Unraveling the Detoxification Mechanism of 2,4-Dichlorophenol by Marine-Derived Mesophotic Symbiotic Fungi Isolated from Marine Invertebrates
Abstract
1. Introduction
2. Results and Discussion
2.1. 2,4-DCP Biotransformation Potential of Isolated Fungi
2.2. Identification of 2,4-DCP Metabolites
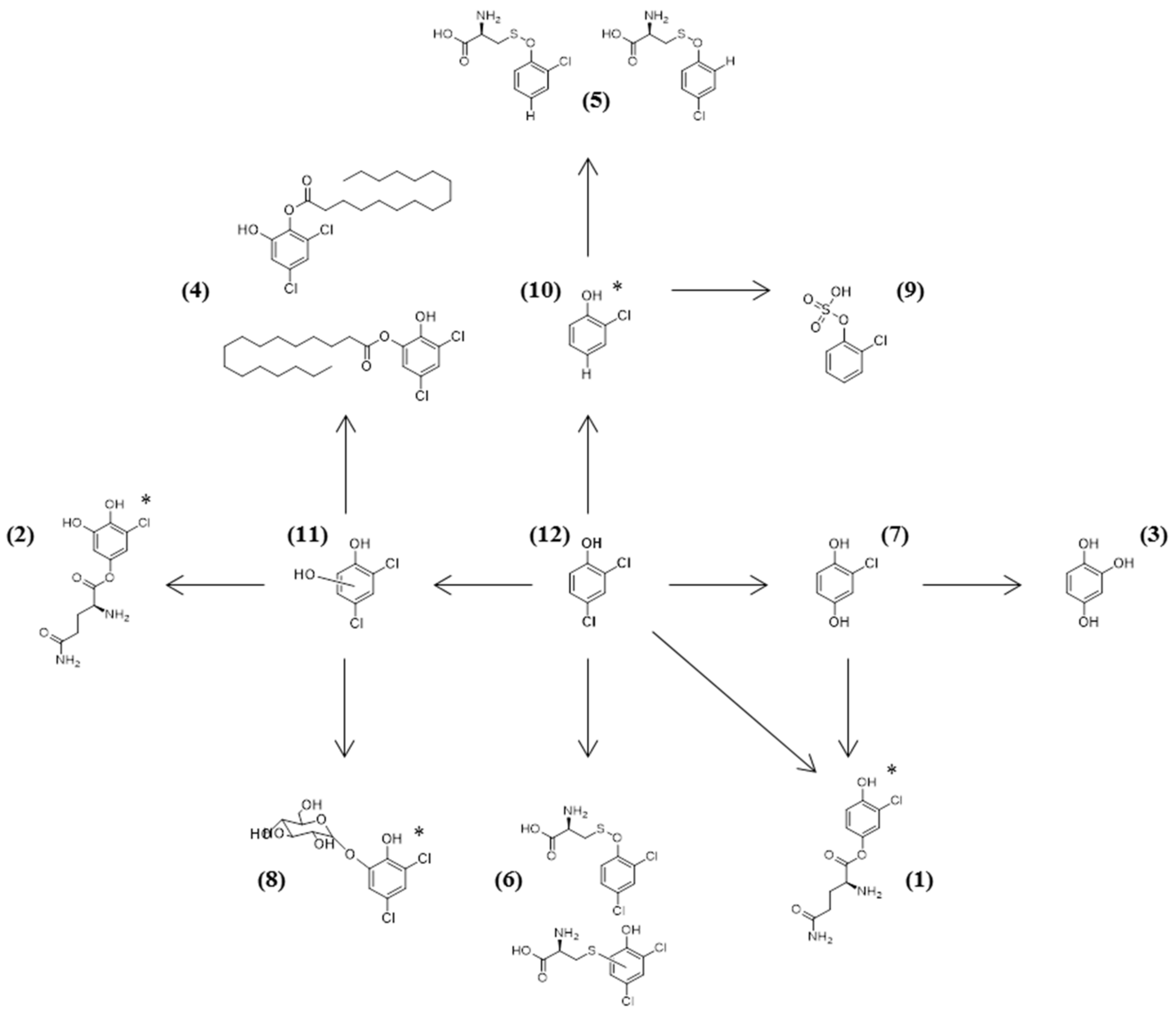
2.3. Fungal Mechanisms for 2,4-DCP Detoxification
3. Materials and Methods
3.1. Chemicals
3.2. Culture Conditions and Resting-Cell Reactions
3.3. Identification of Fungal Strains
3.4. Culture Conditions and Resting-Cell Reactions
3.5. Identification of 2,4-DCP Metabolites by UHPLC-HRMS/MS
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Das, S.; Dash, H.R. Microbial bioremediation: A potential tool for restoration of contaminated areas. In Microbial Biodegradation and Bioremediation; Surajit Das, Ed.; Elsevier: Amsterdam, The Netherlands, 2014; pp. 1–21. [Google Scholar]
- Honda, M.; Kannan, K. Biomonitoring of chlorophenols in human urine from several Asian countries, Greece and the United States. Environ. Pollut. 2018, 232, 487–493. [Google Scholar] [CrossRef] [PubMed]
- El-Naas, M.H.; Mousa, H.A.; Gamal, M. El Microbial degradation of chlorophenols. In Microbe-Induced Degradation of Pesticides; Singh, S.N., Ed.; Springer: New York, NY, USA, 2017; pp. 23–58. [Google Scholar]
- Huang, Z.; Chen, G.; Zeng, G.; Chen, A.; Zuo, Y.; Guo, Z.; Tan, Q.; Song, Z.; Niu, Q. Polyvinyl alcohol-immobilized Phanerochaete chrysosporium and its application in the bioremediation of composite-polluted wastewater. J. Hazard. Mater. 2015, 289, 174–183. [Google Scholar] [CrossRef] [PubMed]
- Patel, B.P.; Kumar, A. Biodegradation and co-metabolism of monochlorophenols and 2,4-dichlorophenol by microbial consortium. CLEAN Soil Air Water 2017, 45, 1700329. [Google Scholar] [CrossRef]
- Arora, P.; Bae, H. Bacterial degradation of chlorophenols and their derivatives. Microb. Cell Fact. 2014, 13, 31. [Google Scholar] [CrossRef] [PubMed]
- Patel, B.P.; Kumar, A. Biodegradation of 2,4-dichlorophenol by Bacillus endophyticus strain: Optimization of experimental parameters using response surface methodology and kinetic study. Desalin. Water Treat. 2016, 57, 15932–15940. [Google Scholar] [CrossRef]
- Ren, H.; Li, Q.; Zhan, Y.; Fang, X.; Yu, D. 2,4-Dichlorophenol hydroxylase for chlorophenol removal: Substrate specificity and catalytic activity. Enzym. Microb. Technol. 2016, 82, 74–81. [Google Scholar] [CrossRef]
