Histone-, Receptor-, and Integrin-Related Gene Products and ADAM28 as Relevant to B-Cell Acute Lymphoblastic Leukemia (B-ALL)
Abstract
1. Introduction
2. Materials and Methods
2.1. Data Acquisition
2.2. Preprocessing of RNA-Sequencing Data
2.3. Downstream Analyses
3. Results
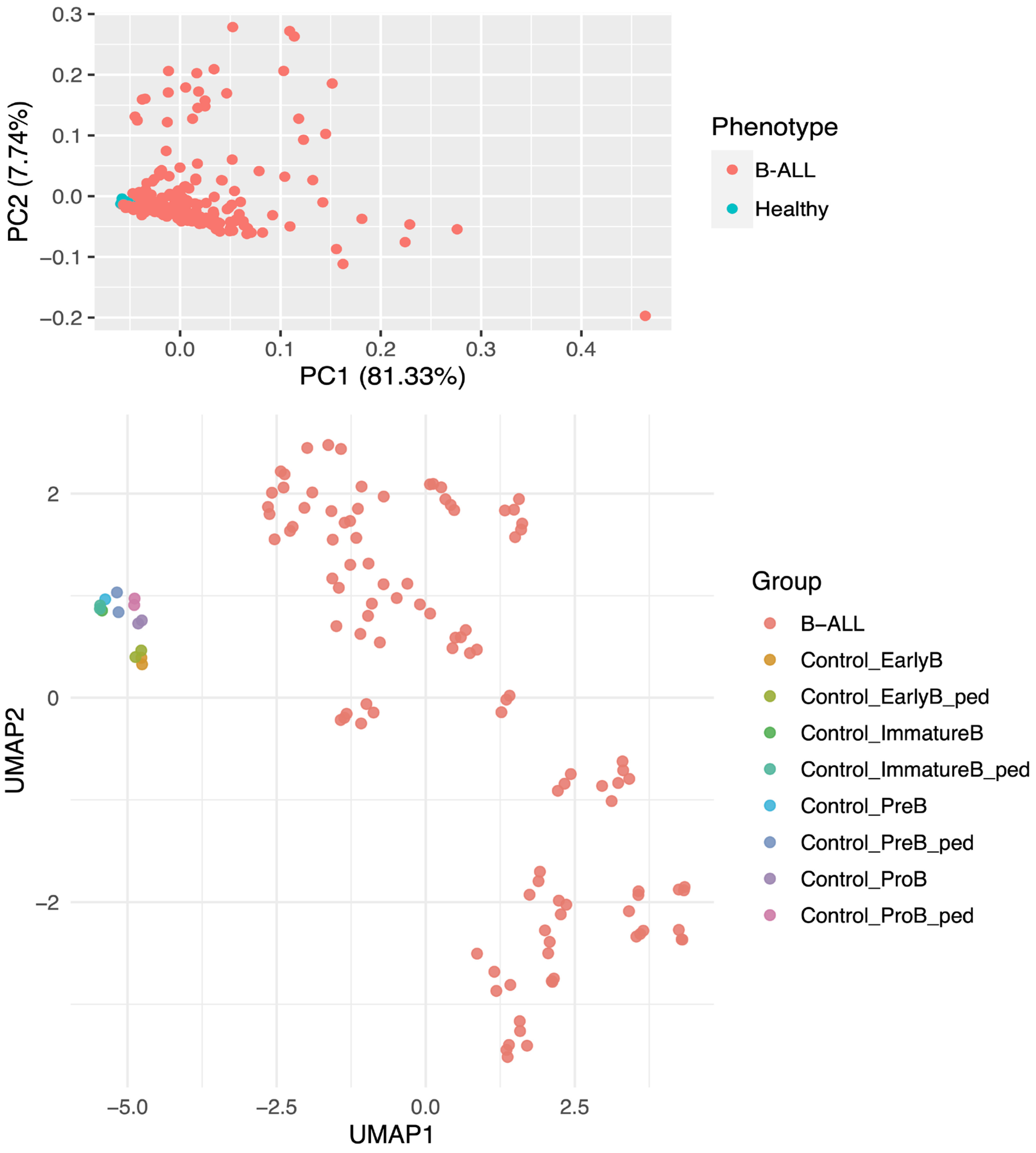
3.1. Unsupervised Clustering Reveals Separation of Case and Control Samples
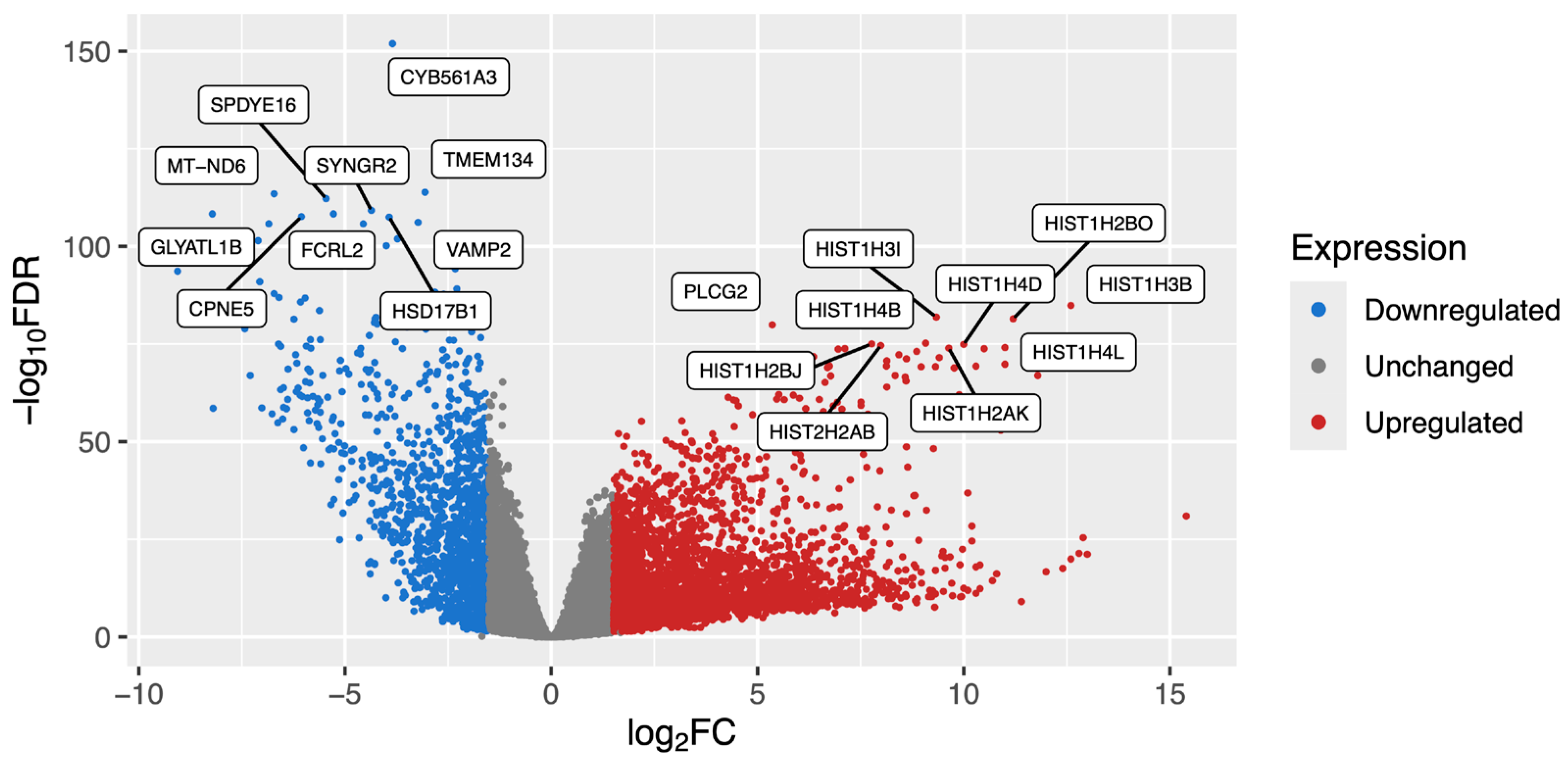
3.2. Differential Gene Expression Highlights Significant Differences in B-ALL
3.3. Signaling Pathway Enrichment Identifies Eight Biologically Relevant Cascades
3.4. Therapeutic Target Prioritization Predicts Known and Novel Targets
3.5. Machine Learning Identified ADAM28 as a Mechanistic Marker
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| B-ALL | B-cell acute lymphoblastic leukemia |
| GEO | Gene Expression Omnibus |
| DEG | Differentially expressed genes |
| SRA | Sequence read archive |
| ARMOR | Automated Reproducible Modular workflow for preprocessing and differential analysis of RNA-seq data |
| SPIA | Signaling pathway impact analysis |
| PCA | Principal component analysis |
| UMAP | Uniform manifold approximation and projection |
| FDR | false discovery rate-adjusted p-value |
| HIST2H2AA3 | Histone cluster 2 H2A family member A3 |
| HIST1H3B | H3 clustered histone 2 |
| CLEC14A | C-type lectin domain containing 14A |
| EPHA7 | Ephrin receptor A7 |
| NPR1 | natriuretic peptide receptor 1 |
| ANGPTL1 | Angiopoietin-like 1 |
| IGHA1 | Immunoglobulin chain formation-associated immunoglobulin heavy constant alpha 1 |
| IGKV3D-11 | Immunoglobulin kappa variable 3D-11 |
| CHAD | Adhesion-associated chondroadherin |
| GLYATL1B | Glycine-N-acyltransferase like 1B |
| CHURC1-FNTB | Churchill domain containing 1-farnesyltransferase fusion |
| TNF | Tumor necrosis factor alpha |
| IL6 | Interleukin 6 |
| p53 | Tumor protein p53 |
| AKT1 | AKT serine/threonine kinase 1 |
| FLT1 | FMS-related receptor tyrosine kinase 1 |
| CD40 | Cluster of differentiation 40 |
| TLR8 | Toll-like receptor 8 |
| GRM5 | Glutamate metabotropic receptor 5 |
| ADAM28 | ADAM metallopeptidase domain 28 |
| DIPK1C | Divergent protein kinase domain 1C |
| BCP-ALL | Precursor B-ALL |
| BAALC | BAALC binder of MAP3K1 and KLF4 |
| MLL-R | mixed-lineage leukemia gene rearrangement |
| AML | Acute myeloid leukemia |
| BCL2 | BCL2 apoptosis regulator |
References
- Inaba, H.; Pui, C.H. Advances in the Diagnosis and Treatment of Pediatric Acute Lymphoblastic Leukemia. J. Clin. Med. 2021, 10, 1926. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Bhojwani, D.; Yang, J.J.; Pui, C.H. Biology of childhood acute lymphoblastic leukemia. Pediatr. Clin. N. Am. 2015, 62, 47–60. [Google Scholar] [CrossRef] [PubMed Central]
- Terwilliger, T.; Abdul-Hay, M. Acute lymphoblastic leukemia: A comprehensive review and 2017 update. Blood Cancer J. 2017, 7, e577. [Google Scholar] [CrossRef] [PubMed Central]
- Sasaki, K.; Jabbour, E.; Short, N.J.; Jain, N.; Ravandi, F.; Pui, C.H.; Kantarjian, H. Acute lymphoblastic leukemia: A population-based study of outcome in the United States based on the surveillance, epidemiology, and end results (SEER) database, 1980–2017. Am. J. Hematol. 2021, 96, 650–658, Erratum in Am. J. Hematol. 2021, 96, 1344. [Google Scholar] [CrossRef] [PubMed Central]
