Chalcomycins from Marine-Derived Streptomyces sp. and Their Antimicrobial Activities
Abstract
:1. Introduction
2. Results and Discussion
3. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Aminov, R. History of Antimicrobial Drug Discovery: Major Classes and Health Impact. Biochem. Pharmacol. 2017, 133, 4–19. [Google Scholar] [CrossRef] [PubMed]
- Chaudhary, A.S. A Review of Global Initiatives to Fight Antibiotic Resistance and Recent Antibiotics Discovery. Acta Pharm. Sin. B 2016, 6, 552–556. [Google Scholar] [CrossRef] [PubMed]
- Gamerdinger, M.; Deuerling, E. Macrolides: The Plug Is Out. Cell 2012, 151, 469–471. [Google Scholar] [CrossRef] [PubMed]
- Kwiatkowska, B.; Maslinska, M. Macrolide Therapy in Chronic Inflammatory diseases. Mediat. Inflamm. 2012, 2012, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Hansen, J.L.; Ippolito, J.A.; Ban, N.; Nissen, P.; Moore, P.B.; Steitz, T.A. The Structures of Four Macrolide Antibiotics Bound to the Large Ribosomal Subunit. Mol. Cell 2002, 10, 117–128. [Google Scholar] [CrossRef]
- Blondeau, J.M.; DeCarolis, E.; Metzler, K.L.; Hansen, G.T. The macrolides. Expert Opin. Investig. Drugs 2002, 11, 189–215. [Google Scholar] [CrossRef] [PubMed]
- Zuckerman, J.M.; Qamar, F.; Bono, B.R. Review of Macrolides (Azithromycin, Clarithromycin), Ketolids (Telithromycin) and Glycylcyclines (Tigecycline). Med. Clin. N. Am. 2011, 95, 761–791. [Google Scholar] [CrossRef] [PubMed]
- Elshahawi, S.I.; Shaaban, K.A.; Kharel, M.K.; Thorson, J.S. A comprehensive review of glycosylated bacterial natural products. Chem. Soc. Rev. 2015, 44, 7591–7697. [Google Scholar] [CrossRef] [PubMed]
- Omura, S.; Nakagawa, A. Chemical and biological studies on 16-membered macrolide antibiotics. J. Antibiot. 1975, 28, 401–433. [Google Scholar] [CrossRef] [PubMed]
- Corcoran, J.W.; Hahn, F.E. Antibiotics, Vol. 3: Mechanism of Action of Antimicrobial and Antitumor Agents; Springer: Berlin/Heidelberg, Germany, 1975; pp. 459–479. [Google Scholar]
