Polyketides from the Mangrove-Derived Endophytic Fungus Cladosporium cladosporioides
Abstract
1. Introduction
2. Results and Discussion
2.1. Structure Elucidation of the New Compounds
2.2. Biological Activities of the Isolated Compounds
3. Experimental Section
3.1. General Experimental Procedures
3.2. Fungal Material
3.3. Fermentation
3.4. Extraction and Isolation
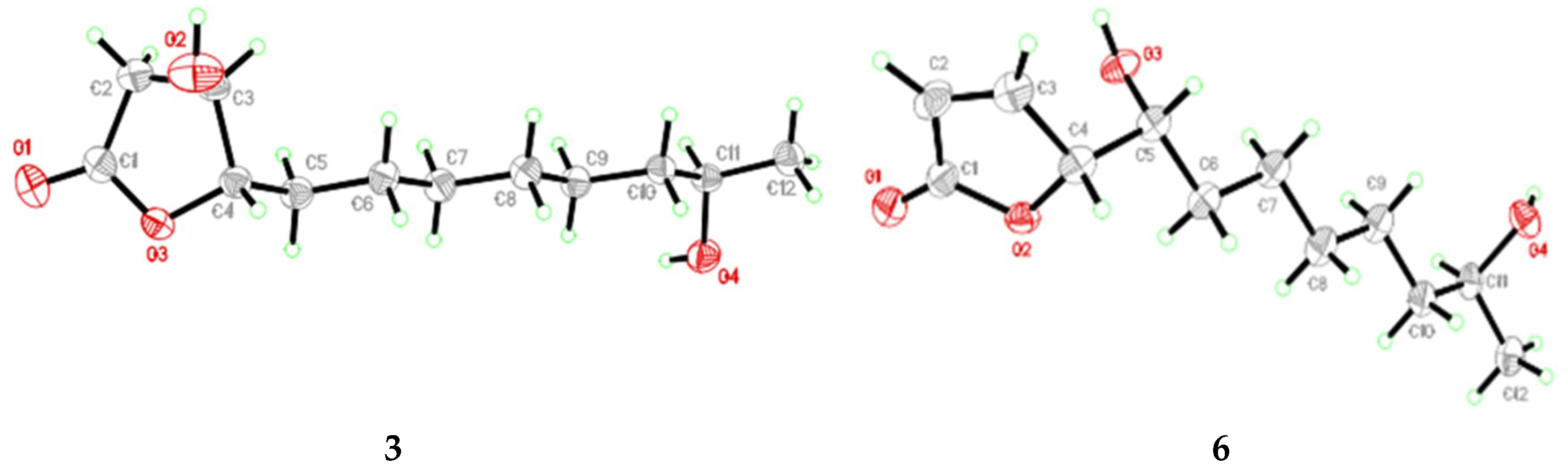
3.5. X-Ray Crystallographic Analysis of Compounds 3 and 6
3.6. Acetylation of Compounds 3 and 4
3.7. Antimicrobial Assay
3.8. Enzyme inhibitory Assay
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Bensch, K.; Groenewald, J.Z.; Braun, U.; Dijksterhuis, J.; de Jesus Yáñez-Morales, M.; Crous, P.W. Common but different: The expanding realm of Cladosporium. Stud. Mycol. 2015, 82, 23–74. [Google Scholar] [CrossRef]
- Imhoff, J.F. Natural products from marine fungi—still an underrepresented resource. Mar. Drugs 2016, 14, 19. [Google Scholar] [CrossRef] [PubMed]
- Jacyno, J.M.; Harwood, J.S.; Cutler, H.G.; Lee, M.K. Isocladosporin, a biologically active isomer of cladosporin from Cladosporium cladosporioides. J. Nat. Prod. 1993, 56, 1397–1401. [Google Scholar] [CrossRef]
- Shigemori, H.; Kasai, Y.; Komatsu, K.; Tsuda, M.; Mikami, Y.; Kobayashi, J. Sporiolides A and B, new cytotoxic twelve-membered macrolides from a marine-derived fungus Cladosporium species. Mar. Drugs 2004, 2, 164–169. [Google Scholar] [CrossRef]
- Gu, B.B.; Zhang, Y.Y.; Ding, L.J.; He, S.; Wu, B.; Dong, J.D.; Zhu, P.; Chen, J.J.; Zhang, J.R.; Yan, X.J. Preparative separation of sulfur-containing diketopiperazines from marine fungus Cladosporium sp. using high-speed counter-current chromatography in stepwise elution mode. Mar. Drugs 2015, 13, 354–365. [Google Scholar] [CrossRef] [PubMed]
- Peng, J.X.; Lin, T.; Wang, W.; Xin, Z.H.; Zhu, T.J.; Gu, Q.Q.; Li, D.H. Antiviral alkaloids produced by the mangrove-derived fungus Cladosporium sp. PJX-41. J. Nat. Prod. 2013, 76, 1133–1140. [Google Scholar] [CrossRef]
- Wu, G.W.; Sun, X.H.; Yu, G.H.; Wang, W.; Zhu, T.J.; Gu, Q.Q.; Li, D.H. Cladosins A–E, hybrid polyketides from a deep-sea-derived fungus, Cladosporium sphaerospermum. J. Nat. Prod. 2014, 77, 270–275. [Google Scholar] [CrossRef]
- Sassa, T.; Ooi, T.; Nukina, M.; Ikeda, M.; Kato, N. Structural confirmation of cotylenin A, a novel fusicoccane-diterpene glycoside with potent plant growth-regulating activity from Cladosporium fungus sp. 501-7w. Biosci. Biotechnol. Biochem. 1998, 62, 1815–1818. [Google Scholar] [CrossRef]
- Li, X.D.; Li, X.; Li, X.M.; Xu, G.M.; Liu, Y.; Wang, B.G. 20-Nor-isopimarane epimers produced by Aspergillus wentii SD-310, a fungal strain obtained from deep sea sediment. Mar. Drugs 2018, 16, 440. [Google Scholar] [CrossRef]