- Zhang, R.; Zhu, H.; Yuan, Y.; Zhao, J.; Yang, X.; Tian, Z. Risk Factors for Relapse of Childhood B Cell Acute Lymphoblastic Leukemia. Med. Sci. Monit. 2020, 26, e923271. [Google Scholar] [CrossRef] [PubMed Central]
- Junk, S.V.; Förster, A.; Schmidt, G.; Zimmermann, M.; Fedders, B.; Haermeyer, B.; Bergmann, A.K.; Möricke, A.; Cario, G.; Auber, B.; et al. Germline variants in patients developing second malignant neoplasms after therapy for pediatric acute lymphoblastic leukemia-a case-control study. Leukemia 2024, 38, 887–892. [Google Scholar] [CrossRef] [PubMed Central]
- Choed-Amphai, C.; Khorana, J.; Sathitsamitphong, L.; Natesirinilkul, R.; Charoenkwan, P. Predictive factors for L-asparaginase hypersensitivity in pediatric acute lymphoblastic leukemia. Int. J. Hematol. 2024, 119, 442–449. [Google Scholar] [CrossRef] [PubMed]
- Daub, J.T.; Amini, S.; Kersjes, D.J.E.; Ma, X.; Jäger, N.; Zhang, J.; Pfister, S.M.; Holstege, F.C.P.; Kemmeren, P. A systematic analysis of genetic interactions and their underlying biology in childhood cancer. Commun. Biol. 2021, 4, 1139. [Google Scholar] [CrossRef] [PubMed Central]
- Al-Mahayri, Z.N.; AlAhmad, M.M.; Ali, B.R. Long-Term Effects of Pediatric Acute Lymphoblastic Leukemia Chemotherapy: Can Recent Findings Inform Old Strategies? Front. Oncol. 2021, 11, 710163. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Clough, E.; Barrett, T.; Wilhite, S.E.; Ledoux, P.; Evangelista, C.; Kim, I.F.; Tomashevsky, M.; Marshall, K.A.; Phillippy, K.H.; Sherman, P.M.; et al. NCBI GEO: Archive for gene expression and epigenomics data sets: 23-year update. Nucleic Acids Res. 2024, 52, D138–D144. [Google Scholar] [CrossRef] [PubMed Central]
- Katz, K.; Shutov, O.; Lapoint, R.; Kimelman, M.; Brister, J.R.; O’Sullivan, C. The Sequence Read Archive: A decade more of explosive growth. Nucleic Acids Res. 2022, 50, D387–D390. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Saint Fleur-Lominy, S.; Evensen, N.A.; Bhatla, T.; Sethia, G.; Narang, S.; Choi, J.H.; Ma, X.; Yang, J.J.; Kelly, S.; Raetz, E.; et al. Evolution of the Epigenetic Landscape in Childhood B Acute Lymphoblastic Leukemia and Its Role in Drug Resistance. Cancer Res. 2020, 80, 5189–5202. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Autry, R.J.; Paugh, S.W.; Carter, R.; Shi, L.; Liu, J.; Ferguson, D.C.; Lau, C.E.; Bonten, E.J.; Yang, W.; McCorkle, J.R.; et al. Integrative genomic analyses reveal mechanisms of glucocorticoid resistance in acute lymphoblastic leukemia. Nat. Cancer 2020, 1, 329–344. [Google Scholar] [CrossRef] [PubMed Central]
- Neveu, B.; Spinella, J.F.; Richer, C.; Lagacé, K.; Cassart, P.; Lajoie, M.; Jananji, S.; Drouin, S.; Healy, J.; Hickson, G.R.X.; et al. CLIC5: A novel ETV6 target gene in childhood acute lymphoblastic leukemia. Haematologica 2016, 101, 1534–1543. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Black, K.L.; Naqvi, A.S.; Asnani, M.; Hayer, K.E.; Yang, S.Y.; Gillespie, E.; Bagashev, A.; Pillai, V.; Tasian, S.K.; Gazzara, M.R.; et al. Aberrant splicing in B-cell acute lymphoblastic leukemia. Nucleic Acids Res. 2018, 46, 11357–11369. [Google Scholar] [CrossRef] [PubMed Central]
