ABC Transporters in Prorocentrum lima and Their Expression Under Different Environmental Conditions Including Okadaic Acid Production
Abstract
1. Introduction
2. Results
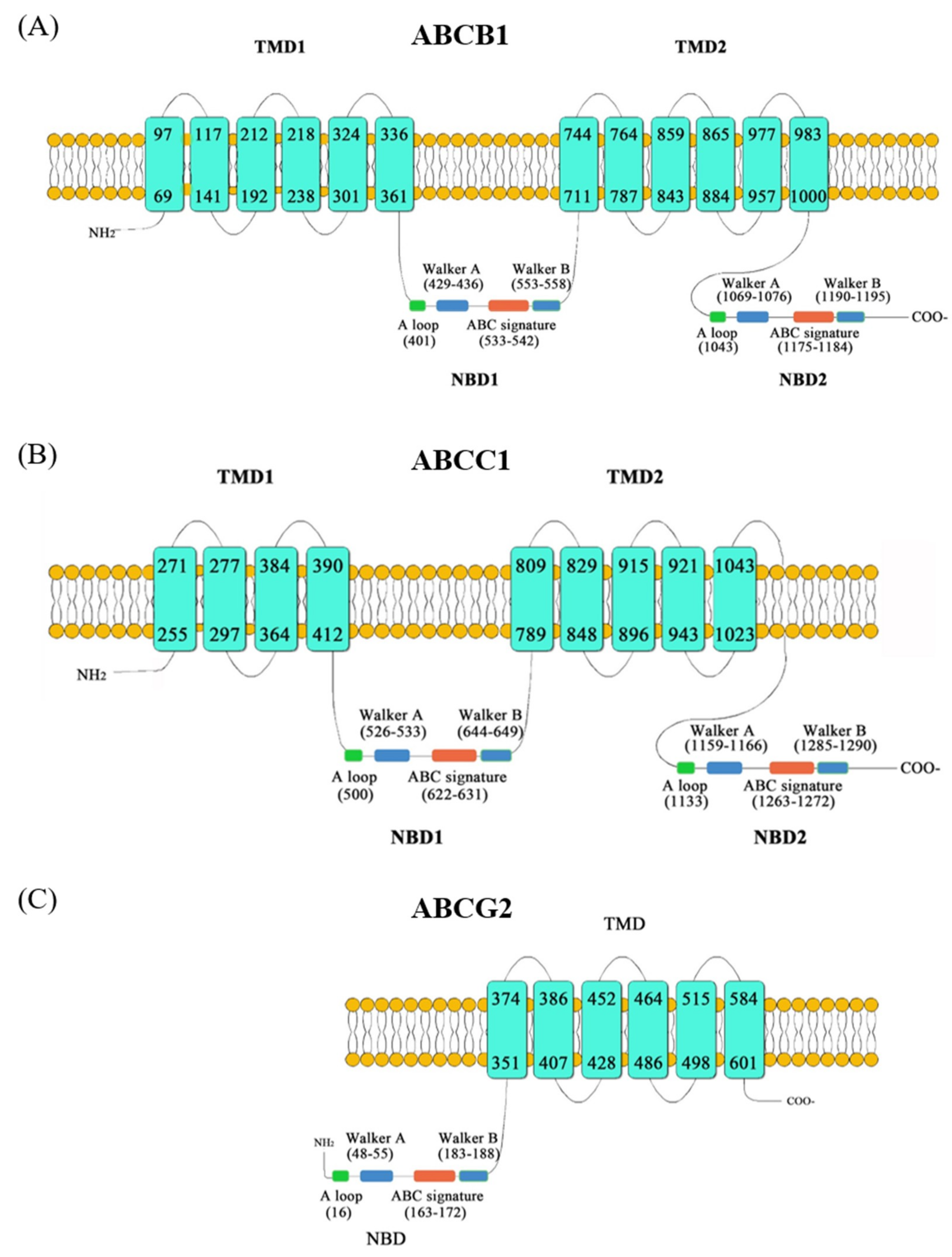
2.1. Sequence Analysis of ABCs Gene
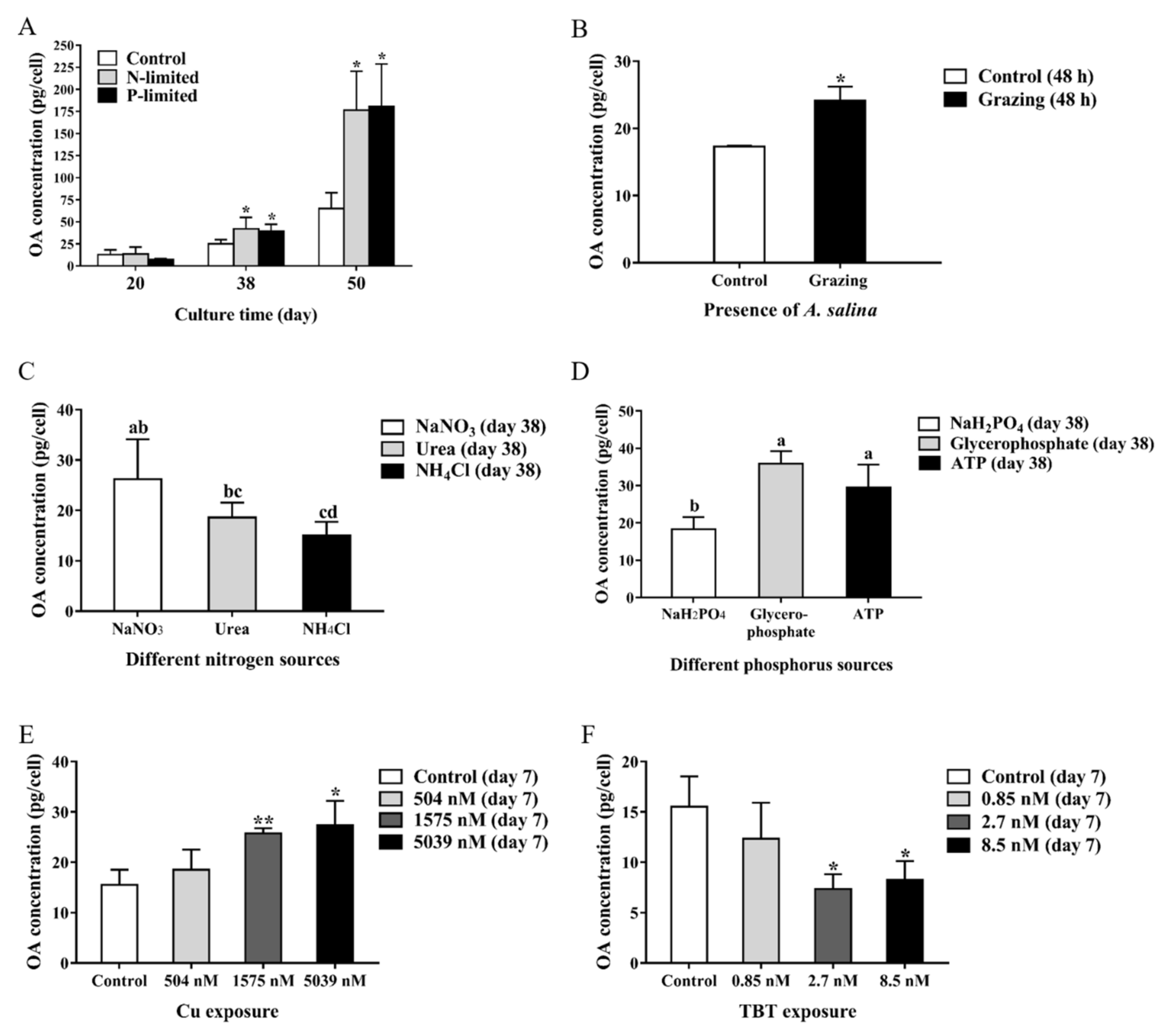
2.2. Content of OA under Different Conditions
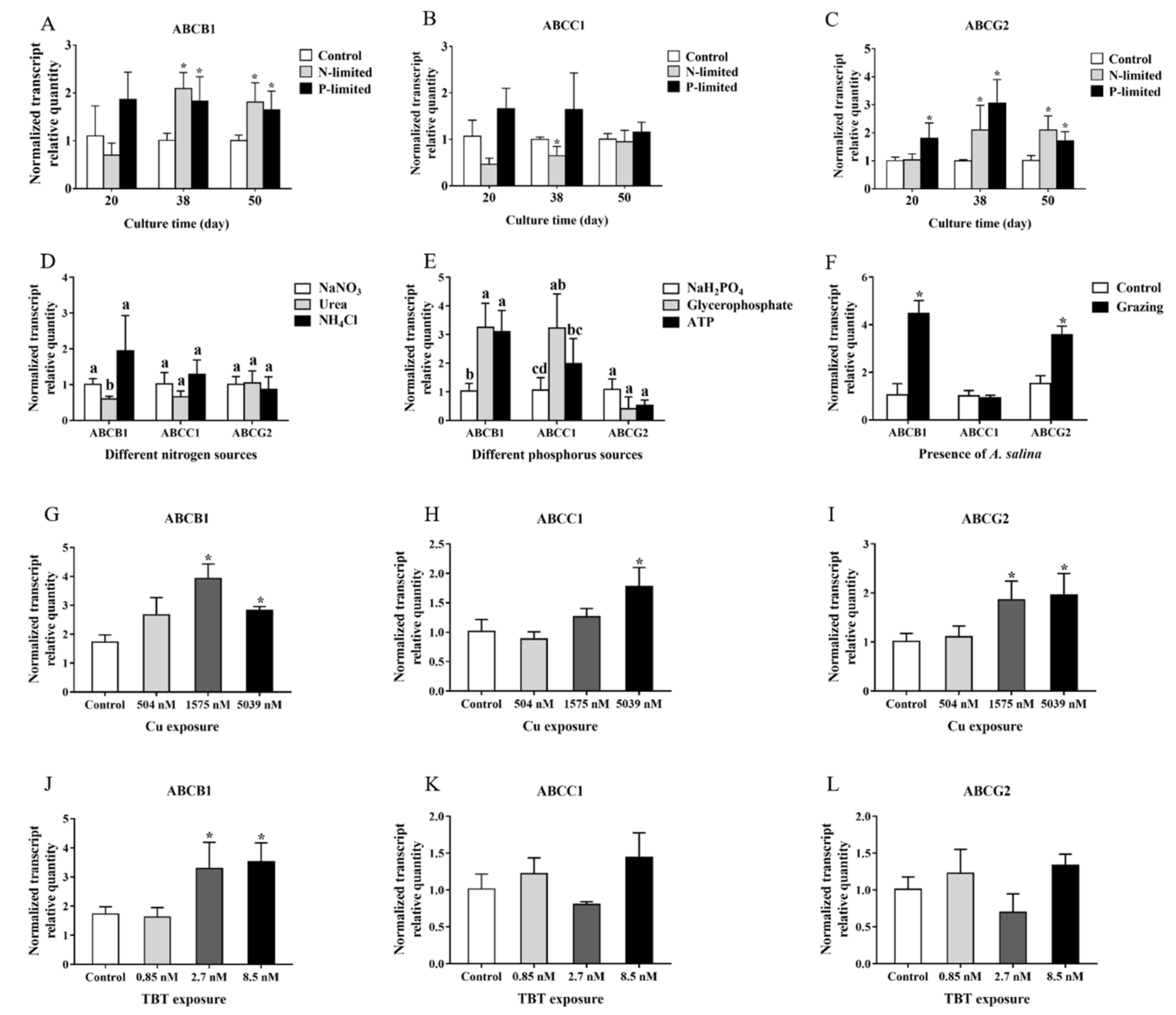