- Yang, S.Q.; Li, X.M.; Li, X.; Chi, L.P.; Wang, B.G. Two New Diketomorpholine Derivatives and a New Highly Conjugated Ergostane-Type Steroid from the Marine Algal-Derived Endophytic Fungus Aspergillus alabamensis EN-547. Mar. Drugs 2018, 16, 114. [Google Scholar] [CrossRef]
- Li, H.L.; Li, X.M.; Mándi, A.; Antus, S.; Li, X.; Zhang, P.; Liu, Y.; Kurtán, T.; Wang, B.G. Characterization of cladosporols from the marine algal-derived endophytic fungus Cladosporium cladosporioides EN-399 and configurational revision of the previously reported cladosporol derivatives. J. Org. Chem. 2017, 82, 9946–9954. [Google Scholar] [CrossRef] [PubMed]
- Zhu, M.L.; Gao, H.Q.; Wu, C.M.; Zhu, T.J.; Che, Q.; Gu, Q.Q.; Guo, P.; Li, D.H. Lipid-lowering polyketides from a soft coral-derived fungus Cladosporium sp. TZP29. Bioorg. Med. Chem. Lett. 2015, 25, 3606–3609. [Google Scholar] [CrossRef]
- Smith, C.J.; Abbanat, D.; Bernan, V.S.; Maiese, W.M.; Greenstein, M.; Jompa, J.; Tahir, A.; Ireland, C.M. Novel polyketide metabolites from a species of marine fungi. J. Nat. Prod. 2000, 63, 142–145. [Google Scholar] [CrossRef]
- Franck, X.; Araujo, M.E.V.; Jullian, J.C.; Hocquemiller, R.; Figadère, B. Synthesis and structure determination of iso-cladospolide B. Tetrahedron Lett. 2001, 42, 2801–2803. [Google Scholar] [CrossRef]
- Gesner, S.; Cohen, N.; Ilan, M.; Yarden, O.; Carmeli, S. Pandangolide 1a, a metabolite of the sponge-associated fungus Cladosporium sp., and the absolute stereochemistry of pandangolide 1 and iso-cladospolide B. J. Nat. Prod. 2005, 68, 1350–1353. [Google Scholar] [CrossRef]
- Corey, E.J.; Ulrich, P.; Fitzpatrick, J.M. A stereoselective synthesis of (±)-11-Hydroxy-trans-8-dodecenoic acid lactone, a naturally occurring macrolide from Cephalosporium recifei. J. Am. Chem. Soc. 1976, 98, 222–224. [Google Scholar] [CrossRef]
- Rodphaya, D.; Sekiguchi, J.; Yamada, Y. New macrolides from Penicillium Urticae mutant S11R59. J. Antibiot. 1986, 39, 629–635. [Google Scholar] [CrossRef]
- Sun, P.; Xu, D.X.; Mándi, A.; Kurtán, T.; Li, T.J.; Schulz, B.; Zhang, W. Structure, absolute configuration, and conformational study of 12-membered macrolides from the fungus Dendrodochium sp. associated with the sea cucumber Holothuria nobilis selenka. J. Org. Chem. 2013, 78, 7030–7047. [Google Scholar] [CrossRef]
- Ohtani, I.; Kusumi, T.; Kashman, Y.; Kakisawa, H. High-field FT NMR application of Mosher’s method. The absolute configurations of marine terpenoids. J. Am. Chem. Soc. 1991, 113, 4092–4096. [Google Scholar] [CrossRef]
- Calculator Plugins Were Used for Structure Property Prediction and Calculation, Marvin 5.9.2, 2012, ChemAxon. Available online: http://www.chemaxon.com (accessed on 31 August 2018).
