The Relationship between MALAT1 Polymorphism rs3200401 C > T and the Risk of Overall Cancer: A Meta-Analysis
Abstract
:1. Introduction
2. Materials and Methods
2.1. Search Strategy
2.2. Inclusion and Exclusion Criteria
2.3. Data Extraction and Quality Assessment
2.4. Statistical Analysis
3. Results
3.1. Characteristics of Studies
3.2. Quantitative Analysis
3.3. Sensitivity Analysis and Publication Bias
3.4. Meta-Regression Analysis
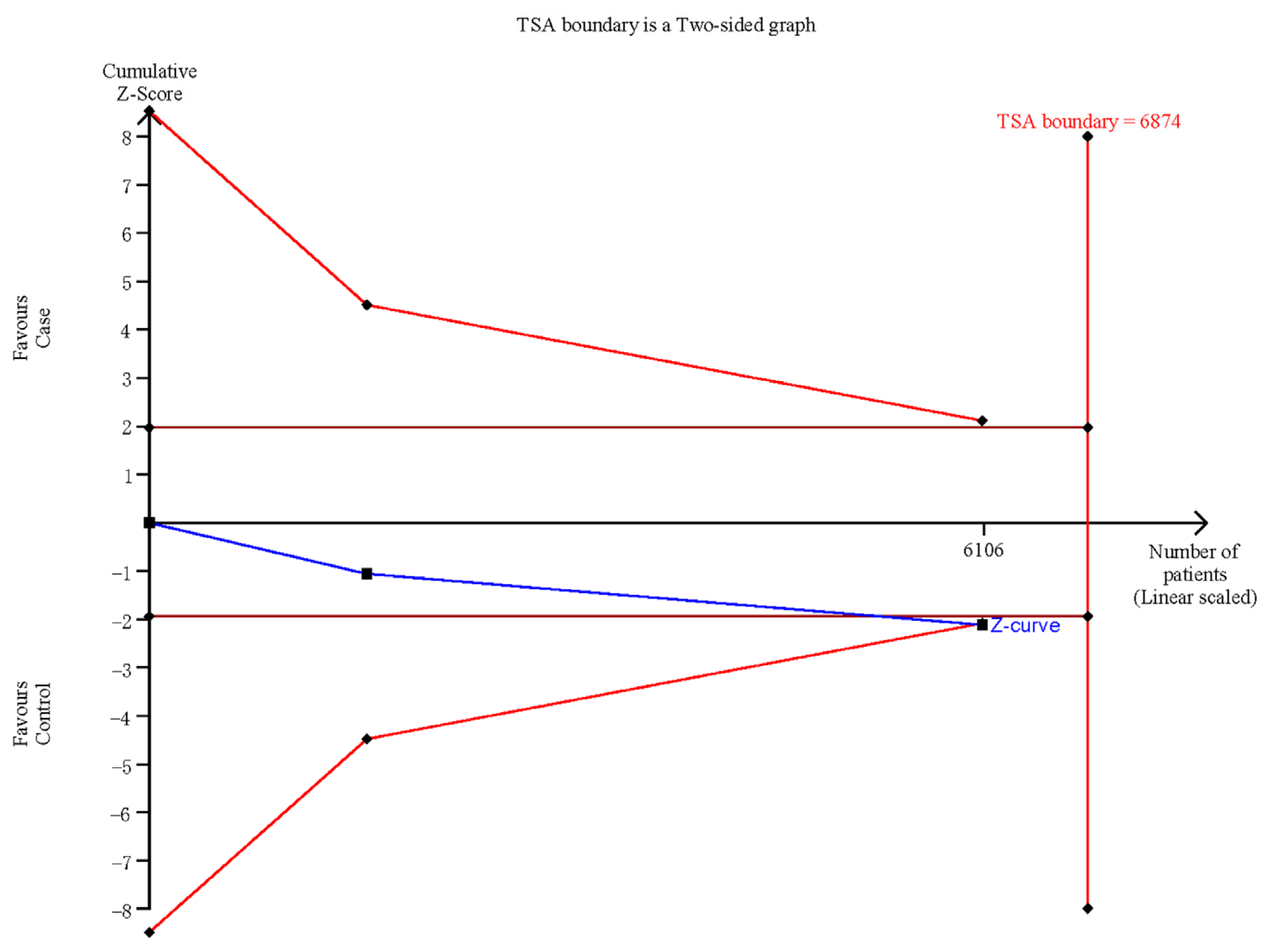
3.5. Trial Sequential Analysis
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- Keum, N.; Giovannucci, E. Global burden of colorectal cancer: Emerging trends, risk factors and prevention strategies. Nat. Rev. Gastroenterol. Hepatol. 2019, 16, 713–732. [Google Scholar] [CrossRef] [PubMed]
- Lin, L.; Yan, L.; Liu, Y.; Qu, C.; Ni, J.; Li, H. The Burden and Trends of Primary Liver Cancer Caused by Specific Etiologies from 1990 to 2017 at the Global, Regional, National, Age, and Sex Level Results from the Global Burden of Disease Study 2017. Liver Cancer 2020, 9, 563–582. [Google Scholar] [CrossRef] [PubMed]
- Lao, V.V.; Grady, W.M. Epigenetics and colorectal cancer. Nat. Rev. Gastroenterol. Hepatol. 2011, 8, 686–700. [Google Scholar] [CrossRef] [PubMed]
- Villanueva, L.; Álvarez-Errico, D.; Esteller, M. The Contribution of Epigenetics to Cancer Immunotherapy. Trends Immunol. 2020, 41, 676–691. [Google Scholar] [CrossRef]
- Mei, H.; Liu, Y.; Zhou, Q.; Hu, K.; Liu, Y. Long noncoding RNA MALAT1 acts as a potential biomarker in cancer diagnosis and detection: A meta-analysis. Biomark. Med. 2019, 13, 45–54. [Google Scholar] [CrossRef]
- Peng, W.X.; Koirala, P.; Mo, Y.Y. LncRNA-mediated regulation of cell signaling in cancer. Oncogene 2017, 36, 5661–5667. [Google Scholar] [CrossRef]
- Fu, S.; Wang, Y.; Li, H.; Chen, L.; Liu, Q. Regulatory Networks of LncRNA MALAT-1 in Cancer. Cancer Manag. Res. 2020, 12, 10181–10198. [Google Scholar] [CrossRef]
- Guru, S.C.; Agarwal, S.K.; Manickam, P.; Olufemi, S.E.; Crabtree, J.S.; Weisemann, J.M.; Kester, M.B.; Kim, Y.S.; Wang, Y.; Emmert-Buck, M.R.; et al. A transcript map for the 2.8-Mb region containing the multiple endocrine neoplasia type 1 locus. Genome Res. 1997, 7, 725–735. [Google Scholar] [CrossRef] [Green Version]
- Goyal, B.; Yadav, S.R.M.; Awasthee, N.; Gupta, S.; Kunnumakkara, A.B.; Gupta, S.C. Diagnostic, prognostic, and therapeutic significance of long non-coding RNA MALAT1 in cancer. Biochim. Biophys. Acta Rev. Cancer 2021, 1875, 188502. [Google Scholar] [CrossRef]