- Anku, W.W.; Mamo, M.A.; Govender, P.P. Phenolic compounds in water: Sources, reactivity, toxicity and treatment methods. In Phenolic Compounds—Natural Sources, Importance and Applications; Soto-Hernandez, M., Palma-Tenango, M., del Rosario Garcia-Mateos, M., Eds.; InTech: Vienna, Austria, 2017; pp. 419–443. [Google Scholar]
- Srivastava, S. Bioremediation Technology: A greener and sustainable approach for restoration of environmental pollution. In Applied Environmental Biotechnology: Present Scenario and Future Trends; Kaushik, G., Ed.; Springer: New Delhi, India, 2015; pp. 1–18. [Google Scholar]
- Nikolaivits, E.; Dimarogona, M.; Fokialakis, N.; Topakas, E. Marine-derived biocatalysts: Importance, accessing and application in aromatic pollutant bioremediation. Front. Microbiol. 2017, 8, 265. [Google Scholar] [CrossRef] [PubMed]
- Turner, J.A.; Babcock, R.C.; Hovey, R.; Kendrick, G.A. Deep thinking: A systematic review of mesophotic coral ecosystems. ICES J. Mar. Sci. 2017, 74, 2309–2320. [Google Scholar] [CrossRef]
- Gribble, G. Biological activity of recently discovered halogenated marine natural products. Mar. Drugs 2015, 13, 4044–4136. [Google Scholar] [CrossRef]
- Atashgahi, S.; Häggblom, M.M.; Smidt, H. Organohalide respiration in pristine environments: Implications for the natural halogen cycle. Environ. Microbiol. 2018, 20, 934–948. [Google Scholar] [CrossRef]
- Sinniger, F.; Ballantine, D.L.; Bejarano, I.; Colin, P.L.; Pochon, X.; Pomponi, S.A.; Puglise, K.A.; Pyle, R.L.; Reaka, M.; Spalding, H.L.; et al. Biodiversity of mesophotic coral ecosystems. In Mesophotic Coral Ecosystems: A Lifeboat for Coral Reefs; Baker, K.E., Puglise, K.A., Harris, P.T.H., Eds.; The United Nations Environment Programme and GRID-Arendal: Arendal, Norway, 2016; pp. 50–62. [Google Scholar]
- Nicoletti, R.; Andolfi, A. The marine-derived filamentous fungi in biotechnology. In Grand Challenges in Marine Biotechnology; Rampelotto, P.H., Trincone, A., Eds.; Springer: Champaign, IL, USA, 2018; pp. 157–189. [Google Scholar]
- Birolli, W.G.; de Santos, D.A.; Alvarenga, N.; Garcia, A.C.F.S.; Romão, L.P.C.; Porto, A.L.M. Biodegradation of anthracene and several PAHs by the marine-derived fungus Cladosporium sp. CBMAI 1237. Mar. Pollut. Bull. 2018, 129, 525–533. [Google Scholar] [CrossRef] [PubMed]
