HLA-G 14bp Ins/Del Polymorphism in the 3′UTR Region and Acute Rejection in Kidney Transplant Recipients: An Updated Meta-Analysis
Abstract
:1. Introduction
2. Materials and Methods
2.1. Search Strategy
2.2. Study Selection
2.3. Data Extraction
2.4. Statistical Analysis
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Conflicts of Interest
References
- WHO. Constitution of the world health organization. Am. J. Public Health Nations Health 1946, 36, 1315–1323. [Google Scholar] [CrossRef] [Green Version]
- Cella, D.; Webster, K. Linking outcomes management to quality-of-life measurement. Oncology 1997, 11, 232–235. [Google Scholar]
- Abbasi, M.A.; Chertow, G.M.; Hall, Y.N. End-stage renal disease. BMJ Clin. Evid. 2010, 2010, 2002. [Google Scholar] [PubMed]
- Levey, A.S.; de Jong, P.E.; Coresh, J.; El Nahas, M.; Astor, B.C.; Matsushita, K.; Gansevoort, R.T.; Kasiske, B.L.; Eckardt, K.U. The definition, classification, and prognosis of chronic kidney disease: A kdigo controversies conference report. Kidney Int. 2011, 80, 17–28. [Google Scholar] [CrossRef] [Green Version]
- Farbiszewski, R.; Radecka, A.; Chwiecko, M.; Holownia, A. The effect of heparegen on antioxidant enzyme activities in ethanol-induced liver injury in rats. Alcohol 1992, 9, 403–407. [Google Scholar] [CrossRef]
- Fiebiger, W.; Mitterbauer, C.; Oberbauer, R. Health-related quality of life outcomes after kidney transplantation. Health Qual. Life Outcomes 2004, 2, 2. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Abu Jawdeh, B.G.; Govil, A. Acute kidney injury in transplant setting: Differential diagnosis and impact on health and health care. Adv. Chronic Kidney Dis. 2017, 24, 228–232. [Google Scholar] [CrossRef] [PubMed]
- Kasiske, B.L.; Anjum, S.; Shah, R.; Skogen, J.; Kandaswamy, C.; Danielson, B.; O’Shaughnessy, E.A.; Dahl, D.C.; Silkensen, J.R.; Sahadevan, M.; et al. Hypertension after kidney transplantation. Am. J. Kidney Dis. 2004, 43, 1071–1081. [Google Scholar] [CrossRef] [PubMed]