- Shangguan, S.; Ehrenberg, P.K.; Geretz, A.; Yum, L.; Kundu, G.; May, K.; Fourati, S.; Nganou-Makamdop, K.; Williams, L.D.; Sawant, S.; et al. Monocyte-derived transcriptome signature indicates antibody-dependent cellular phagocytosis as a potential mechanism of vaccine-induced protection against HIV-1. eLife 2021, 10, e69577. [Google Scholar] [CrossRef] [PubMed Central]
- Panwar, B.; Schmiedel, B.J.; Liang, S.; White, B.; Rodriguez, E.; Kalunian, K.; McKnight, A.J.; Soloff, R.; Seumois, G.; Vijayanand, P.; et al. Multi-cell type gene coexpression network analysis reveals coordinated interferon response and cross-cell type correlations in systemic lupus erythematosus. Genome Res. 2021, 31, 659–676. [Google Scholar] [CrossRef] [PubMed Central]
- Orjuela, S.; Huang, R.; Hembach, K.M.; Robinson, M.D.; Soneson, C. ARMOR: An utomated eproducible MOdular Workflow for Preprocessing and Differential Analysis of NA-seq Data. G3 2019, 9, 2089–2096. [Google Scholar] [CrossRef] [PubMed Central]
- Patro, R.; Duggal, G.; Love, M.I.; Irizarry, R.A.; Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 2017, 14, 417–419. [Google Scholar] [CrossRef] [PubMed Central]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef] [PubMed Central]
- Wu, D.; Smyth, G.K. Camera: A competitive gene set test accounting for inter-gene correlation. Nucleic Acids Res. 2012, 40, e133. [Google Scholar] [CrossRef] [PubMed Central]
- Tarca, A.L.; Draghici, S.; Khatri, P.; Hassan, S.S.; Mittal, P.; Kim, J.S.; Kim, C.J.; Kusanovic, J.P.; Romero, R. A novel signaling pathway impact analysis. Bioinformatics 2009, 25, 75–82. [Google Scholar] [CrossRef] [PubMed Central]
- Rapier-Sharman, N.; Clancy, J.; Pickett, B.E. Joint Secondary Transcriptomic Analysis of Non-Hodgkin’s B-Cell Lymphomas Predicts Reliance on Pathways Associated with the Extracellular Matrix and Robust Diagnostic Biomarkers. J. Bioinform. Syst. Biol. 2022, 5, 119–135. [Google Scholar] [CrossRef] [PubMed Central]
- Scott, T.M.; Solis-Leal, A.; Lopez, J.B.; Robison, R.A.; Berges, B.K.; Pickett, B.E. Comparison of Intracellular Transcriptional Response of NHBE Cells to Infection with SARS-CoV-2 Washington and New York Strains. Front. Cell. Infect. Microbiol. 2022, 12, 1009328. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Dobbs Spendlove, M.; MGibson, T.; McCain, S.; Stone, B.C.; Gill, T.; Pickett, B.E. Pathway2Targets: An open-source pathway-based approach to repurpose therapeutic drugs and prioritize human targets. PeerJ 2023, 11, e16088. [Google Scholar] [CrossRef] [PubMed Central]
- Scott, T.M.; Jensen, S.; Pickett, B.E. A signaling pathway-driven bioinformatics pipeline for predicting therapeutics against emerging infectious diseases. F1000Research 2021, 10, 330. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- McHale, C.M.; Zhang, L.; Lan, Q.; Li, G.; Hubbard, A.E.; Forrest, M.S.; Vermeulen, R.; Chen, J.; Shen, M.; Rappaport, S.M.; et al. Changes in the peripheral blood transcriptome associated with occupational benzene exposure identified by cross-comparison on two microarray platforms. Genomics 2009, 93, 343–349. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Kuang, S.Q.; Bai, H.; Fang, Z.H.; Lopez, G.; Yang, H.; Tong, W.; Wang, Z.Z.; Garcia-Manero, G. Aberrant DNA methylation and epigenetic inactivation of Eph receptor tyrosine kinases and ephrin ligands in acute lymphoblastic leukemia. Blood 2010, 115, 2412–2419. [Google Scholar] [CrossRef] [PubMed Central]
