Abstract
Microglia belong to tissue-resident macrophages of the central nervous system (CNS), representing the primary innate immune cells. This cell type constitutes ~7% of non-neuronal cells in the mammalian brain and has a variety of biological roles integral to homeostasis and pathophysiology from the late embryonic to adult brain. Its unique identity that distinguishes its “glial” features from tissue-resident macrophages resides in the fact that once entering the CNS, it is perennially exposed to a unique environment following the formation of the blood–brain barrier. Additionally, tissue-resident macrophage progenies derive from various peripheral sites that exhibit hematopoietic potential, and this has resulted in interpretation issues surrounding their origin. Intensive research endeavors have intended to track microglial progenitors during development and disease. The current review provides a corpus of recent evidence in an attempt to disentangle the birthplace of microglia from the progenitor state and underlies the molecular elements that drive microgliogenesis. Furthermore, it caters towards tracking the lineage spatiotemporally during embryonic development and outlining microglial repopulation in the mature CNS. This collection of data can potentially shed light on the therapeutic potential of microglia for CNS perturbations across various levels of severity.
1. Introduction
The central nervous system’s (CNS) principal innate immune cells are tissue-resident macrophages, which include microglia [1,2]. While microglia are the parenchymal brain macrophages, the perivascular, meningeal, and choroid plexus macrophages constitute the non-parenchymal tissue-resident macrophages of the CNS [3]. Microglia have a range of biological activities in both the developing and adult mammalian brain, although this population of cells makes up the lowest percentage of non-neuronal cells in the mammalian brain [4]. The release of mediators (e.g., trophic factors, cytokines) and phagocytosis are the two main mechanisms by which microglia shape brain development and perform key functions across life [5]. These microglial activities are implicated in developmental processes such as synaptic patterning, myelinogenesis, axonal dynamics, cell positioning, and survival [6]. In the adult brain, microglial activities are central to the regulation of acute and chronic immune responses and maintenance of CNS homeostasis through the removal of viruses, bacteria, and other foreign particles, but also cellular debris and synapses, mediation of neurogenesis following CNS injury, and protection of neural tissue [7,8,9]. However, microglia as innate immune cells are sensitive to chronic inflammation, which can impair their beneficial functions and participate in the etiology of protracted neurodegenerative diseases such as Parkinson’s and Alzheimer’s diseases, multiple sclerosis, and amyotrophic lateral sclerosis (ALS) [10,11,12].
The term microglia—micro (small) and glia (glue)—was first introduced in 1919 by Pío del Río Hortega, who proposed that microglia adopt a malleable morphology, transforming from a resting to an activated state during disease exhibiting phagocytic properties [13]. This view was recently considered too simplistic, as microglia can adopt a wide variety of morphological and functional states [5]. Under normal physiological conditions, microglia display a ramified morphology with multiple branches and processes constantly surveilling the CNS parenchyma. Inflammatory stimuli can change microglial morphology, for instance converting microglia from a ramified to an amoeboid form characterized by an enlarged cell body and retracted processes. In contrast to the ramified state, amoeboid microglia with an amoeboid morphology are generally considered to exhibit a high phagocytic and proinflammatory phenotype. The activated microglial cells were previously categorized as: classical (M1) or alternative (M2), corresponding to either a proinflammatory and neurotoxic state or an anti-inflammatory state, respectively. However, it is now suggested that the M1/M2 phenotypes are not representative of in vivo microglia states because microglia rarely appear with a distinct M1 or M2 phenotype [5,14].
Tissue-resident macrophages are present in the CNS and across different organs, such as osteoclasts in bone, intestinal macrophages in the gastrointestinal tract, Kupffer cells in liver, alveolar macrophages in lungs, and Langerhans cells in skin [1]. Microglial cells are unique tissue-resident macrophages that differ from their hematogenous origins due to their surrounding environment, which is immune-privileged owing to the formation of the blood–brain barrier (BBB). The timing of BBB formation is vital for the invasion of microglia progenitors during embryonic development. Many studies have delineated that closure is vitally important at specific embryonic days. The permeability of BBB was found to decrease for large molecules from E12.5, and it became impermeable to small molecules as early as E14.5 [15]. This tight regulation showcases the importance of CNS master regulator elements to protect the central environment from pathogens and other harmful agents.
Each tissue-resident macrophage type has a distinct embryonic origin, as their progenitors derive from different waves of hemopoiesis. Consequently, the understanding of embryonic hematopoiesis is vital for delineating the microglial origin. Regarding hematopoiesis in embryonic life, three waves have been described in mice. The primitive hematopoiesis starts at E7.5 in the yolk sac (YS), generating primitive erythroid, megakaryocyte, and macrophage progenitors such as early c-MYB-independent erythro-myeloid precursors (EMPs). The second hematopoietic wave, called transient definitive hematopoiesis, originates from YS hemogenic endothelium, giving rise to late c-MYB-dependent EMPs at E8.25 and progenitors with lymphoid potentials at E9, which additionally emerge from the developing aorta-gonad-mesonephros (AGM) region. The definitive hematopoiesis occurs at E10.5 with the generation of hematopoietic stem cells (HSCs) that originate from the embryonic AGM region, colonizing slightly later the fetal liver [16,17,18,19].
The present review aims to define existing data on the origin of microglia because there has been controversy over their ontogeny. The developmental milestones that are being covered herein are the primary cues that direct microgliogenesis. The ontogeny of microglia is investigated thoroughly, as it is of prime importance considering that this cell type is involved in the pathogenesis of many diseases and is presented as a target for therapies that are being developed to control the associated phenotypes.
2. Discovery and Ontogeny of Microglia
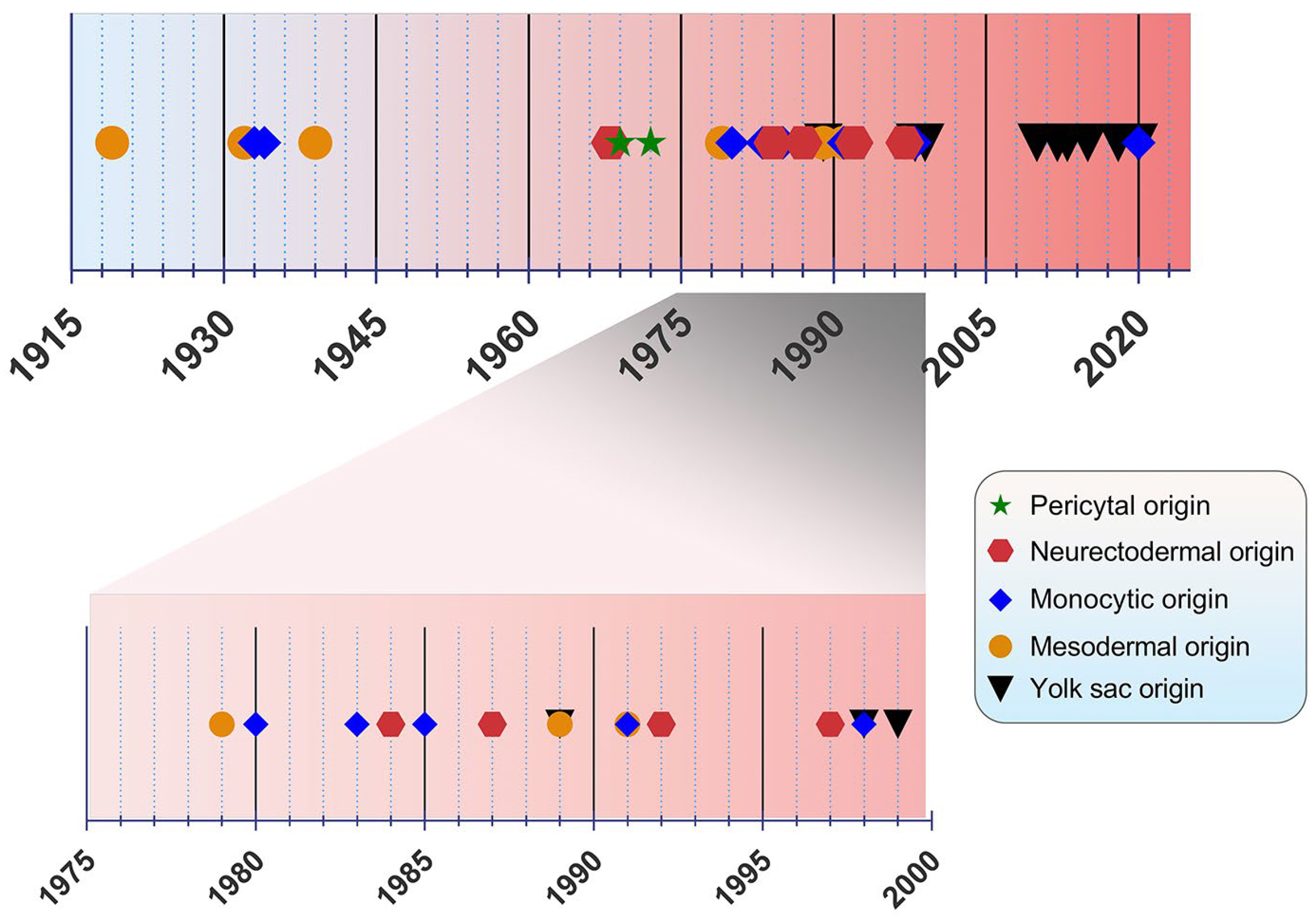
The origin of microglia has been a debated topic for years. In the past, four main origin concepts have been proposed as a source of microglia: (i) the mesodermal-associated mater elements, (ii) the neuroectodermal matrix cells, (iii) the pericytes, and (iv) the invasion of monocytes especially during early development (Figure 1) [20]. Río Hortega was hailed as the “Father of Microglia”, because their discovery supported the mesodermal origin, after observing the invasion of the pial elements within the CNS parenchyma [13,21]. Using comparable staining methods, John Kershman agreed on the mesenchymal origin of microglia, which were found to be genetically related to histiocytes, a stationary phagocytic cell present in connective tissue [22]. With reference to Boya et al., the meningeal envelope was proposed to be the source of microglia, which sustains a mesodermal nature in agreement with the classical experiments by Río Hortega [23,24]. Later, the theory of multiple mesodermal sources of microglia depending on time and localization was posited [25]. However, another study proposed vascular pericytes as the parent cells of microglia [26,27]. The first reports on the monocytic origin of microglia came to the fore in 1933 and 1934 from Santha and Juba, respectively, who hypothesized that ramified microglia originated from circulating monocytes because the initial appearance of these cells coincided with the vascularization of the brain [28,29].
Figure 1.
Timeline view of microglial origin since their discovery by Río Hortega.
In the following decades, many researchers accepted this view, demonstrating microglial monocytic identity when investigating their origin [30,31,32,33], while others rejected the possibility that microglia are derived from mononuclear blood cells [34]. In 1968, autoradiography experiments performed with tritiated thymidine were conducted in adult rats, showing that cells of the subependymal layer give rise to a number of glial cell types, such as astrocytes and microglia, offering a different perspective regarding microglial origin [35]. The neuroectodermal origin was also supported by Kitamura et al., implying that glioblasts are the source of both astrocytes and microglia in mice [36]. It was also proposed that microglia and astroglia have a common progenitor cell developing from neuroepithelial cells [37]. Performing non-radioactive in situ hybridization and immunoperoxidase techniques, only a small population of microglia were found to be derived from bone marrow progenitors, because most of the cells were shown to be generated from locally residing precursors with a neuroectodermal ontogeny [38]. The non-monocytic origin of microglia favored by Schelper and Adrian implicated that these cells are CNS intrinsic ones, enforcing the above theory of a neuroectodermal origin [39]. This perception was also put forward by other researchers, but began to lose ground from the 2000’s onwards. [40,41].
The view of the origin of microglia from the YS was first introduced in 1989 [42]. A nucleoside diphosphatase histochemical study was conducted to evaluate the distribution of microglia in the developing human CNS, implying that mesenchymal cells with haemopoietic potential migrate into neural tissues and then give rise to cells resembling microglia [43]. Likewise, primitive macrophages of YS were found to be derived from fetal macrophages before the appearance of pro-monocytes/monocytes colonizing the embryonic tissues in mice [42]. In an avian model, microglia precursors were demonstrated to invade neural tissue from the pial surface and proliferated inside the CNS, indicating that their penetration through the embryonic CNS vessels is not possible [44]. However, a human embryogenesis study using lectin+ and CD68+ markers revealed two populations of microglia, indicating two different potential origins, specifically from the YS and bone marrow. Different routes of entry were also proposed: one through the mesenchyme and the other via the blood circulation [45]. Alliot et al., aiming to delineate the origin of microglia in mice, detected these cells in the brain from Ε8 being derived from YS progenitors, which proliferate in situ [46].