2.3. Transcriptional Responses of ABCB1, ABCC1 and ABCG2 in P. lima under Different Conditions
3. Discussion
4. Materials and Methods
4.1. Algal Culture
4.2. Cloning of ABC Transporter Genes
4.3. Detection of OA in P. lima
4.4. Reverse Transcription-Quantitative PCR (RT-qPCR)
4.5. Expression of ABC Transports in P. lima under Different Conditions
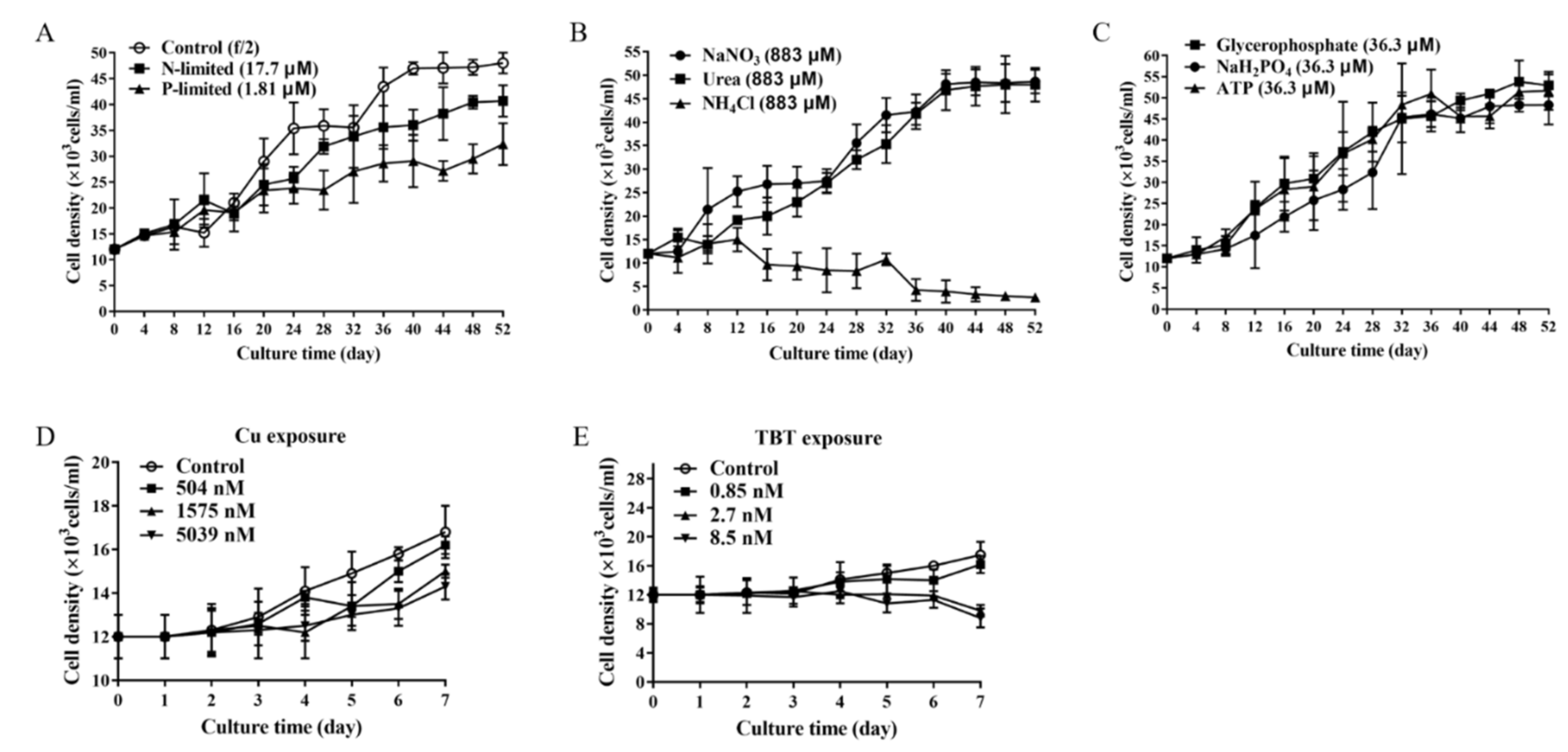
- P. lima under nutrient-limited conditions. P. lima in an initial cell density of 1.2 × 107 cells/L was grown under N-limited conditions (with 17.7 μM NO3−), P-limited condition (1.81 μM PO43−) or nutrient-sufficient conditions (with 883.0 μM NO3−, 36.3 μM PO43−). Trace metals and EDTA were at the levels corresponding to f/2 medium in all cultures. Algal cells were collected on days 20, 38, and 50 for OA extraction, RT-qPCR analysis and morphologic observation, etc.
