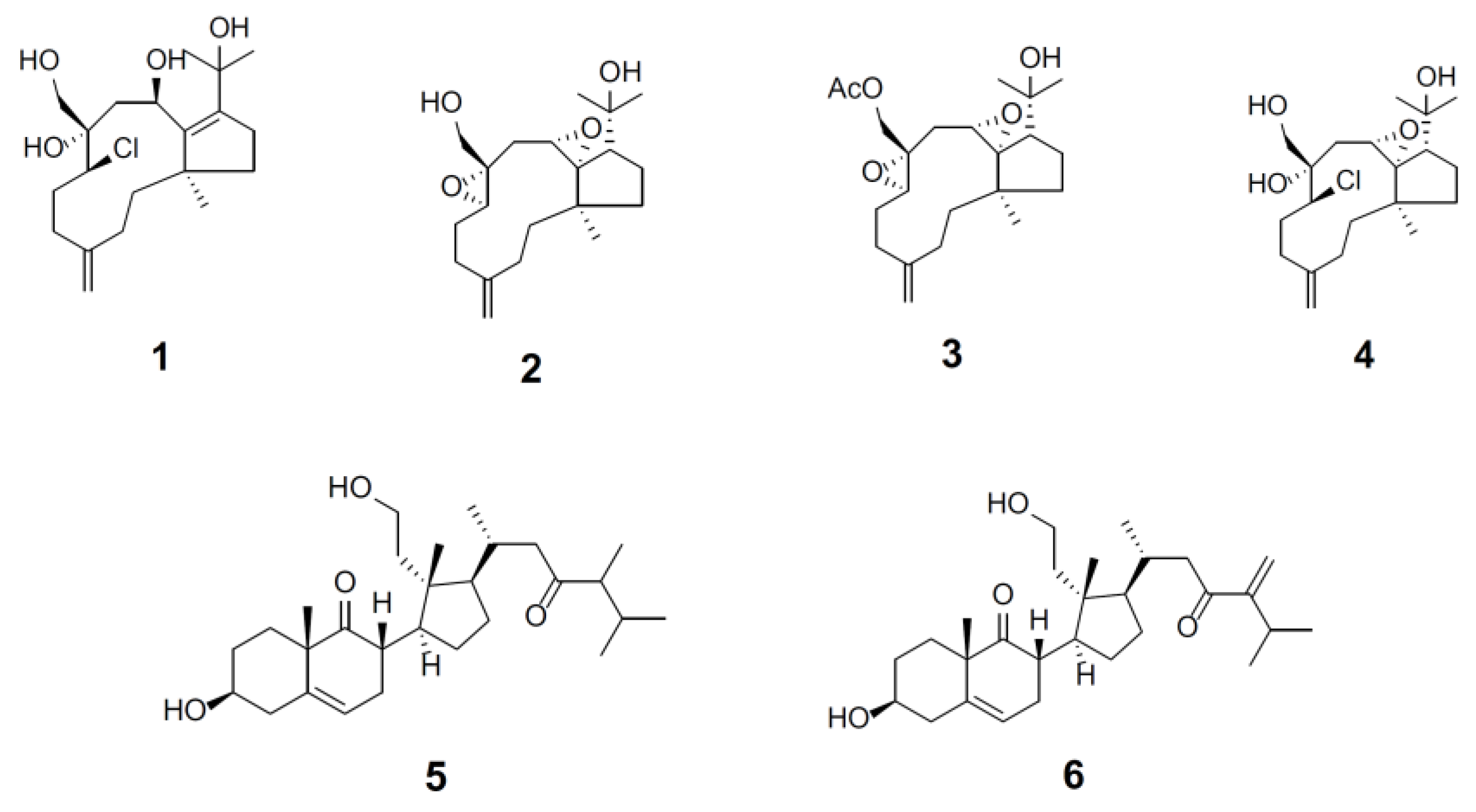
2.1. Compound Purification and Structure Elucidation
Chromatographic fractionation of ethyl acetate solubles from
C. flava afforded four dolabellane diterpenes,
1–
4, as well as two secosteroids,
5 and
6. The three known dolabellane diterpenes,
2–
4 were identified by comparison of their spectral data with those of reported literatures [
5,
6]. The structures of new compounds,
1,
5, and
6 were elucidated by analysis of 1D and 2D spectral data (
Figures S1–S24).