- Phan, N.N.; Liu, S.; Wang, C.Y.; Hsu, H.P.; Lai, M.D.; Li, C.Y.; Chen, C.F.; Chiao, C.C.; Yen, M.C.; Sun, Z.; et al. Overexpressed gene signature of EPH receptor A/B family in cancer patients-comprehensive analyses from the public high-throughput database. Int. J. Clin. Exp. Pathol. 2020, 13, 1220–1242. [Google Scholar] [PubMed Central]
- Xia, J.; Wang, M.; Zhu, Y.; Bu, C.; Li, T. Differential mRNA and long noncoding RNA expression profiles in pediatric B-cell acute lymphoblastic leukemia patients. BMC Pediatr. 2022, 22, 10. [Google Scholar] [CrossRef] [PubMed Central]
- Figueroa, M.E.; Chen, S.C.; Andersson, A.K.; Phillips, L.A.; Li, Y.; Sotzen, J.; Kundu, M.; Downing, J.R.; Melnick, A.; Mullighan, C.G. Integrated genetic and epigenetic analysis of childhood acute lymphoblastic leukemia. J. Clin. Investig. 2013, 123, 3099–3111. [Google Scholar] [CrossRef] [PubMed Central]
- Yin, Z.; Liao, M.; Yan, R.; Li, G.; Ou, R.; Liu, Z.; Zhong, Q.; Shen, H.; Zhu, Y.; Xie, S.; et al. Transcriptome- and metabolome-based candidate mechanism of BCR-ABL-independent resistance to olverembatinib in Philadelphia chromosome-positive acute lymphoblastic leukemia. Funct. Integr. Genom. 2023, 23, 53. [Google Scholar] [CrossRef] [PubMed]
- Messina, M.; Chiaretti, S.; Tavolaro, S.; Peragine, N.; Vitale, A.; Elia, L.; Sica, S.; Levis, A.; Guarini, A.; Foà, R. Protein kinase gene expression profiling and in vitro functional experiments identify novel potential therapeutic targets in adult acute lymphoblastic leukemia. Cancer 2010, 116, 3426–3437. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.-L.; Tang, X.; Zhou, G.; Liu, S.; Wang, Y.; Chen, F.; Li, T.; Wen, F.; Liu, S.; Mai, H. PROM1 and CTGF Expression in Childhood -Rearrangement Acute Lymphoblastic Leukemia. J. Oncol. 2022, 2022, 5896022. [Google Scholar] [CrossRef] [PubMed Central]
- Zinngrebe, J.; Schlichtig, F.; Kraus, J.M.; Meyer, M.; Boldrin, E.; Kestler, H.A.; Meyer, L.H.; Fischer-Posovszky, P.; Debatin, K.M. Biomarker profile for prediction of response to SMAC mimetic monotherapy in pediatric precursor B-cell acute lymphoblastic leukemia. Int. J. Cancer 2020, 146, 3219–3231. [Google Scholar] [CrossRef] [PubMed]
- Petrov, I.; Suntsova, M.; Mutorova, O.; Sorokin, M.; Garazha, A.; Ilnitskaya, E.; Spirin, P.; Larin, S.; Kovalchuk, O.; Prassolov, V.; et al. Molecular pathway activation features of pediatric acute myeloid leukemia (AML) and acute lymphoblast leukemia (ALL) cells. Aging 2016, 8, 2936–2947. [Google Scholar] [CrossRef] [PubMed Central]
- Mura, M.; Swain, R.K.; Zhuang, X.; Vorschmitt, H.; Reynolds, G.; Durant, S.; Beesley, J.F.J.; Herbert, J.M.J.; Sheldon, H.; Andre, M.; et al. Identification and angiogenic role of the novel tumor endothelial marker CLEC14A. Oncogene 2012, 31, 293–305. [Google Scholar] [CrossRef] [PubMed]
