Epigenetic Marks, DNA Damage Markers, or Both? The Impact of Desiccation and Accelerated Aging on Nucleobase Modifications in Plant Genomic DNA
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Material and Treatments
2.2. Viability Assessment
2.3. ROS Detection
2.4. Measurement of Total and Non-Protein Antioxidant Capacity
2.5. DNA Isolation and Measurement of DNA Modifications
2.6. Statistical Analysis
3. Results
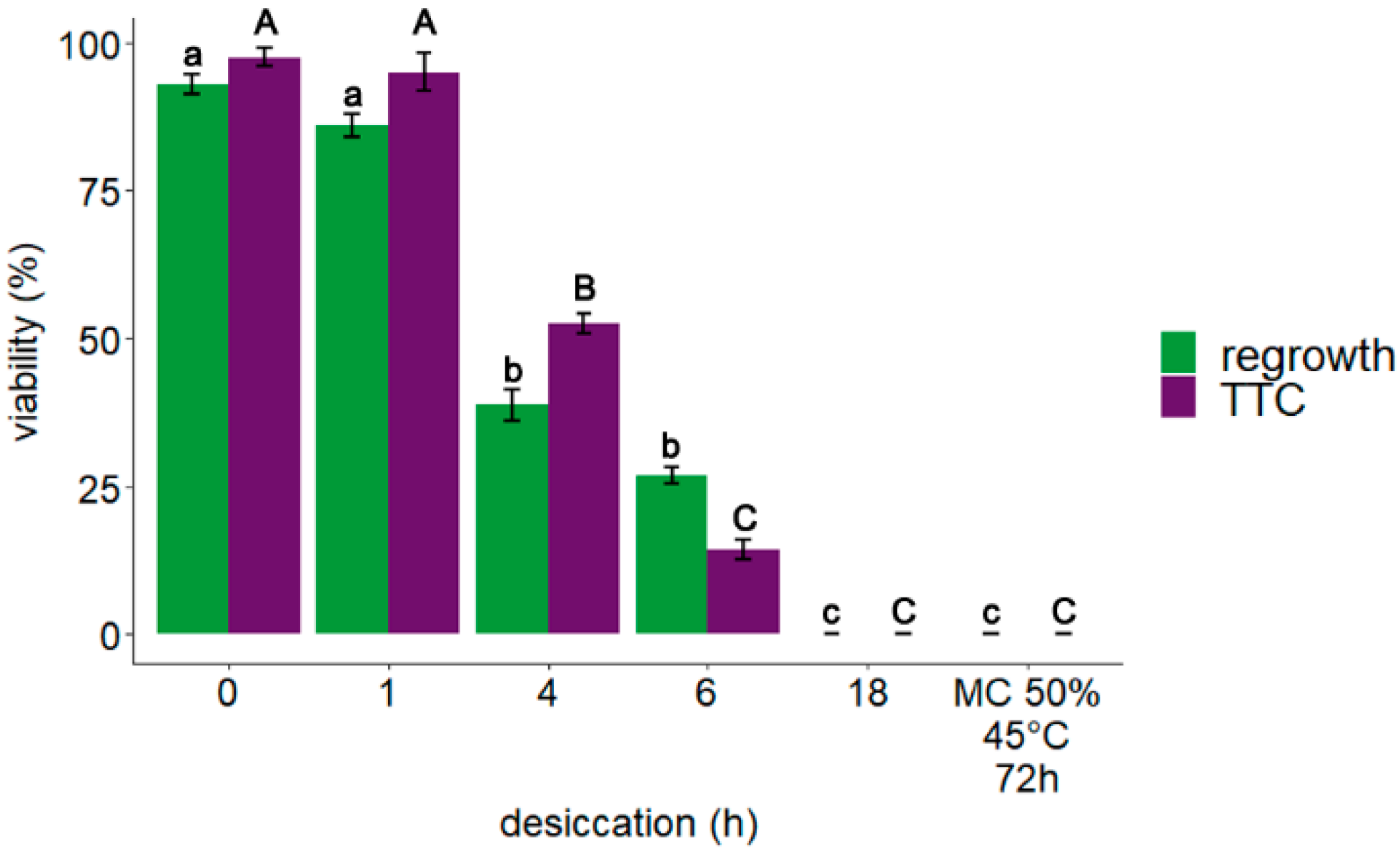
3.1. Explant Viability: TTC Assay and Regrowth
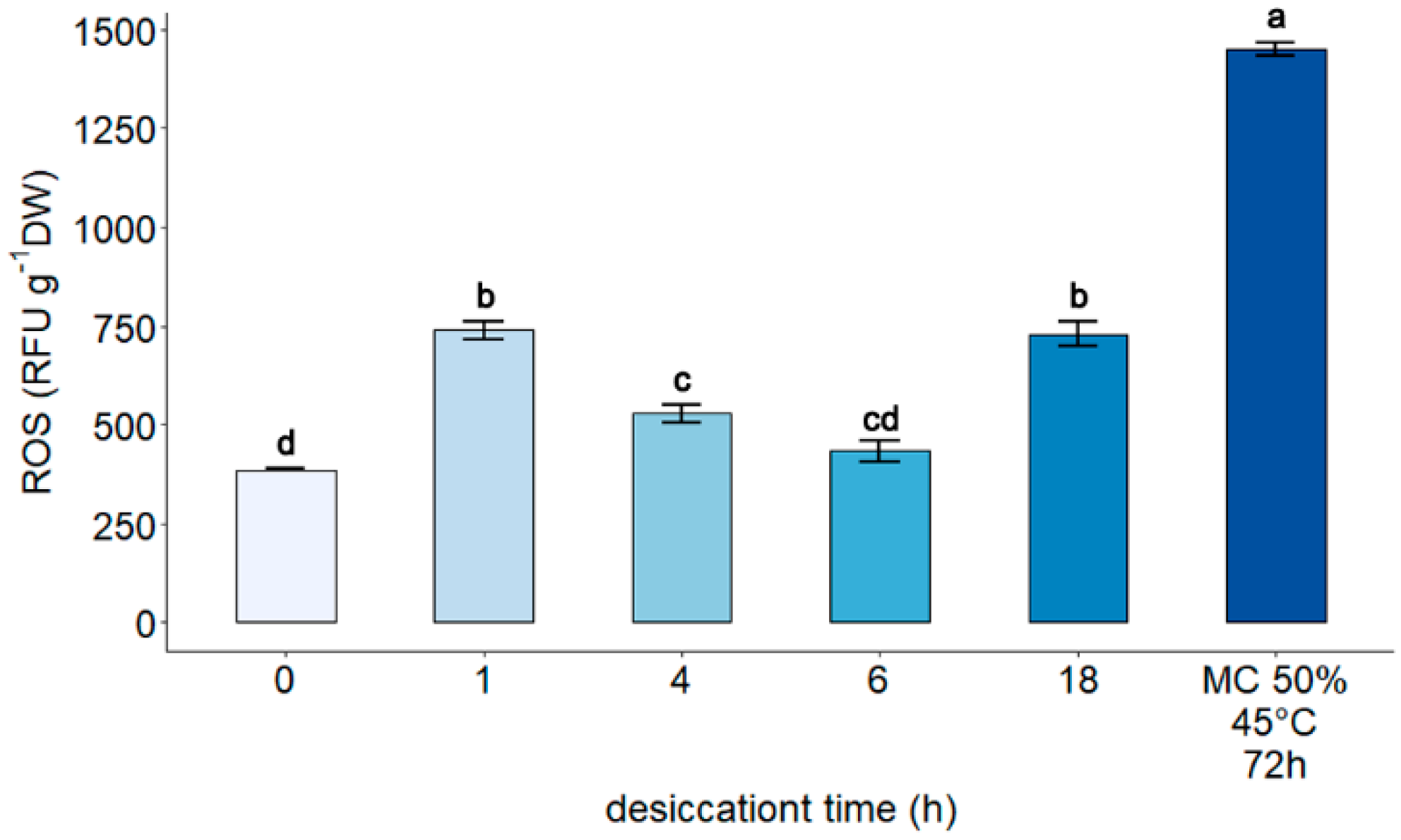
3.2. Measurement of Reactive Oxygen Species
3.3. Measurement of Total Antioxidant Capacity and Non-Protein Antioxidant Capacity
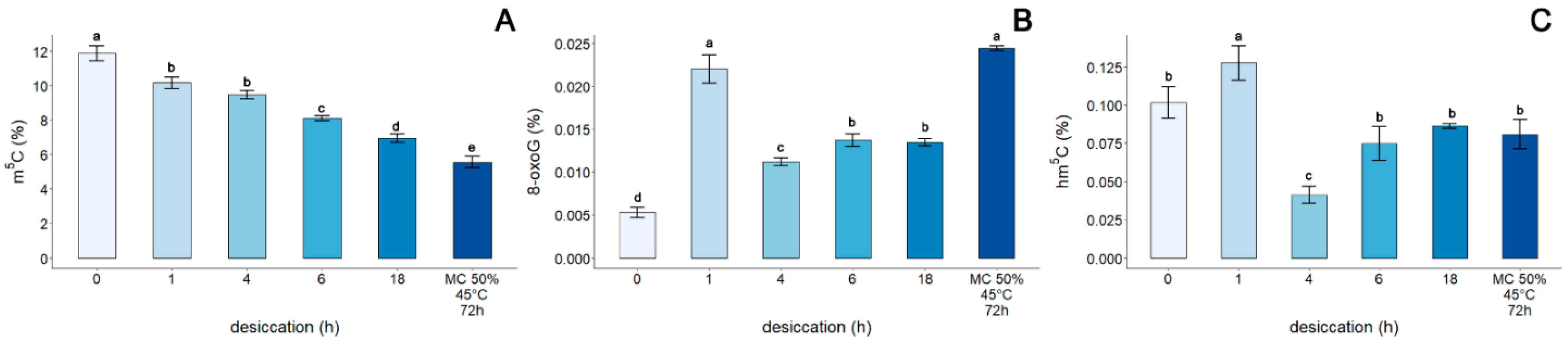
3.4. Assessment of the Genomic Level of m5C, 8-oxoG, and hm5C in the DNA of Embryonic Axes of Acer pseudoplatanus
3.5. Correlation and Principal Component Analyses
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Gorini, F.; Scala, G.; Cooke, M.S.; Majello, B.; Amente, S. Towards a Comprehensive View of 8-Oxo-7,8-Dihydro-2′-Deoxyguanosine: Highlighting the Intertwined Roles of DNA Damage and Epigenetics in Genomic Instability. DNA Repair 2021, 97, 103027. [Google Scholar] [CrossRef] [PubMed]
- Bilyard, M.K.; Becker, S.; Balasubramanian, S. Natural, Modified DNA Bases. Curr. Opin Chem. Biol. 2020, 57, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Ruppel, W.G. Zur Chemie der Tuberkelbacillen. Hoppe-Seyler’S Z. Für Physiol. Chem. 1899, 26, 218–232. [Google Scholar] [CrossRef] [Green Version]
- Hotchkiss, R.D. The Quantitative Separation Of Purines, Pyrimidines, and Nucleosides by Paper Chromatography. J. Biol. Chem. 1948, 175, 315–332. [Google Scholar] [CrossRef]
- Wang, X.; Song, S.; Wu, Y.-S.; Li, Y.-L.; Chen, T.; Huang, Z.; Liu, S.; Dunwell, T.L.; Pfeifer, G.P.; Dunwell, J.M.; et al. Genome-Wide Mapping of 5-Hydroxymethylcytosine in Three Rice Cultivars Reveals Its Preferential Localization in Transcriptionally Silent Transposable Element Genes. J. Exp. Bot. 2015, 66, 6651–6663. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shi, D.-Q.; Ali, I.; Tang, J.; Yang, W.-C. New Insights into 5hmC DNA Modification: Generation, Distribution and Function. Front. Genet. 2017, 8, 100. [Google Scholar] [CrossRef]
- Tang, Y.; Xiong, J.; Jiang, H.-P.; Zheng, S.-J.; Feng, Y.-Q.; Yuan, B.-F. Determination of Oxidation Products of 5-Methylcytosine in Plants by Chemical Derivatization Coupled with Liquid Chromatography/Tandem Mass Spectrometry Analysis. Anal. Chem. 2014, 86, 7764–7772. [Google Scholar] [CrossRef]
