Molecular and Morphological Toxicity of Diatom-Derived Hydroxyacid Mixtures to Sea Urchin Paracentrotus lividus Embryos
Abstract
1. Introduction
2. Results
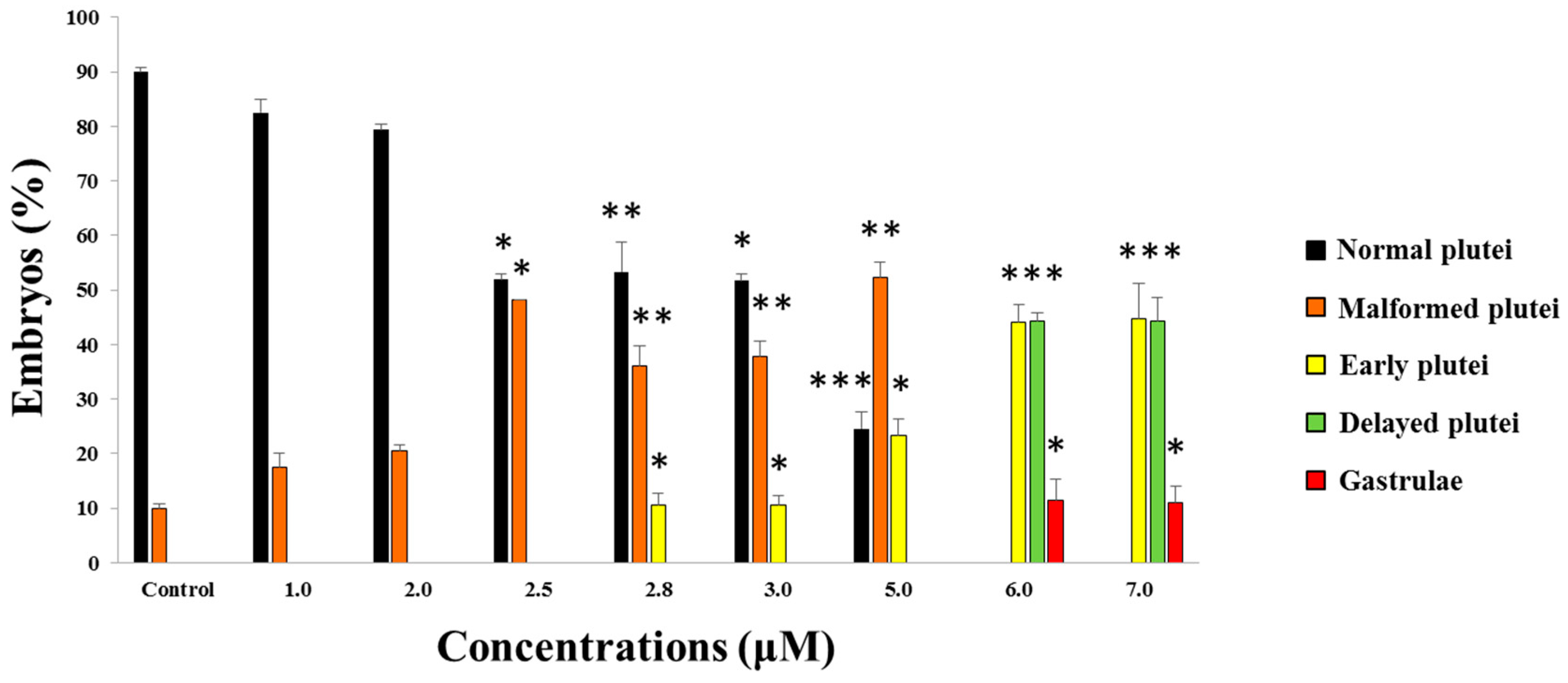
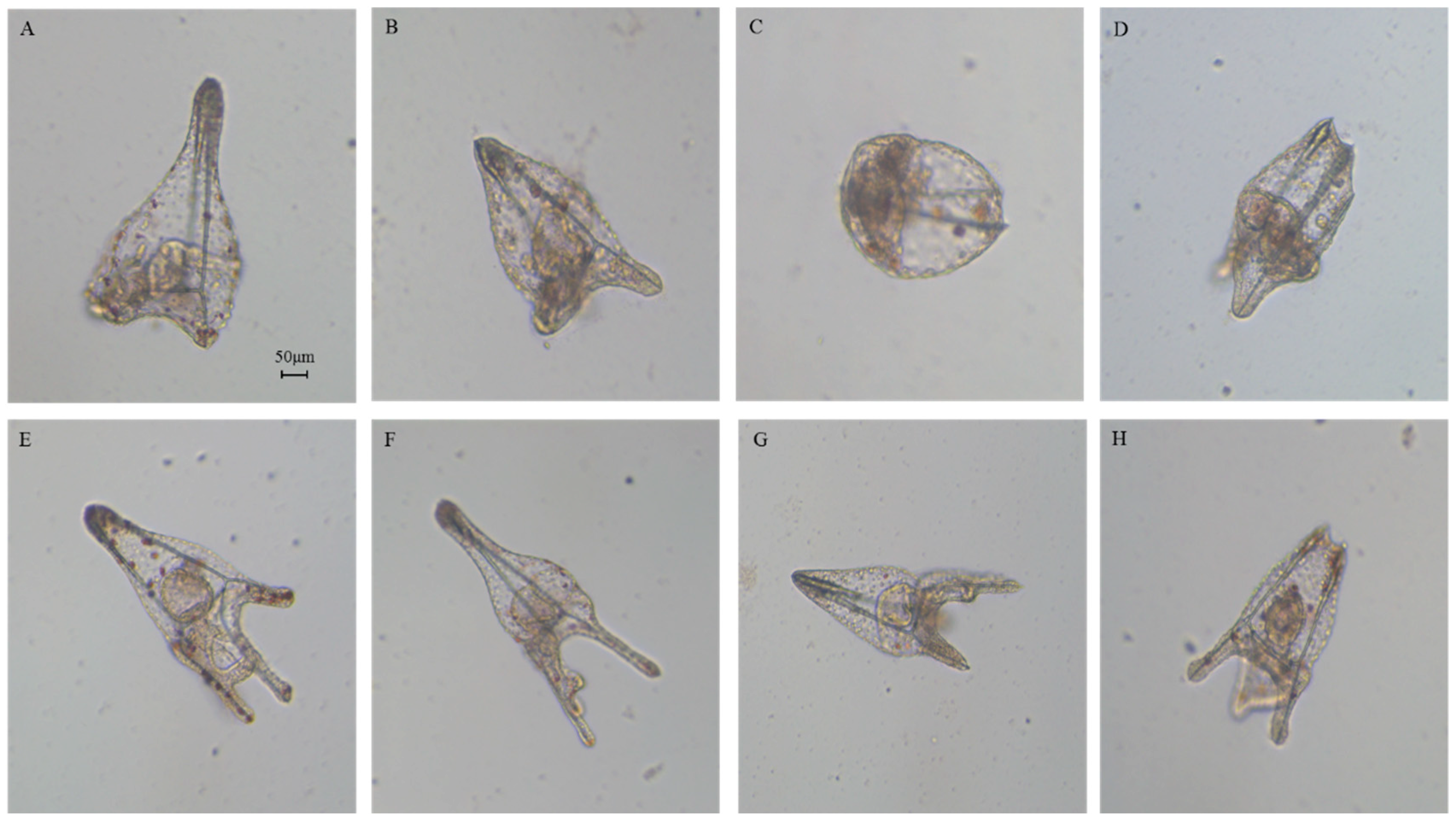
2.1. Effects of 5-, 9-, 11-, and 15-HEPE Mixtures on Sea Urchin Development
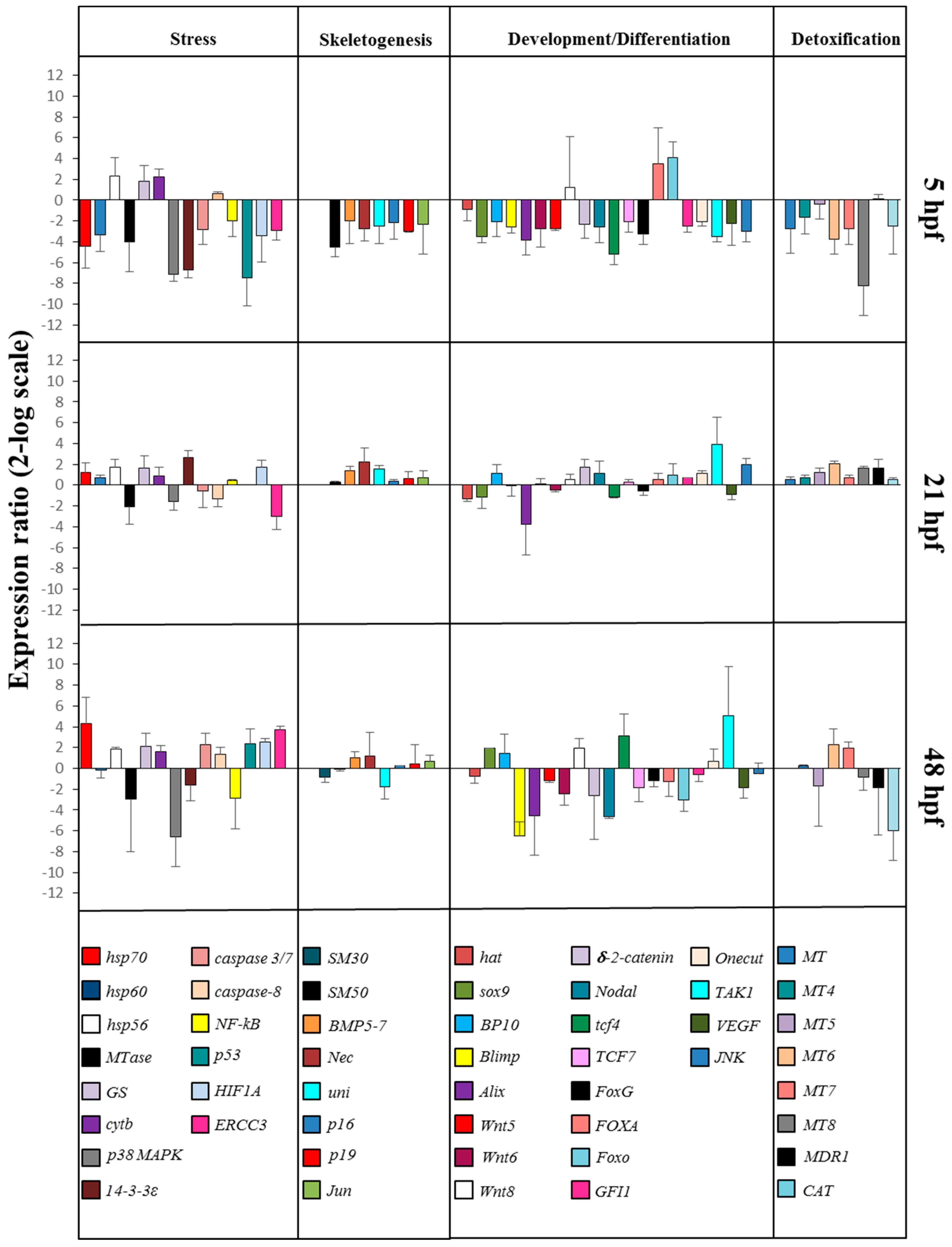
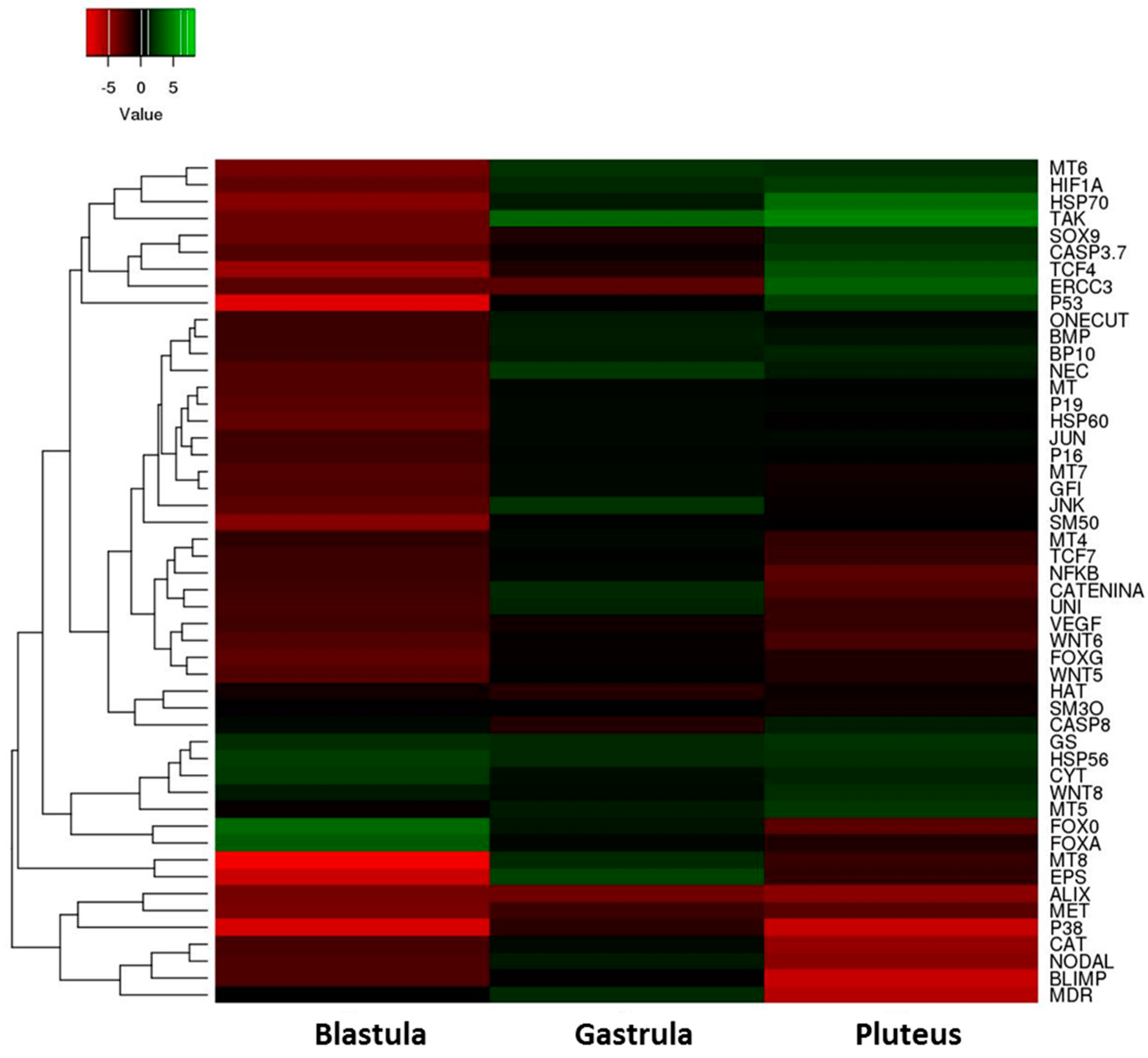
2.2. Gene Response to HEPE Mixtures
2.2.1. Stress Genes
2.2.2. Skeletogenic Genes
2.2.3. Genes Involved in Development/Differentiation
2.2.4. Genes Involved in Detoxification
3. Discussion
4. Materials and Methods
4.1. HEPE Mixtures and Morphological Analysis
4.2. Gene Expression by Real-Time qPCR
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Pohnert, G.; Boland, W. The oxylipin chemistry of attraction and defence in brown algae and diatoms. Nat. Prod. Rep. 2002, 19, 108–122. [Google Scholar] [PubMed]
- Blée, E. Impact of phyto-oxylipins in plant defense. Trends Plant Sci. 2002, 7, 315–322. [Google Scholar] [CrossRef]
- Howe, G.A.; Schimiller, A.L. Oxylipin metabolism in response to stress. Curr. Opin. Plant Biol. 2002, 5, 230–236. [Google Scholar] [CrossRef]
- Pohnert, G. Phospholipase A2 activity triggers the wound-activated chemical defense in diatom Thalassiosira rotula. Plant Physiol. 2002, 129, 103–111. [Google Scholar] [CrossRef] [PubMed]
