Convergent and Divergent Mitochondrial Pathways as Causal Drivers and Therapeutic Targets in Neurological Disorders
Abstract
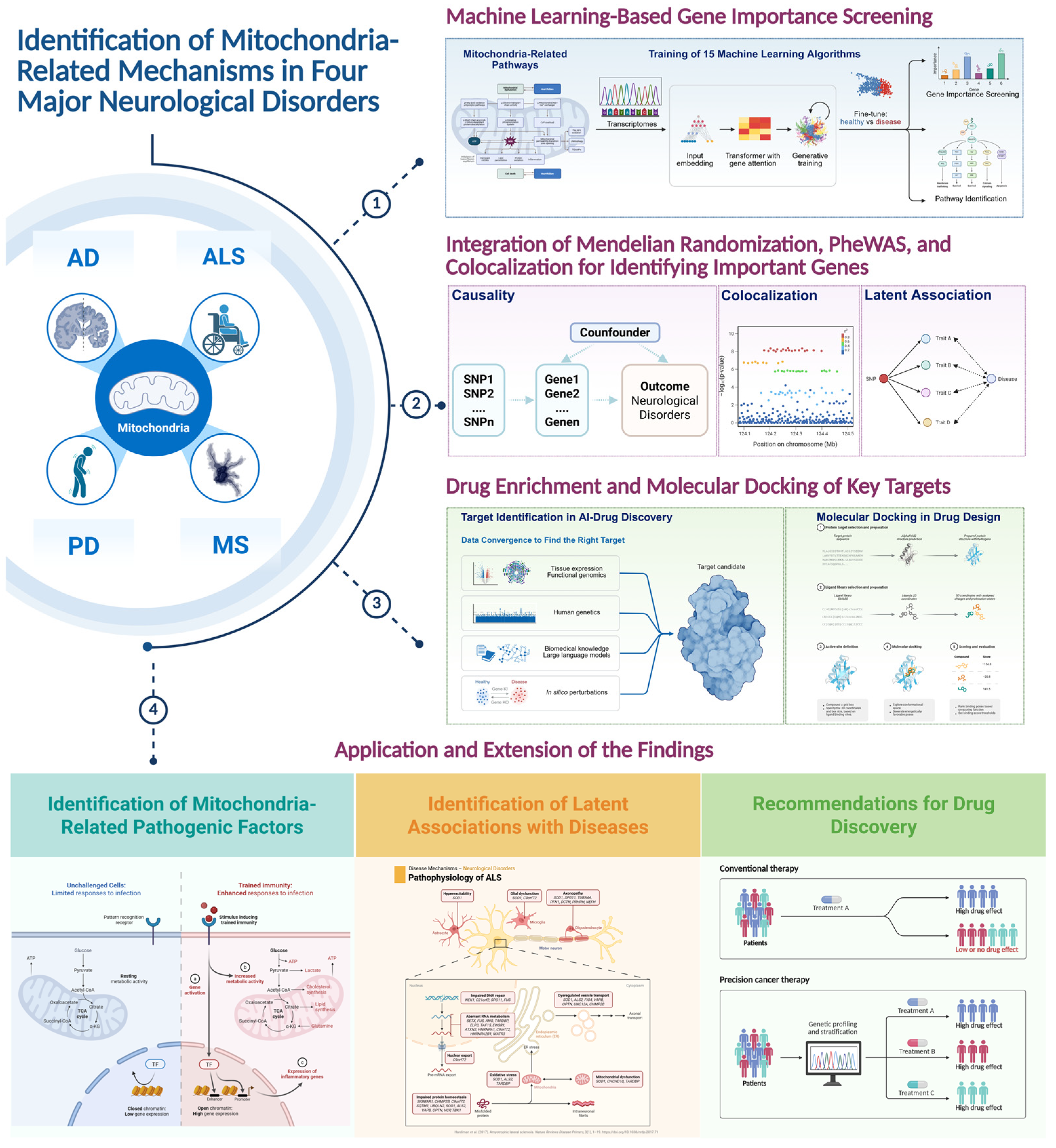
1. Introduction
2. Materials and Methods
2.1. Study Participants and Data Source
2.2. Mitochondrial Pathway-Based Gene Expression Modeling and Evaluation
- Logistic Regression: A linear model for binary classification, optimized by tuning the regularization parameter to control overfitting. Performance was evaluated using the area under the ROC curve (ROC-AUC).
- Support Vector Machines (SVM): A powerful classification algorithm that maximizes the margin between classes, optimized by tuning the regularization parameter C.
- Naive Bayes: A probabilistic classifier assuming conditional independence between features, applied here using the Gaussian Naive Bayes model based on the normal distribution of gene expression data.
- Decision Tree: A non-linear model that partitions feature space based on feature values, with the tree’s depth constrained to prevent overfitting.
- Random Forest: An ensemble method that aggregates predictions from multiple decision trees trained on bootstrapped data subsets to improve generalization.
- Gradient Boosting: A sequential ensemble method that builds models to correct the errors of previous ones, optimized by tuning the number of estimators to prevent overfitting.
- XGBoost: An optimized gradient boosting algorithm incorporating techniques like regularization and tree pruning, tuned for the number of estimators.
- CatBoost: A gradient boosting algorithm particularly effective for categorical features, optimized by adjusting tree depth.
- AdaBoost: An ensemble method that adjusts the weights of misclassified instances, with the number of estimators tuned for optimal performance.
- SGDClassifier: A model based on stochastic gradient descent for optimizing convex loss functions, applied with a log-loss function and regularization parameter tuning.
- K-Nearest Neighbors (KNN): A non-parametric method based on the majority class of nearest neighbors, with the number of neighbors optimized for performance.
- Multi-Layer Perceptron (MLP): A deep learning model with multiple layers of neurons, optimized with a single hidden layer of 100 neurons and regularization to prevent overfitting.
- FTTransformer: A deep learning model based on the Transformer architecture, designed to capture long-range dependencies in sequential data, optimized by adjusting hidden layer architecture.
- ExtraTreesClassifier: An ensemble method that builds multiple decision trees with random feature splits, aimed at increasing model robustness and avoiding overfitting.
- Quadratic Discriminant Analysis (QDA): A generative model assuming Gaussian distributions for each class, used to assess the performance of probabilistic models with quadratic decision boundaries.
2.3. Mendelian Randomization Analysis to Investigate Causal Relationships Between Mitochondrial Pathway-Associated Genes and Psychiatric Disorders
2.4. Colocalization Analysis of Mitochondrial Pathway-Associated Genes from MR Results with Risk Loci for Neurological Diseases
- P_0: No association with either the mitochondrial pathway gene or neurodegenerative disease.
- P_1: Association with the mitochondrial pathway gene only.
- P_2: Association with neurodegenerative disease only.
- P_3: Both traits are associated but driven by distinct causal variants.
- P_4: Both traits share a common causal variant (strong evidence for colocalization).