- Chen, T.; Li, E. Structure and function of eukaryotic DNA methyltransferases. Curr. Top. Dev. Biol. 2004, 60, 55–89. [Google Scholar] [CrossRef]
- Plitta, B.P.; Michalak, M.; Bujarska-Borkowska, B.; Barciszewska, M.Z.; Barciszewski, J.; Chmielarz, P. Effect of Desiccation on the Dynamics of Genome-Wide DNA Methylation in Orthodox Seeds of Acer platanoides L. Plant Physiol. Biochem. 2014, 85, 71–77. [Google Scholar] [CrossRef]
- Plitta-Michalak, B.P.; Naskręt-Barciszewska, M.Z.; Kotlarski, S.; Tomaszewski, D.; Tylkowski, T.; Barciszewski, J.; Chmielarz, P.; Michalak, M. Changes in Genomic 5-Methylcytosine Level Mirror the Response of Orthodox (Acer platanoides L.) and Recalcitrant (Acer pseudoplatanus L.) Seeds to Severe Desiccation. Tree Physiol. 2018, 38, 617–629. [Google Scholar] [CrossRef] [Green Version]
- Michalak, M.; Plitta-Michalak, B.; Nasktęt-Barciszewska, M.; Barciszewski, J.; Bujarska-Borkowska, B.; Chmielarz, P. Global 5-Methylcytosine Alterations in DNA during Ageing of Quercus robur Seeds. Ann. Bot-London 2015, 116. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Plitta, B.P.; Michalak, M.; Naskręt-Barciszewska, M.Z.; Barciszewski, J.; Chmielarz, P. DNA Methylation of Quercus robur L. Plumules Following Cryo-Pretreatment and Cryopreservation. Plant Cell Tiss. Organ Cult. 2014, 117, 31–37. [Google Scholar] [CrossRef]
- Zhang, H.; Lang, Z.; Zhu, J.-K. Dynamics and Function of DNA Methylation in Plants. Nat. Rev. Mol. Cell Biol. 2018, 19, 489–506. [Google Scholar] [CrossRef] [PubMed]
- Smolikova, G.; Leonova, T.; Vashurina, N.; Frolov, A.; Medvedev, S. Desiccation Tolerance as the Basis of Long-Term Seed Viability. Int. J. Mol. Sci. 2021, 22, 101. [Google Scholar] [CrossRef]
- Kurek, K.; Plitta-Michalak, B.; Ratajczak, E. Reactive Oxygen Species as Potential Drivers of the Seed Aging Process. Plants 2019, 8, 174. [Google Scholar] [CrossRef] [Green Version]
- Sharma, P.; Jha, A.B.; Dubey, R.S.; Pessarakli, M. Reactive Oxygen Species, Oxidative Damage, and Antioxidative Defense Mechanism in Plants under Stressful Conditions. J. Bot. 2012, 2012, 1–26. [Google Scholar] [CrossRef] [Green Version]
- Adetunji, A.E.; Adetunji, T.L.; Varghese, B.; Sershen; Pammenter, N.W. Oxidative Stress, Ageing and Methods of Seed Invigoration: An Overview and Perspectives. Agronomy 2021, 11, 2369. [Google Scholar] [CrossRef]
- Lee, T.-H.; Kang, T.-H. DNA Oxidation and Excision Repair Pathways. Int. J. Mol. Sci. 2019, 20, 6092. [Google Scholar] [CrossRef] [Green Version]