- Frisch, M.J.; Trucks, G.W.; Schlegel, H.B.; Scuseria, G.E.; Robb, M.A.; Cheeseman, J.R.; Scalmani, G.; Barone, V.; Mennucci, B.; Petersson, G.A.; et al. Gaussian 09, Revision, C.01; Gaussian, Inc.: Wallingford, CT, USA, 2010. [Google Scholar]
- Bruhn, T.; Hemberger, Y.; Schaumloffel, A.; Bringmann, G. SpecDis, Version 1.51; University of Wuerzburg: Würzburg, Germany, 2011. [Google Scholar]
- Trost, B.M.; Aponick, A. Palladium-catalyzed asymmetric allylic alkylation of meso- and dl-1,2-divinylethylene carbonate. J. Am. Chem. Soc. 2006, 128, 3931–3933. [Google Scholar] [CrossRef]
- Reddy, C.R.; Rao, N.N.; Sujitha, P.; Kumar, C.G. Protecting group-free syntheses of (4S,5S,11R)- and (4S,5S,11S)-iso-Cladospolide B and their biological evaluation. Synthesis 2012, 44, 1663–1666. [Google Scholar] [CrossRef]
- Crystallographic data of compounds 3 and 6 have been deposited in the Cambridge Crystallographic Data Centre as CCDC 1889697 and 1889726, respectively. 2019. Available online: http://www.ccdc.cam.ac.uk/ data request/cif (accessed on 31 March 2019).
- Sheldrick, G.M. SADABS, Software for Empirical Absorption Correction; University of Gottingen: Gottingen, Germany, 1996. [Google Scholar]
- Sheldrick, G.M. SHELXTL, Structure Determination Software Programs; Bruker Analytical X-ray System Inc.: Madison, WI, USA, 1997. [Google Scholar]