The YS origin of microglia was confirmed by Ginhoux et al. by performing a fate mapping analysis in mice and showing that YS primitive myeloid progenitors generated before E7.5 can contribute to the CNS microglial population [47]. Moreover, in this study, RUNX1+ YS progenitors were found to migrate into the brain through blood vessels between E8.5 and E9.5 [47]. The YS origin was further supported by identifying the transcription factor MYB, which is required for the development of HSCs as well as CD11bhigh monocytes and macrophages [48], contrary to YS-derived macrophages, which are the potential precursors of CNS microglia [49]. Specifically, primitive c-kit+ EMPs detected from E8 in the YS were proposed to serve as the precursors of microglia in mice [50]. As the progenitors of microglia were identified to be the EMPs of YS, the vast majority of other tissue-resident macrophages arise from fetal monocytes that derive from late c-MYB+ EMPs of the YS [51]. The HSC-derived hematopoiesis that takes place for monocytes at E14.5 and granulocytes at E16.5 in mice advocates that these progenitors only seldom replace parenchymal microglia, which mainly emanates from CSF-1R+ EMPs. [52]. This view was re-evaluated by Sheng et al., who developed the KitMercreMer fate mapping mouse strain and suggested that all resident-tissue macrophages, except microglia and Langerhans cells of the epidermis, are derived from HSCs [53].
In 2018, De et al. identified two distinct microglial cell populations, namely canonical (non-HOXB8) and HOXB8 microglia using a transgenic strategy, fluorescence-activated cell sorting technique in YS and qRT-PCR in HOXB8 cells in the different hematopoietic tissues [54]. The HOXB8 population was suggested to be derived from the second wave of YS hematopoiesis populating the AGM and fetal liver. Besides the YS, an additional source of microglia was proposed by Fehrenbach et al., who considered the definitive hemopoiesis as responsible for microglial development and recruitment to the mouse CNS, especially at the post-YS phase [55]. Besides parenchymal microglia, a genetic distinct population of macrophages was identified, namely the border-associated macrophages (BAMs) residing among the meninges, choroid plexus, and perivascular spaces. Like microglia, these cells are generated by early EMPs; however, microglia require TGF-β for their development, whereas BAMs are TGF-β-independent. Additionally, in the mouse YS, two distinctive primitive populations were observed: the CD206− and CD206+ macrophages. The differentiation of these populations after their final colonization is mediated by environmental drivers [56]. Interestingly, tamoxifen dosing in CCR2-CreER transgenic mice suggested that not only YS EMPs, but also fetal HSC-derived monocytes participate in the generation of IBA1+TMEM119+P2RY12+ parenchymal microglia, IBA1+, and isolectin+ BAMs in the mouse brain [57]. Lastly, a recent study in eight aborted human embryos proposed that tissue-resident macrophages development is very similar to other mammalian species, highlighting the presence of two distinct waves of YS-derived macrophages. Specifically for microglia, they were found to be derived from the early first wave along with a minor contribution from the second one [58].
To recapitulate, the YS is the main site of microglial origin. The suggested microglial progenitors in mice are the early, c-ΜΥΒ-independent, CSF-1R+ EMPs of the YS. However, the definite nomenclature of the progenitors and the confirmation in human models are still under consideration.
3. Molecular Cues Orchestrating Microgliogenesis
Upon birth, the phenotype of microglia corresponds to an amoeboid shape, phagocytically and mitotically active, while in later developmental stages, microglia become ramified. The RUNX1, a transcription factor expressed during the first two postnatal weeks at the forebrain by amoeboid microglia, downregulates the proliferation of these cells and assists in their transformation towards a ramified morphology [59]. During embryonic development, RUNX1 controls the expression of the transcription factor PU.1 [60]. In Irf8-deficient YS, the number of A1 cells (CD45+ c-kitlo CX3CR1− immature cells) remained unchanged, while the A2 population (CD45+ c-kit− CX3CR1+ cells) decreased [50]. Additionally, Pu.1 deficiency provoked an impairment of A1 and A2 progenitors. From A2 cells, microglia were generated and expanded in the developing brain under the influence of specific matrix metalloproteinases, such as MMP-9 and MMP-8. Factors such as MYB, BATF3, ID2, Klf4, and NR4A1 were not necessary for the development of microglia from their progenitors [50,61]. While PU.1 was essential for terminal myeloid differentiation, early myeloid genes such as Gm-csfr, G-csfr, and Mpo were maintained in Pu.1-/- embryos, whereas myeloid genes associated with terminal differentiation (etc. Cd11b, Cd64, and M-csfr) were found to be impaired [62].
The CSF-1R is a vital receptor for microglial cell development expressed on YS macrophages and microglia at E9.5 and throughout brain development. In contrast to many tissue macrophages, adult microglia can still be replenished, albeit at reduced levels in Csf-1op/op mice. Although the microglia presented—even in small amounts—in a null mutation model of the Csf-1 in Csf-1op/op mice, microglia were fully depleted in mice lacking CSF-1R [47]. This was a strong clue that a second ligand of CSF-1R, namely the IL-34, was implicated in microgliogenesis. As the microglial phenotype in Csf-1r-/- mice was more severe than that observed in Csf-1op/op mice, it was evident that IL-34 plays a significant role in the regulation of microglial homeostasis. Its mRNA expression in the brain is also significantly higher than that of CSF-1 during early postnatal development [47]. In addition, in il-34- and csf-1ra-deficient zebrafish larva, the migration and colonization of CNS by embryonic macrophages was impaired, indicating a role for the Il34-Csf1ra pathway during microglial cell expansion throughout the CNS [63].
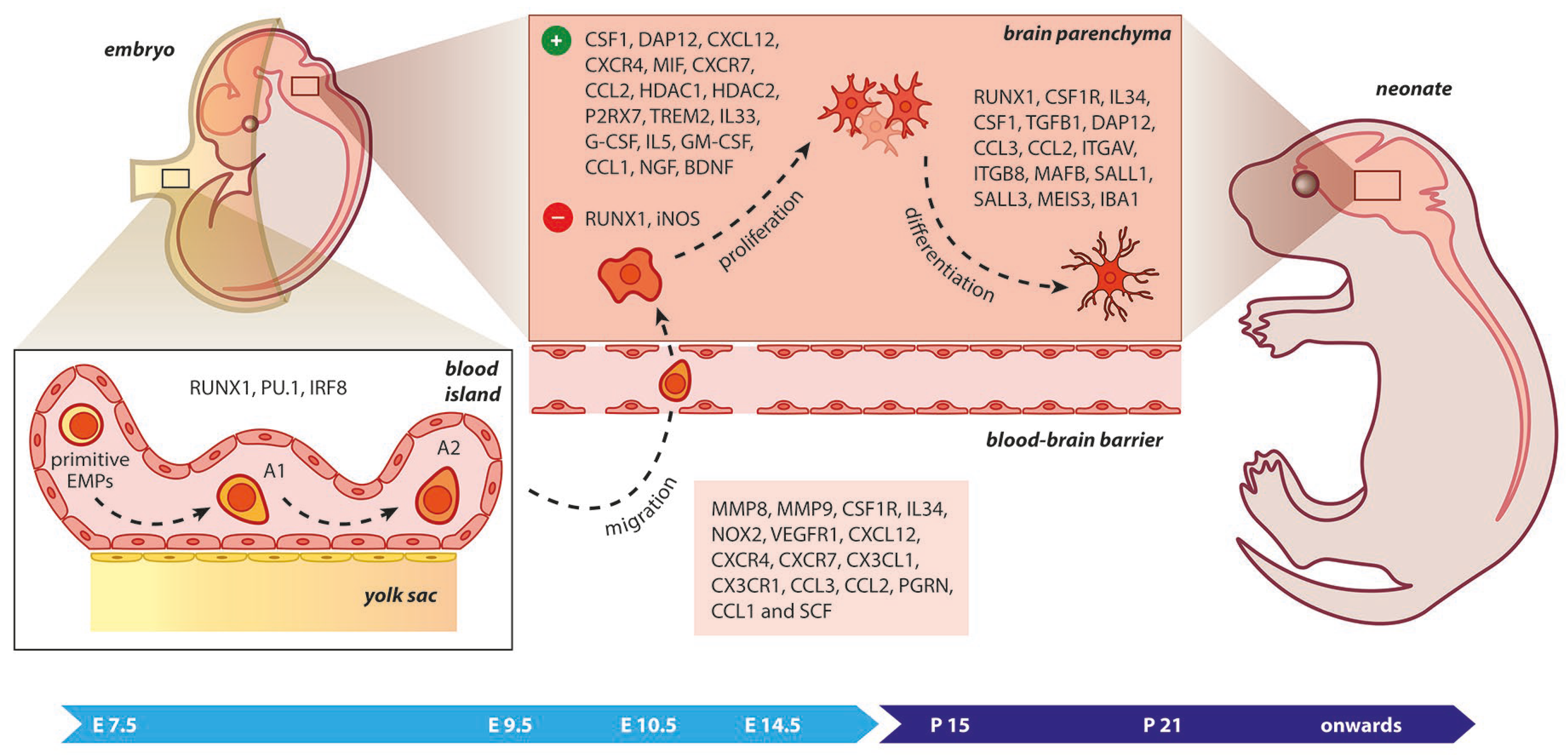
Microglia require TGF-β signaling to be maintained in their surveillant state, but not for their survival. The absence of TGF-β1 was found to have an impact on microglial development from E14.5, but not on microglial progenitors at E10.5 [64,65]. Other transcription factors that include SALL1, SALL3, and MEIS3 are involved in the specification of tissue-resident macrophages during organogenesis and ensure microglial function [66]. In fact, when the SALL1 locus was inducibly inactivated, microglial cells transformed from a ramified morphology to pro-inflammatory deregulating tissue homeostasis [64]. A sharp decline in the number of microglial cells was observed in postnatal Dap12-deficient mice that was comparable to the in vitro impairment of microglial cell differentiation [67]. This may be due to M-CSF’s role in inducing microglial proliferation and survival via a pathway requiring DAP12 and β-catenin. However, another study showed that microglial populations remained unaffected in Dap12-deficient mice similar to young (embryonic and early postnatal) wild-type mice, while a reduction in their numbers was observed in specific CNS regions of deficient adult mice [50,68]. In Nox2 gene deficiency, treatment with apocynin, which is a NOX2 inhibitor, or impairment of the VEGFR1 kinase resulted in microglia that could not migrate efficiently into the caudal subventricular zone (SVZ) of the cerebral cortex, suggesting that chemotaxis of microglia was under the influence of NOX2 and VEGFR1 activation (Figure 2) [69].
Figure 2.
Microgliogenesis at a glance. The primitive erythro-myeloid progenitors (EMPs; early, c−MYB−independent, CSF−1R+ EMPs) arise from the yolk sac (YS) as early as embryonic day 7.5 (E7.5). These cells give rise to CD45+ c−kitlo CX3CR1− immature (A1) cells that develop into CD45+ c−kit− CX3CR1+ (A2) cells. The early differentiation of microglial progenitors is regulated by the expression of RUNX1, PU.1, and IRF8. The invasion of progenitors into the neural tube begins at E9.5 through blood circulation and is followed by proliferation and terminal differentiation. As the blood–brain barrier becomes impermeable to small molecules at E14.5, the microglia invasion may be prevented. The transformation of immature microglia into ramified (mature form) occurs between the second and third postnatal weeks. The migration, proliferation, and terminal differentiation of microglia are also orchestrated from the depicted molecular cues. Light blue arrow timeline represents prenatal period, dark blue arrow timeline refers to postnatal days.
The depletion of Cxcl12 seems to block microglial cell invasion into the SVZ, whereas the ectopic Cxcl12 expression or pharmacological impairment of CXCR4 demonstrated that the CXCL12/CXCR4 signaling is involved in microglial cell recruitment assisting cortical development. In the same context, cell death occurring in the developing forebrain stimulates microglial cell proliferation mediated via MIF activation [70]. Treatment with CXCL12 activates Erk1/2 and Akt signaling, which are necessary for microglial proliferation mediated by CXCL12. Similarly, Erk1/2 signaling was found to be important for CXCL12-depedent migration of microglial populations. Pharmacological blockade of CXCR4 or CXCR7 induced a decline in CXCL12-mediated proliferation and migration of microglial cells, suggesting that CXCR4 and CXCR7 form a receptor unit for CXCL12 in the rodent microglia required for the aforementioned developmental processes, both in vitro and in vivo [71]. Furthermore, CX3CL1/CX3CR1 signaling may regulate microglial invasion within CNS parenchyma during postnatal life [72]. Interestingly, the transformation of microglia from an amoeboid to a ramified morphology was proposed to be mediated by cues released from astrocytes. Utilizing time-lapse video microscopy in co-cultures of human fetal microglial cells and astrocytic cells, the chemokines MIP-1α and MCP-1 were identified as regulators of microglial motility and differentiation [73].