- Carosella, E.D.; Rouas-Freiss, N.; Tronik-Le Roux, D.; Moreau, P.; LeMaoult, J. Hla-g: An immune checkpoint molecule. Adv. Immunol. 2015, 127, 33–144. [Google Scholar]
- Kovats, S.; Main, E.K.; Librach, C.; Stubblebine, M.; Fisher, S.J.; DeMars, R. A class i antigen, hla-g, expressed in human trophoblasts. Science 1990, 248, 220–223. [Google Scholar] [CrossRef]
- LeMaoult, J.; Zafaranloo, K.; Le Danff, C.; Carosella, E.D. Hla-g up-regulates ilt2, ilt3, ilt4, and kir2dl4 in antigen presenting cells, nk cells, and t cells. FASEB J. 2005, 19, 662–664. [Google Scholar] [CrossRef]
- Qiu, J.; Terasaki, P.I.; Miller, J.; Mizutani, K.; Cai, J.; Carosella, E.D. Soluble hla-g expression and renal graft acceptance. Am. J. Transpl. 2006, 6, 2152–2156. [Google Scholar] [CrossRef]
- Luque, J.; Torres, M.I.; Aumente, M.D.; Marin, J.; Garcia-Jurado, G.; Gonzalez, R.; Pascual, D.; Guerra, N.; Lopez-Rubio, F.; Alvarez-Lopez, M.R.; et al. Soluble hla-g in heart transplantation: Their relationship to rejection episodes and immunosuppressive therapy. Hum. Immunol. 2006, 67, 257–263. [Google Scholar] [CrossRef]
- Creput, C.; Le Friec, G.; Bahri, R.; Amiot, L.; Charpentier, B.; Carosella, E.; Rouas-Freiss, N.; Durrbach, A. Detection of hla-g in serum and graft biopsy associated with fewer acute rejections following combined liver-kidney transplantation: Possible implications for monitoring patients. Hum. Immunol. 2003, 64, 1033–1038. [Google Scholar] [CrossRef]
- Hviid, T.V.; Rizzo, R.; Melchiorri, L.; Stignani, M.; Baricordi, O.R. Polymorphism in the 5’ upstream regulatory and 3’ untranslated regions of the hla-g gene in relation to soluble hla-g and il-10 expression. Hum. Immunol. 2006, 67, 53–62. [Google Scholar] [CrossRef] [PubMed]
