A | Phospholipase A2 catalytic activity | [15] |
| Halcurias carlgreni | Halcurin | ~5 | N/A | [112] |
| Heteractis crispa (=Radianthus crispus = R. macrodactyla) | Neurotoxin I | N/A | Type II sodium channel toxin | [113] |
| Neurotoxin II | N/A | Type II sodium channel toxin | [114] | |
| Neurotoxin III | N/A | Type II sodium channel toxin | [115] | |
| Neurotoxin IV and V | N/A | Type II sodium channel toxins | [116] | |
| Rc I | N/A | Lethal against crabs, Type I sodium channel toxin | [117] | |
| Kunitz-type Trypsin inhibitor IV | N/A | Kunitz-type inhibitor | [118] | |
| Actinoporin RTX-A | N/A | Hemolytic | [119] | |
| Actinoporin RTX-S II | 19 | Hemolytic | [120] | |
| HCIQ2c1 | 6 | Neuroprotective, Kunitz-type inhibitor | [121] | |
| Hcr 1b-1 | 4.5 | ASIC3 channel inhibitor | [122] | |
| Hcr1b-2, Hcr1b-3, Hcr1b-4 | 5 | Neurotoxic (ASIC1 channel inhibitors) | [123] | |
| HCRG1, HCRG2 | 6 | Anti-inflammatory, Kunitz-type inhibitors | [124] | |
| InhVJ | 6 | Kunitz-type inhibitor | [125] | |
| HGRC21 | 6 | Analgesic, inhibitor of TRPV1, Kunitz-type inhibitor | [126] | |
| APHC1, APHC2, APHC3 | 6 | Analgesic (TRPV1 modulation) | [127,128,129] | |
| HCGS 1.10 | 6 | Analgesic | [130] | |
| HCGS 1.20 | 6 | Anti-inflammatory | [131] | |
| rHCGS1.19 and rHCGS1.36 | 6 | Antihistamine, Kunitz-type serine protease inhibitors | [132] | |
| Heteractis magnifica (=Radianthus magnifica = R. paumotensis = R. ritteri) | Magnificalysins I and II | ~19 | Hemolytic and lethal activity on mice | [133] |
| HMGS1 | 6 | Kunitz-type inhibitor | [134] | |
| HMgIII | Hemolytic and cytolytic | [135] | ||
| HmK | ~4 | Potassium channel inhibitor | [136] | |
| RpI, RpII, RpIII, and RpIV | ~5 | Toxic in mice and crabs, Type II sodium channel toxins | [137] | |
| HMIQ3c1 | N/A | Neuroprotective (Kunitz-type inhibitor) | [138] | |
| Magnificamide | 4.7 | Alpha-amylase inhibitor | [139] | |
| Heterodactyla hemprichi | δ-TLTX-Hh1a and δ-TLTX-Hh1c | N/A | Lethal on crabs, Type II sodium channel toxin | [111] |
| Metridium senile | Metridin | 3.97 | Hemolytic | [140] |
| Phospholipase A2 | N/A | Phospholipase A2 catalytic activity | [15] | |
| Ms 9a-1 | 3.6 | Analgesic by potentiating TRPA1 | [141] | |
| Nematostella vectensis | Nv1116.25.1, 116.27.1, 116.28.1, 116.37.1, 116.39.1, 116.40.1, 116.41.1, 116.45.1 | N/A | Type II sodium channel toxins | [142] |
| Nv4 – Nv8 | N/A | Sodium channel inhibitors | [143] | |
| NEP1 – NEP20 | N/A | N/A | [144] | |
| Oulactis orientalis | Actinoporin Or-A and Or-G | ~18 | Cytolytic | [145] |
| NEP1 to NEP 20 | N/A | N/A | [144] | |
| Parasicyonis actinostoloides | PA-TX | N/A | N/A | [146] |
| Phyllodiscus semoni | PsTX-115 | N/A | Renal injury | [147] |
| PsTX-60A; PsTX-60B | 60 | Cytolytic, hemolytic | [148] | |
| Phymanthus crucifer | PhcrTx1 | 3.47 | ASIC inhibitor | [149] |
| Pocillopora damicornis | Phospholipase A2 | N/A | Phospholipase A2 catalytic activity | [15] |
| Sagartia rosea | Cytolysin Src-I | 19.6 | Cytolysis | [150] |
| Sarcophyton elegans | Phospolipase A2 | N/A | Phospholipase A2 catalytic activity | [15] |
| Stichodactyla helianthus | Sh1 | ~5 | Neurotoxic on crabs, Type II sodium channel inhibitor | [151] |
| Helianthamide | 4.7 | Alpha-amylase inhibitor | [152] | |
| SHPI-1 | ~6 | Kunitz-type proteinase inhibitor | [153] | |
| ShK | ~5 | Potassium channel inhibitor | [154] | |
| Sticholysin-I and II | ~19 | Hemolytic | [155] | |
| Stoichactis sp. | Phospholipase A2 | N/A | Phospholipase A2 catalytic activity | [15] |
| Stichodactyla gigantea | Gigantoxins I-III | N/A | Crab toxicity, Sodium channel inhibitors | [156] |
| Stichodactyla haddoni | SHTX I–III | ~7 | Crab-paralyzing activity, SHTX-III is a Kunitz-type proteinase inhibitor | [157] |
| SHTX IV | ~5 | Crab lethality, Type II sodium channel toxin | [157] | |
| EGF-like peptide SHTX-5 | N/A | N/A | [157] | |
| Thalassianthus aster | Ta I3,8-7,6 | N/A | Type II sodium channel toxin | [111] |
