LASSO-Driven Selection of Biochemical and Clinical Markers for Primary Resistance to PD-1 Inhibitors in Metastatic Melanoma
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Population
2.2. Data Collection
2.3. Statistical Analysis
2.4. Management of Missing Data
3. Results
3.1. Patient Characteristics
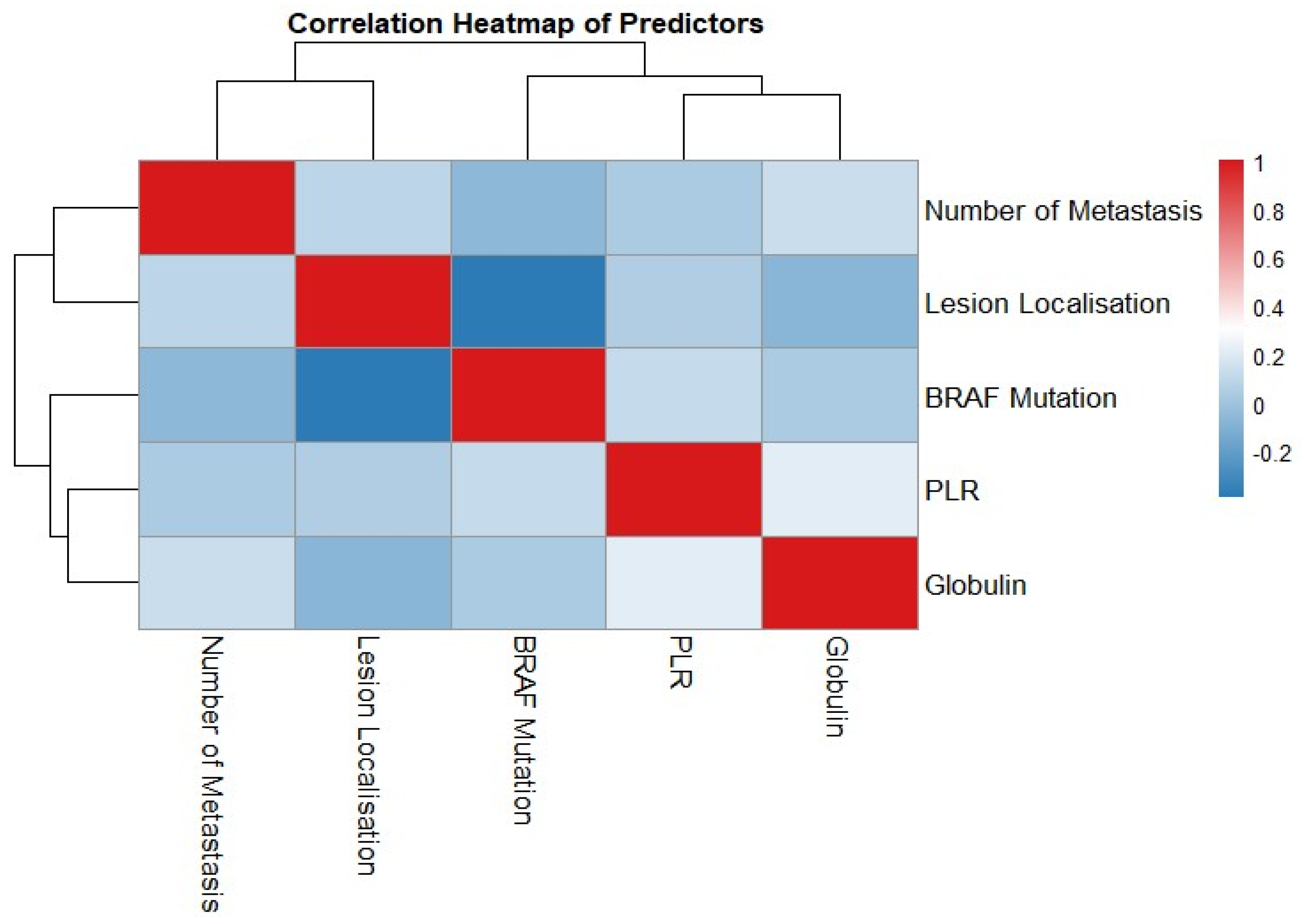
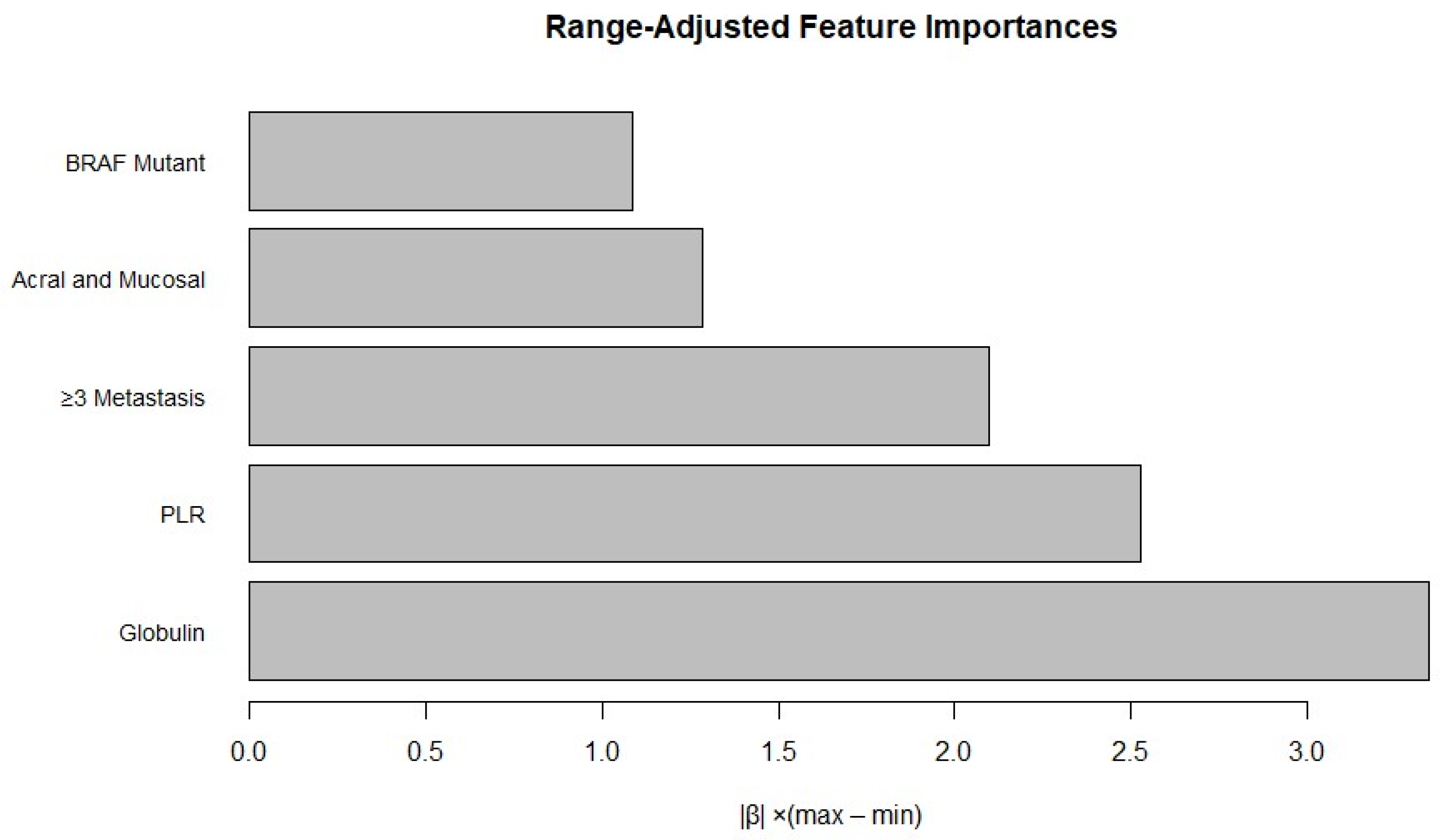
3.2. Markers Selection
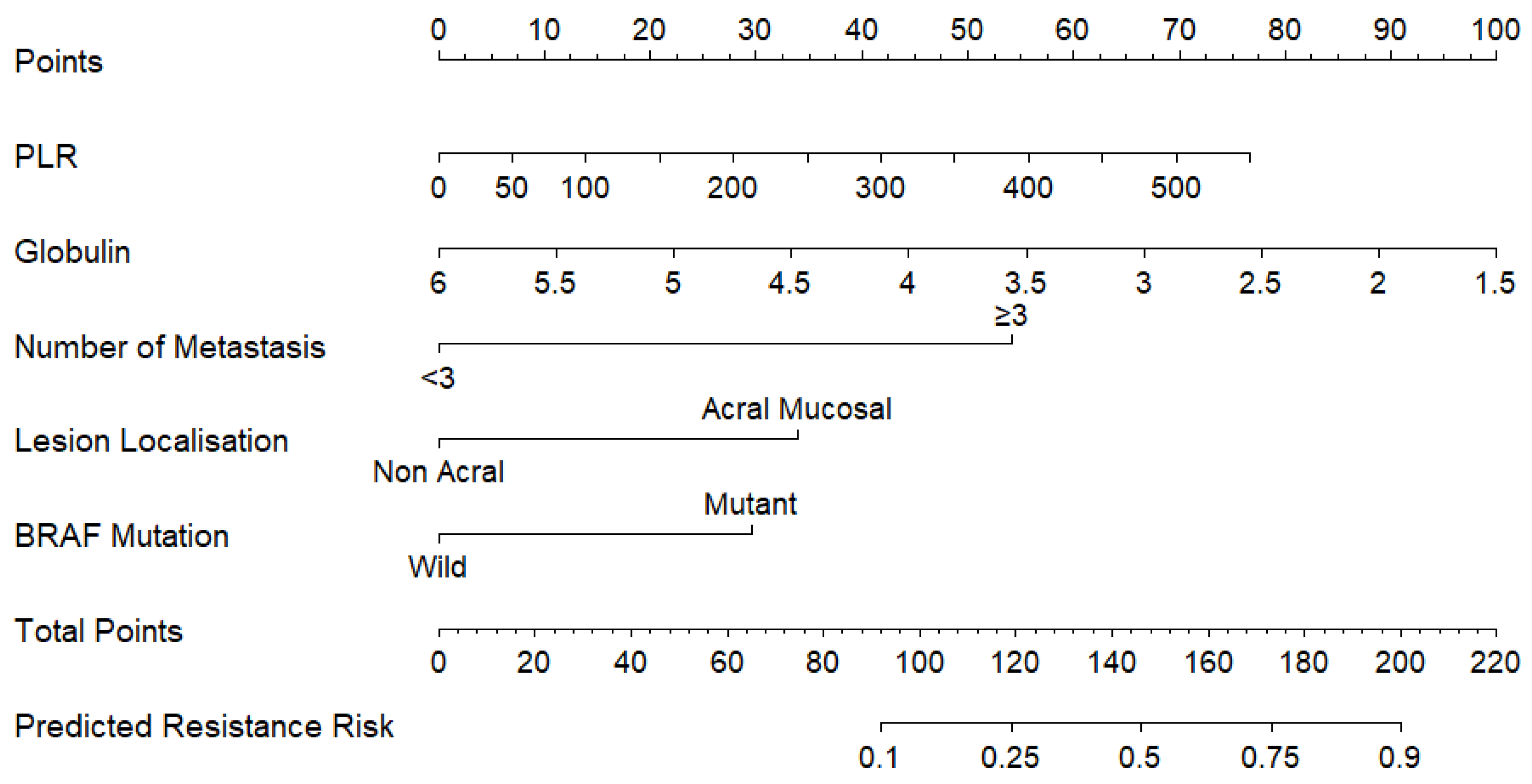
3.3. Construction of Nomogram
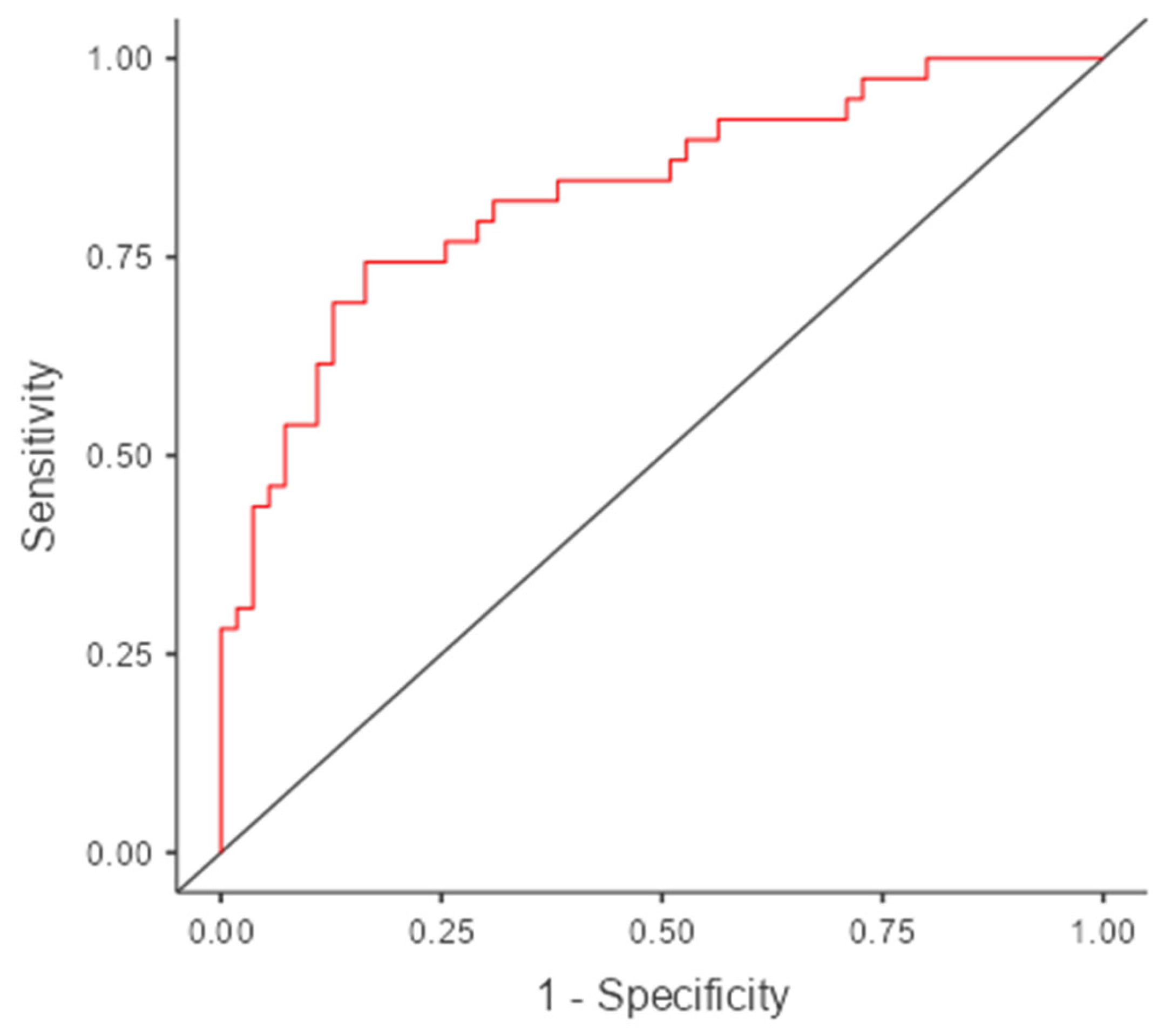
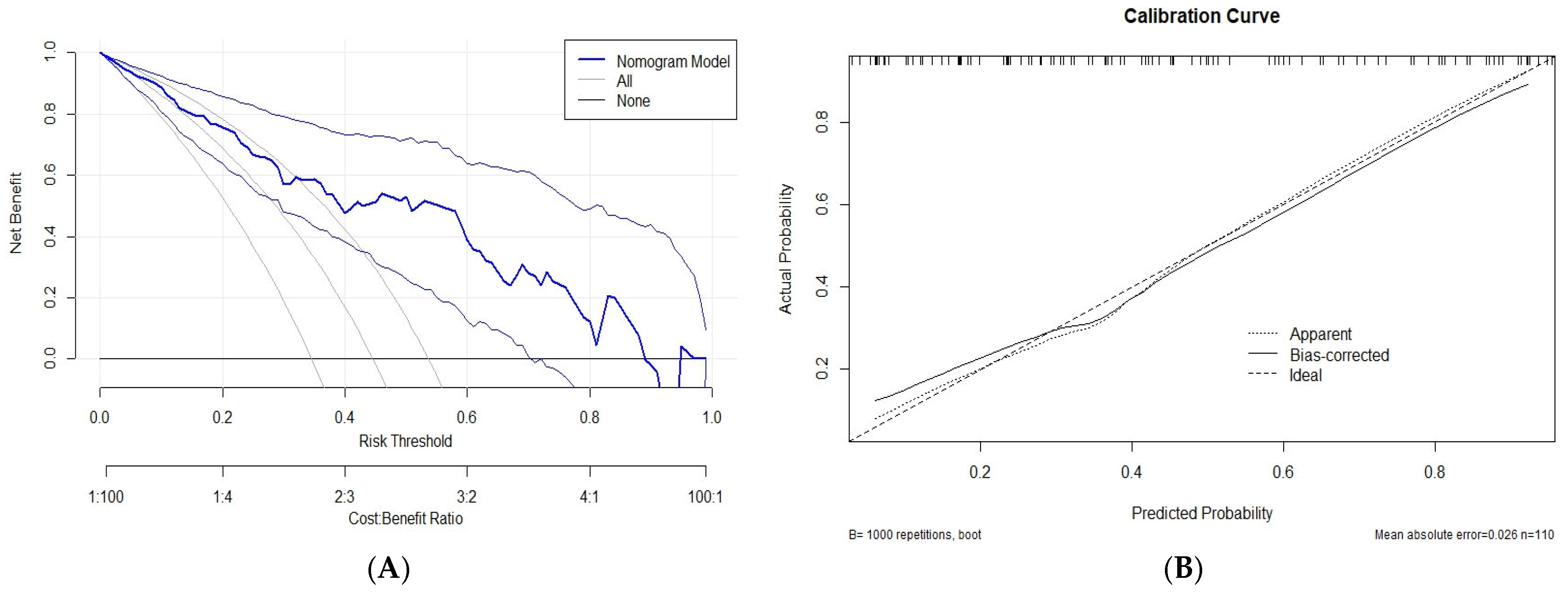
3.4. Model Performance Metrics
4. Discussion
5. Limitation
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
- Lymphocyte
- Monocyte
- Eosinophil
- Mean Platelet Volume
- Lactate Dehydrogenase