- Darbas, S.; Yilmaz, V.T.; Kocak, H.; Kisaoglu, A.; Demiryilmaz, I.; Aydinli, B.; Arslan, H.S.; Ucar, F. New markers for predictions of acute and chronic rejection and graft outcomes in kidney transplant recipients; hla-g gene 3′utr 14 bp polymorphism and shla-g. Gene 2021, 790, 145712. [Google Scholar] [CrossRef] [PubMed]
- Xu, C.X.; Zhang, Y.L.; Huang, X.Y.; Han, F.; Jin, Z.K.; Tian, P.X.; Dou, M. Prediction of acute renal allograft rejection by combined hla-g 14-bp insertion/deletion genotype analysis and detection of kidney injury molecule-1 and osteopontin in the peripheral blood. Transpl. Immunol. 2021, 65, 101371. [Google Scholar] [CrossRef] [PubMed]
- Durmanova, V.; Bandzuchova, H.; Zilinska, Z.; Tirpakova, J.; Kuba, D.; Buc, M.; Polakova, K. Association of hla-g polymorphisms in the 3’utr region and soluble hla-g with kidney graft outcome. Immunol. Investig. 2019, 48, 644–658. [Google Scholar] [CrossRef]
- Janssen, M.; Thaiss, F.; Nashan, B.; Koch, M.; Thude, H. Donor derived hla-g polymorphisms have a significant impact on acute rejection in kidney transplantation. Hum. Immunol. 2019, 80, 176–183. [Google Scholar] [CrossRef]
- Misra, M.K.; Prakash, S.; Kapoor, R.; Pandey, S.K.; Sharma, R.K.; Agrawal, S. Association of hla-g promoter and 14-bp insertion-deletion variants with acute allograft rejection and end-stage renal disease. Tissue Antigens 2013, 82, 317–326. [Google Scholar] [CrossRef]
- Littera, R.; Piredda, G.; Pani, A.; Frongia, M.; Onano, B.; Michittu, M.B.; Murgia, M.G.; Lai, S.; Alba, F.; Valentini, M.D.; et al. Role of human leukocyte antigen-g 14-base pair polymorphism in kidney transplantation outcomes. J. Nephrol. 2013, 26, 1170–1178. [Google Scholar] [CrossRef]
- Jin, Z.K.; Xu, C.X.; Tian, P.X.; Xue, W.J.; Ding, X.M.; Zheng, J.; Ding, C.G.; Ge, G.Q.; Mao, T.C.; Lin, Y. Impact of hla-g 14-bp polymorphism on acute rejection and cytomegalovirus infection in kidney transplant recipients from northwestern China. Transpl. Immunol. 2012, 27, 69–74. [Google Scholar] [CrossRef]