The overexpression of miR-124 in microglia accelerated the transformation of these cells to an inactivated state through inhibition of the C/EBP-α and PU.1, while the depletion of miR-124 led to microglial activation both in vitro and in vivo. These findings underscored the potential role of miR-124 as a regulator of microglial surveillance in the CNS [74]. Microglial polarization is regulated by ARID1A, an epigenetic subunit of the SWI/SNF chromatin-remodeling complex, through alterations of the chromatin state in microglia [75,76]. The migration of microglial cells also seemed to be affected by PGRN, because its knockdown resulted in a failure of microglial precursors to colonize the embryonic retina [77]. The absence of integrin αVβ8 from the CNS prevents microglial transition from immature precursors to a mature state. As αVβ8 controls TGFβ signaling to microglia, these “dysmature” microglial populations are expanded as a consequence of impaired TGFβ signaling during the perinatal period, leading to disrupted oligodendrocyte development, interneuron loss, and neuromotor dysfunction [78]. Epigenetic factors may also affect microglial development. Embryonic HDAC1 and HDAC2 absence disrupts microgliogenesis, altering the crucial acetylation marks implicated in morphology, reactivity, cell cycle, and apoptosis. Specifically, reduced proliferation and induced apoptosis were observed after ablation of the above epigenetic regulators, resulting in the hyperacetylation of specific pro-apoptotic and cell cycle genes [79].
Fate-mapping strategies remain the best way to track cells from the embryonic YS (microglia) versus bone-marrow (monocyte-derived macrophages). In terms of markers, the exact distinction between microglia and periphery-originated macrophages is challenging as they express common markers such as CD11b, CX3CR1, CD45, F4/80, and IBA-1 [80]. Nevertheless, TMEM119 has been recognized as a trans-membranous molecule that is abundantly produced only by microglia, along with P2RY12, but both markers can be downregulated in disease [5,81,82]. However, recently it was proposed that TMEM119 is neither a specific nor a reliable marker for microglial cells [83]. Siglec-H was also found to be a specific marker for microglia in rodents, as it was almost absent in CNS-infiltrating monocytes and CNS-associated macrophages [84]. Recently, HexB has also emerged as a promising marker, but the characterization is still largely lacking [85]. On the contrary, CD44 and CD169 are markers expressed only in peripheral-divided cells and not on resident microglia [86,87].
TREM2, as a protein involved in intracellular signals, interacts with transmembrane protein DAP12, thus activating the Wnt/β-catenin pathway and stabilizing β-catenin via blocking GSK3β activation. Thus, TREM2 promotes the survival and proliferation of primary microglial cells [88]. In addition, the transcription factor MAFB may be involved in regulating microglial cell development and homeostasis [89]. The homeostasis is further preserved by the epigenetic regulator MECP2, which controls microglial responsiveness to external stimuli [90,91]. In the postnatal developing brain, the absence of microglial EED, a Polycomb protein vital for synaptic pruning, led to the upregulation of phagocytosis-related genes [92]. Contrariwise, the deletion of microglial Tgm2 in mice resulted in the downregulation of microglial phagocytic-related genes accompanied by synaptic pruning and cognitive impairment [93]. A P2RX7-induced proliferation of embryonic spinal cord microglia was proposed after comparison of wild-type and P2rx7-/- embryos. The ablation of P2rx7 also affected microglial density, while Pannexin-1-/- embryos showed unaltered proliferation rates. Altogether, microglial proliferation may be regulated by P2RX7 receptors in a Pannexin-1-independent way during early development [94].
Another in vitro study confirmed that IL-33, which is released by astrocytes and endothelial cells, enhances the proliferation of microglial populations [95]. Similarly, in the uninjured CNS, G-CSF increased microglial numbers [96]. However, the GM-CSF was a stronger stimulus for microglial proliferation in human brain cultures [97]. The increasing microglial populations were correlated with a direct effect of GM-CSF upon treatment with IL-5, whereas IL-5 induced an intense cellular metabolism in contrast with GM-CSF treatment in microglial cell cultures [98]. Moreover, 1 ng/mL of CCL-1 mediated chemotaxis, while 100 ng/mL increased motility, proliferation, and phagocytosis of microglial cells in culture [99]. An induction of microglial cell proliferation was mediated in vitro by CCL2 along with miR-10 [100]. Neurotrophins have a potential role in modulating the proliferation and survival of microglial populations in vitro. Specifically, NGF and BDNF increased microglial proliferation, contrary to NT-3 and NT-4 [101]. Lastly, SCF was identified as a promoter of microglial cell proliferation, migration, and phagocytosis in culture (Table 1) [102].

Table 1.
Molecular drivers of microglial early differentiation, migration, proliferation, and terminal differentiation.
Summarizing, microgliogenesis is a complex biological process strictly regulated by multiple molecular drivers in a similar pattern to other CNS cells, such as oligodendrocytes [103].
4. Spatiotemporal Distribution in Various Species
In rodents, microglia were observed in the fetal forebrain at E11, when the telencephalic vesicles form [108]. Other studies identified E12 as the initial point of brain colonization [109,110]. Using in vivo immunohistochemistry and ex vivo time-lapse analysis of microglia, E11.5 was identified as the first day of the microglial entrance in the cortex [111]. The route includes in turn the pial surface, lateral ventricle, and cortical wall, moving over towards the cortical plate in the later embryonic phases. Three invasion phases in the cortical parenchyma have been proposed: (a) between E10.5 and E14.5, a gradual increase in the number of microglial cells takes place, succeeded by (b) a rapid phase with a significant rise in microglia from E14.5 to E15.5, followed by (c) the last slow wave of entry from E15.5 to E17.5. Before the invasion in the parenchyma, the peripheral microglia proliferates, especially at early phases [111]. Stremmel et al. demonstrated that, from E8.5, the CX3CR1+ pre-macrophages were detectable in the YS proliferating and preparing to enter the blood circulation for their migration to the brain parenchyma, while Kierdorf et al. suggested that E9.5 is the starting point for the migration of microglial progenitors into the neural tube [50]. The invading wave of YS progenitors to the tissue peaks around E10.5, then excessively decreases towards E12.5 and disappears at E14.5. Consequently, microglial progenitors are dependent on the vascular system for their migration [112]. Finally, the transformation of immature microglia into ramified, mature cells occurs between the second and third postnatal week (Figure 2) [113,114].
In humans, well-differentiated microglia were observed after 35 weeks of gestation (GW) [115]. However, Rezaie and Male suggested that colonization of the spinal cord starts around 9 GW, with the major influx of microglial cell populations estimated around 16 GW. In the second trimester, the cerebrum is colonized by microglial populations widely dispersed in the intermediate zone at 20–22 GW [116]. In the initial phase of microglial colonization between 12 and 14 GW, two cell populations were identified by Rezaie et al., namely CD68++ RCA-1+ MHC II- amoeboid cells located in the subplate and RCA-1++ CD68- MHC II- progenitors first observed in the marginal layer and lower cortical plate and which ramified within the subplate [117]. In 2006, the first intracerebral microglial populations were described close to the meninges and choroid plexus, next to the di-telencephalic fissure at 5.5 GW, whereas the cortical anlagen was populated with cells starting at 10–12 GW [118]. Routes of entry were found to be different for the cerebral cortex compared with the diencephalon. Microglial cells invaded the cerebrum from the ventricular lumen and the leptomeninges, starting at 4.5 GW. From 12 GW, the intraparenchymal vascular route of entry could be determined [119]. In 2010, Verney et al. suggested that the invasion of amoeboid microglia occurred between 4.5 and 5.5 GW into the human forebrain; this is in accordance with the data from other animal models such as rodents, regarding the spatiotemporal patterns observed for microglial development. Ultimately, the meninges, choroid plexus, and ventricles were identified as the three early routes of microglial entry [120].
In avians, the first microglial population was found to be located within the brain independently of vascularization, reaching the nervous system parenchyma by passing through the pial basal lamina [121]. More specifically, before E9, the cerebellar anlage contained only a small number of microglial precursors. Microglial precursors cross the pial surface at the basal region of the peduncles to enter the cerebellar anlage. Then, microglia proceed radially to the various cortical layers by migrating via the white matter. Following the ultimate settlement of microglial cells, differentiation then ensues [122].
5. Proliferation in the Adult Compromised CNS
As the BBB and microglial cell maturation are established, the question arises as to how microglia are renewed in the adult brain. The participation of bone marrow-derived cells in the repopulation of microglial cell niches was proposed in various conditions, especially after bone marrow transplantation [123,124,125,126,127,128], and in diseases such as stroke [129], cerebral ischemia [130], bacterial meningitis [131], entorhinal cortex lesions [132], Parkinson’s disease [133], Alzheimer’s disease [134,135], multiple sclerosis [136], facial nerve axotomy and autoimmune encephalitis [137], scrapie [138], and brain and peripheral nerve injury [139,140,141]. During aging and the transition from plasticity to proinflammatory activation in primary neurodegeneration, the latest data also suggest that many metabolic byproducts and mitochondrial components can serve as damage-associated molecules, creating an extracellular gradient and accumulation of reactive oxygen species, which in turn propagate the inflammatory neurodegeneration [142,143]. Under acute situations such as when a stab wound inflicts damage to a brain region, the resident microglia need the contribution of circulating monocytes to efficiently respond to the extra load of detritus [144]. It has been suggested that even after recovering from severe brain inflammation, resident microglia form a remarkably stable cell pool that is seldom replenished by hematogenous cells in adult animals [145].
A physiological process that aids in the development of the adult microglial cell population is the proliferation of microglial precursors in the developing brain [146]. Lawson et al. suggested that resident microglia synthesize DNA and go on to divide in situ. Additionally, cells were found to be recruited from the circulating monocyte pool through an intact BBB and rapidly differentiated into resident microglia. These two processes contributed almost equally to the steady-state turnover of resident microglia [147]. In a mouse model of ALS, the local proliferation of resident microglia had the greatest contribution to the observing microgliosis, while the effects of bone marrow-derived cells were limited among the microglia populations [148]. Strong evidence for the local self-renewal of CNS microglia as the main source of repopulation of adult microglia were obtained from a model using chimeric animals obtained by parabiosis showing that these cells could be maintained independently from bone marrow–derived cells during adulthood in ALS and facial nerve axotomy [149]. However, Ly-6ChiCCR2+ monocytes were found to be recruited to the lesioned brain differentiating into mature microglia. Remarkably, monocyte invasion during CNS pathology with an intact BBB or in non-diseased adult CNS required previous conditioning of brains, such as direct tissue irradiation [150]. Indeed, brain conditioning with lethal irradiation and alkylating agents such as busulfan was found to be vital for an efficient microglial cell repopulation after hematopoietic stem cell transplantation [151].
In 2013, Li et al. observed that after ischemic stroke, a small number of blood-derived CX3CR1GFP/+ cells invaded the brain parenchyma; however, these cells were phenotypically different from resident microglia with distinct kinetics. This study delineated the greatest impact of local resident microglia on the repopulation of parenchymal cells compared to recruited blood-derived cells after ischemic stroke [152]. The efficiency of microglia for self-renewal arising from a CNS-resident pool independently from peripheral myeloid cells was also supported by another experimental study that investigated the repopulation of brain parenchyma using a model of conditional depletion of microglial cells [153]. During the process of cellular restoration, the proliferation of local microglia was found to be dependent on the IL-1 receptor, which was highly expressed by local cell pools. Bone-marrow-derived macrophages populated the brain only after irradiation and bone marrow transplantation, and did not express the IL-1 receptor [153].
In zebrafish, using temporal-spatial resolution fate mapping analysis, embryonic microglia emerged from the rostral blood island in a RUNX1-independent and PU.1-dependent manner, while adult microglia originated from the ventral wall of the dorsal aorta in a RUNX1-dependent, c-MYB- and PU.1-independent manner [154]. The microglial self-renewal was shown to resemble a stochastic process at steady state, whereas clonal microglial expansion seems to predominate under unilateral facial nerve axotomy [155]. In another study, the partial microglial depletion resulted in the engraftment of peripherally derived macrophages independently of irradiation. These newly-engrafted cell populations differ transcriptionally from microglia [156]. Similarly, another depletion study showed that the microglial niche is filled with new cells via local proliferation of CX3CR1+F4/80lowClec12a– microglia and invasion of CX3CR1+F4/80hiClec12a+ macrophages derived from Ly6Chi monocytes. This engraftment was associated with vascular activation and type I interferon, while it was shown to be independent of BBB integrity [157]. These peripherally engrafted cells were transcriptionally distinct from microglia, showcasing different surface marker expression, phagocytic capacity, and cytokine release [157,158].