- Bovio, E.; Gnavi, G.; Prigione, V.; Spina, F.; Denaro, R.; Yakimov, M.; Calogero, R.; Crisafi, F.; Varese, G.C. The culturable mycobiota of a Mediterranean marine site after an oil spill: Isolation, identification and potential application in bioremediation. Sci. Total Environ. 2017, 576, 310–318. [Google Scholar] [CrossRef] [PubMed]
- Vroumsia, T.; Steiman, R.; Seigle-Murandi, F.; Benoit-Guyod, J.-L. Groupe pour l’Étude du Devenir des Xénobiotiques dans l’Environnement (GEDEXE) Fungal bioconversion of 2,4-dichlorophenoxyacetic acid (2,4-D) and 2,4-dichlorophenol (2,4-DCP). Chemosphere 2005, 60, 1471–1480. [Google Scholar] [CrossRef] [PubMed]
- Matafonova, G.; Shirapova, G.; Zimmer, C.; Giffhorn, F.; Batoev, V.; Kohring, G.-W. Degradation of 2,4-dichlorophenol by Bacillus sp. isolated from an aeration pond in the Baikalsk pulp and paper mill (Russia). Int. Biodeterior. Biodegrad. 2006, 58, 209–212. [Google Scholar] [CrossRef]
- Kargi, F.; Eker, S. Kinetics of 2,4-dichlorophenol degradation by Pseudomonas putida CP1 in batch culture. Int. Biodeterior. Biodegrad. 2005, 55, 25–28. [Google Scholar] [CrossRef]
- Chen, A.; Zeng, G.; Chen, G.; Fan, J.; Zou, Z.; Li, H.; Hu, X.; Long, F. Simultaneous cadmium removal and 2,4-dichlorophenol degradation from aqueous solutions by Phanerochaete chrysosporium. Appl. Microbiol. Biotechnol. 2011, 91, 811–821. [Google Scholar] [CrossRef]
- Marco-Urrea, E.; Reddy, C.A. Degradation of chloro-organic pollutants by white rot fungi. In Microbial Degradation of Xenobiotics. Environmental Science and Engineering; Singh, S., Ed.; Springer: Berlin/Heidelberg, Germany, 2012; pp. 31–66. [Google Scholar]
- Aranda, E. Promising approaches towards biotransformation of polycyclic aromatic hydrocarbons with Ascomycota fungi. Curr. Opin. Biotechnol. 2016, 38, 1–8. [Google Scholar] [CrossRef]
- Nakagawa, A.; Osawa, S.; Hirata, T.; Yamagishi, Y.; Hosoda, J.; Horikoshi, T. 2,4-Dichlorophenol degradation by the soil fungus Mortierella sp. Biosci. Biotechnol. Biochem. 2006, 70, 525–527. [Google Scholar] [CrossRef]
- Zhang, D.; Yang, Y.; Leakey, J.E.; Cerniglia, C.E. Phase I and phase II enzymes produced by Cunninghamella elegans for the metabolism of xenobiotics. FEMS Microbiol. Lett. 1996, 138, 221–226. [Google Scholar] [CrossRef][Green Version]
- Mountfield, R.J.; Hopper, D.J. The formation of 1-hydroxymethylnaphthalene and 6-hydroxymethylquinoline by both oxidative and reductive routes in Cunninghamella elegans. Appl. Microbiol. Biotechnol. 1998, 50, 379–383. [Google Scholar] [CrossRef]