- Sheldrick, G.M. SHELXL-97 and SHELXS-97, Program for X-ray Crystal Structure Solution and Refinement; University of Göttingen: Göttingen, Germany, 1997. [Google Scholar]
- Ellman, G.L.; Courtney, K.D.; Andres Jr, V.; Featherstone, R.M. A new and rapid colorimetric determination of acetylcholinesterase activity. Biochem. Pharmacol. 1961, 7, 88–95. [Google Scholar] [CrossRef]
- Pierce, C.G.; Uppuluri, P.; Tristan, A.R.; Wormley Jr, F.L.; Mowat, E.; Ramage, G.; Lopez-Ribot, J.L. A simple and reproducible 96-well plate-based method for the formation of fungal biofilms and its application to antifungal susceptibility testing. Nat. Protoc. 2008, 3, 1494–1500. [Google Scholar] [CrossRef]
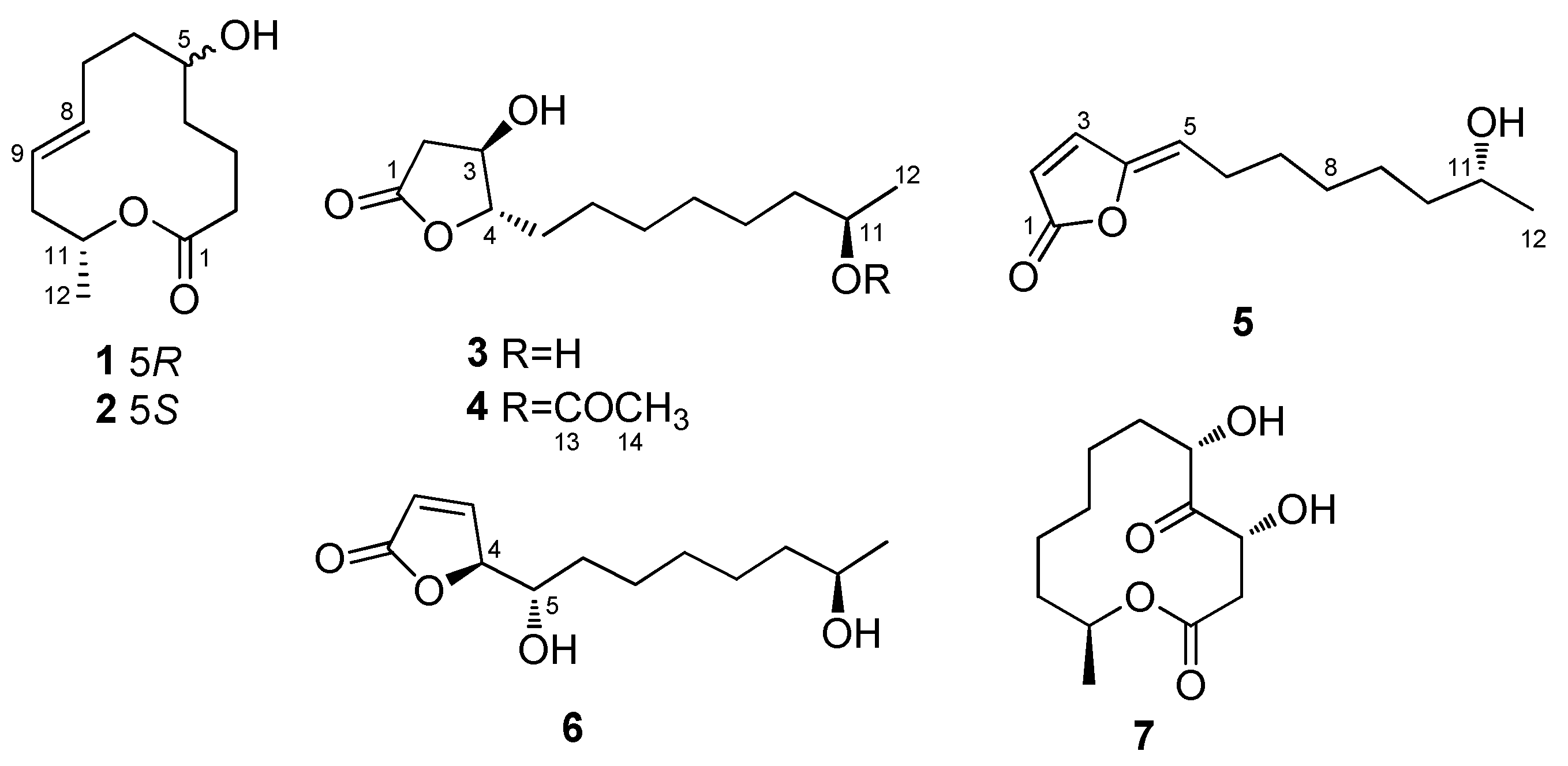
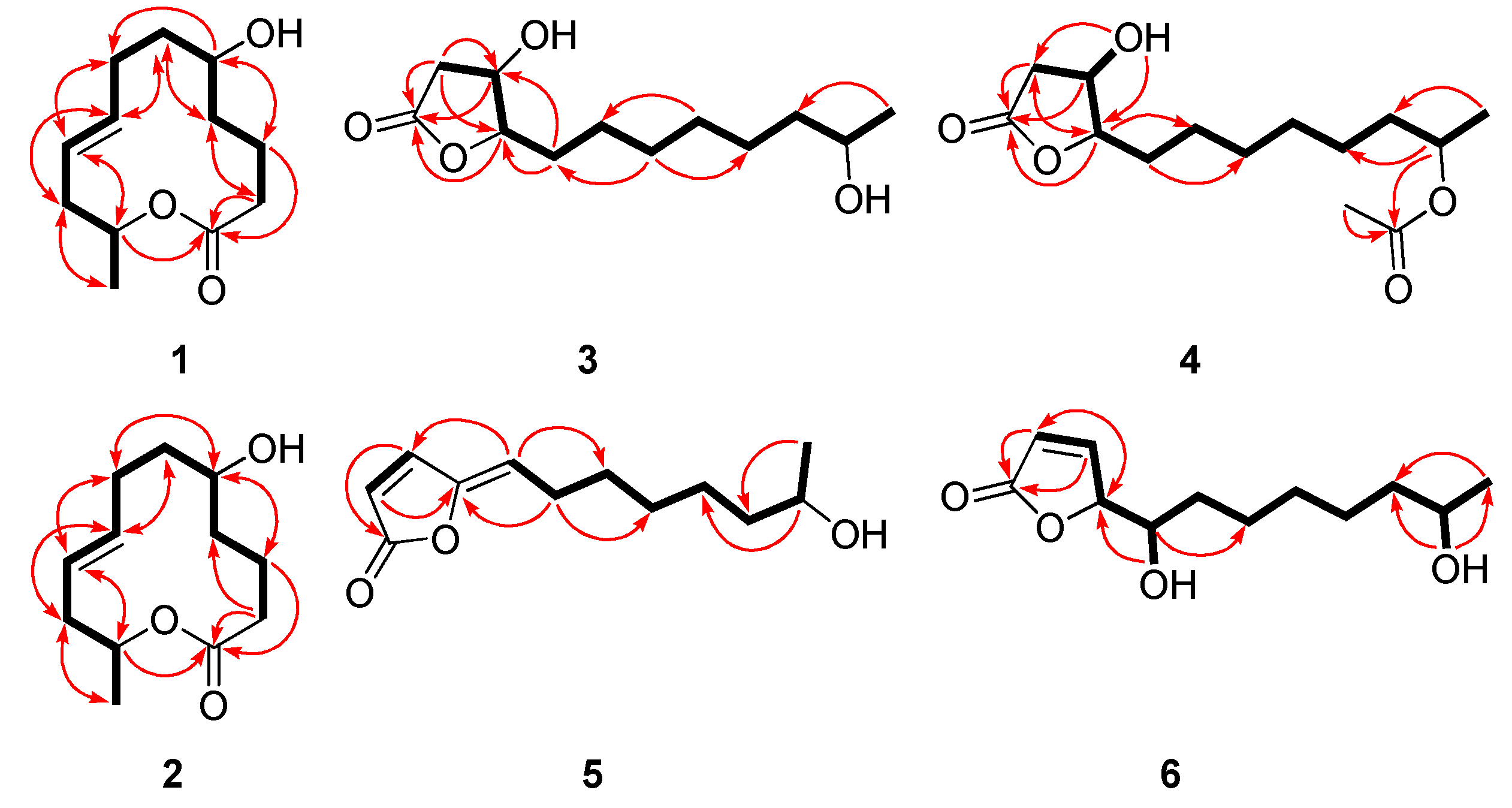
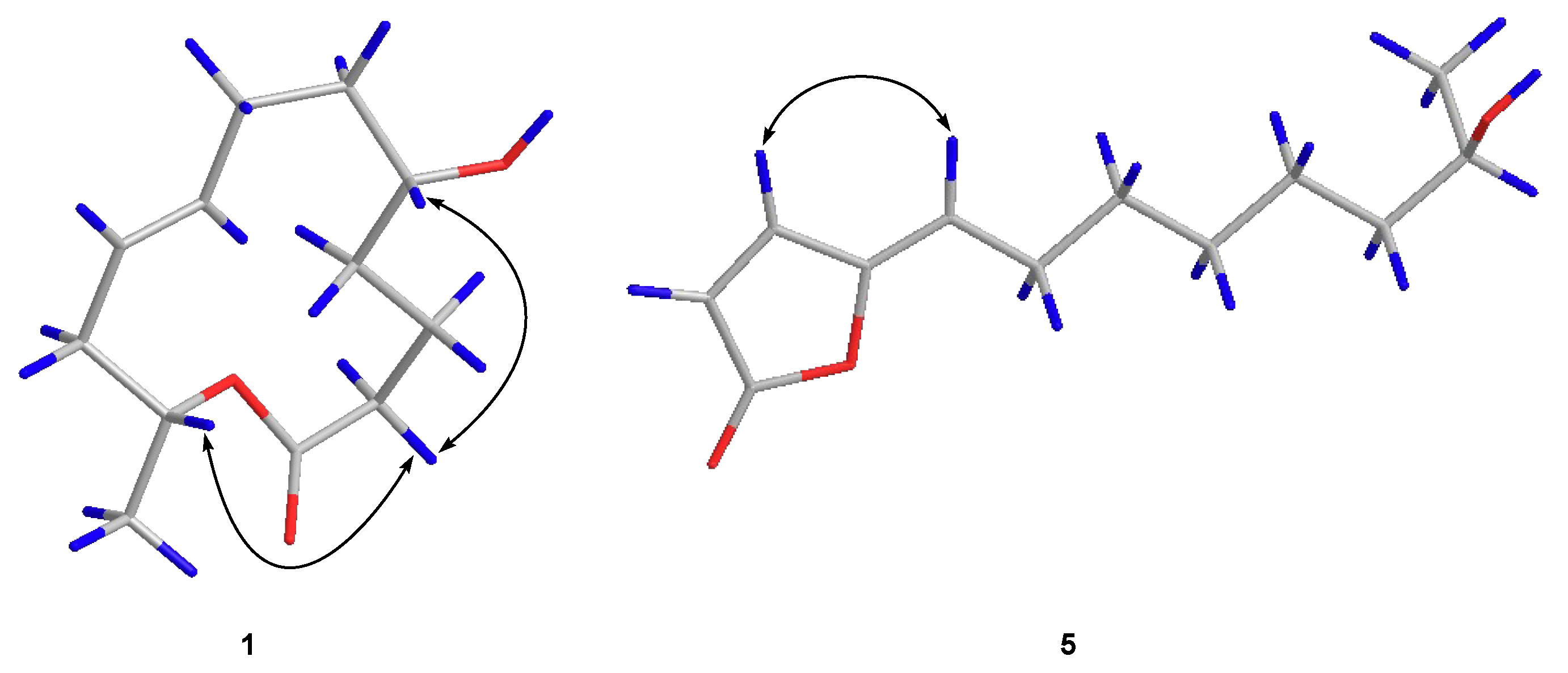
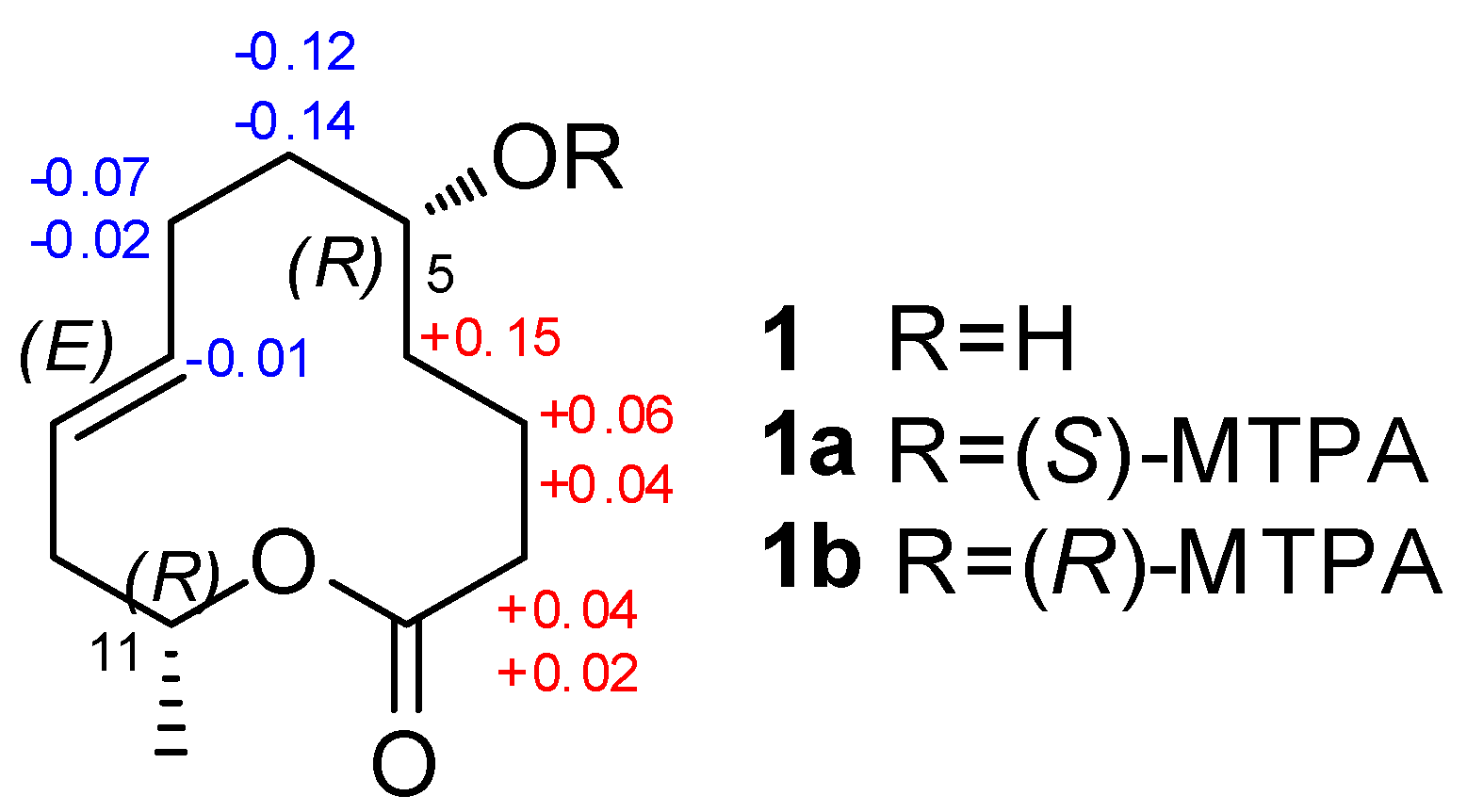
| Pos. | 1 | 2 | 7 | ||||
|---|---|---|---|---|---|---|---|
| a δH (J in Hz) | b δH (J in Hz) | d δC | b δH (J in Hz) | e δC | c δH (J in Hz) | f δC | |
| 1 | 172.3, C | 172.2, C | 174.7, C | ||||
| 2 | α 2.40, ddd (13.0, 5.6, 3.9) β 1.93, ddd (13.1, 11.0, 4.4) | α 2.40, ddd (13.0, 5.6, 3.9) β 1.93, ddd (13.1, 11.0, 4.4) | 35.2, CH2 | α 2.25, m (overlap) β 2.16, m | 31.8, CH2 | α 3.23, dd (18.5, 8.5) β 2.90, dd (18.5, 1.7) | 43.0, CH2 |