- C-Reactive Protein
- Albumin
- Globulin
- Neutrophil-to-Lymphocyte Ratio (NLR)
- Lymphocyte-to-Monocyte Ratio (LMR)
- Platelet-to-Lymphocyte Ratio (PLR)
- Mean Platelet Volume to Lymphocyte Ratio
- Albumin-to-Globulin Ratio
- Scottish Inflammatory Prognostic Score (SIPS)
- Royal Marsden Hospital (RMH) Prognostic Score
- Global Risk Index Model Score (GRIm-Score)
- Combine Immun Index (CII)
- Haemoglobin, Albumin, Lymphocyte, and Platelet (HALP) Score
- MD Anderson Immune Checkpoint Inhibitor Score (MD Anderson Score)
- Prognostic Inflammatory and Nutritional Index (PIV score)
- Pan-Immune Inflammation Value and ECOG (PILE score)
- Prognostic Nutritional Index (PNI)
- Systemic Immune Inflammation Index (SII)
- Systemic Inflammatory Response Index (SIRI)
Appendix B
- Neutrophil-to-Lymphocyte Ratio (NLR): Neutrophil Count/Lymphocyte Count
- Lymphocyte-to-Monocyte Ratio (LMR): Lymphocyte Count/Monocyte Count
- Platelet-to-Lymphocyte Ratio (PLR): Platelet Count/Lymphocyte Count
- Mean Platelet Volume to Lymphocyte Ratio: Mean Platelet Volume (MPV)/Lymphocyte Count
- Albumin-to-Globulin Ratio: Albumin/Globulin
- Scottish Inflammatory Prognostic Score (SIPS):
- If Albumin < 35 g/L and CRP > 10 mg/L, score = 2
- If one is abnormal, score = 1
- If both are normal, score = 0
- Royal Marsden Hospital (RMH) Prognostic Score:
- Albumin < 35 g/L
- Lactate Dehydrogenase (LDH) > Upper Limit of Normal (ULN)
- More than two sites of metastases
- Global Risk Index Model Score (GRIm-Score): Combines Albumin, Haemoglobin, NLR, and BMI.