- Ciliao Alves, D.C.; de Oliveira Crispim, J.C.; Castelli, E.C.; Mendes-Junior, C.T.; Deghaide, N.H.; Barros Silva, G.E.; Costa, R.S.; Saber, L.T.; Moreau, P.; Donadi, E.A. Human leukocyte antigen-g 3′ untranslated region polymorphisms are associated with better kidney allograft acceptance. Hum. Immunol 2012, 73, 52–59. [Google Scholar] [CrossRef]
- Aghdaie, M.H.; Azarpira, N.; Kazemi, K.; Geramizadeh, B.; Darai, M.; Malekhoseini, S.A. Frequency of hla-g exon 8 polymorphisms and kidney allograft outcome in iranian population. Mol. Biol. Rep. 2011, 38, 3593–3597. [Google Scholar] [CrossRef]
- Hou, W.; Huang, Z.; Ji, Z.; Zhou, J. Human leukocyte antigen-g-14-base-pair-insertion/deletion polymorphism and graft survival in kidney transplant recipients. Exp. Clin. Transpl. 2014, 12, 89–94. [Google Scholar]
- Genomes Project, C.; Auton, A.; Brooks, L.D.; Durbin, R.M.; Garrison, E.P.; Kang, H.M.; Korbel, J.O.; Marchini, J.L.; McCarthy, S.; McVean, G.A.; et al. A global reference for human genetic variation. Nature 2015, 526, 68–74. [Google Scholar]
- Katznelson, S.; Gjertson, D.W.; Cecka, J.M. The effect of race and ethnicity on kidney allograft outcome. Clin. Transpl. 1995, 1995, 379–394. [Google Scholar]
- Cohen, O.; De La Zerda, D.; Beygui, R.E.; Hekmat, D.; Laks, H. Ethnicity as a predictor of graft longevity and recipient mortality in heart transplantation. Transpl. Proc. 2007, 39, 3297–3302. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; Han, Y.; Huang, H.; Bi, L.; Kong, X.; Ma, X.; Shi, B.; Xiao, L. Circulating nk cell subsets and nktlike cells in renal transplant recipients with acute tcellmediated renal allograft rejection. Mol. Med. Rep. 2019, 19, 4238–4248. [Google Scholar] [PubMed] [Green Version]
- Kroemer, A.; Edtinger, K.; Li, X.C. The innate natural killer cells in transplant rejection and tolerance induction. Curr. Opin. Organ. Transpl. 2008, 13, 339–343. [Google Scholar] [CrossRef] [PubMed]