- Karanjawala, Z.E.; Murphy, N.; Hinton, D.R.; Hsieh, C.-L.; Lieber, M.R. Oxygen Metabolism Causes Chromosome Breaks and Is Associated with the Neuronal Apoptosis Observed in DNA Double-Strand Break Repair Mutants. Curr. Biol. 2002, 12, 397–402. [Google Scholar] [CrossRef] [Green Version]
- Fenton, H.J.H. Oxidation of Tartaric Acid in Presence of Iron. J. Chem. Soc. Trans. 1894, 65, 899–910. [Google Scholar] [CrossRef] [Green Version]
- Fleming, A.M.; Burrows, C. On the Irrelevancy of Hydroxyl Radical to DNA Damage from Oxidative Stress and Implications for Epigenetics. Chem. Soc. Rev. 2020, 49, 6524–6528. [Google Scholar] [CrossRef] [PubMed]
- Fleming, A.M.; Burrows, C.J. Iron Fenton Oxidation of 2′-Deoxyguanosine in Physiological Bicarbonate Buffer Yields Products Consistent with the Reactive Oxygen Species Carbonate Radical Anion Not the Hydroxyl Radical. Chem. Commun. 2020, 56, 9779–9782. [Google Scholar] [CrossRef] [PubMed]
- Chatgilialoglu, C.; Ferreri, C.; Krokidis, M.G.; Masi, A.; Terzidis, M.A. On the Relevance of Hydroxyl Radical to Purine DNA Damage. Free Radic. Res. 2021, 55, 384–404. [Google Scholar] [CrossRef] [PubMed]
- Yin, R.; Zhang, D.; Song, Y.; Zhu, B.-Z.; Wang, H. Potent DNA Damage by Polyhalogenated Quinones and H2O2 via a Metal-Independent and Intercalation-Enhanced Oxidation Mechanism. Sci. Rep. 2013, 3, 1269. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jena, N.R.; Mishra, P.C. Formation of 8-Nitroguanine and 8-Oxoguanine due to Reactions of Peroxynitrite with Guanine. J. Comput. Chem. 2007, 28, 1321–1335. [Google Scholar] [CrossRef] [PubMed]
- Halliwell, B.; Adhikary, A.; Dingfelder, M.; Dizdaroglu, M. Hydroxyl Radical Is a Significant Player in Oxidative DNA Damage in Vivo. Chem. Soc. Rev. 2021, 50, 8355–8360. [Google Scholar] [CrossRef]
- Illés, E.; Mizrahi, A.; Marks, V.; Meyerstein, D. Carbonate-Radical-Anions, and Not Hydroxyl Radicals, Are the Products of the Fenton Reaction in Neutral Solutions Containing Bicarbonate. Free Radical Biol. Med. 2019, 131, 1–6. [Google Scholar] [CrossRef]
- Illés, E.; Patra, S.G.; Marks, V.; Mizrahi, A.; Meyerstein, D. The FeII(Citrate) Fenton Reaction under Physiological Conditions. J. Inorg. Biochem. 2020, 206, 111018. [Google Scholar] [CrossRef]
- Giorgio, M.; Dellino, G.I.; Gambino, V.; Roda, N.; Pelicci, P.G. On the Epigenetic Role of Guanosine Oxidation. Redox Biol. 2020, 29, 101398. [Google Scholar] [CrossRef]
- Wyatt, G.R.; Cohen, S.S. A New Pyrimidine Base from Bacteriophage Nucleic Acids. Nature 1952, 170, 1072–1073. [Google Scholar] [CrossRef]