2.5. PheWAS and Bidirectional Mendelian Randomization Analysis of Colocalized SNPs with Neurological Diseases
2.6. Molecular Docking of Drug Candidates Genes
2.7. Statistical Analysis
3. Results
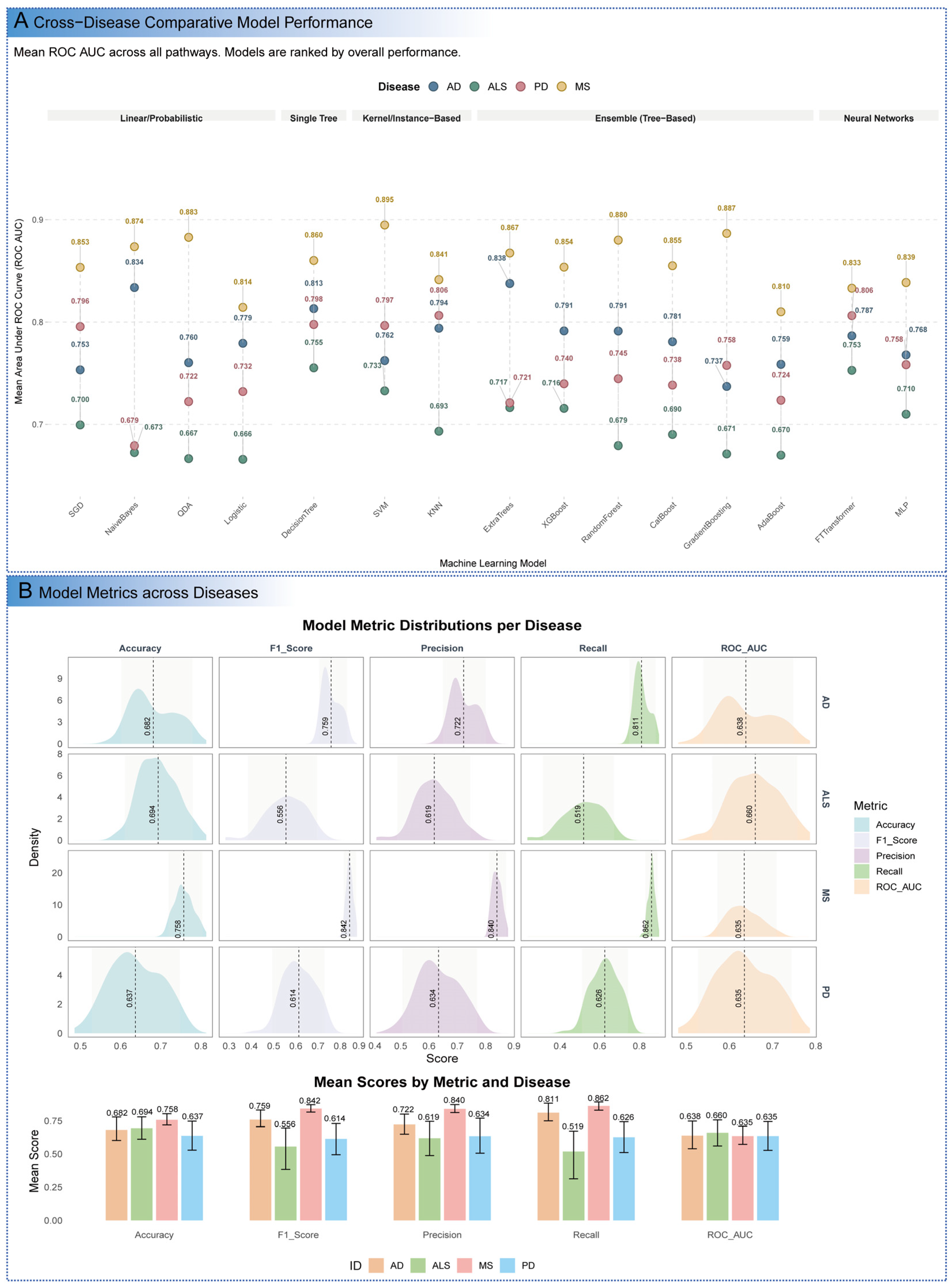
3.1. Machine Learning Model Performance for Disease Classification
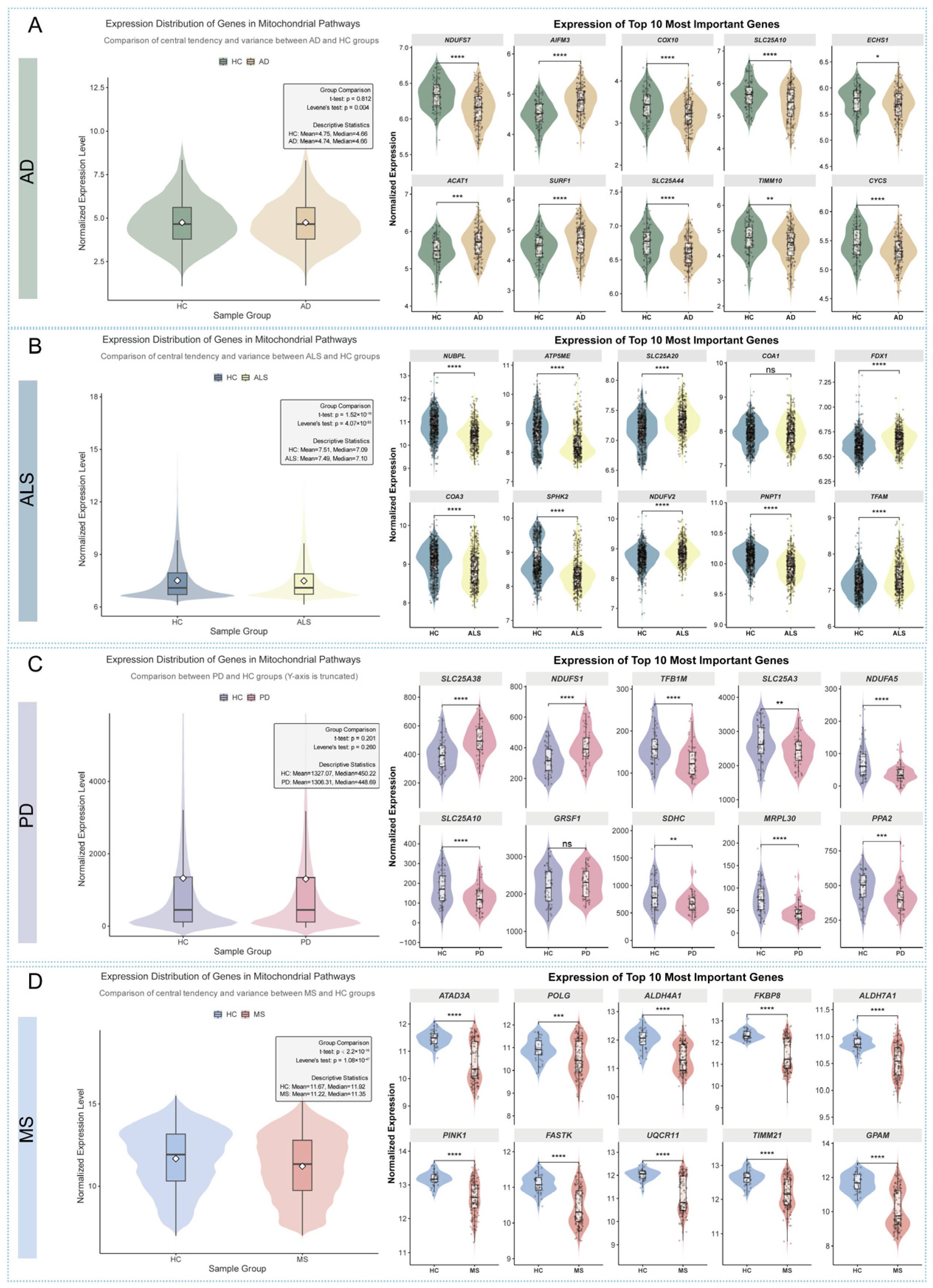
3.2. Mitochondrial Gene Expression Profiles Reveal a Uniquely Distinct Signature in MS
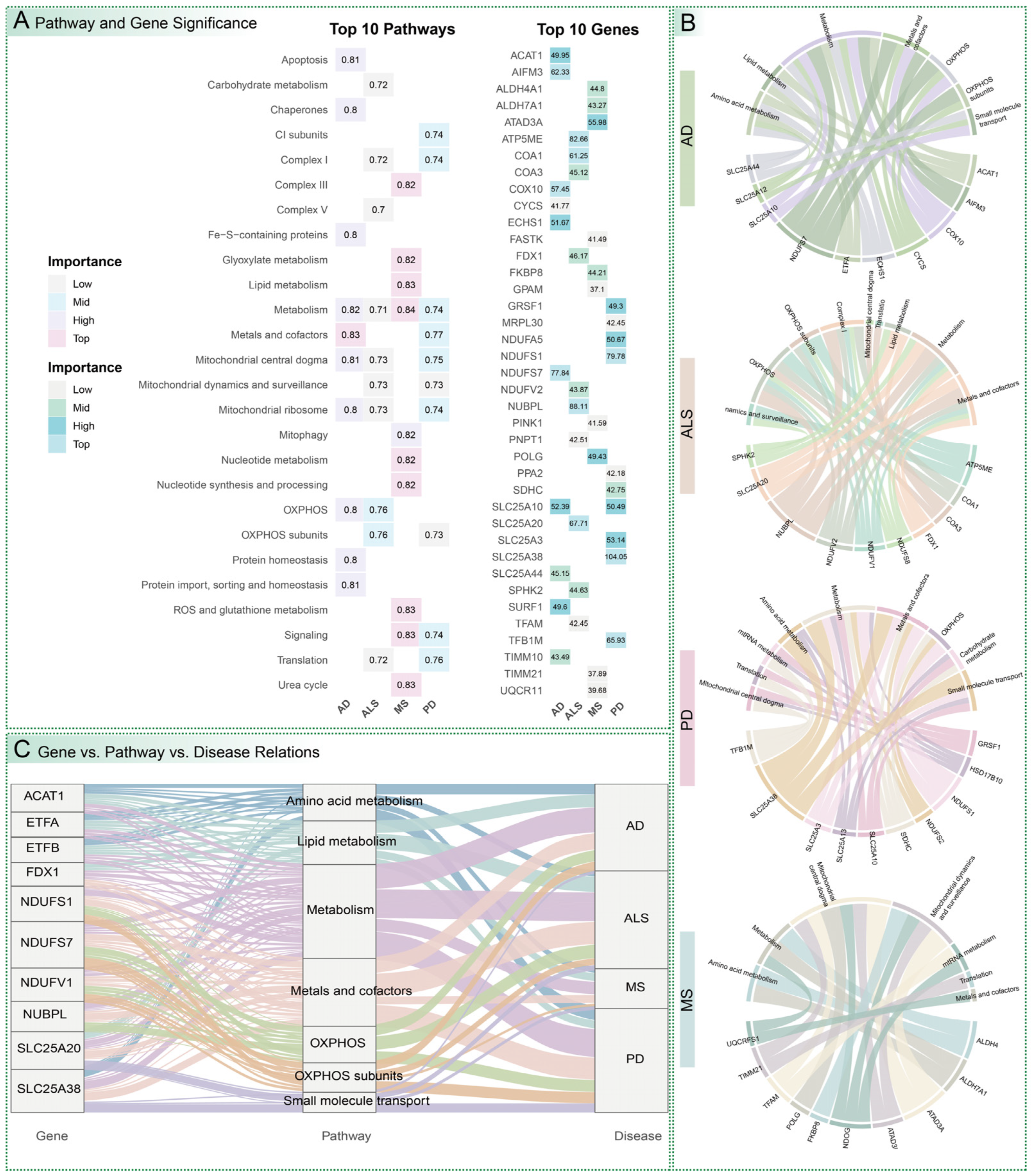
3.3. Identification of Key Mitochondrial Pathways and Genes
3.4. Gene–Pathway Interactions and Cross-Disease Comparisons
3.5. Mendelian Randomization Analysis of Causal Gene Associations
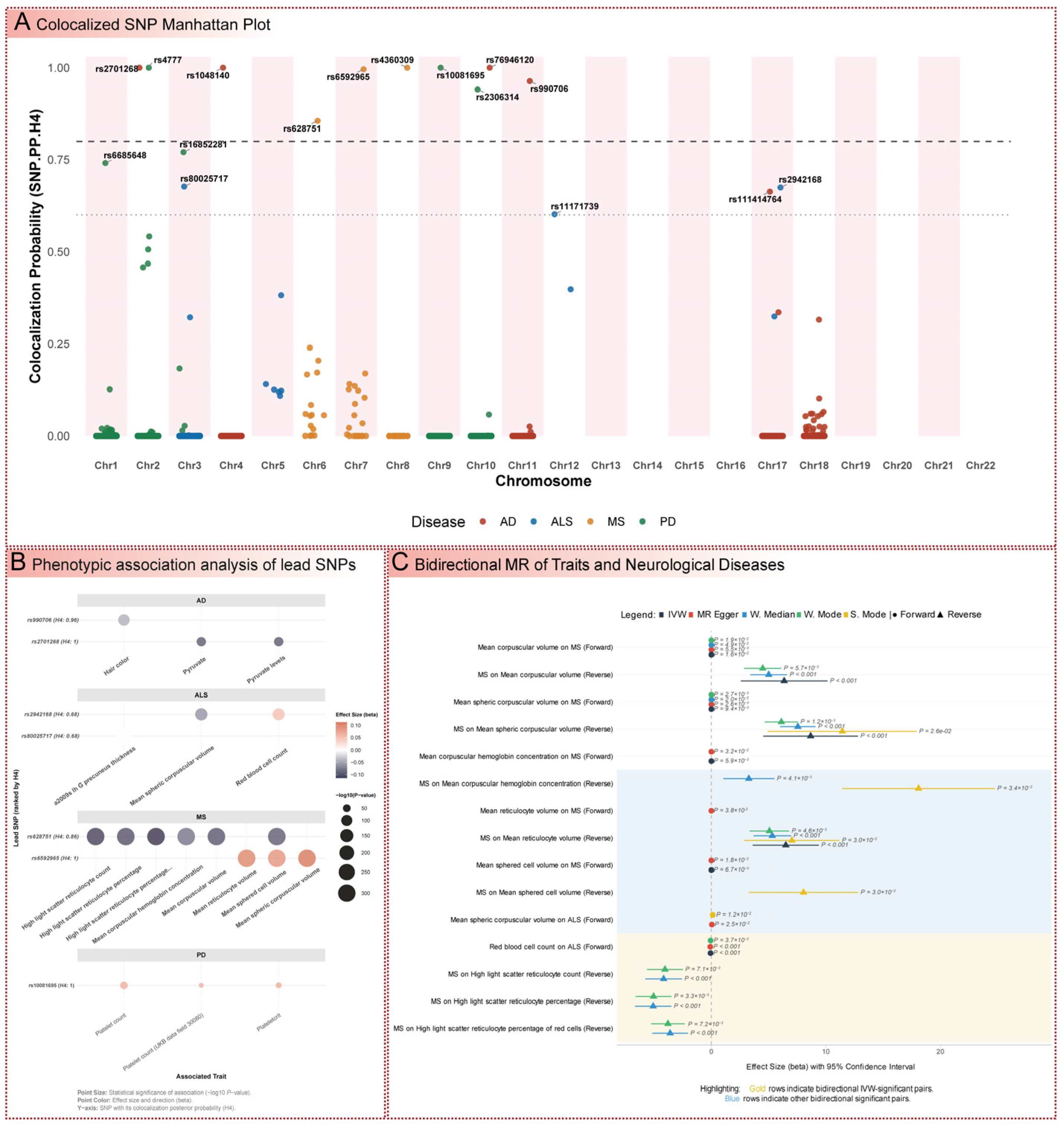
3.6. Colocalization Analysis of eQTLs and Disease Risk Variants
3.7. Phenome-Wide Association Study of Colocalized Variants
3.8. Bidirectional Mendelian Randomization of Phenotypic Traits
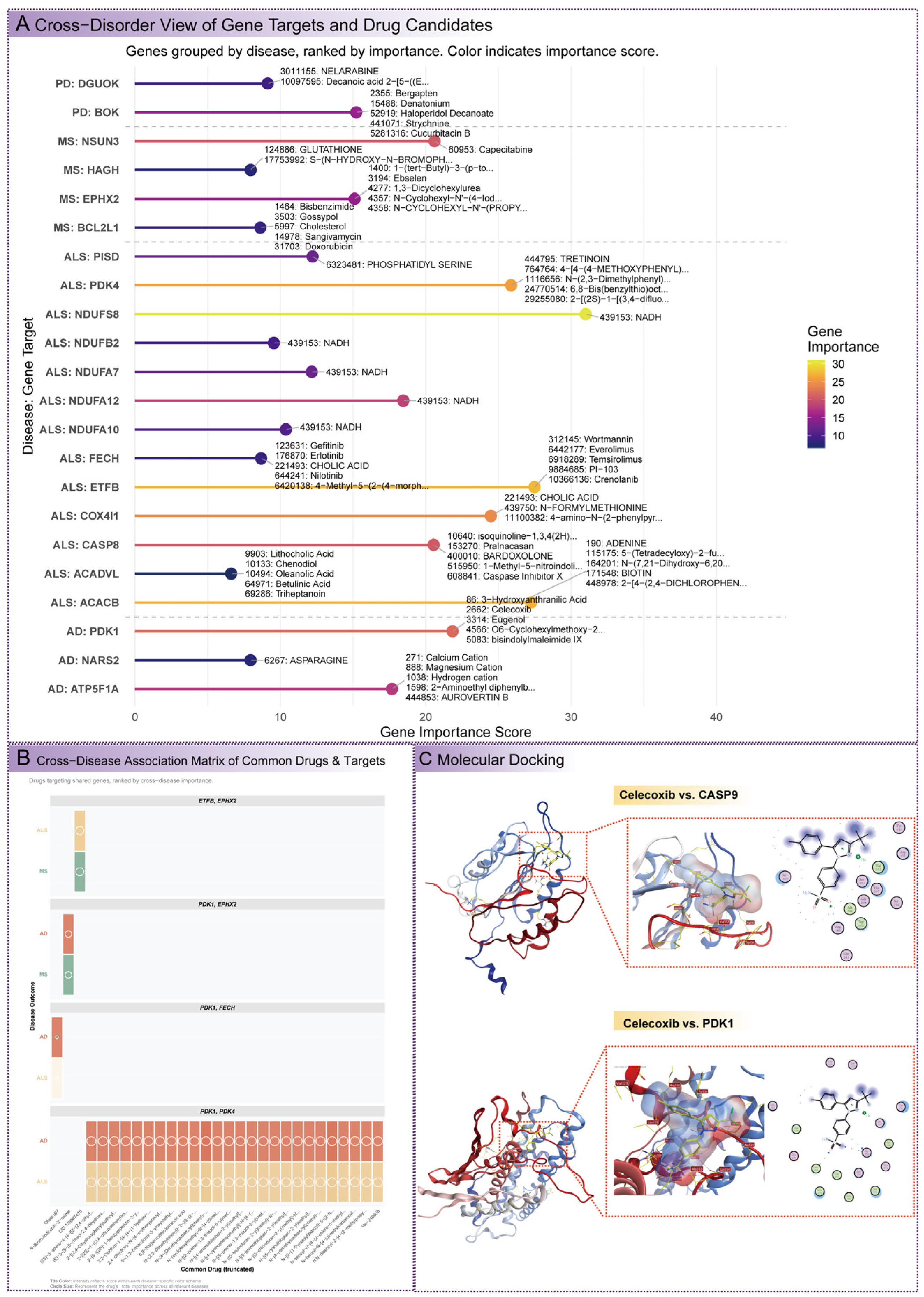
3.9. Drug Enrichment Analysis and Therapeutic Implications
3.10. Molecular Docking Analysis of Celecoxib with Multi-Disease Targets
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AD | Alzheimer’s Disease |
| ALS | ALS Amyotrophic Lateral Sclerosis |
| AUC | Area Under the Curve |
| COLOC | Colocalization |
| eQTL | Expression Quantitative Trait Locus |
| GWAS | Genome-Wide Association Study |
| IV | Instrumental Variable |
| IVW | Inverse Variance Weighted |
| KNN | K-Nearest Neighbors |
| LD | Linkage Disequilibrium |
| LASSO | Least Absolute Shrinkage and Selection Operator |
| MLP | Multi-Layer Perceptron |
| PD | Parkinson’s Disease |
| PheWAS | Phenome-Wide Association Study |
| ROC-AUC | Receiver Operating Characteristic—Area Under the Curve |
| SNP | Single Nucleotide Polymorphism |
| SVM | Support Vector Machine |
| TCA | Tricarboxylic Acid |
References
- Ashleigh, T.; Swerdlow, R.H.; Beal, M.F. The Role of Mitochondrial Dysfunction in Alzheimer’s Disease Pathogenesis. Alzheimers Dement. J. Alzheimers Assoc. 2023, 19, 333–342. [Google Scholar] [CrossRef]
- Chen, W.; Zhao, H.; Li, Y. Mitochondrial Dynamics in Health and Disease: Mechanisms and Potential Targets. Signal Transduct. Target. Ther. 2023, 8, 333. [Google Scholar] [CrossRef] [PubMed]
- Johri, A.; Beal, M.F. Mitochondrial Dysfunction in Neurodegenerative Diseases. J. Pharmacol. Exp. Ther. 2012, 342, 619–630. [Google Scholar] [CrossRef]
- Collier, J.J.; Oláhová, M.; McWilliams, T.G.; Taylor, R.W. Mitochondrial Signalling and Homeostasis: From Cell Biology to Neurological Disease. Trends Neurosci. 2023, 46, 137–152. [Google Scholar] [CrossRef] [PubMed]
- Clemente-Suárez, V.; Redondo-Flórez, L.; Beltrán-Velasco, A.; Ramos-Campo, D.; Belinchón-deMiguel, P.; Martinez-Guardado, I.; Dalamitros, A.; Yáñez-Sepúlveda, R.; Martín-Rodríguez, A.; Tornero-Aguilera, J. Mitochondria and Brain Disease: A Comprehensive Review of Pathological Mechanisms and Therapeutic Opportunities. Biomedicines 2023, 11, 2488. [Google Scholar] [CrossRef]
- Johnson, J.; Mercado-Ayon, E.; Mercado-Ayon, Y.; Dong, Y.N.; Halawani, S.; Ngaba, L.; Lynch, D.R. Mitochondrial Dysfunction in the Development and Progression of Neurodegenerative Diseases. Arch. Biochem. Biophys. 2021, 702, 108698. [Google Scholar] [CrossRef]
- Lucchesi, M.; Biso, L.; Bonaso, M.; Longoni, B.; Buchignani, B.; Battini, R.; Santorelli, F.M.; Doccini, S.; Scarselli, M. Mitochondrial Dysfunction in Genetic and Non-Genetic Parkinson’s Disease. Int. J. Mol. Sci. 2025, 26, 4451. [Google Scholar] [CrossRef]