- Kühnl, A.; Gökbuget, N.; Stroux, A.; Burmeister, T.; Neumann, M.; Heesch, S.; Haferlach, T.; Hoelzer, D.; Hofmann, W.K.; Thiel, E.; et al. High BAALC expression predicts chemoresistance in adult B-precursor acute lymphoblastic leukemia. Blood 2010, 115, 3737–3744. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Chang, T.Y.; Lan, K.C.; Chiu, C.Y.; Sheu, M.L.; Liu, S.H. ANGPTL1 attenuates cancer migration, invasion, and stemness through regulating FOXO3a-mediated SOX2 expression in colorectal cancer. Clin. Sci. 2022, 136, 657–673. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Kuo, T.C.; Tan, C.T.; Chang, Y.W.; Hong, C.C.; Lee, W.J.; Chen, M.W.; Jeng, Y.M.; Chiou, J.; Yu, P.; Chen, P.S.; et al. Angiopoietin-like protein 1 suppresses SLUG to inhibit cancer cell motility. J. Clin. Investig. 2013, 123, 1082–1095. [Google Scholar] [CrossRef] [PubMed Central]
- Chen, H.A.; Kuo, T.C.; Tseng, C.F.; Ma, J.T.; Yang, S.T.; Yen, C.J.; Yang, C.Y.; Sung, S.Y.; Su, J.L. Angiopoietin-like protein 1 antagonizes MET receptor activity to repress sorafenib resistance and cancer stemness in hepatocellular carcinoma. Hepatology 2016, 64, 1637–1651, Erratum in Hepatology 2017, 65, 760. [Google Scholar] [CrossRef] [PubMed]
- Bacher, U.; Schnittger, S.; Grüneisen, A.; Haferlach, T.; Kern, W.; Haferlach, C. Inverted duplication dup(1)(q32q21) as sole aberration in lymphoid and myeloid malignancies. Cancer Genet. Cytogenet. 2009, 188, 108–111. [Google Scholar] [CrossRef] [PubMed]
- Goud, T.M.; Al Salmani, K.K.; Al Harasi, S.M.; Al Musalhi, M.; Wasifuddin, S.M.; Rajab, A. Importance of FISH combined with Morphology, Immunophenotype and Cytogenetic Analysis of Childhood/ Adult Acute Lymphoblastic Leukemia in Omani Patients. Asian Pac. J. Cancer Prev. 2015, 16, 7343–7350. [Google Scholar] [CrossRef] [PubMed]
- Camper, L.; Heinegârd, D.; Lundgren-Akerlund, E. Integrin alpha2beta1 is a receptor for the cartilage matrix protein chondroadherin. J. Cell Biol. 1997, 138, 1159–1167. [Google Scholar] [CrossRef] [PubMed Central]
- Lee, B.M.; Buck-Koehntop, B.A.; Martinez-Yamout, M.A.; Dyson, H.J.; Wright, P.E. Embryonic neural inducing factor churchill is not a DNA-binding zinc finger protein: Solution structure reveals a solvent-exposed beta-sheet and zinc binuclear cluster. J. Mol. Biol. 2007, 371, 1274–1289. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Sousa, S.F.; Fernandes, P.A.; Ramos, M.J. Farnesyltransferase--new insights into the zinc-coordination sphere paradigm: Evidence for a carboxylate-shift mechanism. Biophys. J. 2005, 88, 483–494. [Google Scholar] [CrossRef] [PubMed Central]
- Hu, X.; Cao, P.; Wang, F.; Zhang, H.; Wang, T.; Liu, H.; Wu, X. Lengthening of the CHURC1 3′ untranslated region by alternative polyadenylation is associated with the progression of acute myeloid leukemia. Genes Dis. 2025, 12, 101248. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Liu, Z.; Zhou, J.; Wu, S.; Chen, Z.; Wu, S.; Chen, L.; Zhu, X.; Li, Z. Why Treg should be the focus of cancer immunotherapy: The latest thought. Biomed. Pharmacother. 2023, 168, 115142. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Martínez, J.L.; DeAndrés-Galiana, E.J.; Cernea, A. The Effect of NOP16 Mutation in Chronic Lymphocytic Leukemia. J. Mol. Genet. Med. 2017, 11, 1000295. Available online: https://www.omicsonline.org/open-access/the-effect-of-nop16-mutation-in-chronic-lymphocytic-leukemia-1747-0862-1000295-95280.html (accessed on 3 February 2025).