- D’Ippolito, G.; Cutignano, A.; Briante, R.; Febbraio, F.; Cimmino, G.; Fontana, A. New C16 fatty-acid-based oxylipin pathway in the marine diatom Thalassiosira rotula. Org. Biomol. Chem. 2005, 3, 4065–4070. [Google Scholar] [CrossRef] [PubMed]
- D’Ippolito, G.; Lamari, N.; Montresor, M.; Romano, G.; Cutignano, A.; Gerecht, A.; Cimino, G.; Fontana, A. 15S-lipoxygenase metabolism in the marine diatom Pseudonitzchia delicatissima. New Phytol. 2009, 183, 1064–1071. [Google Scholar] [CrossRef] [PubMed]
- D’Ippolito, G.; Iadicicco, I.; Romano, G.; Fontana, A.; Iadicicco, O. New birth-control aldehydes from the marine diatom Skeletonema costatum: Characterization and biogenesis. Tetrahedron Lett. 2002, 43, 6133–6136. [Google Scholar] [CrossRef]
- D’Ippolito, G.; Iadicicco, I.; Romano, G.; Fontana, A.; Iadicicco, O. Detection of short-chain aldehydes in marine organisms: The diatom Thalassiosira rotula. Tetrahedron Lett. 2002, 43, 6137–6140. [Google Scholar]
- Nanjappa, D.; d’Ippolito, G.; Gallo, C.; Zingone, A.; Fontana, A. Oxylipin diversity in the diatom family Lepto Cilindraceae reveals DHA derivatives in marine diatom. Mar. Drugs 2014, 12, 368–384. [Google Scholar] [CrossRef] [PubMed]
