Mass Spectrometry-Based Characterization of New Spirolides from Alexandrium ostenfeldii (Dinophyceae)
Abstract
1. Introduction
2. Results and Discussion
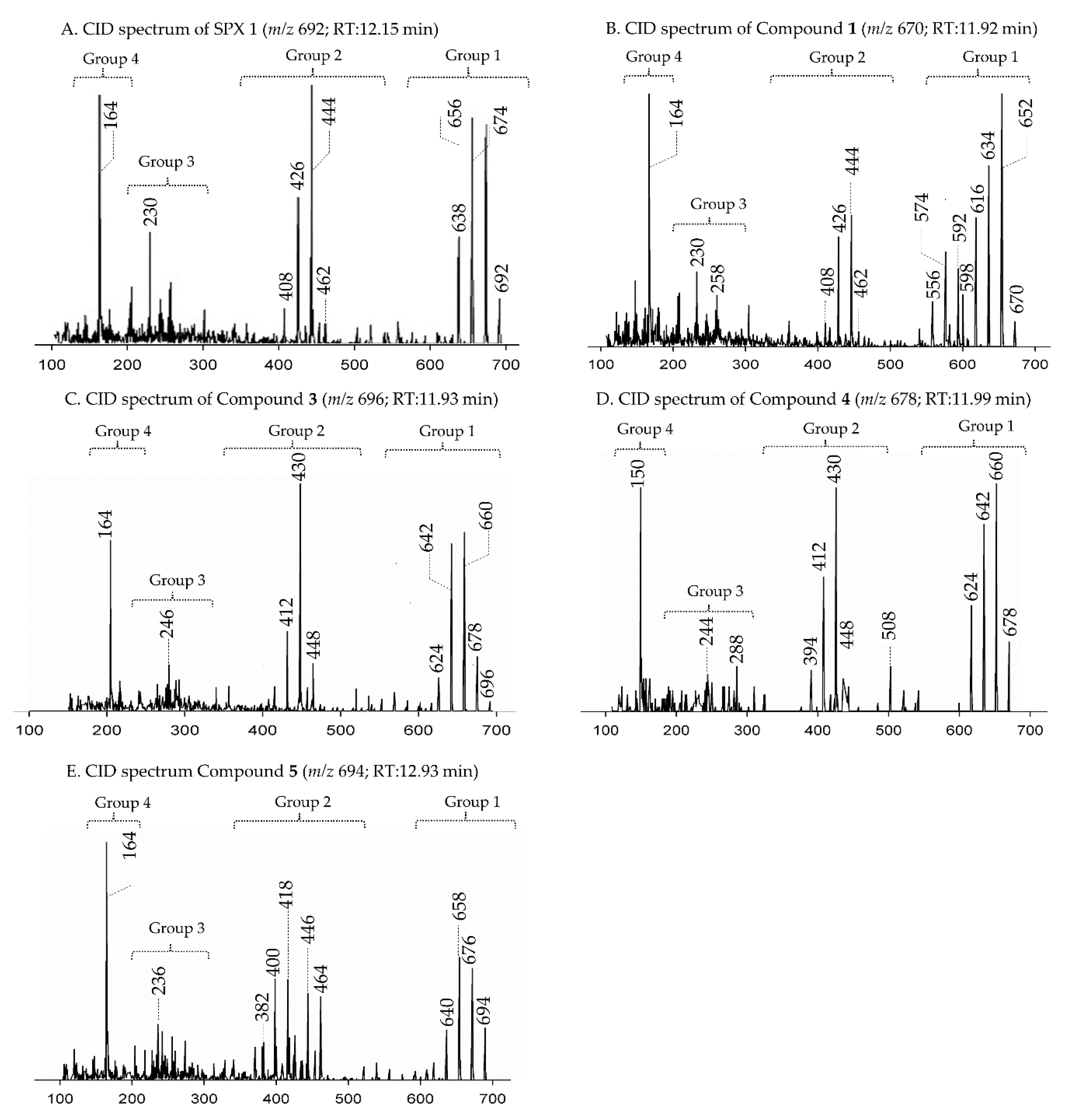
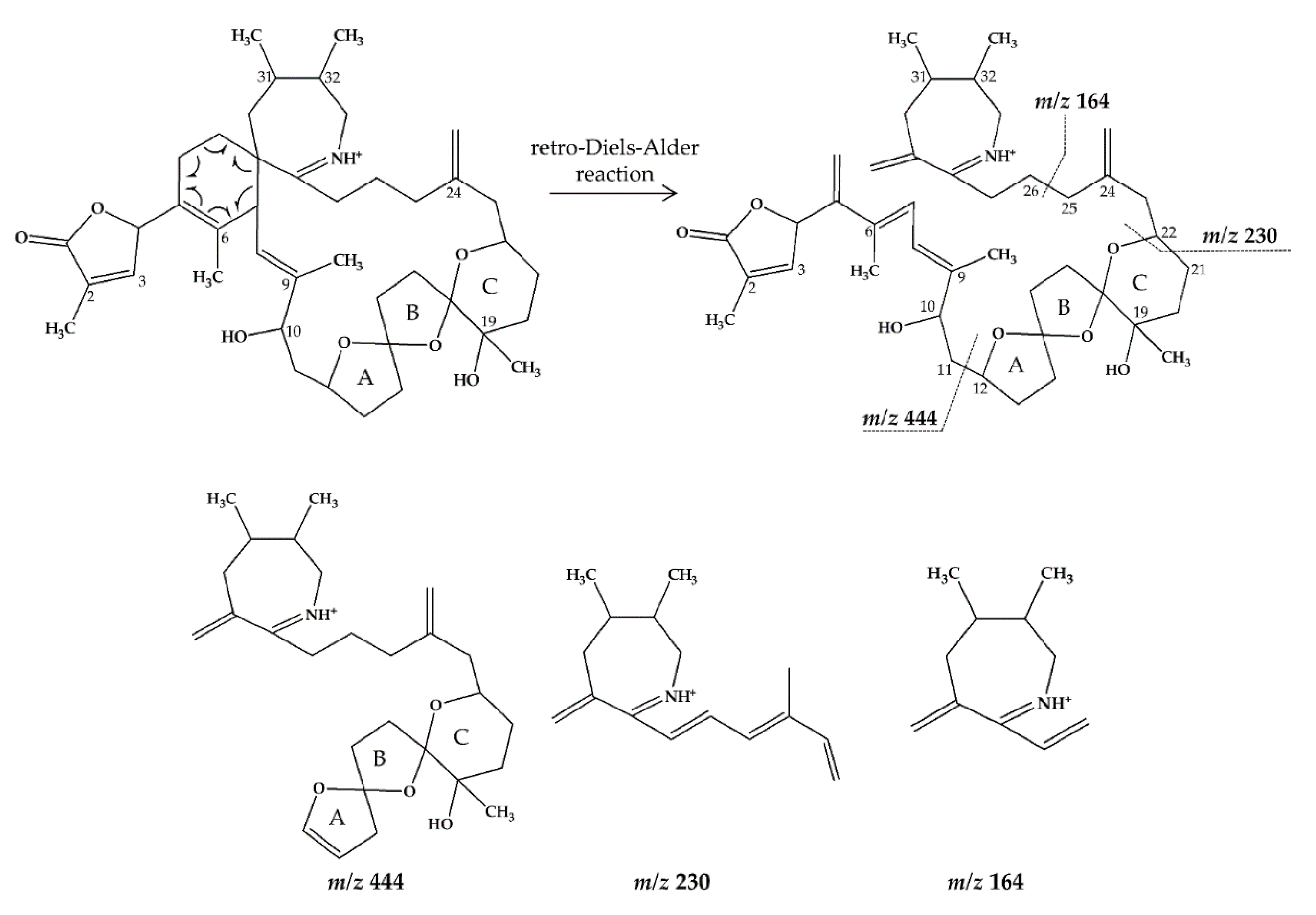
2.1. Interpretation of Collision-Induced Dissociation (CID) Spectra
2.2. Characterization of Novel Spirolides
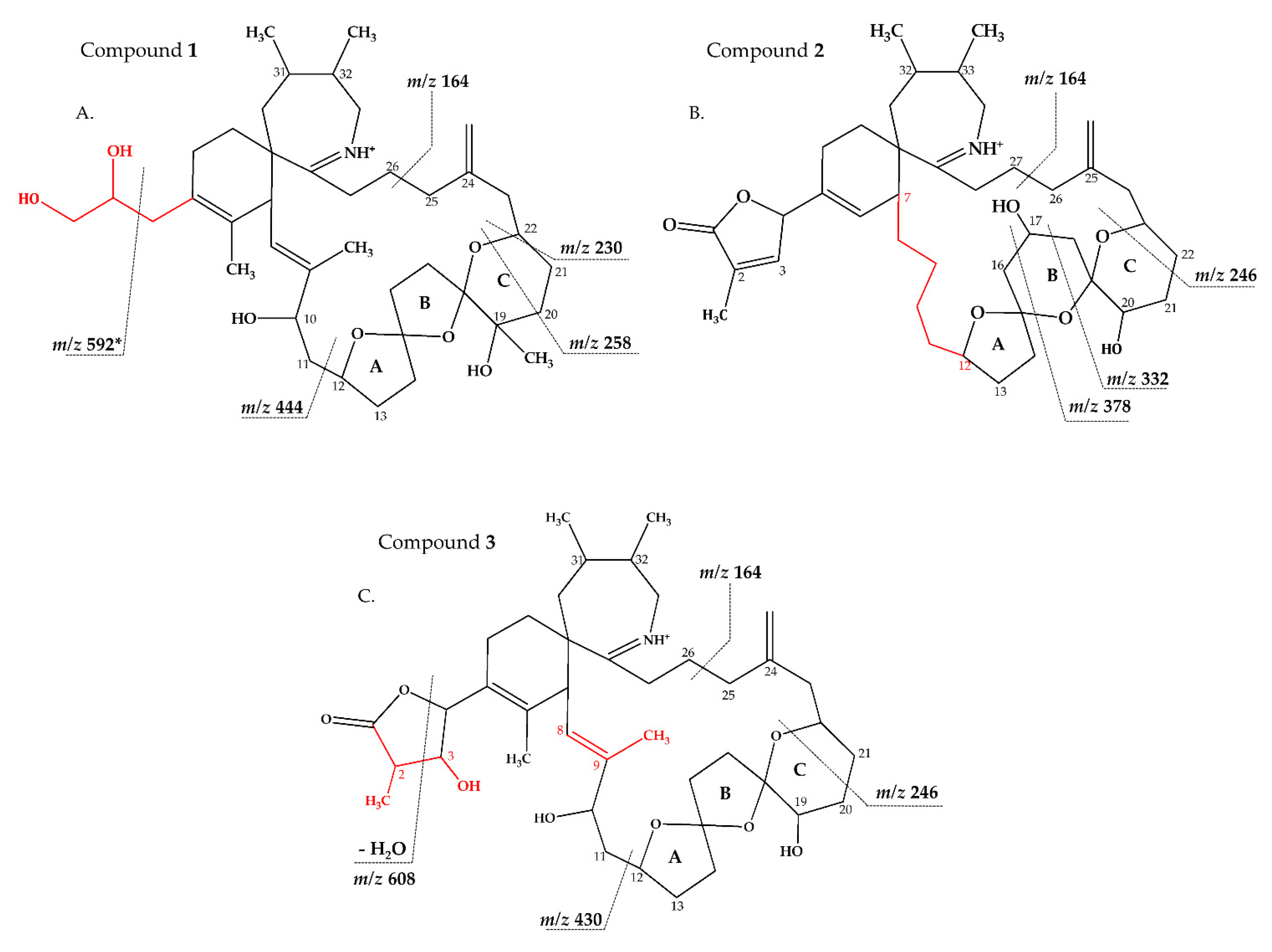
2.2.1. Compound 1
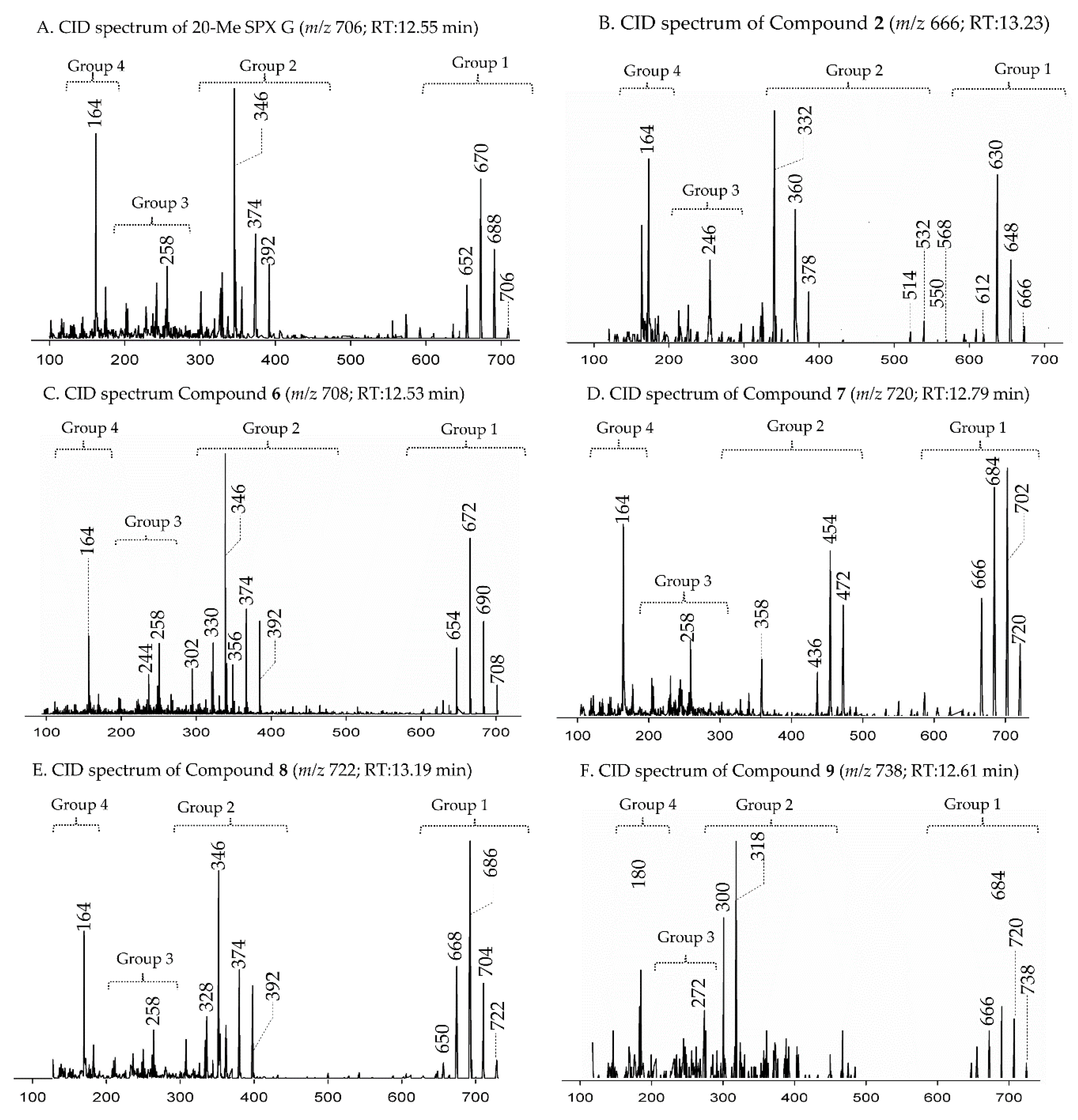
2.2.2. Compound 2–3
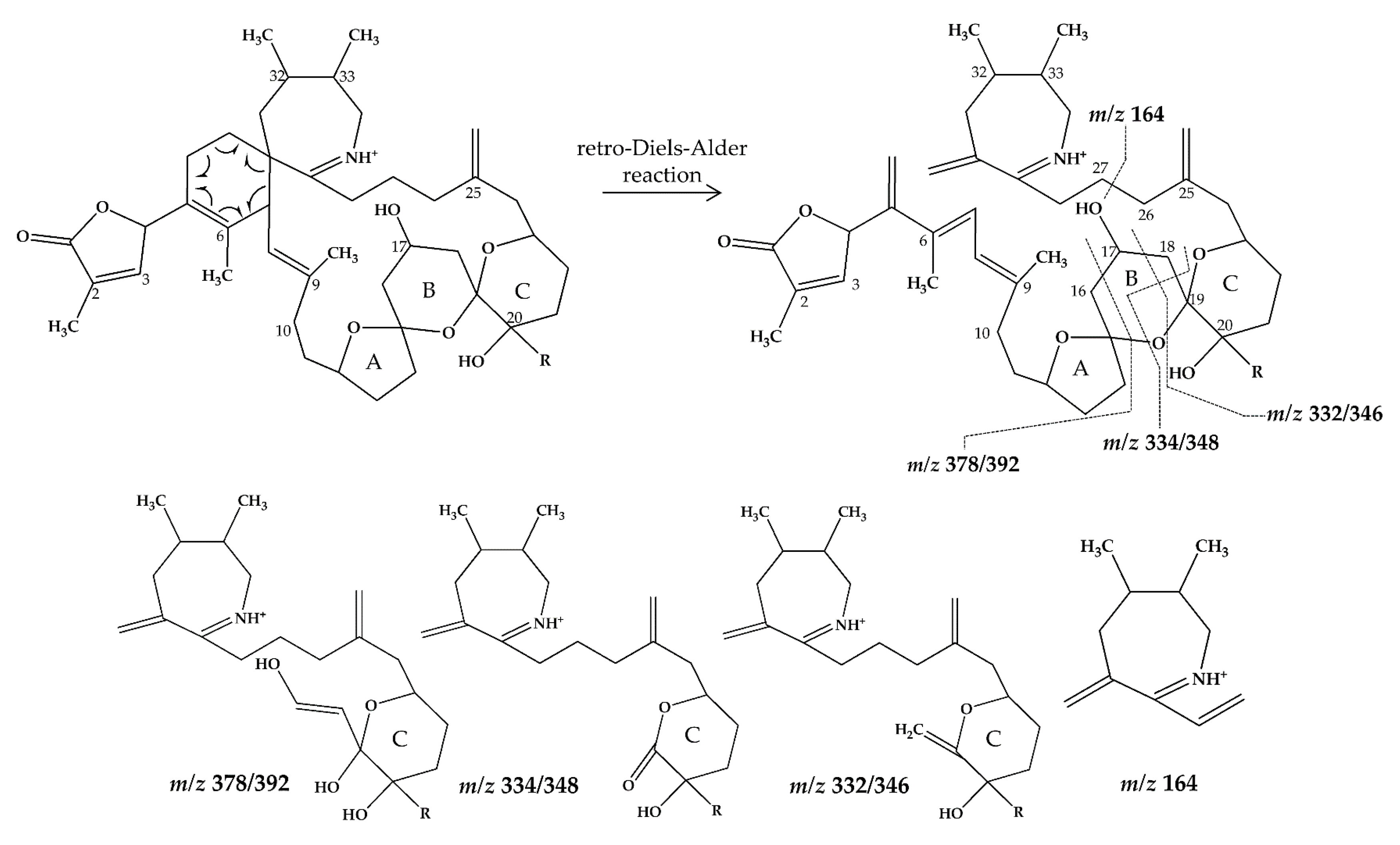
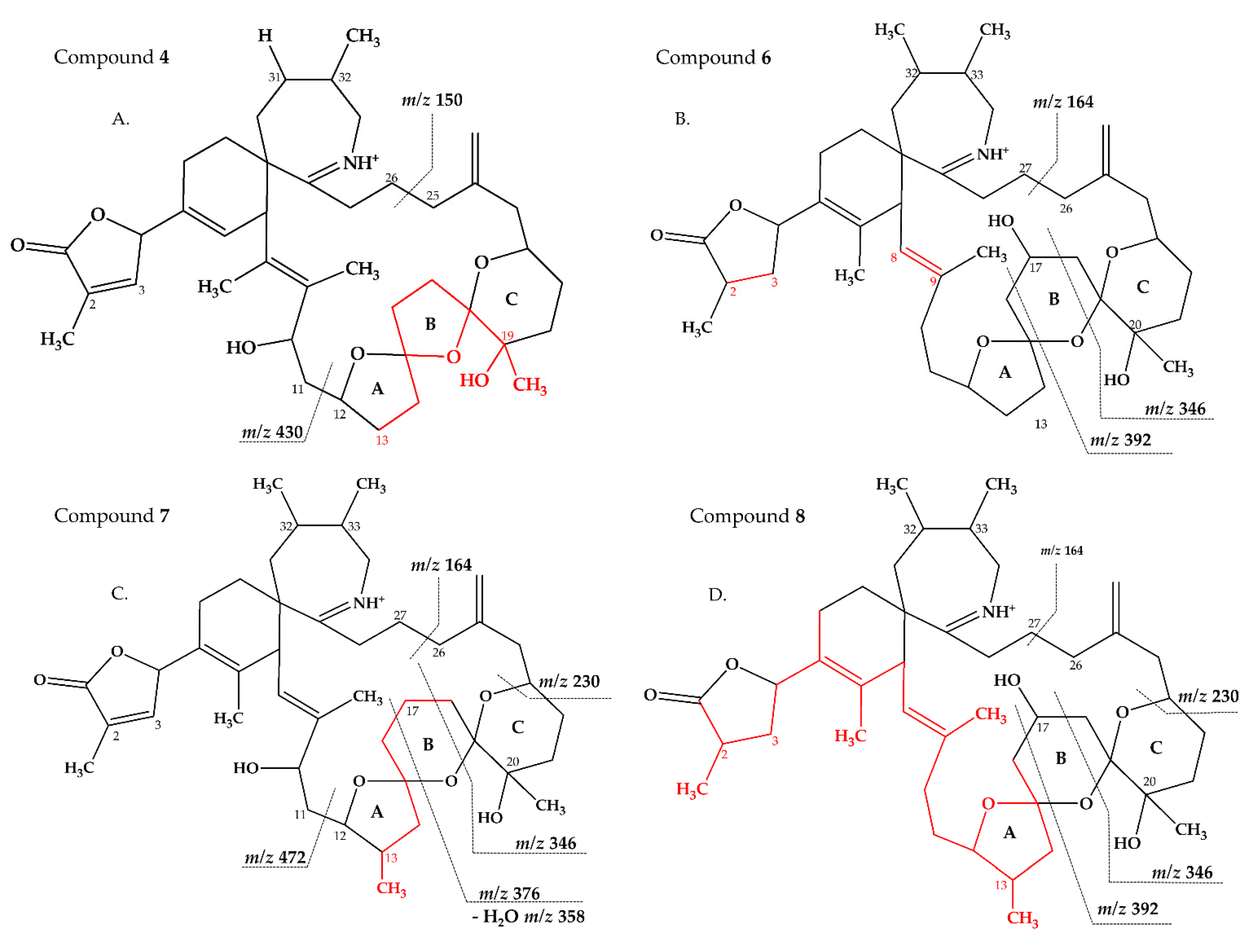
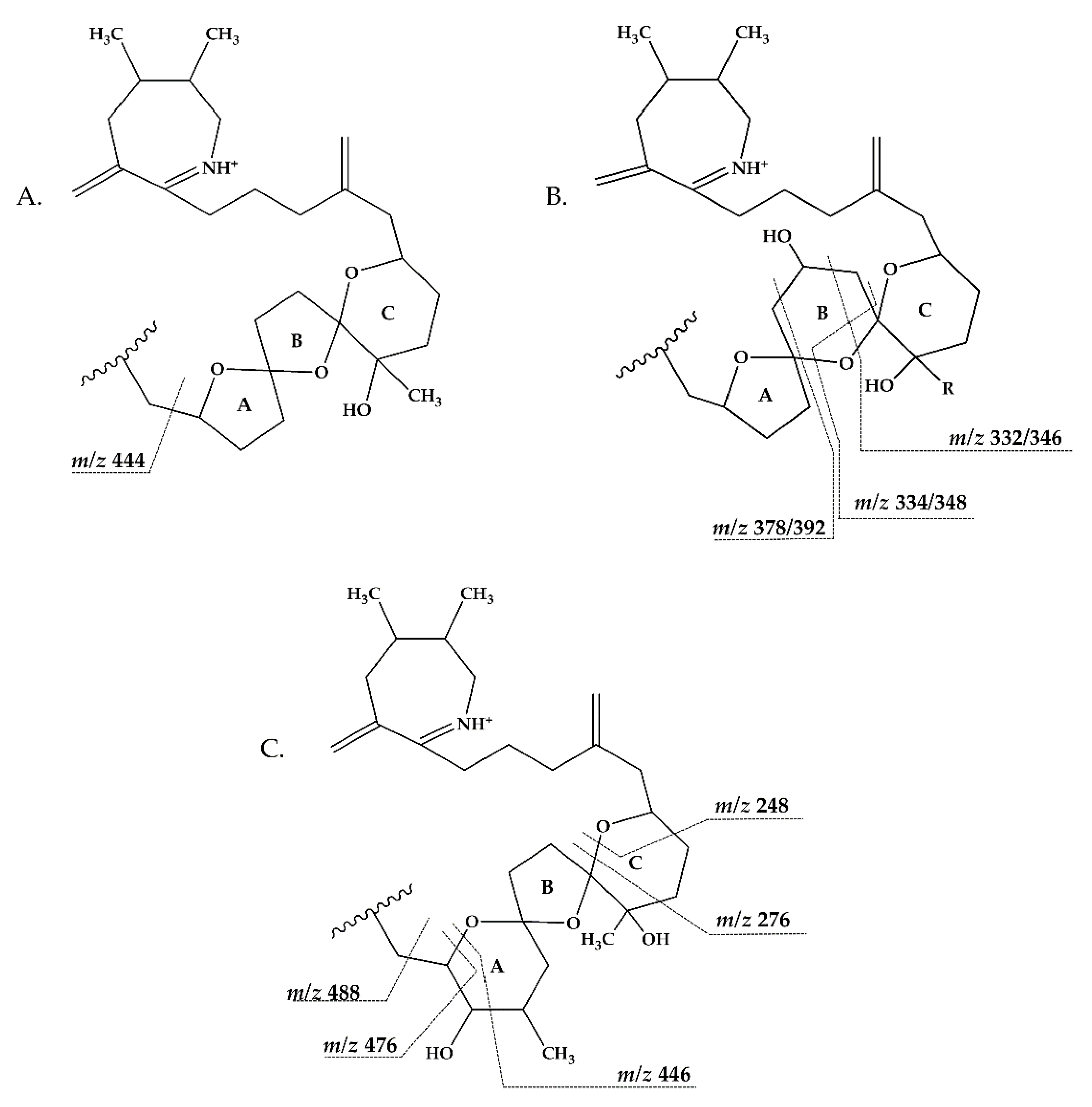
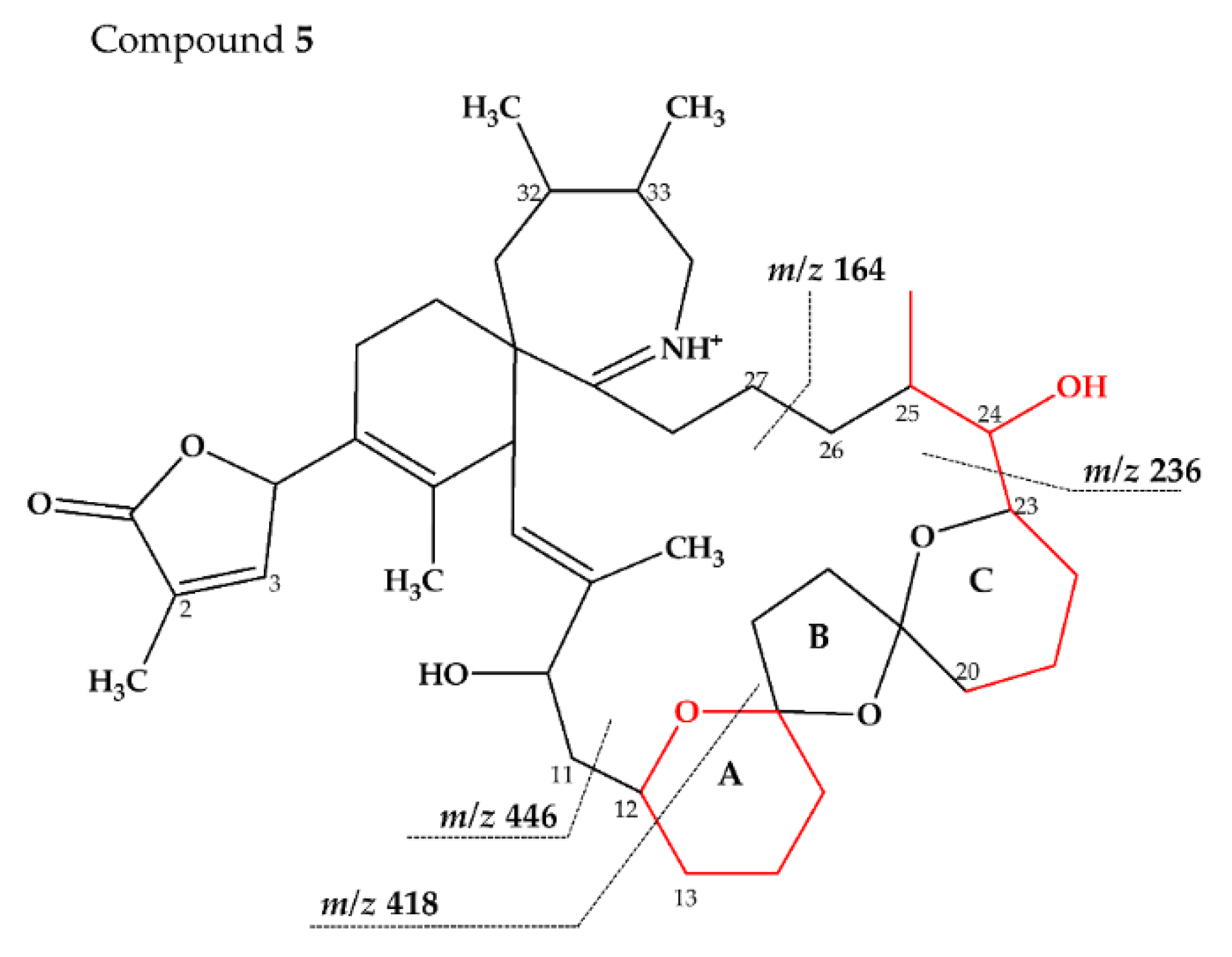
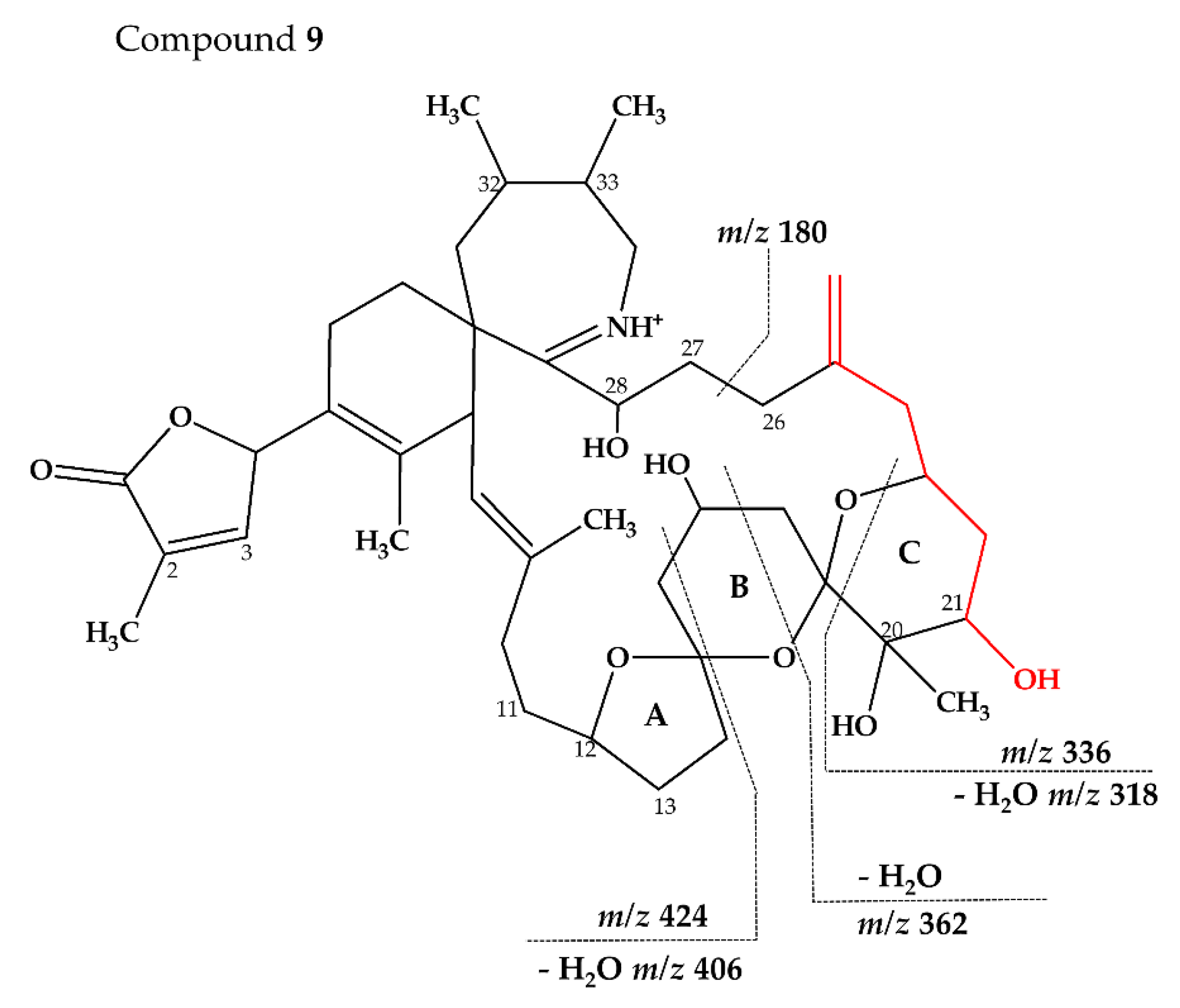
2.2.3. Compounds 4–9
2.3. Structural Variability of Spirolides
3. Materials and Methods
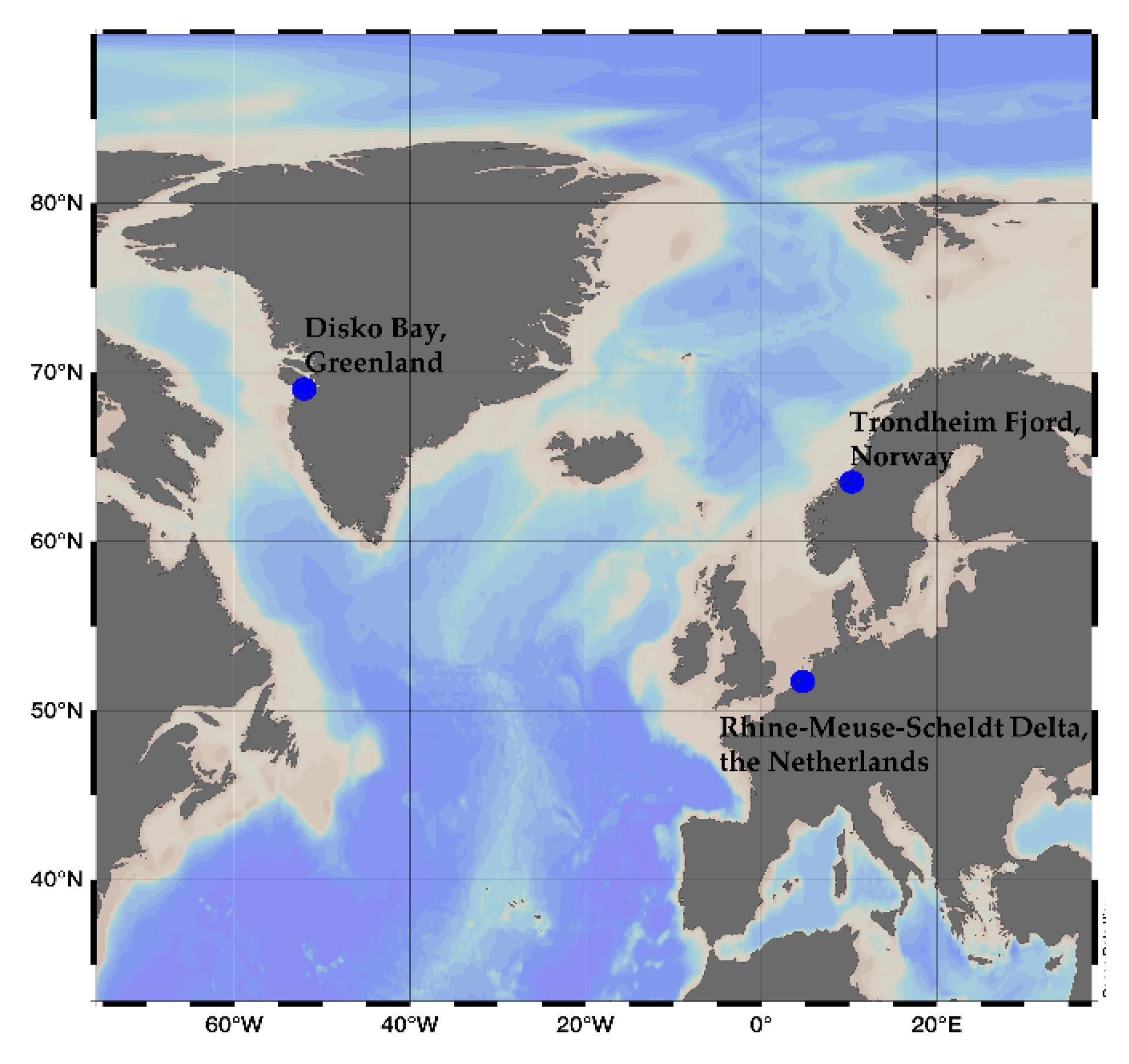
3.1. Culture Conditions and Cell Harvest
3.2. Spirolide Extraction
3.3. Parameters of Liquid Chromatography-Tandem Mass Spectrometry (LC-MS/MS)
3.3.1. Liquid Chromatography (LC)
3.3.2. Scan Modes in Tandem Mass Spectrometry (MS/MS)
3.4. Analyses of Spirolides by High Resolution Tandem Mass Spectrometry (HR-MS/MS)
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Glibert, P.M.; Anderson, D.M.; Gentien, P.; Granéli, E.; Sellner, K.G. The Global, Complex Phenomena of Harmful Algal Blooms. Oceanography 2005, 18, 136–147. [Google Scholar] [CrossRef]
- Sellner, K.G.; Doucette, G.J.; Kirkpatrick, G.J. Harmful algal blooms: Causes, impacts and detection. J. Ind. Microbiol. Biotechnol. 2003, 30, 383–406. [Google Scholar] [CrossRef] [PubMed]
- Hu, T.; Curtis, J.M.; Oshima, Y.; Quilliam, M.A.; Walter, J.A.; Watson-Wright, W.M.; Wright, J.L.C. Spirolides B and D, two novel macrocycles isolated from the digestive glands of shellfish. J. Chem. Soc. Chem. Commun. 1995, 2159–2161. [Google Scholar] [CrossRef]
- Hu, T.; Curtis, J.M.; Walter, J.A.; Wright, J.L.C. Characterization of biologically inactive spirolides E and F: Identification of the spirolide pharmacophore. Tetrahedron Lett. 1996, 37, 7671–7674. [Google Scholar] [CrossRef]
- Cembella, A.D.; Lewis, N.I.; Quilliam, M.A. The marine dinoflagellate Alexandrium ostenfeldii (Dinophyceae) as the causative organism of spirolide shellfish toxins. Phycologia 2000, 39, 67–74. [Google Scholar] [CrossRef]
- Paulsen, O. Plankton-investigations in the waters round Iceland in 1903. Medd. Kommn. Havunders. København Ser Plankt 1904, 1, 1–40. [Google Scholar]