- Oliveira, T.; Zhang, M.; Joo, E.J.; Abdel-Azim, H.; Chen, C.W.; Yang, L.; Chou, C.H.; Qin, X.; Chen, J.; Alagesan, K.; et al. Glycoproteome remodeling in MLL-rearranged B-cell precursor acute lymphoblastic leukemia. Theranostics 2021, 11, 9519–9537. [Google Scholar] [CrossRef] [PubMed Central]
- Perez-Atayde, A.R.; Sallan, S.E.; Tedrow, U.; Connors, S.; Allred, E.; Folkman, J. Spectrum of tumor angiogenesis in the bone marrow of children with acute lymphoblastic leukemia. Am. J. Pathol. 1997, 150, 815–821. [Google Scholar] [PubMed Central]
- Ruan, Y.; Kim, H.N.; Ogana, H.A.; Gang, E.J.; Li, S.; Liu, H.C.; Bhojwani, D.; Wayne, A.S.; Yang, M.; Kim, Y.M. In vitro and in vivo effects of AVA4746, a novel competitive antagonist of the ligand binding of VLA-4, in B-cell acute lymphoblastic leukemia. Exp. Ther. Med. 2022, 23, 47. [Google Scholar] [CrossRef] [PubMed Central]
- Gang, E.J.; Kim, H.N.; Hsieh, Y.T.; Ruan, Y.; Ogana, H.A.; Lee, S.; Pham, J.; Geng, H.; Park, E.; Klemm, L.; et al. Integrin α6 mediates the drug resistance of acute lymphoblastic B-cell leukemia. Blood 2020, 136, 210–223. [Google Scholar] [CrossRef] [PubMed Central]
- Rapier-Sharman, N.; Kim, S.; Mudrow, M.; Told, M.T.; Fischer, L.; Fawson, L.; Parry, J.; Poole, B.D.; O’Neill, K.L.; Piccolo, S.R.; et al. Comparison of B-Cell Lupus and Lymphoma Using a Novel Immune Imbalance Transcriptomics Algorithm Reveals Potential Therapeutic Targets. Genes 2024, 15, 1215. [Google Scholar] [CrossRef] [PubMed Central]
- Yu, X.; Chen, Y.; Chen, J.; Fan, Y.; Lu, H.; Wu, D.; Xu, Y. Shared genetic architecture between autoimmune disorders and B-cell acute lymphoblastic leukemia: Insights from large-scale genome-wide cross-trait analysis. BMC Med. 2024, 22, 161. [Google Scholar] [CrossRef] [PubMed Central]
- Deng, W.; Xu, Y.; Yuan, X. Clinical features and prognosis of acute lymphoblastic leukemia in children with Epstein-Barr virus infection. Transl. Pediatr. 2022, 11, 642–650. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Cairo, M.S. EBV: The virus that keeps on giving! Blood 2023, 141, 689–691. [Google Scholar] [CrossRef] [PubMed]
- Gallardo-Zapata, J.; Pérez-Figueroa, E.; Olivar-López, V.; Medina-Sansón, A.; Jiménez-Hernández, E.; Ortega, E.; Maldonado-Bernal, C. TLR Agonists Modify NK Cell Activation and Increase Its Cytotoxicity in Acute Lymphoblastic Leukemia. Int. J. Mol. Sci. 2024, 25, 7500. [Google Scholar] [CrossRef] [PubMed Central]
- Bai, L.; Zhou, L.; Han, W.; Chen, J.; Gu, X.; Hu, Z.; Yang, Y.; Li, W.; Zhang, X.; Niu, C.; et al. BAX as the mediator of C-MYC sensitizes acute lymphoblastic leukemia to TLR9 agonists. J. Transl. Med. 2023, 21, 108. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Sánchez-Herrera, D.; Contreras-Ramos, A.; Jiménez-Hernández, E.; Medina-Sansón, A.; Giono-Cerezo, S.; Maldonado-Bernal, C. NK Cell Subpopulation Is Altered and the Expression of TLR1 and TLR9 Is Decreased in Patients with Acute Lymphoblastic Leukemia. J. Oncol. 2021, 2021, 5528378. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Canevarolo, R.R.; Cury, N.M.; Yunes, J.A. The Expression and Activation of the NF-κB Pathway Correlate with Methotrexate Resistance and Cell Proliferation in Acute Lymphoblastic Leukemia. Genes 2023, 14, 1880. [Google Scholar] [CrossRef] [PubMed Central]
- Dai, Q.; Zhang, G.; Wang, Y.; Ye, L.; Shi, R.; Peng, L.; Guo, S.; He, J.; Yang, H.; Zhang, Y.; et al. Cytokine network imbalance in children with B-cell acute lymphoblastic leukemia at diagnosis. Cytokine 2023, 169, 156267. [Google Scholar] [CrossRef] [PubMed]
- Lejman, M.; Kuśmierczuk, K.; Bednarz, K.; Ostapińska, K.; Zawitkowska, J. Targeted Therapy in the Treatment of Pediatric Acute Lymphoblastic Leukemia-Therapy and Toxicity Mechanisms. Int. J. Mol. Sci. 2021, 22, 9827. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Ivanov, A.V.; Alecsa, M.S.; Popescu, R.; Starcea, M.I.; Mocanu, A.M.; Rusu, C.; Miron, I.C. Pediatric Acute Lymphoblastic Leukemia Emerging Therapies-From Pathway to Target. Int. J. Mol. Sci. 2023, 24, 4661. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Jędraszek, K.; Malczewska, M.; Parysek-Wójcik, K.; Lejman, M. Resistance Mechanisms in Pediatric B-Cell Acute Lymphoblastic Leukemia. Int. J. Mol. Sci. 2022, 23, 3067. [Google Scholar] [CrossRef] [PubMed Central]
- Dander, E.; Palmi, C.; D’Amico, G.; Cazzaniga, G. The Bone Marrow Niche in B-Cell Acute Lymphoblastic Leukemia: The Role of Microenvironment from Pre-Leukemia to Overt Leukemia. Int. J. Mol. Sci. 2021, 22, 4426. [Google Scholar] [CrossRef] [PubMed Central]
- Kugler, E. Charting a path through resistance: Histone deacetylase inhibitors for TP53-mutated B-cell acute lymphoblastic leukemia. Haematologica 2024, 109, 1643–1645. [Google Scholar] [CrossRef] [PubMed Central]