- Wang, C.X.; Ding, R.; Jiang, S.T.; Tang, J.S.; Hu, D.; Chen, G.D.; Lin, F.; Hong, K.; Yao, X.S.; Gao, H. Aldgamycins J–O, 16-membered Macrolides with a Branched Octose Unit from Streptomycetes sp. and their Antibacterial Activities. J. Nat. Prod. 2016, 79, 2446–2454. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Chen, G.D.; Zou, J.; He, R.R.; Qin, S.Y.; Hu, D.; Li, G.Q.; Guo, L.D.; Yao, X.S.; Gao, H. Dimericbiscognienyne A: A meroterpenoid dimer from Biscogniauxia sp. with new skeleton and its activity. Org. Lett. 2017, 19, 38–41. [Google Scholar] [CrossRef] [PubMed]
- Sun, T.Y.; Zou, J.; Chen, G.D.; Hu, D.; Wu, B.; Liu, X.Z.; Yao, X.S.; Gao, H. A set of interesting sequoiatones stereoisomers from a wetland soil-derived fungus Talaromyces flavus. Acta Pharm. Sin. B 2017. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.X.; Chen, G.D.; Feng, C.C.; He, R.R.; Qin, S.Y.; Hu, D.; Chen, H.R.; Liu, X.Z.; Yao, X.Z.; Gao, H. Same Data, different structures: diastereoisomers with substantiallyidentical NMR data from nature. Chem. Commun. 2016, 52, 1250–1253. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.M.; Sun, T.Y.; Ma, M.; Chen, G.D.; Zhou, Z.Q.; Wang, C.X.; Hu, D.; Chen, L.G.; Yao, X.S.; Gao, H. Adeninealkylresorcinol, the first alkylresorcinol tethered with nucleobase from Lasiodiplodia sp. Fitoterapia 2016, 112, 254–259. [Google Scholar] [CrossRef] [PubMed]
- Sun, T.Y.; Kuang, R.Q.; Chen, G.D.; Qin, S.Y.; Wang, C.X.; Hu, D.; Wu, B.; Liu, X.Z.; Yao, X.S.; Gao, H. Three pairs of new isopentenyl dibenzo[b,e]oxepinone enantiomers from Talaromyces flavus, a wetland soil-derived fungus. Molecules 2016, 21, 1184. [Google Scholar] [CrossRef] [PubMed]
- He, J.W.; Wang, C.X.; Yang, L.; Chen, G.D.; Hu, D.; Guo, L.D.; Yao, X.S.; Gao, H. A pair of new polyketide enantiomers from three endolichenic fungal strains Nigrospora sphaerica, Alternaria alternata, and Phialophora sp. Nat. Prod. Commun. 2016, 11, 829–831. [Google Scholar] [PubMed]
- Zhao, Q.; Wang, C.X.; Yu, Y.; Wang, G.Q.; Zheng, Q.C.; Chen, G.D.; Lian, Y.Y.; Lin, F.; Guo, L.D.; Gao, H. Nodulisporipyrones A-D, new bioactive α-pyrone derivatives from Nodulisporium sp. J. Asian Nat. Prod. Res. 2015, 17, 567–575. [Google Scholar] [CrossRef] [PubMed]