- Steiert, J.G.; Crawford, R.L. Catabolism of pentachlorophenol by a Flavobacterium sp. Biochem. Biophys. Res. Commun. 1986, 141, 825–830. [Google Scholar] [CrossRef]
- Orser, C.S.; Dutton, J.; Lange, C.; Jablonski, P.; Xun, L.; Hargis, M. Characterization of a Flavobacterium glutathione S-transferase gene involved reductive dechlorination. J. Bacteriol. 1993, 175, 2640–2644. [Google Scholar] [CrossRef] [PubMed]
- Xun, L.; Topp, E.; Orser, C.S. Purification and characterization of a tetrachloro-p-hydroquinone reductive dehalogenase from a Flavobacterium sp. J. Bacteriol. 1992, 174, 8003–8007. [Google Scholar] [CrossRef] [PubMed]
- Vacondio, B.; Birolli, W.G.; Ferreira, I.M.; Seleghim, M.H.R.; Gonçalves, S.; Vasconcellos, S.P.; Porto, A.L.M. Biodegradation of pentachlorophenol by marine-derived fungus Trichoderma harzianum CBMAI 1677 isolated from ascidian Didemnun ligulum. Biocatal. Agric. Biotechnol. 2015, 4, 266–275. [Google Scholar] [CrossRef]
- Hiratsuka, N.; Wariishi, H.; Tanaka, H. Degradation of diphenyl ether herbicides by the lignin-degrading basidiomycete Coriolus versicolor. Appl. Microbiol. Biotechnol. 2001, 57, 563–571. [Google Scholar] [PubMed]
- Öberg, L.G.; Rappe, C. Biochemical formation of PCDD/Fs from chlorophenols. Chemosphere 1992, 25, 49–52. [Google Scholar] [CrossRef]
- Tsujiyama, S.; Muraoka, T.; Takada, N. Biodegradation of 2,4-dichlorophenol by shiitake mushroom (Lentinula edodes) using vanillin as an activator. Biotechnol. Lett. 2013, 35, 1079–1083. [Google Scholar] [CrossRef] [PubMed]
- Leontievsky, A.A.; Myasoedova, N.M.; Baskunov, B.P.; Evans, C.S.; Golovleva, L.A. Transformation of 2,4,6-trichlorophenol by the white rot fungi Panus tigrinus and Coriolus versicolor. Biodegradation 2000, 11, 331–340. [Google Scholar] [CrossRef]
- Rubilar, O.; Diez, M.C.; Gianfreda, L. Transformation of chlorinated phenolic compounds by white rot fungi. Crit. Rev. Environ. Sci. Technol. 2008, 38, 227–268. [Google Scholar] [CrossRef]
- Stoilova, I.; Krastanov, A.; Stanchev, V.; Daniel, D.; Gerginova, M.; Alexieva, Z. Biodegradation of high amounts of phenol, catechol, 2,4-dichlorophenol and 2,6-dimethoxyphenol by Aspergillus awamori cells. Enzym. Microb. Technol. 2006, 39, 1036–1041. [Google Scholar] [CrossRef]
| Isolate Code | Invertebrate of Origin | Location | Depth (m) | Isolate Identification | % DCP Removal |