- Hu, T.; Burton, I.W.; Cembella, A.D.; Curtis, J.M.; Quilliam, M.A.; Walter, J.A.; Wright, J.L.C. Characterization of spirolides A, C, and 13-desmethyl C, new marine toxins isolated from toxic plankton and contaminated shellfish. J. Nat. Prod. 2001, 64, 308–312. [Google Scholar] [CrossRef]
- Sleno, L.; Windust, A.J.; Volmer, D.A. Structural study of spirolide marine toxins by mass spectrometry Part II. Mass spectrometric characterization of unknown spirolides and related compounds in a cultured phytoplankton extract. Anal. Bioanal. Chem. 2004, 378, 977–986. [Google Scholar] [CrossRef]
- Ciminiello, P.; Dell’Aversano, C.; Fattorusso, E.; Forino, M.; Grauso, L.; Tartaglione, L.; Guerrini, F.; Pistocchi, R. Spirolide Toxin Profile of Adriatic Alexandrium ostenfeldii Cultures and Structure Elucidation of 27-Hydroxy-13,19-Didesmethyl Spirolide, C.J. Nat. Prod. 2007, 70, 1878–1883. [Google Scholar] [CrossRef]
- Ciminiello, P.; Dell’Aversano, C.; Iacovo, E.D.; Fattorusso, E.; Forino, M.; Grauso, L.; Tartaglione, L.; Guerrini, F.; Pezzolesi, L.; Pistocchi, R. Characterization of 27-hydroxy-13-desmethyl spirolide C and 27-oxo-13,19-didesmethyl spirolide C. Further insights into the complex Adriatic Alexandrium ostenfeldii toxin profile. Toxicon 2010, 56, 1327–1333. [Google Scholar] [CrossRef] [PubMed]
- Zurhelle, C.; Nieva, J.; Tillmann, U.; Harder, T.; Krock, B.; Tebben, J. Identification of Novel Gymnodimines and Spirolides from the Marine Dinoflagellate Alexandrium ostenfeldii. Mar. Drugs 2018, 16, 446. [Google Scholar] [CrossRef] [PubMed]
- Aasen, J.; MacKinnon, S.L.; LeBlanc, P.; Walter, J.A.; Hovgaard, P.; Aune, T.; Quilliam, M.A. Detection and identification of spirolides in Norwegian shellfish and plankton. Chem. Res. Toxicol. 2005, 18, 509–515. [Google Scholar] [CrossRef] [PubMed]
- Roach, J.S.; LeBlanc, P.; Lewis, N.I.; Munday, R.; Quilliam, M.A.; MacKinnon, S.L. Characterization of a Dispiroketal Spirolide Subclass from Alexandrium ostenfeldii. J. Nat. Prod. 2009, 72, 1237–1240. [Google Scholar] [CrossRef] [PubMed]
- Hummert, C.; Rühl, A.; Reinhardt, K.; Gerdts, G.; Luckas, B. Simultaneous analysis of different algal toxins by LC-MS. Chromatographia 2002, 55, 673–680. [Google Scholar] [CrossRef]
- MacKinnon, S.L.; Walter, J.A.; Quilliam, M.A.; Cembella, A.D.; LeBlanc, P.; Burton, I.W.; Hardstaff, W.R.; Lewis, N.I. Spirolides Isolated from Danish Strains of Toxigenic Dinoflagellate Alexandrium ostenfeldi. J. Nat. Prod. 2006, 69, 983–987. [Google Scholar] [CrossRef]
- Villar Gonzáles, A.; Rodríguez-Velasco, L.M.; Ben-Gigirey, B.; Botana, L.M. First evidence of spirolides in Spanish shellfish. Toxicon 2006, 48, 1068–1075. [Google Scholar] [CrossRef]