- Harrison, G.A.; Humphrey, K.E.; Jakobsen, I.B.; Cooper, D.W. A 14 bp deletion polymorphism in the hla-g gene. Hum. Mol. Genet. 1993, 2, 2200. [Google Scholar] [CrossRef]
- Martelli-Palomino, G.; Pancotto, J.A.; Muniz, Y.C.; Mendes-Junior, C.T.; Castelli, E.C.; Massaro, J.D.; Krawice-Radanne, I.; Poras, I.; Rebmann, V.; Carosella, E.D.; et al. Polymorphic sites at the 3’ untranslated region of the hla-g gene are associated with differential hla-g soluble levels in the brazilian and french population. PLoS ONE 2013, 8, e71742. [Google Scholar]
- Rousseau, P.; Le Discorde, M.; Mouillot, G.; Marcou, C.; Carosella, E.D.; Moreau, P. The 14 bp deletion-insertion polymorphism in the 3′ ut region of the hla-g gene influences hla-g mrna stability. Hum. Immunol. 2003, 64, 1005–1010. [Google Scholar] [CrossRef] [PubMed]
- Polakova, K.; Bandzuchova, H.; Zilinska, Z.; Chrenova, S.; Kuba, D.; Russ, G. Analysis of hla-g expression in serum and biopsy samples of kidney transplant recipients. Immunobiology 2015, 220, 533–537. [Google Scholar] [CrossRef]
- LeMaoult, J.; Krawice-Radanne, I.; Dausset, J.; Carosella, E.D. Hla-g1-expressing antigen-presenting cells induce immunosuppressive cd4+ t cells. Proc. Natl. Acad. Sci. USA 2004, 101, 7064–7069. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ge, Y.Z.; Ge, Q.; Li, M.H.; Shi, G.M.; Xu, X.; Xu, L.W.; Xu, Z.; Lu, T.Z.; Wu, R.; Zhou, L.H.; et al. Association between human leukocyte antigen-g 14-bp insertion/deletion polymorphism and cancer risk: A meta-analysis and systematic review. Hum. Immunol. 2014, 75, 827–832. [Google Scholar] [CrossRef]
- Fabris, A.; Segat, L.; Catamo, E.; Morgutti, M.; Vendramin, A.; Crovella, S. Hla-g 14 bp deletion/insertion polymorphism in celiac disease. Am. J. Gastroenterol. 2011, 106, 139–144. [Google Scholar] [CrossRef]