- Elfawy, H.A.; Das, B. Crosstalk between Mitochondrial Dysfunction, Oxidative Stress, and Age Related Neurodegenerative Disease: Etiologies and Therapeutic Strategies. Life Sci. 2019, 218, 165–184. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.; Wang, L.; Liu, J.; Xie, F.; Su, B.; Wang, X. Abnormalities of Mitochondrial Dynamics in Neurodegenerative Diseases. Antioxidants 2017, 6, 25. [Google Scholar] [CrossRef]
- Kathiresan, D.S.; Balasubramani, R.; Marudhachalam, K.; Jaiswal, P.; Ramesh, N.; Sureshbabu, S.G.; Puthamohan, V.M.; Vijayan, M. Role of Mitochondrial Dysfunctions in Neurodegenerative Disorders: Advances in Mitochondrial Biology. Mol. Neurobiol. 2025, 62, 6827–6855. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Granado, J.; Piñero, J.; Furlong, L.I. Benchmarking Post-GWAS Analysis Tools in Major Depression: Challenges and Implications. Front. Genet. 2022, 13, 1006903. [Google Scholar] [CrossRef] [PubMed]
- Gallagher, M.D.; Chen-Plotkin, A.S. The Post-GWAS Era: From Association to Function. Am. J. Hum. Genet. 2018, 102, 717–730. [Google Scholar] [CrossRef] [PubMed]
- Hardiman, O.; Al-Chalabi, A.; Chio, A.; Corr, E.M.; Logroscino, G.; Robberecht, W.; Shaw, P.J.; van den Berg, L.H. Amyotrophic lateral sclerosis. Nat. Rev. Dis. Primers. 2017, 3, 17071. [Google Scholar] [CrossRef] [PubMed]
- Bellenguez, C.; Küçükali, F.; Jansen, I.E.; Kleineidam, L.; Moreno-Grau, S.; Amin, N.; Naj, A.C.; Campos-Martin, R.; Grenier-Boley, B.; Andrade, V.; et al. New Insights into the Genetic Etiology of Alzheimer’s Disease and Related Dementias. Nat. Genet. 2022, 54, 412–436. [Google Scholar] [CrossRef]
- Nicolas, A.; Kenna, K.P.; Renton, A.E.; Ticozzi, N.; Faghri, F.; Chia, R.; Dominov, J.A.; Kenna, B.J.; Nalls, M.A.; Keagle, P.; et al. Genome-Wide Analyses Identify KIF5A as a Novel ALS Gene. Neuron 2018, 97, 1267–1288. [Google Scholar] [CrossRef]
- Sudlow, C.; Gallacher, J.; Allen, N.; Beral, V.; Burton, P.; Danesh, J.; Downey, P.; Elliott, P.; Green, J.; Landray, M.; et al. UK Biobank: An Open Access Resource for Identifying the Causes of a Wide Range of Complex Diseases of Middle and Old Age. PLoS Med. 2015, 12, e1001779. [Google Scholar] [CrossRef]
- Patel, H.; Hodges, A.K.; Curtis, C.; Lee, S.H.; Troakes, C.; Dobson, R.J.B.; Newhouse, S.J. Transcriptomic Analysis of Probable Asymptomatic and Symptomatic Alzheimer Brains. Brain. Behav. Immun. 2019, 80, 644–656. [Google Scholar] [CrossRef]
- van Rheenen, W.; Diekstra, F.P.; Harschnitz, O.; Westeneng, H.-J.; van Eijk, K.R.; Saris, C.G.J.; Groen, E.J.N.; van Es, M.A.; Blauw, H.M.; van Vught, P.W.J.; et al. Whole Blood Transcriptome Analysis in Amyotrophic Lateral Sclerosis: A Biomarker Study. PLoS ONE 2018, 13, e0198874. [Google Scholar] [CrossRef] [PubMed]
- Swindell, W.R.; Kruse, C.P.S.; List, E.O.; Berryman, D.E.; Kopchick, J.J. ALS Blood Expression Profiling Identifies New Biomarkers, Patient Subgroups, and Evidence for Neutrophilia and Hypoxia. J. Transl. Med. 2019, 17, 170. [Google Scholar] [CrossRef] [PubMed]
- Enz, L.S.; Zeis, T.; Schmid, D.; Geier, F.; van der Meer, F.; Steiner, G.; Certa, U.; Binder, T.M.C.; Stadelmann, C.; Martin, R.; et al. Increased HLA-DR Expression and Cortical Demyelination in MS Links with HLA-DR15. Neurol. Neuroimmunol. Neuroinflammation 2020, 7, e656. [Google Scholar] [CrossRef]