- Zou, J.; Li, J.; Wu, Z.Y.; Zhao, Q.; Wang, G.Q.; Zhao, H.; Chen, G.D.; Sun, X.; Guo, L.D.; Gao, H. New α-pyrone and phthalide from the Xylariaceae fungus. J. Asian Nat. Prod. Res. 2015, 17, 705–710. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.H.; Chen, G.D.; Wang, C.X.; Hu, D.; Li, X.X.; Lian, Y.Y.; Lin, F.; Guo, L.D.; Gao, H. Pericoterpenoid A, a new bioactive cadinane-type sesquiterpene from Periconia sp. J. Asian Nat. Prod. Res. 2015, 17, 671–675. [Google Scholar] [CrossRef] [PubMed]
- Tang, X.L.; Dai, P.; Gao, H.; Wang, C.X.; Chen, G.D.; Hong, K.; Hu, D.; Yao, X.S. A Single Gene Cluster for Chalcomycins and Aldgamycins: Genetic Basis for Bifurcation of Their Biosynthesis. ChemBioChem 2016, 17, 1241–1249. [Google Scholar] [CrossRef] [PubMed]
- Frohardt, R.P.; Pittillo, R.F.; Ehrlich, J. Chalcomycin and Its Fermentative Production. U.S. Patent US 3065137, 20 November 1962. [Google Scholar]
- Hauske, J.R.; Dibrino, J.; Guadliana, M.; Kostek, G. Structure Elucidation of a New Neutral Macrolide Antibiotic. J. Org. Chem. 1986, 51, 2808–2814. [Google Scholar] [CrossRef]
- Kim, S.D.; Ryoo, I.J.; Kim, C.J.; Kim, W.G.; Kim, J.P.; Kong, J.Y.; Koshino, H.; Uramoto, M.; Yoo, I.D. GERI-155, a New Macrolide Antibiotic Related to Chalcomycin. J. Antibiot. 1996, 49, 955–957. [Google Scholar] [CrossRef] [PubMed]
- Goo, Y.M.; Lee, Y.Y.; Kim, B.T. A New 16-membered Chalcomycin Type Macrolide Antibiotic, 250-144C. J. Antibiot. 1997, 50, 85–88. [Google Scholar] [CrossRef] [PubMed]
- Asolkar, R.N.; Maskey, R.P.; Helmke, E.; Laatsch, H. Chalcomycin B, a New Macrolide Antibiotic from the Marine Isolate Streptomyces sp. B7064. J. Antibiot. 2002, 55, 893–898. [Google Scholar] [CrossRef] [PubMed]
- Ding, L.; Qin, S.; Li, F.; Zhang, W.; Lachi, H. Method for Preparation of Chalcomycin-like Compounds from Marine Streptomyces and Its Application as Antitumor Agents. C.N. Patent CN 101624413, 2010. [Google Scholar]