| 3 | 1.56, m (overlap) | 1.56, m (overlap) | 18.2, CH2 | α 1.67, m (overlap) β 1.54, m | 19.9, CH2 | 4.72, dd (8.5, 1.7) | 65.7, CH |
| 4 | 1.43, m (overlap) | 1.43, m (overlap) | 32.9, CH2 | α 1.41, m β 0.97, m | 30.0, CH2 | 209.3, C | |
| 5 | 3.51, m | 3.51, m | 70.2, CH | 3.48, m | 65.4, CH | 4.31, d (5.4) | 76.3, CH |
| 6 | α 1.60, m (overlap) β 1.35, m (overlap) | α 1.60, m (overlap) β 1.35, m (overlap) | 32.6, CH2 | α 1.63, m (overlap) β 1.30, m | 33.7, CH2 | α 1.97, m β 1.76, m | 30.7, CH2 |
| 7 | α 2.17, m β 1.72, m | α 2.17, m β 1.72, m | 28.6, CH2 | α 2.05, m β 1.91, m | 27.7, CH2 | α 1.45, m (overlap) β 1.18, m (overlap) | 20.8, CH2 |
| 8 | 5.34, ddd (15.3, 10.7, 2.8) | 5.34, ddd (15.3, 10.7, 2.8) | 135.2, CH | 5.26, m | 132.7, CH | α 1.60, m (overlap) β 1.31, m (overlap) | 26.9, CH2 |
| 9 | 5.20, ddd (15.3, 10.1, 4.1) | 5.20, ddd (15.3, 10.1, 4.1) | 125.6, CH | 5.11, m | 126.7, CH | α 1.51, m (overlap) β 1.10, m (overlap) | 22.7, CH2 |
| 10 | α 2.25, m β 2.06, m | α 2.25, m β 2.06, m | 40.3, CH2 | α 2.29, m (overlap) β 1.99, m (overlap) | 40.3, CH2 | α 1.69, m (overlap) β 1.39, m (overlap) | 33.5, CH2 |
| 11 | 4.84, m | 4.84, m | 68.3, CH | 5.02, m | 67.8, CH | 4.88, m | 75.2, CH |