- Touzet, N.; Franco, J.M.; Raine, R. Morphogenetic Diversity and Biotoxin Composition of Alexandrium (Dinophyceae) in Irish coastal Waters. Harmful Algae 2008, 7, 782–797. [Google Scholar] [CrossRef]
- Katikou, P.; Aligizaki, K.; Zacharaki, T.; Iossifidis, D.; Nikolaidis, G. First Report of Spirolides in Greek Shellfish Associated with Causative Alexandrium Species. In Proceedings of the 14th International Conference on Harmful Algae, Crete Island, Greece, 1–5 November 2010; Pagou, P., Hallegraeff, G., Eds.; International Society for the Study of Harmful Algae and Intergovernmental Oceanographic Commission of UNESCO: Paris, France, 2013. [Google Scholar]
- Tillmann, U.; Kremp, A.; Tahvanainen, P.; Krock, B. Characterization of spirolide producing Alexandrium ostenfeldii (Dinophyceae) from the western Arctic. Harmful Algae 2014, 39, 259–270. [Google Scholar] [CrossRef]
- Van de Waal, D.B.; Tillmann, U.; Martens, H.; Krock, B.; van Scheppingen, Y.; John, U. Characterization of multiple isolates from an Alexandrium ostenfeldii bloom in The Netherlands. Harmful Algae 2015, 49, 94–104. [Google Scholar] [CrossRef]
- Martens, H.; Tillmann, U.; Harju, K.; Dell’Aversano, C.; Tartaglione, L.; Krock, B. Toxin Variability Estimations of 68 Alexandrium ostenfeldii (Dinophyceae) Strains from The Netherlands Reveal a Novel Abundant Gymnodimine. Microorganisms 2017, 5, 29. [Google Scholar] [CrossRef]
- Gribble, K.E.; Keafer, B.A.; Quilliam, M.A.; Cembella, A.D.; Kulis, D.M.; Manahan, A.; Anderson, D.M. Distribution and toxicity of Alexandrium ostenfeldii (Dinophyceae) in the Gulf of Maine, USA. Deep–Sea Res. II. 2005, 52, 2745–2763. [Google Scholar] [CrossRef]
- Almandoz, G.O.; Montoya, N.G.; Hernando, M.P.; Benavides, H.R.; Carignan, M.O.; Ferrario, M.E. Toxic strains of the Alexandrium ostenfeldii complex in southern South America (Beagle Channel, Argentina). Harmful Algae 2014, 37, 100–109. [Google Scholar] [CrossRef]
- Álvarez, G.; Uribe, E.; Ávalos, P.; Mariño, C.; Blanco, J. First identification of azaspiracid and spirolides in Mesodesma donacium and Mulinia edulis from Northern Chile. Toxicon 2010, 55, 638–641. [Google Scholar] [CrossRef] [PubMed]
- Guinder, V.A.; Tillmann, U.; Krock, B.; Delgado, A.L.; Krohn, T.; Garzón Cardona, J.E.; Metfies, K.; Abbate, C.L.; Silva, R.; Lara, R. Plankton Multiproxy Analyses in the Northern Patagonian Shelf, Argentina: Community Structure, Phycotoxins, and Characterization of Toxic Alexandrium Strains. Front. Mar. Sci. 2018, 5, 394. [Google Scholar] [CrossRef]