|---|---|---|---|---|---|
| TM2-S5 | Sponge on top of Spondylus | Med E | 38 | Penicillium steckii | 58.5 |
| TM6-S1 | Dendronephthya sp. | Eilat | 120 | Aspergillus tubingensis | 23.2 |
| TM6-S2 | Dendronephthya sp. | Eilat | 120 | Penicillium kewense | 9.3 |
| TM37-S2 | Spondylus sp. | Eilat | 49 | Penicillium chrysogenum | 49.2 |
| TM38-S1 | Polycarpa sp. | Eilat | 49 | Penicillium sp. | 56.2 |
| TM9-S2 | Theonella sp. | Eilat | 65 | Chrysosporium sp. | 74.0 |
| TM43-S1 | Xestospongia sp. | Andaman | 30–40 | Penicillium sp. | 17.5 |
| TM43-S3 | Xestospongia sp. | Andaman | 30–40 | Cladosporium halotolerans | 4.5 |
| TM46-S1 | Thrinacophora sp. | Andaman | 30–40 | Cladosporium sp. | 30.4 |
| TM47-S1 | Chondrilla sp. | Andaman | 30–40 | Cladosporium halotolerans | 46.9 |
| TM116-S2 | Xestospongia sp. | Andaman | 30–40 | Aspergillus sp. | 28.4 |
| TM116-S3 | Xestospongia sp. | Andaman | 30–40 | Hortaea sp. (yeast) | 9.5 |
| TM122-S1 | Iotrochota sp. | Andaman | 30–40 | Aspergillus sp. | 21.7 |
| TM122-S2 | Iotrochota sp. | Andaman | 30–40 | Aspergillus fumigatus | 29.5 |
| TM124-S1 | Echinaster sepositus | Med W | 30–40 | Aspergillus sp. | 69.0 |
| TM125-S2 | Clavelina dellavallei | Med W | 30–40 | Penicillium sp. | 51.6 |
| TM125-S3 | Clavelina dellavallei | Med W | 30–40 | Penicillium sp. | 50.2 |
| TM126-S1 | Clayx nicaensis | Med W | 30–40 | Aspergillus sp. | 47.0 |
| TM133-S2 | Cerianthus membranaceus | Med W | 30–40 | Penicillium crustosum | 10.6 |
| TM138-S1 | Didemnum maculosum | Med W | 30–40 | Purpureocillium lilacinum | 26.2 |
| TM138-S3 | Didemnum maculosum | Med W | 30–40 | Cladosporium sp. | 16.8 |
| TM138-S4 | Didemnum maculosum | Med W | 30–40 | Purpureocillium lilacinum | 24.6 |
| TM2-S6 | Sponge on top of Spondylus | Med E | 38 | Aspergillus sp. | 55.6 |
| TM7-S1 | Siphonogorgia sp. | Eilat | 120 | Penicillium sp. | 29.3 |
| TM30-S1 | Diacarnus erythraenus | Eilat | 49 | Penicillium fellutanum | 37.1 |
| TM220-S1 | Gorgonian overgrown by Antipathozoanthus sp. | Eilat | 75 | Cladosporium sphaerospermum | 40.9 |
| TM220-S4 | Gorgonian overgrown by Antipathozoanthus sp. | Eilat | 75 | Penicillium sp. | 47.1 |
| TM225-S2 | Villogorgia nozzolea | Eilat | 68 | Penicillium chrysogenum | 9.0 |
| TM226-S1 | Callyspongia sp. | Eilat | 86 | Cladosporium halotolerans | 36.2 |
| TM226-S2 | Callyspongia sp. | Eilat | 86 | Penicillium chrysogenum | 9.7 |