- Combine Immun Index (CII): (NLR × Platelet Count)/Lymphocyte Count
- Haemoglobin, Albumin, Lymphocyte, and Platelet (HALP) Score: (Haemoglobin × Albumin × Lymphocyte Count)/Platelet Count
- MD Anderson Immune Checkpoint Inhibitor Score: Based on specific biomarkers and inflammatory parameters.
- Prognostic Inflammatory and Nutritional Index (PIV score): (Platelet Count × Neutrophil Count)/(Lymphocyte Count × Albumin)
- Pan-Immune Inflammation Value and ECOG (PILE score): ECOG Score + (AST + ALT + Bilirubin)
- Prognostic Nutritional Index (PNI): (Albumin (g/dL) × 10) + (Lymphocyte Count (per mm3) × 0.005)
- Systemic Immune Inflammation Index (SII): (Platelet Count × Neutrophil Count)/Lymphocyte Count
- Systemic Inflammatory Response Index (SIRI): (Neutrophil Count × Monocyte Count)/Lymphocyte Count
Appendix C
| Variable | Lambda |
|---|---|
| PLR | 0.002256649 |
| MPV/lymphocyte | 0.020427439 |
| PNI | −0.003089101 |
| Globulin | −0.284104028 |
| ICI before ICI | −0.197677588 |
| Metastasis Status | 0.343231699 |
| Lesion localisation | 0.615436046 |
| Oligometastasis Status | −0.143845135 |
| Metastasis Sites | 1.071470755 |
| BRAF mutation | 0.163690698 |
References
- Ascierto, P.A.; Long, G.V.; Robert, C.; Brady, B.; Dutriaux, C.; Di Giacomo, A.M.; Mortier, L.; Hassel, J.C.; Rutkowski, P.; McNeil, C. Survival outcomes in patients with previously untreated BRAF wild-type advanced melanoma treated with nivolumab therapy: Three-year follow-up of a randomized phase 3 trial. JAMA Oncol. 2019, 5, 187–194. [Google Scholar] [CrossRef] [PubMed]
- Larkin, J.; Chiarion-Sileni, V.; Gonzalez, R.; Grob, J.-J.; Rutkowski, P.; Lao, C.D.; Cowey, C.L.; Schadendorf, D.; Wagstaff, J.; Dummer, R. Five-year survival with combined nivolumab and ipilimumab in advanced melanoma. N. Engl. J. Med. 2019, 381, 1535–1546. [Google Scholar] [CrossRef] [PubMed]
- Schachter, J.; Ribas, A.; Long, G.V.; Arance, A.; Grob, J.-J.; Mortier, L.; Daud, A.; Carlino, M.S.; McNeil, C.; Lotem, M. Pembrolizumab versus ipilimumab for advanced melanoma: Final overall survival results of a multicentre, randomised, open-label phase 3 study (KEYNOTE-006). Lancet 2017, 390, 1853–1862. [Google Scholar] [CrossRef] [PubMed]
- Carlino, M.S.; Menzies, A.M.; Atkinson, V.; Cebon, J.S.; Jameson, M.B.; Fitzharris, B.M.; McNeil, C.M.; Hill, A.G.; Ribas, A.; Atkins, M.B.; et al. Long-term Follow-up of Standard-Dose Pembrolizumab Plus Reduced-Dose Ipilimumab in Patients with Advanced Melanoma: KEYNOTE-029 Part 1B. Clin. Cancer Res. 2020, 26, 5086–5091. [Google Scholar] [CrossRef]
- Wolchok, J.D.; Chiarion-Sileni, V.; Rutkowski, P.; Cowey, C.L.; Schadendorf, D.; Wagstaff, J.; Queirolo, P.; Dummer, R.; Butler, M.O.; Hill, A.G.; et al. Final, 10-year outcomes with nivolumab plus ipilimumab in advanced melanoma. N. Engl. J. Med. 2025, 392, 11–22. [Google Scholar] [CrossRef]
- Dummer, R.; Corrie, P.; Gutzmer, R.; Meniawy, T.M.; Del Vecchio, M.; Lebbé, C.; Guida, M.; Dutriaux, C.; Dreno, B.; Meyer, N.; et al. First-Line, Fixed-Duration Nivolumab Plus Ipilimumab Followed by Nivolumab in Clinically Diverse Patient Populations With Unresectable Stage III or IV Melanoma: CheckMate 401. J. Clin. Oncol. 2023, 41, 3917–3929. [Google Scholar] [CrossRef]
- Pires da Silva, I.; Ahmed, T.; McQuade, J.L.; Nebhan, C.A.; Park, J.J.; Versluis, J.M.; Serra-Bellver, P.; Khan, Y.; Slattery, T.; Oberoi, H.K.; et al. Clinical Models to Define Response and Survival With Anti-PD-1 Antibodies Alone or Combined With Ipilimumab in Metastatic Melanoma. J. Clin. Oncol. 2022, 40, 1068–1080. [Google Scholar] [CrossRef]
- Acar, C.; Yüksel, H.Ç.; Şahin, G.; Açar, F.P.; Karaca, B. Exploring the Frequency and Risk Factors of Hyperprogressive Disease in Patients with Advanced Melanoma Treated with Immune Checkpoint Inhibitors. Curr. Oncol. 2024, 31, 6343–6355. [Google Scholar] [CrossRef]
- Zhang, C.; Zhang, C.; Wang, H. Immune-checkpoint inhibitor resistance in cancer treatment: Current progress and future directions. Cancer Lett. 2023, 562, 216182. [Google Scholar] [CrossRef]
- Amaral, T.; Seeber, O.; Mersi, E.; Sanchez, S.; Thomas, I.; Meiwes, A.; Forschner, A.; Leiter, U.; Eigentler, T.K.; Keim, U.; et al. Primary Resistance to PD-1-Based Immunotherapy—A Study in 319 Patients with Stage IV Melanoma. Cancers 2020, 12, 1207. [Google Scholar] [CrossRef]
- Kluger, H.; Barrett, J.C.; Gainor, J.F.; Hamid, O.; Hurwitz, M.; LaVallee, T.; Moss, R.A.; Zappasodi, R.; Sullivan, R.J.; Tawbi, H.; et al. Society for Immunotherapy of Cancer (SITC) consensus definitions for resistance to combinations of immune checkpoint inhibitors. J. Immunother. Cancer 2023, 11, e005921. [Google Scholar] [CrossRef] [PubMed]
- Ziogas, D.C.; Theocharopoulos, C.; Koutouratsas, T.; Haanen, J.; Gogas, H. Mechanisms of resistance to immune checkpoint inhibitors in melanoma: What we have to overcome? Cancer Treat. Rev. 2023, 113, 102499. [Google Scholar] [CrossRef] [PubMed]