| 12 | 1.17, d (6.3) | 1.17, d (6.3) | 20.4, CH3 | 1.15, d (6.3) | 20.2, CH3 | 1.25, d (6.2) | 19.3, CH3 |
| 3-OH | 3.19, s | ||||||
| 5-OH | 4.39, brs | 4.39, brs | 4.28, brs | 3.03, s | |||
| Pos. | 3 | 4 | 5 | 6 | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| a δH (J in Hz) | b δH (J in Hz) | d δC | e δC | c δH (J in Hz) | f δC | a δH (J in Hz) | d δC | a δH (J in Hz) | d δC | |
| 1 | 175.5, C | 175.8, C | 175.5, C | 169.7, C | 173.2, C | |||||
| 2 | α 2.85, dd (17.7, 6.4) β 2.25, dd (17.7, 3.2) | α 2.78, dd (18.0, 6.7) β 2.47, dd (18.0, 3.7) | 37.0, CH2 | 37.8, CH2 | α 2.85, dd (17.7, 6.4) β 2.25, dd (17.7, 3.2) | 37.0, CH2 | 6.37, d (5.4) | 118.4, CH | 6.21, d (5.6) | 121.1, CH |
| 3 | 4.09, m | 4.22, m | 70.1, CH | 71.7, CH | 4.09, m | 70.1, CH | 7.80, d (5.4) | 145.1, CH | 7.71, d (5.6) | 156.6, CH |
| 4 | 4.18, m | 4.31, ddd (8.3, 5.3, 3.0) | 87.5, CH | 88.2, CH | 4.18, ddd (8.1, 5.5, 2.5) | 87.5, CH | 149.4, C | 5.04, d (1.5) | 86.2, CH | |
| 5 | α 1.56, m β 1.48, m | 1.57, m | 32.1, CH2 | 33.2, CH2 | α 1.57, m β 1.48, m (overlap) | 32.0, CH2 | 5.53, t (7.9) | 117.1, CH | 3.66, m | 69.4, CH |
| 6 | 1.26, m (overlap) | 1.34, m (overlap) | 25.2, CH2 | 25.7, CH2 | 1.25, m (overlap) | 24.7, CH2 | 2.30, m | 25.8, CH2 | 1.44, m (overlap) | 40.0, CH2 |