Through additional studies, Huang et al. delineated that repopulated microglial cell populations are entirely generated from residual microglial proliferation after acute depletion [159], instead of nestin-expressing progenitors, as was argued in a CSF1R inhibitor-mediated experiment [160]. In agreement with the previous statement, Zhan et al. demonstrated that after acute ablation, the newborn adult microglia generated via self-renewal from the local CX3CR1+ microglia without any contribution of nestin+ progenitors or peripheral myeloid cells. The repopulated microglia formed stable and distinct clusters with minimum migration capacity via clonal expansion. Although these regenerated microglial cells were presented in an immature state, microglial differentiation was mediated by NF-κB and interferon pathways [161]. A fate mapping study from Chen et al. showed that after neonatal stroke, a monocyte-to-microglia transition is possible [57]. In contrast, a study conducted in 2021 showed that microglia are not replaced by bone-marrow-derived cells in Alzheimer’s disease similar to the BAMs, which seldom replenished the microglial cell pool [162]. Ultimately, microglial cell manipulation is being intensely investigated in the context of immune-mediated diseases such as multiple sclerosis, where microglia are heavily implicated as pathogenic mediators of progressive disease [163,164,165], and targeted therapies are being developed [166,167].
Summarizing the results of the above studies, it is postulated that the greatest contribution to microglial repopulation is based upon its local self-renewal, both in steady state and disease. However, circulating monocytes may also contribute to a lesser extent, especially in disease. The final confirmation of the exact repopulation pattern necessitates further investigation.
6. Conclusions
The widely accepted, contemporary view of the origin of CNS-resident microglia is the YS. However, this was hotly debated until the early 2010s. Although this may pass unnoticed to the majority of the research community in the immune-related neuroscience field, understanding the underlying molecular development they undergo during embryogenesis may aid towards developing novel therapies that ideally could decelerate, halt, or reverse neurodegeneration by targeting the microglia-mediated repair process. A main challenge now is to elucidate the precise biological identity of each different microglial state as well as the variable microglial activity per CNS region, allowing us to perform selective interventions. Another field of application that can potentially benefit from relevant developmental research is aging, where mechanisms implicating microgliogenesis can be exploited in favor of slowing the senescent progress by, e.g., combating oxidative stress. Finally, the understanding of cellular ontogeny may enable successful lab-approached manipulations aimed at depletion of microglial cells and beneficial microglial renewal in the CNS, in both homeostasis and disorders.
Author Contributions
I.D. and P.T., writing—original draft preparation and visualization; M.E.M., S.M., M.-È.T., S.P. and L.Z., conceptualization, review and editing; M.B., E.K. and C.S., review and editing. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Conflicts of Interest
The authors declare no conflict of interest.
Abbreviations
| AGM | Aorta-gonad-mesonephros |
| Akt | Protein kinase B |
| ALS | Amyotrophic lateral sclerosis |
| ARID1A | AT-rich interaction domain 1A |
| BAMs | Border-associated macrophages |
| BATF3 | Basic leucine zipper transcriptional factor ATF-like 3 |
| BBB | Blood–brain barrier |
| BDNF | Brain-derived neurotrophic factor |
| C/EBP-α | CCAAT/enhancer-binding protein alpha |
| CCL1 | Chemokine (C-C motif) ligand 1 |
| CCL2 | Chemokine (C-C motif) ligand 2 |
| CCL3 | Chemokine (C-C motif) ligand 3 |
| CCR2 | C-C chemokine receptor type 2 |
| CD11b | Cluster of differentiation molecule 11B |
| CD206 | Cluster of differentiation molecule 206 |
| CD45 | Cluster of differentiation molecule 45 |
| CD64 | Cluster of differentiation molecule 64 |
| CD68 | Cluster of differentiation molecule 68 |
| CNS | Central nervous system |
| CSF-1 | Colony stimulating factor-1 |
| CSF1R | Colony stimulating factor 1 receptor |
| CX3CL1 | Chemokine (C-X3-C motif) ligand 1 |
| CX3CR1 | CX3C motif chemokine receptor 1 |
| CXCL12 | C-X-C motif chemokine ligand 12 |
| CXCR4 | C-X-C chemokine receptor type 4 |
| CXCR7 | C-X-C chemokine receptor type 7 |
| DAP12 | DNAX-activating protein of 12 kDa |
| E | Embryonic day |
| EED | Embryonic ectoderm development |
| EMPs | Erythro-myeloid progenitors |
| Erk1/2 | Extracellular signal-regulated kinase 1/2 |
| G-CSF | Granulocyte colony stimulating factor |
| G-CSFR | Granulocyte colony stimulating factor receptor |
| GM-CSF | Granulocyte-macrophage colony-stimulating factor |
| GM-CSFR | Granulocyte-macrophage colony-stimulating factor receptor |
| GSK3β | Glycogen synthase kinase-3 beta |
| GW | Gestational week |
| HDAC1 | Histone Deacetylase 1 |
| HDAC2 | Histone Deacetylase 2 |
| HOXB8 | Homeobox B8 |
| HSC | Hematopoietic stem cells |
| IBA1 | Ionized calcium binding adaptor molecule 1 |
| ID2 | Inhibitor of DNA binding 2 |
| IL-33 | Interleukin 33 |
| IL-34 | Interleukin 34 |
| IL-5 | Interleukin 5 |
| iNOS | Inducible nitric oxide synthase |
| IRF8 | Interferon regulatory factor 8 |
| KLF4 | Krüppel-like factor 4 |
| M-CSF | Macrophage colony-stimulating factor |
| M-CSFR | Macrophage colony-stimulating factor receptor |
| MAFB | MAF bZIP transcription factor B |
| MCP-1 | Monocyte chemoattractant protein-1 |
| MECP2 | Methyl-CpG binding protein 2 |
| MEIS3 | Meis homeobox 3 |
| MHC II | Major histocompatibility complex class II |
| MIF | Macrophage migration inhibitory factor |
| MIP-1α | Macrophage inflammatory protein-1 alpha |
| miR-124 | microRNA 124 |
| MMP8 | Matrix Metallopeptidase 8 |
| MMP9 | Matrix Metallopeptidase 9 |
| MPO | Myeloperoxidase |
| MYB | MYB proto-oncogene, transcription factor |
| NF-kB | Nuclear factor kappa B |
| NGF | Nerve growth factor |
| NOX2 | NADPH oxidase 2 |
| NR4A1 | Nuclear receptor subfamily 4 group A member 1 |
| NT-3 | Neurotrophin-3 |
| NT-4 | Neurotrophin-4 |
| P2RX7 | P2X purinoceptor 7 |
| P2RY12 | Purinergic receptor P2Y12 |
| PGRN | Progranulin |
| qRT-PCR | Quantitative reverse transcriptase polymerase chain reaction |
| RCA-1 | Ricinus communis agglutinin-1 |
| RUNX1 | Runt-related transcription factor 1 |
| SALL1 | Spalt like transcription factor 1 |
| SALL3 | Spalt like transcription factor 3 |
| SCF | Stem cell factor |
| SVZ | Subventricular zone |
| TGF-β1 | Transforming growth factor beta 1 |
| TGM2 | Transglutaminase 2 |
| TMEM119 | Transmembrane Protein 119 |
| TREM2 | Triggering receptor expressed on myeloid cells 2 |
| VEGFR1 | Vascular endothelial growth factor receptor-1 |
| YS | Yolk sac |
| αVβ8 integrin | Integrin subunit alpha V and beta 8 |
References
- Davies, L.C.; Jenkins, S.J.; Allen, J.E.; Taylor, P.R. Tissue-Resident Macrophages. Nat. Immunol. 2013, 14, 986–995. [Google Scholar] [CrossRef] [PubMed]
- Soulet, D.; Rivest, S. Microglia. Curr. Biol. CB 2008, 18, R506–R508. [Google Scholar] [CrossRef] [PubMed]
- Lee, E.; Eo, J.-C.; Lee, C.; Yu, J.-W. Distinct Features of Brain-Resident Macrophages: Microglia and Non-Parenchymal Brain Macrophages. Mol. Cells 2021, 44, 281–291. [Google Scholar] [CrossRef] [PubMed]
- Dos Santos, S.E.; Medeiros, M.; Porfirio, J.; Tavares, W.; Pessôa, L.; Grinberg, L.; Leite, R.E.P.; Ferretti-Rebustini, R.E.L.; Suemoto, C.K.; Filho, W.J.; et al. Similar Microglial Cell Densities across Brain Structures and Mammalian Species: Implications for Brain Tissue Function. J. Neurosci. 2020, 40, 4622–4643. [Google Scholar] [CrossRef] [PubMed]
- Paolicelli, R.C.; Sierra, A.; Stevens, B.; Tremblay, M.-E.; Aguzzi, A.; Ajami, B.; Amit, I.; Audinat, E.; Bechmann, I.; Bennett, M.; et al. Microglia States and Nomenclature: A Field at Its Crossroads. Neuron 2022, 110, 3458–3483. [Google Scholar] [CrossRef]
- Lenz, K.M.; Nelson, L.H. Microglia and Beyond: Innate Immune Cells As Regulators of Brain Development and Behavioral Function. Front. Immunol. 2018, 9, 698. [Google Scholar] [CrossRef]
- Tremblay, M.-È.; Stevens, B.; Sierra, A.; Wake, H.; Bessis, A.; Nimmerjahn, A. The Role of Microglia in the Healthy Brain. J. Neurosci. 2011, 31, 16064–16069. [Google Scholar] [CrossRef]
- Neurons and Neuroglia-ClinicalKey. Available online: https://www.clinicalkey.com/#!/content/book/3-s2.0-B978032328782100438X (accessed on 28 December 2022).