| TM230-S1 | Siphonogorgia sp. | Eilat | 115 | Penicillium sp. | 55.0 |
| TM230-S6 | Siphonogorgia sp. | Eilat | 115 | Penicillium sp. | 32.5 |
| TM242-S1 | Orange/beige gorgonian | Eilat | 152 | Penicillium chrysogenum | 44.0 |
| TM53-S1 | Plakina sp. | Andaman | 30–40 | Cladosporium sp. | 16.3 |
| TM54-S1 | Terpios sp. | Andaman | 30–40 | Aspergillus sp. | 40.0 |
| TM58-S1 | Biemna sp. | Andaman | 30–40 | Penicillium sp. | 30.0 |
| TM58-S2 | Biemna sp. | Andaman | 30–40 | Aspergillus niger | 46.0 |
| TM65-S2 | Padina sp | Andaman | 30–40 | Pseudocercosporella sp. | 46.4 |
| TM65-S4 | Padina sp | Andaman | 30–40 | Acremonium sp. | 28.3 |
| TM75-S1 | Ulva sp. | Andaman | 30–40 | Aspergillus sp. | 21.6 |
| TM83-S1 | Bryozoan | Andaman | 30–40 | Cladosporium halotolerans | 15.6 |
| TM83-S2 | Bryozoan | Andaman | 30–40 | Cladosporium halotolerans | 33.7 |
| TM116-S1 | Xestospongia sp. | Andaman | 30–40 | Aspergillus aculeolatus | 24.6 |
| TM122-S3 | Iotrochota sp. | Andaman | 30–40 | Aspergillus creber | 62.0 |
| TM122-S4 | Iotrochota sp. | Andaman | 30–40 | Obolarina sp. | 40.1 |
| TM124-S4 | Echinaster sepositus | Med W | 30–40 | Aspergillus sp. | 36.0 |
| TM124-S7 | Echinaster sepositus | Med W | 30–40 | Aspergillus sp. | 46.4 |
| TM132-S1 | Haliclona mediterranea | Med W | 30–40 | Aspergillus sp. | 24.9 |
| TM141-S1 | Cliona celata | Med W | 30–40 | Alternaria sp. | 22.3 |
| TM141-S2 | Cliona celata | Med W | 30–40 | Alternaria sp. | 16.3 |
| TM141-S3 | Cliona celata | Med W | 30–40 | Alternaria sp. | 39.8 |
| TM148-S1 | Astropecten bispinosus | Med W | 30–40 | Aspergillus sp. | 24.9 |
| A/A | Rt (Min) | [M − H]− | EC | MS2 | Found in |
|---|---|---|---|---|---|
| (1) | 4.90 | 271.0480 | C11H13ClN2O4 | 114.9515 (C4O2Cl), 191.0924 (C11H13O2N) | Penicillium steckii TM2-S5, Chrysosporium sp. ΤΜ9-S2, Aspergillus creber TM122-S3, Aspergillus sp. TM124-S1 |
| (2) | 6.98 | 287.0429 | C11H13O5N2Cl | - | Chrysosporium sp. ΤΜ9-S2, Aspergillus sp. TM124-S1, Penicilluim sp. TM38-S1 |
| (3) | 7.01 | 125.0233 | C6H6O3 | - | Penicillium steckii TM2-S5, Penicilluim sp.TM38-S1, Aspergillus creber TM122-S3, Aspergillus sp. TM124-S1 |
| (4) | 8.87 | 415.1801 | C22H34Cl2O3 | - | All |
| (4) | 9.06 | 415.1801 | C22H34Cl2O3 | - | All |
| (5) | 9.42 | 245.9986 | C9H10NClO3S | 158.9675 (C6H4OClS) | Chrysosporium sp. ΤΜ9-S2 |
| (5) | 9.63 | 245.9986 | C9H10NClO3S | 158.9675 (C6H4OClS) | Chrysosporium sp. ΤΜ9-S2 |