- Fontana, A.; d’Ippolito, G.; Cutignano, A.; Romano, G.; Lamari, N.; Massa Gallucci, A.; Cimino, G.; Miralto, A.; Ianora, A. LOX-induced lipid peroxidation mechanism responsible for the detrimental effect of marine diatoms on zooplankton grazers. ChemBioChem 2007, 8, 1810–1818. [Google Scholar] [CrossRef] [PubMed]
- Cutignano, A.; Lamari, N.; d’Ippolito, G.; Manzo, E.; Cimino, G.; Fontana, A. Lipoxygenase products in marine diatoms: A concise analytical method to explore the functional potential of oxylipins. Am. J. Physiol. Cell Physiol. 2011, 47, 233–243. [Google Scholar] [CrossRef] [PubMed]
- Miralto, A.; Barone, G.; Romano, G.; Poulet, S.A.; Ianora, A.; Russo, G.L.; Buttino, I.; Mazzarella, G.; Laabir, M.; Cabrini, M.; et al. The insidious effect of diatoms on copepod reproduction. Nature 1999, 402, 173–176. [Google Scholar] [CrossRef]
- Ianora, A.; Miralto, A. Toxigenic effects of diatoms on grazers, phytoplankton and other microbes: A review. Ecotoxicology 2010, 19, 493–511. [Google Scholar] [CrossRef] [PubMed]
- Russo, E.; Ianora, A.; Carotenuto, I. Re-shaping marine plankton communities: Effects of diatom oxylipins on copepods and beyong. Mar. Biol. 2019, 166, 9. [Google Scholar] [CrossRef]
- Pinsino, A.; Matranga, V. Sea urchin immune cells as sentinels of environmental stress. Dev. Comp. Immunol. 2015, 49, 198–205. [Google Scholar] [CrossRef] [PubMed]