- Penn, N.W.; Suwalski, R.; O’Riley, C.; Bojanowski, K.; Yura, R. The Presence of 5-Hydroxymethylcytosine in Animal Deoxyribonucleic Acid. Biochem. J. 1972, 126, 781–790. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kriaucionis, S.; Heintz, N. The Nuclear DNA Base 5-Hydroxymethylcytosine Is Present in Purkinje Neurons and the Brain. Science 2009, 324, 929–930. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tahiliani, M.; Koh, K.P.; Shen, Y.; Pastor, W.A.; Bandukwala, H.; Brudno, Y.; Agarwal, S.; Iyer, L.M.; Liu, D.R.; Aravind, L.; et al. Conversion of 5-Methylcytosine to 5-Hydroxymethylcytosine in Mammalian DNA by MLL Partner TET1. Science 2009, 324, 930–935. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yakovlev, I.A.; Gackowski, D.; Abakir, A.; Viejo, M.; Ruzov, A.; Olinski, R.; Starczak, M.; Fossdal, C.G.; Krutovsky, K. Mass Spectrometry Reveals the Presence of Specific Set of Epigenetic DNA Modifications in the Norway Spruce Genome. Sci. Rep. 2019, 9, 19314. [Google Scholar] [CrossRef] [PubMed]
- Rausch, C.; Hastert, F.D.; Cardoso, M.C. DNA Modification Readers and Writers and Their Interplay. J. Mol. Biol. 2020, 432, 1731–1746. [Google Scholar] [CrossRef] [PubMed]
- Yao, Q.; Song, C.-X.; He, C.; Kumaran, D.; Dunn, J.J. Heterologous Expression and Purification of Arabidopsis Thaliana VIM1 Protein: In Vitro Evidence for Its Inability to Recognize Hydroxymethylcytosine, a Rare Base in Arabidopsis DNA. Protein Expr. Purif. 2012, 83, 104–111. [Google Scholar] [CrossRef]
- Terragni, J.; Bitinaite, J.; Zheng, Y.; Pradhan, S. Correction to Biochemical Characterization of Recombinant β-Glucosyltransferase and Analysis of Global 5-Hydroxymethylcytosine in Unique Genomes. Biochemistry 2012, 51, 6261. [Google Scholar] [CrossRef]
- Liu, S.; Dunwell, T.L.; Pfeifer, G.P.; Dunwell, J.M.; Ullah, I.; Wang, Y. Detection of Oxidation Products of 5-Methyl-2′-Deoxycytidine in Arabidopsis DNA. PLoS ONE 2013, 8, e84620. [Google Scholar] [CrossRef]
- Wang, W.; Huang, F.; Qin, Q.; Zhao, X.; Li, Z.; Fu, B. Comparative Analysis of DNA Methylation Changes in Two Rice Genotypes under Salt Stress and Subsequent Recovery. Biochem. Biophys. Res. Commun. 2015, 465, 790–796. [Google Scholar] [CrossRef] [Green Version]
- Mahmood, A.M.; Dunwell, J.M. Evidence for Novel Epigenetic Marks within Plants. AIMS Genet. 2019, 06, 070–087. [Google Scholar] [CrossRef]