Clavinflol C (
1) had a molecular formula of C
20H
33O
4Cl as deduced from HR-ESI-MS (
Figure S1) and NMR data. Its IR bands (
Figure S2) indicated the presence of exo-methylene (1,639, 961 cm
−1) and hydroxyl (3343 cm
−1) groups. The
1H NMR data of
1 (
Figure S3) showed a pair of exo-methylene singlets (
4.91, 5.00) and an AB quartet for hydroxy-methyl group (
3.39, 3.82, J = 11.6 Hz), three methyl singlets (
1.03, 1.32, 1.40), an oxygenated methine proton (
4.18), and a chlorinated methine proton (
4.00). To determine the proton sequence of
1, a COSY spectrum (
Figure S6) revealed the connectiveness of H-2/H-3, H-5/H-6/H-7, H-9/H-10, and H-13/H-14. The
13C NMR (
Figure S4) and HSQC spectra (
Figure S5) of
1 showed signals for three methyl carbons, eight methylene carbons including the exomethylene (
113.9, 147.7), two methine carbons, and five quaternary carbons. Detailed analyses of the
1H,
13C NMR, and HSQC spectra revealed that
1 is a dolabellane diterpene with a 5/11 membered ring and a tetrasubstituted olefin at the C-11/C-12 positions. This type of skeleton was further confirmed from the observation of long range correlations of H
2-16/C-3, C-5; H
2-2/C-4; H
2-7/C-8, C-17; H-10/C-1, C-7, C-8, C-9, C-11, H
3-15/C-1, C-2, C-11, C-13; H
2-13/C-1, C-11, C-12; H
2-14/ C-11, C-12; H
3-19/C-12, C-18, C-20; H
3-20/C-12, C-18, C-19 in the HMBC spectrum (
Figure S7). The relative stereochemistry of
1 was determined from NOESY experiments as illustrated in
Figure S8. Assuming that H
3-15 is α-oriented, key NOESY correlations from H
3-15 to H-10 and from H-7 to H-10 suggested that H-10 and H-7 were in the α-orientation. NOESY correlations between H-9a/H-6a and H-9b/H-17b suggested that OH-8 was in the
β-orientation.
Compound
5 was isolated as a white amorphous powder, showing a pseudo-molecular ion peak at
m/
z 469.32880 [M + Na]
+ in the HR-ESI-MS (
Figure S9), consistent with the molecular formula C
28H
46 NaO
4 (calculated for 469.32883), requiring six degrees of unsaturation. The presence of an oxymethylene and a keto carbonyl carbon was confirmed by the
1H NMR (
Figure S11) (δ
H 3.88 (m, H-11a) and 3.74 (m, H-11b)) and
13C NMR (
Figure S12) (δ
C 59.2 (CH
2), 212.5 (qC), and 216.2 (qC)) data, as well as from the IR absorption (
Figure S10) at 3396 and 1704 cm
−1. The diagnostic NMR signals of a 9,11-secosterol were confirmed by the
1H–
1H COSY correlation (
Figure S14) from H
2-11 to H
2-12 as well as HMBC correlations (
Figure S15) from H3-18 to C-12, C-13, C-14, and C-17; from H3-19 to C-1, C-5, C-9, and C-10. The NMR features of
5 were analogous to those of 3,11-dihydroxy-24-methyl-9,11-secocholest-5-en-9-one [
7], except for the presence of a ketone (δ
C 201.1 (qC)) at C-23. Based on NOESY correlations (
Figure S16) of H
3-19/H-1, H
3-19/H-2, H
3-19/H-4, H
3-19/H-8, H-3/H-1, H-3/H-2, H-3/H-4, H-8/H
3-18, H-8/H-7, H
3-18/H-15, H
3-18/H-16, H
3-18/H-20, and H-14/H-7, the relative stereochemistry at C-3, C-8, C-10, C-13, C-14, C-17, and C-20 in
5 was found to be the same as those of 3
β,11-dihydroxy-24-methyl-9,11-secocholest-5-en-9-one [
7]. On the basis of the above-mentioned findings, the structure of
5 was consistent with the structure shown as 3
β,11-dihydroxy-24-methyl-9,11-secocholest-5-en-9,23-dione.