- Todgham, A.E.; Hofmann, G.E. Transcriptomic response of sea urchin larvae Strongylocentrotus purpuratus to CO2-driven seawater acidification. J. Exp. Biol. 2009, 212, 2579–2594. [Google Scholar] [CrossRef] [PubMed]
- Roepke, T.A.; Snyder, M.J.; Cherr, G.N. Estradiol and endocrine disrupting compounds adversely affect development of sea urchin embryos at environmentally relevant concentrations. Aquat. Toxicol. 2005, 71, 155–173. [Google Scholar] [CrossRef] [PubMed]
- Varrella, S.; Romano, G.; Ianora, A.; Bentley, M.G.; Ruocco, N.; Costantini, M. Molecular response to toxic diatom-derived aldehydes in the sea urchin Paracentrotus lividus. Mar. Drugs 2014, 12, 2089–2113. [Google Scholar] [CrossRef] [PubMed]
- Varrella, S.; Romano, G.; Ruocco, N.; Ianora, A.; Bentley, M.G.; Costantini, M. First morphological and molecular evidence of the negative impact of diatom-derived hydroxyacids on the sea urchin Paracentrotus lividus. Toxicol. Sci. 2016, 151, 419–433. [Google Scholar] [CrossRef] [PubMed]
- Ruocco, N.; Varrella, S.; Romano, G.; Ianora, A.; Bentley, M.G.; Somma, D.; Leonardi, A.; Mellone, S.; Zuppa, A.; Costantini, M. Diatom-derived oxylipins induce cell death in sea urchin embryos activating caspase-8 and caspase 3/7. Aquat. Toxicol. 2016, 176, 128–140. [Google Scholar] [CrossRef] [PubMed]