- Rath, S.; Sharma, R.; Gupta, R.; Ast, T.; Chan, C.; Durham, T.J.; Goodman, R.P.; Grabarek, Z.; Haas, M.E.; Hung, W.H.W.; et al. MitoCarta3.0: An Updated Mitochondrial Proteome Now with Sub-Organelle Localization and Pathway Annotations. Nucleic Acids Res. 2021, 49, D1541–D1547. [Google Scholar] [CrossRef]
- DiMauro, S. Mitochondrial Diseases. Biochim. Biophys. Acta BBA—Bioenerg. 2004, 1658, 80–88. [Google Scholar] [CrossRef]
- Atamna, H.; Frey, W.H. Mechanisms of Mitochondrial Dysfunction and Energy Deficiency in Alzheimer’s Disease. Mitochondrion 2007, 7, 297–310. [Google Scholar] [CrossRef] [PubMed]
- Sayehmiri, F.; Motamedi, F.; Batool, Z.; Naderi, N.; Shaerzadeh, F.; Zoghi, A.; Rezaei, O.; Khodagholi, F.; Pourbadie, H.G. Mitochondrial Plasticity and Synaptic Plasticity Crosstalk; in Health and Alzheimer’s Disease. CNS Neurosci. Ther. 2024, 30, e14897. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Du, Y.; Zhao, X.; Wu, C.; Yu, P. Reducing PDK1/Akt Activity: An Effective Therapeutic Target in the Treatment of Alzheimer’s Disease. Cells 2022, 11, 1735. [Google Scholar] [CrossRef] [PubMed]
- Wood, H. Neurodegenerative Disease: PDK1—A Common Therapeutic Target for AD and Prion Disease? Nat. Rev. Neurol. 2013, 9, 543. [Google Scholar] [CrossRef] [PubMed]
- Liu, T.; Yin, H. PDK1 Promotes Tumor Cell Proliferation and Migration by Enhancing the Warburg Effect in Non-Small Cell Lung Cancer. Oncol. Rep. 2017, 37, 193–200. [Google Scholar] [CrossRef]
- Fukushi, A.; Kim, H.-D.; Chang, Y.-C.; Kim, C.-H. Revisited Metabolic Control and Reprogramming Cancers by Means of the Warburg Effect in Tumor Cells. Int. J. Mol. Sci. 2022, 23, 10037. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Jiang, Y.; Meisenhelder, J.; Yang, W.; Hawke, D.H.; Zheng, Y.; Xia, Y.; Aldape, K.; He, J.; Hunter, T.; et al. Mitochondria-Translocated PGK1 Functions as a Protein Kinase to Coordinate Glycolysis and the TCA Cycle in Tumorigenesis. Mol. Cell 2016, 61, 705–719. [Google Scholar] [CrossRef]
- Reiter, R.J.; Sharma, R.N.; Manucha, W.; Rosales-Corral, S.; Almieda Chuffa, L.G.D.; Loh, D.; Luchetti, F.; Balduini, W.; Govitrapong, P. Dysfunctional Mitochondria in Age-Related Neurodegeneration: Utility of Melatonin as an Antioxidant Treatment. Ageing Res. Rev. 2024, 101, 102480. [Google Scholar] [CrossRef]
- Bogenhagen, D.F.; Ostermeyer-Fay, A.G.; Haley, J.D.; Garcia-Diaz, M. Kinetics and Mechanism of Mammalian Mitochondrial Ribosome Assembly. Cell Rep. 2018, 22, 1935–1944. [Google Scholar] [CrossRef]
- Jia, J.; Yin, J.; Zhang, Y.; Xu, G.; Wang, M.; Jiang, H.; Li, L.; Zeng, X.; Zhu, D. Thioredoxin-1 Promotes Mitochondrial Biogenesis through Regulating AMPK/Sirt1/PGC1α Pathway in Alzheimer’s Disease. ASN Neuro 2023, 15, 17590914231159226. [Google Scholar] [CrossRef]
- Chaytow, H.; Carroll, E.; Gordon, D.; Huang, Y.-T.; Van Der Hoorn, D.; Smith, H.L.; Becker, T.; Becker, C.G.; Faller, K.M.E.; Talbot, K.; et al. Targeting Phosphoglycerate Kinase 1 with Terazosin Improves Motor Neuron Phenotypes in Multiple Models of Amyotrophic Lateral Sclerosis. Ebiomedicine 2022, 83, 104202. [Google Scholar] [CrossRef] [PubMed]
- Reyna-Bolaños, I.; Solís-García, E.P.; Vargas-Vargas, M.A.; Peña-Montes, D.J.; Saavedra-Molina, A.; Cortés-Rojo, C.; Calderón-Cortés, E. Polydatin Prevents Electron Transport Chain Dysfunction and ROS Overproduction Paralleled by an Improvement in Lipid Peroxidation and Cardiolipin Levels in Iron-Overloaded Rat Liver Mitochondria. Int. J. Mol. Sci. 2024, 25, 11104. [Google Scholar] [CrossRef] [PubMed]