Compound
6 appeared as a white amorphous powder like
5. Careful inspection of the 2D NMR spectroscopic data (
Figures S21–23) of
6 led to the establishment of the same nucleus as that of
5. The NMR spectroscopic data (
Figures S19 and S20) of
6 were analogous to those of
5, except for NMR signals due to the conjugated enone in
6. The location of the conjugated enone was identified by the HMBC correlations (
Figure S23) from the methylene protons (H
2-22) to the carbonyl carbon (C-23) and from H
3-26, 27 to C-24, securing the structure of
6, which was shown as 3
β,11-dihydroxy-24-methylene-9,11-secocholest-5-en-9,23-dione.
2.2. Identification of Marine Compounds Showed High-Content Characteristics of Proteasome Inhibition
The proteasome inhibition assay was performed by following the standard operation protocol of high-content screening (HCS) of EGFP-UL76 aggresome as described previously [
4]. A stringent proteasome inhibition was considered as the HCS measurements of marine compounds with an increase greater than 0.2-fold relative to those of the control without treatment. Under this validity criterion, we demonstrated the identification of six compounds with proteasome inhibition and their effects were statistically significant. Four compounds with dolabellane-based structures designated clavinflol C (
1), stolonidiol (
2), stolonidiol-17-acetate (
3), and clavinflol B (
4) (
Figure 2 and
Figure 3) [
5,
8]. Additionally, two unprecedent compounds with secosteroid-based structures designated compound
5 and
6 (
Figure 4 and
Figure 5). Prior to HCS experiments, the in vitro cell-based MTT (3-(4,5-Dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide) cytotoxicity assays were performed against four cell lines: A549 (human lung adenocarcinoma), HT-29 (human colon adenocarcinoma), and P-388 (mouse lymphocytic leukemia). Plasmid pEGFP-UL76 transfected HEK293T (human embryonic kidney large-T antigen-transformed) cell expressing EGFP-UL76 for HCS assay was assessed the ED
50 using both MTT and high-content nuclear count measurements (
Table S1). The ED
50 values for respective compounds were as follows: compound
1, >50 μg/mL, >50 μg/mL, >50 μg/mL, >25 μg/mL, and 6.14 μg/mL; stolonidiol (
2), 3.9 μg/mL, >50 μg/mL, 0.6 μg/mL, >25 μg/mL, and >25 μg/mL; stolonidiol-17-acetate (
3), >50 μg/mL, >50 μg/mL, >50 μg/mL, >25 μg/mL, and 19.56 μg/mL; clavinflol B (
4), >50 μg/mL, >50 μg/mL, >50 μg/mL, >25 μg/mL, and 21.33 μg/mL; compound
5, >50 μg/mL, 3.2 μg/mL, 4.6 μg/mL, 12.28 μg/mL, and 12.13 μg/mL; compound
6, 5.3 μg/mL, >50 μg/mL, 4.8 μg/mL, >25 μg/mL, and 10.92 μg/mL. Both the MTT assay and high-content nucleus counts were performed to assess the ED
50 values of HEK293T cells expressing EGFP-UL76 for bortezomib which were 11.95 nM and 24.29 nM and for MG132 were 1.18 μM and 1.91 μM, respectively. Clavinflol B (
4) showed moderate cytotoxicity in previous reports, which was consistent with our data (
Table S1) [
6].