- Qui, J.; Rafuse, C.; Lewis, N.I.; Li, A.; Meng, F.; Beach, D.G.; McCarron, P. Screening of cyclic imine and paralytic shellfish toxins in isolates of the genus Alexandrium (Dinophyceae) from Atlantic Canada. Harmful Algae 2018, 77, 108–118. [Google Scholar] [CrossRef]
- Munday, R.; Quilliam, M.A.; Leblanc, P.; Lewis, N.; Gallant, P.; Sperker, S.A.; Ewart, H.S.; MacKinnon, S.L. Investigation into the toxicology of spirolides, a group of marine phycotoxins. Toxins 2012, 4, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Otero, P.; Alfonso, A.; Rodríguez, P.; Rubiolo, J.A.; Cifuentes, J.M.; Bermúdez, R.; Vieytes, M.R.; Botana, L.M. Pharmacokinetic and toxicological data of spirolides after oral and intraperitoneal administration. Food Chem. Toxicol. 2012, 50, 232–237. [Google Scholar] [CrossRef] [PubMed]
- Aráoz, R.; Ouanounou, G.; Iorga, B.I.; Goudet, A.; Alili, D.; Amar, M.; Benoit, E.; Molgó, J.; Servent, D. The Neurotoxic Effect of 13,19-Didesmethyl and 13-Desmethyl Spirolide C Phycotoxins Is Mainly Mediated by Nicotinic Rather Than Muscarinic Acetylcholine Receptors. Toxicol. Sci. 2015, 147, 156–157. [Google Scholar] [CrossRef]
- Couesnon, A.; Aráoz, R.; Iorga, B.I.; Benoit, E.; Reynaud, M.; Servent, D.; Molgó, J. The Dinoflagellate Toxin 20-Methyl Spirolide-G Potently Blocks Skeletal Muscle and Neuronal Nicotinic Acetylcholine Receptors. Toxins 2016, 8, 249. [Google Scholar] [CrossRef]
- EFSA Panel on Contaminants in the Food Chain (CONTAM). Scientific Opinion on marine biotoxins in shellfish—Cyclic imines (spirolides, gymnodimines, pinnatoxins and pteriatoxins). EFSA J. 2010, 8, 1628–1667. [Google Scholar] [CrossRef]
- Nielsen, L.T.; Krock, B.; Hansen, P.J. Production and excretion of okadaic acid, pectenotoxin-2 and a novel dinophysistoxin from the DSP-causing marine dinoflagellate Dinophysis acuta—Effects of light, food availability and growth phase. Harmful Algae 2013, 23, 34–45. [Google Scholar] [CrossRef]
- Krock, B.; Busch, J.A.; Tillmann, U.; García-Camacho, F.; Sánchez-Mirón, A.; Gallardo-Rodríguez, J.J.; López-Rosales, L.; Andree, K.B.; Fernández-Tejedor, M.; Witt, M.; et al. LC-MS/MS Detection of Karlotoxins Reveals New Variants in Strains of the Marine Dinoflagellate Karlodinium veneficum from the Ebro Delta (NW Mediterranean). Mar. Drugs 2017, 15, 391. [Google Scholar] [CrossRef] [PubMed]