- Musatov, A.; Robinson, N.C. Susceptibility of Mitochondrial Electron-Transport Complexes to Oxidative Damage. Focus on Cytochrome c Oxidase. Free Radical Res. 2012, 46, 1313–1326. [Google Scholar] [CrossRef]
- Valko, M.; Jomova, K.; Rhodes, C.J.; Kuča, K.; Musílek, K. Redox- and Non-Redox-Metal-Induced Formation of Free Radicals and Their Role in Human Disease. Arch. Toxicol. 2016, 90, 1–37. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Shen, Q.; Liu, Y.; Zhang, Y.; Sun, L.; Ma, X.; Song, N.; Xie, J. Homeostasis and Metabolism of Iron and Other Metal Ions in Neurodegenerative Diseases. Signal Transduct. Target. Ther. 2025, 10, 31. [Google Scholar] [CrossRef] [PubMed]
- Emamnejad, R.; Pagnin, M.; Petratos, S. The Iron Maiden: Oligodendroglial Metabolic Dysfunction in Multiple Sclerosis and Mitochondrial Signaling. Neurosci. Biobehav. Rev. 2024, 164, 105788. [Google Scholar] [CrossRef]
- Wang, P.-F.; Jiang, F.; Zeng, Q.-M.; Yin, W.-F.; Hu, Y.-Z.; Li, Q.; Hu, Z.-L. Mitochondrial and Metabolic Dysfunction of Peripheral Immune Cells in Multiple Sclerosis. J. Neuroinflammation 2024, 21, 28. [Google Scholar] [CrossRef]
- Kumari, S.; Dhapola, R.; Reddy, D.H. Apoptosis in Alzheimer’s Disease: Insight into the Signaling Pathways and Therapeutic Avenues. Apoptosis 2023, 28, 943–957. [Google Scholar] [CrossRef]
- Haack, T.B.; Jackson, C.B.; Murayama, K.; Kremer, L.S.; Schaller, A.; Kotzaeridou, U.; De Vries, M.C.; Schottmann, G.; Santra, S.; Büchner, B.; et al. Deficiency of ECHS 1 Causes Mitochondrial Encephalopathy with Cardiac Involvement. Ann. Clin. Transl. Neurol. 2015, 2, 492–509. [Google Scholar] [CrossRef]
- Burgin, H.J.; McKenzie, M. Understanding the Role of OXPHOS Dysfunction in the Pathogenesis of ECHS1 Deficiency. FEBS Lett. 2020, 594, 590–610. [Google Scholar] [CrossRef]
- Schmitt, K.; Grimm, A.; Kazmierczak, A.; Strosznajder, J.B.; Götz, J.; Eckert, A. Insights into Mitochondrial Dysfunction: Aging, Amyloid-β, and Tau–a Deleterious Trio. Antioxid. Redox Signal. 2012, 16, 1456–1466. [Google Scholar] [CrossRef]
- Palomo, G.M.; Manfredi, G. Exploring New Pathways of Neurodegeneration in ALS: The Role of Mitochondria Quality Control. Brain Res. 2015, 1607, 36–46. [Google Scholar] [CrossRef] [PubMed]
- Iacoangeli, A.; Fogh, I.; Selvackadunco, S.; Topp, S.D.; Shatunov, A.; van Rheenen, W.; Al-Khleifat, A.; Opie-Martin, S.; Ratti, A.; Calvo, A.; et al. SCFD1 Expression Quantitative Trait Loci in Amyotrophic Lateral Sclerosis Are Differentially Expressed. Brain Commun. 2021, 3, fcab236. [Google Scholar] [CrossRef] [PubMed]
- Huang, N.; Li, S.; Xie, Y.; Han, Q.; Xu, X.-M.; Sheng, Z.-H. Reprogramming an Energetic AKT-PAK5 Axis Boosts Axon Energy Supply and Facilitates Neuron Survival and Regeneration after Injury and Ischemia. Curr. Biol. 2021, 31, 3098–3114.e7. [Google Scholar] [CrossRef] [PubMed]
- Peralta, S.; Goffart, S.; Williams, S.L.; Diaz, F.; Garcia, S.; Nissanka, N.; Area-Gomez, E.; Pohjoismäki, J.; Moraes, C.T. ATAD3 Controls Mitochondrial Cristae Structure in Mouse Muscle, Influencing mtDNA Replication and Cholesterol Levels. J. Cell Sci. 2018, 131, jcs217075. [Google Scholar] [CrossRef]