- Rebmann, V.; van der Ven, K.; Passler, M.; Pfeiffer, K.; Krebs, D.; Grosse-Wilde, H. Association of soluble hla-g plasma levels with hla-g alleles. Tissue Antigens 2001, 57, 15–21. [Google Scholar] [CrossRef]
- Chiusolo, P.; Bellesi, S.; Piccirillo, N.; Giammarco, S.; Marietti, S.; De Ritis, D.; Metafuni, E.; Stignani, M.; Baricordi, O.R.; Sica, S.; et al. The role of hla--g 14-bp polymorphism in allo-hsct after short-term course mtx for gvhd prophylaxis. Bone Marrow Transpl. 2012, 47, 120–124. [Google Scholar] [CrossRef]
- Rizzo, R.; Rubini, M.; Govoni, M.; Padovan, M.; Melchiorri, L.; Stignani, M.; Carturan, S.; Ferretti, S.; Trotta, F.; Baricordi, O.R. Hla-g 14-bp polymorphism regulates the methotrexate response in rheumatoid arthritis. Pharm. Genom. 2006, 16, 615–623. [Google Scholar] [CrossRef]
- Park, Y.; Park, Y.; Kim, Y.S.; Kwon, O.J.; Kim, H.S. Allele frequencies of human leukocyte antigen-g in a korean population. Int. J. Immunogenet. 2012, 39, 39–45. [Google Scholar] [CrossRef] [PubMed]
- Kramer, C.S.M.; Roelen, D.L.; Heidt, S.; Claas, F.H.J. Defining the immunogenicity and antigenicity of hla epitopes is crucial for optimal epitope matching in clinical renal transplantation. HLA 2017, 90, 5–16. [Google Scholar] [CrossRef] [PubMed]

| Author | Nationality | Immunosuppressive Treatment | Acute Rejection | Non-Acute Rejection | HWE p | ||||
|---|---|---|---|---|---|---|---|---|---|
| (Publish Year) | Ins/Ins | Ins/Del | Del/Del | Ins/Ins | Ins/Del | Del/Del | |||
| Ahgdaie et al. (2011) | Iran | cyclosporine, azathioprine, prednisone, mycophenolate mofetil | 6 | 33 | 33 | 8 | 38 | 26 | 0.86 |
| Ciliao Alves et al. (2012) | Brazil | (cyclosporine or tacrolimus), methylprednisone, and mycophenolate mofetil | 1 | 10 | 4 | 8 | 19 | 14 | 1.00 |
| Jin et al. (2012) | China | (tacrolimus or cyclosporine), mycophenolate mofetil and prednisone | 17 | 18 | 7 | 18 | 49 | 35 | 1.00 |