- Slominski, R.M.; Sarna, T.; Płonka, P.M.; Raman, C.; Brożyna, A.A.; Slominski, A.T. Melanoma, Melanin, and Melanogenesis: The Yin and Yang Relationship. Front. Oncol. 2022, 12, 842496. [Google Scholar] [CrossRef] [PubMed]
- Slominski, R.M.; Kim, T.-K.; Janjetovic, Z.; Brożyna, A.A.; Podgorska, E.; Dixon, K.M.; Mason, R.S.; Tuckey, R.C.; Sharma, R.; Crossman, D.K.; et al. Malignant Melanoma: An Overview, New Perspectives, and Vitamin D Signaling. Cancers 2024, 16, 2262. [Google Scholar] [CrossRef]
- Slominski, R.M.; Raman, C.; Chen, J.Y.; Slominski, A.T. How cancer hijacks the body’s homeostasis through the neuroendocrine system. Trends Neurosci. 2023, 46, 263–275. [Google Scholar] [CrossRef]
- Byeon, S.; Cho, H.J.; Jang, K.T.; Kwon, M.; Lee, J.; Lee, J.; Kim, S.T. Molecular profiling of Asian patients with advanced melanoma receiving check-point inhibitor treatment. ESMO Open 2021, 6, 100002. [Google Scholar] [CrossRef]
- van Not, O.J.; Blokx, W.A.M.; van den Eertwegh, A.J.M.; de Meza, M.M.; Haanen, J.B.; Blank, C.U.; Aarts, M.J.B.; van den Berkmortel, F.W.P.J.; de Groot, J.W.B.; Hospers, G.A.P.; et al. BRAF and NRAS Mutation Status and Response to Checkpoint Inhibition in Advanced Melanoma. JCO Precis. Oncol. 2022, 6, e2200018. [Google Scholar] [CrossRef]
- Borch, T.H.; Harbst, K.; Rana, A.H.; Andersen, R.; Martinenaite, E.; Kongsted, P.; Pedersen, M.; Nielsen, M.; Kjeldsen, J.W.; Kverneland, A.H. Clinical efficacy of T-cell therapy after short-term BRAF-inhibitor priming in patients with checkpoint inhibitor-resistant metastatic melanoma. J. Immunother. Cancer 2021, 9, e002703. [Google Scholar] [CrossRef]
- Zhang, W.; Kong, Y.; Li, Y.; Shi, F.; Lyu, J.; Sheng, C.; Wang, S.; Wang, Q. Novel molecular determinants of response or resistance to immune checkpoint inhibitor therapies in melanoma. Front. Immunol. 2022, 12, 798474. [Google Scholar] [CrossRef]
- Hossain, S.M.; Gimenez, G.; Stockwell, P.A.; Tsai, P.; Print, C.G.; Rys, J.; Cybulska-Stopa, B.; Ratajska, M.; Harazin-Lechowska, A.; Almomani, S. Innate immune checkpoint inhibitor resistance is associated with melanoma sub-types exhibiting invasive and de-differentiated gene expression signatures. Front. Immunol. 2022, 13, 955063. [Google Scholar] [CrossRef]
- Greten, F.R.; Grivennikov, S.I. Inflammation and cancer: Triggers, mechanisms, and consequences. Immunity 2019, 51, 27–41. [Google Scholar] [CrossRef]
- Kim, R.; An, M.; Lee, H.; Mehta, A.; Heo, Y.J.; Kim, K.-M.; Lee, S.-Y.; Moon, J.; Kim, S.T.; Min, B.-H. Early tumor–immune microenvironmental remodeling and response to first-line fluoropyrimidine and platinum chemotherapy in advanced gastric cancer. Cancer Discov. 2022, 12, 984–1001. [Google Scholar] [CrossRef] [PubMed]
- Ou, Y.; Liang, S.; Gao, Q.; Shang, Y.; Liang, J.; Zhang, W.; Liu, S. Prognostic value of inflammatory markers NLR, PLR, LMR, dNLR, ANC in melanoma patients treated with immune checkpoint inhibitors: A meta-analysis and systematic review. Front. Immunol. 2024, 15, 1482746. [Google Scholar] [CrossRef] [PubMed]
- Acar, C.; Yüksel, H.Ç.; Şahin, G.; Açar, F.P.; Gunenc, D.; Karaca, B. Prognostic utility of the CALLY index in metastatic melanoma: Building a nomogram for Patients on Anti-PD-1 therapy. Clin. Transl. Oncol. 2025. [Google Scholar] [CrossRef] [PubMed]
- Paolino, G.; Huber, V.; Camerini, S.; Casella, M.; Macone, A.; Bertuccini, L.; Iosi, F.; Moliterni, E.; Cecchetti, S.; Ruspantini, I.; et al. The Fatty Acid and Protein Profiles of Circulating CD81-Positive Small Extracellular Vesicles Are Associated with Disease Stage in Melanoma Patients. Cancers 2021, 13, 4157. [Google Scholar] [CrossRef]
- Lauss, M.; Phung, B.; Borch, T.H.; Harbst, K.; Kaminska, K.; Ebbesson, A.; Hedenfalk, I.; Yuan, J.; Nielsen, K.; Ingvar, C.; et al. Molecular patterns of resistance to immune checkpoint blockade in melanoma. Nat. Commun. 2024, 15, 3075. [Google Scholar] [CrossRef]
- Tao, W.; Sun, Q.; Xu, B.; Wang, R. Towards the Prediction of Responses to Cancer Immunotherapy: A Multi-Omics Review. Life 2025, 15, 283. [Google Scholar] [CrossRef]
- Zhao, S.; Li, Z.; Liu, K.; Wang, G.; Wang, Q.; Yu, H.; Chen, W.; Dai, H.; Li, Y.; Xie, J.; et al. Combining multi-omics analysis with machine learning to uncover novel molecular subtypes, prognostic markers, and insights into immunotherapy for melanoma. BMC Cancer 2025, 25, 630. [Google Scholar] [CrossRef]
- Gschwind, A.; Ossowski, S. AI Model for Predicting Anti-PD1 Response in Melanoma Using Multi-Omics Biomarkers. Cancers 2025, 17, 714. [Google Scholar] [CrossRef]
- van Duin, I.A.J.; Verheijden, R.J.; van Diest, P.J.; Blokx, W.A.M.; El-Sharouni, M.-A.; Verhoeff, J.J.C.; Leiner, T.; van den Eertwegh, A.J.M.; de Groot, J.W.B.; van Not, O.J.; et al. A prediction model for response to immune checkpoint inhibition in advanced melanoma. Int. J. Cancer 2024, 154, 1760–1771. [Google Scholar] [CrossRef]
- Nosrati, A.; Tsai, K.K.; Goldinger, S.M.; Tumeh, P.; Grimes, B.; Loo, K.; Algazi, A.P.; Nguyen-Kim, T.D.L.; Levesque, M.; Dummer, R. Evaluation of clinicopathological factors in PD-1 response: Derivation and validation of a prediction scale for response to PD-1 monotherapy. Br. J. Cancer 2017, 116, 1141–1147. [Google Scholar] [CrossRef]