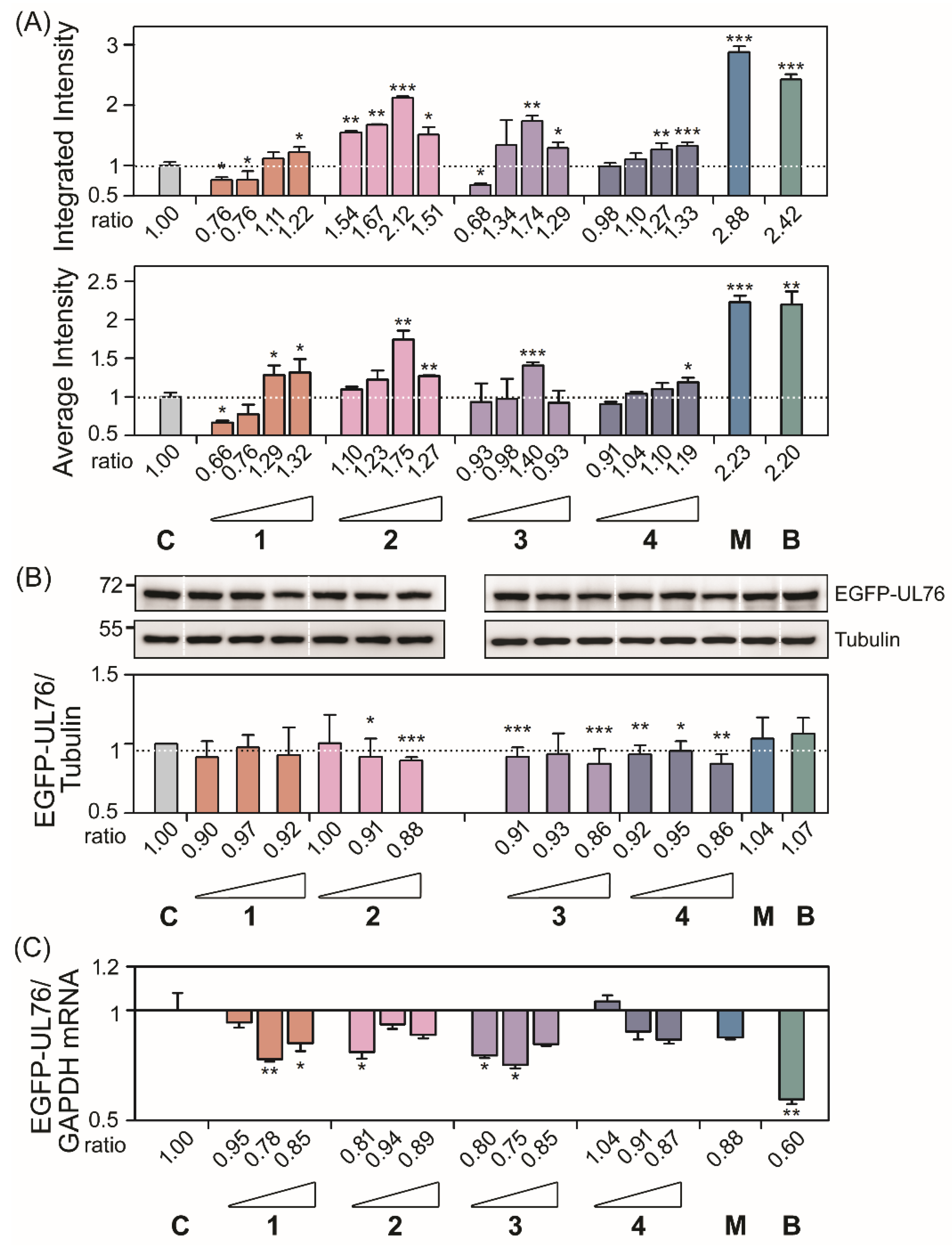
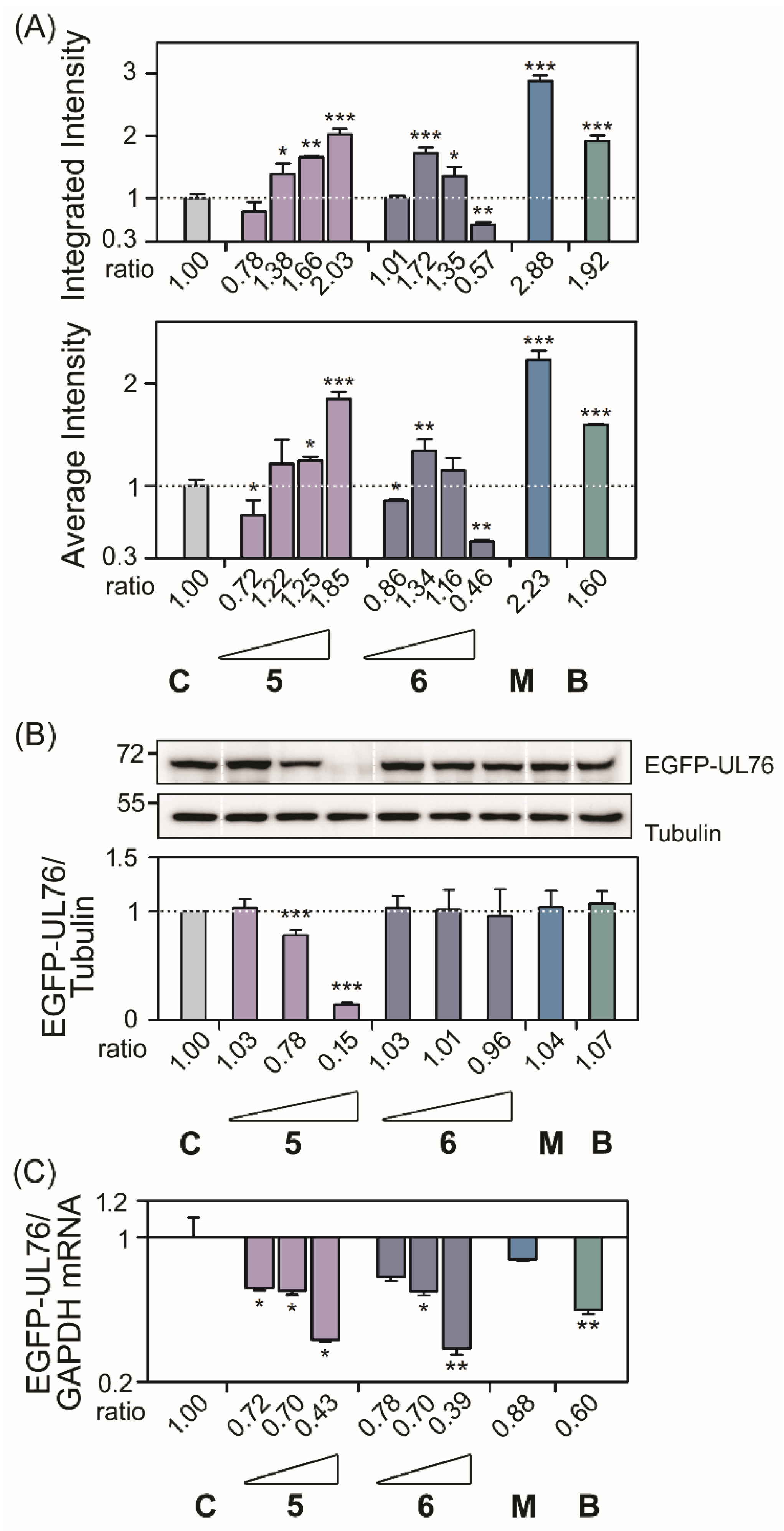
Following the HCS assay, the high-content EGFP-UL76 aggresome measurements integrated intensity and average intensity per cell were analyzed and the relative ratios were obtained by normalization to the control. For the ratio of EGFP-UL76 aggresome integrated intensity per cell (
Figure 2A and
Figure 3A, top panels), the highest ratios for compounds
1,
2,
3,
4,
5, and
6 were 1.22 (
p = 0.0390), 2.12 (
p < 0.0010), 1.74 (
p = 0.0020), 1.33 (
p < 0.0010), 2.03 (
p < 0.0010), and 1.72 (
p < 0.0010), respectively. The highest ratios of average intensity per cell presented for compounds
1,
2,
3,
4,
5, and
6 were 1.32 (
p = 0.0371), 1.75 (
p = 0.0021), 1.40 (
p < 0.0010), 1.19 (
p = 0.0117), 1.85 (
p < 0.0010), and 1.34 (
p = 0.0089), respectively (
Figure 2A and
Figure 3A, bottom panels). Furthermore, all these increases in ratios achieved statistical significance.
