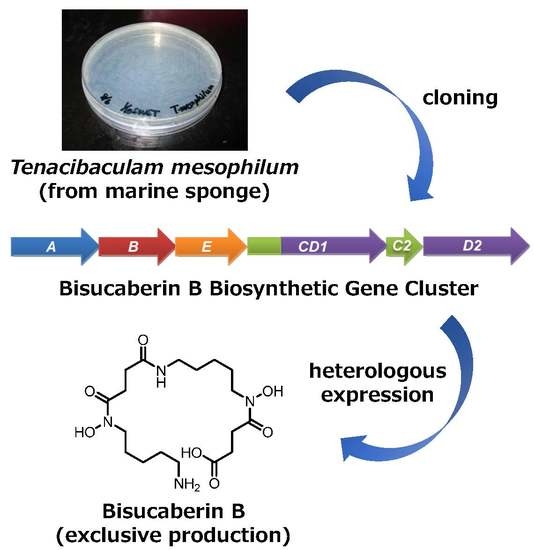
Cloning of the Bisucaberin B Biosynthetic Gene Cluster from the Marine Bacterium Tenacibaculum mesophilum, and Heterologous Production of Bisucaberin B
Abstract
1. Introduction
2. Results and Discussion
2.1. Cloning of the Bisucaberin B Biosynthetic Gene Cluster
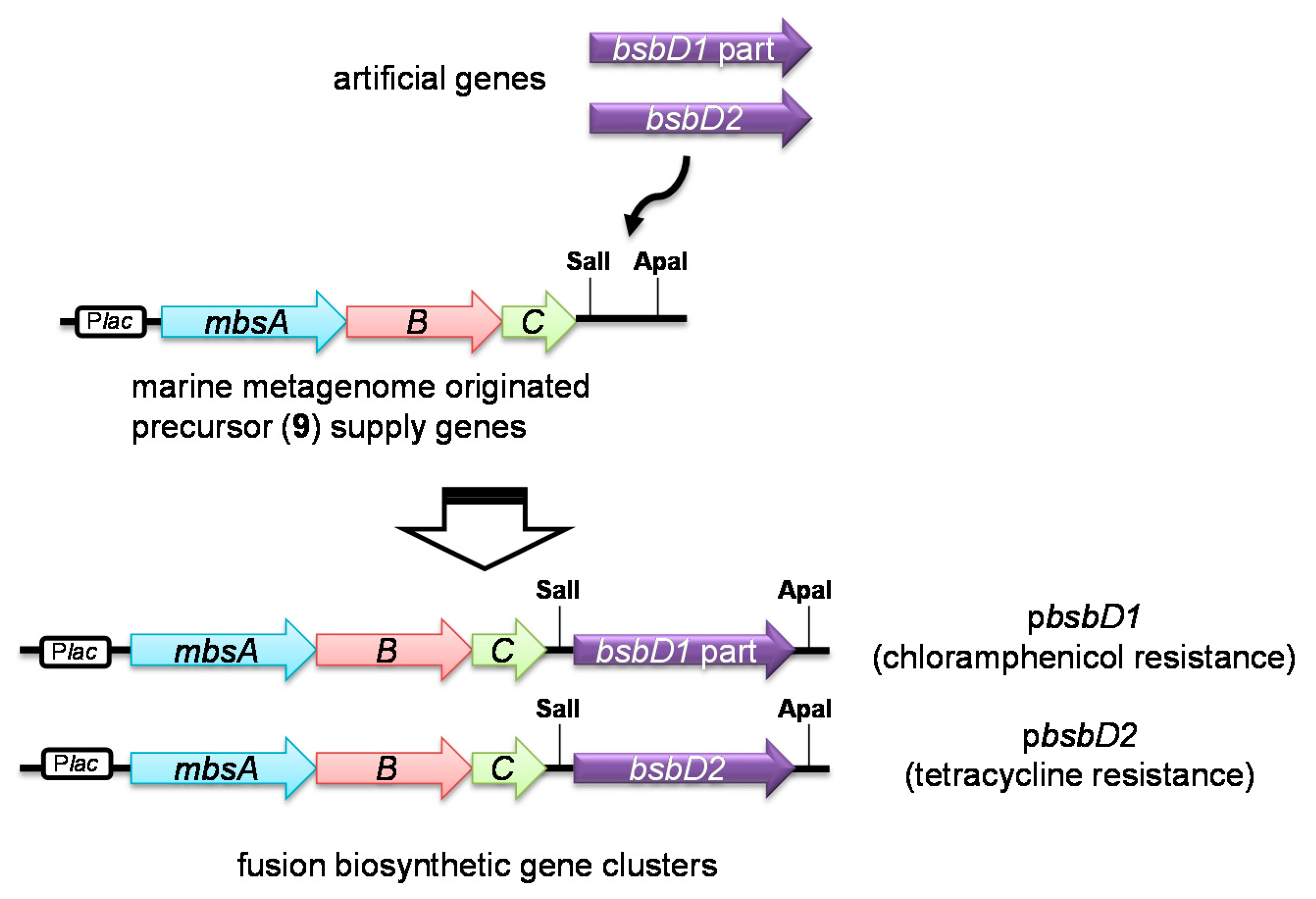
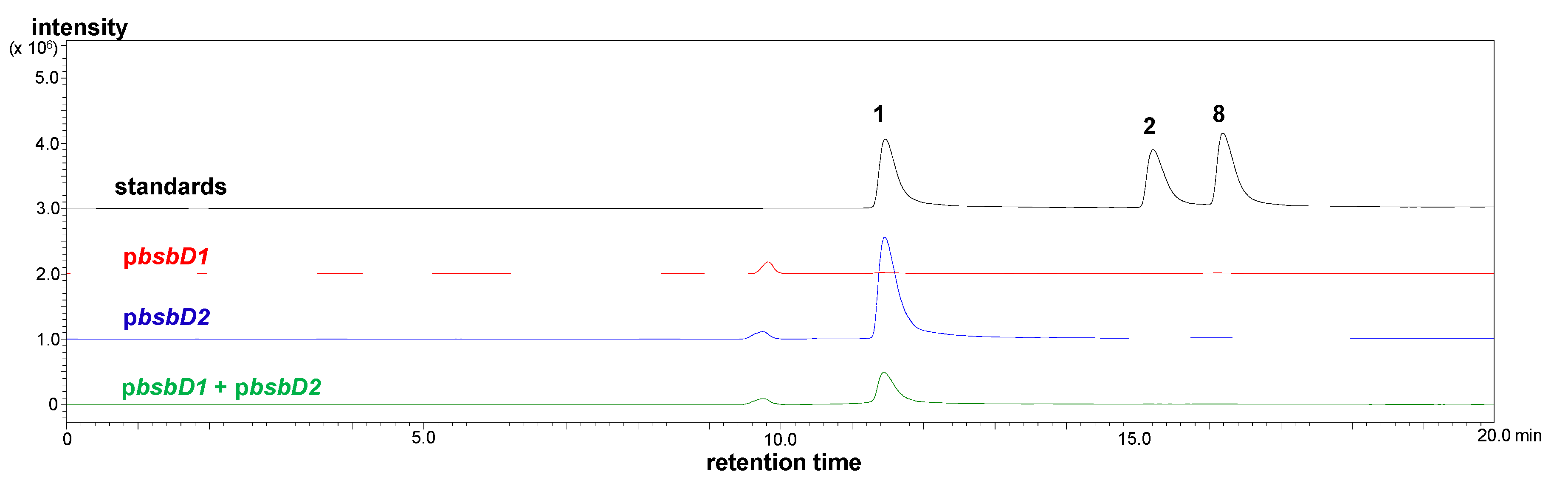
2.2. Heterologous Production of Bisucaberin B by a Fusion Gene Cluster System
3. Materials and Methods
3.1. General Experimental Procedures
3.2. T. mesophilum Genomic Library Construction
3.3. Amplification of the Fragment of the Biosynthetic Genes by Degenerate Primer PCR
3.4. Cloning of the Bisucaberin B Biosynthetic Gene Cluster (Bsb Cluster)
3.5. Synthesis of Artificial Genes
3.6. Construction of the Fusion Gene Clusters
3.7. Preparation of Single and Double Transformants
3.8. CAS Solution Assay
3.9. Heterologous Production and Identification of Bisucaberin B
3.10. LC-MS Analysis of Metabolites
3.11. Quantification of Bisucaberin B by MRM Analysis
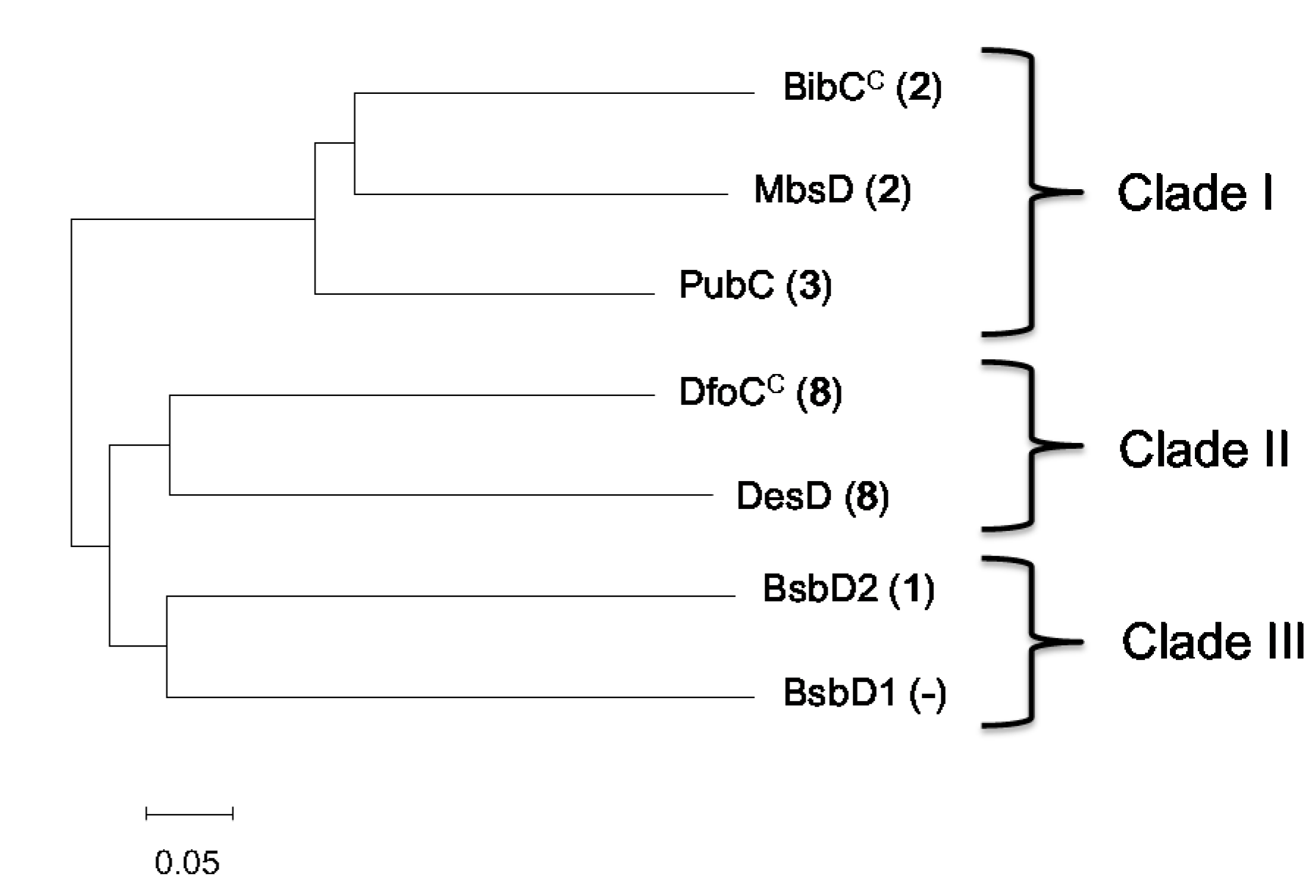
3.12. Phylogenetic Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Sandy, M.; Butler, A. Microbial iron acquisition: Marine and terrestrial siderophores. Chem. Rev. 2009, 109, 4580–4595. [Google Scholar] [CrossRef] [PubMed]
- D’Onofrio, A.; Crawford, J.M.; Stewart, E.J.; Witt, K.; Gavrish, E.; Epstein, S.; Clardy, J.; Lewis, K. Siderophores from neighboring organisms promote the growth of uncultured bacteria. Chem. Biol. 2010, 17, 254–264. [Google Scholar] [CrossRef] [PubMed]
- Seyedsayamdost, M.R.; Traxler, M.F.; Zheng, S.-L.; Kolter, R.; Clardy, J. Structure and biosynthesis of amychelin, an unusual mixed-ligand siderophore from Amycolatopsis sp. AA4. J. Am. Chem. Soc. 2011, 133, 11434–11437. [Google Scholar] [CrossRef] [PubMed]