- Chen, Z.; Trapp, B.D. Microglia and Neuroprotection. J. Neurochem. 2016, 136 (Suppl. 1), 10–17. [Google Scholar] [CrossRef]
- Bachiller, S.; Jiménez-Ferrer, I.; Paulus, A.; Yang, Y.; Swanberg, M.; Deierborg, T.; Boza-Serrano, A. Microglia in Neurological Diseases: A Road Map to Brain-Disease Dependent-Inflammatory Response. Front. Cell. Neurosci. 2018, 12, 488. [Google Scholar] [CrossRef]
- Guerrero, B.L.; Sicotte, N.L. Microglia in Multiple Sclerosis: Friend or Foe? Front. Immunol. 2020, 11, 374. [Google Scholar] [CrossRef]
- Clarke, B.E.; Patani, R. The Microglial Component of Amyotrophic Lateral Sclerosis. Brain J. Neurol. 2020, 143, 3526–3539. [Google Scholar] [CrossRef]
- Sierra, A.; de Castro, F.; Del Río-Hortega, J.; Rafael Iglesias-Rozas, J.; Garrosa, M.; Kettenmann, H. The “Big-Bang” for Modern Glial Biology: Translation and Comments on Pío Del Río-Hortega 1919 Series of Papers on Microglia. Glia 2016, 64, 1801–1840. [Google Scholar] [CrossRef]
- Colonna, M.; Butovsky, O. Microglia Function in the Central Nervous System During Health and Neurodegeneration. Annu. Rev. Immunol. 2017, 35, 441–468. [Google Scholar] [CrossRef]
- Sohet, F.; Lin, C.; Munji, R.N.; Lee, S.Y.; Ruderisch, N.; Soung, A.; Arnold, T.D.; Derugin, N.; Vexler, Z.S.; Yen, F.T.; et al. LSR/Angulin-1 Is a Tricellular Tight Junction Protein Involved in Blood–Brain Barrier Formation. J. Cell Biol. 2015, 208, 703–711. [Google Scholar] [CrossRef]
- Hoeffel, G.; Ginhoux, F. Ontogeny of Tissue-Resident Macrophages. Front. Immunol. 2015, 6, 486. [Google Scholar] [CrossRef]
- Wu, Y.; Hirschi, K.K. Tissue-Resident Macrophage Development and Function. Front. Cell Dev. Biol. 2020, 8, 617879. [Google Scholar] [CrossRef]
- Vink, C.S.; Mariani, S.A.; Dzierzak, E. Embryonic Origins of the Hematopoietic System: Hierarchies and Heterogeneity. HemaSphere 2022, 6, e737. [Google Scholar] [CrossRef]
- Yoder, M.C. Inducing Definitive Hematopoiesis in a Dish. Nat. Biotechnol. 2014, 32, 539–541. [Google Scholar] [CrossRef]
- Ling, E.A. Transformation of Monocytes into Amoeboid Microglia in the Corpus Callosum of Postnatal Rats, as Shown by Labelling Monocytes by Carbon Particles. J. Anat. 1979, 128, 847–858. [Google Scholar]
- Rio-Hortega, P. The Microglia. Lancet 1939, 233, 1023–1026. [Google Scholar] [CrossRef]
- Kershman, J. Genesis of Microglia in the Human Brain. Arch. Neurol. Psychiatry 1939, 41, 24–50. [Google Scholar] [CrossRef]
- Boya, J.; Calvo, J.; Prado, A. The Origin of Microglial Cells. J. Anat. 1979, 129, 177–186. [Google Scholar] [PubMed]
- Boya, J.; Calvo, J.L.; Carbonell, A.L.; Borregon, A. A Lectin Histochemistry Study on the Development of Rat Microglial Cells. J. Anat. 1991, 175, 229–236. [Google Scholar] [PubMed]
- Dalmau, I.; Finsen, B.; Tønder, N.; Zimmer, J.; González, B.; Castellano, B. Development of Microglia in the Prenatal Rat Hippocampus. J. Comp. Neurol. 1997, 377, 70–84. [Google Scholar] [CrossRef]
- Barón, M.; Gallego, A. The Relation of the Microglia with the Pericytes in the Cat Cerebral Cortex. Z. Zellforsch. Mikrosk. Anat. 1972, 128, 42–57. [Google Scholar] [CrossRef]
- Mori, S.; Leblond, C.P. Identification of Microglia in Light and Electron Microscopy. J. Comp. Neurol. 1969, 135, 57–80. [Google Scholar] [CrossRef]
- Juba, A. Untersuchungen Über Die Entwicklung Der Hortegaschen Microglia Des Menschen. Arch. Psychiat. Νervenkr. 1933, 101, 577–592. [Google Scholar] [CrossRef]
- Santha, K.; Juba, A. Weitere Untersuchungen Über Die Entwicklung Der Hortegaschen Mikroglia. Arch. Psychiat. Νervenkr. 1933, 98, 598–613. [Google Scholar] [CrossRef]
- Perry, V.H.; Hume, D.A.; Gordon, S. Immunohistochemical Localization of Macrophages and Microglia in the Adult and Developing Mouse Brain. Neuroscience 1985, 15, 313–326. [Google Scholar] [CrossRef]
- Ling, E.A.; Penney, D.; Leblond, C.P. Use of Carbon Labeling to Demonstrate the Role of Blood Monocytes as Precursors of the “ameboid Cells” Present in the Corpus Callosum of Postnatal Rats. J. Comp. Neurol. 1980, 193, 631–657. [Google Scholar] [CrossRef]
- Murabe, Y.; Sano, Y. Morphological Studies on Neuroglia. VII. Distribution of “Brain Macrophages” in Brains of Neonatal and Adult Rats, as Determined by Means of Immunohistochemistry. Cell Tissue Res. 1983, 229, 85–95. [Google Scholar] [CrossRef]
- Ling, E.A.; Kaur, C.; Wong, W.C. Expression of Major Histocompatibility Complex and Leukocyte Common Antigens in Amoeboid Microglia in Postnatal Rats. J. Anat. 1991, 177, 117–126. [Google Scholar]
- Oehmichen, M.; Wiethölter, H.; Greaves, M.F. Immunological Analysis of Human Microglia: Lack of Monocytic and Lymphoid Membrane Differentiation Antigens. J. Neuropathol. Exp. Neurol. 1979, 38, 99–103. [Google Scholar] [CrossRef]
- Lewis, P.D. The Fate of the Subependymal Cell in the Adult Rat Brain, with a Note on the Origin of Microglia. Brain J. Neurol. 1968, 91, 721–736. [Google Scholar] [CrossRef]
- Kitamura, T.; Miyake, T.; Fujita, S. Genesis of Resting Microglia in the Gray Matter of Mouse Hippocampus. J. Comp. Neurol. 1984, 226, 421–433. [Google Scholar] [CrossRef]
- Fedoroff, S.; Zhai, R.; Novak, J.P. Microglia and Astroglia Have a Common Progenitor Cell. J. Neurosci. Res. 1997, 50, 477–486. [Google Scholar] [CrossRef]
- de Groot, C.J.; Huppes, W.; Sminia, T.; Kraal, G.; Dijkstra, C.D. Determination of the Origin and Nature of Brain Macrophages and Microglial Cells in Mouse Central Nervous System, Using Non-Radioactive in Situ Hybridization and Immunoperoxidase Techniques. Glia 1992, 6, 301–309. [Google Scholar] [CrossRef]
- Schelper, R.L.; Adrian, E.K. Monocytes Become Macrophages; They Do Not Become Microglia: A Light and Electron Microscopic Autoradiographic Study Using 125-Iododeoxyuridine. J. Neuropathol. Exp. Neurol. 1986, 45, 1–19. [Google Scholar] [CrossRef]
- Matsumoto, Y.; Fujiwara, M. Absence of Donor-Type Major Histocompatibility Complex Class I Antigen-Bearing Microglia in the Rat Central Nervous System of Radiation Bone Marrow Chimeras. J. Neuroimmunol. 1987, 17, 71–82. [Google Scholar] [CrossRef]
- Hutchins, K.D.; Dickson, D.W.; Rashbaum, W.K.; Lyman, W.D. Localization of Morphologically Distinct Microglial Populations in the Developing Human Fetal Brain: Implications for Ontogeny. Brain Res. Dev. Brain Res. 1990, 55, 95–102. [Google Scholar] [CrossRef]
- Takahashi, K.; Yamamura, F.; Naito, M. Differentiation, Maturation, and Proliferation of Macrophages in the Mouse Yolk Sac: A Light-Microscopic, Enzyme-Cytochemical, Immunohistochemical, and Ultrastructural Study. J. Leukoc. Biol. 1989, 45, 87–96. [Google Scholar] [CrossRef] [PubMed]
- Fujimoto, E.; Miki, A.; Mizoguti, H. Histochemical Study of the Differentiation of Microglial Cells in the Developing Human Cerebral Hemispheres. J. Anat. 1989, 166, 253–264. [Google Scholar] [PubMed]
- Kurz, H.; Christ, B. Embryonic CNS Macrophages and Microglia Do Not Stem from Circulating, but from Extravascular Precursors. Glia 1998, 22, 98–102. [Google Scholar] [CrossRef]
- Andjelkovic, A.V.; Nikolic, B.; Pachter, J.S.; Zecevic, N. Macrophages/Microglial Cells in Human Central Nervous System during Development: An Immunohistochemical Study. Brain Res. 1998, 814, 13–25. [Google Scholar] [CrossRef]
- Alliot, F.; Godin, I.; Pessac, B. Microglia Derive from Progenitors, Originating from the Yolk Sac, and Which Proliferate in the Brain. Brain Res. Dev. Brain Res. 1999, 117, 145–152. [Google Scholar] [CrossRef]
- Ginhoux, F.; Greter, M.; Leboeuf, M.; Nandi, S.; See, P.; Gokhan, S.; Mehler, M.F.; Conway, S.J.; Ng, L.G.; Stanley, E.R.; et al. Fate Mapping Analysis Reveals That Adult Microglia Derive from Primitive Macrophages. Science 2010, 330, 841–845. [Google Scholar] [CrossRef]
- Mizutani, M.; Pino, P.A.; Saederup, N.; Charo, I.F.; Ransohoff, R.M.; Cardona, A.E. The Fractalkine Receptor but Not CCR2 Is Present on Microglia from Embryonic Development throughout Adulthood. J. Immunol. 2012, 188, 29–36. [Google Scholar] [CrossRef]
- Schulz, C.; Gomez Perdiguero, E.; Chorro, L.; Szabo-Rogers, H.; Cagnard, N.; Kierdorf, K.; Prinz, M.; Wu, B.; Jacobsen, S.E.W.; Pollard, J.W.; et al. A Lineage of Myeloid Cells Independent of Myb and Hematopoietic Stem Cells. Science 2012, 336, 86–90. [Google Scholar] [CrossRef]
- Kierdorf, K.; Erny, D.; Goldmann, T.; Sander, V.; Schulz, C.; Perdiguero, E.G.; Wieghofer, P.; Heinrich, A.; Riemke, P.; Hölscher, C.; et al. Microglia Emerge from Erythromyeloid Precursors via Pu.1- and Irf8-Dependent Pathways. Nat. Neurosci. 2013, 16, 273–280. [Google Scholar] [CrossRef]
- Hoeffel, G.; Chen, J.; Lavin, Y.; Low, D.; Almeida, F.F.; See, P.; Beaudin, A.E.; Lum, J.; Low, I.; Forsberg, E.C.; et al. C-Myb(+) Erythro-Myeloid Progenitor-Derived Fetal Monocytes Give Rise to Adult Tissue-Resident Macrophages. Immunity 2015, 42, 665–678. [Google Scholar] [CrossRef]
- Gomez Perdiguero, E.; Klapproth, K.; Schulz, C.; Busch, K.; Azzoni, E.; Crozet, L.; Garner, H.; Trouillet, C.; de Bruijn, M.F.; Geissmann, F.; et al. Tissue-Resident Macrophages Originate from Yolk-Sac-Derived Erythro-Myeloid Progenitors. Nature 2015, 518, 547–551. [Google Scholar] [CrossRef]
- Sheng, J.; Ruedl, C.; Karjalainen, K. Most Tissue-Resident Macrophages Except Microglia Are Derived from Fetal Hematopoietic Stem Cells. Immunity 2015, 43, 382–393. [Google Scholar] [CrossRef]
- De, S.; Van Deren, D.; Peden, E.; Hockin, M.; Boulet, A.; Titen, S.; Capecchi, M.R. Two Distinct Ontogenies Confer Heterogeneity to Mouse Brain Microglia. Dev. Camb. Engl. 2018, 145, dev152306. [Google Scholar] [CrossRef]
- Fehrenbach, M.K.; Tjwa, M.; Bechmann, I.; Krueger, M. Decreased Microglial Numbers in Vav1-Cre+:Dicer Knock-out Mice Suggest a Second Source of Microglia beyond Yolk Sac Macrophages. Ann. Anat. Anat. Anz. 2018, 218, 190–198. [Google Scholar] [CrossRef]