- Ruocco, N.; Annunziata, C.; Ianora, A.; Libralato, G.; Manfra, L.; Costantini, S.; Costantini, M. Toxicity of diatom-derived polyunsaturated aldehyde mixtures on sea urchin Paracentrotus lividus development. Sci. Rep. 2019, 9, 517. [Google Scholar] [CrossRef] [PubMed]
- Varrella, S.; Romano, G.; Costantini, S.; Ruocco, N.; Ianora, A.; Bentley, M.G.; Costantini, M. Toxic diatom aldehydes affect defence gene networks in sea urchins. PLoS ONE 2016, 11, e0149734. [Google Scholar] [CrossRef] [PubMed]
- Ruocco, N.; Fedele, A.M.; Costantini, S.; Romano, G.; Ianora, A. New inter-correlated genes targeted by diatom-derived polyunsaturated aldehydes in the sea urchin Paracentrotus lividus. Ecotoxicol. Environ. Saf. 2017, 142, 355–362. [Google Scholar] [CrossRef] [PubMed]
- Ianora, A.; Romano, G.; Carotenuto, Y.; Esposito, F.; Roncalli, V.; Buttino, I. Impact of the diatom oxylipin 15S-HEPE on the reproductive success of the copepod Temora stylifera. Hydrobiologia 2011, 666, 265–275. [Google Scholar] [CrossRef]
- Goldstone, J.V.; Hamdoun, A.; Cole, B.J.; Howard-Ashby, M.; Nebert, D.W.; Scally, M.; Dean, M.; Epel, D.; Hahn, M.E.; Stegeman, J.J. The chemical defensome: Environmental sensing and responses genes in the Strongylocentrotus purpuratus genome. Dev. Biol. 2006, 300, 366–384. [Google Scholar] [CrossRef] [PubMed]
- Marrone, V.; Piscopo, M.; Romano, G.; Ianora, A.; Palumbo, A.; Costantini, M. Defensome against toxic diatom aldehydes in the sea urchin Paracentrotus lividus. PLoS ONE 2012, 7, e31750. [Google Scholar] [CrossRef] [PubMed]