- Traxler, M.F.; Seyedsayamdost, M.R.; Clardy, J.; Kolter, R. Interspecies modulation of bacterial development through iron competition and siderophore piracy. Mol. Microbiol. 2012, 86, 628–644. [Google Scholar] [CrossRef] [PubMed]
- Boettcher, T.; Clardy, J. A chimeric siderophore halts swarming vibrio. Angew. Chem. Int. Ed. 2014, 53, 3510–3513. [Google Scholar] [CrossRef] [PubMed]
- Bickel, H.; Bosshardt, R.; Gaeumann, E.; Reusser, P.; Vischer, E.; Voser, W.; Wettstein, A.; Zaehner, H. Metabolic products of actinomycetaceae. XXVI. Isolation and properties of ferrioxamines A to F, representing new sideramine compounds. Helv. Chim. Acta 1960, 43, 2118–2128. [Google Scholar] [CrossRef]
- Takahashi, A.; Nakamura, H.; Kameyama, T.; Kurasawa, S.; Naganawa, H.; Okami, Y.; Takeuchi, T.; Umezawa, H.; Iitaka, Y. Bisucaberin, a new siderophore, sensitizing tumor cells to macrophage-mediated cytolysis. II. Physico-chemical properties and structure determination. J. Antibiot. 1987, 40, 1671–1676. [Google Scholar] [CrossRef] [PubMed]
- Moore, C.H.; Foster, L.A.; Gerbig, D.G., Jr.; Dyer, D.W.; Gibson, B.W. Identification of alcaligin as the siderophore produced by Bordetella pertussis and B. bronchiseptica. J. Bacteriol. 1995, 177, 1116–1118. [Google Scholar] [CrossRef] [PubMed]
- Ledyard, K.M.; Butler, A. Structure of putrebactin, a new dihydroxamate siderophore produced by Shewanella putrefaciens. J. Biol. Inorg. Chem. 1997, 2, 93–97. [Google Scholar] [CrossRef]
- Fujita, M.J.; Nakano, K.; Sakai, R. Bisucaberin B, a linear hydroxamate class siderophore from the marine bacterium Tenacibaculum mesophilum. Molecules 2013, 18, 3917–3926. [Google Scholar] [CrossRef] [PubMed]