Consequently after the assay procedure, we performed Western blotting analysis and q-PCR experiments to examine the levels of EGFP-UL76 protein and mRNA transcript under the same experimental conditions (
Figure 2B,C and
Figure 3B,C). In these two experiments, cells treated with bortezomib 25 nM and MG132 1 μM were used in parallel as proteasome inhibitory controls. We obtained similar results that the ratios of EGFP-UL76/tubulin under treatment with bortezomib and MG132 showed no difference from the control level, which was consistent with a previous report [
4].
Compound
1 did not affect protein ratios of EGFP-UL76/tubulin proteins at 1, 5, and 25 μg/mL of any kind. However, for cells treated with compound
2 at 1, 5, and 25 μg/mL, the ratios were 1.00 (
p = 0.9466), 0.91 (
p = 0.0359), and 0.88 (
p < 0.0010), respectively (
Figure 2B and
Figure 3B). Nevertheless, the cytotoxic ED
50 for compound
2 was greater than 25 μg/mL for HEK293T cell in both assays (
Table S1). The protein ratios for compound
3 were 0.91 (
p < 0.0010), 0.96 (
p = 0.1484), and 0.86 (
p < 0.0010), respectively. The protein ratios for compound
4 were 0.92 (
p = 0.0045), 0.95 (
p = 0.0420), and 0.86 (