| (6) | 10.24 | 279.9596 | C9H9O3NSCl2 | - | Chrysosporium sp. ΤΜ9-S2 |
| (6) | 10.42 | 279.9596 | C9H9O3NSCl2 | - | Chrysosporium sp. ΤΜ9-S2 |
| (7) | 11.68 | 142.9903 | C6H5ClO2 | 114.9515 (C4O2Cl) | Chrysosporium sp. ΤΜ9-S2 |
| (8) | 12.33 | 339.0033 | C12H14Cl2O7 | 160.9565 (C6H3OCl2), 124.9798 (C6H2OCl) | Chrysosporium sp. ΤΜ9-S2 |
| (9) | 12.43 | 206.9513 | C6H5O4ClS | - | Penicillium steckii TM2-S5, Aspergillus creber TM122-S3, Aspergillus sp. TM124-S1 |
| (10) | 13.37 | 126.9945 | C6H5OCl | - | Penicillium steckii TM2-S5, Chrysosporium sp. ΤΜ9-S2, Aspergillus creber TM122-S3 |
| (11) | 13.81 | 176.9505 | C6H4Cl2O2 | - | Chrysosporium sp. ΤΜ9-S2 |
| (11) | 13.94 | 176.9505 | C6H4Cl2O2 | - | Chrysosporium sp. ΤΜ9-S2 |
| (12) | 15.12 | 160.9565 | C6H4Cl2O | 124.9798 (C6H2OCl); 88.9943 (C3H2OCl) | All |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nikolaivits, E.; Agrafiotis, A.; Termentzi, A.; Machera, K.; Le Goff, G.; Álvarez, P.; Chavanich, S.; Benayahu, Y.; Ouazzani, J.; Fokialakis, N.; et al. Unraveling the Detoxification Mechanism of 2,4-Dichlorophenol by Marine-Derived Mesophotic Symbiotic Fungi Isolated from Marine Invertebrates. Mar. Drugs 2019, 17, 564. https://doi.org/10.3390/md17100564
Nikolaivits E, Agrafiotis A, Termentzi A, Machera K, Le Goff G, Álvarez P, Chavanich S, Benayahu Y, Ouazzani J, Fokialakis N, et al. Unraveling the Detoxification Mechanism of 2,4-Dichlorophenol by Marine-Derived Mesophotic Symbiotic Fungi Isolated from Marine Invertebrates. Marine Drugs. 2019; 17(10):564. https://doi.org/10.3390/md17100564
Chicago/Turabian StyleNikolaivits, Efstratios, Andreas Agrafiotis, Aikaterini Termentzi, Kyriaki Machera, Géraldine Le Goff, Pedro Álvarez, Suchana Chavanich, Yehuda Benayahu, Jamal Ouazzani, Nikolas Fokialakis, and et al. 2019. "Unraveling the Detoxification Mechanism of 2,4-Dichlorophenol by Marine-Derived Mesophotic Symbiotic Fungi Isolated from Marine Invertebrates" Marine Drugs 17, no. 10: 564. https://doi.org/10.3390/md17100564
APA StyleNikolaivits, E., Agrafiotis, A., Termentzi, A., Machera, K., Le Goff, G., Álvarez, P., Chavanich, S., Benayahu, Y., Ouazzani, J., Fokialakis, N., & Topakas, E. (2019). Unraveling the Detoxification Mechanism of 2,4-Dichlorophenol by Marine-Derived Mesophotic Symbiotic Fungi Isolated from Marine Invertebrates. Marine Drugs, 17(10), 564. https://doi.org/10.3390/md17100564