- Romano, G.; Miralto, A.; Ianora, A. Teratogenic effects of diatom metabolites on sea urchin Paracentrotus lividus embryos. Mar. Drugs 2010, 8, 950–967. [Google Scholar] [CrossRef] [PubMed]
- Ruocco, N.; Costantini, S.; Zupo, V.; Lauritano, C.; Caramiello, D.; Ianora, A.; Budillon, A.; Romano, G.; Nuzzo, G.; D’Ippolito, G.; et al. Toxigenic effects of two benthic diatoms upon grazing activity of the sea urchin: Morphological, metabolomic and de novo transcriptomic analysis. Sci. Rep. 2018, 8, 5622. [Google Scholar] [CrossRef] [PubMed]
- Ruocco, N.; Costantini, S.; Zupo, V.; Romano, G.; Ianora, A.; Fontana, A.A. High-quality RNA extraction from the sea urchin Paracentrotus lividus embryos. PLoS ONE 2017, 12, e0172171. [Google Scholar] [CrossRef] [PubMed]
- Costa, C.; Karakostis, K.; Zito, F.; Matranga, V. Phylogenetic analysis and expression patterns of p16 and p19 in Paracentrotus lividus embryos. Dev. Genes Evol. 2012, 222, 245–251. [Google Scholar] [CrossRef] [PubMed]
- Pfaffl, M.W. A new mathematical model for relative quantification in realtime RT-PCR. Nucleic Acid Res. 2001, 29, e45. [Google Scholar] [CrossRef] [PubMed]