- Olivieri, N.F.; Brittenham, G.M. Iron-chelating therapy and the treatment of thalassemia. Blood 1997, 89, 739–761. [Google Scholar] [PubMed]
- Ishida, S.; Arai, M.; Nikawa, H.; Kobayashi, M. Inhibitory effect of cyclic trihydroxamate siderophore, desferrioxamine E, on the biofilm formation of Mycobacterium species. Biol. Pharm. Bull. 2011, 34, 917–920. [Google Scholar] [CrossRef] [PubMed]
- Kameyama, T.; Takahashi, A.; Kurasawa, S.; Ishizuka, M.; Okami, Y.; Takeuchi, T.; Umezawa, H. Bisucaberin, a new siderophore, sensitizing tumor cells to macrophage-mediated cytolysis. I. Taxonomy of the producing organism, isolation and biological properties. J. Antibiot. 1987, 40, 1664–1670. [Google Scholar] [CrossRef] [PubMed]
- Kang, H.Y.; Brickman, T.J.; Beaumont, F.C.; Armstrong, S.K. Identification and characterization of iron-regulated Bordetella pertussis alcaligin siderophore biosynthesis genes. J. Bacteriol. 1996, 178, 4877–4884. [Google Scholar] [CrossRef] [PubMed]
- Barona-Gomez, F.; Wong, U.; Giannakopulos, A.E.; Derrick, P.J.; Challis, G.L. Identification of a cluster of genes that directs desferrioxamine biosynthesis in Streptomyces coelicolor M145. J. Am. Chem. Soc. 2004, 126, 16282–16283. [Google Scholar] [CrossRef] [PubMed]
- Kadi, N.; Song, L.; Challis, G.L. Bisucaberin biosynthesis: An adenylating domain of the BibC multi-enzyme catalyzes cyclodimerization of N-hydroxy-N-succinylcadaverine. Chem. Commun. 2008, 5119–5121. [Google Scholar] [CrossRef] [PubMed]
- Kadi, N.; Arbache, S.; Song, L.; Oves-Costales, D.; Challis, G.L. Identification of a gene clusterthat directs putrebactin biosynthesis in Shewanella Species: PubC catalyzes cyclodimerization of N-Hydroxy-N-succinylputrescine. J. Am. Chem. Soc. 2008, 130, 10458–10459. [Google Scholar] [CrossRef] [PubMed]