- P. lima under different nutrients sources (nitrogen or phosphorus) conditions. P. lima in an initial cell density of 1.2 × 107 cells/L was grown under 883 μM of NaNO3, urea or NH4Cl, where other nutrients were the same as the f/2 medium. Algal cells were collected on the 38th day for OA detection and RT-qPCR analysis. Similar experiments were performed by employing NaH2PO4, glycerophosphate or ATP (36.3 μM) as sources of phosphorus.
- P. lima incubated in f/2 medium with Cu2+ or TBT. P. lima in an initial cell density of 1.2 × 107 cells/L was incubated in f/2 medium with Cu2+ (504 nM, 1575 nM and 5039 nM) or TBT (0.85 nM, 2.7 nM and 8.5 nM) as described Couet et al. [43]. As for the control, f/2 medium was employed. After 7 days of culture, algal cells were collected for OA detection and RT-qPCR analysis.
- P. lima exposure to A. salina. P. lima in an initial cell density of 1.2 × 107 cells/L was cultured in f/2 medium with or without 180 of A. salina as described by Dang et al. [44]. The A. salina individuals were hatched from the spawns, which were purchased from Ocean University of China in Qingdao, China. The animals were cultured in 2000 mL of seawater sterilized with a salinity of 27 at 20–25 °C for 7 days, and fed with the green alga Tetraselmis subcordiformis.
4.6. Statistical Analyses
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Rees, D.C.; Johnson, E.; Lewinson, O. ABC transporters: The power to change. Nat. Rev. Mol. Cell Biol. 2009, 10, 218–227. [Google Scholar] [CrossRef] [PubMed]
- Davidson, A.L.; Dassa, E.; Orelle, C.; Chen, J. Structure, function, and evolution of bacterial ATP-binding cassette systems. Microbiol. Mol. Biol. Rev. 2008, 72, 317–364. [Google Scholar] [CrossRef]
- Gottesman, M.M.; Pastan, I. Biochemistry of multidrug resistance mediated by the multidrug transporter. Ann. Rev. Biochem. 1993, 62, 385–427. [Google Scholar] [CrossRef] [PubMed]
- Verrier, P.J.; Bird, D.; Burla, B.; Dassa, E.; Forestier, C.; Geisler, M.; Klein, M.; Kolukisaoglu, U.; Lee, Y.; Martinoia, E.; et al. Plant ABC proteins—A unified nomenclature and updated inventory. Trends Plant Sci. 2008, 13, 151–159. [Google Scholar] [CrossRef] [PubMed]
- Ofori, P.A.; Mizuno, A.; Suzuki, M.; Martinoia, E.; Reuscher, S.; Aoki, K.; Shibata, D.; Otagaki, S.; Matsumoto, S.; Shiratake, K. Genome-wide analysis of ATP binding cassette (ABC) transporters in tomato. PLoS ONE 2018, 13, e0200854. [Google Scholar] [CrossRef] [PubMed]
- Litman, T.; Druley, T.E.; Stein, W.D.; Bates, S.E. From MDR to MXR: New understanding of multidrug resistance systems, their properties and clinical significance. Cell. Mol. Life Sci. 2001, 58, 931–959. [Google Scholar] [CrossRef] [PubMed]
- Stukkens, Y.A.; Bultreys, A.; Grec, S.; Trombik, T.; Vanham, D.; Boutry, M. NpPDR1, a pleiotropic drug resistance-type ATP-binding cassette transporter from Nicotiana plumbaginifolia, plays a major role in plant pathogen defense. Plant Physiol. 2005, 139, 341–352. [Google Scholar] [CrossRef]
- Nuruzzaman, M.; Zhang, R.; Cao, H.Z.; Luo, Z.Y. Plant pleiotropic drug resistance transporters: Transport mechanism, gene expression, and function. J. Integr. Plant. Biol. 2014, 56, 729–740. [Google Scholar] [CrossRef]
- Gaillard, S.; Jacquet, H.; Vavasseur, A.; Leonhardt, N.; Forestier, C. AtMRP6/AtABCC6, an ATP-binding cassette transporter gene expressed during early steps of seedling development and up-regulated by cadmium in Arabidopsis thaliana. BMC Plant Biol. 2008, 8, 22. [Google Scholar] [CrossRef]
- Kim, D.Y.; Bovet, L.; Maeshima, M.; Martinoia, E.; Lee, Y. The ABC transporter AtPDR8 is a cadmium extrusion pump conferring heavy metal resistance. Plant J. 2007, 50, 207–218. [Google Scholar] [CrossRef]
- Schulz, B.; Kolukisaoglu, H.U. Genomics of plant ABC transporters: The alphabet of photosynthetic life forms or just holes in membranes? FEBS Lett. 2006, 580, 1010–1016. [Google Scholar] [CrossRef] [PubMed]