- Fidyt, K.; Pastorczak, A.; Cyran, J.; Crump, N.T.; Goral, A.; Madzio, J.; Muchowicz, A.; Poprzeczko, M.; Domka, K.; Komorowski, L.; et al. Potent, p53-independent induction of NOXA sensitizes MLL-rearranged B-cell acute lymphoblastic leukemia cells to venetoclax. Oncogene 2022, 41, 1600–1609. [Google Scholar] [CrossRef] [PubMed Central]
- Grüninger, P.K.; Uhl, F.; Herzog, H.; Gentile, G.; Andrade-Martinez, M.; Schmidt, T.; Han, K.; Morgens, D.W.; Bassik, M.C.; Cleary, M.L.; et al. Functional characterization of the PI3K/AKT/MTOR signaling pathway for targeted therapy in B-precursor acute lymphoblastic leukemia. Cancer Gene Ther. 2022, 29, 1751–1760. [Google Scholar] [CrossRef] [PubMed Central]
- Hohtari, H.; Kankainen, M.; Adnan-Awad, S.; Yadav, B.; Potdar, S.; Ianevski, A.; Dufva, O.; Heckman, C.; Sexl, V.; Kytölä, S.; et al. Targeting Apoptosis Pathways with BCL2 and MDM2 Inhibitors in Adult B-cell Acute Lymphoblastic Leukemia. Hemasphere 2022, 6, e701. [Google Scholar] [CrossRef] [PubMed Central]
- Narang, S.; Ghebrechristos, Y.; Evensen, N.A.; Murrell, N.; Jasinski, S.; Ostrow, T.H.; Teachey, D.T.; Raetz, E.A.; Lionnet, T.; Witkowski, M.; et al. Clonal evolution of the 3D chromatin landscape in patients with relapsed pediatric B-cell acute lymphoblastic leukemia. Nat. Commun. 2024, 15, 7425. [Google Scholar] [CrossRef] [PubMed Central]
- Isobe, T.; Takagi, M.; Sato-Otsubo, A.; Nishimura, A.; Nagae, G.; Yamagishi, C.; Tamura, M.; Tanaka, Y.; Asada, S.; Takeda, R.; et al. Multi-omics analysis defines highly refractory RAS burdened immature subgroup of infant acute lymphoblastic leukemia. Nat. Commun. 2022, 13, 4501. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Khanolkar, A.; Liu, G.; Simpson Schneider, B.M. Defining the Basal and Immunomodulatory Mediator-Induced Phosphoprotein Signature in Pediatric B Cell Acute Lymphoblastic Leukemia (B-ALL) Diagnostic Samples. Int. J. Mol. Sci. 2023, 24, 13937. [Google Scholar] [CrossRef] [PubMed Central]
- Bellaye, P.S.; Georgievski, A.; Ballerini, P.; Bouslama, B.; Richard, C.; Boidot, R.; Chevreux, G.; Legros, V.; Guy, J.; Racine, J.; et al. CD40 Agonist on Patient-Derived Xenograft Mice for the Treatment of B-Cell Acute Lymphoblastic Leukemia. Clin. Cancer Res. 2025, 31, 181–196. [Google Scholar] [CrossRef] [PubMed]
- Rios de Los Rios, J.; Enciso, J.; Vilchis-Ordoñez, A.; Vázquez-Ramírez, R.; Ramirez-Ramirez, D.; Balandrán, J.C.; Rodríguez-Martínez, A.; Ruiz-Tachiquín, M.; Pompa-Mera, E.; Mendoza, L.; et al. Acute lymphoblastic leukemia-secreted miRNAs induce a proinflammatory microenvironment and promote the activation of hematopoietic progenitors. J. Leukoc. Biol. 2022, 112, 31–45. [Google Scholar] [CrossRef] [PubMed]
- Fu, Z.; Li, S.; Han, S.; Shi, C.; Zhang, Y. Antibody drug conjugate: The “biological missile” for targeted cancer therapy. Signal Transduct. Target. Ther. 2022, 7, 93. [Google Scholar] [CrossRef] [PubMed Central]
- Qi, H.; Chi, L.; Wang, X.; Jin, X.; Wang, W.; Lan, J. Identification of a Seven-lncRNA-mRNA Signature for Recurrence and Prognostic Prediction in Relapsed Acute Lymphoblastic Leukemia Based on WGCNA and LASSO Analyses. Anal. Cell. Pathol. 2021, 2021, 6692022. [Google Scholar] [CrossRef] [PubMed Central]
- Ohtsuka, T.; Shiomi, T.; Shimoda, M.; Kodama, T.; Amour, A.; Murphy, G.; Ohuchi, E.; Kobayashi, K.; Okada, Y. ADAM28 is overexpressed in human non-small cell lung carcinomas and correlates with cell proliferation and lymph node metastasis. Int. J. Cancer 2006, 118, 263–273. [Google Scholar] [CrossRef] [PubMed]
- Wei, L.; Wen, J.-Y.; Chen, J.; Ma, X.-K.; Wu, D.-H.; Chen, Z.-H.; Huang, J.-L. Oncogenic ADAM28 induces gemcitabine resistance and predicts a poor prognosis in pancreatic cancer. World J. Gastroenterol. 2019, 25, 5590–5603. [Google Scholar] [CrossRef] [PubMed Central]
- Yin, Q.; Gu, J.; Qi, Y.; Lu, Y.; Yang, L.; Liu, J.; Liang, X. ADAM28 from both endothelium and gastric cancer cleaves von Willebrand Factor to eliminate von Willebrand Factor-induced apoptosis of gastric cancer cells. Eur. J. Pharmacol. 2021, 898, 173994. [Google Scholar] [CrossRef] [PubMed]
- Gérard, C.; Hubeau, C.; Carnet, O.; Bellefroid, M.; Sounni, N.E.; Blacher, S.; Bendavid, G.; Moser, M.; Fässler, R.; Noel, A.; et al. Microenvironment-derived ADAM28 prevents cancer dissemination. Oncotarget 2018, 9, 37185–37199. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Wang, J.; Li, H.; Wang, Y.; Wang, L.; Yan, X.; Zhang, D.; Ma, X.; Du, Y.; Liu, X.; Yang, Y. MicroRNA-552 enhances metastatic capacity of colorectal cancer cells by targeting a disintegrin and metalloprotease 28. Oncotarget 2016, 7, 70194–70210. [Google Scholar] [CrossRef] [PubMed Central]