- Birnboim-Perach, R.; Benhar, I. Using Combination therapy to overcome diverse challenges of Immune Checkpoint Inhibitors treatment. Int. J. Biol. Sci. 2024, 20, 3911. [Google Scholar] [CrossRef]
- Pavlick, A.C.; Ariyan, C.E.; Buchbinder, E.I.; Davar, D.; Gibney, G.T.; Hamid, O.; Hieken, T.J.; Izar, B.; Johnson, D.B.; Kulkarni, R.P.; et al. Society for Immunotherapy of Cancer (SITC) clinical practice guideline on immunotherapy for the treatment of melanoma, version 3.0. J. Immunother. Cancer 2023, 11, e006947. [Google Scholar] [CrossRef]
- Atkins, M.B.; Lee, S.J.; Chmielowski, B.; Ribas, A.; Tarhini, A.A.; Truong, T.-G.; Davar, D.; O’Rourke, M.A.; Curti, B.D.; Brell, J.M. DREAMseq (doublet, randomized evaluation in advanced melanoma sequencing): A phase III trial—ECOG-ACRIN EA6134. J. Clin. Oncol. 2021, 39, 356154. [Google Scholar] [CrossRef]
- Ilieva, K.M.; Correa, I.; Josephs, D.H.; Karagiannis, P.; Egbuniwe, I.U.; Cafferkey, M.J.; Spicer, J.F.; Harries, M.; Nestle, F.O.; Lacy, K.E. Effects of BRAF mutations and BRAF inhibition on immune responses to melanoma. Mol. Cancer Ther. 2014, 13, 2769–2783. [Google Scholar] [CrossRef] [PubMed]
- Haas, L.; Elewaut, A.; Gerard, C.L.; Umkehrer, C.; Leiendecker, L.; Pedersen, M.; Krecioch, I.; Hoffmann, D.; Novatchkova, M.; Kuttke, M. Acquired resistance to anti-MAPK targeted therapy confers an immune-evasive tumor microenvironment and cross-resistance to immunotherapy in melanoma. Nat. Cancer 2021, 2, 693–708. [Google Scholar] [CrossRef] [PubMed]
- Hribernik, N.; Boc, M.; Ocvirk, J.; Knez-Arbeiter, J.; Mesti, T.; Ignjatovic, M.; Rebersek, M. Retrospective analysis of treatment-naive Slovenian patients with metastatic melanoma treated with pembrolizumab–real-world experience. Radiol. Oncol. 2020, 54, 119–127. [Google Scholar] [CrossRef]
- Seremet, T.; Jansen, Y.; Planken, S.; Njimi, H.; Delaunoy, M.; El Housni, H.; Awada, G.; Schwarze, J.K.; Keyaerts, M.; Everaert, H. Undetectable circulating tumor DNA (ctDNA) levels correlate with favorable outcome in metastatic melanoma patients treated with anti-PD1 therapy. J. Transl. Med. 2019, 17, 1–13. [Google Scholar] [CrossRef]
- Gangadhar, T.C.; Hwu, W.-J.; Postow, M.A.; Hamid, O.; Daud, A.; Dronca, R.; Joseph, R.; O’Day, S.J.; Hodi, F.; Pavlick, A.C. Efficacy and safety of pembrolizumab in patients enrolled in KEYNOTE-030 in the United States: An expanded access program. J. Immunother. 2017, 40, 334–340. [Google Scholar] [CrossRef]
- Kreft, S.; Gesierich, A.; Eigentler, T.; Franklin, C.; Valpione, S.; Ugurel, S.; Utikal, J.; Haferkamp, S.; Blank, C.; Larkin, J. Efficacy of PD-1–based immunotherapy after radiologic progression on targeted therapy in stage IV melanoma. Eur. J. Cancer 2019, 116, 207–215. [Google Scholar] [CrossRef]
- Guven, D.C.; Sahin, T.K.; Erul, E.; Rizzo, A.; Ricci, A.D.; Aksoy, S.; Yalcin, S. The association between albumin levels and survival in patients treated with immune checkpoint inhibitors: A systematic review and meta-analysis. Front. Mol. Biosci. 2022, 9, 1039121. [Google Scholar] [CrossRef]
- Nakanishi, Y.; Masuda, T.; Yamaguchi, K.; Sakamoto, S.; Horimasu, Y.; Mimae, T.; Nakashima, T.; Miyamoto, S.; Tsutani, Y.; Iwamoto, H. Albumin–globulin ratio is a predictive biomarker of antitumor effect of anti-PD-1 antibody in patients with non-small cell lung cancer. Int. J. Clin. Oncol. 2020, 25, 74–81. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Shang, K.; Wu, S.; Wang, J.; Cao, B. The prognostic value of albumin-globulin ratio and eosinophil-neutrophil ratio in patients with advanced tumors undergoing treatment with PD-1/PD-L1 inhibitors. Nutr. Cancer 2022, 74, 2815–2828. [Google Scholar] [CrossRef] [PubMed]
- Shebrain, A.; Idris, O.A.; Jawad, A.; Zhang, T.; Xing, Y. Advancements and Challenges in Personalized Therapy for BRAF-Mutant Melanoma: A Comprehensive Review. J. Clin. Med. 2024, 13, 5409. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Chen, Y.; Xie, Y.; Ma, X.; Liu, Y. Deciphering and overcoming Anti-PD-1 resistance in Melanoma: A comprehensive review of Mechanisms, biomarker Developments, and therapeutic strategies. Int. Immunopharmacol. 2024, 132, 111989. [Google Scholar] [CrossRef]
- Liu, C.; Wang, W.; Meng, X.; Sun, B.; Cong, Y.; Liu, J.; Wang, Q.; Liu, G.; Wu, S. Albumin/globulin ratio is negatively correlated with PD-1 and CD25 mRNA levels in breast cancer patients. OncoTargets Ther. 2018, 11, 2131–2139. [Google Scholar] [CrossRef]
- Diem, S.; Fässler, M.; Bomze, D.; Ali, O.H.; Berner, F.; Niederer, R.; Hillmann, D.; Mangana, J.; Levesque, M.P.; Dummer, R.; et al. Immunoglobulin G and Subclasses as Potential Biomarkers in Metastatic Melanoma Patients Starting Checkpoint Inhibitor Treatment. J. Immunother. 2019, 42, 89–93. [Google Scholar] [CrossRef]