- Fujita, M.J.; Kimura, N.; Yokose, H.; Otsuka, M. Heterologous production of bisucaberin using a biosynthetic gene cluster cloned from a deep sea metagenome. Mol. Biosyst. 2012, 8, 482–485. [Google Scholar] [CrossRef] [PubMed]
- Fujita, M.J.; Sakai, R. Heterologous production of desferrioxamines with a fusion biosynthetic gene cluster. Biosci. Biotechnol. Biochem. 2013, 77, 2467–2472. [Google Scholar] [CrossRef] [PubMed]
- Kadi, N.; Oves-Costales, D.; Barona-Gomez, F.; Challis, G.L. A new family of ATP-dependent oligomerization-macrocyclization biocatalysts. Nat. Chem. Biol. 2007, 3, 652–656. [Google Scholar] [CrossRef] [PubMed]
- Ruetschlin, S.; Gunesch, S.; Boettcher, T. One enzyme to build them all: Ring-size engineered siderophores inhibit the swarming motility of vibrio. ACS Chem. Biol. 2018, 13, 1153–1158. [Google Scholar] [CrossRef] [PubMed]
- Telfer, T.J.; Gotsbacher, M.P.; Soe, C.Z.; Codd, R. Mixing up the pieces of the desferrioxamine B jigsaw defines the biosynthetic sequence catalyzed by DesD. ACS Chem. Biol. 2016, 11, 1452–1462. [Google Scholar] [CrossRef] [PubMed]
- Soe, C.Z.; Codd, R. Unsaturated macrocyclic dihydroxamic acid siderophores produced by Shewanella putrefaciens using precursor-directed biosynthesis. ACS Chem. Biol. 2014, 9, 945–956. [Google Scholar] [CrossRef] [PubMed]
- Soe, C.Z.; Telfer, T.J.; Levina, A.; Lay, P.A.; Codd, R. Simultaneous biosynthesis of putrebactin, avaroferrin and bisucaberin by Shewanella putrefaciens and characterisation of complexes with iron(III), molybdenum(VI) or chromium(V). J. Inorg. Biochem. 2016, 162, 207–215. [Google Scholar] [CrossRef] [PubMed]