- Lei, Y.; Guerra Martinez, C.; Torres-Odio, S.; Bell, S.L.; Birdwell, C.E.; Bryant, J.D.; Tong, C.W.; Watson, R.O.; West, L.C.; West, A.P. Elevated Type I Interferon Responses Potentiate Metabolic Dysfunction, Inflammation, and Accelerated Aging in mtDNA Mutator Mice. Sci. Adv. 2021, 7, eabe7548. [Google Scholar] [CrossRef]
- Newman, L.E.; Weiser Novak, S.; Rojas, G.R.; Tadepalle, N.; Schiavon, C.R.; Grotjahn, D.A.; Towers, C.G.; Tremblay, M.-È.; Donnelly, M.P.; Ghosh, S.; et al. Mitochondrial DNA Replication Stress Triggers a Pro-Inflammatory Endosomal Pathway of Nucleoid Disposal. Nat. Cell Biol. 2024, 26, 194–206. [Google Scholar] [CrossRef] [PubMed]
- Chujo, T.; Tomizawa, K. Neurological Diseases Caused by Loss of Transfer RNA Modifications: Commonalities in Their Molecular Pathogenesis. J. Mol. Biol. 2025, 437, 169047. [Google Scholar] [CrossRef]
- Eulenburg, V.; Hülsmann, S. Synergistic Control of Transmitter Turnover at Glycinergic Synapses by GlyT1, GlyT2, and ASC-1. Int. J. Mol. Sci. 2022, 23, 2561. [Google Scholar] [CrossRef]
- Cavarischia-Rega, C.; Sharma, K.; Fitzgerald, J.C.; Macek, B. Proteome Dynamics in iPSC-Derived Human Dopaminergic Neurons. Mol. Cell. Proteomics 2024, 23, 100838. [Google Scholar] [CrossRef]
- Pena, I.A.; Shi, J.S.; Chang, S.M.; Yang, J.; Block, S.; Adelmann, C.H.; Keys, H.R.; Ge, P.; Bathla, S.; Witham, I.H.; et al. SLC25A38 Is Required for Mitochondrial Pyridoxal 5′-Phosphate (PLP) Accumulation. Nat. Commun. 2025, 16, 978. [Google Scholar] [CrossRef] [PubMed]
- Tan, S.; Dengler, A.S.; Darawsheh, R.Z.; Kory, N. The iAAA-Mitochondrial Protease YME1L1 Regulates the Degradation of the Short-Lived Mitochondrial Transporter SLC25A38. bioRxiv 2024. [Google Scholar] [CrossRef]
- Xiao, Y.; Karam, C.; Yi, J.; Zhang, L.; Li, X.; Yoon, D.; Wang, H.; Dhakal, K.; Ramlow, P.; Yu, T.; et al. ROS-Related Mitochondrial Dysfunction in Skeletal Muscle of an ALS Mouse Model during the Disease Progression. Pharmacol. Res. 2018, 138, 25–36. [Google Scholar] [CrossRef]
- Choi, E.-H.; Kim, M.-H.; Park, S.-J. Targeting Mitochondrial Dysfunction and Reactive Oxygen Species for Neurodegenerative Disease Treatment. Int. J. Mol. Sci. 2024, 25, 7952. [Google Scholar] [CrossRef] [PubMed]
- Jadiya, P.; Garbincius, J.F.; Elrod, J.W. Reappraisal of Metabolic Dysfunction in Neurodegeneration: Focus on Mitochondrial Function and Calcium Signaling. Acta Neuropathol. Commun. 2021, 9, 124. [Google Scholar] [CrossRef]
- Gill, A.J.; Smith, M.D.; Galleguillos, D.; Garton, T.; Mace, J.W.; Gadani, S.P.; Kumar, S.; Pokharel, A.; Solem, K.; Potluri, S.; et al. NLRX1 Limits Inflammatory Neurodegeneration in the Anterior Visual Pathway. J. Neuroinflammation 2025, 22, 21. [Google Scholar] [CrossRef]
- Gharagozloo, M.; Mahmoud, S.; Simard, C.; Yamamoto, K.; Bobbala, D.; Ilangumaran, S.; Smith, M.D.; Lamontagne, A.; Jarjoura, S.; Denault, J.-B.; et al. NLRX1 Inhibits the Early Stages of CNS Inflammation and Prevents the Onset of Spontaneous Autoimmunity. PLoS Biol. 2019, 17, e3000451. [Google Scholar] [CrossRef]
- Lin, M.; Liu, N.; Qin, Z.; Wang, Y. Mitochondrial-Derived Damage-Associated Molecular Patterns Amplify Neuroinflammation in Neurodegenerative Diseases. Acta Pharmacol. Sin. 2022, 43, 2439–2447. [Google Scholar] [CrossRef]
- Mishra, Y.; Kumar, A.; Kaundal, R.K. Mitochondrial Dysfunction Is a Crucial Immune Checkpoint for Neuroinflammation and Neurodegeneration: mtDAMPs in Focus. Mol. Neurobiol. 2025, 62, 6715–6747. [Google Scholar] [CrossRef] [PubMed]