- Pires da Silva, I.; Lo, S.; Quek, C.; Gonzalez, M.; Carlino, M.S.; Long, G.V.; Menzies, A.M. Site-specific response patterns, pseudoprogression, and acquired resistance in patients with melanoma treated with ipilimumab combined with anti–PD-1 therapy. Cancer 2020, 126, 86–97. [Google Scholar] [CrossRef]
- Lee, J.H.J.; Lyle, M.; Menzies, A.M.; Chan, M.M.K.; Lo, S.; Clements, A.; Carlino, M.S.; Kefford, R.F.; Long, G.V. Metastasis-specific patterns of response and progression with anti-PD-1 treatment in metastatic melanoma. Pigment. Cell Melanoma Res. 2018, 31, 404–410. [Google Scholar] [CrossRef]
- Seeber, T.O. Primary Resistance to Immune Checkpoint Inhibitors in Patients with Metastatic Melanoma: Prognosis, Subsequent Therapies and Survival. Ph.D. Thesis, Universität Tübingen, Tübingen, Germany, 2021. [Google Scholar]
- Champiat, S.; Matte, P.; Baldini, C.; Vasseur, D.; Ouali, K.; Gazzah, A.; Bahleda, R.; Bayle, A.; Postel-Vinay, S.; Smolenschi, C.; et al. Abstract C068: Factors associated with primary resistance to immune checkpoint blockade in early phase clinical trials. Mol. Cancer Ther. 2023, 22, C068. [Google Scholar] [CrossRef]
- Maugeais, M.; Peron, J.; Dalle, S.; Duruisseaux, M.; Corbaux, P.; Reverdy, T.; Lopez, J.; Souquet, P.J.; Maillet, D. 1038P Impact of metastatic location on immune-checkpoint inhibitors efficacy in patients with different solid tumours. Ann. Oncol. 2020, 31, S714. [Google Scholar] [CrossRef]
- Horst, A.K.; Neumann, K.; Diehl, L.; Tiegs, G. Modulation of liver tolerance by conventional and nonconventional antigen-presenting cells and regulatory immune cells. Cell Mol. Immunol. 2016, 13, 277–292. [Google Scholar] [CrossRef]
- Shui, I.M.; Liu, X.Q.; Zhao, Q.; Kim, S.T.; Sun, Y.; Yearley, J.H.; Choudhury, T.; Webber, A.L.; Krepler, C.; Cristescu, R.; et al. Baseline and post-treatment biomarkers of resistance to anti-PD-1 therapy in acral and mucosal melanoma: An observational study. J. Immunother. Cancer 2022, 10, e004879. [Google Scholar] [CrossRef] [PubMed]
- Gui, J.; Guo, Z.; Wu, D. Clinical features, molecular pathology, and immune microenvironmental characteristics of acral melanoma. J. Transl. Med. 2022, 20, 367. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Zhao, J.J.; Soon, Y.Y.; Kee, A.; Tay, S.H.; Aminkeng, F.; Ang, Y.; Wong, A.S.C.; Bharwani, L.D.; Goh, B.C.; et al. Factors Predictive of Primary Resistance to Immune Checkpoint Inhibitors in Patients with Advanced Non-Small Cell Lung Cancer. Cancers 2023, 15, 2733. [Google Scholar] [CrossRef] [PubMed]
- Splendiani, E.; Besharat, Z.M.; Covre, A.; Maio, M.; Di Giacomo, A.M.; Ferretti, E. Immunotherapy in melanoma: Can we predict response to treatment with circulating biomarkers? Pharmacol. Ther. 2024, 256, 108613. [Google Scholar] [CrossRef]
- Li, J.; Liu, J.; Yang, S.; Xia, Y.; Meng, Q.; Sun, B.; Liu, Y.; Zhao, B.; Jin, J.; Xu, H.; et al. PD-L1 positive platelets mediate resistance to immune checkpoint inhibitors in patients with colorectal cancer. Cell Commun. Signal 2025, 23, 29. [Google Scholar] [CrossRef]
- Young, R.; Panchal, H.; Ma, W.; Chen, S.; Steele, A.P.; Iannucci, A.A.; Li, T. Low lymphocyte counts and association with clinical outcomes in hospitalized patients with cancer receiving immune checkpoint inhibitor therapy. J. Clin. Oncol. 2022, 40, e14600. [Google Scholar] [CrossRef]
- Ménétrier-Caux, C.; Ray-Coquard, I.; Blay, J.-Y.; Caux, C. Lymphopenia in cancer patients and its effects on response to immunotherapy: An opportunity for combination with cytokines? J. Immunother. Cancer 2019, 7, 85. [Google Scholar] [CrossRef]
- Shui, I.M.; Scherrer, E.; Frederickson, A.; Li, J.W.; Mynzhassarova, A.; Druyts, E.; Tawbi, H. Resistance to anti-PD1 therapies in patients with advanced melanoma: Systematic literature review and application of the Society for Immunotherapy of Cancer Immunotherapy Resistance Taskforce anti-PD1 resistance definitions. Melanoma Res. 2022, 32, 393–404. [Google Scholar] [CrossRef]
| Resistance Status | ||||
|---|---|---|---|---|
| Patient Characteristics | Non-Resistant | Resistant | Total | |
| Total N (%) | 61 (55.5) | 49 (44.5) | 110 | |
| Age | Median (IQR) | 61.0 (48.0–68.0) | 59.0 (46.0–67.0) | 60.0 (47.2–68.0) |
| Sex | Male | 34 (55.7) | 24 (49.0) | 58 (52.7) |
| Female | 27 (44.3) | 25 (51.0) | 52 (47.3) | |
| ECOG PS | Poor (0–1) | 5 (8.2) | 8 (16.3) | 13 (11.8) |
| Good (2–3) | 56 (91.8) | 41 (83.7) | 97 (88.2) | |
| BRAF/MEKi treatment before PD1 | No | 51 (83.6) | 39 (79.6) | 90 (81.8) |
| Yes | 10 (16.4) | 10 (20.4) | 20 (18.2) | |
| Chemotherapy Before PD1 | No | 48 (78.7) | 36 (73.5) | 84 (76.4) |
| Yes | 13 (21.3) | 13 (26.5) | 26 (23.6) | |
| ICI before ICI | No | 40 (65.6) | 35 (71.4) | 75 (68.2) |
| İpilimumab | 18 (29.5) | 14 (28.6) | 32 (29.1) | |
| PD1 | 3 (4.9) | 0 (0.0) | 3 (2.7) | |
| Metastasis Status | De Novo | 28 (45.9) | 15 (30.6) | 43 (39.1) |
| Recurrent | 33 (54.1) | 34 (69.4) | 67 (60.9) | |
| Anti-PD1 Treatment Line | First | 35 (57.4) | 23 (46.9) | 58 (52.7) |
| Second and Beyond | 26 (42.6) | 26 (53.1) | 52 (47.3) | |
| Lesion Localization | Non-Acral Cutaneous | 44 (72.1) | 24 (49.0) | 68 (61.8) |