- Jen, J.; Tang, Y.A.; Lu, Y.H.; Lin, C.C.; Lai, W.W.; Wang, Y.C. Oct4 transcriptionally regulates the expression of long non-coding RNAs NEAT1 and MALAT1 to promote lung cancer progression. Mol. Cancer 2017, 16, 104. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, J.; Li, Q.; Xue, B.; He, R. MALAT1 inhibits the Wnt/β-catenin signaling pathway in colon cancer cells and affects cell proliferation and apoptosis. Bosn. J. Basic Med. Sci. 2020, 20, 357–364. [Google Scholar] [CrossRef] [PubMed]
- Fu, Q.; Tan, X.; Tang, H.; Liu, J. CCL21 activation of the MALAT1/SRSF1/mTOR axis underpins the development of gastric carcinoma. J. Transl. Med. 2021, 19, 210. [Google Scholar] [CrossRef]
- Malakar, P.; Shilo, A.; Mogilevsky, A.; Stein, I.; Pikarsky, E.; Nevo, Y.; Benyamini, H.; Elgavish, S.; Zong, X.; Prasanth, K.V.; et al. Long Noncoding RNA MALAT1 Promotes Hepatocellular Carcinoma Development by SRSF1 Upregulation and mTOR Activation. Cancer Res. 2017, 77, 1155–1167. [Google Scholar] [CrossRef] [Green Version]
- Wang, Y.; Zhang, Q. Long Noncoding RNA MALAT1 Knockdown Inhibits Proliferation, Migration, and Invasion and Promotes Apoptosis in Non-Small-Cell Lung Cancer Cells Through Regulating miR-515-3p/TRIM65 Axis. Cancer Biother. Radiopharm. 2020. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Zhang, X.; Wang, H.; Wang, L.; Liu, T.; Du, L.; Yang, Y.; Wang, C. MALAT1 Is Associated with Poor Response to Oxaliplatin-Based Chemotherapy in Colorectal Cancer Patients and Promotes Chemoresistance through EZH2. Mol. Cancer Ther. 2017, 16, 739–751. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lv, Z.; Xu, Q.; Yuan, Y. A systematic review and meta-analysis of the association between long non-coding RNA polymorphisms and cancer risk. Mutat. Res. Rev. Mutat. Res. 2017, 771, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Peng, R.; Luo, C.; Guo, Q.; Cao, J.; Yang, Q.; Dong, K.; Wang, S.; Wang, K.; Song, C. Association analyses of genetic variants in long non-coding RNA MALAT1 with breast cancer susceptibility and mRNA expression of MALAT1 in Chinese Han population. Gene 2018, 642, 241–248. [Google Scholar] [CrossRef]
- Ding, Y.F.; Wen, Y.C.; Chuang, C.Y.; Lin, C.W.; Yang, Y.C.; Liu, Y.F.; Chang, W.M.; Chang, L.C.; Yang, S.F.; Chien, M.H. Combined Impacts of Genetic Variants of Long Non-Coding RNA MALAT1 and the Environmental Carcinogen on the Susceptibility to and Progression of Oral Squamous Cell Carcinoma. Front. Oncol. 2021, 11, 684941. [Google Scholar] [CrossRef]