- Zendong, Z.; McCarron, P.; Herrenknecht, C.; Sibat, M.; Amzil, Z.; Cole, R.B.; Hess, P. High resolution mass spectrometry for quantitative analysis and untargeted screening of algal toxins in mussels and passive samplers. J. Chromatogr. A 2015, 1416, 10–21. [Google Scholar] [CrossRef] [PubMed]
- McLafferty, F.W.; Tureček, F. Interpretation of Mass Spectra, 4th ed.; University Science Books: Mill Valley, CA, USA, 1993; pp. 12–13. [Google Scholar]
- Sleno, L.; Windust, A.J.; Volmer, D.A. Structural study of spirolide marine toxins by mass spectrometry Part I. Fragmentation pathways of 13-desmethyl spirolide C by collision-induced dissociation and infrared multiphoton dissociation mass spectrometry. Anal. Bioanal. Chem. 2004, 378, 969–976. [Google Scholar] [CrossRef]
- Christian, B.; Below, A.; Dreßler, N.; Scheibner, O.; Luckas, B.; Gerdts, G. Are spirolides converted in biological systems?—A study. Toxicon 2008, 51, 934–940. [Google Scholar] [CrossRef]
- Selwood, A.I.; Miles, C.O.; Wilkinson, A.L.; Van Ginkel, R.; Munday, R.; Rise, F.; McNabb, P. Isolation, Structural Determination and Acute Toxicity of Pinnatoxins E, F and G. J. Agric. Food Chem. 2010, 58, 6532–6542. [Google Scholar] [CrossRef]
- Van Wagoner, R.M.; Satake, M.; Wright, J.L.C. Polyketide biosynthesis in dinoflagellate: What makes it different? Nat. Prod. Rep. 2014, 31, 1101. [Google Scholar] [CrossRef]
- Rhodes, L.; Smith, K.; Selwood, A.; McNabb, P.; Munday, R.; Suda, S.; Molenaar, S.; Hallegraeff, G. Dinoflagellate Vulcanodinium rugosum identified as the causative organism of pinnatoxins in Australia, New Zealand and Japan. Phycologia 2011, 50, 624–628. [Google Scholar] [CrossRef]
- Seki, T.; Satake, M.; MacKenzie, L.; Kaspar, H.F.; Yasumoto, T. Gymnodimine, a new marine toxin of unprecedented structure isolated from New Zealand oysters and the dinoflagellate, Gymnodinium sp. Tetrahedron Lett. 1995, 36, 7093–7096. [Google Scholar] [CrossRef]
- Miles, C.O.; Samdal, I.A.; Aasen, J.A.G.; Jensen, D.J.; Quilliam, M.A.; Petersen, D.; Briggs, L.M.; Wilkins, A.L.; Rise, F.; Cooney, J.M.; et al. Evidence for numerous analogs of yessotoxin in Protoceratium reticulatum. Harmful Algae 2005, 4, 1075–1091. [Google Scholar] [CrossRef]
- Binzer, S.B.; Svenssen, D.K.; Daugbjerg, N.; Alves-de-Souza, C.; Pinto, E.; Hansen, P.J.; Larsen, T.O.; Varga, E. A-, B- and C-type prymnesins are clade specific compounds and chemotaxonomic markers in Prymnesium parvum. Harmful Algae 2019, 81, 10–17. [Google Scholar] [CrossRef] [PubMed]