- Hrvatin, S.; Piel, J. Rapid isolation of rare clones from highly complex DNA libraries by PCR analysis of liquid gel pools. J. Microbiol. Methods 2007, 68, 434–436. [Google Scholar] [CrossRef] [PubMed]
- Pao, S.S.; Paulsen, I.T.; Saier, M.H., Jr. Major facilitator superfamily. Microbiol. Mol. Biol. Rev. 1998, 62, 1–34. [Google Scholar] [PubMed]
- Lesuisse, E.; Simon-Casteras, M.; Labbe, P. Siderophore-mediated iron uptake in Saccharomyces cerevisiae: The SIT1 gene encodes a ferrioxamine B permease that belongs to the major facilitator superfamily. Microbiology 1998, 144, 3455–3462. [Google Scholar] [CrossRef] [PubMed]
- Chatfield, C.H.; Mulhern, B.J.; Viswanathan, V.K.; Cianciotto, N.P. The major facilitator superfamily-type protein LbtC promotes the utilization of the legiobactin siderophore by Legionella pneumophila. Microbiology 2012, 158, 721–735. [Google Scholar] [CrossRef] [PubMed]
- Hannauer, M.; Sheldon, J.R.; Heinrichs, D.E. Involvement of major facilitator superfamily proteins SfaA and SbnD in staphyloferrin secretion in Staphylococcus aureus. FEBS Lett. 2015, 589, 730–737. [Google Scholar] [CrossRef] [PubMed]
- Fujita, M.J.; Sakai, R. Production of avaroferrin and putrebactin by heterologous expression of a deep-sea metagenomic DNA. Mar. Drugs 2014, 12, 4799–4809. [Google Scholar] [CrossRef] [PubMed]
- Schwyn, B.; Neilands, J.B. Universal chemical assay for the detection and determination of siderophores. Anal. Biochem. 1987, 160, 47–56. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Making Your Own Electrocompetent Cells. Available online: https://www.neb.com/protocols/2012/06/21/making-your-own-electrocompetent-cells (accessed on 8 August 2018).

| Proteins | Length | Annotated Functions | Identity 3 |
|---|---|---|---|
| BsbA | 502 aa | lysine decarboxylase | 43% |