- Merchant, S.S.; Prochnik, S.E.; Vallon, O.; Harris, E.H.; Karpowicz, S.J.; Witman, G.B.; Terry, A.; Salamov, A.; Fritz-laylin, L.K.; Maréchal-Drouard, L.; et al. The Chlamydomonas genome reveals the evolution of key animal and plant functions. Science 2007, 318, 245–250. [Google Scholar] [CrossRef] [PubMed]
- Lane, T.S.; Rempe, C.S.; Davitt, J.; Staton, M.E.; Peng, Y.; Soltis, D.E.; Michael, M.; Deyholos, M.; Leebens-Mack, J.H.; Chase, M.; et al. Diversity of ABC transporter genes across the plant kingdom and their potential utility in biotechnology. BMC Biotechnol. 2016, 16, 47. [Google Scholar] [CrossRef] [PubMed]
- Hanikenne, M.; Motte, P.; Wu, M.C.S.; Wang, T.; Loppes, R.; Matagne, R.F. A mitochondrial half-size ABC transporter is involved in cadmium tolerance in Chlamydomonas reinhardtii. Plant Cell Environ. 2005, 28, 863–873. [Google Scholar] [CrossRef]
- Benning, C.; Hurlock, A.; Warakanont, J. Lipid transport involving chloroplast envelope membranes in plants and algae. FASEB J. 2015, 29, 366. [Google Scholar]
- Scherer, C.; Wiltshire, K.; Bickmeyer, U. Inhibition of multidrug resistance transporters in the diatom Thalassiosira rotula facilitates dye staining. Plant Physiol. Biochem. 2008, 46, 100–103. [Google Scholar] [CrossRef]
- Hou, D.Y.; Liang, J.J.; Zou, C.; Li, H.Y.; Liu, J.S.; Yang, W.D. MRP functional activity and character in the dinoflagellate Prorocentrum lima. J. Appl. Phycol. 2016, 28, 1667–1676. [Google Scholar] [CrossRef]
- Yang, I.; Beszteri, S.; Tillmann, U.; Cembella, A.; John, U. Growth- and nutrient-dependent gene expression in the toxigenic marine dinoflagellate Alexandrium minutum. Harmful Algae 2011, 12, 55–69. [Google Scholar] [CrossRef]
- Wang, D.Z.; Li, C.; Zhang, Y.; Wang, Y.Y.; He, Z.P.; Lin, L.; Hong, H.S. Quantitative proteomic analysis of differentially expressed proteins in the toxicity-lost mutant of Alexandrium catenella (Dinophyceae) in the exponential phase. J. Proteom. 2012, 75, 5564–5577. [Google Scholar] [CrossRef]
- Carvalho, R.N.; Bopp, S.K.; Lettieri, T. Transcriptomics responses in marine diatom Thalassiosira pseudonana exposed to the polycyclic aromatic hydrocarbon benzo[a]pyrene. PLoS ONE 2011, 6, e26985. [Google Scholar] [CrossRef]
- Wang, T.L.; Wu, M. An ATP-binding cassette transporter related to yeast vacuolar ScYCF1 is important for Cd sequestration in Chlamydomonas reinhardtii. Plant Cell Environ. 2006, 29, 1901–1912. [Google Scholar] [CrossRef]
- Tillett, D.; Dittmann, E.; Erhard, M.; von Döhren, H.; Borner, T.; Neilan, B.A. Structural organization of microcystin biosynthesis in Microcystis aeruginosa PCC7806: An integrated peptide-polyketide synthetase system. Chem. Biol. 2000, 7, 753–764. [Google Scholar] [CrossRef]
- Pearson, L.A.; Hisbergues, M.; Borner, T.; Dittmann, E.; Neilan, B.A. Inactivation of an ABC transporter gene, mcyH, results in loss of microcystin production in the cyanobacterium Microcystis aeruginosa PCC 7806. Appl. Environ. Microbiol. 2004, 70, 6370–6378. [Google Scholar] [CrossRef]
- Gong, W.; Browne, J.; Hall, N.; Schruth, D.; Paerl, H.; Marchetti, A. Molecular insights into a dinoflagellate bloom. ISME J. 2017, 11, 439–452. [Google Scholar] [CrossRef] [PubMed]
- Nascimento, S.M.; Purdie, D.A.; Morris, S. Morphology, toxin composition and pigment content of Prorocentrum lima strains isolated from a coastal lagoon in southern UK. Toxicon 2005, 45, 633–649. [Google Scholar] [CrossRef] [PubMed]
- Morton, S.L.; Leighfield, T.A.; Haynes, B.L.; Petitpain, D.L.; Busman, M.A.; Moeller, P.D.R.; Bean, L.; Mcgowan, J.; Hurst, J.W., Jr.; Van Dolah, F.M. Evidence of diarrhetic shellfish poisoning along the coast of Maine. J. Shellfish Res. 1999, 18, 681–686. [Google Scholar]