- Morisaki, N.; Hashimoto, Y.; Furihata, K.; Yazawa, K.; Tamura, M.; Mikami, Y. Glycosylative Inactivation of Chalcomycin and Tylosin by a Clinically Isolated Nocardia asteroides Strain. J. Antibiot. 2001, 54, 157–165. [Google Scholar] [CrossRef] [PubMed]
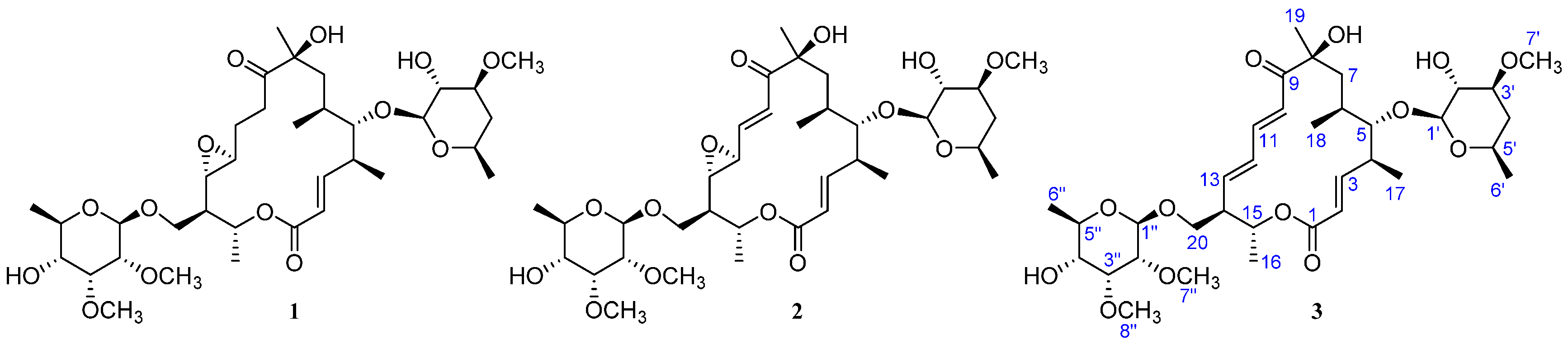
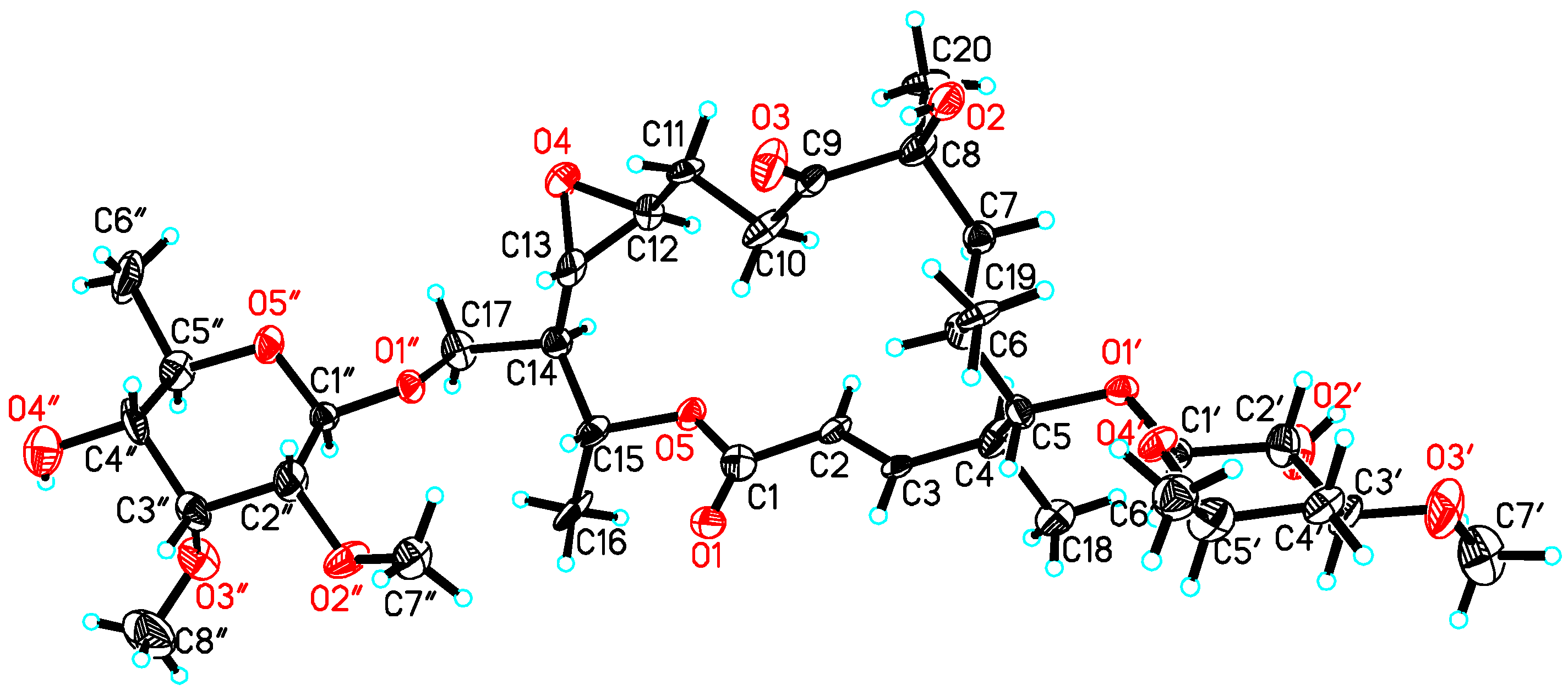

| Position | δC, Mult. | δH (J in Hz) § | 1H, 1H-COSY | HMBC | ROESY |
|---|---|---|---|---|---|
| aglycone | |||||
| 1 | 165.6, C | – | – | – | – |
| 2 | 121.4, CH | 5.75 d (15.4) | 3 | 4 | 4, 17 |
| 3 | 151.7, CH | 6.62 dd (15.4, 9.5) | 2, 4 | 1 | 5, 6, 17 |
| 4 | 41.0, CH | 2.66 | 3, 5, 17 | 2, 3 | 2, 6, 7a, 7b |
| 5 | 88.1, CH | 3.19 | 4, 6 | 3, 4, 6, 7, 17, 18, 1′ | 3, 6, 17, 18, 1′ |
| 6 | 34.0, CH | 1.30 | 5, 7a, 7b, 18 | – | 3, 4, 5, 7a, 10 |
| 7 | 37.4, CH2 | 1.89, Ha | 6, 7b | 6, 8, 9, 18 | 4, 6, 10, 19 |
| 1.83, Hb | 6, 7a | 6, 18 | 4, 18, 19 | ||
| 8 | 78.3, C | – | – | – | – |
| 9 | 202.0, C | – | – | – | – |
| 10 | 122.0, CH | 6.18 d (15.1) | 11 | 8, 9, 11, 12 | 6, 7a, 19 |
| 11 | 144.1, CH | 7.30 dd (15.1, 10.1) | 10, 12 | 9, 12, 13 | – |
| 12 | 133.0, CH | 6.15 dd (14.1, 10.1) | 11, 13 | 10, 11, 13, 14 | – |
| 13 | 143.3, CH | 6.14 dd (14.1, 9.2) | 12, 14 | 11, 12, 14, 20 | 15, 20b |
| 14 | 51.2, CH | 2.47 | 13, 15, 20a, 20b | 12, 13, 15 | 16, 20a |
| 15 | 69.2, CH | 5.06 dq (10.2, 6.2) | 14, 16 | 1, 13, 14 | 13, 20a, 20b |
| 16 | 18.6, CH3 | 1.36 d (6.3) | 15 | 14, 15 | 14, 20a, 20b |
| 17 | 19.2, CH3 | 1.18 d (6.9) | 4 | 3, 4, 5 | 2, ,3, 5, 1′ |
| 18 | 19.3, CH3 | 1.00 d (6.9) | 6 | 5, 6, 7 | 5, 7b |
| 19 | 27.9, CH3 | 1.38 s | – | 7, 8, 9 | 7a, 7b, 10 |
| 20 | 68.4, CH2 | 4.04 dd (9.6, 3.7), Ha | 14, 20b | 13, 14, 15, 1″ | 14, 15, 16, 20b, 1″ |
| 3.57 dd (9.6, 6.1), Hb | 14, 20a | 13, 14, 15, 1″ | 13, 15, 16, 20a, 1″ | ||
| β-d-chalcose unit | |||||
| 1′ | 103.0, CH | 4.19 d (7.6) | 2′ | 5, 5′ | 5, 17, 3′, 5′ |
| 2′ | 75.1, CH | 3.32 dd (8.8, 7.6) | 1′, 3′ | 1′, 3′, 4′ | 4′b |
| 3′ | 80.4, CH | 3.22 | 2′, 4′a, 4′b | 1′, 2′, 4′, 7′ | 1′, 4′a, 5′ |
| 4′ | 36.8, CH2 | 2.04 ddd (12.7, 4.9, 1.9), Ha | 3′, 4′b, 5′ | 2′, 3′ | 3′, 5′, 6′ |