| 7 | 1.33, m (overlap) | 1.41, m (overlap) | 29.0, CH2 | 29.5, CH2 | 1.33, m (overlap) | 28.6, CH2 | 1.44, m | 28.3, CH2 | 1.34, m (overlap) | 32.8, CH2 |
| 8 | 1.31, m (overlap) | 1.28, m (overlap) | 28.8, CH2 | 29.3, CH2 | 1.31, m (overlap) | 28.5, CH2 | 1.23, m (overlap) | 28.7, CH2 | 1.29, m (overlap) | 29.1, CH2 |
| 9 | 1.24, m (overlap) | 1.26, m (overlap) | 24.8, CH2 | 25.3, CH2 | 1.24, m (overlap) | 24.6, CH2 | 1.27, m (overlap) | 25.0, CH2 | 1.25, m (overlap) | 25.3, CH2 |
| 10 | 1.36, m (overlap) | 1.45, m (overlap) | 39.0, CH2 | 39.3, CH2 | 1.45, m (overlap) | 35.2, CH2 | 1.32, m (overlap) | 38.9, CH2 | 1.39, m (overlap) | 39.0, CH2 |
| 11 | 3.55, m | 3.75, m | 65.7, CH | 68.3, CH | 4.78, m | 70.1, CH | 3.55, m (overlap) | 65.6, CH | 3.55, m | 65.7, CH |
| 12 | 1.02, d (6.1) | 1.15, d (6.2) | 23.6, CH3 | 23.7, CH3 | 1.15, d (6.3) | 19.7, CH3 | 1.02, d (6.1) | 23.6, CH3 | 1.02, d (6.2) | 23.6, CH3 |
| 13 | 169.9, C | |||||||||
| 14 | 1.97, s | 21.0, CH3 | ||||||||
| 3-OH | 5.49, s | 5.50, d (4.0) | ||||||||
| 5-OH | 4.95, d (6.3) | |||||||||
| 11-OH | 4.27, d (4.2) | 3.41, brs | 4.29, d (4.7) | |||||||
| Strains | Compounds | ||||||
|---|---|---|---|---|---|---|---|
| 1 | 2 | 3 | 4 | 6 | 7 | Positive control | |
| E. colib | – | – | – | 32 | 32 | – | 2.0 |
| S. aureusb | – | – | 8.0 | – | – | 32 | 1.0 |