- Pfaffl, M.W.; Horgan, G.W.; Dempfle, L. Relative expression software tool (REST) for group-wise comparison and statistical analysis of relative expression results in real-time PCR. Nucleic Acid Res. 2002, 30, e36. [Google Scholar] [CrossRef] [PubMed]

© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Albarano, L.; Ruocco, N.; Ianora, A.; Libralato, G.; Manfra, L.; Costantini, M. Molecular and Morphological Toxicity of Diatom-Derived Hydroxyacid Mixtures to Sea Urchin Paracentrotus lividus Embryos. Mar. Drugs 2019, 17, 144. https://doi.org/10.3390/md17030144
Albarano L, Ruocco N, Ianora A, Libralato G, Manfra L, Costantini M. Molecular and Morphological Toxicity of Diatom-Derived Hydroxyacid Mixtures to Sea Urchin Paracentrotus lividus Embryos. Marine Drugs. 2019; 17(3):144. https://doi.org/10.3390/md17030144
Chicago/Turabian StyleAlbarano, Luisa, Nadia Ruocco, Adrianna Ianora, Giovanni Libralato, Loredana Manfra, and Maria Costantini. 2019. "Molecular and Morphological Toxicity of Diatom-Derived Hydroxyacid Mixtures to Sea Urchin Paracentrotus lividus Embryos" Marine Drugs 17, no. 3: 144. https://doi.org/10.3390/md17030144
APA StyleAlbarano, L., Ruocco, N., Ianora, A., Libralato, G., Manfra, L., & Costantini, M. (2019). Molecular and Morphological Toxicity of Diatom-Derived Hydroxyacid Mixtures to Sea Urchin Paracentrotus lividus Embryos. Marine Drugs, 17(3), 144. https://doi.org/10.3390/md17030144