p = 0.0014), respectively. However, HEK293T cells treated with compound
3 and
4 at 25 μg/mL exhibited a lighter toxicity with ED
50 values of 19.56 and 21.33 μg/mL, respectively. Compound
5 exhibited significant reduction at 5 and 25 μg/mL; the ratios were 0.78 (
p < 0.0010) and 0.15 (
p < 0.0010), respectively, which is consistent with cytotoxicity ED
50 values (
Table S1). Compound
6 did not affect protein ratios at any tested concentrations. The results from both the quantitative PCR and Western blotting analyses revealed that in the tested compound treatment neither the protein nor the mRNA ratios for EGFP-UL76/GADPH were elevated (
Figure 2C and
Figure 3C). Taking all these results in account, we suggested that the increase in EGFP-UL76 high-content measurement was likely due to the modulation of protein conformation.
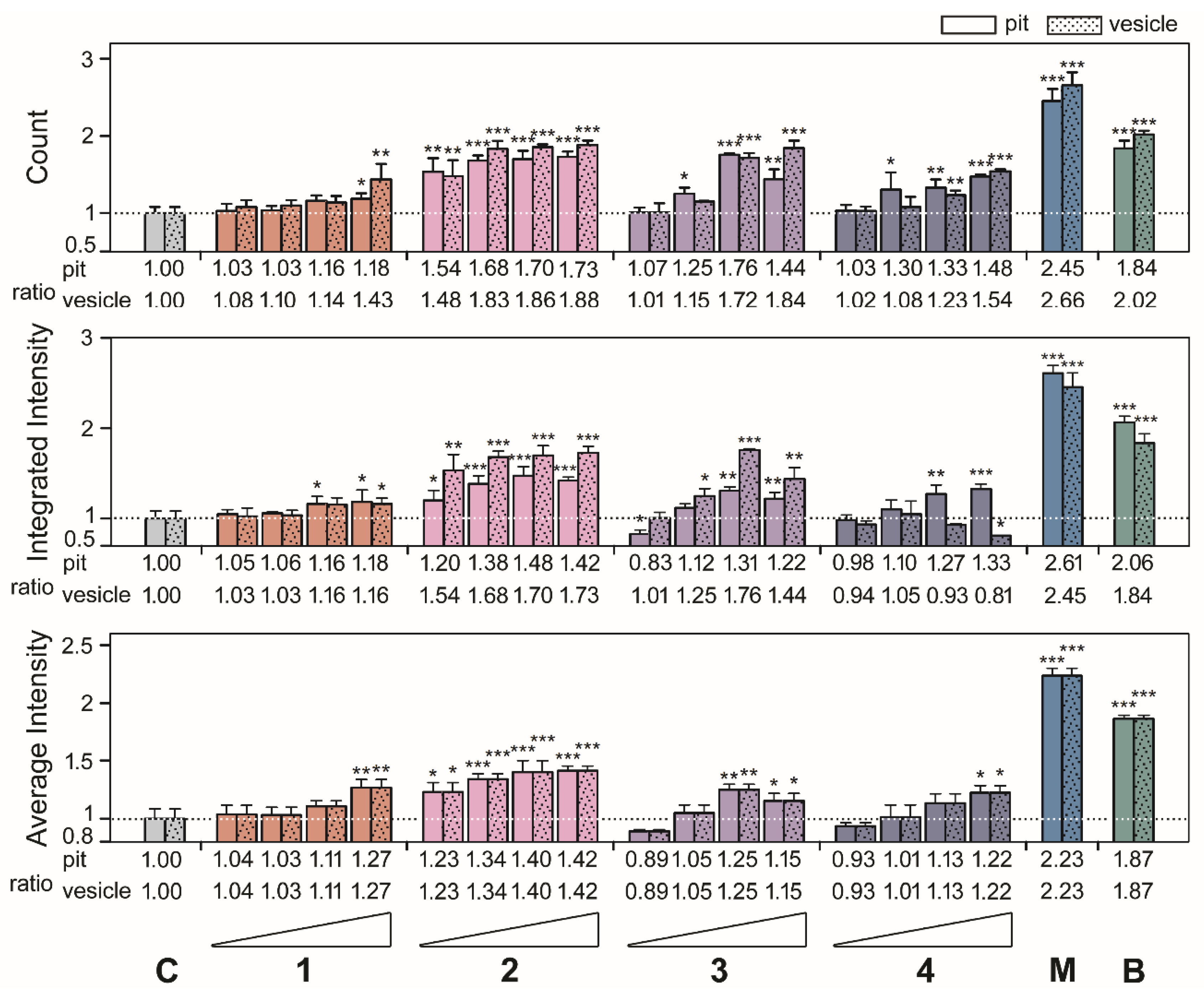
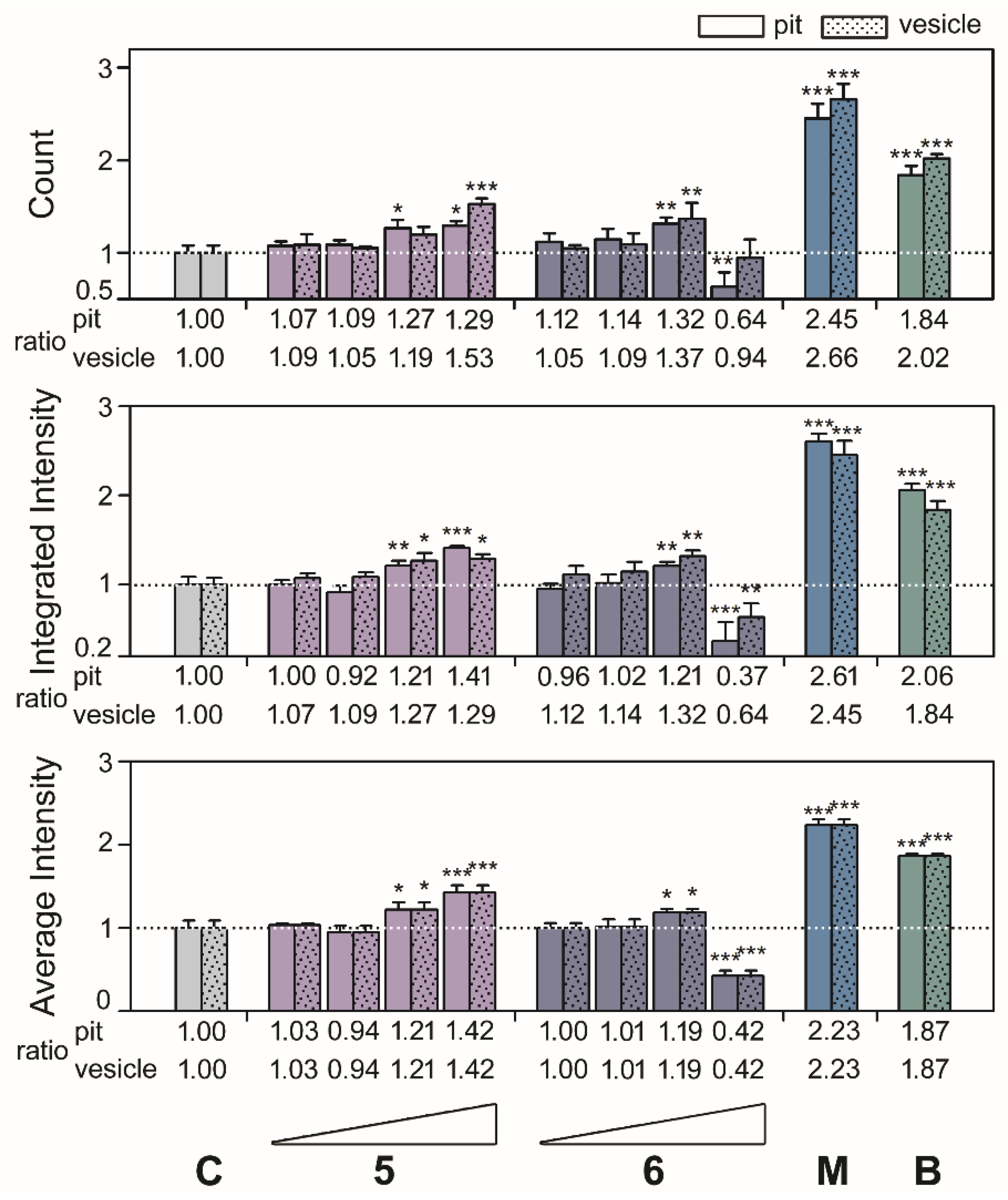
After the results, we investigated the phenotypic size of aggresomes and analyzed the high-content data from
Figure 2 and
Figure 3 by diameter with methods described previously [
4]. As shown in the top panels of
Figure 4 and
Figure 5, compounds
1,
2,
3,
4,
5, and
6 exhibited the highest ratio increases for count, which were as follows: for pit aggresomes: 1.18 (