- Utz, S.G.; See, P.; Mildenberger, W.; Thion, M.S.; Silvin, A.; Lutz, M.; Ingelfinger, F.; Rayan, N.A.; Lelios, I.; Buttgereit, A.; et al. Early Fate Defines Microglia and Non-Parenchymal Brain Macrophage Development. Cell 2020, 181, 557–573.e18. [Google Scholar] [CrossRef]
- Chen, H.-R.; Sun, Y.-Y.; Chen, C.-W.; Kuo, Y.-M.; Kuan, I.S.; Li, Z.-R.T.; Short-Miller, J.C.; Smucker, M.R.; Kuan, C.-Y. Fate Mapping via CCR2-CreER Mice Reveals Monocyte-to-Microglia Transition in Development and Neonatal Stroke. Sci. Adv. 2020, 6, eabb2119. [Google Scholar] [CrossRef]
- Bian, Z.; Gong, Y.; Huang, T.; Lee, C.Z.W.; Bian, L.; Bai, Z.; Shi, H.; Zeng, Y.; Liu, C.; He, J.; et al. Deciphering Human Macrophage Development at Single-Cell Resolution. Nature 2020, 582, 571–576. [Google Scholar] [CrossRef]
- Zusso, M.; Methot, L.; Lo, R.; Greenhalgh, A.D.; David, S.; Stifani, S. Regulation of Postnatal Forebrain Amoeboid Microglial Cell Proliferation and Development by the Transcription Factor Runx1. J. Neurosci. 2012, 32, 11285–11298. [Google Scholar] [CrossRef]
- Huang, G.; Zhang, P.; Hirai, H.; Elf, S.; Yan, X.; Chen, Z.; Koschmieder, S.; Okuno, Y.; Dayaram, T.; Growney, J.D.; et al. PU. 1 Is a Major Downstream Target of AML1 (RUNX1) in Adult Mouse Hematopoiesis. Nat. Genet. 2008, 40, 51–60. [Google Scholar] [CrossRef]
- Goldmann, T.; Wieghofer, P.; Jordão, M.J.C.; Prutek, F.; Hagemeyer, N.; Frenzel, K.; Amann, L.; Staszewski, O.; Kierdorf, K.; Krueger, M.; et al. Origin, Fate and Dynamics of Macrophages at Central Nervous System Interfaces. Nat. Immunol. 2016, 17, 797–805. [Google Scholar] [CrossRef]
- Olson, M.C.; Scott, E.W.; Hack, A.A.; Su, G.H.; Tenen, D.G.; Singh, H.; Simon, M.C. PU. 1 Is Not Essential for Early Myeloid Gene Expression but Is Required for Terminal Myeloid Differentiation. Immunity 1995, 3, 703–714. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Xue, R.; Hassan, S.; Nguyen, T.M.L.; Wang, T.; Pan, H.; Xu, J.; Liu, Q.; Zhang, W.; Wen, Z. Il34-Csf1r Pathway Regulates the Migration and Colonization of Microglial Precursors. Dev. Cell 2018, 46, 552–563.e4. [Google Scholar] [CrossRef] [PubMed]
- Buttgereit, A.; Lelios, I.; Yu, X.; Vrohlings, M.; Krakoski, N.R.; Gautier, E.L.; Nishinakamura, R.; Becher, B.; Greter, M. Sall1 Is a Transcriptional Regulator Defining Microglia Identity and Function. Nat. Immunol. 2016, 17, 1397–1406. [Google Scholar] [CrossRef] [PubMed]
- Butovsky, O.; Jedrychowski, M.P.; Moore, C.S.; Cialic, R.; Lanser, A.J.; Gabriely, G.; Koeglsperger, T.; Dake, B.; Wu, P.M.; Doykan, C.E.; et al. Identification of a Unique TGF-β–Dependent Molecular and Functional Signature in Microglia. Nat. Neurosci. 2014, 17, 131–143. [Google Scholar] [CrossRef] [PubMed]
- Mass, E.; Ballesteros, I.; Farlik, M.; Halbritter, F.; Günther, P.; Crozet, L.; Jacome-Galarza, C.E.; Händler, K.; Klughammer, J.; Kobayashi, Y.; et al. Specification of Tissue-Resident Macrophages during Organogenesis. Science 2016, 353, aaf4238. [Google Scholar] [CrossRef]
- Nataf, S.; Anginot, A.; Vuaillat, C.; Malaval, L.; Fodil, N.; Chereul, E.; Langlois, J.-B.; Dumontel, C.; Cavillon, G.; Confavreux, C.; et al. Brain and Bone Damage in KARAP/DAP12 Loss-of-Function Mice Correlate with Alterations in Microglia and Osteoclast Lineages. Am. J. Pathol. 2005, 166, 275–286. [Google Scholar] [CrossRef]
- Otero, K.; Turnbull, I.R.; Poliani, P.L.; Vermi, W.; Cerutti, E.; Aoshi, T.; Tassi, I.; Takai, T.; Stanley, S.L.; Miller, M.; et al. Macrophage Colony-Stimulating Factor Induces the Proliferation and Survival of Macrophages via a Pathway Involving DAP12 and Beta-Catenin. Nat. Immunol. 2009, 10, 734–743. [Google Scholar] [CrossRef]
- Lelli, A.; Gervais, A.; Colin, C.; Chéret, C.; de Almodovar, C.R.; Carmeliet, P.; Krause, K.-H.; Boillée, S.; Mallat, M. The NADPH Oxidase Nox2 Regulates VEGFR1/CSF-1R-Mediated Microglial Chemotaxis and Promotes Early Postnatal Infiltration of Phagocytes in the Subventricular Zone of the Mouse Cerebral Cortex. Glia 2013, 61, 1542–1555. [Google Scholar] [CrossRef]
- Arnò, B.; Grassivaro, F.; Rossi, C.; Bergamaschi, A.; Castiglioni, V.; Furlan, R.; Greter, M.; Favaro, R.; Comi, G.; Becher, B.; et al. Neural Progenitor Cells Orchestrate Microglia Migration and Positioning into the Developing Cortex. Nat. Commun. 2014, 5, 5611. [Google Scholar] [CrossRef]
- Lipfert, J.; Ödemis, V.; Wagner, D.-C.; Boltze, J.; Engele, J. CXCR4 and CXCR7 Form a Functional Receptor Unit for SDF-1/CXCL12 in Primary Rodent Microglia. Neuropathol. Appl. Neurobiol. 2013, 39, 667–680. [Google Scholar] [CrossRef]
- Hoshiko, M.; Arnoux, I.; Avignone, E.; Yamamoto, N.; Audinat, E. Deficiency of the Microglial Receptor CX3CR1 Impairs Postnatal Functional Development of Thalamocortical Synapses in the Barrel Cortex. J. Neurosci. 2012, 32, 15106–15111. [Google Scholar] [CrossRef]
- Rezaie, P.; Trillo-Pazos, G.; Greenwood, J.; Everall, I.P.; Male, D.K. Motility and Ramification of Human Fetal Microglia in Culture: An Investigation Using Time-Lapse Video Microscopy and Image Analysis. Exp. Cell Res. 2002, 274, 68–82. [Google Scholar] [CrossRef]
- Ponomarev, E.D.; Veremeyko, T.; Barteneva, N.; Krichevsky, A.M.; Weiner, H.L. MicroRNA-124 Promotes Microglia Quiescence and Suppresses EAE by Deactivating Macrophages via the C/EBP-α-PU.1 Pathway. Nat. Med. 2011, 17, 64–70. [Google Scholar] [CrossRef]
- Gong, M.; Shi, R.; Liu, Y.; Ke, J.; Liu, X.; Du, H.-Z.; Liu, C.-M. Abnormal Microglial Polarization Induced by Arid1a Deletion Leads to Neuronal Differentiation Deficits. Cell Prolif. 2022, 55, e13314. [Google Scholar] [CrossRef]
- Su, L.; Zhang, M.; Ji, F.; Zhao, J.; Wang, Y.; Wang, W.; Zhang, S.; Ma, H.; Wang, Y.; Jiao, J. Microglia Homeostasis Mediated by Epigenetic ARID1A Regulates Neural Progenitor Cells Response and Leads to Autism-like Behaviors. Mol. Psychiatry 2022, 5, 1–5. [Google Scholar] [CrossRef]
- Walsh, C.E.; Hitchcock, P.F. Progranulin Regulates Neurogenesis in the Developing Vertebrate Retina. Dev. Neurobiol. 2017, 77, 1114–1129. [Google Scholar] [CrossRef]
- Arnold, T.D.; Lizama, C.O.; Cautivo, K.M.; Santander, N.; Lin, L.; Qiu, H.; Huang, E.J.; Liu, C.; Mukouyama, Y.; Reichardt, L.F.; et al. Impaired AVβ8 and TGFβ Signaling Lead to Microglial Dysmaturation and Neuromotor Dysfunction. J. Exp. Med. 2019, 216, 900–915. [Google Scholar] [CrossRef]
- Datta, M.; Staszewski, O.; Raschi, E.; Frosch, M.; Hagemeyer, N.; Tay, T.L.; Blank, T.; Kreutzfeldt, M.; Merkler, D.; Ziegler-Waldkirch, S.; et al. Histone Deacetylases 1 and 2 Regulate Microglia Function during Development, Homeostasis, and Neurodegeneration in a Context-Dependent Manner. Immunity 2018, 48, 514–529.e6. [Google Scholar] [CrossRef]
- Jurga, A.M.; Paleczna, M.; Kuter, K.Z. Overview of General and Discriminating Markers of Differential Microglia Phenotypes. Front. Cell. Neurosci. 2020, 14, 198. [Google Scholar] [CrossRef]
- Bohnert, S.; Seiffert, A.; Trella, S.; Bohnert, M.; Distel, L.; Ondruschka, B.; Monoranu, C.-M. TMEM119 as a Specific Marker of Microglia Reaction in Traumatic Brain Injury in Postmortem Examination. Int. J. Leg. Med. 2020, 134, 2167–2176. [Google Scholar] [CrossRef]
- van Wageningen, T.A.; Vlaar, E.; Kooij, G.; Jongenelen, C.A.M.; Geurts, J.J.G.; van Dam, A.-M. Regulation of Microglial TMEM119 and P2RY12 Immunoreactivity in Multiple Sclerosis White and Grey Matter Lesions Is Dependent on Their Inflammatory Environment. Acta Neuropathol. Commun. 2019, 7, 206. [Google Scholar] [CrossRef] [PubMed]
- Vankriekelsvenne, E.; Chrzanowski, U.; Manzhula, K.; Greiner, T.; Wree, A.; Hawlitschka, A.; Llovera, G.; Zhan, J.; Joost, S.; Schmitz, C.; et al. Transmembrane Protein 119 Is Neither a Specific nor a Reliable Marker for Microglia. Glia 2022, 70, 1170–1190. [Google Scholar] [CrossRef] [PubMed]
- Konishi, H.; Kobayashi, M.; Kunisawa, T.; Imai, K.; Sayo, A.; Malissen, B.; Crocker, P.R.; Sato, K.; Kiyama, H. Siglec-H Is a Microglia-Specific Marker That Discriminates Microglia from CNS-Associated Macrophages and CNS-Infiltrating Monocytes. Glia 2017, 65, 1927–1943. [Google Scholar] [CrossRef] [PubMed]
- Masuda, T.; Amann, L.; Sankowski, R.; Staszewski, O.; Lenz, M.; Errico, P.D.; Snaidero, N.; Costa Jordão, M.J.; Böttcher, C.; Kierdorf, K.; et al. Novel Hexb-Based Tools for Studying Microglia in the CNS. Nat. Immunol. 2020, 21, 802–815. [Google Scholar] [CrossRef] [PubMed]
- Bennett, M.L.; Bennett, F.C.; Liddelow, S.A.; Ajami, B.; Zamanian, J.L.; Fernhoff, N.B.; Mulinyawe, S.B.; Bohlen, C.J.; Adil, A.; Tucker, A.; et al. New Tools for Studying Microglia in the Mouse and Human CNS. Proc. Natl. Acad. Sci. USA 2016, 113, E1738–E1746. [Google Scholar] [CrossRef]
- Butovsky, O.; Siddiqui, S.; Gabriely, G.; Lanser, A.J.; Dake, B.; Murugaiyan, G.; Doykan, C.E.; Wu, P.M.; Gali, R.R.; Iyer, L.K.; et al. Modulating Inflammatory Monocytes with a Unique MicroRNA Gene Signature Ameliorates Murine ALS. J. Clin. Investig. 2012, 122, 3063–3087. [Google Scholar] [CrossRef]
- Zheng, H.; Jia, L.; Liu, C.-C.; Rong, Z.; Zhong, L.; Yang, L.; Chen, X.-F.; Fryer, J.D.; Wang, X.; Zhang, Y.-W.; et al. TREM2 Promotes Microglial Survival by Activating Wnt/β-Catenin Pathway. J. Neurosci. 2017, 37, 1772–1784. [Google Scholar] [CrossRef]
- Matcovitch-Natan, O.; Winter, D.R.; Giladi, A.; Vargas Aguilar, S.; Spinrad, A.; Sarrazin, S.; Ben-Yehuda, H.; David, E.; Zelada González, F.; Perrin, P.; et al. Microglia Development Follows a Stepwise Program to Regulate Brain Homeostasis. Science 2016, 353, aad8670. [Google Scholar] [CrossRef]