- Varkitzi, I.; Pagou, K.; Granéli, E.; Hatzianestis, I.; Pyrgaki, C.; Pavlidou, A.; Montesanto, B.; Economou-Amilli, A. Unbalanced N:P ratios and nutrient stress controlling growth and toxin production of the harmful dinoflagellate Prorocentrum lima (Ehrenberg) Dodge. Harmful Algae 2010, 9, 304–311. [Google Scholar] [CrossRef]
- Lee, T.C.; Fong, F.L.; Ho, K.C.; Lee, F.W. The mechanism of diarrhetic shellfish poisoning toxin production in Prorocentrum spp.: Physiological and molecular perspectives. Toxins 2016, 8, 272. [Google Scholar] [CrossRef]
- Amar, M.; Aráoz, R.; Iorga, B.I.; Yasumoto, T.; Servent, D.; Molgó, J. Prorocentrolide-A from cultured Prorocentrum lima dinoflagellates collected in Japan blocks sub-types of nicotinic acetylcholine receptors. Toxins 2018, 10, 97. [Google Scholar] [CrossRef]
- Sugg, L.M.; VanDolah, F.M. No evidence for an allelopathic role of okadaic acid among ciguatera-associated dinoflagellates. J. Phycol. 1999, 35, 93–103. [Google Scholar] [CrossRef]
- Windust, A.J.; Wright, J.L.C.; McLachlan, J.L. The effects of the diarrhetic shellfish poisoning toxins, okadaic acid and dinophysistoxin-1, on the growth of microalgae. Mar. Biol. 1996, 126, 19–25. [Google Scholar] [CrossRef]
- Windust, A.J.; Quilliam, M.A.; Wright, J.L.; McLachlan, J.L. Comparative toxicity of the diarrhetic shellfish poisons, okadaic acid, okadaic acid diol-ester and dinophysistoxin-4, to the diatom Thalassiosira weissflogii. Toxicon 1997, 35, 1591–1603. [Google Scholar] [CrossRef]
- Perreault, F.; Matias, M.S.; Oukarroum, A.; Matias, W.G.; Popovic, R. Okadaic acid inhibits cell growth and photosynthetic electron transport in the alga Dunaliella tertiolecta. Sci. Total Environ. 2012, 414, 198–204. [Google Scholar] [CrossRef]
- Kamimoto, Y.; Terasaka, K.; Hamamoto, M.; Takanashi, K.; Fukuda, S.; Shitan, N.; Sugiyama, A.; Suzuki, H.; Shibata, D.; Wang, B.; et al. Arabidopsis ABCB21 is a facultative auxin importer/exporter regulated by cytoplasmic auxin concentration. Plant Cell Physiol. 2012, 53, 2090–2100. [Google Scholar] [CrossRef]
- Park, J.; Song, W.Y.; Ko, D.; Eom, Y.; Hansen, T.H.; Schiller, M.; Lee, T.G.; Martinoia, E.; Lee, Y. The phytochelatin transporters AtABCC1 and AtABCC2 mediate tolerance to cadmium and mercury. Plant J. 2012, 69, 278–288. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Fritz, L. Okadaic acid antibody localizes to chloroplasts in the DSP-toxin-producing dinoflagellates Prorocentrum lima and Prorocentrum maculosum. Phycologia 1994, 33, 455–461. [Google Scholar] [CrossRef]
- Barbier, M.; Amzil, Z.; Mondeguer, F.; Bhaud, Y.; Soyer-Gobillard, M.O.; Lassus, P. Okadaic acid and PP2A cellular immunolocalization in Prorocentrum lima (Dinophyceae). Phycologia 1999, 38, 41–46. [Google Scholar] [CrossRef]
- Lawrence, J.E.; Cembella, A.D. An immunolabeling technique for the detection of diarrhetic shellfish toxins in individual dinoflagellate cells. Phycologia 1999, 38, 60–65. [Google Scholar] [CrossRef]
- Neves, R.A.F.; Fernandes, T.; dos Santos, L.N.; Nascimento, S.M. Toxicity of benthic dinoflagellates on grazing, behavior and survival of the brine shrimp Artemia salina. PLoS ONE 2017, 12, e0175168. [Google Scholar] [CrossRef]
- Mclachlan, J.L.; Marr, J.C.; Conlon-Keily, A.; Adamson, A. Effects of nitrogen concentration and cold temperature on DSP-toxin concentrations in the dinoflagellate Prorocentrum lima (prorocentrales, dinophyceae). Nat. Toxins 1994, 2, 263–270. [Google Scholar] [CrossRef] [PubMed]
- Maldonado, M.T.; Hughes, M.P.; Rue, E.L.; Wells, M.L. The effect of Fe and Cu on growth and domoic acid production by Pseudo-nitzschia multiseries and Pseudo-nitzschia australis. Limnol. Oceanogr. 2002, 47, 515–526. [Google Scholar] [CrossRef]