- Ji, X.; Zhang, J.; Liu, L.; Lin, Z.; Pi, L.; Lin, Z.; Tian, N.; Lin, X.; Chen, S.; Yu, X.; et al. Association of tagSNPs at lncRNA MALAT-1 with HCC Susceptibility in a Southern Chinese Population. Sci. Rep. 2019, 9, 10895. [Google Scholar] [CrossRef]
- Yuan, L.T.; Chang, J.H.; Lee, H.L.; Yang, Y.C.; Su, S.C.; Lin, C.L.; Yang, S.F.; Chien, M.H. Genetic Variants of lncRNA MALAT1 Exert Diverse Impacts on the Risk and Clinicopathologic Characteristics of Patients with Hepatocellular Carcinoma. J. Clin. Med. 2019, 8, 1406. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Qu, Y.; Shao, N.; Yang, W.; Wang, J.; Cheng, Y. Association of polymorphisms in MALAT1 with the risk of esophageal squamous cell carcinoma in a Chinese population. Oncotargets Ther. 2019, 12, 2495–2503. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stang, A. Critical evaluation of the Newcastle-Ottawa scale for the assessment of the quality of nonrandomized studies in meta-analyses. Eur. J. Epidemiol. 2010, 25, 603–605. [Google Scholar] [CrossRef] [Green Version]
- Wei, Y.; Zhao, Z.; Wang, Z.; Zhang, K.; Tang, Z.; Tao, C. Relationships between IL-1β, TNF-α genetic polymorphisms and HBV infection: A meta-analytical study. Gene 2021, 791, 145617. [Google Scholar] [CrossRef] [PubMed]
- Zhao, K.; Jin, S.; Wei, B.; Cao, S.; Xiong, Z. Association study of genetic variation of lncRNA MALAT1 with carcinogenesis of colorectal cancer. Cancer Manag. Res. 2018, 10, 6257–6261. [Google Scholar] [CrossRef] [PubMed]
- Orlandi, E.; Zanot, C.; Poli, A.; Nicolis, M.; Rodolfo, M.; Turco, A.; Sangalli, A.; Gomez-Lira, M. Lack of association of metastasis-associated lung adenocarcinoma transcript 1 variants with melanoma skin cancer risk. Melanoma Res. 2019, 29, 660–663. [Google Scholar] [CrossRef] [PubMed]
- Wen, J.; Chen, L.; Tian, H.; Li, J.; Zhang, M.; Cao, Q.; Zhang, W.; Chen, S.; Shi, L. Effect of MALAT1 Polymorphisms on Papillary Thyroid Cancer in a Chinese Population. J. Cancer 2019, 10, 5714–5721. [Google Scholar] [CrossRef]
- Wu, S.; Sun, H.; Wang, Y.; Yang, X.; Meng, Q.; Yang, H.; Zhu, H.; Tang, W.; Li, X.; Aschner, M.; et al. MALAT1 rs664589 Polymorphism Inhibits Binding to miR-194-5p, Contributing to Colorectal Cancer Risk, Growth, and Metastasis. Cancer Res. 2019, 79, 5432–5441. [Google Scholar] [CrossRef] [Green Version]
- Petkevicius, V.; Streleckiene, G.; Balciute, K.; Link, A.; Leja, M.; Malfertheiner, P.; Skieceviciene, J.; Kupcinskas, J. Association of Long Non-Coding RNA Polymorphisms with Gastric Cancer and Atrophic Gastritis. Genes 2020, 11, 1505. [Google Scholar] [CrossRef]