p = 0.0331), 1.73 (
p < 0.0010), 1.76 (
p < 0.0010), 1.48 (
p < 0.0010), 1.29 (
p = 0.0236), and 1.32 (
p = 0.0062), respectively; for vesicle aggresomes, 1.43 (
p = 0.0028), 1.88 (
p < 0.0010), 1.84 (
p < 0.0010), 1.54 (
p < 0.0010), 1.53 (
p < 0.0010), and 1.37 (
p = 0.0071), respectively. Similar profiles were observed for the ratios of integrated intensity per cell (
Figure 4 and
Figure 5, middle panels). Compounds
1,
2,
3,
4,
5, and
6 showed the highest ratio increases which were as follows: for pit aggresomes, 1.18 (
p = 0.0130), 1.48 (
p < 0.0010), 1.31 (
p = 0.0055), 1.33 (
p < 0.0010), 1.41 (
p < 0.0010), and 1.21 (
p = 0.0097), respectively; for vesicle aggresomes, 1.16 (
p = 0.0220), 1.73 (
p < 0.0010), 1.76 (
p < 0.0010), 1.05 (
p = 0.5352), 1.29 (
p = 0.0236), and 1.32 (
p =0.0063), respectively. The ratios of average intensity per cell were the same for pit and vesicle aggresomes (
Figure 4 and
Figure 5, bottom panels). For compounds
1,
2,
3,
4,
5, and
6 showed the highest increases in ratios observed which were as follows: 1.27 (
p = 0.0099), 1.42 (
p < 0.0010), 1.25 (
p = 0.0034), 1.22 (
p = 0.0628), 1.42 (
p < 0.0010), and 1.19 (
p = 0.0135), respectively.