| E. tardab | – | – | – | – | 32 | – | 0.5 |
| E. ictardab | 32 | – | 16 | – | 16 | 4.0 | 0.5 |
| G. cingulatec | – | 16 | – | 1.0 | 64 | 1.0 | 0.5 |
| B. sorokinianac | – | – | – | 32 | – | – | 0.5 |
| P. aeruginosac | 32 | – | 64 | – | – | 32 | 2.0 |
| F.oxysporum f. sp. Cucumerinum c | – | – | – | 1.0 | – | – | 0.5 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, F.-Z.; Li, X.-M.; Li, X.; Yang, S.-Q.; Meng, L.-H.; Wang, B.-G. Polyketides from the Mangrove-Derived Endophytic Fungus Cladosporium cladosporioides. Mar. Drugs 2019, 17, 296. https://doi.org/10.3390/md17050296
Zhang F-Z, Li X-M, Li X, Yang S-Q, Meng L-H, Wang B-G. Polyketides from the Mangrove-Derived Endophytic Fungus Cladosporium cladosporioides. Marine Drugs. 2019; 17(5):296. https://doi.org/10.3390/md17050296
Chicago/Turabian StyleZhang, Fan-Zhong, Xiao-Ming Li, Xin Li, Sui-Qun Yang, Ling-Hong Meng, and Bin-Gui Wang. 2019. "Polyketides from the Mangrove-Derived Endophytic Fungus Cladosporium cladosporioides" Marine Drugs 17, no. 5: 296. https://doi.org/10.3390/md17050296
APA StyleZhang, F.-Z., Li, X.-M., Li, X., Yang, S.-Q., Meng, L.-H., & Wang, B.-G. (2019). Polyketides from the Mangrove-Derived Endophytic Fungus Cladosporium cladosporioides. Marine Drugs, 17(5), 296. https://doi.org/10.3390/md17050296