| Misra et al. (2013) | India | tacrolimus, mycophenolate mofetil, prednisolone, cyclosporine, everolimus, basiliximab, antithymocyte globulin | 14 | 12 | 9 | 23 | 69 | 56 | 1.00 |
| Littera et al. (2013) | Italy | rapamycin, steroids, and cyclosporine and/or mycophenolate mofetil | 2 | 4 | 4 | 66 | 163 | 97 | 1.00 |
| Durmanova et al. (2019) | Slovakia | basiliximab/daclizumab, tacrolimus, mycophenolate mofetil, corticosteroid, cyclosporine, antithymocyte globulin | 7 | 20 | 10 | 8 | 10 | 14 | 0.42 |
| Janssen et al. (2019) | Germany | calcineurin inhibitors, (mycophenolate mofetil or mTOR inhibitor) and steroids | 6 | 22 | 13 | 28 | 59 | 47 | 0.86 |
| Xu et al. (2021) | China | tacrolimus, mycophenolate mofetil, prednisone, cyclosporine | 13 | 14 | 5 | 7 | 22 | 16 | 1.00 |
| Darbas et al. (2021) | Turkey | antithymocyte globulin, basiliximab, mycophenolate mofetil, sirolimus, tacrolimus, everolimus, cyclosporine | 23 | 23 | 6 | 3 | 19 | 30 | 1.00 |
| Genetic Models | Association Test | Heterogeneity | Model | Publication Bias | |||
|---|---|---|---|---|---|---|---|
| OR | 95% CI | p | p | I2 | Egger’s Test p-Val | ||
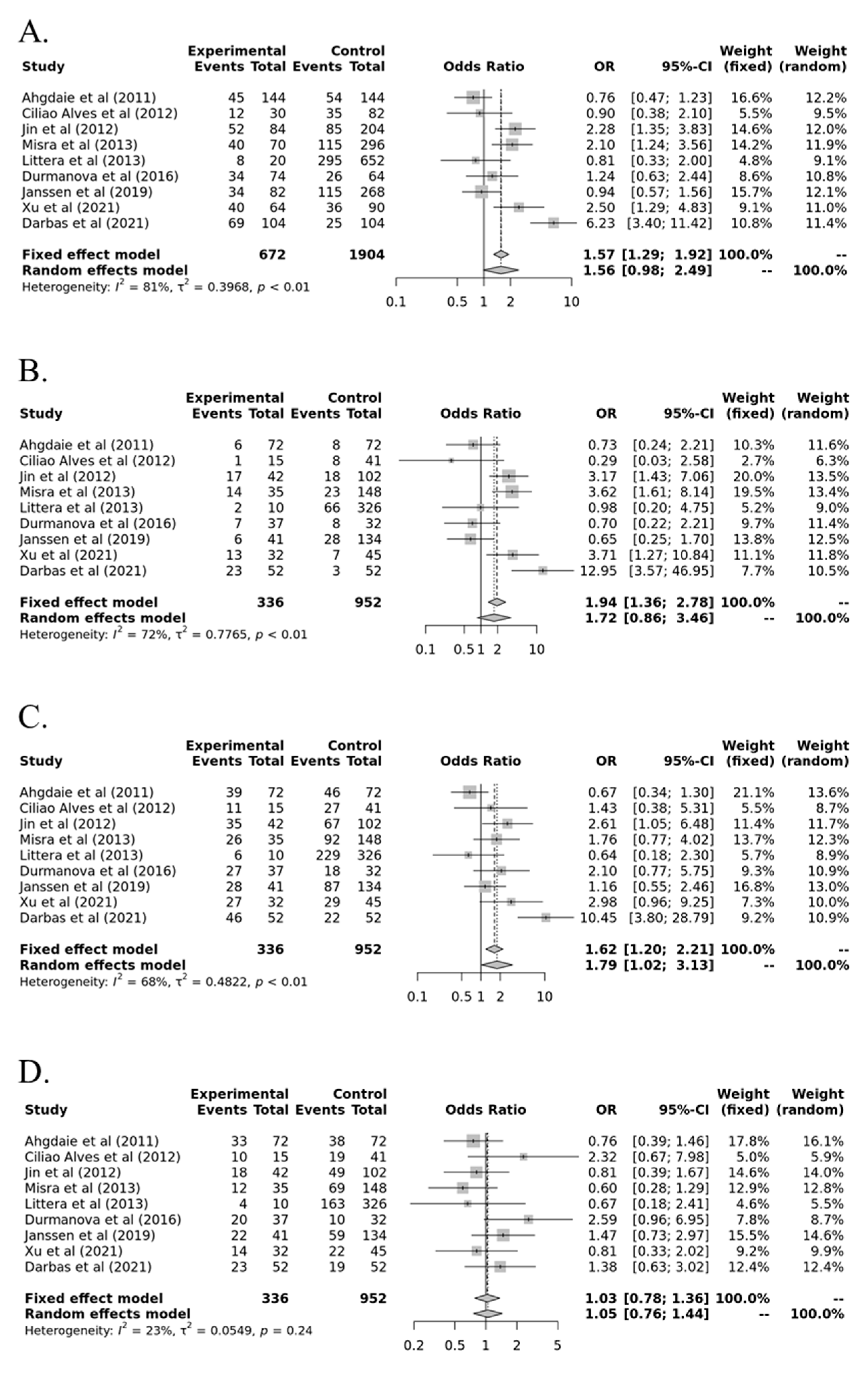
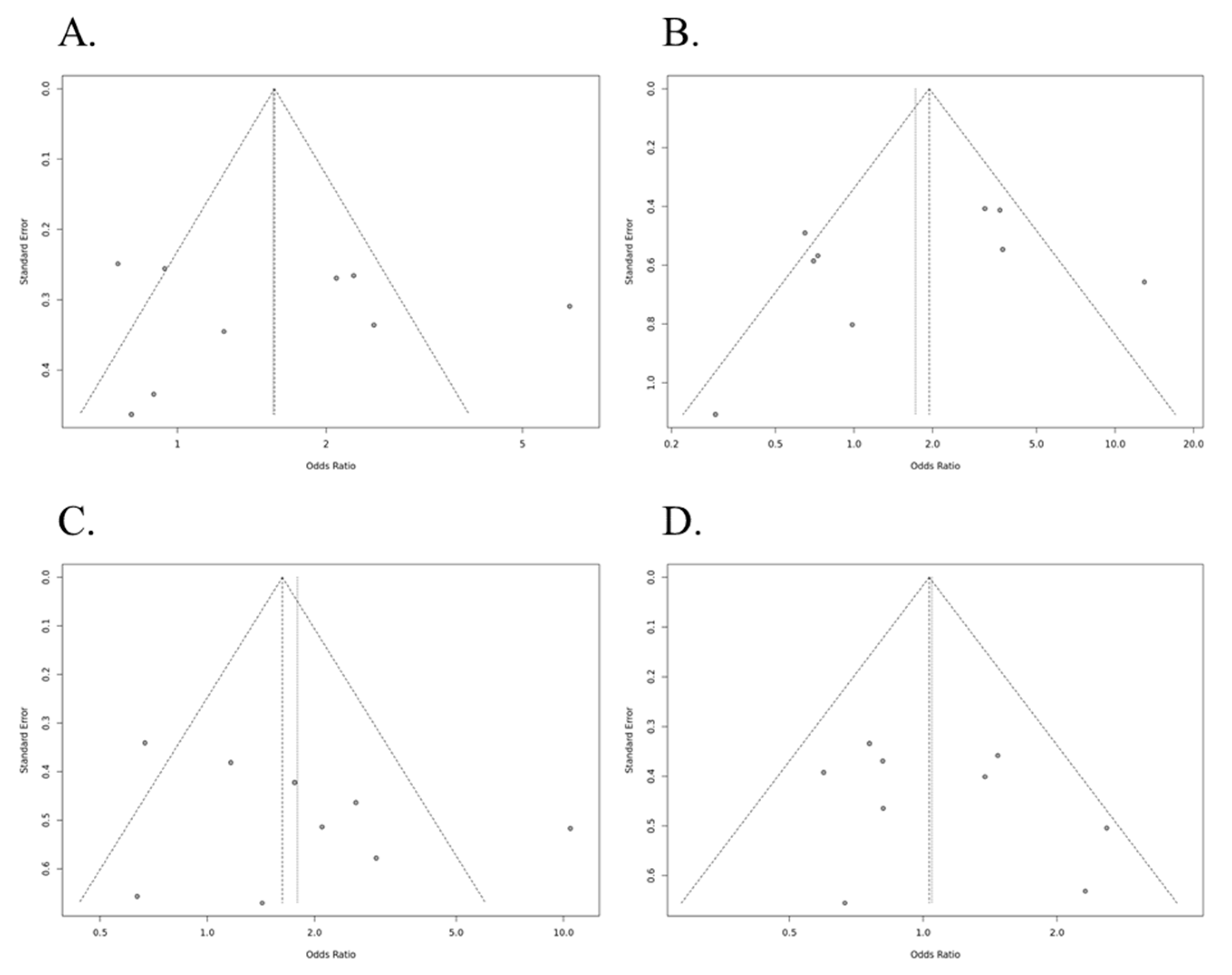
| Allele (Ins vs. Del) | 1.564 | 0.983–2.486 | 0.411 | <0.01 | 0.81 | Random | 0.964 |
| Recessive (Ins/Ins vs. Ins/Del+Del/Del) | 1.721 | 0.856–3.462 | 0.895 | <0.01 | 0.72 | Random | 0.351 |
| Dominant (Ins/Ins+ Ins/Del vs. Del/Del) | 1.790 | 1.023–3.131 | 0.290 | <0.01 | 0.68 | Random | 0.306 |
| Overdominant (Ins/Del vs. Ins/Ins+Del/Del) | 1.032 | 0.783–1.361 | 1.000 | 0.23 | 0.24 | Fixed | 0.392 |
| Genetic Models | Nationality | Association Test | Heterogeneity | Model | Publication Bias | |||
|---|---|---|---|---|---|---|---|---|
| OR | 95% CI | p | p | I2 | Egger’s Test p-Val | |||
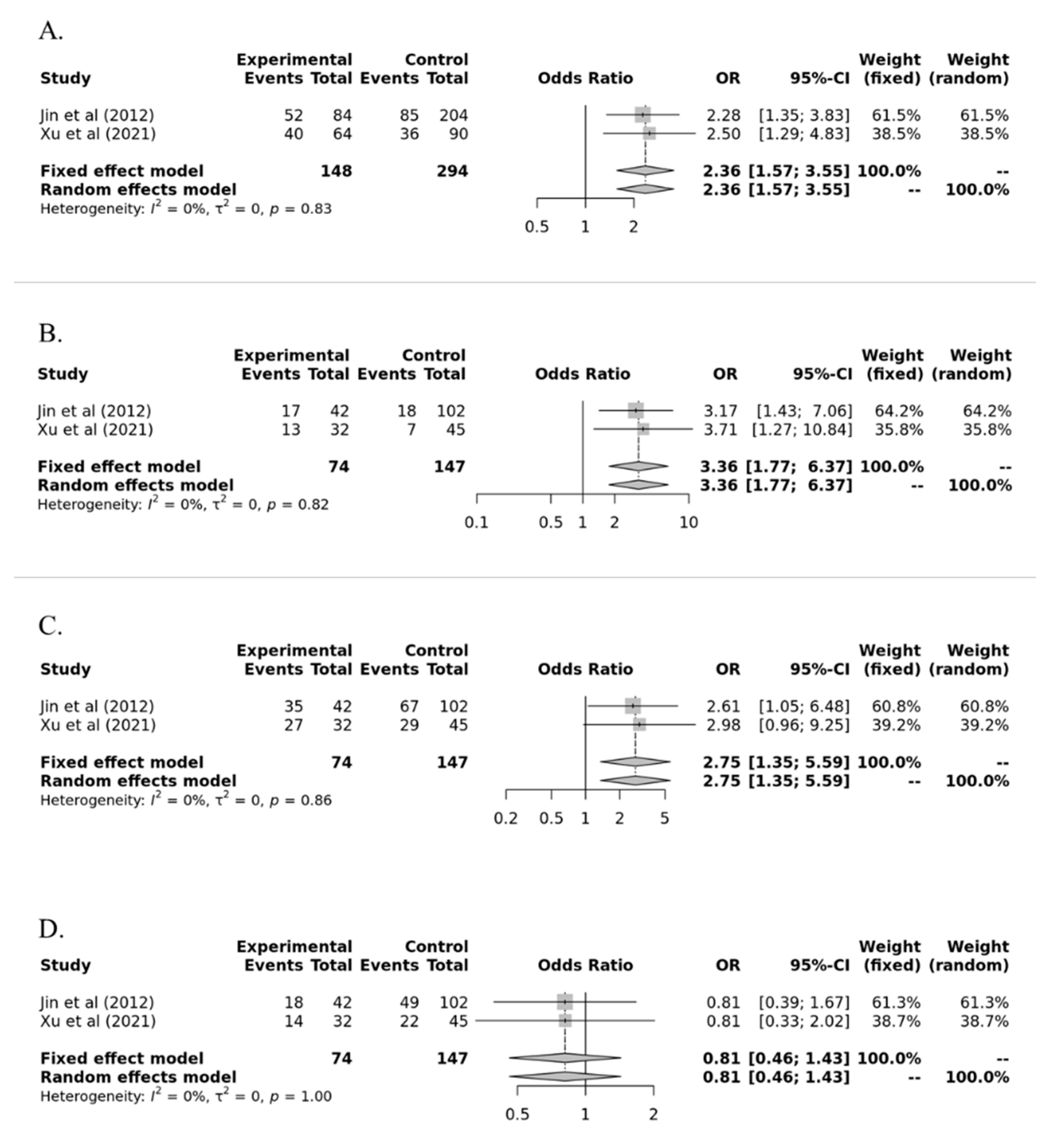
| Allele | Asian | 2.359 | 1.568–3.550 | 3.8 × 10−5 | 0.83 | 0.00 | Fixed | NA |