| Acral | 9 (14.7) | 15 (30.5) | 24 (21.8) | |
| Mucosal | 8 (13.1) | 10 (20.4) | 18 (16.4) | |
| Oligometastasis Status | No | 26 (42.6) | 36 (73.5) | 62 (56.4) |
| Yes | 35 (57.4) | 13 (26.5) | 48 (43.6) | |
| Stage | M1a | 26 (42.6) | 8 (16.3) | 34 (30.9) |
| M1b | 14 (23.0) | 10 (20.4) | 24 (21.8) | |
| M1c | 17 (27.9) | 24 (49.0) | 41 (37.3) | |
| M1d | 4 (6.6) | 7 (14.3) | 11 (10.0) | |
| Metastasis Site | <3 | 49 (80.3) | 20 (40.8) | 69 (62.7) |
| ≥3 | 12 (19.7) | 29 (59.2) | 41 (37.3) | |
| BRAF mutation | Wild | 47 (77.0) | 34 (69.4) | 81 (73.6) |
| Mutant | 14 (23.0) | 15 (30.6) | 29 (26.4) | |
| SIPS | Good | 50 (86.2) | 33 (80.5) | 83 (83.8) |
| Poor | 8 (13.8) | 8 (19.5) | 16 (16.2) | |
| Royal Marsden Hospital Score | Good | 29 (50.9) | 10 (24.4) | 39 (39.8) |
| Poor | 28 (49.1) | 31 (75.6) | 59 (60.2) | |
| MD Anderson ICI Score | Good | 17 (28.8) | 5 (11.9) | 22 (21.8) |
| Intermediate | 38 (64.4) | 26 (61.9) | 64 (63.4) | |
| Poor | 4 (6.8) | 11 (26.2) | 15 (14.9) | |
| Lymphocyte | Median (IQR) | 1820.0 (1315.0–2315.0) | 1430.0 (1080.0–1780.0) | 1635.0 (1255.0–2125.0) |
| Monocyte | Median (IQR) | 560.0 (440.0–785.0) | 560.0 (439.5–710.0) | 560.0 (440.0–737.5) |
| Eosinophil | Median (IQR) | 180.0 (102.5–290.0) | 100.0 (50.0–205.0) | 130.0 (60.0–250.0) |
| Albumin | Median (IQR) | 4.3 (4.0–4.6) | 4.1 (3.9–4.4) | 4.2 (3.9–4.5) |
| Globulin | Median (IQR) | 2.8 (2.5–3.2) | 2.8 (2.4–3.0) | 2.8 (2.5–3.1) |
| CRP | Median (IQR) | 0.2 (0.1–1.1) | 0.5 (0.2–1.4) | 0.3 (0.1–1.4) |
| LDH | Median (IQR) | 201.0 (175.0–274.0) | 227.0 (177.0 t–290.0) | 208.5 (175.0–278.2) |
| MPV | Median (IQR) | 10.1 (9.6–10.7) | 9.8 (9.2–10.3) | 10.0 (9.5–10.7) |
| NLR | Median (IQR) | 2.4 (1.7–3.3) | 3.0 (2.5–3.7) | 2.7 (2.0-3.6) |
| LMR | Median (IQR) | 3.0 (2.5–4.2) | 2.6 (2.0–3.3) | 2.9 (2.2- 3.8) |
| PLR | Median (IQR) | 153.3 (112.2–196.9) | 196.4 (127.0–294.2) | 162.9 (113.8–236.8) |
| MPV/Lymphocyte | Median (IQR) | 5.4 (4.6–7.5) | 6.5 (5.7–8.8) | 6.0 (4.7–8.3) |
| HALP Score | Median (IQR) | 39.1 (26.8–50.3) | 26.4 (16.0–49.2) | 34.3 (19.9–50.2) |
| PIV Score | Median (IQR) | 383.3 (206.1–651.0) | 474.0 (276.6–936.6) | 438.1 (225.7–822.8) |
| PNI Score | Median (IQR) | 51.6 (46.5–54.7) | 48.7 (44.5–51.6) | 50.2 (45.6–54.4) |
| SII Score | Median (IQR) | 676.0 (421.2–982.4) | 889.7 (526.4–1365.1) | 724.2 (465.3–1123.7) |
| Primary Resistance | No | Yes | OR (Univariable) | OR (Multivariable) | |
|---|---|---|---|---|---|
| MPV/lymphocyte | Mean (SD) | 6.4 (2.8) | 7.8 (3.8) | 1.16 (1.01–1.35, p = 0.043) | |
| PNI | Mean (SD) | 51.9 (5.8) | 48.7 (5.8) | 0.91 (0.84–0.98, p = 0.015) | |
| ICI before ICI | No | 40 (53.3) | 35 (46.7) | - | |
| Yes | 21 (60) | 14 (40) | 0.59 (0.22–1.47, p = 0.264) | ||
| Metastasis Status | Denovo | 28 (65.1) | 15 (34.9) | - | |
| Recurrent | 33 (49.3) | 34 (50.7) | 2.28 (0.97–5.63, p = 0.065) | ||
| Oligometastasis Status | No | 49 (71.0) | 20 (29.0) | - | |
| Yes | 12 (29.3) | 29 (70.7) | 0.30 (0.12–0.72, p = 0.008) | ||
| Lesion Localisation | Non Acral | 44 (64.7) | 24 (35.3) | - | |
| Acral + Mucosal | 17 (40.5) | 25 (59.5) | 2.32 (0.99–5.53, p = 0.055) | 3.82 (1.16–12.6, p = 0.027) | |
| BRAF Mutation | Wild | 28 (65.1) | 15 (34.9) | - | - |
| Mutant | 33 (49.3) | 34 (50.7) | 1.78 (0.69–4.65, p = 0.234) | 5.22 (1.31–20.82, p = 0.019) | |
| Metastasis Grup | <3 | 47 (58.0) | 34 (42.0) | - | |
| ≥3 | 14 (48.3) | 15 (51.7) | 5.75 (2.35–14.90, p < 0.001) | 11.4 (3.43–37.55, p < 0.001) | |
| PLR | Mean (SD) | 164.2 (72.5) | 223.7 (118.3) | 1.01 (1.00–1.01, p = 0.006) | 1.007 (1.01–1.01, p = 0.019) |
| Globulin | Mean (SD) | 2.9 (0.7) | 2.8 (0.5) | 0.75 (0.36–1.49, p = 0.429) | 0.35 (0.14–0.88, p = 0.027) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Published by MDPI on behalf of the Lithuanian University of Health Sciences. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yuksel, H.C.; Acar, C.; Sahin, G.; Celebi, G.; Tunbekici, S.; Karaca, B.S. LASSO-Driven Selection of Biochemical and Clinical Markers for Primary Resistance to PD-1 Inhibitors in Metastatic Melanoma. Medicina 2025, 61, 1559. https://doi.org/10.3390/medicina61091559
Yuksel HC, Acar C, Sahin G, Celebi G, Tunbekici S, Karaca BS. LASSO-Driven Selection of Biochemical and Clinical Markers for Primary Resistance to PD-1 Inhibitors in Metastatic Melanoma. Medicina. 2025; 61(9):1559. https://doi.org/10.3390/medicina61091559
Chicago/Turabian StyleYuksel, Haydar C., Caner Acar, Gokhan Sahin, Gulcin Celebi, Salih Tunbekici, and Burcak S. Karaca. 2025. "LASSO-Driven Selection of Biochemical and Clinical Markers for Primary Resistance to PD-1 Inhibitors in Metastatic Melanoma" Medicina 61, no. 9: 1559. https://doi.org/10.3390/medicina61091559
APA StyleYuksel, H. C., Acar, C., Sahin, G., Celebi, G., Tunbekici, S., & Karaca, B. S. (2025). LASSO-Driven Selection of Biochemical and Clinical Markers for Primary Resistance to PD-1 Inhibitors in Metastatic Melanoma. Medicina, 61(9), 1559. https://doi.org/10.3390/medicina61091559