| 1.25, Hb | 3′, 4′a, 5′ | 2′, 3′, 5′ | 2′, 6′ | ||
| 5′ | 67.8, CH | 3.48 | 4′a, 4′b, 6′ | 1′ | 1′, 3′, 4′a |
| 6′ | 20.9, CH3 | 1.23 d (6.2) | 5′ | 4′, 5′ | 4′a, 4′b |
| 7′ | 56.7, CH3 | 3.41 s | – | 3′ | – |
| β-d-mycinose unit | |||||
| 1″ | 101.1, CH | 4.58 d (7.8) | 2″ | 20, 3″, 5″ | 20a, 20b, 5″, 8″ |
| 2″ | 81.9, CH | 3.04 dd (7.8, 3.1) | 1″, 3″ | 1″, 7″ | 3″, 4″, 7″ |
| 3″ | 79.8, CH | 3.76 t (3.1) | 2″, 4″ | 1″, 2″, 4″, 5″, 8″ | 2″, 4″, 8″ |
| 4″ | 72.7, CH | 3.18 | 3″, 5″ | 2″ | 2″, 3″, 6″ |
| 5″ | 70.6, CH | 3.52 | 4″, 6″ | 3″, 4″ | 1″ |
| 6″ | 17.8, CH3 | 1.27 d (6.2) | 5″ | 4″, 5″ | 4″ |
| 7″ | 59.8, CH3 | 3.52 s | – | 2″ | 2″ |
| 8″ | 61.8, CH3 | 3.62 s | – | 3″ | 1″, 3″ |
| Compound | Bacteria | Fungi | ||
|---|---|---|---|---|
| S. aureus | E. coli | C. albicans | A. niger | |
| 1 | 32 | >512 | >512 | >512 |
| 2 | 4 | >512 | >512 | >512 |
| 3 | >512 | >512 | >512 | >512 |
| Tobramycin | 0.4 | 2 | NT | NT |
| Actidione | NT | NT | 64 | 32 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jiang, S.; Zhang, L.; Pei, X.; Deng, F.; Hu, D.; Chen, G.; Wang, C.; Hong, K.; Yao, X.; Gao, A.H. Chalcomycins from Marine-Derived Streptomyces sp. and Their Antimicrobial Activities. Mar. Drugs 2017, 15, 153. https://doi.org/10.3390/md15060153
Jiang S, Zhang L, Pei X, Deng F, Hu D, Chen G, Wang C, Hong K, Yao X, Gao AH. Chalcomycins from Marine-Derived Streptomyces sp. and Their Antimicrobial Activities. Marine Drugs. 2017; 15(6):153. https://doi.org/10.3390/md15060153
Chicago/Turabian StyleJiang, Shutai, Lili Zhang, Xuechang Pei, Fang Deng, Dan Hu, Guodong Chen, Chuanxi Wang, Kui Hong, Xinsheng Yao, and And Hao Gao. 2017. "Chalcomycins from Marine-Derived Streptomyces sp. and Their Antimicrobial Activities" Marine Drugs 15, no. 6: 153. https://doi.org/10.3390/md15060153
APA StyleJiang, S., Zhang, L., Pei, X., Deng, F., Hu, D., Chen, G., Wang, C., Hong, K., Yao, X., & Gao, A. H. (2017). Chalcomycins from Marine-Derived Streptomyces sp. and Their Antimicrobial Activities. Marine Drugs, 15(6), 153. https://doi.org/10.3390/md15060153