- Wittrahm, R.; Takalo, M.; Marttinen, M.; Kuulasmaa, T.; Mäkinen, P.; Kemppainen, S.; Martiskainen, H.; Rauramaa, T.; Pike, I.; Leinonen, V.; et al. MECP2 Increases the Pro-Inflammatory Response of Microglial Cells and Phosphorylation at Serine 423 Regulates Neuronal Gene Expression upon Neuroinflammation. Cells 2021, 10, 860. [Google Scholar] [CrossRef]
- Cronk, J.C.; Derecki, N.C.; Ji, E.; Xu, Y.; Lampano, A.E.; Smirnov, I.; Baker, W.; Norris, G.T.; Marin, I.; Coddington, N.; et al. Methyl-CpG Binding Protein 2 Regulates Microglia and Macrophage Gene Expression in Response to Inflammatory Stimuli. Immunity 2015, 42, 679–691. [Google Scholar] [CrossRef]
- Wang, Y.-Y.; Deng, Y.-S.; Dai, S.-K.; Mi, T.-W.; Li, R.-Y.; Liu, P.-P.; Liu, C.; He, B.-D.; He, X.-C.; Du, H.-Z.; et al. Loss of Microglial EED Impairs Synapse Density, Learning, and Memory. Mol. Psychiatry 2022, 27, 2999–3009. [Google Scholar] [CrossRef]
- Liu, C.; Gao, X.; Shi, R.-X.; Wang, Y.-Y.; He, X.-C.; Du, H.-Z.; Hu, B.; Jiao, J.; Liu, C.-M.; Teng, Z.-Q. Microglial Transglutaminase 2 Deficiency Causes Impaired Synaptic Remodelling and Cognitive Deficits in Mice. Cell Prolif. 2023, e13439. [Google Scholar] [CrossRef]
- Rigato, C.; Swinnen, N.; Buckinx, R.; Couillin, I.; Mangin, J.-M.; Rigo, J.-M.; Legendre, P.; Le Corronc, H. Microglia Proliferation Is Controlled by P2X7 Receptors in a Pannexin-1-Independent Manner during Early Embryonic Spinal Cord Invasion. J. Neurosci. 2012, 32, 11559–11573. [Google Scholar] [CrossRef]
- Yasuoka, S.; Kawanokuchi, J.; Parajuli, B.; Jin, S.; Doi, Y.; Noda, M.; Sonobe, Y.; Takeuchi, H.; Mizuno, T.; Suzumura, A. Production and Functions of IL-33 in the Central Nervous System. Brain Res. 2011, 1385, 8–17. [Google Scholar] [CrossRef]
- Bartolini, A.; Vigliani, M.-C.; Magrassi, L.; Vercelli, A.; Rossi, F. G-CSF Administration to Adult Mice Stimulates the Proliferation of Microglia but Does Not Modify the Outcome of Ischemic Injury. Neurobiol. Dis. 2011, 41, 640–649. [Google Scholar] [CrossRef]
- Lee, S.C.; Liu, W.; Brosnan, C.F.; Dickson, D.W. GM-CSF Promotes Proliferation of Human Fetal and Adult Microglia in Primary Cultures. Glia 1994, 12, 309–318. [Google Scholar] [CrossRef]
- Liva, S.M.; de Vellis, J. IL-5 Induces Proliferation and Activation of Microglia via an Unknown Receptor. Neurochem. Res. 2001, 26, 629–637. [Google Scholar] [CrossRef]
- Akimoto, N.; Ifuku, M.; Mori, Y.; Noda, M. Effects of Chemokine (C–C Motif) Ligand 1 on Microglial Function. Biochem. Biophys. Res. Commun. 2013, 436, 455–461. [Google Scholar] [CrossRef]
- Zhang, L.; Ma, P.; Guan, Q.; Meng, L.; Su, L.; Wang, L.; Yuan, B. Effect of Chemokine CC Ligand 2 (CCL2) on A-synuclein-induced Microglia Proliferation and Neuronal Apoptosis. Mol. Med. Rep. 2018, 18, 4213–4218. [Google Scholar] [CrossRef]
- Zhang, J.; Geula, C.; Lu, C.; Koziel, H.; Hatcher, L.M.; Roisen, F.J. Neurotrophins Regulate Proliferation and Survival of Two Microglial Cell Lines in Vitro. Exp. Neurol. 2003, 183, 469–481. [Google Scholar] [CrossRef]
- Terashima, T.; Nakae, Y.; Katagi, M.; Okano, J.; Suzuki, Y.; Kojima, H. Stem Cell Factor Induces Polarization of Microglia to the Neuroprotective Phenotype in Vitro. Heliyon 2018, 4, e00837. [Google Scholar] [CrossRef] [PubMed]
- Dermitzakis, I.; Manthou, M.E.; Meditskou, S.; Miliaras, D.; Kesidou, E.; Boziki, M.; Petratos, S.; Grigoriadis, N.; Theotokis, P. Developmental Cues and Molecular Drivers in Myelinogenesis: Revisiting Early Life to Re-Evaluate the Integrity of CNS Myelin. Curr. Issues Mol. Biol. 2022, 44, 3208–3237. [Google Scholar] [CrossRef] [PubMed]
- Lituma, P.J.; Woo, E.; O’Hara, B.F.; Castillo, P.E.; Sibinga, N.E.S.; Nandi, S. Altered Synaptic Connectivity and Brain Function in Mice Lacking Microglial Adapter Protein Iba1. Proc. Natl. Acad. Sci. USA 2021, 118, e2115539118. [Google Scholar] [CrossRef] [PubMed]
- Maksoud, M.J.E.; Tellios, V.; Lu, W.-Y. Nitric Oxide Attenuates Microglia Proliferation by Sequentially Facilitating Calcium Influx through TRPV2 Channels, Activating NFATC2, and Increasing P21 Transcription. Cell Cycle 2021, 20, 417–433. [Google Scholar] [CrossRef] [PubMed]
- The Human Protein Atlas. Available online: https://www.proteinatlas.org/ (accessed on 15 January 2023).
- Home-Gene-NCBI. Available online: https://www.ncbi.nlm.nih.gov/gene/ (accessed on 15 January 2023).
- Ashwell, K. The Distribution of Microglia and Cell Death in the Fetal Rat Forebrain. Brain Res. Dev. Brain Res. 1991, 58, 1–12. [Google Scholar] [CrossRef]
- Sorokin, S.P.; Hoyt, R.F.; Blunt, D.G.; McNelly, N.A. Macrophage Development: II. Early Ontogeny of Macrophage Populations in Brain, Liver, and Lungs of Rat Embryos as Revealed by a Lectin Marker. Anat. Rec. 1992, 232, 527–550. [Google Scholar] [CrossRef]
- Wang, C.C.; Wu, C.H.; Shieh, J.Y.; Wen, C.Y.; Ling, E.A. Immunohistochemical Study of Amoeboid Microglial Cells in Fetal Rat Brain. J. Anat. 1996, 189, 567–574. [Google Scholar]
- Swinnen, N.; Smolders, S.; Avila, A.; Notelaers, K.; Paesen, R.; Ameloot, M.; Brône, B.; Legendre, P.; Rigo, J.-M. Complex Invasion Pattern of the Cerebral Cortex Bymicroglial Cells during Development of the Mouse Embryo. Glia 2013, 61, 150–163. [Google Scholar] [CrossRef]
- Stremmel, C.; Schuchert, R.; Wagner, F.; Thaler, R.; Weinberger, T.; Pick, R.; Mass, E.; Ishikawa-Ankerhold, H.C.; Margraf, A.; Hutter, S.; et al. Yolk Sac Macrophage Progenitors Traffic to the Embryo during Defined Stages of Development. Nat. Commun. 2018, 9, 75. [Google Scholar] [CrossRef]
- Kaur, C.; Rathnasamy, G.; Ling, E.-A. Biology of Microglia in the Developing Brain. J. Neuropathol. Exp. Neurol. 2017, 76, 736–753. [Google Scholar] [CrossRef]
- Ling, E.A.; Wong, W.C. The Origin and Nature of Ramified and Amoeboid Microglia: A Historical Review and Current Concepts. Glia 1993, 7, 9–18. [Google Scholar] [CrossRef]
- Esiri, M.M.; al Izzi, M.S.; Reading, M.C. Macrophages, Microglial Cells, and HLA-DR Antigens in Fetal and Infant Brain. J. Clin. Pathol. 1991, 44, 102–106. [Google Scholar] [CrossRef]
- Rezaie, P.; Male, D. Colonisation of the Developing Human Brain and Spinal Cord by Microglia: A Review. Microsc. Res. Tech. 1999, 45, 359–382. [Google Scholar] [CrossRef]
- Rezaie, P.; Dean, A.; Male, D.; Ulfig, N. Microglia in the Cerebral Wall of the Human Telencephalon at Second Trimester. Cereb. Cortex 2005, 15, 938–949. [Google Scholar] [CrossRef]
- Monier, A.; Evrard, P.; Gressens, P.; Verney, C. Distribution and Differentiation of Microglia in the Human Encephalon during the First Two Trimesters of Gestation. J. Comp. Neurol. 2006, 499, 565–582. [Google Scholar] [CrossRef]
- Monier, A.; Adle-Biassette, H.; Delezoide, A.-L.; Evrard, P.; Gressens, P.; Verney, C. Entry and Distribution of Microglial Cells in Human Embryonic and Fetal Cerebral Cortex. J. Neuropathol. Exp. Neurol. 2007, 66, 372–382. [Google Scholar] [CrossRef]
- Verney, C.; Monier, A.; Fallet-Bianco, C.; Gressens, P. Early Microglial Colonization of the Human Forebrain and Possible Involvement in Periventricular White-Matter Injury of Preterm Infants. J. Anat. 2010, 217, 436–448. [Google Scholar] [CrossRef]
- Cuadros, M.A.; Martin, C.; Coltey, P.; Almendros, A.; Navascués, J. First Appearance, Distribution, and Origin of Macrophages in the Early Development of the Avian Central Nervous System. J. Comp. Neurol. 1993, 330, 113–129. [Google Scholar] [CrossRef]
- Cuadros, M.A.; Rodríguez-Ruiz, J.; Calvente, R.; Almendros, A.; Marín-Teva, J.L.; Navascués, J. Microglia Development in the Quail Cerebellum. J. Comp. Neurol. 1997, 389, 390–401. [Google Scholar] [CrossRef]
- Krall, W.J.; Challita, P.M.; Perlmutter, L.S.; Skelton, D.C.; Kohn, D.B. Cells Expressing Human Glucocerebrosidase from a Retroviral Vector Repopulate Macrophages and Central Nervous System Microglia after Murine Bone Marrow Transplantation. Blood 1994, 83, 2737–2748. [Google Scholar] [CrossRef]
- Kennedy, D.W.; Abkowitz, J.L. Kinetics of Central Nervous System Microglial and Macrophage Engraftment: Analysis Using a Transgenic Bone Marrow Transplantation Model. Blood 1997, 90, 986–993. [Google Scholar] [CrossRef] [PubMed]
- Eglitis, M.A.; Mezey, É. Hematopoietic Cells Differentiate into Both Microglia and Macroglia in the Brains of Adult Mice. Proc. Natl. Acad. Sci. USA 1997, 94, 4080–4085. [Google Scholar] [CrossRef] [PubMed]
- Priller, J.; Flügel, A.; Wehner, T.; Boentert, M.; Haas, C.A.; Prinz, M.; Fernández-Klett, F.; Prass, K.; Bechmann, I.; de Boer, B.A.; et al. Targeting Gene-Modified Hematopoietic Cells to the Central Nervous System: Use of Green Fluorescent Protein Uncovers Microglial Engraftment. Nat. Med. 2001, 7, 1356–1361. [Google Scholar] [CrossRef] [PubMed]
- Simard, A.R.; Rivest, S. Bone Marrow Stem Cells Have the Ability to Populate the Entire Central Nervous System into Fully Differentiated Parenchymal Microglia. FASEB J. 2004, 18, 998–1000. [Google Scholar] [CrossRef]
- Asheuer, M.; Pflumio, F.; Benhamida, S.; Dubart-Kupperschmitt, A.; Fouquet, F.; Imai, Y.; Aubourg, P.; Cartier, N. Human CD34+ Cells Differentiate into Microglia and Express Recombinant Therapeutic Protein. Proc. Natl. Acad. Sci. USA 2004, 101, 3557–3562. [Google Scholar] [CrossRef]
- Beck, H.; Voswinckel, R.; Wagner, S.; Ziegelhoeffer, T.; Heil, M.; Helisch, A.; Schaper, W.; Acker, T.; Hatzopoulos, A.K.; Plate, K.H. Participation of Bone Marrow-Derived Cells in Long-Term Repair Processes after Experimental Stroke. J. Cereb. Blood Flow Metab. 2003, 23, 709–717. [Google Scholar] [CrossRef]
- Hess, D.C.; Abe, T.; Hill, W.D.; Studdard, A.M.; Carothers, J.; Masuya, M.; Fleming, P.A.; Drake, C.J.; Ogawa, M. Hematopoietic Origin of Microglial and Perivascular Cells in Brain. Exp. Neurol. 2004, 186, 134–144. [Google Scholar] [CrossRef]