- Erdmann, R.M.; Souza, A.L.; Clish, C.B.; Gehring, M. 5-Hydroxymethylcytosine Is Not Present in Appreciable Quantities in Arabidopsis DNA. G3-Genes Genomes Genet. 2015, 5, 1–8. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Madugundu, G.S.; Cadet, J.; Wagner, J.R. Hydroxyl-Radical-Induced Oxidation of 5-Methylcytosine in Isolated and Cellular DNA. Nucleic Acids Res. 2014, 42, 7450–7460. [Google Scholar] [CrossRef] [PubMed]
- Plitta-Michalak, B.P.; Ramos, A.A.; Pupel, P.; Michalak, M. Oxidative Damage and DNA Repair in Desiccated Recalcitrant Embryonic Axes of Acer pseudoplatanus L. BMC Plant Biol. 2022, 22, 40. [Google Scholar] [CrossRef] [PubMed]
- Michalak, M.; Barciszewska, M.Z.; Barciszewski, J.; Plitta, B.P.; Chmielarz, P. Global Changes in DNA Methylation in Seeds and Seedlings of Pyrus Communis after Seed Desiccation and Storage. PLoS ONE 2013, 8, e70693. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- International Rules for Seed Testing 2021-ISTA Online-International Seed Testing Association. Available online: https://www.seedtest.org/en/international-rules-for-seed-testing-_content---1--1083.html (accessed on 1 March 2022).
- Tylkowski, T. Short-Term Storage of after Ripened Seeds of Acer platanoides L. and Acer pseudoplatanus L. Arbor. Kórnickie 1989, 34, 135–141. [Google Scholar]
- Tylkowski, T. Przedsiewne Traktowanie Nasion Drzew, Krzewów, Pnączy i Krzewinek; Centrum Informacyjne Lasów Państwowych: Warszawa, Poland, 2016; ISBN 978-83-63895-19-8. [Google Scholar]
- Hong, T.D.; Ellis, R.H. A Comparison of Maturation Drying, Germination, and Desiccation Tolerance between Developing Seeds of Acer pseudoplatanus L. and Acer platanoides L. New Phytol. 1990, 116, 589–596. [Google Scholar] [CrossRef]
- Daws, M.I.; Cleland, H.; Chmielarz, P.; Gorian, F.; Leprince, O.; Mullins, C.E.; Thanos, C.A.; Vandvik, V.; Pritchard, H.W.; Daws, M.I.; et al. Variable Desiccation Tolerance in Acer Pseudoplatanus Seeds in Relation to Developmental Conditions: A Case of Phenotypic Recalcitrance? Funct. Plant Biol. 2006, 33, 59–66. [Google Scholar] [CrossRef] [Green Version]
- Berjak, P.; Pammenter, N.W. Implications of the Lack of Desiccation Tolerance in Recalcitrant Seeds. Front. Plant. Sci. 2013, 4, 478. [Google Scholar] [CrossRef] [Green Version]
- Roberts, E.H. Predicting the Storage Life of Seeds. Seed Sci. Technol. 1973, 1, 499–514. [Google Scholar]
- Plitta-Michalak, B.; Michalak, M.; Kotlarski, S.; Chmielarz, P. Cryogenic Storage of Seeds. Sylwan 2013, 157, 723–729. [Google Scholar]