- Hong, J.H.; Jin, E.-H.; Chang, I.A.; Kang, H.; Lee, S.-I.; Sung, J.K. Association of long noncoding RNA MALAT1 polymorphisms with gastric cancer risk in Korean individuals. Mol. Genet. Genom. Med. 2020, 8, e1541. [Google Scholar] [CrossRef]
- Tripathi, V.; Ellis, J.D.; Shen, Z.; Song, D.Y.; Pan, Q.; Watt, A.T.; Freier, S.M.; Bennett, C.F.; Sharma, A.; Bubulya, P.A.; et al. The nuclear-retained noncoding RNA MALAT1 regulates alternative splicing by modulating SR splicing factor phosphorylation. Mol. Cell 2010, 39, 925–938. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, J.Z.; Xiang, J.J.; Wu, L.G.; Bai, Y.S.; Chen, Z.W.; Yin, X.Q.; Wang, Q.; Guo, W.H.; Peng, Y.; Guo, H.; et al. A genetic variant in long non-coding RNA MALAT1 associated with survival outcome among patients with advanced lung adenocarcinoma: A survival cohort analysis. BMC Cancer 2017, 17, 167. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Toraih, E.A.; Ellawindy, A.; Fala, S.Y.; Al Ageeli, E.; Gouda, N.S.; Fawzy, M.S.; Hosny, S. Oncogenic long noncoding RNA MALAT1 and HCV-related hepatocellular carcinoma. Biomed. Pharmacother. Biomed. Pharmacother. 2018, 102, 653–669. [Google Scholar] [CrossRef] [PubMed]
- Konishi, H.; Ichikawa, D.; Yamamoto, Y.; Arita, T.; Shoda, K.; Hiramoto, H.; Hamada, J.; Itoh, H.; Fujita, Y.; Komatsu, S.; et al. Plasma level of metastasis-associated lung adenocarcinoma transcript 1 is associated with liver damage and predicts development of hepatocellular carcinoma. Cancer Sci. 2016, 107, 149–154. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Wang, G.; Tao, K.; Cai, K.; Wu, K.; Ye, L.; Bai, J.; Yin, Y.; Wang, J.; Shuai, X.; et al. Long noncoding RNA metastasis-associated lung adenocarcinoma transcript 1 cooperates with enhancer of zeste homolog 2 to promote hepatocellular carcinoma development by modulating the microRNA-22/Snail family transcriptional repressor 1 axis. Cancer Sci. 2020, 111, 1582–1595. [Google Scholar] [CrossRef] [PubMed]
- Radwan, A.F.; Shaker, O.G.; El-Boghdady, N.A.; Senousy, M.A. Association of MALAT1 and PVT1 Variants, Expression Profiles and Target miRNA-101 and miRNA-186 with Colorectal Cancer: Correlation with Epithelial-Mesenchymal Transition. Int. J. Mol. Sci. 2021, 22, 6147. [Google Scholar] [CrossRef]
- Fathy, N.; Kortam, M.A.; Shaker, O.G.; Sayed, N.H. Long Noncoding RNAs MALAT1 and ANRIL Gene Variants and the Risk of Cerebral Ischemic Stroke: An Association Study. ACS Chem. Neurosci. 2021, 12, 1351–1362. [Google Scholar] [CrossRef]