- Djukic, M.; Mildner, A.; Schmidt, H.; Czesnik, D.; Brück, W.; Priller, J.; Nau, R.; Prinz, M. Circulating Monocytes Engraft in the Brain, Differentiate into Microglia and Contribute to the Pathology Following Meningitis in Mice. Brain J. Neurol. 2006, 129, 2394–2403. [Google Scholar] [CrossRef]
- Bechmann, I.; Goldmann, J.; Kovac, A.D.; Kwidzinski, E.; Simbürger, E.; Naftolin, F.; Dirnagl, U.; Nitsch, R.; Priller, J. Circulating Monocytic Cells Infiltrate Layers of Anterograde Axonal Degeneration Where They Transform into Microglia. FASEB J. 2005, 19, 647–649. [Google Scholar] [CrossRef]
- Rodriguez, M.; Alvarez-Erviti, L.; Blesa, F.J.; Rodríguez-Oroz, M.C.; Arina, A.; Melero, I.; Ramos, L.I.; Obeso, J.A. Bone-Marrow-Derived Cell Differentiation into Microglia: A Study in a Progressive Mouse Model of Parkinson’s Disease. Neurobiol. Dis. 2007, 28, 316–325. [Google Scholar] [CrossRef]
- Malm, T.M.; Koistinaho, M.; Pärepalo, M.; Vatanen, T.; Ooka, A.; Karlsson, S.; Koistinaho, J. Bone-Marrow-Derived Cells Contribute to the Recruitment of Microglial Cells in Response to Beta-Amyloid Deposition in APP/PS1 Double Transgenic Alzheimer Mice. Neurobiol. Dis. 2005, 18, 134–142. [Google Scholar] [CrossRef]
- Simard, A.R.; Soulet, D.; Gowing, G.; Julien, J.-P.; Rivest, S. Bone Marrow-Derived Microglia Play a Critical Role in Restricting Senile Plaque Formation in Alzheimer’s Disease. Neuron 2006, 49, 489–502. [Google Scholar] [CrossRef]
- Ye, S.; Theotokis, P.; Lee, J.Y.; Kim, M.J.; Nheu, D.; Ellen, O.; Bedford, T.; Ramanujam, P.; Wright, D.K.; McDonald, S.J.; et al. NgR-Fc Delivered by Hematopoietic Cells Enhances Neurorepair in a Multiple Sclerosis Model. Brain Commun. 2023, in press. [Google Scholar]
- Flügel, A.; Bradl, M.; Kreutzberg, G.W.; Graeber, M.B. Transformation of Donor-Derived Bone Marrow Precursors into Host Microglia during Autoimmune CNS Inflammation and during the Retrograde Response to Axotomy. J. Neurosci. Res. 2001, 66, 74–82. [Google Scholar] [CrossRef]
- Priller, J.; Prinz, M.; Heikenwalder, M.; Zeller, N.; Schwarz, P.; Heppner, F.L.; Aguzzi, A. Early and Rapid Engraftment of Bone Marrow-Derived Microglia in Scrapie. J. Neurosci. 2006, 26, 11753–11762. [Google Scholar] [CrossRef]
- Vallières, L.; Sawchenko, P.E. Bone Marrow-Derived Cells That Populate the Adult Mouse Brain Preserve Their Hematopoietic Identity. J. Neurosci. 2003, 23, 5197–5207. [Google Scholar] [CrossRef]
- Massengale, M.; Wagers, A.J.; Vogel, H.; Weissman, I.L. Hematopoietic Cells Maintain Hematopoietic Fates upon Entering the Brain. J. Exp. Med. 2005, 201, 1579–1589. [Google Scholar] [CrossRef]
- Zhang, J.; Shi, X.Q.; Echeverry, S.; Mogil, J.S.; De Koninck, Y.; Rivest, S. Expression of CCR2 in Both Resident and Bone Marrow-Derived Microglia Plays a Critical Role in Neuropathic Pain. J. Neurosci. 2007, 27, 12396–12406. [Google Scholar] [CrossRef]
- Marchi, S.; Guilbaud, E.; Tait, S.W.G.; Yamazaki, T.; Galluzzi, L. Mitochondrial Control of Inflammation. Nat. Rev. Immunol. 2023, 23, 159–173. [Google Scholar] [CrossRef]
- Joshi, A.U.; Minhas, P.S.; Liddelow, S.A.; Haileselassie, B.; Andreasson, K.I.; Dorn, G.W.; Mochly-Rosen, D. Fragmented Mitochondria Released from Microglia Trigger A1 Astrocytic Response and Propagate Inflammatory Neurodegeneration. Nat. Neurosci. 2019, 22, 1635–1648. [Google Scholar] [CrossRef]
- Leong, S.K.; Ling, E.A. Amoeboid and Ramified Microglia: Their Interrelationship and Response to Brain Injury. Glia 1992, 6, 39–47. [Google Scholar] [CrossRef] [PubMed]
- Lassmann, H.; Schmied, M.; Vass, K.; Hickey, W.F. Bone Marrow Derived Elements and Resident Microglia in Brain Inflammation. Glia 1993, 7, 19–24. [Google Scholar] [CrossRef] [PubMed]
- Dalmau, I.; Vela, J.M.; González, B.; Finsen, B.; Castellano, B. Dynamics of Microglia in the Developing Rat Brain. J. Comp. Neurol. 2003, 458, 144–157. [Google Scholar] [CrossRef]
- Lawson, L.J.; Perry, V.H.; Gordon, S. Turnover of Resident Microglia in the Normal Adult Mouse Brain. Neuroscience 1992, 48, 405–415. [Google Scholar] [CrossRef] [PubMed]
- Solomon, J.N.; Lewis, C.-A.B.; Ajami, B.; Corbel, S.Y.; Rossi, F.M.V.; Krieger, C. Origin and Distribution of Bone Marrow-Derived Cells in the Central Nervous System in a Mouse Model of Amyotrophic Lateral Sclerosis. Glia 2006, 53, 744–753. [Google Scholar] [CrossRef]
- Ajami, B.; Bennett, J.L.; Krieger, C.; Tetzlaff, W.; Rossi, F.M.V. Local Self-Renewal Can Sustain CNS Microglia Maintenance and Function throughout Adult Life. Nat. Neurosci. 2007, 10, 1538–1543. [Google Scholar] [CrossRef]
- Mildner, A.; Schmidt, H.; Nitsche, M.; Merkler, D.; Hanisch, U.-K.; Mack, M.; Heikenwalder, M.; Brück, W.; Priller, J.; Prinz, M. Microglia in the Adult Brain Arise from Ly-6ChiCCR2+ Monocytes Only under Defined Host Conditions. Nat. Neurosci. 2007, 10, 1544–1553. [Google Scholar] [CrossRef]
- Capotondo, A.; Milazzo, R.; Politi, L.S.; Quattrini, A.; Palini, A.; Plati, T.; Merella, S.; Nonis, A.; di Serio, C.; Montini, E.; et al. Brain Conditioning Is Instrumental for Successful Microglia Reconstitution Following Hematopoietic Stem Cell Transplantation. Proc. Natl. Acad. Sci. USA 2012, 109, 15018–15023. [Google Scholar] [CrossRef]
- Li, T.; Pang, S.; Yu, Y.; Wu, X.; Guo, J.; Zhang, S. Proliferation of Parenchymal Microglia Is the Main Source of Microgliosis after Ischaemic Stroke. Brain J. Neurol. 2013, 136, 3578–3588. [Google Scholar] [CrossRef]
- Bruttger, J.; Karram, K.; Wörtge, S.; Regen, T.; Marini, F.; Hoppmann, N.; Klein, M.; Blank, T.; Yona, S.; Wolf, Y.; et al. Genetic Cell Ablation Reveals Clusters of Local Self-Renewing Microglia in the Mammalian Central Nervous System. Immunity 2015, 43, 92–106. [Google Scholar] [CrossRef]
- Xu, J.; Zhu, L.; He, S.; Wu, Y.; Jin, W.; Yu, T.; Qu, J.Y.; Wen, Z. Temporal-Spatial Resolution Fate Mapping Reveals Distinct Origins for Embryonic and Adult Microglia in Zebrafish. Dev. Cell 2015, 34, 632–641. [Google Scholar] [CrossRef]
- Tay, T.L.; Mai, D.; Dautzenberg, J.; Fernández-Klett, F.; Lin, G.; Sagar, N.; Datta, M.; Drougard, A.; Stempfl, T.; Ardura-Fabregat, A.; et al. A New Fate Mapping System Reveals Context-Dependent Random or Clonal Expansion of Microglia. Nat. Neurosci. 2017, 20, 793–803. [Google Scholar] [CrossRef]
- Cronk, J.C.; Filiano, A.J.; Louveau, A.; Marin, I.; Marsh, R.; Ji, E.; Goldman, D.H.; Smirnov, I.; Geraci, N.; Acton, S.; et al. Peripherally Derived Macrophages Can Engraft the Brain Independent of Irradiation and Maintain an Identity Distinct from Microglia. J. Exp. Med. 2018, 215, 1627–1647. [Google Scholar] [CrossRef]
- Lund, H.; Pieber, M.; Parsa, R.; Han, J.; Grommisch, D.; Ewing, E.; Kular, L.; Needhamsen, M.; Espinosa, A.; Nilsson, E.; et al. Competitive Repopulation of an Empty Microglial Niche Yields Functionally Distinct Subsets of Microglia-like Cells. Nat. Commun. 2018, 9, 4845. [Google Scholar] [CrossRef]
- Shemer, A.; Grozovski, J.; Tay, T.L.; Tao, J.; Volaski, A.; Süß, P.; Ardura-Fabregat, A.; Gross-Vered, M.; Kim, J.-S.; David, E.; et al. Engrafted Parenchymal Brain Macrophages Differ from Microglia in Transcriptome, Chromatin Landscape and Response to Challenge. Nat. Commun. 2018, 9, 5206. [Google Scholar] [CrossRef]
- Huang, Y.; Xu, Z.; Xiong, S.; Sun, F.; Qin, G.; Hu, G.; Wang, J.; Zhao, L.; Liang, Y.-X.; Wu, T.; et al. Repopulated Microglia Are Solely Derived from the Proliferation of Residual Microglia after Acute Depletion. Nat. Neurosci. 2018, 21, 530–540. [Google Scholar] [CrossRef]
- Elmore, M.R.P.; Najafi, A.R.; Koike, M.A.; Dagher, N.N.; Spangenberg, E.E.; Rice, R.A.; Kitazawa, M.; Matusow, B.; Nguyen, H.; West, B.L.; et al. Colony-Stimulating Factor 1 Receptor Signaling Is Necessary for Microglia Viability, Unmasking a Microglia Progenitor Cell in the Adult Brain. Neuron 2014, 82, 380–397. [Google Scholar] [CrossRef]
- Zhan, L.; Krabbe, G.; Du, F.; Jones, I.; Reichert, M.C.; Telpoukhovskaia, M.; Kodama, L.; Wang, C.; Cho, S.-H.; Sayed, F.; et al. Proximal Recolonization by Self-Renewing Microglia Re-Establishes Microglial Homeostasis in the Adult Mouse Brain. PLoS Biol. 2019, 17, e3000134. [Google Scholar] [CrossRef]
- Wu, X.; Saito, T.; Saido, T.C.; Barron, A.M.; Ruedl, C. Microglia and CD206+ Border-Associated Mouse Macrophages Maintain Their Embryonic Origin during Alzheimer’s Disease. eLife 2021, 10, e71879. [Google Scholar] [CrossRef]
- Luo, C.; Jian, C.; Liao, Y.; Huang, Q.; Wu, Y.; Liu, X.; Zou, D.; Wu, Y. The Role of Microglia in Multiple Sclerosis. Neuropsychiatr. Dis. Treat. 2017, 13, 1661–1667. [Google Scholar] [CrossRef]
- Schetters, S.T.T.; Gomez-Nicola, D.; Garcia-Vallejo, J.J.; Van Kooyk, Y. Neuroinflammation: Microglia and T Cells Get Ready to Tango. Front. Immunol. 2018, 8, 1905. [Google Scholar] [CrossRef] [PubMed]
- Goddery, E.N.; Fain, C.E.; Lipovsky, C.G.; Ayasoufi, K.; Yokanovich, L.T.; Malo, C.S.; Khadka, R.H.; Tritz, Z.P.; Jin, F.; Hansen, M.J.; et al. Microglia and Perivascular Macrophages Act as Antigen Presenting Cells to Promote CD8 T Cell Infiltration of the Brain. Front. Immunol. 2021, 12, 726421. [Google Scholar] [CrossRef] [PubMed]
- O’Loughlin, E.; Madore, C.; Lassmann, H.; Butovsky, O. Microglial Phenotypes and Functions in Multiple Sclerosis. Cold Spring Harb. Perspect. Med. 2018, 8, a028993. [Google Scholar] [CrossRef] [PubMed]
- Nheu, D.; Ellen, O.; Ye, S.; Ozturk, E.; Pagnin, M.; Kertadjaja, S.; Theotokis, P.; Grigoriadis, N.; McLean, C.; Petratos, S. Modulation of the Microglial Nogo-A/NgR Signaling Pathway as a Therapeutic Target for Multiple Sclerosis. Cells 2022, 11, 3768. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).