- Harel, T.; Yoon, W.H.; Garone, C.; Gu, S.; Coban-Akdemir, Z.; Eldomery, M.K.; Posey, J.E.; Jhangiani, S.N.; Rosenfeld, J.A.; Cho, M.T.; et al. Recurrent de Novo and Biallelic Variation of ATAD3A, Encoding a Mitochondrial Membrane Protein, Results in Distinct Neurological Syndromes. Am. J. Hum. Genet. 2016, 99, 831–845. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Hu, D.; Wang, R.; Sun, X.; Ropelewski, P.; Hubler, Z.; Lundberg, K.; Wang, Q.; Adams, D.J.; Xu, R.; et al. ATAD3A Oligomerization Promotes Neuropathology and Cognitive Deficits in Alzheimer’s Disease Models. Nat. Commun. 2022, 13, 1121. [Google Scholar] [CrossRef] [PubMed]
- Balog, J.; Mehta, S.L.; Vemuganti, R. Mitochondrial Fission and Fusion in Secondary Brain Damage after CNS Insults. J. Cereb. Blood Flow Metab. 2016, 36, 2022–2033. [Google Scholar] [CrossRef] [PubMed]
- Jacobs, M.; Geiger, M.; Summers, S.; Janes, T.; Boyea, R.; Zinn, K.; Aburashed, R.; Spence, D. Interferon-β Decreases the Hypermetabolic State of Red Blood Cells from Patients with Multiple Sclerosis. ACS Chem. Neurosci. 2022, 13, 2658–2665. [Google Scholar] [CrossRef]
- Sun, D.; Wang, R.; Du, Q.; Chen, H.; Shi, Z.; Zhang, Y.; Zhang, N.; Wang, X.; Zhou, H. Integrating Genetic and Proteomic Data to Elucidate the Association between Immune System and Blood-Brain Barrier Dysfunction with Multiple Sclerosis Risk and Severity. J. Affect. Disord. 2024, 362, 652–660. [Google Scholar] [CrossRef]
- Beura, S.K.; Panigrahi, A.R.; Yadav, P.; Singh, S.K. Role of Platelet in Parkinson’s Disease: Insights into Pathophysiology & Theranostic Solutions. Ageing Res. Rev. 2022, 80, 101681. [Google Scholar] [CrossRef]
- Ma, H.; Jiang, T.; Tang, W.; Ma, Z.; Pu, K.; Xu, F.; Chang, H.; Zhao, G.; Gao, W.; Li, Y.; et al. Transplantation of Platelet-Derived Mitochondria Alleviates Cognitive Impairment and Mitochondrial Dysfunction in Db/Db Mice. Clin. Sci. 2020, 134, 2161–2175. [Google Scholar] [CrossRef]
- Ganjam, G.K.; Bolte, K.; Matschke, L.A.; Neitemeier, S.; Dolga, A.M.; Höllerhage, M.; Höglinger, G.U.; Adamczyk, A.; Decher, N.; Oertel, W.H.; et al. Mitochondrial Damage by α-Synuclein Causes Cell Death in Human Dopaminergic Neurons. Cell Death Dis. 2019, 10, 865. [Google Scholar] [CrossRef]
- Gallardo-Pérez, J.C.; De Guevara, A.A.-L.; García-Amezcua, M.A.; Robledo-Cadena, D.X.; Pacheco-Velázquez, S.C.; Belmont-Díaz, J.A.; Vargas-Navarro, J.L.; Moreno-Sánchez, R.; Rodríguez-Enríquez, S. Celecoxib and Dimethylcelecoxib Block Oxidative Phosphorylation, Epithelial-Mesenchymal Transition and Invasiveness in Breast CancerStem Cells. Curr. Med. Chem. 2022, 29, 2719–2735. [Google Scholar] [CrossRef]
- Ward, K.; Reddel, S.; Leocani, L.; Barnett, M. Monitoring Multiple Sclerosis: Digital and Fluid Phase Biomarkers. Curr. Opin. Neurol. 2025, 38, 243–248. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Yang, Y.; Zhao, A.; Luo, N.; Niu, M.; Kang, W.; Xie, A.; Lu, H.; Chen, L.; Liu, J. Parkinson’s Disease Peripheral Immune Biomarker Profile: A Multicentre, Cross-Sectional and Longitudinal Study. J. Neuroinflammation 2022, 19, 116. [Google Scholar] [CrossRef] [PubMed]

| Disease | GWAS ID | Sample Size (Case/Control) | Description | PMID |
|---|---|---|---|---|
| Alzheimer’s Disease (AD) | ebi-a-GCST90027158 | 39,106 clinically diagnosed cases, 46,828 proxy cases, 401,577 controls | A two-stage GWAS identified 75 AD risk loci, 42 of which are novel. Pathway analyses revealed amyloid/tau involvement and microglia implications, with a genetic risk score showing a 1.6- to 1.9-fold increased AD risk across deciles. | 35379992 |