- Trautmann, T. Intraspecific Trait Diversity in the Marine Dinoflagellate Alexandrium ostenfeldii and Resulting Effects on the Community Composition. Master’s Thesis, University of Bremen, Bremen, Germany, 2017. [Google Scholar]
- Nagai, S.; Yasuike, M.; Nakamura, Y.; Tahvanainen, P.; Kremp, A. Development of ten microsatellite markers for Alexandrium ostenfeldii, a bloom-forming dinoflagellate producing diverse phycotoxins. J. Appl. Phycol. 2015, 27, 2333–2339. [Google Scholar] [CrossRef]
| Spirolide | R1 | R2 | R3 | R4 | R5 | R6 | ΔC2,3 | Mass Transition | Structure | Reference |
|---|---|---|---|---|---|---|---|---|---|---|
| A | H | CH3 | CH3 | H | H | OH | + | 692 > 150 | 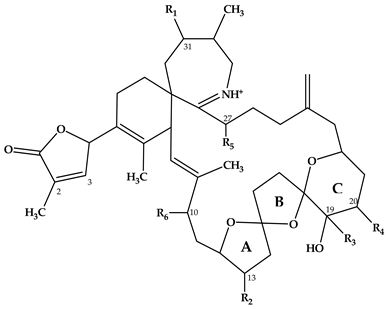 | [7] |
| B | H | CH3 | CH3 | H | H | OH | - | 694 > 150 | [3,4] | |
| C | CH3 | CH3 | CH3 | H | H | OH | + | 706 > 164 | [7] | |
| 13-desMethyl C | CH3 | H | CH3 | H | H | OH | + | 692 > 164 | [7] | |
| 13,19-didesMethyl C | CH3 | H | H | H | H | OH | + | 678 > 164 | [9,15] | |
| 20-Hydroxy-13,19-didesMethyl C | CH3 | H | H | OH | H | OH | + | 694 > 164 | [11] | |
| 27-Hydroxy-13-desMethyl C | CH3 | H | CH3 | H | OH | OH | + | 708 > 180 | [10] | |
| 27-Hydroxy-13,19-didesMethyl C | CH3 | H | H | H | OH | OH | + | 694 > 180 | [10] | |
| 27-Oxo-13,19-didesMethyl C | CH3 | H | H | H | O | OH | + | 692 > 178 | [10] | |
| D | CH3 | CH3 | CH3 | H | H | OH | - | 708 > 164 | [3] | |
| 13-desMethyl D | CH3 | H | CH3 | H | H | OH | - | 694 > 164 | [8] | |
| 20-Hydroxy-13,19-didesMethyl D | CH3 | H | H | OH | H | OH | - | 696 > 164 | [11] | |
| G | CH3 | H | H | H | H | H | + | 692 > 164 |  | [12] |
| 20-Methyl G | CH3 | H | CH3 | H | H | H | + | 706 > 164 | [12] | |
| H | CH3 | H | CH3 | H | H | OH | + | 650 > 164 | 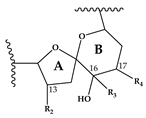 | [13] |
| I | CH3 | H | CH3 | H | H | OH | - | 652 > 164 | [13] |
| Strain | Compound | Elemental Formula | Measured | Calculated | Δ/ppm |
|---|---|---|---|---|---|
| AON 24 (The Netherlands) | 1 (670 > 164) | C40H64NO7+ | 670.4678 | 670.4677 | 0.07 |
| C38H58NO4+ | 592.4363 | 592.4360 | 0.51 | ||
| C27H44NO5+ | 462.3219 | 462.3214 | 1.01 | ||
| C27H42NO4+ | 444.3111 | 444.3108 | 0.66 | ||
| C18H28N+ | 258.2217 | 258.2216 | 0.21 | ||
| C16H24N+ | 230.1904 | 230.1903 | 0.32 | ||
| C11H18N+ | 164.1435 | 164.1434 | 0.81 | ||
| NX-56-10 (Norway) | 2 (666 > 164) | C40H60NO7+ | 666.4365 | 666.4364 | 0.14 |
| C35H50NO3+ | 532.3787 | 532.3785 | 0.38 | ||
| C22H36NO4+ | 378.2640 | 378.2639 | 0.32 | ||
| C22H34NO3+ | 360.2533 | 360.2533 | −0.07 | ||
| C21H34NO2+ | 332.2584 | 332.2584 | −0.13 | ||