- Mitrovic, S.M.; Amandi, M.F.; Mckenzie, L.; Furey, A.; James, K.J. Effects of selenium, iron and cobalt addition to growth and yessotoxin production of the toxic marine dinoflagellate Lane Protoceratium reticulatum in culture. J. Exp. Mar. Biol. Ecol. 2004, 313, 337–351. [Google Scholar] [CrossRef]
- Couet, D.; Pringault, O.; Bancon-Montigny, C.; Briant, N.; Elbaz, F.P.; Delpoux, S.; Kefidaly, O.Y.; Hela, B.; Charaf, M.; Hervé, F.; et al. Effects of copper and butyltin compounds on the growth, photosynthetic activity and toxin production of two HAB dinoflagellates: The planktonic Alexandrium catenella and the benthic Ostreopsis cf. ovata. Aquat. Toxicol. 2018, 196, 154–167. [Google Scholar] [CrossRef]
- Dang, L.; Li, Y.; Liu, F.; Zhang, Y.; Yang, W.; Li, H.; Liu, J. Chemical response of the toxic dinoflagellate Karenia mikimotoi against grazing by three species of zooplankton. J. Eukaryot. Microbiol. 2015, 62, 470–480. [Google Scholar] [CrossRef]
- Selander, E.; Cervin, G.; Pavia, H. Effects of nitrate and phosphate on grazer-induced toxin production in “Alexandrium minutum”. Limnol. Oceanogr. 2008, 53, 523–530. [Google Scholar] [CrossRef]
- Yu, C.S.; Cheng, C.W.; Su, W.C.; Chang, K.C.; Huang, S.W.; Hwang, J.K.; Lu, C.H. CELLO2GO: A web server for protein subCELlular LOcalization prediction with functional gene ontology annotation. PLoS ONE 2014, 9, e99368. [Google Scholar] [CrossRef] [PubMed]
- Emanuelsson, O.; Nielsen, H.; Brunak, S.; Heijne, G. Predicting subcellular localization of proteins based on their N-terminal amino acid sequence. J. Mol. Biol. 2000, 300, 1005–1016. [Google Scholar] [CrossRef]
- Chou, K.C.; Shen, H.B. A new method for predicting the subcellular localization of eukaryotic proteins with both single and multiple sites: Euk-mPLoc 2.0. PLoS ONE 2010, 5, e9931. [Google Scholar] [CrossRef]
- Antizar-Ladislao, B. Environmental levels, toxicity and human exposure to tributyltin (TBT)-contaminated marine environment. A review. Environ. Int. 2008, 34, 292–308. [Google Scholar] [CrossRef]
- Fortibuoni, T.; Noventa, S.; Rampazzo, F.; Gion, C.; Formalewicz, M.; Berto, D.; Raicevich, S. Evidence of butyltin biomagnification along the Northern Adriatic food-web (Mediterranean Sea) elucidated by stable isotope ratios. Environ. Sci. Technol. 2013, 47, 3370–3377. [Google Scholar] [CrossRef]
- Hwang, J.U.; Song, W.Y.; Hong, D.; Ko, D.; Yamaoka, Y.; Jang, S.; Yim, S.; Lee, E.; Khare, D.; Kim, K.; Palmgren, M.; et al. Plant ABC transporters enable many unique aspects of a terrestrial plant’s lifestyle. Mol. Plant 2016, 9, 338–355. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.S.; Tiwari, A.K. Multidrug resistance proteins (MRPs/ABCCs) in cancer chemotherapy and genetic diseases. FEBS J. 2011, 278, 3226–3245. [Google Scholar] [CrossRef] [PubMed]
- Pang, K.; Li, Y.; Liu, M.; Meng, Z.; Yu, Y. Inventory and general analysis of the ATP-binding cassette (ABC) gene superfamily in maize (Zea mays L.). Gene 2013, 526, 411–428. [Google Scholar] [CrossRef]
- Yadav, V.; Molina, I.; Ranathunge, K.; Castillo, I.Q.; Rothstein, S.J.; Reed, J.W. ABCG transporters are required for suberin and pollen wall extracellular barriers in Arabidopsis. Plant Cell 2014, 26, 3569–3588. [Google Scholar] [CrossRef] [PubMed]
- Yu, R.L.; Yuang, X.; Zhao, Y.H.; Hu, G.R.; Tu, X.L. Heavy metal pollution in intertidal sediments from Quanzhou Bay. Chin. J. Environ. Sci. 2008, 20, 664–669. [Google Scholar] [CrossRef]
- Ianora, A.; Bentley, M.G.; Caldwell, G.S.; Casotti, R.; Cembella, A.D.; Engström-Öst, J.; Halsband, C.; Sonnenschein, E.; Legrand, C.; Llewellyn, C.A.; et al. The relevance of marine chemical ecology to plankton and ecosystem function: An emerging field. Mar. Drugs 2011, 9, 1625–1648. [Google Scholar] [CrossRef]
- Turner, J.T. Planktonic marine copepods and harmful algae. Harmful Algae 2014, 32, 81–93. [Google Scholar] [CrossRef]
- Bergkvist, J.; Selander, E.; Pavia, H. Induction of toxin production in dinoflagellates: The grazer makes a difference. Oecologia 2008, 156, 147–154. [Google Scholar] [CrossRef]