| Amyotrophic Lateral Sclerosis (ALS) | ebi-a-GCST005647 | 20,806 ALS cases, 59,804 controls | A GWAS identified KIF5A mutations in the C-terminal domain as a novel ALS risk factor, implicating cytoskeletal defects. The study contrasted these with SPG10 and CMT2 mutations, linking them to ALS pathogenesis. | 29566793 |
| Multiple Sclerosis (MS) | ukb-b-17670 | 462,933 (1679 cases, 461,254 controls) | A large-scale GWAS from the UK Biobank analyzed genetic factors contributing to MS risk. | -- |
| Parkinson’s Disease (PD) | ieu-b-7 | 482,730 (33,674 cases, 449,056 controls) | The International Parkinson’s Disease Genomics Consortium conducted a GWAS to investigate genetic factors in PD. | -- |
| Disease | GEO NO. | Sample Size (case/control) | Description | PMID |
| AD | GSE118553 | 33 AsymAD, 52 AD, 27 Control | Differential and co-expression analysis on brain tissue samples identified significant transcriptomic changes in the frontal cortex of AsymAD subjects. A total of 14 genes, including GPM6B and ANKEF1, were linked to AD neuropathology. | 31063847 |
| ALS | GSE112681 | 396 ALS patients, 75 ALS mimic diseases, 645 healthy controls | Microarray analysis identified 752 ALS-increased and 764 ALS-decreased DEGs, highlighting gene expression shifts resembling acute stress responses. A 61-gene signature was linked to improved survival prediction. | 29939990/ 31118040 |
| MS | GSE131282 | 64 MS NAGM samples, 42 control gray matter samples | Gene expression microarray analysis found elevated HLA-DRB1 expression in MS NAGM, especially in cases with the HLA-DR15 haplotype, suggesting a role in MS pathogenesis. | 31882398 |
| PD | GSE28894 | 14–15 control brains, 11–15 PD brains across four regions | Gene expression profiling identified modest differences between PD and control brains. RNA was generated from 500 ng of total RNA from these regions (medulla: 15 control brains, 14 PD brains; striatum: 15 control brains, 15 PD brains; frontal cortex: 15 control brains, 11 PD brains; cerebellum: 14 control brains, 15 PD brains). | -- |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Du, Y.; Fan, S.-S.; Wu, H.; He, J.; He, Y.; Meng, X.-Y.; Xu, X. Convergent and Divergent Mitochondrial Pathways as Causal Drivers and Therapeutic Targets in Neurological Disorders. Curr. Issues Mol. Biol. 2025, 47, 636. https://doi.org/10.3390/cimb47080636
Du Y, Fan S-S, Wu H, He J, He Y, Meng X-Y, Xu X. Convergent and Divergent Mitochondrial Pathways as Causal Drivers and Therapeutic Targets in Neurological Disorders. Current Issues in Molecular Biology. 2025; 47(8):636. https://doi.org/10.3390/cimb47080636
Chicago/Turabian StyleDu, Yanan, Sha-Sha Fan, Hao Wu, Junwen He, Yang He, Xiang-Yu Meng, and Xuan Xu. 2025. "Convergent and Divergent Mitochondrial Pathways as Causal Drivers and Therapeutic Targets in Neurological Disorders" Current Issues in Molecular Biology 47, no. 8: 636. https://doi.org/10.3390/cimb47080636
APA StyleDu, Y., Fan, S.-S., Wu, H., He, J., He, Y., Meng, X.-Y., & Xu, X. (2025). Convergent and Divergent Mitochondrial Pathways as Causal Drivers and Therapeutic Targets in Neurological Disorders. Current Issues in Molecular Biology, 47(8), 636. https://doi.org/10.3390/cimb47080636