| Urticina coriacea | U1 | N/A | Hemolytic, cytotoxic | [124] |
| U2 | 3–10.5 | Analgesic (ASIC1 channel inhibitor) | [124] | |
| Urticina crassicornis | Uc-I | 30 | Cytolysis | [158] |
| Urticina eques | Ueq 12-1 | 4.79 | Antibacterial, analgesic by potentiating TRPA1 | [159] |
| Urticina grebenyi | UGR9a-1 | N/A | Analgesic, ASICS channel inhibitor | [160] |
| Urticina piscivora | Up-I | 28 | Hemolytic | [161] |
| Virgularia nidularis | Phospholipase A2 | N/A | Phospholipase A2 catalytic activity | [15] |
| Anthozoa (Octocorallia) | ||||
| Alcyonum digitatum | Phospholipase A2 | N/A | Phospholipase A2 catalytic activity | [15] |
| Paramuricea spp. | Phospholipase A2 | N/A | Phospholipase A2 catalytic activity | [15] |
| Sinularia flexibilis | Phospholipase A2 | N/A | Phospholipase A2 catalytic activity | [15] |
| Species | Accession Number | Reference |
|---|---|---|
| Hydrozoa | ||
| Clytia hemisphaerica | GCA_902728285.1 | N.A. |
| Craspedacusta sowerbii | GCA_003687565.1 | N.A. |
| Hydra oligactis | GCA_004118135.1 | [176] |
| Hydra viridissima | GCA_004118115.1 | [176] |
| Hydra vulgaris | GCA_000219015.1 | [177] |
| Hydra vulgaris | GCA_000004095.1 | [177] |
| Scyphozoa | ||
| Aurelia aurita | GCA_004194415.1 | N.A. |
| Aurelia aurita complex sp. pacific | GCA_004194395.1 | N.A. |
| Aurelia coerulea | GCA_011634815.1 | N.A. |
| Cassiopea xamachana | GCA_900291935.1 | N.A. |
| Chrysaora quinquecirrha | GCA_012295145.1 | N.A. |
| Chrysaora chesapeakei | GCA_011763395.1 | N.A. |
| Chrysaora fuscescens | GCA_009936425.1 | N.A. |
| Nemopilema nomurai | GCA_003864495.1 | N.A. |
| Rhopilema esculentum | GCA_013076305.1 | N.A. |
| Sanderia malayensis | GCA_013076295.1 | N.A. |
| Cubozoa | ||
| Alatina alata | GCA_008930755.1 | [178] |
| Alatinida sp. Z8VKAUB7J3 | GCA_010016025.1 | N.A. |
| Carybdea marsupialis auct. non (Linnaeus, 1758) | GCA_010016065.1 | N.A. |
| Morbakka virulenta | GCA_003991215.1 | N.A. |
| Anthozoa | ||
| Actinia equina | GCA_011057435.1 | [179] |
| Actinia tenebrosa | GCA_009602425.1 | [180] |
| Acropora digitifera | GCA_000222465.2 | [181] |
| Acropora millepora | GCF_004143615.1 | [182] |
| Anemonia viridis | GCA_900234385.1 | N.A. |
| Dendronephthya gigantea | GCA_004324835.1 | [183] |
| Exaiptasia diaphana | GCA_001417965.1 | [184] |
| Heteractis magnifica | GCA_011763375.1 | N.A. |
| Montipora capitata | GCA_006542545.1 | [185] |
| Nematostella vectensis | GCA_000209225.1 | [186] |
| Orbicella faveolata | GCA_002042975.1 | N.A. |
| Orbicella faveolata | GCA_001896105.1 | [187] |
| Phymanthus crucifer | GCA_009858155.1 | N.A. |
| Pocillopora damicornis | GCA_003704095.1 | [188] |
| Porites rus | GCA_900290455.1 | N.A. |
| Renilla reniformis | GCA_900177555.1 | N.A. |
| Stichodactyla mertensii | GCA_011800005.1 | N.A. |
| Stylophora pistillata | GCA_002571385.1 | [189] |
| Staurozoa | ||
| Calvadosia cruxmelitensis | GCA_900245855.1 | N.A. |
| Myxozoa | ||
| Enteromyxum leei | GCA_001455295.2 | [190] |
| Henneguya salminicola | GCA_009887335.1 | N.A. |
| Kudoa iwatai | GCA_001407235.2 | [190,191] |
| Kudoa iwatai | GCA_001407335.1 | [191] |
| Myxobolus squamalis | GCA_010108815.1 | N.A. |
| Sphaeromyxa zaharoni | GCA_001455285.1 | [191] |
| Thelohanellus kitauei | GCA_000827895.1 | [192] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
D’Ambra, I.; Lauritano, C. A Review of Toxins from Cnidaria. Mar. Drugs 2020, 18, 507. https://doi.org/10.3390/md18100507
D’Ambra I, Lauritano C. A Review of Toxins from Cnidaria. Marine Drugs. 2020; 18(10):507. https://doi.org/10.3390/md18100507
Chicago/Turabian StyleD’Ambra, Isabella, and Chiara Lauritano. 2020. "A Review of Toxins from Cnidaria" Marine Drugs 18, no. 10: 507. https://doi.org/10.3390/md18100507
APA StyleD’Ambra, I., & Lauritano, C. (2020). A Review of Toxins from Cnidaria. Marine Drugs, 18(10), 507. https://doi.org/10.3390/md18100507