- Kretzschmar, T.; Burla, B.; Lee, Y.; Martinoia, E.; Nagy, R. Functions of ABC transporters in plants. Essays Biochem. 2011, 50, 145–160. [Google Scholar] [CrossRef]
- Ehlers, A.; These, A.; Hessel, S.; Preiss-Weigert, A.; Lampen, A. Active elimination of the marine biotoxin okadaic acid by P-glycoprotein through an in vitro gastrointestinal barrier. Toxicol. Lett. 2014, 225, 311–317. [Google Scholar] [CrossRef]
- Windust, A.J.; Hu, T.; Wright, J.L.C.; Quilliam, M.A.; McLachlan, J.L. Oxidative metabolism by Thalassiosira weissflogii (Bacillariophyceae) of a diol-ester of okadaic acid, the diarrhetic shellfish poisoning. J. Phycol. 2000, 36, 342–350. [Google Scholar] [CrossRef]
- Lee, M.; Lee, K.; Lee, J.; Noh, E.W.; Lee, Y. AtPDR12 contributes to lead resistance in Arabidopsis. Plant Physiol. 2005, 138, 827–836. [Google Scholar] [CrossRef]
- Bienert, M.D.; Baijot, A.; Boutry, M. ABCG transporters and their role in the biotic stress response. In Plant ABC Transporters. Signaling and Communication in Plants; Geisler, M., Ed.; Springer: Cham, Switzerland, 2014; Volume 22, pp. 131–162. [Google Scholar]
- Hellemans, J.; Mortier, G.; De Paepe, A.; Speleman, F.; Vandesompele, J. qBase relative quantification framework and software for management and automated analysis of real-time quantitative PCR data. Genome Biol. 2007, 8, R19. [Google Scholar] [CrossRef] [PubMed]
| Primer Name | Primer Sequence (5′–3′) | Target |
|---|---|---|
| M13-47 | CGCCAGGGTTTTCCCAGTCACGAC | 5′-RACE |
| RV-M | GAGCGGATAACAATTTCACACAGG | 5′-RACE |
| AAP | CGCGTCGACTAGTACGGGGGGGGGG | 5′-RACE |
| AUAP | CGCGTCGACTAGTAC | 5′-RACE |
| ABCB1-5GSP1 | GCAGAACCGCACGACGACCT | 5′-RACE |
| ABCB1-5GSP2 | GCTCAGACCATGCGCCACAG | 5′-RACE |
| ABCC1-5GSP1 | CTGTCATACAGGGGTTGTGGCTCGCTG | 5′-RACE |
| ABCC1-5GSP2 | GCTCCAGCTTGCCGGTATCAG | 5′-RACE |
| ABCG2-5GSP1 | GCATCCTCTTGTTCCACATACGC | 5′-RACE |
| ABCG2-5GSP2 | GACAGTGTTCGCATAGAAGGTG | 5′-RACE |
| Gene Name | Primer Sequence (5′–3′) | Length (bp) |
|---|---|---|
| ABCB1 | F: ACGTCGGTGAAGGACAACATC | 128 |
| R: CCGACGAAGGTGTTGAACTTC | ||
| ABCC1 | F: ATGTTGACGGCAACGCAGCTC | 121 |
| R: AGAAGCTGGAACGTCCACGTG | ||
| ABCG2 | F: AGCATGGTCCGATTGCCATG | 123 |
| R: TTAGCTGCCCTGGATCACAC | ||
| β-tubulin | F: GTTGCCTCGTTGTAGTAGACG | 106 |
| R: TTTGGGAGGTGATTTCCGACG | ||
| β-actin | F: GCCGACTTCATCTCTGTGTC | 101 |
| R: GGCTACTCCTTCACCACTACG | ||
| 18S | F: CCGACTTAGCAGAAGGGTTG | 104 |
| R: CAGCAGACGCCATACGACTA | ||
| Calm | F: GCCATGAGGGACAGGAACT | 131 |
| R: AAGGAGCTTGGAACCGTGAT | ||
| GAPDH | F: CCCACTCGTTGTCGTACCAG | 105 |
| R: CGGATTTCGTGAGCAACAA |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gu, S.; Xiao, S.-W.; Zheng, J.-W.; Li, H.-Y.; Liu, J.-S.; Yang, W.-D. ABC Transporters in Prorocentrum lima and Their Expression Under Different Environmental Conditions Including Okadaic Acid Production. Mar. Drugs 2019, 17, 259. https://doi.org/10.3390/md17050259
Gu S, Xiao S-W, Zheng J-W, Li H-Y, Liu J-S, Yang W-D. ABC Transporters in Prorocentrum lima and Their Expression Under Different Environmental Conditions Including Okadaic Acid Production. Marine Drugs. 2019; 17(5):259. https://doi.org/10.3390/md17050259
Chicago/Turabian StyleGu, Song, Shao-Wen Xiao, Jian-Wei Zheng, Hong-Ye Li, Jie-Sheng Liu, and Wei-Dong Yang. 2019. "ABC Transporters in Prorocentrum lima and Their Expression Under Different Environmental Conditions Including Okadaic Acid Production" Marine Drugs 17, no. 5: 259. https://doi.org/10.3390/md17050259
APA StyleGu, S., Xiao, S.-W., Zheng, J.-W., Li, H.-Y., Liu, J.-S., & Yang, W.-D. (2019). ABC Transporters in Prorocentrum lima and Their Expression Under Different Environmental Conditions Including Okadaic Acid Production. Marine Drugs, 17(5), 259. https://doi.org/10.3390/md17050259