- Pence, V.C.; Ballesteros, D.; Walters, C.; Reed, B.M.; Philpott, M.; Dixon, K.W.; Pritchard, H.W.; Culley, T.M.; Vanhove, A.-C. Cryobiotechnologies: Tools for Expanding Long-Term Ex Situ Conservation to All Plant Species. Biol. Conserv. 2020, 250, 108736. [Google Scholar] [CrossRef]
- Walters, C.; Pence, V.C. The Unique Role of Seed Banking and Cryobiotechnologies in Plant Conservation. Plants People Planet 2021, 3, 83–91. [Google Scholar] [CrossRef]
- Moricová, P.; Ondřej, V.; Navrátilová, B.; Luhová, L. Changes of DNA Methylation and Hydroxymethylation in Plant Protoplast Cultures. Acta Biochim. Pol. 2013, 60, 33–36. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Visscher, A.M.; Latorre Frances, A.; Yeo, M.; Yan, J.; Colville, L.; Gomez Barreiro, P.; Pritchard, H.W. Comparative Analyses of Extreme Dry Seed Thermotolerance in Five Cactaceae Species. Environ. Exp. Bot. 2021, 188, 104514. [Google Scholar] [CrossRef]
- Golubov, A.; Kovalchuk, I. Analysis of DNA Hydroxymethylation Using Colorimetric Assay. In Plant Epigenetics: Methods and Protocols, 2nd ed.; Kovalchuk, I., Ed.; Humana Press Inc.: Totowa, Japan, 2017; Volume 1456, pp. 89–97. ISBN 978-1-4899-7708-3. [Google Scholar]
- Bernstein, B.E.; Meissner, A.; Lander, E.S. The Mammalian Epigenome. Cell 2007, 128, 669–681. [Google Scholar] [CrossRef] [Green Version]
- Mira, S.; Pirredda, M.; Martín-Sánchez, M.; Marchessi, J.; Martín, C. DNA methylation and integrity in aged seeds and regenerated plants. Seed Sci. Res. 2020, 30, 92–100. [Google Scholar] [CrossRef]
- Pirredda, M.; González-Benito, M.E.; Martín, C.; Mira, S. Genetic and Epigenetic Stability in Rye Seeds under Different Storage Conditions: Ageing and Oxygen Effect. Plants 2020, 9, 393. [Google Scholar] [CrossRef] [Green Version]
- Plitta-Michalak, B.P.; Naskręt-Barciszewska, M.Z.; Barciszewski, J.; Chmielarz, P.; Michalak, M. Epigenetic Integrity of Orthodox Seeds Stored under Conventional and Cryogenic Conditions. Forests 2021, 12, 288. [Google Scholar] [CrossRef]
- Domej, W.; Oettl, K.; Renner, W. Oxidative Stress and Free Radicals in COPD–Implications and Relevance for Treatment. Int. J. Chron. Obstruct. Pulmon. Dis. 2014, 9, 1207–1224. [Google Scholar] [CrossRef] [Green Version]
- Dianov, G.L.; Souza-Pinto, N.; Nyaga, S.G.; Thybo, T.; Stevnsner, T.; Bohr, V.A. Base Excision Repair in Nuclear and Mitochondrial DNA. Prog. Nucleic Acid Res. Mol. Biol. 2001, 68, 285–297. [Google Scholar] [CrossRef]
- Kiran, K.R.; Deepika, V.B.; Swathy, P.S.; Prasad, K.; Kabekkodu, S.P.; Murali, T.S.; Satyamoorthy, K.; Muthusamy, A. ROS-dependent DNA damage and repair during germination of NaCl primed seeds. J. Photochem. Photobiol. B Biol. 2020, 213, 112050. [Google Scholar] [CrossRef] [PubMed]