- Li, Y.; Bao, C.; Gu, S.; Ye, D.; Jing, F.; Fan, C.; Jin, M.; Chen, K. Associations between novel genetic variants in the promoter region of MALAT1 and risk of colorectal cancer. Oncotarget 2017, 8, 92604–92614. [Google Scholar] [CrossRef] [Green Version]
- Zhou, J.; Wang, M.; Mao, A.; Zhao, Y.; Wang, L.; Xu, Y.; Jia, H.; Wang, L. Long noncoding RNA MALAT1 sponging miR-26a-5p to modulate Smad1 contributes to colorectal cancer progression by regulating autophagy. Carcinogenesis 2021, 42, 1370–1379. [Google Scholar] [CrossRef]
- Zhang, C.; Yao, K.; Zhang, J.; Wang, C.; Wang, C.; Qin, C. Long Noncoding RNA MALAT1 Promotes Colorectal Cancer Progression by Acting as a ceRNA of miR-508-5p to Regulate RAB14 Expression. BioMed Res. Int. 2020, 2020, 4157606. [Google Scholar] [CrossRef]
- Lampropoulou, D.I.; Aravantinos, G.; Katifelis, H.; Lazaris, F.; Laschos, K.; Theodosopoulos, T.; Papadimitriou, C.; Gazouli, M. Long non-coding RNA polymorphisms and prediction of response to chemotherapy based on irinotecan in patients with metastatic colorectal cancer. Cancer Biomark. Sect. A Dis. Markers 2019, 25, 213–221. [Google Scholar] [CrossRef] [PubMed]






| First Author | Year | Region | Ethnicity | Source of Controls | Type of Cancer | Sample Size (Case/Control) | Genotyping Method | NOS |
|---|---|---|---|---|---|---|---|---|
| Zhao | 2018 | China | Asian | HB | CRC | 400/400 | TaqMan | 7 |
| Yuan | 2019 | Taiwan | Asian | PB | HCC | 394/1199 | TaqMan | 7 |
| Ji | 2019 | China | Asian | HB | HCC | 624/618 | TaqMan | 7 |
| Orlandi | 2019 | Italy | Caucasian | PB | melanoma | 334/291 | PCR-RFLP | 6 |
| Qu | 2019 | China | Asian | HB | ESCC | 245/490 | TaqMan | 7 |
| Wen | 2019 | China | Asian | HB | PTC | 140/100 | TaqMan | 7 |
| Wu | 2019 | China | Asian | PB | CRC | 1078/1175 | TaqMan | 7 |
| Petkevicius | 2020 | Lithuania | Caucasian | HB | GC | 613/476 | TaqMan | 5 |
| Hong | 2020 | Korea | Asian | HB | GC | 1134/1228 | TaqMan | 5 |
| Ding | 2021 | Taiwan | Asian | PB | OSCC | 1350/1199 | TaqMan | 8 |
| First Author | Case | Control | HWE | ||||
|---|---|---|---|---|---|---|---|
| CC | CT | TT | CC | CT | TT | ||
| Zhao | 283 | 102 | 15 | 294 | 96 | 10 | Y |
| Yuan | 263 | 117 | 14 | 802 | 347 | 50 | Y |
| Ji | 464 | 149 | 9 | 453 | 152 | 12 | Y |
| Orlandi | 190 | 125 | 19 | 174 | 96 | 21 | Y |
| Qu | 148 | 79 | 18 | 338 | 133 | 19 | Y |
| Wen | 808 | 302 | 23 | 872 | 322 | 31 | Y |
| Wu | 751 | 294 | 33 | 856 | 292 | 27 | Y |