- Zhang, J.-M.; Wang, C.-C.; Zhang, G.-C.; Jiang, Q.; Yang, S.-M.; Fu, H.-X.; Wang, Q.-M.; Zhu, X.-L.; Zhu, H.-H.; Jiang, H.; et al. ADAM28 promotes tumor growth and dissemination of acute myeloid leukemia through IGFBP-3 degradation and IGF-I-induced cell proliferation. Cancer Lett. 2019, 442, 193–201. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.-H.; Wang, C.-C.; Jiang, Q.; Yang, S.-M.; Jiang, H.; Lu, J.; Wang, Q.-M.; Feng, F.-E.; Zhu, X.-L.; Zhao, T.; et al. ADAM28 overexpression regulated via the PI3K/Akt pathway is associated with relapse in de novo adult B-cell acute lymphoblastic leukemia. Leuk. Res. 2015, 39, 1229–1238. [Google Scholar] [CrossRef] [PubMed]
| Gene Symbol | Gene Name | log2FC 1 | FDR 2 |
|---|---|---|---|
| HIST2H2AA3 | histone cluster 2 H2A family member a3 | 15.4 | 1.20 × 10−31 |
| EPHA7 | EPH receptor A7 | 13 | 7.63 × 10−22 |
| NPR1 | natriuretic peptide receptor 1 | 12.9 | 3.69 × 10−26 |
| DIPK1C | divergent protein kinase domain 1C | 12.8 | 4.31 × 10−22 |
| HIST1H3B | histone cluster 1 H3 family member b | 12.6 | 1.49 × 10−85 |
| CLEC14A | C-type lectin domain containing 14A | 12.6 | 1.28 × 10−20 |
| S100A16 | S100 calcium binding protein A16 | 12.4 | 3.06 × 10−18 |
| OVCH2 | ovochymase 2 (gene/pseudogene) | 12 | 2.14 × 10−17 |
| ERG | ETS transcription factor ERG | 11.8 | 1.16 × 100−67 |
| GREM1 | gremlin 1, DAN family BMP antagonist | 11.4 | 9.29 × 100−10 |
| Gene Symbol | Gene Name | log2FC 1 | FDR 2 |
|---|---|---|---|
| IGHA1 | immunoglobulin heavy constant alpha 1 | −9.56 | 9.11 × 10−91 |
| ANGPTL1 | angiopoietin like 1 | −9.06 | 2.38 × 10−94 |
| IGKV1-39 | immunoglobulin kappa variable 1–39 (gene/pseudogene) | −8.48 | 2.43 × 10−65 |
| GLYATL1B | glycine-N-acyltransferase like 1B | −8.22 | 4.93 × 10−109 |
| CHAD | chondroadherin | −8.22 | 1.48 × 10−87 |
| CHURC1-FNTB | CHURC1-FNTB readthrough | −8.2 | 3.11 × 10−59 |
| IGKV3D-11 | immunoglobulin kappa variable 3D-11 | −8.08 | 3.67 × 10−75 |
| AC011511.4 | novel transcript | −7.94 | 6.39 × 10−72 |
| IGKV2D-28 | immunoglobulin kappa variable 2D-28 | −7.87 | 2.33 × 10−73 |
| ABHD16B | abhydrolase domain containing 16B | −7.73 | 1.25 × 10−102 |
| Name | Number of Proteins in Pathway | Number of DEGs in Pathway | FDR p-Value | Pathway Regulation Status | Source Database |
|---|---|---|---|---|---|
| Integrins in angiogenesis | 54 | 40 | 8.565 × 10−3 | Activated | NCI |
| Antigen processing and presentation | 63 | 55 | 7.797 × 10−4 | Inhibited | KEGG |
| Calcium signaling pathway | 134 | 102 | 3.862 × 10−3 | Activated | KEGG |
| Systemic lupus erythematosus | 119 | 109 | 2.521 × 10−2 | Activated | KEGG |
| NF-kappa B signaling pathway | 85 | 76 | 4.272 × 10−2 | Inhibited | KEGG |
| Toll-like receptor signaling pathway | 85 | 76 | 4.354 × 10−2 | Inhibited | KEGG |
| Tight junction | 105 | 97 | 4.354 × 10−2 | Activated | KEGG |
| Epstein–Barr virus infection | 195 | 159 | 4.449 × 10−2 | Inhibited | KEGG |
| Target Symbol | Target Name | Subcellular Location | Weighted Score |
|---|---|---|---|
| TNF | tumor necrosis factor | Cell membrane | 3870 |
| IL6 | interleukin 6 | Secreted | 3287.5 |
| TP53 | tumor protein p53 | Cytoplasm | 3177 |
| EGFR | epidermal growth factor receptor | Cell membrane | 2562 |
| CD40LG | CD40 ligand | Cell membrane | 2490 |
| AKT1 | AKT serine/threonine kinase 1 | Cytoplasm | 2323.5 |
| PIK3CA | phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha | Cytosol | 2301 |
| PTGS2 | prostaglandin-endoperoxide synthase 2 | Microsome membrane | 2288 |
| PIK3CB | phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit beta | Cytoplasm | 2228 |
| PIK3CD | phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit delta | Cytoplasm | 2142 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wilkins, M.R.K.; Pickett, B.E. Histone-, Receptor-, and Integrin-Related Gene Products and ADAM28 as Relevant to B-Cell Acute Lymphoblastic Leukemia (B-ALL). Curr. Issues Mol. Biol. 2025, 47, 699. https://doi.org/10.3390/cimb47090699
Wilkins MRK, Pickett BE. Histone-, Receptor-, and Integrin-Related Gene Products and ADAM28 as Relevant to B-Cell Acute Lymphoblastic Leukemia (B-ALL). Current Issues in Molecular Biology. 2025; 47(9):699. https://doi.org/10.3390/cimb47090699
Chicago/Turabian StyleWilkins, Makayla R. K., and Brett E. Pickett. 2025. "Histone-, Receptor-, and Integrin-Related Gene Products and ADAM28 as Relevant to B-Cell Acute Lymphoblastic Leukemia (B-ALL)" Current Issues in Molecular Biology 47, no. 9: 699. https://doi.org/10.3390/cimb47090699
APA StyleWilkins, M. R. K., & Pickett, B. E. (2025). Histone-, Receptor-, and Integrin-Related Gene Products and ADAM28 as Relevant to B-Cell Acute Lymphoblastic Leukemia (B-ALL). Current Issues in Molecular Biology, 47(9), 699. https://doi.org/10.3390/cimb47090699