| C17H28N+ | 246.2216 | 246.2216 | 0.09 | ||
| C11H18N+ | 164.1435 | 164.1434 | 0.62 | ||
| 3 (696 > 164) | C41H62NO8+ | 696.4471 | 696.4470 | 0.13 | |
| C38H54NO3+ | 572.4100 | 572.4098 | 0.37 | ||
| C26H42NO5+ | 448.3060 | 448.3057 | 0.56 | ||
| C26H40NO4+ | 430.2954 | 430.2952 | 0.52 | ||
| C17H28N+ | 246.2217 | 246.2216 | 0.34 | ||
| C11H18N+ | 164.1435 | 164.1434 | 0.90 |
| Strain | Compound | Elemental Formula | Measured | Calculated | Δ/ppm |
|---|---|---|---|---|---|
| MX-S-B11 (Greenland) | 4 (678 > 150) | C41H60NO7+ | 678.4363 | 678.4364 | −0.22 |
| C26H42NO5+ | 448.3059 | 448.3057 | 0.42 | ||
| C26H40NO4+ | 430.2953 | 430.2952 | 0.38 | ||
| C17H26N+ | 244.2061 | 244.2060 | 0.32 | ||
| C10H16N | 150.1278 | 150.1277 | 0.64 | ||
| 5 (694 > 164) | C42H64NO7+ | 694.4679 | 694.4677 | 0.24 | |
| C29H48NO4+ | 474.3581 | 474.3578 | 0.65 | ||
| C27H46NO5+ | 464.3374 | 464.3370 | 0.76 | ||
| C26H44NO3+ | 418.3318 | 418.3316 | 0.59 | ||
| C15H26NO+ | 236.2010 | 236.2009 | 0.35 | ||
| C11H18N+ | 164.1435 | 164.1434 | 0.90 | ||
| 6 (708 > 164) | C43H66NO7+ | 708.4836 * | 708.4834 | 1.06 | |
| C23H38NO4+ | 392.2798 | 392.2795 | 0.56 | ||
| C22H36NO2+ | 346.2744 | 346.2741 | 0.85 | ||
| C11H18N+ | 164.1439 | 164.1434 | −0.08 |
| Strain | Compound | Elemental Formula | Measured | Calculated | Δ/ppm |
|---|---|---|---|---|---|
| MX-S-B11 (Greenland) | 7 (720 > 164) | C44H66NO7+ | 720.4836 | 720.4834 | 0.37 |
| C29H46NO5+ | 490.3530 | 490.3527 | 0.60 | ||
| C29H44NO4+ | 472.3425 | 472.3421 | 0.70 | ||
| C23H38NO3+ | 376.2849 | 376.2846 | 0.77 | ||
| C23H36NO2+ | 358.2741 | 358.2741 | 0.05 | ||
| C22H38NO2+ | 346.2743 | 346.2741 | 0.21 | ||
| C16H24N+ | 230.1905 | 230.1903 | 0.58 | ||
| C11H18N+ | 164.1435 | 164.1434 | 0.72 | ||
| 8 (722 > 164) | C44H68NO7+ | 722.4994 | 722.4990 | 0.50 | |
| C23H38NO4+ | 392.2796 | 392.2795 | 0.17 | ||
| C23H36NO3+ | 374.2691 | 374.2690 | 0.28 | ||
| C22H38NO3+ | 364.2691 | 364.2846 | −0.12 | ||
| C22H36NO2+ | 346.2741 | 346.2741 | 0.24 | ||
| C18H28N+ | 258.2217 | 258.2216 | 0.32 | ||
| C16H24N+ | 230.1904 | 230.1903 | 0.45 | ||
| C11H18N+ | 164.1435 | 164.1434 | 0.72 | ||
| 9 (738–180) | C43H64NO9+ | 738.4579 | 738.4576 | 0.44 | |
| C23H38NO6+ | 424.2697 | 424.2694 | 0.68 | ||
| C22H36NO3+ | 362.2693 | 362.2690 | 0.79 | ||
| C20H32NO3+ | 336.2537 | 336.2533 | 1.10 | ||
| C20H32NO2+ | 318.2429 | 318.2428 | 0.42 | ||
| C15H26NO+ | 236.2010 | 236.2009 | 0.67 | ||
| C11H18NO+ | 180.1384 | 180.1383 | 0.59 |