| Petkevicius | 416 | 171 | 21 | 335 | 126 | 14 | Y |
| Hong | 312 | 133 | 13 | 280 | 92 | 9 | Y |
| Ding | 948 | 363 | 39 | 807 | 347 | 45 | Y |
| Rs3200401 | N | Allelic Model (C vs. T) | Homozygote Model (CC vs. TT) | Heterozygote Model (CC vs. CT) | Dominant Model (CC vs. CT + TT) | Recessive Model (CC + CT vs. TT) | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| I2 | p | OR (95% CI) | pZ | I2 | p | OR (95% CI) | pZ | I2 | p | OR (95% CI) | pZ | I2 | p | OR (95% CI) | pZ | I2 | p | OR (95% CI) | pZ | ||
| Overall | 10 | 48.8% | 0.040 | 1.06 (0.97–1.16) | 0.215 | 24.0% | 0.222 | 1.03 (0.84–1.25) | 0.795 | 6.4% | 0.382 | 1.06 (0.98–1.14) | 0.167 | 43.1% | 0.071 | 1.04 (0.96–1.12) | 0.322 | 11.3% | 0.339 | 1.01 (0.83–1.22) | 0.951 |
| Cancer type | |||||||||||||||||||||
| CRC | 2 | 0.0% | 0.982 | 1.16 (1.01–1.33) | 0.033 | 0.0% | 0.820 | 1.44 (0.93–2.23) | 0.103 | 0.0% | 0.838 | 1.14 (0.97–1.34) | 0.125 | 0.0% | 0.918 | 1.16 (0.99–1.36) | 0.060 | 0.0% | 0.801 | 1.39 (0.90–2.15) | 0.137 |
| HCC | 2 | 0.0% | 0.729 | 0.96 (0.82–1.12) | 0.605 | 0.0% | 0.777 | 0.81 (0.49–1.34) | 0.414 | 0.0% | 0.698 | 0.99 (0.83–1.19) | 0.941 | 0.0% | 0.706 | 0.97 (0.82–1.16) | 0.772 | 0.0% | 0.804 | 0.81 (0.49,1.33) | 0.407 |
| Others | 4 | 72.2% | 0.013 | 1.04 (0.86–1.25) | 0.096 | 60.1% | 0.057 | 0.98 (0.62–1.53) | 0.918 | 48.7% | 0.119 | 1.01 (0.90,1.13) | 0.872 | 67.7% | 0.026 | 1.03 (0.84,1.27) | 0.746 | 52.1% | 0.100 | 0.95 (0.63,1.42) | 0.789 |
| GC | 2 | 0.0% | 0.479 | 1.16 (0.97–1.38) | 0.096 | 0.0% | 0.901 | 1.24 (0.72–2.13) | 0.432 | 0.0% | 0.415 | 1.18 (0.96–1.44) | 0.118 | 0.0% | 0.428 | 1.18 (0.97–1.44) | 0.094 | 0.0% | 0.965 | 1.19 (0.70,2.04) | 0.527 |
| Source of control | |||||||||||||||||||||
| HB | 6 | 46.3% | 0.097 | 1.11 (0.98-1.26) | 0.113 | 25.2% | 0.245 | 1.17 (0.88-1.56) | 0.281 | 0.0% | 0.494 | 1.09 (0.98,1.21) | 0.129 | 47.3% | 0.091 | 1.06 (0.93,1.18) | 0.261 | 12.0% | 0.339 | 1.14 (0.85,1.52) | 0.378 |
| PB | 4 | 54.5% | 0.086 | 1.01 (0.88–1.15) | 0.943 | 16.5% | 0.309 | 0.92 (0.70–1.20) | 0.526 | 35.4% | 0.200 | 1.02 (0.92,1.14) | 0.665 | 49.8% | 0.113 | 1.01 (0.91,1.13) | 0.786 | 3.5% | 0.375 | 0.91 (0.70,1.18) | 0.469 |
| Ethnicity | |||||||||||||||||||||
| Asian | 8 | 59.7% | 0.015 | 1.06 (0.95–1.19) | 0.437 | 37.6% | 0.129 | 1.03 (0.83–1.28) | 0.762 | 22.1% | 0.254 | 1.04 (0.96,1.13) | 0.300 | 53.7% | 0.035 | 1.05 (0.93,1.19) | 0.392 | 24.6% | 0.233 | 1.02 (0.82–1.26) | 0.858 |