| BsbB | 440 aa | lysine-6-monooxygenase | 51% |
| BsbE | 391 aa | siderophore secretion | - |
| BsbCD1 | 810 aa | acyl transferase | 32% 1 |
| peptide synthetase | 42% 2 | ||
| BsbC2 | 196 aa | acyl transferase | 32% |
| BsbD2 | 606 aa | peptide synthetase | 46% |
| Proteins | BsbD2 (1) | BibCC (2) | MbsD (2) | PubC (4) | DesD (8) | DfoCC (8) |
|---|---|---|---|---|---|---|
| BsbD1 (-) | 50/71 | 47/66 | 43/64 | 42/64 | 49/69 | 47/67 |
| BsbD2 (1) | - | 47/65 | 46/66 | 45/63 | 49/68 | 45/66 |
| BibCC (2) | - | 59/75 | 63/76 | 50/69 | 48/65 | |
| MbsD (2) | - | 61/75 | 49/69 | 44/63 | ||
| PubC (4) | - | 47/68 | 45/64 | |||
| DesD (8) | - | 56/72 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fujita, M.J.; Goto, Y.; Sakai, R. Cloning of the Bisucaberin B Biosynthetic Gene Cluster from the Marine Bacterium Tenacibaculum mesophilum, and Heterologous Production of Bisucaberin B. Mar. Drugs 2018, 16, 342. https://doi.org/10.3390/md16090342
Fujita MJ, Goto Y, Sakai R. Cloning of the Bisucaberin B Biosynthetic Gene Cluster from the Marine Bacterium Tenacibaculum mesophilum, and Heterologous Production of Bisucaberin B. Marine Drugs. 2018; 16(9):342. https://doi.org/10.3390/md16090342
Chicago/Turabian StyleFujita, Masaki J., Yusuke Goto, and Ryuichi Sakai. 2018. "Cloning of the Bisucaberin B Biosynthetic Gene Cluster from the Marine Bacterium Tenacibaculum mesophilum, and Heterologous Production of Bisucaberin B" Marine Drugs 16, no. 9: 342. https://doi.org/10.3390/md16090342
APA StyleFujita, M. J., Goto, Y., & Sakai, R. (2018). Cloning of the Bisucaberin B Biosynthetic Gene Cluster from the Marine Bacterium Tenacibaculum mesophilum, and Heterologous Production of Bisucaberin B. Marine Drugs, 16(9), 342. https://doi.org/10.3390/md16090342