- Balestrazzi, A.; Confalonieri, M.; Macovei, A.; Carbonera, D. Seed imbibition in Medicago truncatula Gaertn.: Expression profiles of DNA repair genes in relation to PEG-mediated stress. J. Plant Physiol. 2011, 168, 706–713. [Google Scholar] [CrossRef] [PubMed]
- Chandra, J.; Parkhey, S.; Keshavkant, S. Ageing-regulated changes in genetic integrity of two recalcitrant seeded species having contrasting longevity. Trees 2018, 32, 109–123. [Google Scholar] [CrossRef]
- Chen, H.; Osuna, D.; Colville, L.; Lorenzo, O.; Graeber, K.; Küster, H.; Leubner-Metzger, G.; Kranner, I. Transcriptome-wide mapping of pea seed ageing reveals a pivotal role for genes related to oxidative stress and programmed cell death. PLoS ONE 2013, 8, e78471. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sershen; Varghese, B.; Naidoo, C.; Pammenter, N.W. The Use of Plant Stress Biomarkers in Assessing the Effects of Desiccation in Zygotic Embryos from Recalcitrant Seeds: Challenges and Considerations. Plant Biol. 2016, 18, 433–444. [Google Scholar] [CrossRef] [PubMed]
- Gruber, D.R.; Toner, J.J.; Miears, H.L.; Shernyukov, A.V.; Kiryutin, A.S.; Lomzov, A.A.; Endutkin, A.V.; Grin, I.R.; Petrova, D.V.; Kupryushkin, M.S.; et al. Oxidative Damage to Epigenetically Methylated Sites Affects DNA Stability, Dynamics and Enzymatic Demethylation. Nucleic Acids Res. 2018, 46, 10827–10839. [Google Scholar] [CrossRef]
- Guz, J.; Foksinski, M.; Siomek, A.; Gackowski, D.; Rozalski, R.; Dziaman, T.; Szpila, A.; Olinski, R. The Relationship between 8-Oxo-7,8-Dihydro-2′-Deoxyguanosine Level and Extent of Cytosine Methylation in Leukocytes DNA of Healthy Subjects and in Patients with Colon Adenomas and Carcinomas. Mutat. Res. 2008, 640, 170–173. [Google Scholar] [CrossRef] [PubMed]
- Bhusari, S.S.; Dobosy, J.R.; Fu, V.; Almassi, N.; Oberley, T.; Jarrard, D.F. Superoxide Dismutase 1 Knockdown Induces Oxidative Stress and DNA Methylation Loss in the Prostate. Epigenetics 2010, 5, 402–409. [Google Scholar] [CrossRef] [Green Version]
- Martignago, D.; Bernardini, B.; Polticelli, F.; Salvi, D.; Cona, A.; Angelini, R.; Tavladoraki, P. The Four FAD-Dependent Histone Demethylases of Arabidopsis Are Differently Involved in the Control of Flowering Time. Front. Plant Sci. 2019, 10, 669. [Google Scholar] [CrossRef]