| Caucasian | 2 | 0.0% | 0.753 | 1.07 (0.90–1.27) | 0.428 | 0.0% | 0.437 | 0.99 (0.62–1.59) | 0.971 | 0.0% | 0.693 | 1.13 (0.92–1.40) | 0.253 | 0.0% | 0.845 | 1.12 (0.92–1.37) | 0.261 | 0.0% | 0.383 | 0.94 (0.59–1.50) | 0.807 |
| Study quality | |||||||||||||||||||||
| High | 8 | 54.5% | 0.032 | 1.04 (0.94,1.16) | 0.449 | 38.1% | 0.126 | 1.00 (0.81–1.23) | 0.976 | 8.9% | 0.361 | 1.04 (0.95,1.13) | 0.392 | 47.0% | 0.067 | 1.02 (0.94,1.10) | 0.695 | 28.0% | 0.205 | 0.98 (0.80,1.21) | 0.855 |
| Medium | 2 | 0.0% | 0.479 | 1.16 (0.97–1.38) | 0.096 | 0.0% | 0.901 | 1.24 (0.72–2.13) | 0.432 | 0.0% | 0.415 | 1.18 (0.96–1.44) | 0.118 | 0.0% | 0.428 | 1.18 (0.97–1.44) | 0.094 | 0.0% | 0.965 | 1.19 (0.70,2.04) | 0.527 |
| SNP | Allelic Model | Homozygote Model | Heterozygote Model | Dominant Model | Recessive Model | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| Coef (95% CI) | p | Coef (95% CI) | p | Coef (95% CI) | p | Coef (95% CI) | p | Coef (95% CI) | p | |
| rs3200401 | Ethnicity | |||||||||
| 0.01 (−0.29,0.31) | 0.947 | −0.07 (−0.82,0.67) | 0.830 | 0.07 (−0.21,0.36) | 0.569 | 0.07 (−0.26,0.40) | 0.649 | −0.01 (−0.79,0.60) | 0.755 | |
| Source of control | ||||||||||
| −0.10 (−0.32,0.12) | 0.337 | −0.26 (−0.81,0.30) | 0.316 | −0.06 (−0.27,0.15) | 0.521 | −0.06 (−0.31,0.19) | 0.586 | −0.24 (−0.75,0.28) | 0.318 | |
| Study quality | ||||||||||
| 0.11 (−0.18,0.40) | 0.402 | 0.20 (−0.59,0.99) | 0.577 | 0.12 (−0.15,0.40) | 0.330 | 0.14 (−0.18,0.45) | 0.343 | 0.18 (−0.56,0.92) | 0.592 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, K.; Han, Z.; Wu, J.; Ye, H.; Sun, G.; Shi, J.; Zhang, J.; Wang, P. The Relationship between MALAT1 Polymorphism rs3200401 C > T and the Risk of Overall Cancer: A Meta-Analysis. Medicina 2022, 58, 176. https://doi.org/10.3390/medicina58020176
Li K, Han Z, Wu J, Ye H, Sun G, Shi J, Zhang J, Wang P. The Relationship between MALAT1 Polymorphism rs3200401 C > T and the Risk of Overall Cancer: A Meta-Analysis. Medicina. 2022; 58(2):176. https://doi.org/10.3390/medicina58020176
Chicago/Turabian StyleLi, Keming, Zhuo Han, Jinyu Wu, Hua Ye, Guiying Sun, Jianxiang Shi, Jianying Zhang, and Peng Wang. 2022. "The Relationship between MALAT1 Polymorphism rs3200401 C > T and the Risk of Overall Cancer: A Meta-Analysis" Medicina 58, no. 2: 176. https://doi.org/10.3390/medicina58020176
APA StyleLi, K., Han, Z., Wu, J., Ye, H., Sun, G., Shi, J., Zhang, J., & Wang, P. (2022). The Relationship between MALAT1 Polymorphism rs3200401 C > T and the Risk of Overall Cancer: A Meta-Analysis. Medicina, 58(2), 176. https://doi.org/10.3390/medicina58020176