| Caucasian | 1.377 | 0.773–2.453 | 0.278 | <0.01 | 0.84 | Random | 0.9832 | |
| Recessive | Asian | 3.357 | 1.769–6.370 | 0.0002 | 0.82 | 0.00 | Fixed | NA |
| Caucasian | 1.347 | 0.5456–3.3237 | 0.518 | <0.01 | 0.75 | Random | 0.5363 | |
| Dominant | Asian | 2.750 | 1.354–5.587 | 0.0052 | 0.86 | 0.00 | Fixed | NA |
| Caucasian | 1.590 | 0.804–3.144 | 0.183 | <0.01 | 0.73 | Random | 0.4594 | |
| Overdominant | Asian | 0.812 | 0.461–1.431 | 0.471 | 1.00 | 0.00 | Fixed | NA |
| Caucasian | 1.112 | 0.810–1.527 | 0.511 | 0.15 | 0.37 | Fixed | 0.4574 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kang, S.W.; Oh, E.; Cho, W.; Kim, M.; Park, E.J.; Kwack, K.H.; Chung, K.; Nam, O.H.; Chae, Y.K.; Ban, J.Y. HLA-G 14bp Ins/Del Polymorphism in the 3′UTR Region and Acute Rejection in Kidney Transplant Recipients: An Updated Meta-Analysis. Medicina 2021, 57, 1007. https://doi.org/10.3390/medicina57101007
Kang SW, Oh E, Cho W, Kim M, Park EJ, Kwack KH, Chung K, Nam OH, Chae YK, Ban JY. HLA-G 14bp Ins/Del Polymorphism in the 3′UTR Region and Acute Rejection in Kidney Transplant Recipients: An Updated Meta-Analysis. Medicina. 2021; 57(10):1007. https://doi.org/10.3390/medicina57101007
Chicago/Turabian StyleKang, Sang Wook, Eunkyung Oh, Wonwoo Cho, Minseok Kim, Eo Jin Park, Kyu Hwan Kwack, Kang Chung, Ok Hyung Nam, Yong Kwon Chae, and Ju Yeon Ban. 2021. "HLA-G 14bp Ins/Del Polymorphism in the 3′UTR Region and Acute Rejection in Kidney Transplant Recipients: An Updated Meta-Analysis" Medicina 57, no. 10: 1007. https://doi.org/10.3390/medicina57101007
APA StyleKang, S. W., Oh, E., Cho, W., Kim, M., Park, E. J., Kwack, K. H., Chung, K., Nam, O. H., Chae, Y. K., & Ban, J. Y. (2021). HLA-G 14bp Ins/Del Polymorphism in the 3′UTR Region and Acute Rejection in Kidney Transplant Recipients: An Updated Meta-Analysis. Medicina, 57(10), 1007. https://doi.org/10.3390/medicina57101007