| Spirolide | MX-S-B11 (Greenland) | NX-56-10 (Norway) | AON 24 (The Netherlands) | Ketal Ring System | |||
|---|---|---|---|---|---|---|---|
| Cell Quota (fg cell−1) | Relative Content (%) | Cell Quota (fg cell−1) | Relative Content (%) | Cell Quota (fg cell−1) | Relative Content (%) | (A:B:C) | |
| Known | |||||||
| 13-desMethyl spirolide C | 34.5 | 0.73 | 146.1 | 6.37 | 740.3 | 99.90 | 5:5:6 |
| 20-Methyl spirolide G | 2086.4 | 44.41 | 5.1 | 0.22 | 0.4 | 0.05 | 5:6:6 |
| Spirolide A | 384.3 | 8.18 | 0.3 | 0.01 | <LOD (0.06) | Nd | 5:5:6 |
| Spirolide C | 33.9 | 0.72 | 0.1 | 0.01 | 0.1 | 0.02 | 5:5:6 |
| 13,19-didesMethyl spirolide C | <LOD (0.16) | Nd | 2037.9 | 88.85 | <LOD (0.06) | Nd | 5:5:6 |
| Spirolide H | 257.3 | 5.48 | 0.4 | 0.02 | <LOD (0.06) | Nd | 5:6 |
| Spirolide I | 158.4 | 3.37 | 0.3 | 0.01 | <LOD (0.06) | Nd | 5:6 |
| Unknown | |||||||
| Compound 1 | <LOD (0.16) | Nd | <LOD (0.07) | Nd | 0.2 | 0.03 | 5:5:6 |
| Compound 2 | <LOD (0.16) | Nd | 81.0 | 3.53 | <LOD (0.06) | Nd | 5:6:6 |
| Compound 3 | <LOD (0.16) | Nd | 22.3 | 0.97 | <LOD (0.06) | Nd | 5:5:6 |
| Compound 4 | 19.6 | 0.42 | <LOD (0.07) | Nd | <LOD (0.06) | Nd | 5:5:6 |
| Compound 5 | 233.7 | 4.98 | <LOD (0.07) | Nd | <LOD (0.06) | Nd | 6:5:6 |
| Compound 6 | 883.2 | 18.80 | <LOD (0.07) | Nd | <LOD (0.06) | Nd | 5:6:6 |
| Compound 7 | 423.5 | 9.02 | <LOD (0.07) | Nd | <LOD (0.06) | Nd | 5:6:6 |
| Compound 8 | 182.0 | 3.87 | <LOD (0.07) | Nd | <LOD (0.06) | Nd | 5:6:6 |
| Compound 9 | 0.8 | 0.02 | <LOD (0.07) | Nd | <LOD (0.06) | Nd | 5:6:6 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nieva, J.A.; Tebben, J.; Tillmann, U.; Wohlrab, S.; Krock, B. Mass Spectrometry-Based Characterization of New Spirolides from Alexandrium ostenfeldii (Dinophyceae). Mar. Drugs 2020, 18, 505. https://doi.org/10.3390/md18100505
Nieva JA, Tebben J, Tillmann U, Wohlrab S, Krock B. Mass Spectrometry-Based Characterization of New Spirolides from Alexandrium ostenfeldii (Dinophyceae). Marine Drugs. 2020; 18(10):505. https://doi.org/10.3390/md18100505
Chicago/Turabian StyleNieva, Joyce A., Jan Tebben, Urban Tillmann, Sylke Wohlrab, and Bernd Krock. 2020. "Mass Spectrometry-Based Characterization of New Spirolides from Alexandrium ostenfeldii (Dinophyceae)" Marine Drugs 18, no. 10: 505. https://doi.org/10.3390/md18100505
APA StyleNieva, J. A., Tebben, J., Tillmann, U., Wohlrab, S., & Krock, B. (2020). Mass Spectrometry-Based Characterization of New Spirolides from Alexandrium ostenfeldii (Dinophyceae). Marine Drugs, 18(10), 505. https://doi.org/10.3390/md18100505