- Castander-Olarieta, A.; Pereira, C.; Sales, E.; Meijon, M.; Arrillaga, I.; Jesus Canal, M.; Goicoa, T.; Dolores Ugarte, M.; Moncalean, P.; Montalban, I.A. Induction of Radiata Pine Somatic Embryogenesis at High Temperatures Provokes a Long-Term Decrease in DNA Methylation/Hydroxymethylation and Differential Expression of Stress-Related Genes. Plants-Basel 2020, 9, 1762. [Google Scholar] [CrossRef]
- Zarakowska, E.; Czerwinska, J.; Tupalska, A.; Yousefzadeh, M.J.; Gregg, S.Q.; Croix, C.M.S.; Niedernhofer, L.J.; Foksinski, M.; Gackowski, D.; Szpila, A.; et al. Oxidation Products of 5-Methylcytosine Are Decreased in Senescent Cells and Tissues of Progeroid Mice. J. Gerontol. A Biol. Sci. 2018, 73, 1003–1009. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ogneva, Z.V.; Dubrovina, A.S.; Kiselev, K.V. Age-Associated Alterations in DNA Methylation and Expression of Methyltransferase and Demethylase Genes in Arabidopsis thaliana. Biol. Plant. 2016, 60, 628–634. [Google Scholar] [CrossRef]
- Wang, W.-S.; Pan, Y.-J.; Zhao, X.-Q.; Dwivedi, D.; Zhu, L.-H.; Ali, J.; Fu, B.-Y.; Li, Z.-K. Drought-Induced Site-Specific DNA Methylation and Its Association with Drought Tolerance in Rice (Oryza sativa L.). J. Exp. Bot. 2011, 62, 1951–1960. [Google Scholar] [CrossRef] [PubMed]
- Jang, H.; Shin, H.; Eichman, B.F.; Huh, J.H. Excision of 5-Hydroxymethylcytosine by DEMETER Family DNA Glycosylases. Biochem. Biophys. Res. Commun. 2014, 446, 1067–1072. [Google Scholar] [CrossRef] [Green Version]



| Desiccation (h) | MC, % (WC, g H2O g−1 Dry Weight) |
|---|---|
| 0 | 50.2 ± 0.08 |
| (1.01 ± 0.03) | |
| 1 | 19.3 ± 0.07 |
| (0.24 ± 0.01) | |
| 4 | 11.9 ± 0.08 |
| (0.13 ± 0.01) | |
| 6 | 9.2 ± 0.4 |
| (0.10 ± 0.01) | |
| 18 | 5.7 ± 1.2 |
| (0.06 ± 0.01) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Plitta-Michalak, B.P.; Litkowiec, M.; Michalak, M. Epigenetic Marks, DNA Damage Markers, or Both? The Impact of Desiccation and Accelerated Aging on Nucleobase Modifications in Plant Genomic DNA. Cells 2022, 11, 1748. https://doi.org/10.3390/cells11111748
Plitta-Michalak BP, Litkowiec M, Michalak M. Epigenetic Marks, DNA Damage Markers, or Both? The Impact of Desiccation and Accelerated Aging on Nucleobase Modifications in Plant Genomic DNA. Cells. 2022; 11(11):1748. https://doi.org/10.3390/cells11111748
Chicago/Turabian StylePlitta-Michalak, Beata P., Monika Litkowiec, and Marcin Michalak. 2022. "Epigenetic Marks, DNA Damage Markers, or Both? The Impact of Desiccation and Accelerated Aging on Nucleobase Modifications in Plant Genomic DNA" Cells 11, no. 11: 1748. https://doi.org/10.3390/cells11111748






